Abstract
Substrate-dependent cardiac differentiation of induced pluripotent stem cells (iPSCs) has been studied on various extracellular matrix (ECM)-derived substrates, such as collagen type I (Col-I). However, ECM-derived substrates have multiple cell-adhesive amino acid sequences and stimulate various signaling pathways in cells, making it difficult to clarify the mechanism of substrate-dependent stem cell differentiation. A substrate presenting one of these sequences is a powerful tool for elucidating the mechanism. We designed elastin-like proteins (ELPs) composed of repetitive VPGIG sequences with or without the RGD cell adhesion motif (ELP-RGD/ELP-Ctrl) and used a chemical crosslinker to generate hydrogels. By adjusting the ELP and crosslinker concentrations, we obtained ELP-Ctrl and ELP-RGD hydrogels with a Young’s modulus of 0.3 kPa. The ELP-Ctrl and ELP-RGD gels were used as a substrate for the cardiac differentiation of cultured murine iPSCs. Cells on the ELP-RGD gel showed four times higher gene expression of the contractile protein troponin T type 2 than those on a Col-I gel, which is an effective substrate for iPSC cardiac differentiation. The ELP-RGD gel might stimulate integrin-derived signaling pathways in the cells to promote cardiac differentiation. This study showed the potential of ELP hydrogels for studying substrate-dependent iPSC cardiac differentiation by enabling the control of cell-adhesive sequence presentation.
Similar content being viewed by others
Introduction
A large number of cardiomyocytes are required for cell-based myocardial regeneration therapies, screening drugs for cardiac disorders, and the clarification of the pathogenic mechanisms of heart diseases. Among various types of stem cells [1,2,3], induced pluripotent stem cells (iPSCs) have received the greatest focus as a cell source for cardiomyocytes [4,5,6]. Soluble factors in cell culture media, such as 5-azacytidine and trichostatin A (TSA), have been used to induce the cardiac differentiation of iPSCs [7,8,9], and the underlying mechanisms have been clarified. For instance, the recently discovered small-molecule KY02111 stimulates the cardiac differentiation of iPSCs by inhibiting Wnt signaling [9].
In addition to soluble factors, the cellular microenvironment or niche has been recognized as an essential factor for the cardiac differentiation of stem cells [5, 6, 10,11,12]. Burridge et al. [10] showed that iPSCs seeded onto laminin 511- or 521-coated substrates adhered, grew, and differentiated into cardiomyocytes well. Jung et al. [11] found an optimal formulation of three extracellular matrix (ECM) proteins (61% collagen type I (Col-I), 24% laminin 111, and 15% fibronectin) for the cardiac differentiation of iPSCs by culturing them in polyethylene glycol-based hydrogels containing ECM components. Macri-Pellizzeri et al. [12] cultured iPS-derived embryoid bodies on Col-I- or fibronectin-coated polyacrylamide gels with various elasticities and reported that cells grown on soft gels (0.6 kPa) coated with fibronectin effectively expressed cardiac marker genes. We previously induced the cardiac differentiation of iPSCs cultured on polyacrylamide gels coated with Col-I, gelatin, or fibronectin and showed that cardiac marker gene expression in iPSCs on Col-I-coated gels was enhanced when the gel elasticity was 9 kPa [6]. Although the results of these previous studies suggest the importance of the niche, particularly the cell culture substrate, for directing the fate of iPSCs, the mechanisms underlying substrate-dependent cardiac differentiation of iPSCs have not been fully clarified.
Since cells adhere to substrates via transmembrane molecules such as integrins, researchers suspect the involvement of integrin-related signaling pathways in the cardiac differentiation of iPSCs [6, 10, 11]. However, ECM-derived substrates contain multiple cell-adhesive amino acid sequences and various integrin-related signaling pathways can be stimulated when iPSCs are cultured on these substrates. For instance, stem cells bind to Col-I via the integrins α1β1, α2β1, α3β1, and/or αVβ3 [13]. In addition, the purity of ECM-derived substrates can vary among lots [14], and contaminants may have biological functions. These facts make it difficult to identify the key signaling pathway for substrate-dependent cardiac differentiation of iPSCs.
Substrates presenting a single-cell adhesion motif are preferred to simplify the investigation of the substrate-dependent cardiac differentiation of iPSCs. Elastin-like proteins (ELPs) are artificial polypeptides that contain tropoelastin-derived repeated amino acid sequences of VPGXG (where X is any amino acid except proline). Transgenic bacterial expression systems enable the modulation of the ELP amino acid sequence to contain a single-cell adhesion sequence at the gene level and the production of homologous polymers at >100 mg L–1 in shake flask culture [15, 16]. By using chemical crosslinkers, ELPs are transformed into hydrogels, and their elasticity, which is one of the important factors determining stem cell differentiation [17], can be tuned [18,19,20]. Thus ELP hydrogels with or without a single-cell adhesion sequence and of uniform elasticity can be produced, facilitating the identification of a key signaling pathway induced by the particular cell-adhesive sequence. In addition, unlike the synthetic polyacrylamide hydrogels that are widely used as stem cell culture niche [17], ELP-based matrices can potentially be used as scaffolds in vivo. However, to date, no study has reported the cardiac differentiation of iPSCs on/in ELP substrates.
The goal of this study was to establish an ELP hydrogel-based culture system for the cardiac differentiation of iPSCs using a tissue culture polystyrene (TCPS) plate and Col-I gel as a conventional substrate and a cardiac differentiation-stimulating substrate [6], respectively. Since Col-I contains the cell adhesion motif RGD in its primary structure [21], we produced ELPs with or without RGD (ELP-RGD or ELP-Ctrl, respectively) by using a transgene expression system. We used a crosslinker, tetrakis(hydroxymethyl)phosphonium chloride (THPC) [20], to produce soft gels with an elasticity of 0.3 kPa, which mimics the elasticity of embryonic cardiac tissue and reportedly directs the cardiac commitment of pluripotent P19 cells [22]. Murine iPSCs were cultured on the ELP hydrogels in the presence of calcium-rich medium and TSA to induce cardiac differentiation [8]. We quantified the gene expression of the cardiac transcription factor, GATA-binding protein 4 (GATA4) as well as that of contractile proteins alpha-myosin heavy chain (α-MHC) and troponin T type 2 (TnT2) as we aimed to obtain self-beating cardiomyocytes.
Materials and methods
Vector construction
Previously reported plasmids [23] containing DNA fragments coding ELP-Ctrl and ELP-RGD were digested with BamHI and HindIII (New England Biolabs, MA) and the fragments were inserted into the BamHI–HindIII site of the pET30a vector (Novagen, Germany). These expression plasmids were designated pET[ELP-Ctrl] and pET[ELP-RGD], respectively. Prior to expression, the constructs were verified by restriction digest and DNA sequencing. The full amino acid sequences of the expressed proteins are shown in Table 1.
Recombinant ELP production and purification
Escherichia coli strain BL21(DE3) competent cells (Novagen) transformed with pET[ELP-Ctrl] or pET[ELP-RGD] were plated onto Luria-Bertani agar containing 35 µg mL−1 kanamycin. After an overnight incubation at 37 °C, recombinant colonies were transferred to Overnight Express Instant TB medium (Novagen), supplemented with 10 vol% glycerol and 35 µg mL−1 kanamycin in shake flasks and incubated at 180 rpm for 24 h at 30 °C. Cells were pelleted by centrifugation at 1200 × g for 5 min at 4 °C and stored at –80 °C until use.
ELP-Ctrl and ELP-RGD were purified as described previously [24]. Briefly, cells were resuspended and lysed in a lysis buffer containing 8 M urea, and impurities were removed by centrifugation and filtration. The ELPs, tagged with polyhistidine (His) at the amino terminal, were purified with a Ni-chelate column (HisTrap FF crude; GE Healthcare, UK); the proteins trapped on the column were washed with a solution containing 5 mM imidazole and then eluted with a solution containing 250 mM imidazole. The protein composition of the purified solutions was analyzed by sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE) on Mini-PROTEAN TGX gels (Any kD; Bio-Rad Laboratories, CA) under reducing conditions. The proteins separated on a gel were fixed and visualized with EzStain AQua solution (Atto, Japan), which contains Coomassie Brilliant Blue. The proteins on another gel were fixed and stained with a His-tag-specific dye (InVision His-Tag In-Gel Stain Kit; Thermo Fisher Scientific, MA). Images were captured with a ChemiDoc XRS gel imager (Bio-Rad Laboratories). The intensity of peptide bands was measured using ImageJ (http://imagej.nih.gov.ij/) to determine the purity of the ELPs. Eluted fractions containing ELP-Ctrl or ELP-RGD were concentrated by ultrafiltration (nominal molecular weight limit, 10 × 103) and dialyzed against deionized water (molecular weight cutoff, 10 × 103) for 1 day, changing the water every 4–6 h. Then the ELP aqueous solutions were freeze-dried, and the weight of ELP was measured to calculate its yield.
Rheological characterization of hydrogels
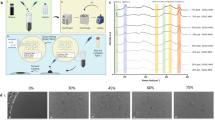
The freeze-dried ELPs were suspended in phosphate-buffered saline (PBS, pH 7.4; Gibco, MA) on ice, and a THPC aqueous solution (Sigma-Aldrich, MO) was added to the suspension, followed by incubation at 37 °C. Within 1 h, opaque ELP hydrogels formed. THPC reacts with primary and secondary amines [20], resulting in crosslinks among ELPs (Fig. 1). To evaluate the elasticity and strain sweep of the ELP hydrogels, their rheological behavior was recorded on a rheometer (MCR301; Anton Paar, Austria), equipped with 8-mm-diameter parallel plates with a 0.5-mm measuring gap distance and a temperature controller. The ELP suspension incubated on ice was transferred onto the rheometer plate and cooled at 4 °C, and the THPC solution was added immediately (final concentrations: 25 mg mL–1 ELP-Ctrl and 68 µg mL–1 THCP; and 50 mg mL–1 ELP-RGD and 260 µg mL–1 THPC). Then the temperature of the plate was raised to 37 °C. The measurement site was chambered with a wet spongy ring surrounding the sample to inhibit the evaporation of water. After the incubation at 37 °C for 1 h, dynamic oscillatory strain sweeps were obtained (strain, 0.1–100%; frequency, 1 rad s–1) at 37 °C. As the shear storage modulus (G’) of the ELP hydrogels was constant at 0.1%–1% strain, the value at 1% strain was determined as G’ of the sample, and its Young’s modulus (E) was calculated using the following equation:
where μ is the Poisson ratio of the gels, which was assumed to be 0.5. Therefore, E was calculated as a function of G’ as follows [22, 25]:
A Col-I gel produced from a porcine Col-I solution (1.65 mg mL–1, CellmatrixTM type I-P; Nitta Gelatin, Japan) in accordance with the manufacturer’s protocol, and its elasticity and strain sweep were evaluated. For each gel type, 3 samples were tested (n = 3).
Determination of the water content of the hydrogels
The water content of the ELP-Ctrl, ELP-RGD, and Col-I hydrogels was determined by weighing wet and dry hydrogels as described previously [24]. Briefly, after gelation at 37 °C for 1 h, the hydrogels were submerged in ultrapure water at room temperature for 3 h, and excess water was removed. The weight of wet hydrogels was measured, and then the hydrogels were lyophilized. The weight of the lyophilized hydrogels was measured, and the water content of the gels was calculated.
Scanning electron microscopy of hydrogels
The ELP-Ctrl, ELP-RGD, and Col-I hydrogels were immersed in ultrapure water, frozen at –80 °C, and freeze-dried. Samples were coated with gold by vacuum vapor deposition and imaged using a scanning electron microscope (JCM-5700; JEOL, Japan).
Cardiac differentiation culture of murine iPSCs
The freeze-dried ELPs were suspended in PBS on ice and sterilized by exposure to ultraviolet (UV) light for 30 min. ELP molecular weight (MW) and gel elasticity were unaffected by this UV light exposure (Supplementary Data Fig. S1). The THPC solution, sterilized by filtering through a polyethersulfone membrane with 0.22-µm pores (Merck Millipore, MA), was added to a 48-well TCPS plate (IWAKI, Japan). Then the ELP solution was poured into the TCPS plate on ice (total volume 100 µL well–1; final concentrations: 25 mg mL–1 ELP-Ctrl and 68 µg mL–1 THCP; and 50 mg mL–1 ELP-RGD and 260 µg mL–1 THPC). After incubation for 1 h under culture conditions (37 °C in a humidified atmosphere of 95% air and 5% CO2) for gelation, the ELP hydrogels were washed twice with PBS and twice with high-calcium Minimum Essential Medium (MEM) alpha (Gibco) supplemented with 10 vol% fetal bovine serum (FBS; Equitech-Bio, TX), 0.1 mM 2-mercaptoethanol, and 0.5 vol% antibiotic mixture (final concentrations, 50 unit mL–1 penicillin; and 50 µg mL–1 streptomycin; Gibco) (CD medium). Col-I gel was also prepared.
After expansion in the maintenance culture with Dulbecco’s modified Eagle’s medium (Gibco) containing 15 vol% FBS, 1 vol% MEM nonessential amino acids (Gibco), 0.1 mM 2-mercaptoethanol, 103 unit mL–1 leukemia inhibitory factor (ESG1106; Merck Millipore), and 0.5 vol% antibiotic mixture (M medium) as described previously [6, 26], iPSCs (iPS-MEF-Ng-175B-5; passage number, 15–17; Riken, Japan), as well as mitomycin C-treated SNL feeder cells (SNL 76/7; passage number, 15; European Collection of Authenticated Cell Cultures, UK), were removed from the dish by mixing with 0.25% trypsin-EDTA (Gibco) and were pelleted by centrifugation at 200 × g for 5 min at 4 °C. The cells were resuspended in M medium, seeded onto the gelatin-coated dish, and incubated for 1 h under the culture condition. This allowed the feeder cells to attach to the dish, and the supernatant containing the iPSCs was collected. The iPSCs were pelleted, resuspended in CD medium, and seeded onto the hydrogels and a 6-well TCPS plate (IWAKI) at 3.2 × 103 cells cm–1. The cells were cultured in CD medium under the culture condition for 15 days. The medium was changed on days 3 and 5. On day 7, the medium was replaced with CD medium containing 10 ng mL–1 TSA (Wako Pure Chemical, Japan) to induce cardiac differentiation [8]. After incubation for 24 h, the medium with TSA was replaced with CD medium; thereafter, the medium was changed daily. iPSCs were cultured on TCPS with M medium as a negative control for cardiac differentiation.
Real-time polymerase chain reaction (qPCR) analysis
On day 11, the iPSCs cultured on the hydrogels and TCPS were washed with PBS and lysed with TRIzol® reagent (Life Technologies, CA). Total RNA was purified using the High Pure RNA Tissue kit (Roche Diagnostics, Germany). cDNA was synthesized by reverse transcription using the High-Capacity cDNA Reverse Transcription Kit (Life Technologies). Using the same procedures, total RNA was extracted from the heart of a C57BL/6N mouse (6-week-old; Japan SLC, Japan), and cDNA was synthesized. A qPCR was carried out using the TaqMan® Gene Expression Master Mix (Applied Biosystems, CA), and probe-based PCR and real-time fluorescence detection were conducted using the StepOneTM Real-Time PCR system (Applied Biosystems). Primers and probes (TaqMan® Gene Expression assays; Applied Biosystems) were designed for the genes coding glyceraldehyde-3-phosphate dehydrogenase (GAPDH) (assay ID, Mm99999915_g1), GATA4 (assay ID, Mm00484689_m1), α-MHC (assay ID, Mm00440359_m1), TnT2 (assay ID, Mm01290256_m1), and (sex determining region Y)-box 2 (Sox2; assay ID, Mm03053810_s1); GAPDH and Sox2 were used as an internal control and a marker for undifferentiated cells, respectively. Gene expression was quantified relative to the level in the heart using the 2−ΔΔCT method [27] as preliminary evaluations had confirmed the validity of this method. Four different samples were used for each time point (n = 4), except for Gata4 and TnT2 levels in the ELP-RGD group (n = 3).
Statistical analysis
A two-sided Student’s t test was used for comparison between two groups. For comparison among three groups, a one-way analysis of variance followed by Tukey or Tukey–Kramer post hoc comparisons was used. A value of p < 0.05 was considered significant.
Ethic statement
All animal experiments were conducted in accordance with the Guidelines for Animal Experiments established by the Ministry of Health, Labour, and Welfare of Japan and by the NCVC Research Institute. The protocol was approved by the Committee on Ethics of Animal Experiments of the NCVC Research Institute (permit numbers 16050 and 17049).
Results and discussion
Production of ELPs
Proteins produced by E. coli transformed with the pET[ELP-Ctrl] or pET[ELP-RGD] vectors were analyzed by SDS-PAGE (Fig. 2). The major protein after purification had a MW of approximately 40 × 103 and 30 × 103 for ELP-Ctrl and ELP-RGD, respectively (Fig. 2a). These values corresponded to the theoretical MWs of ELP-Ctrl (36.1 × 103) and ELP-RGD (30.8 × 103) as shown in Table 1. In addition, the proteins were positive for the recombinant protein-specific His-tag (Fig. 2b), suggesting that the ELPs were successfully synthesized through expression in E. coli cultured in shake flasks. ELP yield was >130 mg L–1 culture medium, and the purity was high (>84%) (Table 1).
SDS-PAGE analysis of proteins extracted from E. coli cells transformed with the pET[ELP-Ctrl] or pET[ELP-RGD] vector. a Crude protein solutions were purified on a Ni-chelate column, and crude, flow-through (FT), wash, and eluted fractions were collected for analysis by SDS-PAGE with Coomassie Brilliant Blue staining. BenchMarkTM Pre-Stained Protein Standard (Invitrogen, CA) was used as a marker. b The eluted fraction was run on the gel and stained with His-tag-specific dye. BenchMarkTM His-tagged Protein Standard (Invitrogen) was used as a marker. Arabic numerals at the side of the photographs represent MW (×103)
Physical properties of ELP hydrogels
Since the elasticity of the substrates affects the cardiac differentiation of iPSCs [6], we optimized the concentrations of the ELPs and THPC (Supplementary Data Tables S1 and S2) to obtain ELP-Ctrl gel, ELP-RGD gel, and Col-I gel with uniform elasticity. At any concentrations of THPC, ELP gels showed a flat G’ at a strain of 0.1–1%; however, at larger strain, the modulus decreased sharply with increasing strain (Fig. 3a), indicating the formation of physical crosslinks. It is well known that polypeptides composed of repeated VPGIG are hydrated and form a random coil-rich structure at low temperature. In contrast, at higher temperature (>30 °C), they undergo a conformational change into a β-spiral-rich structure to aggregate hydrophobically [28]. Therefore, in addition to the chemical crosslinks formed by THPC, physical crosslinks consisting of aggregated poly-VPGIG might be involved in ELP hydrogel formation. The THPC-induced chemical crosslink was necessary for the formation of ELP hydrogels, because without the chemical crosslinker, ELPs in PBS on ice just precipitated when the temperature of the solution increased to room temperature (data not shown).
Physical properties of the hydrogels. a Typical curves of the storage (G’) and loss (G”) moduli of Col-I, ELP-Ctrl, and ELP-RGD hydrogels at 37 °C characterized by oscillatory shear measurements at strains of 0.1–100%. b Young’s modulus of the hydrogels calculated from their storage modulus at 1% strain. Data are shown as the mean ± SEM (n = 3). No significant differences among the hydrogels were detected by one-way ANOVA. c Water content of the hydrogels. Data are shown as the mean ± SEM (n = 3). Asterisks indicate significant differences analyzed by one-way ANOVA followed by Tukey post hoc comparison (**p < 0.01, ***p < 0.001). d Scanning electron micrographs of the hydrogels. Scale bar = 50 µm
ELP-Ctrl and ELP-RGD gels with a Young’s modulus of approximately 0.3 kPa were obtained when their concentration was 25 and 50 mg mL–1, respectively (Fig. 3b). A smaller amount of ELP-Ctrl, which has a larger MW and more primary and secondary amines than ELP-RGD, was required to form a hydrogel with an elasticity of 0.3 kPa. These findings are consistent with a previous study showing that the strengthening of the G’ of chemically crosslinked ELP gels is accompanied by increases in the MW, concentration, and/or number of lysine (primary amine) residues of ELP [29]. Similarly, the Col-I gel also had an elasticity of 0.3 kPa. The Young’s modulus did not significantly differ among the three types of gels (Fig. 3b). Thus the effects of substrate elasticity on the cardiac differentiation of iPSCs cultured on the gels in this study were avoided.
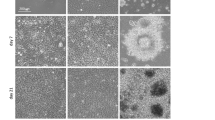
As shown in Fig. 3c, the Col-I, ELP-Ctrl, and ELP-RGD hydrogels had high water content (>95%). While the Col-I gel retained significantly more water than the ELP-Ctrl and ELP-RGD gels, there was no significant difference between the latter. The microstructure of the freeze-dried hydrogels is shown in Fig. 3d. The Col-I gel had a coarse network structure, whereas the ELP-Ctrl and ELP-RGD gels had a dense network structure. This structural difference was likely to be related to the water content of the hydrogels as a coarser network might retain a larger amount of water.
Cardiac gene expression levels in iPSCs cultured on ELP hydrogels
Cardiac differentiation of murine iPSCs cultured on ELP-Ctrl, ELP-RGD, and Col-I gels, as well as on a conventional TCPS substrate with a Young’s modulus of 2 GPa [6], was induced by calcium-rich medium and TSA treatment [8]. The cells were exposed to TSA treatment on day 7 after cell seeding. On day 11 on TCPS, self-beating colonies were observed at 0.5–1 colonies cm–2 (Supplementary Data Movie S1) in addition to a sarcomeric structure (Supplementary Data Fig. S2), indicating that the procedure for cardiac differentiation of iPSCs was successful, which was supported by the qPCR results (Fig. 4a). Compared with iPSCs cultured with CD medium and TSA, cells cultured with M medium showed a significantly lower TnT2 expression level; instead, Sox2 (undifferentiation marker) expression on day 11 was maintained at the same level as that on day 0 in these cells. Since we could not observe cells on the ELP gels because of their opacity, only cardiac gene expression levels were investigated to assess the differentiation.
Gene expression in iPSCs cultured on the substrates. a Sox2 and TnT2 expression levels in cells cultured on TCPS with or without cardiac induction on day 11 post cell seeding. The values are relative to the expression level in iPSCs at day 0 for Sox2 and the level in adult mouse heart for TnT2. Data are shown as the mean ± SEM (n = 4). An asterisk indicates significant differences analyzed by two-sided Student’s t test (p < 0.05). ND not detected. b GATA4, α-MHC, and TnT2 expression levels in iPSCs cultured on the TCPS, Col-I gel, ELP-Ctrl gel, and ELP-RGD gel with cardiac induction at day 11 post-cell seeding. The value is relative to the expression level in adult mouse heart. Data are shown as the mean ± SEM (n = 3 or 4). Asterisks indicate significant differences analyzed by one-way ANOVA followed by Tukey or Tukey–Kramer post hoc comparison (p < 0.05)
Cardiac gene expression levels in iPSCs cultured on the four substrates for 11 days are shown in Fig. 4b. Cells on TCPS, Col-I gel, and ELP-RGD gel showed high GATA4 expression levels, which were comparable to that in the adult mouse heart. Especially, cells on Col-I gel showed the highest GATA4 expression. In addition, Col-I-cultured cells presented significantly higher α-MHC expression than cells in other substrates. These results are consistent with the finding in our previous study that Col-I substrates are the most effective for the cardiac differentiation of iPSCs: gene expression of cardiac transcription factors (e.g., GATA4 and T-box 5) and contractile proteins (e.g., α-MHC and TnT2) in iPSCs cultured on Col-I substrates was the highest among four substrates (Col-I, fibronectin, gelatin, and TCPS) [6]. In contrast, TnT2 expression in cells cultured on ELP-RGD gel was significantly higher than that in cells on TCPS and Col-I gel substrates, with the expression level being four times higher expression than that in Col-I gel. In addition, iPSCs cultured on ELP-RGD hydrogel were negative for undifferentiation (Sox2), hepatic (ALB; endoderm), neural (GFAP; ectoderm), and osteogenic (Runx2; mesoderm other than myocardium) markers (Supplementary Data Table S3). Therefore, similar to Col-I substrates, the ELP-RGD substrate may be an effective substrate for the cardiac differentiation of iPSCs.
Our previous study showed that more cells adhered to an ELP-RGD substrate than to an ELP substrate without RGD [22]. The present study corroborated that more cells adhered to the ELP-RGD gel than to the ELP-Ctrl gel on day 3 (Supplementary Data Fig. S3). This suggests that RGD peptides on the surface of the ELP-RGD gel were presented to transmembrane receptors on the cells. Stem cells express integrins αIIbβ3, α5β1, and αvβ3 [13], which are transmembrane receptors of RGD. A defect in the integrin β1 in embryonic stem cells reportedly causes a delay in cardiac differentiation and an incomplete sarcomeric structure [30]. Blocking integrin β1 decreases GATA4 expression in cardiac progenitor cells [31]. In addition, a signaling pathway involving focal adhesion kinase, Stat3, and Pim1 is triggered by the interaction between fibronectin and integrin α5β1 (i.e., RGD and integrin α5β1), resulting in the promotion of cardiac progenitor cell proliferation [32]. Therefore, signaling pathways derived from integrin β1, which is one of the receptors of RGD, might have affected the cardiac differentiation of iPSCs cultured on the ELP-RGD gel. Based on the amino acid sequence of Col-I α2 chain precursor (PubMed ID, NP_001230584.1), the RGD content in the Col-I gel was calculated to be 40 µM, while that in the ELP-RGD gel was 320 µM. Efficient RGD presentation on the ELP-RGD gel might have stimulated the integrin β1-derived signaling pathway for cardiac differentiation, resulting in higher TnT2 expression in cells on the ELP-RGD gel than in those cultured on Col-I gel (Fig. 4b). Efficient presentation of a cell adhesion motif is one of the advantages of scaffolds made of recombinant protein. We are currently investigating integrin-derived signaling pathways activated in iPSCs on ELP-RGD gel.
Taken together, the results of this study suggest the potential of an ELP hydrogel as a substrate to evaluate substrate-dependent cardiac differentiation of iPSCs, enabling high yield, tuning of substrate elasticity, and presentation of a single-cell adhesion motif.
Conclusion
ELPs with or without the RGD cell adhesion motif (ELP-RGD/ELP-Ctrl) were produced recombinantly and were used to form a hydrogel. The ELP hydrogel was suggested to contain both chemical and physical crosslinks and showed a Young’s modulus of 0.3 kPa, mimicking the elasticity of embryonic cardiac tissue. Murine iPSCs were cultured on the ELP gels, and cardiac differentiation was induced by calcium-rich medium and TSA. Cells cultured on ELP-RGD gel had four times higher TnT2 (encoding a contractile protein) expression than those cultured on Col-I gel, which is reportedly one of the effective substrates for cardiac differentiation of iPSCs. The RGD sequence in ELP-RGD might affect integrin-derived signaling pathways in iPSCs to stimulate their cardiac differentiation. Owing to the ease of designing the presentation of a single-cell adhesion motif, ELP hydrogels are potentially useful substrates for the substrate-dependent cardiac differentiation of iPSCs.
References
Miskon A, Ehashi T, Mahara A, Uyama H, Yamaoka T. Beating behavior of primary neonatal cardiomyocytes and cardiac-differentiated P19CL6 cells on different extracellular matrix components. J Artif Organs. 2009;12:111–7.
Miskon A, Mahara A, Uyama H, Yamaoka T. A suspension induction for myocardial differentiation of rat mesenchymal stem cells on various extracellular matrix proteins. Tissue Eng Part C. 2010;16:979–87.
Yamaoka T, Hirata M, Dan T, Yamashita A, Otaka A, Nakaoki T, et al. Individual evaluation of cardiac marker expression and self-beating during cardiac differentiation of P19CL6 cells on different culture substrates. J Biomed Mater Res A. 2017;105:1166–74.
Hansson EM, Lindsay ME, Chien KR. Regeneration next: toward heart stem cell therapeutics. Cell Stem Cell. 2009;5:364–77.
Seo JH, Hirata M, Kakinoki S, Yamaoka T, Yui N. Dynamic polyrotaxane-coated surfaces for effective differentiation of mouse induced pluripotent stem cells into cardiomyocytes. RSC Adv. 2016;6:35668–76.
Hirata M, Yamaoka T. Effect of stem cell niche elasticity/ECM protein on the self-beating cardiomyocyte differentiation of induced pluripotent stem (iPS) cells at different stages. Acta Biomater. 2018;65:44–52.
Gai H, Leung EL, Costantino PD, Aguila JR, Nguyen DM, Fink LM, et al. Generation and characterization of functional cardiomyocytes using induced pluripotent stem cells derived from human fibroblasts. Cell Biol Int. 2009;33:1184–93.
Kaichi S, Hasegawa K, Takaya T, Yokoo N, Mima T, Kawamura T, et al. Cell line-dependent differentiation of induced pluripotent stem cells into cardiomyocytes in mice. Cardiovasc Res. 2010;88:314–23.
Minami I, Yamada K, Otsuji TG, Yamamoto T, Shen Y, Otsuka S, et al. A small molecular that promotes cardiac differentiation of human pluripotent stem cells under defined, cytokine- and xeno-free conditions. Cell Rep. 2012;2:1448–60.
Burridge PW, Matsa E, Shukla P, Lin ZC, Churko JM, Ebert AD, et al. Chemically defined generation of human cardiomyocytes. Nat Methods. 2014;11:855–60.
Jung JP, Hu D, Domian IJ, Ogle BM. An integrated statistical model for enhanced murine cardiomyocyte differentiation via optimized engagement of 3D extracellular matrices. Sci Rep. 2015;5:18705. https://doi.org/10.1038/srep18705
Macrí-Pellizzeri L, Pelacho B, Sancho A, Lglesias-García O, Simón-Yarza AM, Soriano-Navarro M, et al. Substrate stiffness and composition specifically direct differentiation of induced pluripotent stem cells. Tissue Eng Part A. 2015;21:1633–41.
Higuchi A, Kumar SS, Ling Q, Alarfaj AA, Munusamy MA, Murugan K, et al. Polymeric design of cell culture materials that guide the differentiation of human pluripotent stem cell. Prog Polym Sci. 2017;65:83–126.
Zhang S. Beyond the petri dish. Nat Biotechnol. 2004;22:151–2.
Meyer DE, Chilkoti A. Genetically encoded synthesis of protein-based polymers with precisely specified molecular weight and sequence by recursive directional ligation: examples from the elastin-like polypeptide system. Biomacromolecules. 2002;3:357–67.
Trabbic-Carlson K, Liu L, Kim B, Chilkoti A. Expression and purification of recombinant proteins from Escherichia coli: Comparison of an elastin-like polypeptide fusion with an oligohistidine fusion. Protein Sci. 2004;13:3274–84.
Engler AJ, Sen S, Sweeney HL, Discher DE. Matrix elasticity directs stem cell lineage specification. Cell. 2006;126:677–89.
Zio KD, Tirrell DA. Mechanical properties of artificial protein matrices engineered for control of cell and tissue behavior. Macromolecules. 2003;36:1553–8.
Lim DW, Nettles DL, Setton LA, Chilkoti A. Rapid crosslinking of elastin-like polypeptides with hydroxylmethylphosphines in aqueous solution. Biomacromolecules. 2007;8:1463–70.
Chung C, Lampe KJ, Heilshorn SC. Tetrakis(hydroxylmethyl) phosphonium chloride as a covalent crosslinking agent for cell encapsulation within protein-based hydrogels. Biomacromolecules. 2012;13:3912–6.
Davis GE. Affinity of integrins for damaged extracellular matrix: αVβ3 binds to denatured collagen type I through RGD sites. Biochem Biophys Res Commun. 1992;182:1025–31.
Kraehenbuehl TP, Zammaretti P, Van der Vlies AJ, Schoenmakers RG, Lutolf MP, Hubbell JA. Three-dimensional extracellular matrix-directed cardioprogenitor differentiation: systematic modulation of a synthetic cell-responsive PEG-hydrogel. Biomaterials. 2008;29:2757–66.
Mahara A, Kiicj KL, Yamaoka T. In vivo guided vascular regeneration with a non-porous elastin-like polypeptide hydrogel tubular scaffold. J Biomed Mater Res A. 2017;105:1746–55.
Kambe Y, Murakoshi A, Urakawa H, Kimura Y, Yamaoka T. Vascular induction and cell infiltration into peptide-modified bioactive silk fibroin hydrogels. J Mater Chem B. 2017;5:7557–71.
Kraehenbuehl TP, Zammaretti P, Van der Vlies AJ, Schoenmakers RG, Lutolf MP, Jaconi ME, et al. Three-dimensional extracellular matrix-derived cardioprogenitor differentiation: systematic modulation of a synthetic cell-responsive PEG-hydrogel. Biomaterials. 2008;29:2757–66.
Hirata M, Yamaoka T. Hepatocytic differentiation of iPS cells on decellularized liver tissue. J Artif Organs. 2017;20:318–25.
Livak JJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and 2−ΔΔCT method. Methods. 2001;25:402–8.
Yamaoka T, Tamura T, Seto Y, Tada T, Kunugi S, Tirrell DA. Mechanism for the phase transition of a genetically engineered elastin model peptide (VPGIG)40 in aqueous solution. Biomacromolecules. 2003;4:1680–5.
Trabbic-Carlson K, Setton LA, Chilkoti A. Swelling and mechanical behaviors of chemically cross-linked hydrogels of elastin-like polypeptides. Biomacromolecules. 2003;4:572–80.
Fässler R, Rohwedel J, Maltsev V, Bloch W, Lentini S, Guan K, et al. Differentiation and integrity of cardiac muscle cells are impaired in the absence of β1 integrin. J Cell Sci. 1996;109:2989–99.
Hodgkinson CP, Gomez JA, Payne AJ, Zhang L, Wang X, Dal-Pra S, et al. Abi3bp regulates cardiac progenitor cell proliferation and differentiation. Circ Res. 2014;115:1007–17.
Konstandin MH, Toko H, Gastelum GM, Quijada P, De La Torre A, Quintana M, et al. Fibronectin is essential for regenerative cardiac progenitor cell response following myocardial infarction. Circ Res. 2013;113:115–25.
Acknowledgements
This work was supported financially in part by the Intramural Research Funds from the NCVC (25-2-2 and 29-6-2) and by the Japan Society for the Promotion of Science Grant-in-Aid for Challenging Exploratory Research (26560251).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Kambe, Y., Tokushige, T., Mahara, A. et al. Cardiac differentiation of induced pluripotent stem cells on elastin-like protein-based hydrogels presenting a single-cell adhesion sequence. Polym J 51, 97–105 (2019). https://doi.org/10.1038/s41428-018-0110-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41428-018-0110-2