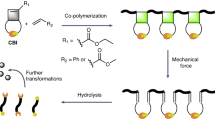
Abstract
A DNA building block with tunable rigidity was constructed, and the corresponding hydrogel formation process was investigated accordingly. A high rigidity was demonstrated to facilitate fast gelation. Different gelation pathways of the rigid and flexible building blocks were revealed, and a cyclized dimer intermediate was proposed. The energy barrier of the ring-opening process was also shown to play a fundamental role in determining the gelation kinetics. Furthermore, the hydrogel molecular network rigidity was also tuned in situ through strand displacement, which also supports the kinetic control mechanism of the formation process of DNA hydrogels.
Similar content being viewed by others
Introduction
Hydrogels are a class of three-dimensional network materials with a high water content1,2,3 that have been widely utilized in biomedical applications4,5,6,7. During the past few decades, DNA has arisen as a promising material for the construction of supramolecular hydrogels through complementary base pairing8,9,10,11,12. The rigid DNA duplex network excludes small size meshes and endows hydrogels with high permeability13,14. Moreover, the dynamic properties that arise from DNA hybridization also endow the hydrogel with excellent injectability and self-healing and shear-thinning properties15,16,17,18. Furthermore, the designability of the DNA sequence has also endowed DNA supramolecular hydrogels with good stimulus responsiveness19,20,21,22,23,24. Therefore, DNA supramolecular hydrogels have been used in three-dimensional cell culture25,26, spinal cord repair27, osteoarthritis treatment28, heavy metal ion detection29,30, and drug delivery31,32,33.
Compared with that in traditional hydrogels, the distance of the crosslinking points in DNA supramolecular hydrogels is usually shorter than the persistence length of the DNA double strand, which endows DNA supramolecular hydrogels with rigid networks13,14,27. It has been well demonstrated that the rigidity of the molecular network plays important roles in the mechanical properties of hydrogels34,35,36,37. For example, Liu et al. introduced the conformational transition of the building block between a flexible single strand and an i-motif quadruplex to tune the rigidity of the molecular network, which increased the mechanical strength from 250 to 1000 Pa34. The same group also reported that the collapse of PPO (polypropylene oxide) segments in the hydrogel network could increase its rigidity, thus improving the mechanical strength35. These examples have successfully demonstrated that the molecular network rigidity determines the mechanical strength of DNA supramolecular hydrogels.
In previous studies, the fast formation of DNA supramolecular hydrogels within seconds was observed, which was explained by the rigidity of the building blocks19. The rigidity potentially determines the gelation process by tuning the dynamic balance between the cyclization and chain extension of the building blocks, which has been proven in linear supramolecular polymerization systems38,39,40. For example, Zhang et al. designed a bifunctional monomer with self-sorting properties and demonstrated that the increased monomer rigidity could further promote supramolecular polymerization38. Liu et al. also reported that the formation of a rigid duplex in the central part of flexible single-stranded DNA facilitated ring-opening polymerization39. While the importance of monomer rigidity regarding supramolecular polymerization has been well recognized, how the rigidity of the building block affects the supramolecular gelation process has rarely been studied.
Herein, we constructed a rigidity-tunable building block to investigate the kinetic process of hydrogel formation through backbone remodeling. Different reaction pathways of the rigid and flexible building blocks were revealed, and the conversion between intramolecular and intermolecular interactions was confirmed. Furthermore, the hydrogel molecular network rigidity was also tuned in situ, supporting the kinetic control mechanism. Our results revealed the rigidity effect of building blocks on hydrogel formation kinetics, which may benefit the application of DNA supramolecular hydrogels in smart devices and biomimicry materials.
Results and discussion
As illustrated in Fig. 1, the branched DNA molecule (B-i) was composed of three identical arms, which were chemically bonded to a central unit. A half i-motif sequence was integrated at the end of each arm as a crosslinker (11 nt, blue), which could be partially protonated and formed a full quadruplex i-motif structure with another arm under acidic conditions. For B-i, the core region was a 15 nt single-stranded DNA, which was longer than its persistent length of 1 nm and can be defined as a flexible building block41. To tune the rigidity of the building block, a backbone segment (15 nt, orange) was also designed in the core region. With the addition of the complementary sequence cBi, duplex structures were formed, whose lengths were shorter than the estimated persistence length42,43,44, thus transforming into a rigid form (RB). Then, the different gelation kinetic processes between the flexible and rigid building blocks were investigated by changing the order of pH adjustment and DNA hybridization.
As previously reported15, commercially available branched phosphoramidite was employed for the synthesis of branched B-i molecule (the sequence can be found in Table S1). The detailed synthetic process can also be found in Fig. S1, and the product was characterized by 20% denaturing polyacrylamide gel electrophoresis (PAGE). As shown in Fig. 2a, the sharp and clear band (Lane 4) indicated the high purity of the synthesized product. Compared with the linear sequences I, 2I, and 3I (with the same repeated sequence as B-i, illustrated in Table S1), B-i showed a slower migration rate, which was due to the larger hydrodynamic volume of the branched structure. The molecular weight was detected as 23,926 g/mol by matrix-assisted laser desorption/ ionization time-of-flight mass spectrum (MALDI-TOF MS, Fig. S2), which matched the theoretical molecular weight of 23,913 g/mol. These results indicated that the branched DNA molecule B-i was successfully synthesized and purified.
Then, the assembly behavior of B-i was studied. As shown in Fig. 2b, under 10% native PAGE, when cBi (sequence can be found in Table S1) was added to B-i in a molecular ratio of 3:1 at pH 8.00, the clear band suggested efficient hybridization between B-i and cBi (Lane 5). Furthermore, when the molecular ratio decreased to 2:1, four bands appeared (Fig. S3, Lane 3), which corresponded to B-i, B-i+cBi, B-i + 2cBi, and B-i + 3cBi. These results can be explained by the fact that the assemblies of each arm of B-i and cBi were random and independent of each other. The assembly behavior of B-i and cBi under basic conditions suggested that we could construct a rigid building block (RB) through DNA hybridization.
Next, we investigated the formation of the i-motif by circular dichroism (CD) spectroscopy. As shown in Fig. 2c and Fig. S4, there was a positive peak at 282 nm, a negative peak at 246 nm and a crossover at 257 nm under acidic conditions for both B-i and RB, which reflected the formation of the i-motif. It is worth noting that the CD intensity of the i-motif was very similar at different pH values for B-i (Fig. 2c), while the i-motif signal intensity decreased with increasing pH for RB (Fig. S4). As a control, the CD spectrum of linear sequence I was also investigated, which could only form an intermolecular i-motif. The pH-dependent density of I was observed (Fig. S5) to be similar to that of RB, suggesting the potential intermolecular i-motif in RB and the intramolecular i-motif in B-i. These results indicated that the rigidity difference of the branched units would result in different crosslinking types, thus allowing the tuning the gelation kinetic process of hydrogel formation.
To investigate the rigidity effect, we first prepared a DNA supramolecular hydrogel from a rigid building block. After the hybridization of B-i and cBi at a molar ratio of 1:3 at pH 8.00, acidic buffer was added to tune the pH to 5.50 and obtain final concentrations of 750 μM for B-i and 2250 μM for cBi. As shown in Fig. 3a, the clear slope appearing in the tube within seconds after pH adjustment indicated the fast gelation process, which was consistent with the quick formation of the i-motif.
a Digital photographs of the gelation process of the rigid building block. b Rheological test of the gelation process by a time sweep test with a fixed strain of 1% and frequency of 1 Hz at 25 °C. c Rheological test of strain scanning of the DNA hydrogel with strains ranging from 0.1 to 1000% at 25 °C and 1 Hz. d Rheological test of temperature scanning of the DNA hydrogel with temperatures ranging from 25 to 45 °C with a fixed strain of 1% and frequency of 1 Hz.
Rheological tests were then performed to study the properties of this DNA supramolecular hydrogel. As shown in Fig. 3b, the mechanical strength stabilized quickly, and the storage modulus (G’, 221.9 Pa) was higher than the loss modulus (G”, 1.7 Pa), implying a quick gelation process at pH 5.50. G’ was higher than G” over the frequency sweep range (Fig. S6), which is typical hydrogel behavior. In the oscillatory strain-dependent rheology test (Fig. 3c), the G’ of the hydrogel decreased rapidly after 40% strain and had a crossing point with G” at 80% strain, which implied that this hydrogel exhibited a non-Newtonian fluid shear-thinning property. In the temperature mode rheology test (Fig. 3d), as the temperature increased, G’ decreased, and both G’ and G” were lower than 10 Pa after 33 °C, indicating a gel-sol transition. Hence, the rigid building block could quickly form a DNA supramolecular hydrogel with shear-thinning and temperature responsive properties.
Following the same pH adjustment strategy, we also applied a flexible building block to prepare a DNA supramolecular hydrogel. As shown in Fig. S7, small droplets on the inner wall of the EP tube were observed in the invert-vail test, indicating that the system remained in a solution state. Then, the solution was incubated at 4 °C for 24 h, but no hydrogel was formed. Based on the rigidity difference between B-i and RB, we assumed that the molecular rigidity of the building block was critical to gelation and that the increased rigidity of B-i would facilitate gelation under acidic conditions.
Backbone remodeling was then employed to reveal the dynamic effect. After the pH was adjusted to 5.50, cBi was added to increase the rigidity of the flexible building block (B-i). The system gradually lost its fluidity in the invert-vail test and finally formed a hydrogel after several hours (Fig. 4a), which was different from the rapid gelation process in RB. The rheological properties of this DNA supramolecular hydrogel were also studied, and G’ gradually increased from 4 to 120 Pa in ~5 h (Fig. 4b). In subsequent tests, the hydrogel was incubated at 4 °C for 24 h after backbone remodeling to achieve complete gelation. The mechanical strength of this hydrogel (G’, 211.8 Pa) after incubation was similar to that of the hydrogel in RB (G’, 221.9 Pa). Furthermore, the same shear-thinning and temperature responsive properties to the RB hydrogel were also confirmed by rheological tests (Fig. 4c, d). We also applied field emission scanning electron microscopy (FE-SEM) to investigate the micromorphology transition during the gelation process. As shown in Fig. S8a, the pore size was very small, and no lamellar structure was observed at 0 h. After incubating for 4 h, the pore size increased, and the walls became thicker (Fig. S8b). After 24 h of incubation, a morphology similar to that of RB hydrogel was observed (Fig. S8c, d). These results indicated that compared with the rigid building block, the backbone remodeling of the flexible building block facilitated a slow formation of the hydrogel through different microscopic processes, but the same molecular topological network could be achieved.
a Digital photographs of the gelation process through backbone remodeling. b Rheological test of the gelation process by a time sweep test with a fixed strain of 1% and frequency of 1 Hz at 4 °C. c Rheological test of strain scanning of the DNA hydrogel with strains ranging from 0.1 to 1000% with a fixed frequency of 1 Hz at 25 °C. d Rheological test of temperature scanning of the DNA hydrogel with temperatures ranging from 20 to 50 °C with a fixed strain of 1% and frequency of 1 Hz.
To reveal the pathways of the gelation process from different building blocks, we further explored the assembly behavior of B-i and RB at different pH values. At pH 5.50, as illustrated in Fig. 5a, large aggregates were formed in the RB sample after pH adjustment (Lane 5), indicating the formation of a network assembly. On the other hand, the single band of B-i (Lanes 2–4) was observed to have a slower migration rate than a random branched molecule (the sequence can be found in Table S1), which indicated that a specific assembly containing at least two B-i was formed. Combined with the CD spectrum of B-i (Fig. 2c), it can be assumed, as illustrated in Fig. 1, that two adjacent arms in the same molecule formed an intramolecular i-motif and the remaining arm formed an intermolecular i-motif with another molecule. The backbone remodeling process from the flexible building block to the rigid building block was also investigated, and as illustrated in Fig. 5a, similar aggregates (Lane 6) were also observed after the introduction of cBi to flexible Bi at pH 5.50 (Lane 5) and subsequent incubation at 4 °C for 24 h, indicating the complete transition from the flexible state to the rigid state.
a Characterization of the assembly behavior of B-i at pH 5.50 by 10% native PAGE. Lane 1 to Lane 7: Random, B-i at 10 μM, 50 μM, 100 μM, RB, backbone remodeling of B-i and subsequently incubated for 24 h, cBi. b Characterization of the assembly behavior of B-i at pH 5.00 by 10% native PAGE. Lane 1 to Lane 8: Random, B-i at 10 μM, 50 μM, 100 μM, RB, backbone remodeling of B-i after subsequently incubating for 10 days, for 24 h, and cBi.
Based on the above experiments, we put forward the following molecular mechanism: Under acidic conditions, for the rigid building block RB, the DNA duplex in the core region could inhibit intramolecular cyclization and promote the quick formation of an intermolecular i-motif network. In contrast, the flexible building block B-i preferred to form a thermodynamically stable dimer structure (as illustrated in Fig. 1), which cannot form a crosslinked hydrogel molecular network. The hybridization of cBi to flexible B-i under acidic conditions could enhance its rigidity, which would potentially break the intramolecular i-motif and subsequently form the intermolecular i-motif network. The stability balance between the i-motif and the duplex played an important role in this backbone remodeling process, and therefore, the corresponding gelation was a slow process.
To further reveal the mechanism of the ring-opening process, the pH-dependent gelation process was investigated. As the pH decreased from 6.00 to 5.00, the stability of the intramolecular i-motif was increased (indicated by the melting temperature, summarized in Table S2), which raised the energy barrier. In the gelation experiments, we found that RB could still form a hydrogel quickly after the pH was adjusted to 5.00 (rheological properties are shown in Fig. S9). However, the flexible Bi was still in the solution state under the same pH even after 24 h at 4 °C with the addition of cBi (Fig. S10), indicating that the stable intramolecular i-motif inhibits ring opening. Native PAGE also supported this conclusion, and at pH 5.00 (Fig. 5b), no larger assembly was formed after 24 h (Lane 7), while most dimers were retained after 10 days (Lane 6). On the other hand, when the pH was tuned to 6.00, the flexible Bi could form the hydrogel ~4 h after the addition of cBi (Fig. S11), which can be explained by the higher stability of the duplex than the i-motif. It should be noted that due to the low stability of the i-motif, the hydrogel could not be maintained at room temperature under pH 6.00, which was consistent with the rigid system (Fig. S12). However, the faster gelation speed at pH 6.00 (<4 h) than that at pH 5.50 (~5 h) at 4 °C could still support the ring-opening mechanism. These results showed the important role of the energy barrier in opening the ring and further indicated the importance of the rigidity of the building blocks.
The molecular network of the hydrogel rigidity was also tuned in situ through strand displacement (Fig. 6a). We have prepared a Lc sequence, which extended toward cBi (the sequence can be found in Table S1). As illustrated in Fig. 6b, this Lc-RB could also form the molecular network (Lane 4) and the hydrogel (Fig. S13). Then, a fuel strand (fully complementary to Lc, Table S1) was added to remove Lc from the network to change its rigidity at 4 °C for 24 h. To demonstrate successful strand displacement, fluorescence-labeled Lc was applied and analyzed by native PAGE. After incubation, we found that the aggregates still existed (Fig. 6b, Lane 5), but no fluorescence signal could be observed in the sample well under fluorescence imaging (Fig. 6c, Lane 3). We found that after strand displacement, the hydrogel state was still maintained (Fig. S11), which indicated that the hydrogel network would not collapse even when Lc was completely removed. We also applied a rheological test to investigate the properties of the hydrogel after strand displacement. As shown in Fig. 6d and e, G’ decreased from 1230 to 443.7 Pa, and the gel-sol transition point increased from 18.0 to 55.4% during the strain sweep after backbone remodeling. These results indicated that the flexible network was thermally stable and that gelation was kinetically controlled by the ring-opening process. Our strategy has also allowed the preparation of hydrogels with both rigid and flexible networks, which determines the rheological properties of the hydrogel.
a Schematic of the network remodeling process. b Characterization of the strand displacement reaction at pH 5.50 by 10% native PAGE. Lane 1 to Lane 7: Random, B-i, Lc, B-i:Lc = 1:3, Bi:Lc:Fuel = 1:3:3, Lc+Fuel, Fuel. c Characterization of the strand displacement reaction at pH 5.50 by fluorescence 10% native PAGE. Lane 1 to Lane 4: Lc, B-i:Lc = 1:3, Bi:Lc:Fuel = 1:3:3, Lc+Fuel. d Rheological test of strain scanning of the DNA hydrogel with strains ranging from 0.1 to 1000% with a fixed frequency of 1 Hz at 4 °C and pH 5.50. e Mechanical strength of the hydrogels before or after strand displacement at 4 °C and pH 5.50.
Conclusion
In summary, the rigidity effect of the building blocks on the formation process of DNA hydrogels has been investigated. By tuning the hybridization state, the rigidity of the branched DNA molecule was adjusted, and different gelation pathways of the rigid and flexible building block systems were revealed. It has been observed that a high rigidity could inhibit the intramolecular cyclization of the branched molecule and promote the quick formation of the hydrogel network. The energy barrier of the ring-opening process can be affected by the stability balance between the rigid and flexible states of the building block, which therefore plays a fundamental role in determining the backbone remodeling process. We also proved that the molecular network could still be maintained after the building block was adjusted to the flexible state, which further demonstrated that gelation is kinetically controlled by the ring-opening process. It is anticipated that our results could further deepen the understanding of the relationship between building block rigidity and the gelation process, which would expand the application of DNA hydrogels in biomimetic materials and smart devices.
Materials and methods
Materials
Acrylamide, N,N-methylene bis-acrylamide, ammonium persulfate (APS), ammonium hydroxide (NH3•H2O), 2-morpholineethanesulfonic acid (MES), trifluoroacetic acid, sodium chloride (NaCl), tris(hydroxymethyl)-aminomethane (Tris), ethylenediaminetetraacetic acid disodium salt (EDTA•2Na), boric acid (B(OH)3), hydrochloric acid (HCl), sodium hydroxide (NaOH), triethylamine, and acetic acid were purchased from Sigma‒Aldrich. A branched phosphonamidite called Long Trebler was purchased from Glen Research. All reagents used for DNA solid-phase synthesis (CPG, four phosphoramidite monomers, oxidant, activator, acetonitrile, Cap A, Cap B) were purchased from Hebei Dinaxingke (China). A 0.22 μm polyether sulfone filter membrane was purchased from Millipore.
DNA synthesis
All the molecules were synthesized by a Mermade 12 DNA synthesizer (Bioautomation Company) using commercially available phosphonamidites and CPG on a 1 μmol scale. For the synthesis of B-i, the coupling time of Long Trebler was increased to 15 min. After the synthesis, the CPG loaded with oligonucleotides was placed in concentrated ammonium hydroxide and heated at 60 °C for 3 h to cleave the DNA from CPG. Then, the oligonucleotides were purified by RP-HPLC (Agilent Technologies) using acetonitrile and triethylammonium acetate buffer for elution. The purified oligonucleotides were treated with trifluoroacetic acid to remove the DMT group at the 5’ end and remove salts by ultrafiltration. All synthesized DNA molecules were characterized by 20% denaturing PAGE and MALDI-TOF (Shimadazu Biotech Axima Performance).
Polyacrylamide gel electrophoresis (PAGE) analysis
The 20% denaturing PAGE (Acrylamide: Bis-acrylamide = 19:1, 7 M Urea) was used to characterize the synthesis of oligonucleotides with 1× TBE (Tris-B(OH)3-EDTA•2Na) as running buffer. For the analysis of the assembly behavior, pH 8.00 10% native PAGE (acrylamide:bis-acrylamide = 19:1) with 1× TBE as running buffer and pH 5.00/5.50 10% native PAGE with MES buffer (50 mM MES and 12.5 mM MgAc2) as running buffer was performed at 4 °C. For fluorescence native PAGE, 2% FAM-modified Lc was added.
Circular dichroism (CD) spectrum measurement
The oligonucleotides were dissolved in MES buffer (50 mM MES and 137 mM NaCl) at different pH values, and then the samples were heated to 95 °C for 5 min and gently cooled to room temperature. The circular dichroism spectrum of the sample in the wavelength range of 225–320 nm was collected by an Applied Photophysics Chirascan Spectropolarimeter at room temperature with a 1.0 cm sample cell. Each sample was measured three times and averaged. The scan of the buffer alone was used as a control and subtracted from the average scan for each sample.
Melting point measurement
The melting profile of the i-motif was measured by an Applied Photophysics Chirascan Spectropolarimeter. The temperature was scanned from 5 to 90 °C at a scan rate of 1 °C/min. For both B-i and I, the CD signal at 282 nm was detected. The melting profile of the DNA duplex was measured by an ultraviolet and visible (UV‒Vis) spectrophotometer (Agilent Cary 100), and the absorbance at 260 nm was detected. The cell length was 1.0 cm in all measurements. The melting point (Tm) was obtained from the peak of the corresponding differential melting curve.
Preparation of DNA supramolecular hydrogels
For rigid building blocks, both B-i and cBi were first dissolved in pH 8.00 MES buffer (5 mM MES and 137 mM NaCl), heated to 95 °C for 5 min and cooled to room temperature naturally. Then, 500 mM MES buffer (pH = 5.00/5.50/6.00) was added to the sample to tune the buffer to acidic with a final MES concentration of 50 mM. The final concentration of B-i was 750 μM, and that of cBi was 2250 μM.
For the flexible building block, B-i was directly dissolved in an acidic MES buffer (50 mM MES and 137 mM NaCl, pH = 5.00/5.50/6.00) and further incubated at 4 °C for 24 h. The concentration of B-i was 750 μM.
For the backbone remodeling process, both B-i and cBi were individually dissolved in an acidic MES buffer (50 mM MES and 137 mM NaCl, pH = 5.00/5.50/6.00) to reach a concentration of 1.5 mM for B-i and 4.5 mM for cBi. Then, these two solutions were mixed in equal volumes and incubated at 4 °C for 24 h to form the DNA hydrogel. The final concentration of B-i was 750 μM, and that of cBi was 2250 μM.
For the in situ tuning of the rigidity of the hydrogel molecular network through strand displacement, both B-i and Lc were first dissolved in pH 8.00 MES buffer (5 mM MES and 137 mM NaCl), heated to 95 °C for 5 min and cooled to room temperature naturally. Then, 500 mM MES buffer (pH = 5.00/5.50/6.00) was added to the sample to tune the buffer to acidic with a final MES concentration of 50 mM. After 24 h of incubation at 4 °C, a fuel strand fully complementary to Lc was added and further incubated at 4 °C for another 24 h to ensure complete strand displacement. The final concentration was 750 μM for B-i and 2250 μM for Lc and the fuel strand.
Rheological tests
A Kinexus Pro+ rheometer (Malvern Instruments) was used to test the rheological properties of the formed DNA hydrogels in the following mode: (i) a time sweep test was carried out at a fixed strain of 1% and frequency of 1 Hz at 25 °C for 5 min; (ii) strain sweep tests were carried out from 0.1 to 100% with a fixed frequency of 1 Hz at 25 °C; (iii) a frequency sweep was performed from 10 to 0.01 Hz with a strain of 1%. (iv) Temperature tests were performed at a frequency of 1 Hz, strain of 1%, and heating rate of 1 °C/min over different temperature ranges.
Field emission scanning electron microscopy (FE-SEM) test
Samples were collected at different incubation time points during gelation. Then, they were lyophilized for 24 h. The freeze-dried samples were broken off with tweezers and fixed on the sample plate with conductive adhesive. The samples were coated with gold before testing.
References
Appel, E. A., del Barrio, J., Loh, X. J. & Scherman, O. A. Supramolecular polymeric hydrogels. Chem. Soc. Rev. 41, 6195–6214 (2012).
Yang, C. H. & Suo, Z. G. Hydrogel ionotronics. Nat. Rev. Mater. 3, 125–142 (2018).
Ahmed, E. M. Hydrogel: preparation, characterization, and applications: a review. J. Adv. Res. 6, 105–121 (2015).
Li, J. et al. Short and simple peptide-based pH-sensitive hydrogel for antitumor drug delivery. Chin. Chem. Lett. 33, 1936–1940 (2022).
Plamper, F. A. & Richtering, W. Functional microgels and microgel systems. Acc. Chem. Res. 50, 131–140 (2017).
Lutolf, M. P. Spotlight on hydrogels. Nat. Mater. 8, 451–453 (2009).
Liu, X. Y., Liu, J., Lin, S. T. & Zhao, X. H. Hydrogel machines. Mater. Today 36, 102–124 (2020).
Nagahara, S. & Matsuda, T. Hydrogel formation via hybridization of oligonucleotides derivatized in water-soluble vinyl polymers. Polym. Gels Netw. 4, 111–127 (1996).
Li, F., Tang, J. P., Geng, J. H., Luo, D. & Yang, D. Y. Polymeric DNA hydrogel: design, synthesis and applications. Prog. Polym. Sci. 98, 101163 (2019).
Li, F. Y., Lyu, D. Y., Liu, S. & Guo, W. W. DNA hydrogels and microgels for biosensing and biomedical applications. Adv. Mater. 32, 1806538 (2020).
Shi, J. Z., Shi, Z. W., Dong, Y. C., Wu, F. & Liu, D. S. Responsive DNA-based supramolecular hydrogels. ACS Appl. Bio Mater. 3, 2827–2837 (2020).
Dong, Y. H. et al. DNA functional materials assembled from branched DNA: design, synthesis, and applications. Chem. Rev. 120, 9420–9481 (2017).
Shao, Y., Jia, H. Y., Cao, T. Y. & Liu, D. S. Supramolecular hydrogels based on DNA self-assembly. Acc. Chem. Res. 50, 659–668 (2017).
Jin, J. et al. A triggered DNA hydrogel cover to envelop and release single cells. Adv. Mater. 25, 4714–4717 (2013).
Yang, B. et al. Shear-thinning and designable responsive supramolecular DNA hydrogels based on chemically branched DNA. ACS Appl. Mater. Interfaces 13, 48414–48422 (2021).
Li, C. et al. Rapid formation of a supramolecular polypeptide-DNA hydrogel for in situ three-dimensional multilayer bioprinting. Angew. Chem. Int. Ed. 54, 3957–3961 (2015).
Li, C. et al. A writable polypeptide–DNA hydrogel with rationally designed multi-modification sites. Small 11, 1138–1143 (2015).
Peng, L. et al. Macroscopic volume change of dynamic hydrogels induced by reversible DNA hybridization. J. Am. Chem. Soc. 134, 12302–12307 (2012).
Chen, E. J. et al. A pH-triggered, fast-responding DNA hydrogel. Angew. Chem. Int. Ed. 48, 7660–7663 (2009).
Liu, H. Y., Cao, T. Y., Xu, Y., Dong, Y. C. & Liu, D. S. Tuning the mechanical properties of a DNA hydrogel in three phases based on ATP aptamer. Int. J. Mol. Sci. 19, 1633 (2018).
Li, C. et al. Responsive double network hydrogels of interpenetrating DNA and CB[8] host-guest supramolecular systems. Adv. Mater. 27, 3298–3304 (2015).
Vázquez-González, M. & Willner, I. Stimuli-responsive biomolecule-based hydrogels and their applications. Angew. Chem. Int. Ed. 59, 15342–15377 (2020).
Lu, C. H., Guo, W. W., Hu, Y. W., Qi, X. J. & Willner, I. Multitriggered shape-memory acrylamide–DNA hydrogels. J. Am. Chem. Soc. 137, 15723–15731 (2015).
Ren, J. T. et al. pH-responsive and switchable triplex-based DNA hydrogels. Chem. Sci. 6, 4190–4195 (2015).
Yang, B. et al. A biostable L-DNA hydrogel with improved stability for biomedical applications. Angew. Chem. Int. Ed. 61, e202202520 (2022).
Wu, J. Y. et al. Self-assembly of dendritic DNA into a hydrogel: application in three-dimensional cell culture. ACS Appl. Mater. Interfaces 13, 49705–49712 (2021).
Yuan, T. Y. et al. Highly permeable DNA supramolecular hydrogel promotes neurogenesis and functional recovery after completely transected spinal cord injury. Adv. Mater. 33, 2102428 (2021).
Yan, X. et al. Anti-friction MSCs delivery system improves the therapy for severe osteoarthritis. Adv. Mater. 33, 2104758 (2021).
Joseph, K. A., Dave, N. & Liu, J. W. Electrostatically directed visual fluorescence response of DNA-functionalized monolithic hydrogels for highly sensitive Hg2+ detection. ACS Appl. Mater. Interfaces 3, 733–739 (2011).
Huang, Y. S. et al. Target-responsive DNAzyme cross-linked hydrogel for visual quantitative detection of lead. Anal. Chem. 86, 11434–11439 (2014).
Li, J. et al. Self-assembly of DNA nanohydrogels with controllable size and stimuli-responsive property for targeted gene regulation therapy. J. Am. Chem. Soc. 137, 1412–1415 (2015).
Ding, F. et al. A crosslinked nucleic acid nanogel for effective siRNA delivery and antitumor therapy. Angew. Chem. Int. Ed. 57, 3064–3068 (2018).
Mo, F. L. et al. DNA hydrogel-based gene editing and drug delivery systems. Adv. Drug Deliv. Rev. 168, 79–98 (2021).
Zhou, X. et al. Reversibly tuning the mechanical properties of a DNA hydrogel by a DNA nanomotor. Chem. Commun. 52, 10668–10671 (2016).
Wu, F. et al. Self-collapsing of single molecular poly-propylene oxide (PPO) in a 3D DNA network. Small 14, 1703462 (2018).
Xing, Z. Y. et al. Microrheology of DNA hydrogels. Proc. Natl Acad. Sci. USA 115, 8137–8142 (2018).
Stoev, I. D. et al. On the role of flexibility in linker-mediated DNA hydrogels. Soft. Mater. 16, 990–1001 (2020).
Huang, Z. H. et al. Supramolecular polymerization promoted and controlled through self-sorting. Angew. Chem. Int. Ed. 53, 5351–5355 (2014).
Jia, H. Y. et al. Controllable supramolecular “ring opening” polymerization based on DNA duplex. Polymer 171, 121–126 (2019).
Zhang, W. et al. Controllable fabrication of a supramolecular polymer incorporating twisted cucurbit[14]uril and cucurbit[8]uril via self-sorting. Chin. Chem. Lett. 33, 2455–2458 (2022).
Yang, D. Y. et al. DNA materials: bridging nanotechnology and biotechnology. Acc. Chem. Res. 47, 1902–1911 (2014).
Smith, S. B., Cui, Y. J. & Bustamante, C. Overstretching B-DNA: the elastic. response of individual double-stranded and single-stranded DNA molecules. Science 271, 795–799 (1996).
Baumann, C. G., Smith, S. B., Bloomfield, V. A. & Bustamante, C. Ionic effects on the elasticity of single DNA molecules. Proc. Natl Acad. Sci. USA 94, 6185–6190 (1997).
Mitchell, J. S., Glowacki, J., Grandchamp, A. E., Manning, R. S. & Maddocks, J. H. Sequence-dependent persistence lengths of DNA. J. Chem. Theory Comput. 13, 1539–1555 (2017).
Acknowledgements
This work was financially supported by the Institute of Chemistry CAS (C220200701), National Natural Science Foundation of China (21890731, 21821001, 21534007, 21971248, and 21890730), and National Basic Research Plan of China (2018YFA0208900).
Author information
Authors and Affiliations
Contributions
D.L. and Y.D. conceived the idea and designed the experiment. Y.P. and B.Y. conducted the experiments and data analysis. Y.P. wrote the manuscript. R.X., X.L., Y.D., and D.L. assisted with writing of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Pan, Y., Yang, B., Xu, R. et al. Rigidity-dependent formation process of DNA supramolecular hydrogels. NPG Asia Mater 14, 92 (2022). https://doi.org/10.1038/s41427-022-00445-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41427-022-00445-w