Abstract
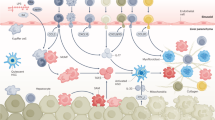
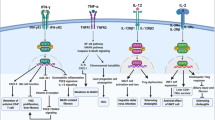
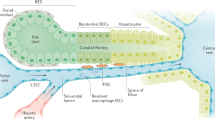
The hepatic immune system is designed to tolerate diverse harmless foreign moieties to maintain homeostasis in the healthy liver. Constant priming and regulation ensure that appropriate immune activation occurs when challenged by pathogens and tissue damage. Failure to accurately discriminate, regulate, or effectively resolve inflammation offsets this balance, jeopardizing overall tissue health resulting from an either overly tolerant or an overactive inflammatory response. Compelling scientific and clinical evidence links dysregulated hepatic immune and inflammatory responses upon sterile injury to several pathological conditions in the liver, particularly nonalcoholic steatohepatitis and ischemia-reperfusion injury. Murine and human studies have described interactions between diverse immune repertoires and nonhematopoietic cell populations in both physiological and pathological activities in the liver, although the molecular mechanisms driving these associations are not clearly understood. Here, we review the dynamic roles of inflammatory mediators in responses to sterile injury in the context of homeostasis and disease, the clinical implications of dysregulated hepatic immune activity and therapeutic developments to regulate liver-specific immunity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Knolle, P. A. & Wohlleber, D. Immunological functions of liver sinusoidal endothelial cells. Cell. Mol. Immunol. 13, 347–353 (2016).
Kubes, P. & Jenne, C. Immune responses in the liver. Annu. Rev. Immunol. 36, 247–277 (2018).
Surewaard, B. G. J. & Kubes, P. Measurement of bacterial capture and phagosome maturation of Kupffer cells by intravital microscopy. Methods 128, 12–19 (2017).
Zeng, Z. et al. CRIg functions as a macrophage pattern recognition receptor to directly bind and capture blood-borne gram-positive bacteria. Cell Host Microbe 20, 99–106 (2016).
Horst, A. K. et al. Modulation of liver tolerance by conventional and nonconventional antigen-presenting cells and regulatory immune cells. Cell. Mol. Immunol. 13, 277–292 (2016).
Thomson, A. W. et al. Immunobiology of liver dendritic cells. Immunol. Cell Biol. 80, 65–73 (2002).
Webb, L. M. et al. Type I interferon is required for T helper (Th) 2 induction by dendritic cells. EMBO J. 36, 2404–2418 (2017).
Kelly, A. et al. CD141+ myeloid dendritic cells are enriched in healthy human liver. J. Hepatol. 60, 135–142 (2014).
Nati, M. et al. The role of immune cells in metabolism-related liver inflammation and development of non-alcoholic steatohepatitis (NASH). Rev. Endocr. Metab. Disord. 17, 29–39 (2016).
Ochando, J. et al. Tolerogenic dendritic cells in organ transplantation. Transpl. Int. 33, 113–127 (2020).
Arrese, M. et al. Innate immunity and inflammation in NAFLD/NASH. Digestive Dis. Sci. 61, 1294–1303 (2016).
Dou, L. et al. Hepatic dendritic cells, the tolerogenic liver environment, and liver disease. Semin Liver Dis. 38, 170–180 (2018).
Tsuda, Y. et al. An immunosuppressive subtype of neutrophils identified in patients with hepatocellular carcinoma. J. Clin. Biochem. Nutr. 51, 12–32 (2012).
Pillay, J. et al. Immune suppression by neutrophils and granulocytic myeloid-derived suppressor cells: similarities and differences. Cell. Mol. Life Sci. 70, 3813–3827 (2013).
Coffelt, S. B. et al. IL-17-producing γδ T cells and neutrophils conspire to promote breast cancer metastasis. Nature 522, 345–348 (2015).
Gabrilovich, D. I. Myeloid-derived suppressor cells. Cancer Immunol. Res. 5, 3–8 (2017).
Yao, L. et al. Characterization of liver monocytic myeloid-derived suppressor cells and their role in a murine model of non-alcoholic fatty liver disease. PLoS One 11, e0149948 (2016).
Veglia, F., Perego, M. & Gabrilovich, D. Myeloid-derived suppressor cells coming of age. Nat. Immunol. 19, 108–119 (2018).
Jenne, C. N. & Kubes, P. Immune surveillance by the liver. Nat. Immunol. 14, 996–1006 (2013).
Doherty, D. G. & O’Farrelly, C. Innate and adaptive lymphoid cells in the human liver. Immunological Rev. 174, 5–20 (2000).
Shen, Y. et al. Ambiguous roles of innate lymphoid cells in chronic development of liver diseases. World J. Gastroenterol. 24, 1962–1977 (2018).
Kenna, T. et al. NKT cells from normal and tumor-bearing human livers are phenotypically and functionally distinct from murine NKT cells. J. Immunol. 171, 1775–1779 (2003).
Dusseaux, M. et al. Human MAIT cells are xenobiotic-resistant, tissue-targeted, CD161hi IL-17-secreting T cells. Blood 117, 1250–1259 (2011).
Nemeth, E., Baird, A. W. & O’Farrelly, C. Microanatomy of the liver immune system. Semin Immunopathol. 31, 333–343 (2009).
Clemente-Casares, X. et al. Expanding antigen-specific regulatory networks to treat autoimmunity. Nature 530, 434–440 (2016).
Curry, M. P. et al. Expansion of peripheral blood CD5+ B cells is associated with mild disease in chronic hepatitis C virus infection. J. Hepatol. 32, 121–125 (2000).
Curry, M. P. et al. Expansion of innate CD5pos B cells expressing high levels of CD81 in hepatitis C virus infected liver. J. Hepatol. 38, 642–650 (2003).
Monteverde, A., Ballarè, M. & Pileri, S. Hepatic lymphoid aggregates in chronic hepatitis C and mixed cryoglobulinemia. Springe. Semin. Immunopathol. 19, 99–110 (1997).
Doherty, D. G. Immunity, tolerance and autoimmunity in the liver: a comprehensive review. J. Autoimmun. 66, 60–75 (2016).
Gao, B., Jeong, W. I. & Tian, Z. Liver: an organ with predominant innate immunity. Hepatology 47, 729–736 (2008).
Krueger, P. D. et al. Regulation of NK cell repertoire and function in the liver. Crit. Rev. Immunol. 31, 43–52 (2011).
Taniguchi, H. et al. Presence of hematopoietic stem cells in the adult liver. Nat. Med. 2, 198–203 (1996).
Watanabe, H. et al. c-kit+ stem cells and thymocyte precursors in the livers of adult mice. J. Exp. Med. 184, 687–693 (1996).
Crosbie, O. M. et al. In vitro evidence for the presence of hematopoietic stem cells in the adult human liver. Hepatology 29, 1193–1198 (1999).
Wolber, F. M. et al. Roles of spleen and liver in development of the murine hematopoietic system. Exp. Hematol. 30, 1010–1019 (2002).
Jiang, X. et al. Characterizing the lymphopoietic kinetics and features of hematopoietic progenitors contained in the adult murine liver in vivo. PLoS One 8, e76762 (2013).
Lysakova‐Devine, T. & O’Farrelly, C. Tissue‐specific NK cell populations and their origin. J. Leukoc. Biol. 96, 981–990 (2014).
Golden‐Mason, L. et al. Differential expression of lymphoid and myeloid markers on differentiating hematopoietic stem cells in normal and tumor‐bearing adult human liver. Hepatology 31, 1251–1256 (2000).
Arnold, B. Parenchymal cells in immune and tolerance induction. Immunol. Lett. 89, 225–228 (2003).
Wong, Y. C. et al. Immune outcomes in the liver: Is CD8 T cell fate determined by the environment? J. Hepatol. 63, 1005–1014 (2015).
Burghardt, S. et al. Hepatocytes contribute to immune regulation in the liver by activation of the Notch signaling pathway in T cells. J. Immunol. 191, 5574–5582 (2013).
MacParland, S. A. et al. Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat. Commun. 9, 1–21 (2018).
Guillot, A. & Tacke, F. Liver macrophages: old dogmas and new insights. Hepatol. Commun. 3, 730–743 (2019).
Halpern, K. B. et al. Single-cell spatial reconstruction reveals global division of labour in the mammalian liver. Nature 542, 352–356 (2017).
Gebhardt, R. & Matz-Soja, M. Liver zonation: novel aspects of its regulation and its impact on homeostasis. World J. Gastroenterol. 20, 8491 (2014).
Aizarani, N. et al. A human liver cell atlas reveals heterogeneity and epithelial progenitors. Nature 572, 199–204 (2019).
Stamataki, Z. & Swadling, L. The liver as an immunological barrier redefined by single-cell analysis. Immunology 160, 157–170 (2020).
Ramachandran, P. et al. Resolving the fibrotic niche of human liver cirrhosis at single-cell level. Nature 575, 512–518 (2019).
Zhao, J. et al. Single-cell RNA sequencing reveals the heterogeneity of liver-resident immune cells in human. Cell Discov. 6, 22 (2020).
Dobie, R. et al. Single-cell transcriptomics uncovers zonation of function in the mesenchyme during liver fibrosis. Cell Rep. 29, 1832–1847.e8 (2019).
Hossain, M. & Kubes, P. Innate immune cells orchestrate the repair of sterile injury in the liver and beyond. Eur. J. Immunol. 49, 831–841 (2019).
Takeuchi, O. & Akira, S. Pattern recognition receptors and inflammation. Cell 140, 805–820 (2010).
Kubes, P. & Mehal, W. Z. Sterile inflammation in the liver. Gastroenterology 143, 1158–1172 (2012).
Nakatani, Y. et al. Endotoxin clearance and its relation to hepatic and renal disturbances in rats with liver cirrhosis. Liver 21, 64–70 (2001).
Robinson, M. W., Harmon, C. & O’Farrelly, C. Liver immunology and its role in inflammation and homeostasis. Cell. Mol. Immunol. 13, 267–276 (2016).
Kumar, V. NKT-cell subsets: promoters and protectors in inflammatory liver disease. J. Hepatol. 59, 618–620 (2013).
Deng, Z. B. et al. Immature myeloid cells induced by a high‐fat diet contribute to liver inflammation. Hepatology 50, 1412–1420 (2009).
Arrenberg, P., Maricic, I. & Kumar, V. Sulfatide-mediated activation of type II natural killer T cells prevents hepatic ischemic reperfusion injury in mice. Gastroenterology 140, 646–655 (2011).
Halder, R. C. et al. Type II NKT cell–mediated anergy induction in type I NKT cells prevents inflammatory liver disease. J. Clin. Investig. 117, 2302–2312 (2007).
Ertunc, M. E. & Hotamisligil, G. S. Lipid signaling and lipotoxicity in metaflammation: indications for metabolic disease pathogenesis and treatment. J. Lipid Res. 57, 2099–2114 (2016).
Ilan, Y., Shailubhai, K. & Sanyal, A. Immunotherapy with oral administration of humanized anti-CD3 monoclonal antibody: a novel gut-immune system-based therapy for metaflammation and NASH. Clin. Exp. Immunol. 193, 275–283 (2018).
Ju, C. & Tacke, F. Hepatic macrophages in homeostasis and liver diseases: from pathogenesis to novel therapeutic strategies. Cell. Mol. Immunol. 13, 316–327 (2016).
Lanthier, N. et al. Kupffer cell activation is a causal factor for hepatic insulin resistance. Am. J. Physiol. Gastrointest. Liver Physiol. 298, G107–G116 (2010).
Rivera, C. A. et al. Toll-like receptor-4 signaling and Kupffer cells play pivotal roles in the pathogenesis of non-alcoholic steatohepatitis. J. Hepatol. 47, 571–579 (2007).
Lefkowitch, J. H., Haythe, J. H. & Regent, N. Kupffer cell aggregation and perivenular distribution in steatohepatitis. Mod. Pathol. 15, 699–704 (2002).
Bieghs, V. & Trautwein, C. The innate immune response during liver inflammation and metabolic disease. Trends Immunol. 34, 446–452 (2013).
Bertola, A. et al. Hepatic expression patterns of inflammatory and immune response genes associated with obesity and NASH in morbidly obese patients. PLoS One 5, e13577 (2010).
Miura, K. et al. Hepatic recruitment of macrophages promotes nonalcoholic steatohepatitis through CCR2. Am. J. Physiol. Gastrointest. Liver Physiol. 302, G1310–G1321 (2012).
Galastri, S. et al. Lack of CC chemokine ligand 2 differentially affects inflammation and fibrosis according to the genetic background in a murine model of steatohepatitis. Clin. Sci. 123, 459–471 (2012).
Wang, H. et al. Immunological mechanisms and therapeutic targets of fatty liver diseases. Cell. Mol. Immunol. 18.1, 73–91 (2021).
Tacke, F. & Zimmermann, H. W. Macrophage heterogeneity in liver injury and fibrosis. J. Hepatol. 60, 1090–1096 (2014).
Dixon, L. J. et al. Kupffer cells in the liver. Compr. Physiol. 3, 785–797 (2013).
Fallowfield, J. A. et al. Scar-associated macrophages are a major source of hepatic matrix metalloproteinase-13 and facilitate the resolution of murine hepatic fibrosis. J. Immunol. 178, 5288–5295 (2007).
Seidman, J. S. et al. Niche-specific reprogramming of epigenetic landscapes drives myeloid cell diversity in nonalcoholic steatohepatitis. Immunity 52, 1057–1074.e7 (2020).
Xiong, X. et al. Landscape of intercellular crosstalk in healthy and NASH liver revealed by single-cell secretome gene analysis. Mol. Cell 75, 644–660.e5 (2019).
Wen, Y. et al. Hepatic macrophages in liver homeostasis and diseases-diversity, plasticity and therapeutic opportunities. Cell. Mol. Immunol. 18, 45–56 (2020).
Van Hul, N. et al. Kupffer cells influence parenchymal invasion and phenotypic orientation, but not the proliferation, of liver progenitor cells in a murine model of liver injury. Am. J. Pathol. 179, 1839–1850 (2011).
Huang, X. et al. Expression changes of activin A in the development of hepatic fibrosis. World J. Gastroenterol. 7, 37 (2001).
Gressner, O. A. et al. Intracrine signalling of activin A in hepatocytes upregulates connective tissue growth factor (CTGF/CCN2) expression. Liver Int. 28, 1207–1216 (2008).
Kiagiadaki, F. et al. Activin-A causes Hepatic stellate cell activation via the induction of TNFα and TGFβ in Kupffer cells. Biochim Biophys. Acta Mol. Basis Dis. 1864, 891–899 (2018).
Inzaugarat, M. E. et al. Altered phenotype and functionality of circulating immune cells characterize adult patients with nonalcoholic steatohepatitis. J. Clin. Immunol. 31, 1120–1130 (2011).
Gadd, V. L. et al. The portal inflammatory infiltrate and ductular reaction in human nonalcoholic fatty liver disease. Hepatology 59, 1393–1405 (2014).
Stockinger, B., Veldhoen, M. & Martin, B. Th17 T cells: linking innate and adaptive immunity. Semin Immunol. 19, 353–361 (2007).
Winer, D. A. et al. B Lymphocytes in obesity-related adipose tissue inflammation and insulin resistance. Cell. Mol. Life Sci. 71, 1033–1043 (2014).
Lee-Montiel, F. T. et al. Control of oxygen tension recapitulates zone-specific functions in human liver microphysiology systems. Exp. Biol. Med. 242, 1617–1632 (2017).
Wilson, G. K., Tennant, D. A. & McKeating, J. A. Hypoxia inducible factors in liver disease and hepatocellular carcinoma: current understanding and future directions. J. Hepatol. 61, 1397–1406 (2014).
Katz, N. & Jungermann, K. Autoregulatory shift from fructolysis to lactate gluconeogenesis in rat hepatocyte suspensions. The problem of metabolic zonation of liver parenchyma. Biol. Chem. 357, 359 (1976).
Kietzmann, T. Metabolic zonation of the liver: the oxygen gradient revisited. Redox Biol. 11, 622–630 (2017).
Bhandari, T. et al. HIF-1α influences myeloid cell antigen presentation and response to subcutaneous OVA vaccination. J. Mol. Med. 91, 1199–1205 (2013).
Acosta-Iborra, B. et al. Macrophage oxygen sensing modulates antigen presentation and phagocytic functions involving IFN-γ production through the HIF-1α transcription factor. J. Immunol. 182, 3155–3164 (2009).
Clambey, E. T. et al. Hypoxia-inducible factor-1 alpha–dependent induction of FoxP3 drives regulatory T-cell abundance and function during inflammatory hypoxia of the mucosa. Proc. Natl Acad. Sci. USA 109, E2784–E2793 (2012).
Ben‐Shoshan, J. et al. Hypoxia controls CD4+ CD25+ regulatory T‐cell homeostasis via hypoxia‐inducible factor‐1α. Eur. J. Immunol. 38, 2412–2418 (2008).
Abu-Amara, M. et al. Liver ischemia/reperfusion injury: processes in inflammatory networks—a review. Liver Transpl. 16, 1016–1032 (2010).
Kuboki, S. et al. Distinct contributions of CD4+ T cell subsets in hepatic ischemia/reperfusion injury. Am. J. Physiol. Gastrointest. Liver Physiol. 296, G1054–G1059 (2009).
Taniai, H. et al. Susceptibility of murine periportal hepatocytes to hypoxia-reoxygenation: role for NO and Kupffer cell-derived oxidants. Hepatology 39, 1544–1552 (2004).
Hanschen, M. et al. Reciprocal activation between CD4+ T cells and Kupffer cells during hepatic ischemia-reperfusion. Transplantation 86, 710–718 (2008).
Quesnelle, K. M., Bystrom, P. V. & Toledo-Pereyra, L. H. Molecular responses to ischemia and reperfusion in the liver. Arch. Toxicol. 89, 651–657 (2015).
Woolbright, B. L. & Jaeschke, H. The impact of sterile inflammation in acute liver injury. J. Clin. Transl. Res. 3(Suppl 1), 170–188 (2017).
Jaeschke, H., Smith, C. V. & Mitchell, J. R. Hypoxic damage generates reactive oxygen species in isolated perfused rat liver. Biochem Biophys. Res Commun. 150, 568–574 (1988).
Jaeschke, H. et al. Superoxide generation by Kupffer cells and priming of neutrophils during reperfusion after hepatic ischemia. Free Radic. Res Commun. 15, 277–284 (1991).
Jaeschke, H. et al. Complement activates Kupffer cells and neutrophils during reperfusion after hepatic ischemia. Am. J. Physiol. 264, G801–G809 (1993).
Tsung, A. et al. The nuclear factor HMGB1 mediates hepatic injury after murine liver ischemia-reperfusion. J. Exp. Med. 201, 1135–1143 (2005).
Hasegawa, T. et al. Reduced inflammatory response and increased microcirculatory disturbances during hepatic ischemia-reperfusion injury in steatotic livers of ob/ob mice. Am. J. Physiol. Gastrointest. Liver Physiol. 292, G1385–G1395 (2007).
Jang, J. H. et al. Ischemic preconditioning and intermittent clamping confer protection against ischemic injury in the cirrhotic mouse liver. Liver Transpl. 14, 980–988 (2008).
Macshut, M. et al. Older donor age is a risk factor for negative outcomes after adult living donor liver transplantation using small‐for‐size grafts. Liver Transplant. 25, 1524–1532 (2019).
Tanemura, A. et al. Donor age affects liver regeneration during early period in the graft liver and late period in the remnant liver after living donor liver transplantation. World J. Surg. 36, 1102–1111 (2012).
Okaya, T. et al. Age-dependent responses to hepatic ischemia/reperfusion injury. Shock 24, 421–427 (2005).
Wang, J. et al. Visualizing the function and fate of neutrophils in sterile injury and repair. Science 358, 111–116 (2017).
Saijou, E. et al. Neutrophils alleviate fibrosis in the CCl4‐induced mouse chronic liver injury model. Hepatol. Commun. 2, 703–717 (2018).
Lu, L. et al. Innate immune regulations and liver ischemia-reperfusion injury. Transplantation 100, 2601–2610 (2016).
Elnaggar, A. S. & Guarrera, J. V. The marginal liver donor and organ preservation strategies, in Liver Anesthesiology and Critical Care Medicine. (ed Wagener, G.) 207–220 (Springer, 2018).
Ilan, Y. Review article: novel methods for the treatment of non-alcoholic steatohepatitis—targeting the gut immune system to decrease the systemic inflammatory response without immune suppression. Alimentary Pharmacol. Therapeutics 44, 1168–1182 (2016).
Ilan, Y. et al. Induction of regulatory T cells decreases adipose inflammation and alleviates insulin resistance in ob/ob mice. Proc. Natl Acad. Sci. USA 107, 9765–9770 (2010).
Mizrahi, M. et al. Oral administration of anti-CD3 MAB to patients with NASH is safe, promotes regulatory T cells, decreases liver enzymes, and alleviates insulin resistance: results of a phase IIA blinded placebo-controlled trial. Hepatology 54, 117 (2011).
Verdam, F. J. et al. Novel evidence for chronic exposure to endotoxin in human nonalcoholic steatohepatitis. J. Clin. Gastroenterol. 45, 149–152 (2011).
Adar, T. et al. Oral administration of immunoglobulin G-enhanced colostrum alleviates insulin resistance and liver injury and is associated with alterations in natural killer T cells. Clin. Exp. Immunol. 167, 252–260 (2012).
Baeck, C. et al. Pharmacological inhibition of the chemokine C-C motif chemokine ligand 2 (monocyte chemoattractant protein 1) accelerates liver fibrosis regression by suppressing Ly-6C(+) macrophage infiltration in mice. Hepatology 59, 1060–1072 (2014).
Ehling, J. et al. CCL2-dependent infiltrating macrophages promote angiogenesis in progressive liver fibrosis. Gut 63, 1960–1971 (2014).
Laing, R. W. et al. Viability testing and transplantation of marginal livers (VITTAL) using normothermic machine perfusion: study protocol for an open-label, non-randomised, prospective, single-arm trial. BMJ Open 7 (2017).
Mergental, H. et al. Transplantation of discarded livers following viability testing with normothermic machine perfusion. Nat. Commun. 11, 1–12 (2020).
Dutkowski, P. et al. Evolving trends in machine perfusion for liver transplantation. Gastroenterology 156, 1542–1547 (2019).
Dutkowski, P. et al. Challenges to liver transplantation and strategies to improve outcomes. Gastroenterology 148, 307–323 (2015).
Nasralla, D. et al. A randomized trial of normothermic preservation in liver transplantation. Nature 557, 50–56 (2018).
Schlegel, A. et al. Outcomes of DCD liver transplantation using organs treated by hypothermic oxygenated perfusion before implantation. J. Hepatol. 70, 50–57 (2019).
Murphy, M. P. How mitochondria produce reactive oxygen species. Biochem J. 417, 1–13 (2009).
Schlegel, A. et al. Protective mechanisms of end-ischemic cold machine perfusion in DCD liver grafts. J. Hepatol. 58, 278–286 (2013).
Iskender, I. et al. Cytokine filtration modulates pulmonary metabolism and edema formation during ex vivo lung perfusion. J. Heart Lung Transplant. 37, 283–291 (2018).
Mills, E. L. et al. Itaconate is an anti-inflammatory metabolite that activates Nrf2 via alkylation of KEAP1. Nature 556, 113–117 (2018).
Ogger, P. P. et al. Itaconate controls the severity of pulmonary fibrosis. Sci. Immunol. 5, eabc1884 (2020).
O’Connell, J. et al. The relationship of omental and subcutaneous adipocyte size to metabolic disease in severe obesity. PloS One 5, e9997 (2010).
Markose, D. et al. Immune cell regulation of liver regeneration and repair. J. Immunol. Regenerative Med. 2, 1–10 (2018).
Jungermann, K. & Kietzmann, T. Zonation of parenchymal and nonparenchymal metabolism in liver. Annu Rev. Nutr. 16, 179–203 (1996).
Fu, P. & W. Li. Chapter 8—nitric oxide in liver ischemia–reperfusion injury, in Liver Pathophysiology (ed. Muriel, P.) 125–127 (Academic Press: Boston, 2017).
Zhang, Q. et al. Landscape and dynamics of single immune cells in hepatocellular carcinoma. Cell 179, 829–845.e20 (2019).
Funding
This research was supported by an SFI Frontiers Grant.
Author information
Authors and Affiliations
Contributions
O.A. drafted the manuscript, designed the figures, and contributed to the main conceptual ideas. M. W. R. contributed to the writing, provided critical feedback, and helped shape the manuscript. C.O.F. designed and directed the review, supervised the work, and contributed to the writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
About this article
Cite this article
Ahmed, O., Robinson, M.W. & O’Farrelly, C. Inflammatory processes in the liver: divergent roles in homeostasis and pathology. Cell Mol Immunol 18, 1375–1386 (2021). https://doi.org/10.1038/s41423-021-00639-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41423-021-00639-2
Keywords
This article is cited by
-
Ghost messages: cell death signals spread
Cell Communication and Signaling (2023)
-
Modeling drug-induced liver injury and screening for anti-hepatofibrotic compounds using human PSC-derived organoids
Cell Regeneration (2023)
-
The immunological function of CXCR2 in the liver during sepsis
Journal of Inflammation (2022)
-
Anti-neoplastic sulfonamides alter the metabolic homeostasis and disrupt the suppressor activity of regulatory T cells
Scientific Reports (2022)
-
Run to survive—how preoperative exercise could prevent major surgical complications
Nature Metabolism (2021)