Abstract
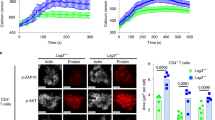
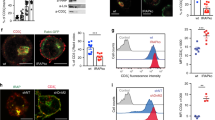
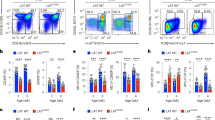
Upon their interaction with cognate antigen, T cells integrate different extracellular and intracellular signals involving basal and induced protein–protein interactions, as well as the binding of proteins to lipids, which can lead to either cell activation or inhibition. Here, we show that the selective T cell expression of CMIP, a new adapter protein, by targeted transgenesis drives T cells toward a naïve phenotype. We found that CMIP inhibits activation of the Src kinases Fyn and Lck after CD3/CD28 costimulation and the subsequent localization of Fyn and Lck to LRs. Video microscopy analysis showed that CMIP blocks the recruitment of LAT and the lipid raft marker cholera toxin B at the site of TCR engagement. Proteomic analysis identified several protein clusters differentially modulated by CMIP and, notably, Cofilin-1, which is inactivated in CMIP-expressing T cells. Moreover, transgenic T cells exhibited the downregulation of GM3 synthase, a key enzyme involved in the biosynthesis of gangliosides. These results suggest that CMIP negatively impacts proximal signaling and cytoskeletal rearrangement and defines a new mechanism for the negative regulation of T cells that could be a therapeutic target.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout









Similar content being viewed by others
References
Brubaker, S. W., Bonham, K. S., Zanoni, I. & Kagan, J. C. Innate immune pattern recognition: a cell biological perspective. Annu. Rev. Immunol. 33, 257–290 (2015).
Chakraborty, A. K. & Weiss, A. Insights into the initiation of TCR signaling. Nat. Immunol. 15, 798–807 (2014).
Viola, A. & Gupta, N. Tether and trap: regulation of membrane-raft dynamics by actin-binding proteins. Nat. Rev. Immunol. 7, 889–896 (2007).
Brownlie, R. J. & Zamoyska, R. T cell receptor signalling networks: branched, diversified and bounded. Nat. Rev. Immunol. 13, 257–269 (2013).
Sahali, D. et al. A novel approach to investigation of the pathogenesis of active minimal- change nephrotic syndrome using subtracted cDNA library screening. J. Am. Soc. Nephrol. 13, 1238–1247 (2002).
Boumediene, A. et al. NEPHRUTIX: a randomized, double-blind, placebo vs Rituximab-controlled trial assessing T-cell subset changes in Minimal Change Nephrotic Syndrome. J. Autoimmun. 88, 91–102 (2018).
Sendeyo, K. et al. Upregulation of c-mip is closely related to podocyte dysfunction in membranous nephropathy. Kidney Int. 83, 414–425 (2013).
Bouachi, K. et al. Expression of CMIP in podocytes is restricted to specific classes of lupus nephritis. PLoS ONE 13, e0207066 (2018).
Audard, V. et al. Occurrence of minimal change nephrotic syndrome in classical Hodgkin lymphoma is closely related to the induction of c-mip in Hodgkin-Reed Sternberg cells and podocytes. Blood 115, 3756–3762 (2010).
Bouatou, Y. et al. Nephrotic syndrome in small cell lung cancer and induction of C-Mip in podocytes. Am. J. Kidney Dis. 2017;69:477–480.
Kamal, M. et al. C-mip interacts physically with RelA and inhibits nuclear factor kappa B activity. Mol. Immunol. 46, 991–998 (2009).
Kamal, M. et al. C-mip interacts with the p85 subunit of PI3 kinase and exerts a dual effect on ERK signaling via the recruitment of Dip1 and DAP kinase. FEBS Lett. 584, 500–506 (2010).
Grimbert, P. et al. The Filamin-A is a partner of Tc-mip, a new adapter protein involved in c-maf-dependent Th2 signaling pathway. Mol. Immunol. 40, 1257–1261 (2004).
Izzedine, H. et al. Expression patterns of RelA and c-mip are associated with different glomerular diseases following anti-VEGF therapy. Kidney Int. 85, 457–470 (2014).
Ory, V. et al. c-mip down-regulates NF-kappaB activity and promotes apoptosis in podocytes. Am. J. Pathol. 180, 2284–2292 (2012).
Izzedine, H. et al. Kidney diseases associated with anti-vascular endothelial growth factor (VEGF): an 8-year observational study at a single center. Medicine 93, 333–339 (2014).
Moktefi, A. et al. Repression of CMIP transcription by WT1 is relevant to podocyte health. Kidney Int. 90, 1298–1311 (2016).
Nakao, A. et al. Blockade of transforming growth factor beta/Smad signaling in T cells by overexpression of Smad7 enhances antigen-induced airway inflammation and airway reactivity. J. Exp. Med. 192, 151–158 (2000).
Ballarin-Gonzalez, B. et al. Protection and systemic translocation of siRNA following oral administration of Chitosan/siRNA nanoparticles. Mol. Ther. Nucleic Acids 2, e76 (2013).
Gao, S. et al. The effect of chemical modification and nanoparticle formulation on stability and biodistribution of siRNA in mice. Mol. Ther. 17, 1225–1233 (2009).
Malissen, B. & Bongrand, P. Early T cell activation: integrating biochemical, structural, and biophysical cues. Annu Rev. Immunol. 33, 539–561 (2015).
Zeng, P., Xu, Y., Zeng, C., Ren, H. & Peng, M. Chitosan-modified poly(D,L-lactide-co-glycolide) nanospheres for plasmid DNA delivery and HBV gene-silencing. Int J. Pharm. 415, 259–266 (2011).
Eibert, S. M. et al. Cofilin peptide homologs interfere with immunological synapse formation and T cell activation. Proc. Natl Acad. Sci. USA 101, 1957–1962 (2004).
Yu, L. et al. cMaf inducing protein inhibits cofilin1 activity and alters podocyte cytoskeleton organization. Mol. Med. Rep. 16, 4955–4963 (2017).
Saraiva, M. & O’Garra, A. The regulation of IL-10 production by immune cells. Nat. Rev. Immunol. 10, 170–181 (2010).
Xie, J. et al. Phosphotyrosine-dependent interaction between the kinases PKCtheta and Zap70 promotes proximal TCR signaling. Sci. Signal. 2019;12:eaar3349.
Zanin-Zhorov, A. et al. Protein kinase C-theta mediates negative feedback on regulatory T cell function. Science 328, 372–376 (2010).
Ghosh, P., Tan, T. H., Rice, N. R., Sica, A. & Young, H. A. The interleukin 2 CD28-responsive complex contains at least three members of the NF kappa B family: c-Rel, p50, and p65. Proc. Natl Acad. Sci. USA 90, 1696–1700 (1993).
Bomsztyk, K. et al. Evidence that interleukin-1 and phorbol esters activate NF-kappa B by different pathways: role of protein kinase C. Cell Regul. 2, 329–335 (1991).
Zamoyska, R. et al. The influence of the src-family kinases, Lck and Fyn, on T cell differentiation, survival and activation. Immunol. Rev. 191, 107–118 (2003).
Tuosto, L. et al. Organization of plasma membrane functional rafts upon T cell activation. Eur. J. Immunol. 31, 345–349 (2001).
Garofalo, T. et al. Association of GM3 with Zap-70 induced by T cell activation in plasma membrane microdomains: GM3 as a marker of microdomains in human lymphocytes. J. Biol. Chem. 277, 11233–11238 (2002).
Nagafuku, M. et al. CD4 and CD8 T cells require different membrane gangliosides for activation. Proc. Natl Acad. Sci. USA 109, E336–E342 (2012).
Dykstra, M., Cherukuri, A., Sohn, H. W., Tzeng, S. J. & Pierce, S. K. Location is everything: lipid rafts and immune cell signaling. Annu. Rev. Immunol. 21, 457–481 (2003).
Davis, D. M. & Dustin, M. L. What is the importance of the immunological synapse? Trends Immunol. 25, 323–327 (2004).
Zumerle, S., Molon, B. & Viola, A. Membrane rafts in T cell activation: a spotlight on CD28 costimulation. Front. Immunol. 8, 1467 (2017).
Ballek, O. et al. TCR triggering induces the formation of Lck-RACK1-Actinin-1 multiprotein network affecting Lck redistribution. Front. Immunol. 7, 449 (2016).
Zhang, S. Y. et al. c-mip impairs podocyte proximal signaling and induces heavy proteinuria. Sci. Signal. 3, ra39 (2010).
Varshney, P., Yadav, V. & Saini, N. Lipid rafts in immune signalling: current progress and future perspective. Immunology 149, 13–24 (2016).
Sahali, D. et al. Immunopathogenesis of idiopathic nephrotic syndrome with relapse. Semin. Immunopathol. 36, 421–429 (2014).
Kassu, A. et al. Regulation of virus-specific CD4+ T cell function by multiple costimulatory receptors during chronic HIV infection. J. Immunol. 185, 3007–3018 (2010).
Hodi, F. S. et al. Biologic activity of cytotoxic T lymphocyte-associated antigen 4 antibody blockade in previously vaccinated metastatic melanoma and ovarian carcinoma patients. Proc. Natl Acad. Sci. USA 100, 4712–4717 (2003).
Velu, V. et al. Enhancing SIV-specific immunity in vivo by PD-1 blockade. Nature 458, 206–210 (2009).
Macdonald, J. L. & Pike, L. J. A simplified method for the preparation of detergent-free lipid rafts. J. Lipid Res. 46, 1061–1067 (2005).
Bourderioux, M. et al. A new workflow for proteomic analysis of urinary exosomes and assessment in cystinuria patients. J. Proteome Res. 14, 567–577 (2015).
Xia, J., Mandal, R., Sinelnikov, I. V., Broadhurst, D. & Wishart, D. S. MetaboAnalyst 2.0–a comprehensive server for metabolomic data analysis. Nucleic Acids Res. 40(Web Server issue), W127–W133 (2012).
Li, F. et al. MPINet: metabolite pathway identification via coupling of global metabolite network structure and metabolomic profile. BioMed. Res. Int. 2014;2014:325697.
Cavill, R. et al. Consensus-phenotype integration of transcriptomic and metabolomic data implies a role for metabolism in the chemosensitivity of tumour cells. PLoS Comput. Biol. 7, e1001113 (2011).
Cox, J. & Mann, M. MaxQuant enables high peptide identification rates, individualized p.p.b.-range mass accuracies and proteome-wide protein quantification. Nat. Biotechnol. 26, 1367–1372 (2008).
Acknowledgements
We are grateful to Prof. Rose Zamoyska (Medical Research Council, London, United Kingdom) for providing us with the Tg(CD2-rtTA) CRza mouse model. This work was supported in part by a grant from the French Kidney Foundation. J.O., P.V., and K.S. were supported by grants from the Ministry of Research.
Author information
Authors and Affiliations
Contributions
J.O., K.S., C.C., P.V., B.S., E.C., C.H., V.F., A.P., G.A. and I.C.G. performed the experiments. M.O., V.A. and D.S. wrote the paper. D.S. and M.O. supervised the project. All authors discussed the results and participated in writing the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
About this article
Cite this article
Oniszczuk, J., Sendeyo, K., Chhuon, C. et al. CMIP is a negative regulator of T cell signaling. Cell Mol Immunol 17, 1026–1041 (2020). https://doi.org/10.1038/s41423-019-0266-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41423-019-0266-5
Keywords
This article is cited by
-
Integrative GWAS and co-localisation analysis suggests novel genes associated with age-related multimorbidity
Scientific Data (2023)
-
Immunopathogenesis of idiopathic nephrotic syndrome
Cellular & Molecular Immunology (2022)
-
FOXC1-induced LINC01123 acts as a mediator in triple negative breast cancer
Cancer Cell International (2020)