Abstract
Breast cancer is a heterogeneous disease. In particular, triple-negative breast cancer (TNBC) comprises various molecular subgroups with unclear identities and currently has few targeted treatment options. Our previous study identified protein C receptor (Procr) as a surface marker on mammary stem cells (MaSCs) located in the basal layer of the normal mammary gland. Given the possible connection of TNBC with basal layer stem cells, we conducted comparative analyses of Procr in breast cancers of mouse and human origin. In mouse mammary tumors, we showed that Procr+ cells are enriched for cancer stem cells (CSCs) in Wnt1 basal-like tumors, but not in Brca1 basal-like tumors or PyVT luminal tumors. In human cancers, PROCR was robustly expressed in half of TNBC cases. Experiments with patient-derived xenografts (PDXs) revealed that PROCR marks CSCs in this discrete subgroup (referred to as PROCR+ TNBC). Interfering with the function of PROCR using an inhibitory nanobody reduced the CSC numbers, arrested tumor growth and prevented rapid tumor recurrence. Our data suggest a key role of MaSC in breast tumorigenesis. Moreover, our work indicates that PROCR can be used as a biomarker to stratify TNBC into clinically relevant subgroups and may provide a novel targeted treatment strategy for this clinically important tumor subtype.
Similar content being viewed by others
Introduction
Breast cancer (BC) comprises many biologically different entities with distinct clinical outcomes. To classify BC, immunohistochemistry (IHC) markers such as estrogen receptor (ER), progesterone receptor (PR) and human epidermal growth factor receptor 2 (HER2), together with clinicopathological features, are conventionally used for patient prognosis and management. IHC markers separate patients into four subgroups, Luminal A (ER+, PR+), Luminal B (ER+, PR+, HER2/Ki67+), HER2 (HER2+) and Triple-negative breast cancer (TNBC, i.e. lack of ER and PR expression, and absence of HER2 amplification or overexpression).1 TNBCs account for 15–20% of newly diagnosed breast cancer cases, and are generally associated with a high risk of disease recurrence and poor patient survival.1,2 TNBC is a difficult and complex disease and its clinical outcome has been unsatisfactory, even when diagnosed at an early stage of the disease.1,3,4 TNBCs are heterogeneous in terms of biology, prognosis and response to treatment.5,6,7 The absence of well-defined molecular targets is a major challenge in treating patients with TNBC. Thus, stratifying TNBCs into well-defined molecular subgroups and identifying molecular drivers on which to base targeted therapy is of utmost importance.
Cancer stem cells (CSCs) play prominent roles in tumor growth, treatment resistance, recurrence, and metastasis of cancers.8,9,10 In breast cancer, a number of markers have been proven useful for the isolation of subsets of cells enriched for CSCs, including CD44, CD24 and ALDH1 activity.11,12 However, they are not universal CSC markers across all subtypes of breast cancer, as shown by the fact that CD44hi, CD24lo and ALDHhi, CSC-enriched subsets bear little overlap within the same breast tumor.12 In addition, CSCs were found in both CD44hi CD24− and CD44hi CD24+ fractions of ER-negative BCs and TNBCs.13 Variability among different breast cancer subtypes is likely extensive, and it may thus be challenging to designate a single CSC population for all breast cancers. CSCs of various subtypes could be distinct in molecular features.8 In certain subtypes, CSCs may share similar features and regulatory mechanisms with normal MaSCs. In this work, we investigated whether MaSCs are associated with a particular subtype of TNBC. Our previous study identified Procr, a Wnt target, as a surface marker for MaSCs (Lin−, CD24+, CD29hi, Procr+) in the normal mouse mammary gland.14 In this work, we investigated the functional role of Procr in mammary tumor models and human breast cancers, and reveal the biological significance of Procr expression. Finally, we explored its potential utility in the diagnosis and therapy of TNBC.
Results
Procr marks CSCs in MMTV-Wnt1 mammary basal-like tumor
To investigate the role of Procr in tumors, we utilized three distinct mammary tumor models: MMTV-Wnt1 tumors are preferentially induced from stem/progenitor cells15 and share transcriptional patterns with human basal-like breast cancer16,17; MMTV-PyVT tumors are closely clustered to the luminal B subtype;17,18 and K14-Cre;Brca1f/+;p53f/+ tumors are associated with human BRCA1 basal-like tumors.17,19 In all three tumor models, Procr+ cells are distributed in a small portion (1–8%) of basal cells and some stromal cells (Fig. 1a–c). To investigate whether Procr+ cells are enriched for CSCs in these tumors, Procr+ cells (Lin−, CD24+, CD29hi, Procr+) and Procr− cells (Lin−, CD24+, CD29hi, Procr−) were isolated from the tumors and xenografted to fat pads of recipients in limiting dilution. After grafted into FVB recipients, Procr+ cells from MMTV-Wnt1/FVB tumors reconstituted tumors robustly, whereas in sharp contrast, Procr− cells failed to form tumors (Fig. 1d). These results suggest that Procr+ cells are enriched for CSCs of MMTV-Wnt1 tumors. Of note, a previous study suggested that transplantation of a mixture of MMTV-Wnt1 basal and luminal cells, but not MMTV-Wnt1 basal cells alone (in a mixed genetic background), can reconstitute tumors in transplantation assays.20 However, our data suggest that MMTV-Wnt1 basal cells alone in the FVB background, in particular Procr+ basal cells, can efficiently generate tumors upon transplantation. The observed discrepancies could be explained by differences in genetic background that have been reported to affect the dynamics of MMTV-Wnt1 tumor formation.21 Similar experiments were carried out using MMTV-PyVT tumors and K14-Cre;Brca1f/+;p53f/+ tumors. Procr+ and Procr− cells were isolated and xenografted to Nude recipients. Interestingly, they displayed no discernable tumor-initiating capacity (Fig. 1e, f), suggesting that Procr expression does not preferentially define CSCs in these tumor subtypes. Collectively, xenograft experiments suggest that Procr+ cells are enriched for CSCs in mouse MMTV-Wnt1 mammary tumors.
Procr marks CSCs of a particular basal-like subtype. a–c FACS analyses showing the distribution of Procr+ cells related to basal, luminal and stromal cell compartments in WT mammary gland, MMTV-Wnt1, MMTV-PyVT and K14-Cre;Brca1f/+;p53f/+ tumors. One of three similar experiments is shown in a. Quantification indicating that Procr+ basal cells consisted of 2.5 ± 0.9% of total basal cells in normal tissue control (WT). There is a significant increase of Procr+ basal cells in MMTV-Wnt1 tumor (7.6 ± 0.5%), and a decrease of Procr+ basal cells in MMTV-PyVT (1.1 ± 0.1%) and K14-Cre;Brca1f/+;p53f/+ (1.1 ± 0.2%) tumors (b). Procr+ cells are absent from luminal compartment in WT and all three tumor models, and the percentages of Procr+ cells in stromal compartment have no significant changes in tumors compared to the WT (c). Data are pooled from three independent experiments and presented as mean ± SEM in b, c. ***P < 0.001, *P < 0.05, ns P > 0.05. d Procr+ basal cells (Lin−, CD24+, CD29hi, Procr+) and Procr− basal cells (Lin−, CD24+, CD29hi, Procr−) isolated from MMTV-Wnt1/FVB tumor were engrafted in limiting dilution to FVB recipient fat pads. Procr+ basal cells formed tumor vigorously with CSC frequency of 1/45, while Procr− basal cells could not. ***P < 0.001. e, f Procr+ cells (Lin−, CD24+, CD29hi, Procr+) and Procr− cells (Lin−, CD24+, CD29hi, Procr−) isolated from MMTV-PyVT tumor (e) or K14-Cre;Brca1f/+;p53f/+ tumor (f) were engrafted in limiting dilution to Nude recipient fat pads. There is no significant difference in tumor formation efficiency between the two populations. ns P > 0.05
Procr is critical for MMTV-Wnt1 mammary tumor growth
To address whether Procr is required for MMTV-Wnt1 tumor formation, we knocked down Procr expression and examined its impact in tumor xenograft experiments. Single cells were dissociated from MMTV-Wnt1 primary tumors and virally infected by sh-Procr with a GFP tag. The infected cells were sorted using GFP and xenografted to fat pads of immunocompromised recipients. We found that inhibition of Procr drastically attenuates tumor formation of the engrafted cells, while cells infected by control scramble shRNA form tumors potently (Supplementary information, Fig. S1a–d). Similar results were observed when MMTV-Wnt1 tumor cells were grafted to syngeneic FVB mammary fat pads (Supplementary information, Fig. S1e–g). Thus, inhibition of Procr diminishes the tumor formation capacity of CSCs in MMTV-Wnt1 tumors, suggesting that Procr is important for formation of certain basal-like mammary tumors.
PROCR is highly expressed in half of TNBC cases that are associated with poorer clinical outcome compared to PROCR-low TNBC patients
We first examined PROCR expression in clinical non-cancerous human breast tissues. Immunohistochemical staining (IHC) showed that, in most areas, PROCR is expressed in individual basal cells sparsely distributed in the ductal epithelium (Fig. 2a). In a few areas, PROCR staining could also be detected in groups of basal cells (Supplementary information, Fig. S2a), an observation similar to a previous report.22 FACS quantification indicated that PROCR is expressed in 3% of basal cells and 3.5% of mesenchymal cells, while absent in the luminal populations (Fig. 2b, Supplementary information, Fig. S2b). Thus, the expression of PROCR in non-cancerous human tissues is reminiscent of its pattern in the normal mouse mammary gland.14
PROCR is expressed in a subset of basal cells in normal breast tissues. a Representative images showing immunostaining of human non-cancerous mammary tissue with antibodies against PROCR and Keratin 14 (Krt14). Scale bars represent 20 μm in lower magnification image, 5 μm in the enlarged image. b FACS analysis of non-cancerous mammary tissue showing that PROCR is expressed in 3% of basal cells, 3.5% of mesenchymal cells and is devoid from mature luminal cells and luminal progenitors. n = 4 patients. Data are presented as mean ± SEM
Next, we performed PROCR staining in a cohort of 80 breast tumors (20 whole-section specimens for each subtype, including Luminal A, Luminal B, HER2+ and TNBC). We found that PROCR expression is markedly more robust in TNBC compared to Luminal and HER2+ subtypes (Supplementary information, Fig. S2c, d). Within TNBC, 13 out of 20 (65%) exhibited an H-score higher than 80 (Supplementary information, Fig. S2d). To increase our sample size, we used tissue microarrays (TMAs) to examine PROCR expression in a larger cohort comprising 443 breast tumors and 71 non-cancerous mammary controls. Consistently, TNBCs exhibited a markedly high prevalence of PROCR-high cases (52.35% PROCR-high (score = 2, 3); 47.64% PROCR-low (score = 0, 1), n = 149). The frequency of PROCR-high cases was drastically lower (ranging from 2–7%) in non-cancerous tissues and other subtypes of carcinomas (Fig. 3a, b). We further investigated the relationship between PROCR expression levels and various clinical characteristics. PROCR expression was inversely associated with ER status (P < 0.001), PR status (P < 0.001) and HER2 status (P < 0.001) in breast cancers, and positively associated with Ki67 (P = 0.020) and Vimentin (Vim) staining (P < 0.001), an indication of epithelial-to-mesenchymal transition (EMT), but there was no other correlation between PROCR levels and other clinicopathological features (Supplementary information, Table S1).
PROCR is highly expressed in a subgroup of TNBC. a PROCR expression was measured by IHC in tissue microarray containing 71 no-cancerous, 99 Luminal A cancers, 105 Luminal B cancers, 90 HER2 cancers and 149 TNBCs. Representative images of negative (score 0), weak (score 1), medium (score 2) and strong (score 3) staining are shown. PROCR-low cases include score 0 and score 1; PROCR-high cases include score 2 and score 3. Scale bars represent 200 μm in lower magnification images, 50 μm in the enlarged images. b Statistical analysis of PROCR expression according to the IHC score. Majority of non-cancerous, Luminal A, Luminal B and HER2+ samples have weak (score 1) or negative (score 0) expression of PROCR, whereas 52.4% of TNBCs display PROCR-high (score 2 + score 3). c–e Kaplan-Meier analysis of disease free survival (DFS) in our cohort. PROCR expression is associated with poor DFS in TNBC patients (c, n = 141 patients). PROCR expression has no significant association with clinical outcome of hormone receptor-positive (d, n = 187 patients) or HER2 enrichment subtypes (e, n = 87 patients). f, g PROCR expression was measured by IHC in human TNBC samples containing 28 cases of BRCA1 mutation carriers and 30 cases of BRCA1 wildtype. Representative images of negative (H-score < 30), weak (H-score 30-75), medium (H-score 80-120) and strong (H-score > 120) staining are shown in f. PROCR staining is in brown; hematoxylin counterstain is in blue. Scale bar represents 200 μm. Data are presented as mean ± SEM in g. ***P < 0.001
The clinical significance of PROCR expression in breast cancer was assessed. In TNBC patients, PROCR-high was correlated with poorer disease-free survival (DFS) compared with PROCR-low by Kaplan-Meier analysis (Fig. 3c). In the hormone-receptor positive and HER2+ subtypes, no significant association was found between PROCR levels and disease events (Fig. 3d, e). In accordance with the results of our cohort, further analysis of a large public clinical database of breast cancer (Kaplan-Meier Plotter) also supported that a high level of PROCR expression correlates with a poorer clinical outcome in patients with hormone receptor-negative breast cancer (Supplementary information, Fig. S2e), whereas PROCR expression has no prognostic value in hormone receptor-positive patients (Supplementary information, Fig. S2f). Additionally, elevated PROCR expression indicated a higher likelihood for disease events in univariate analysis and exhibited a similar trend upon multivariate analysis (Supplementary information, Table S2). Together, these results suggest that PROCR-high BCs make up about half of TNBC cases, and that PROCR expression level as detected by IHC may stratify TNBCs into two subgroups with different prognoses.
The expression of PROCR was also investigated in a panel of human breast cancer cell lines in our previous study.23 qPCR analysis showed that all ER+/PR+ lines (T-47D, ZR75-1, MDA-MB-415 and MCF-7) and HER2+ lines (SK-BR-3, MDA-MB-453 and BT474) exhibited relatively low PROCR expression. A subset of TNBC lines (MDA-MB-231, Hs578T, HCC38, CAL51 and HCC1806) highly expressed PROCR, whereas some TNBC lines (MDA-MB-468, BT549, MDA-MB-436, HCC1937, HCC1599 and HCC2157) exhibited low expression of PROCR. These results are in line with our observations in patient tissue samples (52% of TNBC cases are PROCR-high), supporting the idea that PROCR expression stratifies TNBCs.
BRCA1 mutant carrier TNBC is associated with PROCR-low TNBC subgroup
Intriguingly, cell lines that contain BRCA1 (MDA-MB-436, HCC1937, and HCC2157) or BRCA2 (HCC1599) mutations fell into the PROCR-low TNBC subgroup (Supplementary information, Fig. S3a). This is in line with the observation that MMTV-Cre;Brca1f/f;p53+/− mouse mammary tumors have basal-like phenotype,17 yet exhibit low expression of Procr (Fig. 1a, b). Enlightened by results in TNBC cell lines and mouse models, we further investigated whether PROCR expression level is inversely correlated with BRCA1 mutation status using human TNBC tissue samples. PROCR staining was performed in samples from a cohort of 58 TNBC patients (28 BRCA1 mutant carriers; 30 BRCA1 wild type patients) (Supplementary information, Table S3). We found that the majority of BRCA1 mutant carriers have low expression of PROCR (89.3%, n = 28), whereas the majority of BRCA1 wild type TNBCs are PROCR-high (70%, n = 30), exhibiting an H-score higher than 80 (Fig. 3f, g). Together, these cell line and patient tissue data indicate an inverse correlation of PROCR-high TNBCs and BRCA1 mutant TNBCs (P < 0.001; Supplementary information, Table S3), suggesting an intriguing hypothesis that they are discrete subgroups with distinct biology.
Based on genome-wide gene expression studies, it has been reported that these TNBC cell lines can be subdivided into proliferation-related, mesenchymal-related and immune-related subsets.2 However, PROCR status did not appear to correlate with these stratifications (Supplementary information, Fig. S3a). In another database containing transcription profiles of 51 BC cell lines,24 PROCR status is positively correlated with Claudin-low TNBCs (Supplementary information, Fig. S3b).
PROCR enriches CSCs in human PROCR+ BC
Next we investigated whether PROCR-expressing cells are enriched for CSCs in PROCR-high TNBC (referred to as PROCR+ BC hereafter). Almost all MDA-MB-231 cells express PROCR,23 making this cell line unsuitable for investigating the relationship between PROCR and CSCs (see discussion). Thus, we utilized patient-derived xenografts (PDXs) for a better representation of primary tumor tissues. IHC indicated negative expression of ER, PR and HER2, and robust expression of PROCR in three PDX tumors used (PDX-1/2/3) (Fig. 4a, Supplementary information, Fig. S4a, c, d). FACS analysis with EpCAM and CD49f indicated two populations in these PDXs, the majority was double-positive (87%) and the rest was double negative (13%), indicating that EpCAM status equals CD49f status in these PDXs (Supplementary information, Fig. S4b). Thus, only EpCAM was used thereafter. PROCR+ cells are in the EpCAM+ compartment, comprising 42–45% of the total tumor cells (Fig. 4b, Supplementary information, Fig. S4c, d), representing a drastic increase of PROCR+ cells compared to those (3%) in non-cancerous breast tissues. Both PROCR+ and PROCR− tumor cells were proliferative, but PROCR+ tumor cells exhibited about 2-fold more G2/M phase cells (4 N), EdU+ cells and Ki67+ cells (Fig. 4c–e). Of note, PROCR+ cells made up of a subpopulation (64.6%) of previously reported CD24−, CD44+ CSCs (Supplementary information, Fig. S4e).
PROCR+ cells are enriched for CSCs in PROCR+ BCs. a IHC indicating that PDX-1 is ER−, PR−, HER2−, and PROCR-high. Scale bars represent 100 μm. b FACS analysis showing the PROCR+ cells (Lin−, EpCAM+, PROCR+) are composed of 42.1 ± 0.2% tumor cells in PDX-1. Data are pooled from 3 independent experiments and presented as mean ± SEM. c FACS analysis showing that both PROCR+ and PROCR− tumor cells isolated from PDX-1 are proliferative, containing G2/M phase cells (4 N) (left), while PROCR+ cells have 1.9-fold more 4 N cells compared to PROCR− tumor cells (right) (c). Data are pooled from three independent experiments and presented as mean ± SEM. **P < 0.01. d PROCR+ (Lin−, EpCAM+, PROCR+) and PROCR− (Lin−, EpCAM+, PROCR−) tumor cells isolated from PDX-1 were cultured in vitro and underwent 1 h EdU incorporation. Representative images are shown (left). Quantification indicating that PROCR+ cells had 2.6-fold more EdU-labeling cells (right). Scale bars represent 100 μm. Data are pooled from three independent experiments and presented as mean ± SEM. ***P < 0.001. e PROCR+ (Lin−, EpCAM+, PROCR+) and PROCR− (Lin−, EpCAM+, PROCR−) cells isolated from PDX-1 were stained with Ki67. Representative images are shown (left). Quantification indicating that PROCR+ cells had 2.1-fold more Ki67+ cells (right). Scale bars represent 100 μm. Data are pooled from three independent experiments and presented as mean ± SEM. ***P < 0.001. f PROCR+ (Lin−, EpCAM+, PROCR+) and PROCR− (Lin−, EpCAM+, PROCR−) and Lin−, EpCAM− cells isolated from PDX-1 were engrafted in limiting dilution as indicated. PROCR+ cells formed tumor readily, while PROCR− and EpCAM− cells had drastically lower tumorigenicity. The wait time for post-transplantation of 1 PROCR+ cell experiment was 8 weeks, while the wait time for the rest of experiments was 4 weeks. The CSC frequencies were calculated using the data with the same wait time period. ***P < 0.001. g FACS analysis of the tumor outgrowths derived from engrafted PROCR+ cells indicating that they comprise both PROCR+ and PROCR− cells, at similar percentages as the parental PDX tumor. Data are pooled from three independent experiments and presented as mean ± SEM. h Gene Set Enrichment Analysis indicating that PROCR+ tumor cells (Lin−, EpCAM+, PROCR+) have enriched EMT (Sarrio et al., 2008), Myc_targets (Zeller et al. 2003), MaSCs (Lim et al. 2010) and Wnt signaling signatures relative to PROCR− tumor cells (Lin−, EpCAM+, PROCR−). i Heat map of isolated PROCR+ and PROCR− tumor cells (Lin−, EpCAM+) from PDX-2 and PDX-3 tumors. PROCR+ tumor cells have reduced E-cad level (epithelial feature gene) and increased mesenchymal signature genes. j Western blot indicating that PROCR+ tumor cells have reduced E-cad level and increased Slug level. Tubulin serves as a loading control
To examine their tumor-initiating ability, PROCR+ and PROCR− cells (Lin−, EpCAM+) were isolated from PDX tumors and engrafted into immunocompromised recipients in limiting dilution. PROCR+ cells can potently form tumors (1/114 CSC frequency, tumor harvested at 4 weeks post transplantation) (Fig. 4f). We noted that the size of tumor outgrowth was consistent with the number of cells transplanted, i.e. fewer transplanted cells generated smaller tumor. Thus, when engrafting a single cell, to better visualize the tumor, we waited for an extra 4 weeks (in total 8 weeks), and found that 30% (nine out of 30) of transplants formed tumors (Fig. 4f). FACS analysis showed that tumor outgrowths derived from engrafted PROCR+ cells comprised both PROCR+ and PROCR− cells, at similar percentages as the parental PDX tumor (Fig. 4g), suggesting that some PROCR+ cells have differentiated into PROCR− cells. Although PROCR− cells are proliferative, they displayed drastically lower tumor-initiating capacity (1/29,475 CSC frequency) (Fig. 4f), suggesting that they are likely not the driver cells fueling tumor formation. Similarly, EpCAM− cells also exhibited very low tumorigenic capacity (1/19,782 CSC frequency) (Fig. 4f). The few small tumors that formed (2/8 or 1/8) when transplanting high numbers (10,000 or 2,000) of EpCAM+, PROCR− cells or EpCAM− cells resembled the outgrowth derived from EpCAM+, PROCR+ epithelial cells (Supplementary information, Fig. S4f, g), suggesting that the rare tumor formation from those two populations may be due to contaminating EpCAM+, PROCR+ cells.
Gene Set Enrichment Analysis (GSEA) of two PDXs revealed that signatures of EMT, Myc targets, mammary stem cells and Wnt signaling are significantly enriched in PROCR+ cells (Fig. 4h). Heat maps and Western blotting analyses confirmed decreased E-cad and increased Slug expression in PROCR+ cells compared to PROCR− cells (Fig. 4i, j). Collectively, these results establish that PROCR+ cells are enriched for CSCs in PROCR+ BCs (PROCR-high TNBC).
Inhibition of PROCR potently suppresses PROCR+ BC formation
We next investigated the possibility of targeting PROCR to inhibit PROCR+ BC growth. Tumor cells in the three PROCR+ BC PDXs were dissociated and infected with sh-PROCR viruses with GFP tags, infected cells were FACS-isolated and xenografted to fat pads of recipients (Fig. 5a). We found that knockdown of PROCR dramatically reduced tumor formation in all three PDX tumor xenograft experiments compared with the scramble control (Fig. 5b–g). In addition, the effect of PROCR knockdown was further examined using CRISPR interference.25 PDXs tumor cells were infected with dCas9-KRAB and sgRNA (sg-PROCR) (Fig. 5h). PROCR knockdown with sgRNA also drastically inhibited tumor formation in xenograft experiments (Fig. 5i, j). The impact of PROCR attenuation was also investigated in cell lines. In MDA-MB-231 cells, knockdown of PROCR using two independent shRNAs significantly inhibited cell growth (Supplementary information, Fig. S5a, b). In vivo, sh-PROCR drastically delayed tumor formation and liver metastasis of MDA-MB-231 cells and inhibited tumor growth, when compared with the scramble control (Supplementary information, Fig. S5c, d). In contrast, knockdown of PROCR was ineffective in influencing PROCR-low TNBC (BT549) or ER+ luminal tumor cell (MCF-7) growth (Supplementary information, Fig. S5e, f). Together, these data provide a proof of concept that PROCR can be targeted to inhibit the growth of PROCR+ BCs.
Inhibition of PROCR suppresses PROCR+ BC PDXs tumor growth. a–g Illustration of inhibition of PROCR in PDX by shRNA. Dissociated PDX tumor cells were virally infected by scramble control or sh-PROCR, and the infected cells were sorted based on the GFP tag, followed by engraftment into Nude recipients (a). An aliquot of cells was used for Western analysis and confirmed about 75% PROCR knockdown efficiency in PDX-1 (b), PDX-2 (d) and PDX-3 (f). Xenografts of the infected cells indicating that PROCR knockdown blocks PDXs tumor growth (b–g). n = 4 mice or more in each group. Data are presented as mean ± SEM. ***P < 0.001. h–j Illustration of inhibition of PROCR expression using CRISPR interference (h). Western analysis validating the repression of PROCR by KRAB (i). Xenografts of the infected cells indicating that PROCR knockdown significantly inhibits PDX tumor growth (i, j). n = 10 mice in each group. Data are presented as mean ± SEM. ***P < 0.001
PROCR inhibitory nanobody suppresses PROCR+ BC growth
Next, we evaluated the therapeutic benefit of targeting PROCR using a more clinically applicable approach. A single-domain antibody containing camelid VHHs26 (also referred to as a Nanobody) targeting the extracellular domain of PROCR was developed, which blocks the interaction of PROCR with its ligand Protein C (Fig. 6a, b, Supplementary information, Fig. S6a, see Methods for details). In MDA-MB-231 cell culture, addition of the inhibitory antibody significantly repressed cell proliferation as seen by decreased cell numbers in passaging and reduced EdU incorporation (Fig. 6c, d). Our recent work described PROCR signaling in TNBC cells, in that PROCR activates pSrc, subsequently activates IGF-1R and both MEK-ERK and PI3K-Akt-mTOR pathways; concomitantly PROCR activates RhoA-ROCK-p38 pathway via a surface effector F2R (illustrated in Supplementary information, Fig. S6b).23 Here, we investigated the mechanism of action of the PROCR inhibitory nanobody. In culture, the inhibitory effects on pSrc, pIGF-1R and all of the known PROCR-dependent intracellular signaling activities were evident by 12 h following the nanobody treatment, and became more pronounced by 16 h (Fig. 6e). Phosphorylation of EGFR-Y845, a site known to be phosphorylated by Src,27 was also diminished, whereas other phosphorylated sites of EGFR, e.g. Y1068 and Y1173, were not affected (Supplementary information, Fig. S6c). This also reflected the specificity of the nanobody on targeting the PROCR-dependent signaling. Furthermore, considering the EMT characteristics that PROCR+ cells have, we investigated whether the nanobody affects EMT. As expected, the nanobody blocked mesenchymal traits. The treated cells had increased level of E-cad, and decreased levels of Vim, Slug and Zeb1 (Fig. 6f).
Generation of a nanobody that inhibits PROCR-dependent signaling. a Illustration of single-domain antibody, consisting antigen binding camelid VHHs (devoid of light chains) and human IgG. b Elisa indicating that the inhibitory antibody blocks PROCR binding with its ligand PROC, while the control antibody cannot. Data are pooled from three independent experiments and presented as mean ± SEM. ***P < 0.001. c, d MDA-MB-231 cells were cultured in the presence of Ctrl or PROCR inhibitory nanobody (200 μg/ml) for four passages in complete media. Cell numbers counted in each passage indicating that the antibody inhibited proliferation (c). EdU incorporation experiment (1 h) showing that the antibody markedly inhibited cell proliferation (d). Data are pooled from three independent experiments and presented as mean ± SEM. ***P < 0.001. e In MDA-MB-231 cell culture, PROCR-dependent signaling activities were examined at 8 h, 12 h and 16 h after addition of the nanobody. Western analysis indicating that the antibody attenuated Src and IGF-1R phosphorylation, and inhibited ERK, PI3K-Akt-mTOR and RhoA-ROCK pathway activities of MDA-MB-231 cells in vitro, at 12 h and 16 h. Analyses in this panel are from the same batch of cells using the same loadings, thus only one loading control (Tubulin) is shown at the end of the panel. For a better illustration, they are shown as three separated columns representing ERK, Akt and RhoA pathway respectively. f In MDA-MB-231 cell culture, EMT related proteins were examined by Western analysis at 8 h, 12 h and 16 h after addition of the nanobody. Increased E-cad level, decreased Vim, Slug and Zeb1 levels were apparent at 12 h and 16 h
The anti-tumor efficacy of the antibody was further investigated in vivo. Mice bearing PROCR+ BC PDX were treated with the antibody. The treatment was started soon after the engraftment (at day 5), and the nanobody was injected for a total of six times. We observed marked inhibition of tumor growth (six fold inhibition in PDX-3, Fig. 7a). Similar inhibitory effects were observed for the other two PDXs (PDX-1 and 2, Supplementary information, Fig. S6d, e). The remaining tumors were further analyzed. We found that the percentage of PROCR+ cells (43.4%) post IgG treatment remains similar to that of their parental tumor, while the percentage of PROCR+ cells was decreased to 19.5% post the nanobody treatment, supporting the idea that the nanobody directly targets PROCR+ cells, resulting in growth inhibition (Fig. 7b). As expected, in a PROCR− BC PDX (PDX-4, Supplementary information, Fig. S4a, h), the nanobody treatment was ineffective in suppressing tumor growth (Supplementary information, Fig. S6f).
The PROCR inhibitory nanobody suppresses the growth of PROCR+ BCs. a IgG or PROCR inhibitory antibody (PROCR-mAb) was i.p. administered at d5, d7, d9, d12, d15 and d18 (8 mg/kg body weight) (blue arrows) after engraftment of PDX-3 tumor cells. Tumor sizes were suppressed for 6-fold with PROCR inhibitory antibody. n = 6 mice in each group. Data are presented as mean ± SEM. ***P < 0.001. Similar experiments using PDX-1 and PDX-2 are shown in (Supplementary information, Fig. S6d, e). b FACS analysis indicating decreased percentage of PROCR+ cells (Lin−, EpCAM+, PROCR+) in the remaining tumors treated with PROCR nanobody, compared to tumors treated with IgG (left). The ratios of PROCR+ cells, PROCR− cells (Lin−, EpCAM+) and EpCAM− (Lin−, EpCAM−) cells were quantified (right). Data are pooled from 3 independent experiments and presented as mean ± SEM. ***P < 0.001. c Lin−, EpCAM+ cells were isolated from the remaining tumors post IgG or PROCR-mAb treatment, and transplanted in limiting dilution. The post PROCR-mAb treatment group had significantly lower tumor formation frequency (1/432) compared to the IgG group (1/53). ***P < 0.001. d Mice baring the PDX tumor (~200 mm3) were administered with paclitaxel and doxorubicin (PTX + DOX) or PROCR-mAb alone or in combination. PTX + DOX were administered at d14, d20 and d26 (PTX: 20 mg/kg, DOX: 3 mg/kg body weight) (green arrowheads); PROCR-mAb or IgG were administered at d14, d17, d20, d24, d28 and d32 (8 mg/kg body weight) (blue arrows). Tumor sizes were suppressed for 2-fold with PTX + DOX, 3-fold with PROCR-mAb, 32-fold with the combination treatment. Post d32, all treatments were removed. Tumor sizes were continued measured (during the withdrawal of treatment). Tumor sizes increased quickly when withdrawal of PTX/DOX treatment (green line), which was in contrast to the moderate increase of tumor sizes post antibody treatment (blue line) and combination treatment (black line). n = 10 mice or more in each group. Data are presented as mean ± SEM. ***P < 0.001
We set out to unequivocally prove that growth inhibition of tumors is due to reduced CSCs following nanobody treatment of PROCR+ BC PDX. Total Lin−, EpCAM+ cells were isolated from the remaining tumors and transplanted again in limiting dilution. In the Ctrl group (post-IgG injection), tumor formation frequency was 1/53 (Fig. 7c), with no significant difference with that of parental tumor. However, the post-nanobody treatment group demonstrated markedly lower tumor formation frequency (1/432) compared to the Ctrl (Fig. 7c). These results suggest that CSCs are indeed reduced upon the nanobody treatment in PROCR+ BC PDX.
Next, we asked whether the nanobody is able to affect established tumors. The treatment began when PROCR+ BC PDX tumors had developed to be about 200 mm3. The nanobody or chemotherapeutic agents (paclitaxel and doxorubicin; PTX/DOX) alone or in combination were administered to the mice. We found that the nanobody alone exhibits more pronounced tumor suppressive effects (three-fold inhibition, blue line) than PTX/DOX (two-fold inhibition) (Fig. 7d, green line). Remarkably, combination treatment completely inhibited the growth of the established tumors (32-fold inhibition) (Fig. 7d, black line). The decreased proportion of PROCR+ cells (CSCs) post the antibody treatment forecasted a slower tumor relapse. Indeed, the rapid relapse was not observed upon withdrawal of the antibody treatment (blue and black lines, Fig. 7d). In contrast, withdrawal of PTX/DOX treatment led to a rapid recurrence shown by sharply increasing tumor sizes (green line in Fig. 7d), consistent with the increased proportion of CSCs (PROCR+ cells) upon PTX/DOX treatment (Supplementary information, Fig. S6g). Collectively, these data reinforce the notion that inhibition of PROCR is able to suppress the growth of PROCR+ BC through directly targeting the CSCs, and demonstrate the potential for PROCR inhibitory monoclonal antibodies as targeting agents to inhibit this subgroup of breast cancer.
Discussion
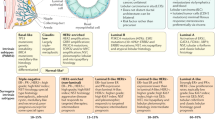
Our study reveals the functional role of PROCR in CSCs of a discrete subgroup of TNBC. TNBC is viewed as a group of different diseases that have similar phenotypes but different genotypes with varying prognoses and responses to chemotherapy. Our study, as illustrated in Fig. 8a, suggests a further stratification of TNBC based on PROCR expression, revealing a PROCR+ BC subgroup, which constitutes about half of TNBC cases in the current study cohort, and is associated with poorer prognosis compared with PROCR-low TNBCs. Furthermore, in PROCR+ BCs, PROCR+ cells are CSCs that can be targeted by PROCR-inhibitory nanobody. The new stratification based on PROCR expression has broad clinical implications. First, PROCR can be measured by IHC, a routine practice in breast cancer diagnosis and treatment planning. Second, providing that the molecular pathways driving tumor-survival in PROCR+ BC have been identified, the new stratification will foster a novel treatment strategy for these patients, potentially employing a combined inhibition of these pathways.
A proposed model for TNBC stratification & targeted therapy based on PROCR expression. a Illustration of PROCR-based stratification and targeted therapy. Our data suggest a further stratification of TNBC based on PROCR expression. The potential PROCR+ BC subgroup (PROCR-high TNBC) constitutes about half of TNBC cases. In PROCR+ BCs, PROCR is expressed at the surface of CSCs and can be targeted by inhibitory antibodies, resulting in inhibition of ERK, PI3K-Akt, RhoA pathways and suppression of EMT in CSCs, rendering tumor inhibition. b Schematic model of the human breast epithelial hierarchy and progression of PROCR+ BC. Hypothetically, PROCR+ BC may result from the acquisition of genetic alterations in MaSCs, which become CSCs fueling the growth of the tumor
There have been seminal studies demonstrating the existence of cancer stem cells in breast cancers using proposed stem cell markers.11,12 Interestingly, PROCR expression has been detected in CSCs populations enriched by one of those markers, CD44.22 However, variability among different breast cancer subtypes is likely to be extensive, and it may thus be challenging and imprecise to designate a single CSC population for all breast cancers. It is noteworthy that using PROCR-high TNBC cell lines, e.g. MDA-MB-231, to investigate the relationship between PROCR and CSCs has been proven not suitable, because almost all MDA-MB-231 cells exhibit high PROCR expression.23 The separation of PROCR+ and PROCR− cells amongst MDA-MB-231 cells in previous studies may have been compromised by the inefficacy of antibody used for FACS analysis.28,29 In this study, using PDX samples, we propose that PROCR+ cells are CSCs in PROCR+ BC subtypes, defined by the following functional assays for CSCs.30,31 They can (1) be prospectively purified (Lin−, EpCAM+, PROCR+); (2) be passaged in a xenograft assay at clonal cell doses; (3) generate a xenograft that is representative of the parent tumor; and (4) give rise to daughter cells (PROCR− cells) that may possess proliferative capacity but are unable to establish the tumor. Our study further demonstrates that PROCR is a druggable target on the surface of CSCs. Inhibition of PROCR by an inhibitory nanobody results in a concomitant blockade of MEK-ERK, PI3K-Akt-mTOR and RhoA-ROCK pathways and repression of EMT, rendering a high degree of tumor suppression. Of note, PROCR inhibition has no effect on growth of PROCR-low TNBC (quadruple-negative BC, QNBC) or ER+/PR+ tumors, highlighting that the identity of CSC and custom intervention will depend on the BC subtype.
The putative connection between MaSCs of normal tissues and CSCs remains unclear despite some recent advances.32,33,34 A recent study demonstrates that mammary tumors with stem cell cell-of-origin exhibit high ZEB1 expression and an EMT signature, and low chromosomal instability.35 Procr expression marks normal mouse MaSCs that exhibit EMT characteristics.14 In this study, our data demonstrate that PROCR expression also marks EMT-featuring CSCs in human PROCR+ BC. Thus, it is tempting to hypothesize that PROCR+ BC is a malignancy originated from normal MaSCs, and that after MaSCs acquire genetic alterations and become CSCs, the expression and molecular mechanism of PROCR are still coupled with these cells (Fig. 8b). It should be noted that CSCs and cell-of-origin are different sides of cancer.36 In this particular subtype, it is plausible that CSCs are originated from the normal MaSCs. On the other hand, accumulating evidence supports the idea that BRCA1 mutant TNBCs originate from luminal progenitors.37,38,39,40,41 It is highly likely that PROCR+ TNBC and BRCA1 TNBC, which are distinct subtypes, have distinct cell-of-origins. An alternative, although not mutually exclusive model is that the PROCR+ tumors are driven by Wnt signaling. Procr is a Wnt target.14 In the current study, Gene Set Enrichment Analysis suggests that Wnt signaling activities are higher in PROCR+ cells, together with Myc and EMT signatures. In the mouse, tumors driven by Wnt1 (MMTV-Wnt1) share similarity with human PROCR+ tumors, in particular their CSCs can be marked by Procr/PROCR expression. It should be noted that MMTV-Wnt1 tumors do not mirror PROCR+ human tumors in the following aspects: (1) MMTV-Wnt1 is an overall basal tumor mixed with low levels of ER expression, whereas PROCR+ human tumors belong to TNBC; (2) MMTV-Wnt1 tumors have only 8% Procr+ cells, while PROCR+ human tumors contain a much higher percentage of PROCR+ cells. Generation of a more relevant mouse model for the investigation of the cell-of-origin and driving signaling of PROCR+ tumor will be the subject of future study.
PDXs have become a prominent model for studying human cancer in vivo. Cancers established as PDXs can retain molecular heterogeneity characteristic of those in patients, which is critical for biomarker specificity and fundamentally impacts therapeutic responses. While PDXs are valuable preclinical models for addressing some of the most challenging barriers to successful drug development, caution in interpreting the potential of a drug tested should be noted. To circumvent immune rejection, human cancers must be transplanted into immunocompromised mice. However, therapeutic responses in general are likely influenced by preexisting cancer-dependent immune phenotypes and immune responses elicited upon therapy-induced tumor perturbation. The extent to which compromised immune systems limit predictive value for a given therapeutic approach remained to be determined.42
In conclusion, our findings identify PROCR as a biomarker to stratify TNBC into clinically relevant subgroups and reveal the key roles of PROCR in tumorigenesis of PROCR+ BCs. Remarkably, a PROCR inhibitory nanobody effectively suppresses the growth of PROCR+ BCs. Our study suggests that PROCR is a promising cell surface target for therapeutic intervention of PROCR+ breast cancers.
Materials and methods
Patients and specimens
Human breast tissues were obtained from Fudan University Shanghai Cancer Center, with approval from the Human Research Ethics Committee of Fudan University Shanghai Cancer Center. Fresh samples were histologically normal para-tumor breast tissue. For the immunohistochemical analysis of PRCOR in breast tumor whole-sections, a total of 80 stage I to III primary breast cancer samples from females with invasive ductal carcinoma were randomly collected at the Department of Breast Surgery at the Fudan University Shanghai Cancer Center between August 2013 and March 2014. The clinical pathologic diagnosis of breast cancer cases was determined by pathologists in the Department of Pathology. In our study, ER, PR, HER2 expression statuses were also determined by IHC staining. Most, but not all, patients with HER2 expression status (IHC, score ≥ 2) were subjected to fluorescence in situ hybridization (FISH) screening for HER2 gene amplification. The HER2 overexpression subgroup was defined as FISH positive or an IHC staining score ≥ 3. As a result, the breast cancer patients were classified into four molecular subtypes according to the ER, PR, and HER2 status, including luminal A subtype (ER+ and/or PR+, low Ki67), luminal B subtype (ER+ and/or PR+, high Ki67 or HER2+), HER2+ subtype (HER2+, ER− and PR−), and triple-negative subtype (ER−, PR−, and HER2−). Total 80 breast cancer samples (20 for each subtypes) were obtained to examine the PROCR protein level by immunohistochemical analysis using breast tumor whole-sections.
To evaluate the prognostic value of PROCR in a large breast cancer patient cohort, we used tissue microarrays (TMAs) containing 450 pathologically proven breast cancer samples and 72 non-cancerous mammary controls to examine the PROCR expression level. The eligibility criteria of breast cancer samples have been described in a previous study.43 Briefly, the breast cancer patients in this cohort fulfilled the following inclusion criteria: (i) female patients diagnosed with stage I to III primary breast cancer; (ii) patients with unilateral invasive ductal carcinoma (IDC); ductal carcinomas in situ were excluded; (iii) patients without any evidence of metastasis at diagnosis; (iv) patients underwent a mastectomy and axillary lymph node dissection or breast conservation surgery followed by adjuvant chemotherapy; the therapeutic regimen decisions were based on the Chinese Anti-Cancer Association guidelines for the diagnosis and treatment of breast cancer.
For tissue microarrays (TMAs), we used the complete random sampling method to collect 207 luminal-like subtype cases, 93 HER2-enriched subtype cases and 150 triple-negative subtype cases from 1709 cases that met the eligibility criteria and were diagnosed as breast cancer at the Department of Breast Surgery in FDSCC between August 2001 and January 2008. In addition, as described previously,43 a total of 72 non-cancerous mammary tissue controls with pathologically confirmed benign mammary diseases were also collected from women who had come to the Outpatient Department at FDSCC for breast cancer screening during the period from January 2013 to February 2013. This study was approved by the institutional review board (IRB) of Fudan University Shanghai Cancer Center (FDSCC), and all participants provided informed consent to participate in this research.
Tissue microarray (TMA)
TMAs were constructed using above 450 paraffin-embedded blocks of breast tumors and 72 blocks of non-cancerous mammary controls using a tissue micro arrayer (UNITMA Instruments, Seoul, Korea). The hematoxylin and eosin (HE)-stained slides from tumors were evaluated to identify representative tumor regions. TMAs were composed of two 1.0 mm tissue cores from different areas of the same tumor to compare staining patterns. TMA sections were subsequently dewaxed in xylene and rehydrated in ethanol for IHC staining.
Evaluation of IHC variables in breast tumor whole-sections and in TMAs
In 80 cases of breast tumor whole-sections, expression of PROCR were semiquantitatively classified according to the immunoreactive H-score (HS; range 0–300) which was calculated as the result of the intensity score (1, faint/week; 2, moderate; 3, strong) multiplied by the distribution score (between 1 [percentage] to 100 [percentage]).
In TMAs, a total of 450 IDC breast cancer cases and 72 non-cancerous mammary tissues were included. Of these cases, 7 breast cancer cases and 1 non-cancerous sample experienced duplicate tissue core loss after IHC staining. Thus, the remaining 443 cancerous and 71 non-cancerous mammary samples were included in the subsequent analysis. The duplicate tissue cores from each case were also stained and scored semi-quantitatively using the same H-score evaluating criteria in breast tumor whole-sections. Subsequently, stratification scoring was conducted according to H-score as follow: HS < 80, scored as 0; 80 < HS < 120, scored as 1; 120 < HS < 200, scored as 2; HS > 200, scored as 3. If the score was equal to or greater than 2, the tumor was considered to have high PROCR expression; otherwise, low PROCR expression was classified. Based on the evaluation standard, scoring was reviewed in parallel by D.W. and F.Q., both examiners were blinded to all clinical data.
Kaplan-Meier analysis using TMAs and Kaplan-Meier Plotter
In the above cohort in TMAs, the breast cancer patients were regularly followed, and the clinical outcome of 415 cases was obtained, with the last update occurring in October 2014. The follow-up period was defined as the time from surgery to the last observation for censored cases or relapse/death for complete observations. Disease-free survival (DFS) was defined as the time from the date of primary surgery to the date of relapse/breast cancer-specific death or October 2014. The categories analyzed for DFS were first recurrence of disease at a local, regional, or distant site and breast cancer-specific death. Patients with study end date and loss of follow-up were considered censored. Thus, these 415 cancerous cases were analyzed in the Kaplan-Meier analysis.
In addition, a large public clinical database (Kaplan-Meier Plotter) of breast cancer was used to explore the association between PROCR expression and clinical outcomes, with the following restricted condition: (1) 140 months of follow-up time, (2) select media cutoff, (3) cases with ER status. Primary purpose of the tool is a meta-analysis-based in silico biomarker assessment. We evaluated the effects of PROCR expression on disease-free survivals (DFS) of 671 hormone receptor-negative patients and 1802 hormone receptor-positive patients with the latest version of this database (2014 version; http://www.kmplot.com/analysis/index.php?p = service).
Experimental animals
MMTV-Wnt1, MMTV-PyVT, K14-Cre, Brca1f/+, p53f/+, Nude, FVB and SCID/Beige mice strains were used. Experimental procedures were approved by Animal Care and Use Committee of Shanghai Institute of Biochemistry and Cell Biology, Chinese Academy of Sciences.
Cell lines and cell culture
The MDA-MB-231, BT549, MCF-7 human breast cancer cell lines and the HEK293T cell line were obtained from the Shanghai Cell Bank Type Culture Collection Committee or American Type Culture Collection (ATCC) and maintained in complete growth medium as recommended by the distributor.
Antibodies
Antibodies used in immunohistochemistry: Mouse anti-human PROCR (1:300, Abcam, cat#ab56689), rabbit anti-human K14 (1:100, Thermo, cat#MA5-132-3), mouse anti-ER (1:50, DAKO, cat#M7047, clone 1D5), mouse anti-PR (1:50 DAKO, cat#M3569, clone PgR 636), rabbit anti-HER2 (1:50, Proteintech, cat#18299-1-AP).
Antibodies used in Western blotting: Rabbit anti-human PROCR (1:200, Novus, cat#H00010544-D01P), rabbit anti-human phosphor-Src (1:1000, Cell Signaling Technology, cat#2101, clone 7G9), rabbit anti-human total Src (1:1000, Cell Signaling Technology, cat#2108), rabbit anti-human phosphor-MEK (1:1000, Cell Signaling Technology, cat#9154, clone 41G9), mouse anti-human total MEK (1:1000, Cell Signaling Technology, cat#4694, clone L38C12), rabbit anti-human phosphor-ERK (1:1000, Cell Signaling Technology, cat#4370, clone D13.14.4E), rabbit anti-human total ERK (1:100, Santa Cruz, cat#Sc-94), rabbit anti-human phosphor-Akt (1:1000, Cell Signaling Technology, cat#4065), rabbit anti-human total Akt (1:1000, Cell Signaling Technology. cat#4691, clone C67E7), rabbit anti-human phosphor-GSK3β (1:1000, Cell Signaling Technology, cat#9331), rabbit anti-human total GSK3β (1:1000, Cell Signaling Technology, cat#12456, clone D5C5Z), rabbit anti-human phosphor-S6K (1:1000, Cell Signaling Technology, cat#9025), rabbit anti-human total S6K (1:1000, Cell Signaling Technology, cat#9202), mouse anti-human c-Myc (1:100, Santa Cruz, cat#Sc-40), mouse anti-human Cyclin D1 (1:100, Santa Cruz, cat#Sc-45-), mouse anti-human RhoA (1:100, Santa Cruz, cat#Sc-418), rabbit anti-human ROCK2 (1:1000, Cell Signaling Technology, cat#9029, clone D1B1), rabbit anti-human phosphor-p38 (1:1000, Cell Signaling Technology, cat#9211), rabbit anti-human total p38 (1:1000, Cell Signaling Technology, cat#9212), rabbit anti-human phosphor-IGF1R (1:1000, Cell Signaling Technology, cat#3024, clone 19H7), rabbit anti-human total IGF-1R (1:1000, Cell Signaling Technology, cat#3027), rabbit anti-human phosphor-EGFR (Tyr1068, Tyr1173, Tyr845) (1:1000, Cell Signaling Technology, cat#3777, 4407, 2231, clone D7A5, 53A5), rabbit anti-human total EGFR (1:1000, Cell Signaling Technology, cat#2232), mouse anti-tubulin (1:5000, Sigma, cat#T5168, clone B-5-1-2), rabbit anti-GAPDH (1:5000, Proteintech, cat#10494-1-AP) and mouse anti-Actin (1:5000, Sigma, cat#A2228, clone AC-74).
Primary cell preparation
The minced primary tumor or mammary tissue was placed in culture medium (RPMI 1640 with 25 mM HEPES, 5% fetal bovine serum (FBS), 1% PS (Penicillin-Streptomycin), 300 U ml−1 Collagenase III (Worthington)) and digested for up to 3 h at 37 °C. After lysis of the red blood cells, a single-cell suspension was obtained by sequential incubation with 0.25% trypsin-EDTA at 37 °C for 5 min and 0.1 mg/ml DNase I (Sigma) for 5 min with gentle pipetting, followed by filtration through 70 μm cell strainers.
Cell labeling and flow cytometry
The following antibodies in 1:200 dilutions were used: PE-cy7 anti-human EpCAM (Biolegned, cat#324222, clone 9C4), APC anti-human CD49f (Biolegned, cat#313616, clone GoH3), FITC anti-human CD31 (Biolegned, cat#303104, clone WM59), FITC anti-human CD45 (Biolegned, cat#304006, clone HI30), FITC anti-human CD235a (Biolegend, cat#329104, clone HI264), APC anti-human PROCR (eBioscience, cat#17-2018-42, clone RCR-227), FITC anti-mouse CD31 (BD, cat#553372, clone MEC13.3), FITC anti-mouse CD45 (BD, cat#553080, clone 30-F11), FITC anti-mouse Ter119 (BD, cat#557915, clone Ter119), PE anti-mouse Procr (eBioscience, cat#12-2012-82, clone 1560), PE-cy7 anti-mouse CD24 (Biolegned, cat#101804, clone M1/69), APC anti-mouse CD29 (Biolegned, cat#102216, clone HMB1-1). Antibody incubation was performed on ice for 20 min in HBSS with 10% fetal bovine serum. For DNA content analysis, Hoechst (1ug/ml) was used. All sortings were performed using an FCASJazz (Becton Dickinson). The purity of sorted population was routinely checked and ensured to be more than 95%.
Immunohistochemistry
Tissue paraffin or frozen sections were incubated with primary antibodies at 4 °C overnight, followed by washes, incubation with secondary antibodies for 2 h at 25 °C, and counterstaining with DAPI (Sigma). For all of the immunofluorescence staining at least 3 independent experiments were conducted. Representative images are shown in the figures.
Immunohistochemistry for PROCR was performed using anti-PROCR antibody and Goat Anti-mouse HRP (1:1000, Santa Cruz) as secondary antibody followed by color development (DAKO) before counterstaining with hematoxylin. The color developing time is tightly controlled as 2 min for all of the specimens and TMAs. All of the reagents are from the same lot number. Two examiners were blinded to all clinical data during PROCR IHC staining and the H-score reading and their results were also blinded to each other.
shRNA and sgRNA constructs
The shRNAs targeting hPROCR sequences were constructed in lentivirus-based pLKO.1-EGFP constructs (Addgene). The efficiency of individual shRNA was validated by western blotting or qPCR. The sequences for hPROCR-shRNA-1 and hPROCR-shRNA-3 are
5’ GCAGCAGCTCAATGCCTACAA 3’ and
5’ TGGCCTCCAAAGACTTCATAT 3’.
If not specified, sh-PROCR represents hPROCR-shRNA-1.
Sequence for mouse Procr-shRNA is
5’ TTGTGTGGAGTTCCTGGAGGA 3’
dCas9-KRAB plasmids are from Addgene. The sgRNAs targeting hPROCR genome sequence were constructed in lentivirus-based plasmid (MP177 from Addgene). The efficiency of individual sgRNA was validated by Western blotting.
Sequence for hPROCR suppression is
5’ CAGACTCCGCCCCTCCCAGA 3’.
Competitive ELISA assays
Purified Protein C (100 μl, 0.2 μg/ml) was pre-coated to the bottom of a 96-well plate at 4 °C overnight. The wells were washed with PBS containing 0.5% Tween-20 and blocked with 1% BSA. A mixture of purified sPROCR (100 μl, 3 μg/ml) and the competing antibody or control antibody (in limiting dilution) was added into the wells and incubated for 2 h at 37 °C. The bound sPROCR was detected after subsequent incubation with a biotin conjugated PROCR primary antibody (R&D Systems) for 1.5 h and Streptavidin HRP secondary antibody (R&D Systems) for 30 min. After HRP color detection, the absorbance was determined with a microplate reader at 450 nm. All tests were performed in triplicate.
In vitro cell proliferation assays
MDA-MB-231 or BT549 cells infected with scramble/PROCR shRNA or treated with PROCR-mAb were plated at similar cell number and passaged every 2 days. During each passage, cell numbers were counted. To evaluate the inhibitory antibody effect on MDA-MB-231 cell proliferation, control non-neutralizing or neutralizing antibodies (200 μg/ml) were added every 24 h.
EdU labeling assays
In PDX samples, the PROCR+ and PROCR− cells were FACS isolated and cultured in 2D overnight in complete growth medium. Cells were then incubated with EdU for 1 h. After 15 min fixation with PFA, the EdU color development was performed following manufacturer’s protocol.
MDA-MB-231 cells were plated at a low density (5 × 104) onto coverslips in 12-well plate and cultured with antibodies in complete culture medium. After 16 h, cells were incubated with EdU for 1 h, followed by PBS washes and fixation with 4% PFA for 10 min. the EdU color development was performed following manufacturer’s protocol (EdU: Life Technology, C10339).
Mammary fat pad xenograft and analysis
Nude mice were used for transplantation with mouse MMTV-PyVT, K14-Cre;Brca1f/+;p53f/+ tumor cells, human breast cancer PDX cells or human breast cancer cell lines. Both Nude and FVB mice were used for transplantation with MMTV-Wnt1 tumor cells. Cells were resuspended in 50% Matrigel (diluted in PBS with 50% FBS) and 0.04% Trypan Blue (Sigma), and injected in 10–20 μl volumes into the fat pads of 8-week-old mice. For in vivo knockdown of PROCR/Procr, MDA-MB-231, MCF-7, human PDXs cells or mouse MMTV-Wnt1 tumor cells were virally infected by ctrl or shPROCR/shProcr/sgPROCR + dCas9-KRAB virus. The infected cells were sorted based on the tagged GFP expression in the vector and resuspended in the above condition for transplantation. Tumor diameters were serially measured with calipers. Tumor volume (mm3) was calculated by the following formula: volume = length × width2 × 0.52. Mouse weight was monitored closely. For tumor inhibition, IgG (control) or neutralizing antibody (8 mg/kg body weight), Doxorubicin (3 mg/kg body weight) and Paclitaxel (20 mg/kg body weight) were intraperitoneal administered following the protocol described in Fig. 7 and Supplementary information, Fig. S6. At least four mice per experimental group were used in animal experiments. All animals were of the same age and sex at the time of mammary epithelial cell or tumor cell injection. No statistical method was used to pre-determine sample size. The experiments were not randomized. There was no blinded allocation during experiments and outcome assessment.
Generation of patient-derived xenografts from human breast cancers
PDX lines were originally initiated by implantation of a fresh patient tumor fragment into the mammary fat pad of recipient SCID/Beige mice and were maintained by serial passage in vivo in Nude mice at intervals characteristic for each line, and in accordance with Institutional Animal Care and Use Committee requirements.
Generation of PROCR monoclonal antibody
The phage display method was used to generate PROCR monoclonal antibody. In brief, the naive llama (camelid) sdAb phagemid library (Genscript) was used for selection of PROCR-specific single domain antibodies. FC-conjugated PROCR extracellular domain (FC-ECD) (1-214aa) protein produced in HEK293T cells was used as target antigen for several rounds of selection to enrich the specific sdAb binders. The plasmids were isolated from the output of phage display and constructed into a vector containing human IgG1 for soluble sdAb screening. Soluble sdAb clones were screened by ELISA using FC-ECD. High affinity sdAbs were selected and their binding to PROCR was further verified by FACS analysis using HEK293 cells stably expressing full length PROCR. The inhibitory activity of the antibody was further examined by competitive Elisa as described above. Clone GS-5 was used in all in vitro cell culture and in vivo studies.
Statistical analysis
Student’s t-test was performed when comparing two groups. One-way ANOVA was performed for comparison among more than three groups. P value was calculated in Prism6 on data represented by bar charts, which consisted of results from three independent experiments unless specified otherwise. For all experiments with error bars, the std. error of mean (SEM) was calculated to indicate the variation within each experiment. The experiments were not randomized. The investigators were not blinded to allocation during experiments and outcome assessment.
References
Foulkes, W. D., Smith, I. E. & Reis-Filho, J. S. Triple-negative breast cancer. N. Engl. J. Med. 363, 1938–1948 (2010).
Lehmann, B. D. et al. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J. Clin. Invest. 121, 2750–2767 (2011).
Metzger-Filho, O. et al. Dissecting the heterogeneity of triple-negative breast cancer. J. Clin. Oncol. 30, 1879–1887 (2012).
Carey, L., Winer, E., Viale, G., Cameron, D. & Gianni, L. Triple-negative breast cancer: disease entity or title of convenience? Nat. Rev. Clin. Oncol. 7, 683–692 (2010).
Perou, C. M. et al. Molecular portraits of human breast tumours. Nature 406, 747–752 (2000).
Prat, A. et al. Phenotypic and molecular characterization of the claudin-low intrinsic subtype of breast cancer. Breast Cancer Res. 12, R68 (2010).
Adamo, B. & Anders, C. K. Stratifying triple-negative breast cancer: which definition(s) to use? Breast Cancer Res. 13, 105 (2011).
Visvader, J. E. & Lindeman, G. J. Cancer stem cells: current status and evolving complexities. Cell Stem Cell 10, 717–728 (2012).
Magee, J. A., Piskounova, E. & Morrison, S. J. Cancer stem cells: impact, heterogeneity, and uncertainty. Cancer Cell 21, 283–296 (2012).
Nguyen, L. V., Vanner, R., Dirks, P. & Eaves, C. J. Cancer stem cells: an evolving concept. Nat. Rev. Cancer 12, 133–143 (2012).
Al-Hajj, M., Wicha, M. S., Benito-Hernandez, A., Morrison, S. J. & Clarke, M. F. Prospective identification of tumorigenic breast cancer cells. Proc. Natl Acad. Sci. USA 100, 3983–3988 (2003).
Ginestier, C. et al. ALDH1 is a marker of normal and malignant human mammary stem cells and a predictor of poor clinical outcome. Cell Stem Cell 1, 555–567 (2007).
Meyer, M. J. et al. CD44posCD49fhiCD133/2hi defines xenograft-initiating cells in estrogen receptor-negative breast cancer. Cancer Res. 70, 4624–4633 (2010).
Wang, D. et al. Identification of multipotent mammary stem cells by protein C receptor expression. Nature 517, 81–84 (2015).
Li, Y. et al. Evidence that transgenes encoding components of the Wnt signaling pathway preferentially induce mammary cancers from progenitor cells. Proc. Natl Acad. Sci. USA 100, 15853–15858 (2003).
Tsukamoto, A. S., Grosschedl, R., Guzman, R. C., Parslow, T. & Varmus, H. E. Expression of the int-1 gene in transgenic mice is associated with mammary gland hyperplasia and adenocarcinomas in male and female mice. Cell 55, 619–625 (1988).
Herschkowitz, J. I. et al. Identification of conserved gene expression features between murine mammary carcinoma models and human breast tumors. Genome Biol. https://doi.org/10.1186/Gb-2007-8-5-R76 (2007).
Guy, C. T., Cardiff, R. D. & Muller, W. J. Induction of mammary tumors by expression of polyomavirus middle T oncogene: a transgenic mouse model for metastatic disease. Mol. Cell Biol. 12, 954–961 (1992).
Xu, X. et al. Conditional mutation of Brca1 in mammary epithelial cells results in blunted ductal morphogenesis and tumour formation. Nat. Genet 22, 37–43 (1999).
Cleary, A. S., Leonard, T. L., Gestl, S. A. & Gunther, E. J. Tumour cell heterogeneity maintained by cooperating subclones in Wnt-driven mammary cancers. Nature 508, 113–117 (2014).
Li, Y., Hively, W. P. & Varmus, H. E. Use of MMTV-Wnt-1 transgenic mice for studying the genetic basis of breast cancer. Oncogene 19, 1002–1009 (2000).
Shipitsin, M. et al. Molecular definition of breast tumor heterogeneity. Cancer Cell 11, 259–273 (2007).
Wang, D. et al. Protein C receptor stimulates multiple signaling pathways in breast cancer cells. J. Biol. Chem. 293, 1413–1424 (2018).
Neve, R. M. et al. A collection of breast cancer cell lines for the study of functionally distinct cancer subtypes. Cancer Cell 10, 515–527 (2006).
Gilbert, L. A. et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154, 442–451 (2013).
Hamers-Casterman, C. et al. Naturally occurring antibodies devoid of light chains. Nature 363, 446–448 (1993).
Tice, D. A., Biscardi, J. S., Nickles, A. L. & Parsons, S. J. Mechanism of biological synergy between cellular Src and epidermal growth factor receptor. Proc. Natl Acad. Sci. USA 96, 1415–1420 (1999).
Hwang-Verslues, W. W. et al. Multiple lineages of human breast cancer stem/progenitor cells identified by profiling with stem cell markers. PLoS ONE 4, e8377 (2009).
Schaffner, F. et al. Endothelial protein C receptor function in murine and human breast cancer development. PLoS ONE 8, e61071 (2013).
Kreso, A. & Dick, J. E. Evolution of the cancer stem cell model. Cell Stem Cell 14, 275–291 (2014).
Clarke, M. F. et al. Cancer stem cells–perspectives on current status and future directions: AACR Workshop on cancer stem cells. Cancer Res. 66, 9339–9344 (2006).
Chakrabarti, R. et al. DeltaNp63 promotes stem cell activity in mammary gland development and basal-like breast cancer by enhancing Fzd7 expression and Wnt signalling. Nat. Cell Biol. 16, 1001–1013 (2014).
Zhang, M. et al. Identification of tumor-initiating cells in a p53-null mouse model of breast cancer. Cancer Res. 68, 4674–4682 (2008).
Su, X. et al. TAp63 suppresses mammary tumorigenesis through regulation of the Hippo pathway. Oncogene, https://doi.org/10.1038/onc.2016.388 (2016).
Morel, A. P. et al. A stemness-related ZEB1-MSRB3 axis governs cellular pliancy and breast cancer genome stability. Nat. Med 23, 568–578 (2017).
Lindeman, G. J. & Visvader, J. E. Insights into the cell of origin in breast cancer and breast cancer stem cells. Asia Pac. J. Clin. Oncol. 6, 89–97 (2010).
Lim, E. et al. Aberrant luminal progenitors as the candidate target population for basal tumor development in BRCA1 mutation carriers. Nat. Med. 15, 907–913 (2009).
Molyneux, G. et al. BRCA1 basal-like breast cancers originate from luminal epithelial progenitors and not from basal stem cells. Cell Stem Cell 7, 403–417 (2010).
Proia, T. A. et al. Genetic predisposition directs breast cancer phenotype by dictating progenitor cell fate. Cell Stem Cell 8, 149–163 (2011).
Nolan, E. et al. RANK ligand as a potential target for breast cancer prevention in BRCA1-mutation carriers. Nat. Med. 22, 933–939 (2016).
Sau, A. et al. Persistent activation of NF-kappaB in BRCA1-deficient mammary progenitors drives aberrant proliferation and accumulation of DNA damage. Cell Stem Cell 19, 52–65 (2016).
Day, C. P., Merlino, G. & Van Dyke, T. Preclinical mouse cancer models: a maze of opportunities and challenges. Cell 163, 39–53 (2015).
Ye, F. G. et al. Cytidine deaminase axis modulated by miR-484 differentially regulates cell proliferation and chemoresistance in breast cancer. Cancer Res. 75, 1504–1515 (2015).
Acknowledgements
We are grateful to Drs. Chi-Chung Hui, Esther Verheyen and Roel Nusse for critical reading of the manuscript. We thank Drs. Dangsheng Li and Weiguo Zou for helpful discussion. We thank Dr. Suling Liu for providing reagents. We thank Dr. Jianhua Sui for advice on antibody development. This work was supported by grants from National Natural Science Foundation of China (31830056, 31861163006, 31625020, 31530045, 31661143043 to Y.A.Z, 81874113 to Z.-M.S; 81672601, 81872137 to X.H.), Chinese Academy of Sciences (XDA12020378 to Y.J. and XDB19000000 to Y.A.Z.), Shanghai Municipal Science and Technology Commission (17XD1404000 to Y.A.Z.) and National Ten Thousand Talents Program (to Y. A. Z.).
Author information
Authors and Affiliations
Contributions
Y.A.Z and Z.-M.S conceived the study. D.W. performed most FACS analyses, genetic, xenograft and in vitro experiments. C.L., R.Y. performed xenograft, antibody and chemo agent administration. D.W., Y.J., C.C. and L.B. performed monoclonal antibody screening and characterization. J.W. performed mouse breeding. X.H., S.L., F.Q., L.Y., L.C. and Z.-M.S provided TMA and PDX samples. Y.B. and Y.-R.L performed bioinformatics analysis. D.W., X.H. and Y.A.Z. analyzed the data and wrote the manuscript. C.L., G.G., H.J., D.L., L.L. and J.C. helped project design and manuscript editing.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Wang, D., Hu, X., Liu, C. et al. Protein C receptor is a therapeutic stem cell target in a distinct group of breast cancers. Cell Res 29, 832–845 (2019). https://doi.org/10.1038/s41422-019-0225-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41422-019-0225-9
This article is cited by
-
LINC00853 contributes to tumor stemness of gastric cancer through FOXP3-mediated transcription of PDZK1IP1
Biological Procedures Online (2023)
-
Protein C receptor maintains cancer stem cell properties via activating lipid synthesis in nasopharyngeal carcinoma
Signal Transduction and Targeted Therapy (2022)
-
MUC1 triggers lineage plasticity of Her2 positive mammary tumors
Oncogene (2022)
-
Lentiviral CRISPR-guided RNA library screening identified Adam17 as an upstream negative regulator of Procr in mammary epithelium
BMC Biotechnology (2021)
-
Predicting cell-to-cell communication networks using NATMI
Nature Communications (2020)