Abstract
Transplantation of hematopoietic stem cells (HSCs) from human umbilical cord blood (hUCB) holds great promise for treating a broad spectrum of hematological disorders including cancer. However, the limited number of HSCs in a single hUCB unit restricts its widespread use. Although extensive efforts have led to multiple methods for ex vivo expansion of human HSCs by targeting single molecules or pathways, it remains unknown whether it is possible to simultaneously manipulate the large number of targets essential for stem cell self-renewal. Recent studies indicate that N6-methyladenosine (m6A) modulates the expression of a group of mRNAs critical for stem cell-fate determination by influencing their stability. Among several m6A readers, YTHDF2 is recognized as promoting targeted mRNA decay. However, the physiological functions of YTHDF2 in adult stem cells are unknown. Here we show that following the conditional knockout (KO) of mouse Ythdf2 the numbers of functional HSC were increased without skewing lineage differentiation or leading to hematopoietic malignancies. Furthermore, knockdown (KD) of human YTHDF2 led to more than a 10-fold increase in the ex vivo expansion of hUCB HSCs, a fivefold increase in colony-forming units (CFUs), and more than an eightfold increase in functional hUCB HSCs in the secondary serial of a limiting dilution transplantation assay. Mapping of m6A in RNAs from mouse hematopoietic stem and progenitor cells (HSPCs) as well as from hUCB HSCs revealed its enrichment in mRNAs encoding transcription factors critical for stem cell self-renewal. These m6A-marked mRNAs were recognized by Ythdf2 and underwent decay. In Ythdf2 KO HSPCs and YTHDF2 KD hUCB HSCs, these mRNAs were stabilized, facilitating HSC expansion. Knocking down one of YTHDF2′s key targets, Tal1 mRNA, partially rescued the phenotype. Our study provides the first demonstration of the function of YTHDF2 in adult stem cell maintenance and identifies its important role in regulating HSC ex vivo expansion by regulating the stability of multiple mRNAs critical for HSC self-renewal, thus identifying potential for future clinical applications.
Similar content being viewed by others
Introduction
Hematopoietic stem cells (HSCs) are responsible for life-long hematopoiesis under homeostatic and stress conditions, relying on an exquisite balance between stem cell self-renewal and differentiation.1,2 Thus, HSC transplantation offers a life-saving therapy for a broad spectrum of disorders, including hematologic, immune, and genetic diseases, as well as cancers.3 However, HSC-based treatment is limited primarily by the lack of human leukocyte antigen (HLA)-matched donor bone marrow (BM). Allogeneic transplantation offers an alternative approach, but graft vs. host disease (GvHD) remains a life-long challenge, as immuno-suppressive medicine has numerous side effects, such as delayed immunological recovery and thrombotic microangiopathy.4,5 Transplantation of HSCs from human umbilical cord blood (hUCB) reduces the risk of GvHD; however, the lower number of HSCs in hUCB than in BM or mobilized peripheral blood limits its application.3 Whereas the targeting of single molecules or pathways has been studied as a possible route toward hUCB HSC expansion,6,7,8,9,10,11,12,13,14,15,16 manipulating the posttranscriptional modification of N6-methyladenosine (m6A,) which controls the fate of a large set of mRNAs, might offer an alternative approach to favor stem cell self-renewal relative to differentiation.17
m6A is a prevalent internal modification of mRNAs that regulates the outcome of gene expression by modulating RNA processing, localization, translation, and eventual decay, all of which can be modulated by “writers,” “erasers,” and “readers” of the mark.18,19 Recent studies have elucidated the roles of m6A modification in stem cell-fate determination and in the endothelial-to-hematopoietic transition during embryogenesis20,21,22,23,24,25 as well as in development of leukemia.26,27,28,29 Interestingly, deficiency of the m6A writer complex, Mettl3 and Mettl14, leads to noticeable outcomes in different types of stem cells. Thus, e.g., Mettl3 or Mettl14 KO promotes differentiation in HSCs,27,28,29 while resulting in impaired priming and thus enhanced stem cell self-renewal and maintenance in mouse embryonic stem cells and in embryonic neuronal stem cells.20,22 The physiological functions of m6A in stem cells and leukemia are mediated through different mechanisms. In stem cells, m6A modifications regulate stem cell-fate determination through the m6A-mediated decay of mRNAs encoding stem cell-fate determinants20,22 whereas in acute myeloid leukemia, Mettl3 and Mettl14 promote leukemogenesis, because m6A modifications stabilize the mRNAs of oncogenes and/or increase their translation.27,28,29 Moreover, previous studies have reported that the leukemogenic functions of FTO and Mettl14 are independent of YTHDF reader proteins.26,28 Bearing this in mind, we have focused on investigating the role of YTHDF2, a well-recognized m6A reader, in promoting targeted mRNA decay30 in the context of HSC maintenance. We hypothesize that manipulation of YTHDF2 might potentially influence the life span of a great number of m6A-marked mRNAs, thus having an impact upon adult HSC self-renewal vs. differentiation and facilitating HSC expansion. Here we show that depletion of YTHDF2 specifically expands mouse and human HSC numbers without skewing lineage fate. Our studies uncover an essential role of YTHDF2 in regulating HSC self-renewal and provide a novel approach to enhance hUCB HSC numbers by ex vivo expansion, thus offering potential for future clinical applications.
Results
Ythdf2 KO leads to an increase in phenotypical HSCs in mice
To investigate the effects of Ythdf2 on phenotypic HSCs, we first used Crispr-Cas9 technology to generate Ythdf2f/f floxed mice. We then used Mx1-Cre mice to achieve conditional knockout (KO) and specifically reduce Ythdf2 expression in hematopoietic cells (hereafter referred to as Ythdf2 KO mice) (Fig. 1a, b). BM HSPCs from these mice showed no discernible differences in the absence of polyinosinic:polycytidylic acid (pI:pC) (Supplementary information, Figure S1a). However, 4 weeks after pI:pC injections, we observed that Ythdf2 KO mice showed a significant increase in both the frequency and absolute number of long-term HSCs (Lin−Sca1+ cKit+ (LSK) CD34−Flk2−; LT-HSCs) and short-term HSCs (LSK CD34+ Flk2−; ST-HSCs) but not in multipotent progenitors (LSK CD34+ Flk2+; MPPs) in comparison with their wild-type (wt) littermates (Fig. 1c to e). Although the Ythdf2 KO led to increased BM cellularity, there was no significant difference in the absolute number of committed progenitors, including common myeloid progenitors, granulocyte-macrophage progenitors, megakaryocyte-erythrocyte progenitors, and common lymphoid progenitors, as well as mature lineage cells, erythrocytes, myeloid cells, B cells, and T cells between Ythdf2 KO and wt mice (Fig. 1f to h). Cell cycle analysis revealed no discernible change in quiescence of HSCs or MPPs after Ythdf2 KO (Supplementary information, Figure S1b). Notably, the proportion of apoptotic cells in Ythdf2 KO LT-, ST-HSCs, and MPPs was significantly reduced compared with wt controls (Supplementary information, Figure S1c). To further identify any potential HSC defects in Ythdf2 KO mice, we examined the number of HSCs, committed progenitors, and mature lineages in the spleen and found no significant differences between wt and Ythdf2 KO mice (Supplementary information, Figures S1d to h). In summary, Ythdf2 KO in primary mice specifically increases HSC numbers with no bias or defects in either progenitor or lineage cells.
Ythdf2 KO leads to an increase in phenotypic HSCs in mice. a Deletion of YTHDF2 in the HSPCs of Mx1-cre;Ythdf2f/f conditional KO (cKO) mice. b Western blotting (left) and histogram (right) showing intracellular flow validation of KO Ythdf2 in mouse HSPCs. c Representative flow plots of HSPCs in BM from wt and Ythdf2 KO mice (n = 5 for each group). d, e Frequency in total nucleated cells (TNCs) (d) and absolute cell number (e) of HSPCs in BM from wt and Ythdf2 KO mice (n = 5 for each group). f Absolute number of BM TNCs from wt and Ythdf2 KO mice (n = 5 for each group). g, h Absolute number of committed progenitors (g) and lineage cells (h) in BM of wt and Ythdf2 KO mice (n = 5 for each group). Data shown as mean ± SEM. Unpaired t-test. n.s. nonsignificant. CMP, common myeloid progenitor; GMP, granulocyte-macrophageprogenitor; MEP, megakaryocyte-erythrocyte progenitor; CLP, common lymphoid progenitor; Erythro, erythrocyte
Ythdf2 KO expands functional HSCs in mice
To determine whether Ythdf2 KO expands functional HSC numbers, we initially performed a short-term homing assay by transplanting 1 × 106 carboxyfluorescein diacetate succinimidyl ester (CFDA SE)-labeled BM cells from KO mice or their control littermates into lethally irradiated recipient mice. This revealed no significant difference in homing capacity between mutant and wt controls (Supplementary information, Figure S1i). We then carried out a limited dilution, competitive repopulation unit assay (LDA) by transplanting 2 × 105, 7.5 × 104, or 2.5 × 104 donor BM cells (CD45.2), together with 2 × 105 recipient BM cells derived from the ptprc mutant strain (CD45.1), into lethally irradiated recipient mice (Fig. 2a). Consistent with the increased number of cells of HSC phenotype in Ythdf2 KO mice, we found that competitive repopulating units (CRUs) increased 2.2-fold in Ythdf2 KO HSCs compared with controls (Fig. 2b). In the group of 2 × 105Ythdf2 KO donor cells we observed a significant increase in the overall repopulation rate compared with controls at 16 weeks post transplantation (Fig. 2c). Moreover, recipients of Ythdf2 KO BM cells exhibited markedly higher frequency and absolute number of donor-derived LT-HSCs and ST-HSCs but not MPPs in BM compared with controls (Fig. 2d, e). Furthermore, we found that the donor-derived committed progenitors and mature lineages in BM from transplantation recipients of mutant and wt cells showed no significant changes (Fig. 2f, g). To determine the long-term repopulation ability of HSCs from Ythdf2 KO mice, we conducted secondary transplantation with BM cells derived from primary recipients. Notably, we found that compared to controls, CRUs from Ythdf2 KO cells showed a 3.5-fold increase (Fig. 2h) and there were no signs of leukemia in BM or spleen 16 weeks after secondary transplantation (Supplementary information, Figures S2a to f). We also investigated the long-term effect of Ythdf2 KO on hematopoiesis under homeostatic conditions by examining stem cells, progenitor cells, and lineages in both BM and spleen at over 5 months following pI:pC injections (Supplementary information, Figure S3a). Although we observed a modest increase in the number of LT-HSCs in BM from Ythdf2 KO mice compared with controls (Supplementary information, Figure S3c), there were no discernible differences between Ythdf2 KO and control mice in total numbers of nucleated cells, progenitors, and lineage cells from either the BM or spleen (Supplementary information, Figures S3b and S3d to j). These observations indicate that the long-term effect of Ythdf2 KO in vivo neither skews lineage differentiation nor facilitates aberrant proliferation. These findings are in line with previous reports that Ythdf2 is not required for leukemogenesis.26,28 To verify the frequency of functional HSCs in the BM 5 months after pI:pC administration, we transplanted 7.5 × 104 BM cells from wt and Ythdf2 KO mice together with competent cells into lethally irradiated recipients. We found that Ythdf2 KO led to significantly higher engraftment compared with wt controls, indicating that the number of functional, although not phenotypical, HSCs remains higher in Ythdf2 KO mice 5 months post treatment than in controls (Supplementary information, Figure S3k). Taken together, these data indicate that Ythdf2 KO results in specific and significant expansion of mouse HSCs in vivo without affecting lineage commitment.
Ythdf2 KO results in expansion of functional HSCs in mice. a Experimental scheme for limiting dilution transplantation assay (LDA) to determine the frequency of functional HSCs. b Primary LDA to determine the CRU frequency by ELDA (Extreme Limiting Dilution Analysis) at 16 weeks post transplantation (n = 10 per group). c Competitive reconstitution assay by transplanting 200 K whole bone marrow (WBM) cells with 200 K rescue cells into irradiated recipients (n = 10 for each group). d, e Frequency in TNC (d) and absolute cell number (e) of donor-derived HSPCs in BM from transplantation recipient mice as c (n = 10 for each group). f, g Absolute cell number of donor-derived (CD45.2+) committed progenitors (f) and lineage cells (g) in BM from primary 200 K BM transplantation recipient mice (n = 9–10 for each group). h Secondary LDA to determine the long-term CRU frequency by ELDA at 16 weeks after secondary transplantation (n = 10). Data shown as mean ± SEM. Unpaired t-test. n.s. nonsignificant
YTHDF2 regulates HSC self-renewal gene expression by m6A-mediated mRNA decay
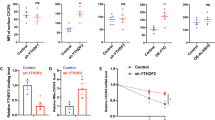
To explore the underlying mechanisms of how Ythdf2 KO expands HSCs, we first mapped the m6A methylome using methylated RNA immunoprecipitation combined with high-throughput sequencing (MeRIP-seq or m6A-seq) in LT-HSCs, ST-HSCs, and MPPs sorted from adult C57BL/6J mice.31,32,33 The m6A peaks were selected as significantly enriched overlapping peaks from two independent replicates. Consistent with previous studies,31,33 we found that m6A peaks were abundant in mRNA open reading frames (ORFs), in 3′-untranslated regions (UTRs), and around the stop codons in all three HSPC populations (Supplementary information, Figures S4a and b). Transcripts of moderately expressed genes were more likely to be methylated (Supplementary information, Figure S4c). Intriguingly, we found that m6A modifications were enriched in the mRNAs of transcription factors (TFs), such as Gata2, Etv6, Stat5, and Tal1, which have been documented to be critical for HSC self-renewal and stem cell state maintenance,34,35,36,37,38,39,40,41 suggesting the m6A modification could play critical roles in the regulation of HSCs (Supplementary information, Table S1). Given the accumulating evidence that m6A mRNA methylation regulates stem cell-fate determination by facilitating the decay of mRNAs encoding transcription factors and genes in key signaling pathways involved in self-renewal and differentiation,20,21,22,23,24,26,42 we measured mRNA degradation rates in LT-, ST-HSCs, and MPPs by monitoring mRNA levels after transcription inhibition with actinomycin D. We found that the degradation rates of methylated mRNAs were significantly higher than those of unmethylated mRNAs in ST-HSCs, and MPPs (Supplementary information, Figure S4d). As YTHDF2 is a well-recognized m6A “reader” that mediates mRNA decay,30 we determined its targets by performing infrared UV-crosslinking immunoprecipitation sequencing (irCLIP-seq) in the mouse multipotent hematopoietic precursor cell line HPC743,44 (Fig. 3a; Supplementary information, Figures S5a to c). The results showed that 57.8% of YTHDF2 target mRNAs contained m6A peaks (Supplementary information, Figure S5d). YTHDF2-binding sites were enriched with the conserved m6A motif and exhibited the characteristic of m6A distribution features (Fig. 3b, c). Gene ontology (GO) analysis of YTHDF2 target transcripts revealed enrichment of genes related to hematopoietic or lymphoid organ development, suggesting the involvement of YTHDF2 in the regulation of hematopoiesis (Fig. 3d). Notably, we found that YTHDF2 bound to transcription factor mRNAs, such as those of Tal1 and Gata2, on sites largely overlapping with m6A peaks (Fig. 3e; Supplementary information, Figure S5e and Table S2). We did not observe any significant difference in the total mass of RNA in LSK Flk2- cells isolated from wt and Ythdf2 KO mice (Supplementary information, Figure S6a). Although m6A modification was present in only 0.1–0.4% of the adenosine nucleotides in mammalian cells,17 we found that Ythdf2 KO led to an increased level of m6A content in total RNA from BM Lin− cells (Supplementary information, Figure S6b). This suggests that YTHDF2 specifically regulates the stability of m6A-marked mRNAs. Accordingly, quantitative PCR (qPCR) analysis of total mRNA revealed increased levels of Tal1, Gata2, Runx1, and Stat5a (representative target mRNAs of Ythdf2) in Ythdf2 KO LSK cells compared with wt controls (Fig. 3f). Single-cell immunofluorescence staining and intracellular flow cytometry further revealed that Ythdf2 KO HSPCs exhibited significant increases in the intensities of m6A-labeled transcription factors involved in stem cell self-renewal, such as TAL1, GATA2, RUNX1, and STAT5, indicative of a suppressive role of YTHDF2 in HSC self-renewal (Fig. 3g; Supplementary information, Figure S6c). A previous study has shown that YTHDF2 regulates RNA metabolism by localizing the bound mRNAs to mRNA decay sites.30 To further explore the mechanism of YTHDF2 in regulating HSC self-renewal, we performed fluorescence in situ hybridization (FISH) of Tal1 mRNA and fluorescence immunostaining of YTHDF2 and DCP1a, a marker of mRNA decay sites,45,46 and analyzed their relative spatial distribution in sorted wt and Ythdf2 KO HSPCs. Co-localization of Tal1 mRNA, DCP1a and YTHDF2 was observed in wt cells but was substantially reduced in Ythdf2 KO cells (Fig. 3h, i). Similar findings were made by co-staining to reveal Gata2 mRNA with YTHDF2 and DCP1a in wt or Ythdf2 KO HSPCs (Supplementary information, Figures S6d and e). To determine whether increased expression of transcription factors, including Tal1, accounted for HSC expansion in Ythdf2 KO mice, we performed rescue experiments using short hairpin (sh) RNA-mediated Tal1 knockdown (KD) in wt and Ythdf2 KO LSK cells followed by transplantation into lethally irradiated recipients. Depletion of Tal1 in HSPCs significantly impaired the reconstitution capacity of wt cells as reported previously47 and also partially rescued the increased engraftment of Ythdf2 KO cells (Supplementary information, Figure S6f). Overall, these data indicate that YTHDF2 regulates HSC self-renewal by enabling the degradation of mRNAs encoding transcription factors essential for stem cell self-renewal.
YTHDF2 functions as an m6A reader and regulates HSC gene expression by mediating mRNA decay. a Schematic of irCLIP-seq workflow. b YTHDF2-binding motif identified by MEME with all irCLIP peaks found in all three replicates. c Pie chart depicting the fraction of YTHDF2-binding peaks in each of five transcript segments. d GO enrichment analysis of YTHDF2 targets from intersect genes of three YTHDF2 irCLIP-seq replicates. e Representative tracks of Tal1 harboring m6A peaks and YTHDF2 irCLIP peaks. Coverage of m6A immunoprecipitation and input fragments indicated in red and grey, respectively. YTHDF2 irCLIP reads highlighted in yellow. Arrows indicate the overlapped peaks between m6A-seq and irCLIP-seq. f qPCR analysis of total mRNA of sorted LSK cells from wt and Ythdf2 KO mice. All Ct values were first normalized to Actb control (not m6A-tagged) and the ratio (Ythdf2 KO over wt) was then calculated. (n = 3). g Representative images (left) and quantification (right) of staining intensity of wt (n = 65) and Ythdf2 KO (n = 54) HSPCs for TAL1 (green). h Fluorescence in situ hybridization of Tal1 mRNA (red) and fluorescence immunostaining of DCP1a (P-body marker) (magenta), YTHDF2 (green) in wt and Ythdf2 KO HSPCs. Arrows indicate colocalized staining. Scale bars, 5 μm. i Quantification of Tal1 mRNA and DCP1a co-localization in sorted LSK cells from wt and Ythdf2 KO mice. Percentage indicates the average frequency of the Tal1 mRNA that colocalized with DCP1a over total Tal1 mRNA level in each LSK cells (n = 9–17). Data shown as mean ± SEM. Unpaired t-test
Dissecting the role of YTHDF2 in human UCB HSPCs by m6A-seq and RNA-seq
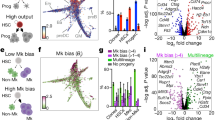
The limited number of HSCs in a single hUBC blood unit has been an obstacle for clinical applications, such as HSC transplantation.3 Our observation that Ythdf2 KO resulted in an increase of phenotypic and functional mouse HSCs prompted us to test whether YTHDF2 KD could facilitate human UCB HSC expansion. We first performed m6A-seq with CD34+ cells isolated from three individual hUCB samples (Supplementary information, Figure S7a). The m6A modifications occurred predominantly in mRNAs (~95%) where they were preferentially found in ORF regions, 3′-UTRs, and close to the stop codon, as expected (~90%) (Fig. 4a, b; Supplementary information, Figure S7b). We further compared m6A landscapes in mouse and hUCB HSPCs, and found that there were 2,239 genes in common that were m6A tagged (Fig. 4c). The m6A-tagged transcripts in common were enriched for genes participating in hematopoiesis and stem cell maintenance (Fig. 4d). The enrichment of m6A labeling in the mRNAs of transcription factors functioning in mouse HSC self-renewal led us to perform GO term analysis upon m6A-marked transcription factor transcripts in hUCB CD34+ cells. Among the 722 m6A-labeled transcription factor mRNAs identified, the major GO terms related to cell-fate commitment and stem cell maintenance (Supplementary information, Figure S7c). For example, HOXB4, whose overexpression has been reported to expand human and mouse HSCs ex vivo,9,10 was marked by m6A in hUCB CD34+ cells (Fig. 4e). Other transcription factors required for HSC self-renewal and critical for inducing HSCs from other cell types,35,48 such as Zfx, RUNX1, and FOSB, were also m6A-tagged in hUCB CD34+ cells (Supplementary information, Figures S7f, g and Table S3). To further dissect the role of YTHDF2 in hUCB HSPCs, we performed RNA sequencing (RNA-seq) with control or YTHDF2 KD hUCB CD34+ cells (Fig. 4f; Supplementary information, Figures S7d and e). Remarkably, transcripts marked by m6A, including HOXB4 and other HSC self-renewal-related transcription factors, showed significant increases of input mRNA reads in YTHDF2 KD cells compared with control cells, whereas there were no notable changes for non-m6A labelled genes (Fig. 4g to i; Supplementary information, Figures S7f and g). These results support the role of YTHDF2 in regulating hUCB HSC self-renewal through RNA degradation.
Role of YTHDF2 in human cord blood HSCs by m6A-seq and RNA-seq analysis. a Metagene profiles depicting sequence coverage in windows surrounding the TSS (left) and stop codon (right). Coverage of m6A IP and control (input) fragments indicated in red and grey, respectively. b Pie chart presenting the fraction of m6A peaks in each of five non-overlapping transcript segments. c Venn diagram showing shared and unique m6A-tagged genes in mouse and hUCB HSPCs. d GO enrichment analysis of m6A-tagged transcripts shared in both mouse and hUCB HSPCs. e Representative tracks of HOXB4 harboring m6A peaks. Color codes are the same as in a. f Lentivirus mediated YTHDF2 KD in hUCB CD34+ HSPCs for RNA-seq. g Cumulative distribution of log2 (fold change) for m6A-marked genes (purple line) and non-m6A-marked genes (black line) with control and YTHDF2 KD hUCB CD34+ cells. h Representative coverage plots from the RNA-seq analysis, showing increased reads of m6A-tagged gene HOXB4 but not a non-m6A-tagged gene ACTB in YTHDF2 KD compared with control hUCB CD34+ cells. i Relative mRNA expression levels of non-m6A labeled ACTB (as control) and m6A-marked transcription factor related to stem cell self-renewal in control and YTHDF2 KD hUCB CD34+ cells. RPKM from RNA-seq analysis were normalized to controls. Adjusted P-value indicated. n.s. nonsignificant
Expansion of hUCB HSCs by YTHDF2 KD
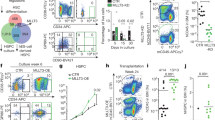
To further explore whether suppression of YTHDF2 can expand human HSCs, we next performed shRNA-induced YTHDF2 KD in hUCB HSPCs as above (Fig. 4f). After 7 days of ex vivo culture, lentiviral-mediated KD of YTHDF2 resulted in an average increase of 14.3-fold and 13.6-fold, respectively, in the frequency and absolute number of CD34+ CD38− CD45RA− EPCR+ HSCs and a 5.1-fold increase in CFUs relative to control cells. Among the different categories of CFUs, the most dramatic increase was seen in the most primitive CFUs representing the granulocyte-erythrocyte-monocyte-megakaryocyte colony type and burst forming unit-erythroid, reflecting a higher expression level of key hematopoietic transcription factors, such as TAL1, in YTHDF2 KD hUCB cells (Fig. 5a to d; Supplementary information, Figure S8a). Interestingly, the apoptotic rate was significantly reduced in YTHDF2 KD hUCB HSPCs compared with control cells, similar to the trend of HSCs in Ythdf2 KO mouse (Fig. 5e). We next compared the effect of overexpression of YTHDF2 on HSPC function and found that this reduced clonogenic potential of hUCB HSPCs by 2.2-fold, confirming that YTHDF2 negatively regulates HSC maintenance ex vivo (Supplementary information, Figures S8b and c).
YTHDF2 KD facilitates expansion of human cord blood HSCs ex vivo. a, b Fold change of frequency (a) and absolute number (b) of indicated cells in YTHDF2 KD over control cells after 7 days culture. c CFU output from transduced CD34+ CD38− hUCB cells and images of CFU the granulocyte-erythrocyte-monocyte-megakaryocytes (GEMMs) (scale bar, 200 μm). d Images of burst forming unit-erythroid (BFU-E) (left) and colony-forming unit-granulocyte/macrophage (CFU-GM) (right) from 7 days cultured control or YTHDF2 KD hUCB cells (scale bar, 200 μm). Three independent cord blood samples were used and this was repeated twice for a to d. e Apoptosis analysis of CD34+ CD38- cells in 7-day cultures of transduced CD34+ hUCB cells by Annexin V staining (n = 3 independent CB samples). Data shown as mean ± SEM. Unpaired t-test. n.s. nonsignificant
To determine whether YTHDF2 KD can expand human HSCs in vivo, we performed LDA by transplanting GFP+ cells sorted from hUCB CD34+ cells infected with control and YTHDF2 shRNA 4 days infection (Fig. 6a). We analyzed BM cells from recipient NOD/SCID Il2rgnull (NSG) mice 10 weeks post transplantation and measured the functional HSC frequency after in vivo expansion. Notably, compared with the control group, recipients of YTHDF2 KD cells displayed a ninefold increase in human hematopoietic cell (hCD45+ GFP+) engraftment in BM without changes in the proportions of the lineages (Fig. 6b, c; Supplementary information, Figures S9a and b). YTHDF2 KD significantly increased the proportion of myeloid, megakaryocytes, and erythrocytes in the BM of primary recipients (Supplementary information, Figure S9c). Accordingly, we found that the HSC frequency in YTHDF2 KD cells was increased 4.4-fold relative to control cells (Fig. 6d; Supplementary information, Table S4). We confirmed the long-term capability of YTHDF2 KD hUCB HSCs to be reconstituted and to undergo self-renewal; 12 weeks after transplantation of BM from primary recipients into sub-lethally irradiated secondary NSG recipient mice, human hematopoietic cell chimerism in the BM was higher in the YTHDF2 KD group compared with the control group (Fig. 6e, f; Supplementary information, Figure S9d). CRUs from YTHDF2 KD cells showed an eightfold increase relative to control cells (Fig. 6g; Supplementary information, Table S4). These data demonstrate that YTHDF2 KD remarkably facilitates the expansion of both phenotypic and long-term functional hUCB HSCs ex vivo.
YTHDF2 KD facilitates expansion of human cord blood functional long-term HSCs. a Experimental scheme for measuring frequency of HSCs after in vivo expansion. b Representative flow plots of hCD45+ GFP+ reconstitution from primary recipient mice receiving the highest two cell doses. hCD45, human CD45 (c) hCD45+ GFP+ engraftment in BM from the primary recipient mice that received the highest two doses (n = 8). d HSC frequency determined by primary LDA. Dashed lines indicate 95% confidence intervals. e Representative flow plots of hCD45+ GFP+ reconstitution from secondary recipient mice receiving the highest two cell doses. f hCD45+ GFP+ engraftment in BM from secondary recipient mice that received the highest two doses (n = 6). g HSC frequency determined by secondary LDA. Dashed lines indicate 95% confidence intervals. Data shown as mean ± SEM. Unpaired t-test. n.s. nonsignificant
Discussion
Although recent studies shed light on the biological functions of mRNA m6A modifications,49,50,51,52,53,54,55,56,57,58,59,60,61,62,63,64,65,66,67 our work identifies Ythdf2 as an important regulator of human and mouse HSC self-renewal by coupling the posttranscriptional m6A modification to the degradation of mRNAs encoding key transcription factors for self-renewal. Repression of Ythdf2 in mouse HSPCs and hUCB HSCs leads to increased expression of multiple key TFs critical for self-renewal, thereby facilitating ex vivo expansion of both phenotypic and functional HSCs without any noticeable lineage bias or leukemic potential. In addition, stem cell niches may, to some extent, contribute to mouse HSC expansion mediated by Ythdf2 suppression as Mx1-cre can be activated in mesenchyme stromal cells.68 It would be intriguing to study the function of Ythdf2 on mesenchymal stem cells (MSCs) and how repression of Ythdf2 in both HSCs and MSCs may synergistically expand HSCs in vivo. Our ex vivo expansion of human HSCs has shown that suppression of Ythdf2 in HSCs is able to expand HSCs as seen in this study.
Given the broad and complicated impact of the m6A writer complex METTL3 and METTL14 on mRNA splicing, translation, and primary miRNA processing,56,67,69 METTL3 or METTL14 depletion results in distinct outcomes in normal stem cells and in leukemia. Recent studies have demonstrated that METTL3 and METTL14 have critical roles in leukemia development and maintenance of leukemia stem cells,27,28,69 while depletion of METTL3 leads to an increase in phenotypic HSCs at the expense of functionality due to lost of stem cell quiescence (cite https://doi.org/10.1038/s41422-018-0062-2). By contrast, Ythdf2 is mainly involved in the influence of m6A-mediated mRNA decay upon stem cells.20,22,23,30 Therefore, manipulating YTHDF2 should extend the half-life of specific m6A-marked mRNAs encoding TFs critical for stem cell self-renewal without affecting other aspects of mRNA processing. In line with this, our data clearly show that YTHDF2 depletion in HSCs neither skews the lineage commitment nor induces hematological malignancies, reducing the risk of leukemogenesis with expanded HSCs. Furthermore, stem cell self-renewal is a complex process that promotes cell division, inhibits apoptosis, prevents differentiation, and retains stemness.70 Our observation that YTHDF2-deficient HSCs exhibited lower apoptotic rates accounts for at least one aspect of the stem cell self-renewal mechanism. We believe that retention of stemness provides another mechanism underlying the YTHDF2-mediated HSC expansion.
A major limitation in using hUCB HSC transplantation is the insufficient number of HSCs in one hUCB unit. Although previous studies have revealed that Dlk1, SR1, Musashi2, and UM171 can expand hUCB HSCs by targeting Notch, AHR signaling, or other unknown pathways,7,8,11,15 our work provides a novel and potent way to target multiple key TFs critical for HSC self-renewal and to enhance the expansion of HSCs. Reducing Ythdf2 level and function during in vitro culture either pharmacologically or through AAV-mediated KD permits the Ythdf2 level and function to be restored after transplantation in vivo, and thus without affecting normal HSC maintenance and function in human patients. Furthermore, we envision that combining our method with previous ones may facilitate the expansion of not only human HSCs, but also other stem cells, offering potential approaches for future stem cell-based therapies.
Materials and methods
Mice
Ythdf2 conditional KO mice were generated by the groups of Chuan He and Bin Shen. Mice were housed in the animal facility at Stowers Institute for Medical Research (SIMR) and handled according to Institute and NIH guidelines. All procedures were approved by the Institutional Animal Care and Use Committee (IACUC) of SIMR.
Flow cytometry and HSPC sorting
Mouse HSPCs, progenitors, and lineage cells were collected from BM (femur and tibia) and spleen. Red blood cells were lysed using a 0.16 M ammonium chloride solution, and the cells were filtered with 70 μm strainers to generate single-cell suspensions. For mouse HSC identification, cells were stained with antibodies against Sca-1 (D7), c-Kit (2B8), CD34 (RAM34), Flk2 (A2F10), CD48 (HM48-1), CD150 (TC15-12F12.2), together with lineage cocktail including CD3 (145-2C11), CD4 (RM4-5), CD8 (53–6.7), Mac-1 (M1/70), Gr1 (RB6-8C5), CD45R (B220, RA3-6B2), IgM (Il-41), and Ter119 (TER-119). To identify progenitors and cell lineages, cells were stained with antibodies as previously described.71 7-Aminoactinomycin D (A1310, Life Technologies) was used to exclude dead cells. hUCB samples were acquired from the St. Louis Cord Blood Bank. Mononuclear cells were isolated with Lymphoprep™ (StemCell Technologies), followed by isolation of human CD34+ cord blood cells by human CD34 MicroBead Kit UltraPure (Miltenyi Biotec). To quantify hUCB HSPCs, cells were stained with antibodies against CD34 (581), CD38 (HIT2), CD45RA (HI100), CD49f (GoH3), and EPCR/CD201 (RCR-401). Cell sorting and analyses were performed on MoFlo (Dako), InFlux Cell Sorter (BD Biosciences), and/or MACSQuant (Miltenyi Biotec). Data analysis was performed using FlowJo software.
Homing assay
In vivo homing assays were performed as previously described.72 Basically, whole BM (WBM) cells from CD45.2 mice were labelled with 5 μM 5-(and -6)-CFDA SE (Molecular Probes) at 37 °C for 10 min and washed three times. WBM cells (1 × 106) were then transplanted into lethally irradiated ptprc mice. After 18 h, the femurs and tibias were flushed, and numbers of CFDA SE + cells were determined.
Competitive reconstitution assay
Competitive reconstitution assays were performed by intravenous transplantation of 2 × 105, 7.5 × 104, or 2.5 × 104 donor-derived WBM cells from wt or Ythdf2 KO mice (CD45.2), together with 2 × 105 rescue cells (CD45.1) into groups of ten lethally irradiated (10 Gy) ptprc recipient mice. For secondary transplantation, primary transplant recipients were sacrificed. BM cells were dissected from femur and tibia, and then transplanted mouse-to-mouse at a dosage of 1 × 106 cells into irradiated secondary recipient mice. Baytril water was given to recipient mice 3 days before irradiation and continued for another 2 weeks after irradiation. Primary and secondary CRU frequencies were measured using ELDA software,73 in which successful engraftment was defined as the presence of a distinct CD45.2+ CD45.1− population ≥ 5% and ≥ 1% of total hematopoietic cells in peripheral blood, respectively.74 Recipient mice undergoing secondary transplantation that had died before 16 weeks post transplantation were also scored as having failed engraftment.
For the rescue experiment, LSK cells were sorted from wt and Ythdf2 KO mouse BM and cultured overnight in StemSpan SFEM medium (Stem Cell Technologies) supplemented with 10 μg/mL heparin (Sigma), 0.5 × penicillin/streptomycin (Sigma), 10 ng/mL recombinant mouse (rm) stem cell factor (SCF) (Biovision, Inc.), and 20 ng/mL Tpo (Cell Sciences, Inc.)75 at 37 °C in a 5% CO2 5% O2 atmosphere. Control shRNA and shRNA targeting mouse Tal1 were delivered into LSK cells by lentivirus at 100 mulitpicity of infection (MOI). To enhance the efficiency of infection, ViroMag Transduction Reagent (OZ Biosciences) was used following manufacturer’s directions. GFP+ cells were sorted at 5 days post infection and cultured for another 3 days to expand their number before transplantation. Equal numbers of GFP+ cells from each group were transplanted with 105 rescue BM cells into lethally irradiated recipients.
Cell cycle and apoptosis assays
Cell cycle analysis was performed with the fluorescein isothiocyanate mouse anti-human Ki-67 set (BD Pharmingen) according to the manufacturer’s instructions. Briefly, 5 × 106 BM cells were isolated and stained with HSC antibodies as described above. Cells were fixed by 4% paraformaldehyde at 4 °C overnight or room temperature (RT) for 1 h and then permeabilized with 0.2% Triton X-100 on ice for 15 mins. Cells were washed with phosphate-buffered saline containing 2% fetal bovine serum, and then were incubated with Ki-67 antibody at RT for 1 h in the dark, and SYTOX Red (Invitrogen) at RT for another 5 mins, followed by flow cytometric analysis with an InFlux Cell Sorter (BD Biosciences). For apoptosis analysis, Annexin V (Invitrogen) and SYTOX Red staining of 5 × 106 BM cells was performed according to the manufacturer’s protocol.
m6A-seq
Two replicates of 105 LT-HSC (LSK CD34− Flk2−), ST-HSC (LSK CD34+ FLK2−), and MPP (LSK CD34+ FLK2+) cells from C57BL/6J mouse were sorted into TRIzol (Invitrogen) and total RNA was isolated according to the manufacturer’s instructions. RNA was fragmented to fragments of approximately 100 nucleotides with the Ambion fragmentation reagent (2 min incubation at 70 °C). The samples were then subjected to Turbo DNase treatment (Ambion), followed by a phenol–chloroform extraction, and resuspension in 85 μl of nuclease-free water; 5 μl was saved to record the input. The remaining 80 μl of RNA fragments was diluted into IPP buffer (150 mM NaCl, 0.1% NP-40, 10 mM Tris-HCl, pH 7.5). RNA was incubated with 25 μl of protein-G magnetic beads, previously bound to 3 μg of anti-m6A polyclonal antibody (Synaptic Systems), for 3 h at 4 °C in IPP buffer. Beads were washed twice with 200 μl IPP buffer, twice with 200 μl low-salt buffer (50 mM NaCl, 0.1% NP-40, 10 mM Tris-HCl, pH 7.5) and twice with 200 μl high-salt buffer (500 mM NaCl, 0.1% NP-40, 10 mM Tris-HCl, pH 7.5). Beads were then treated with 300 μl Elution Buffer (5 mM Tris-HCL pH 7.5, 1 mM EDTA pH 8.0, 0.05% SDS, 4.2 μl Proteinase K (20 mg/ml)) for 1.5 h at 50 °C, and RNA was recovered with phenol:chloroform extraction followed by ethanol precipitation. Three human CD34+ umbilical cord blood cells were isolated as described above and total RNA isolated with TRIzol. RNA was fragmented into fragments of approximately 100 nucleotides with the Ambion fragmentation reagent (2 min 50 s incubation at 70 °C). The samples were then subjected to Turbo DNase treatment (Ambion), followed by phenol:chloroform extraction, and resuspension in 18 μl of nuclease-free water; 1 μl was saved to monitor input. m6A RNA immunoprecipitation was performed with EpiMark® N6-Methyladenosine Enrichment Kit following manufacturer’s instructions.
Following m6A preparation of RNA, quality was assessed on Agilent 2100 Bioanalyzer, and 1 ng (mouse) or 10 ng (human) RNA was used to generate RNAseq libraries according to the manufacturer’s directions for the SMARTer Stranded Total RNA-Seq Kit–Pico Input Mammalian (Takara Bio, Inc.) using 16 cycles (mouse) or 13 cycles (human) PCR2 amplification. This method uses random priming and a template switching oligo to generate complementary DNA, followed by the ligation of barcoded adapters; ribosomal-derived cDNA is then removed through probe-directed enzyme cleavage and subsequent enrichment of non-cleaved fragments. We modified the protocol to retain lower molecular weight sample fragments by using a 1.2 × SPRI bead concentration for the PCR1 cleanup. To remove dimerized adapters, libraries underwent 160–600 bp size selection with a Pippin Prep (Sage Science) 2% gel. The resulting libraries were checked for quality and quantity using the Bioanalyzer and Qubit Fluorometer (Life Technologies). Then equal molar libraries were pooled and requantified. For mouse m6A-seq, libraries were sequenced as 50 bp single read on the Illumina HiSeq 2500 instrument using HiSeq Control Software 2.2.58. Following sequencing, Illumina Primary Analysis version RTA 1.18.64 and Secondary Analysis version bcl2fastq2 v2.18 were run to demultiplex reads for all libraries and generate FASTQ files. For human m6A-seq, libraries were sequenced as a 75 bp single read on the Illumina NextSeq instrument using NextSeq Control Software 2.1.2. Following sequencing, Illumina Primary Analysis version NextSeq RTA 2.4.11 and Secondary Analysis version bcl2fastq2 v2.18 were run to demultiplex reads for all libraries and generate FASTQ files.
Plasmid construction and stable cell line generation
Mouse Ythdf2 (mYthdf2) was re-cloned from a commercial cDNA clone (ORIGENE # MC200730) into vector pcDNA5/FRT/Flag plasmid using primers listed: mYthdf2 ORF Clone BamhI F: 5′-CGC GGA TCC TCG GCC AGC CTC TTG GA-3′ and mYthdf2 ORF Clone NotI R: 5′-ATA AGA ATG CGG CCG CCT ATT TCC CAC GAC CTT GAC GT-3′. Then Flag-mYthdf2 was subcloned under EF1a promoter in pSicoR-EF1a-IRES-EGFP lentiviral construct (Gibson Assembly®, forward primer: 5′-GTC GAC GGT ACC GCG GGC CCA TGG ATT ACA AGG ATG ACG ACG-3′ and reverse primer: 5′-GAG GGA GAG GGG CGG ATC CCC TAT TTC CCA CGA CCT TGA CGT-3′). Human Ythdf2 (hYthdf2) was cloned from plasmid provided by the Chuan He lab using primers indicated: Forward 5′-CGT TCG AAA TGT CGG CCA GCA GCC TCT-3′; Reverse 5′-TCC CCC GGG TTA TTT CCC ACG ACC TT-3′. Then hYthdf2 was cloned into pSicoR-EF1a-IRES-EGFP constructs under EF1a promoter by BstBI and XmaI restriction digestion and ligation. To generate the Flag-mYthdf2 HPC7 stable cell line, lentiviruses were generated by transfection of pSicoR-EF1a-Flag-mYthdf2-IRES-EGFP constructs together with the psPAX2 and pMD2.G plasmids at a ratio of 10:7.5:2.5 into 293T cells using calcium phosphate transfection. The virus particles were harvested 48, 72, and 96 h post transfection, filtered by 0.45 micrometers filter unit (Millipore) and then centrifuged at 18,000 r.p.m., 4 °C for 2 h. HPC7 cells were infected with recombinant lentivirus-transducing units in the presence of 4 μg/mL polybrene (Sigma). GFP+ cells were sorted 48 h after infection and cultured for experiments.
irCLIP-seq and data analysis
For irCLIP-seq, we modified the procedure from previously reported methods.44,76 In brief, irCLIP was performed on ~3 × 108 Flag-Ythdf2 HPC7 cells by UV crosslinking the cells at 0.4 J/cm2 with three treatments. Whole-cell lysates were generated in lysis buffer (150 mM KCl, 10 mM HEPES pH 7.6, 2 mM EDTA, 0.5% NP-40, 0.5 mM dithiothreitol (DTT), 1:100 protease inhibitor cocktail, 400 U/ml RNase inhibitor; 1 ml cell pellet and 2 ml lysis buffer). The lysates were pipetted up and down several times, and then incubated on ice for 5 mins and shock-frozen at −80 °C with liquid nitrogen. The mRNP lysate was thawed on ice and centrifuged at 15,000 g for 15 min to clear the lysate. Flag-Ythdf2 was isolated with 30 μl of protein-G magnetic beads per 1 ml lysate, previously bound to 2 μg of anti-Flag monoclonal antibody (Sigma) for 2 h at 4 °C on rotation. The beads were collected, washed eight times with 1 ml ice-cold NT2 buffer (200 mM NaCl, 50 mM HEPES pH 7.6, 2 mM EDTA, 0.05% NP-40, 0.5 mM DTT, 200 U/ml RNase inhibitor) and once with 200 μl irCLIP NT2 buffer (50 mM Tris, pH 7.5; 150 mM NaCl; 1 mM MgCl2; 0.0005% NP-40). mRNP complex was digested with RNase 1 (Thermo Fisher #AM2294) at 0.4 U/μl in irCLIP NT2 buffer (aqueous volume of 30 μl and supplemented with 6 μl of PEG400 (16.7% final)). The nuclease reaction was incubated at 30 °C for 15 min in an Eppendorf Thermomixer, 15 s 1400 r.p.m., 90 s rest. Nuclease digestions were stopped by addition of 0.5 mL of ice-cold high-stringency buffer (20 mM Tris, pH 7.5; 120 mM NaCl; 25 mM KCl; 5 mM EDTA; 1% Trition X-100; 1% Na-deoxycholate). Immunoprecipitates were then quickly rinsed with 0.25 mL then with 0.05 mL of ice-cold irCLIP NT2 buffer. The irCLIP adaptor ligation and library construction followed a previously reported protocol.44
Data were demultiplexed using FAST-iCLIP version 0.9.3 and aligned to mouse genome mm10 from UCSC using STAR (2.4.2a) with parameters “–outFilterScoreMinOverLread 0 –outFilterMatchNminOverLread 0 –outFilterMatchNmin 0”. RPM-normalized genome browser tracks were created in R (3.4.1) and plotted using the Gviz package (1.20.0). Enriched motifs were identified by taking midpoints of each binding site found in all three replicates, adding 20 bases up and downstream, and running MEME (4.11.1) with parameters “-dna -mod zoops -revcomp -minw 5 -maxw 10 -nmotifs 10 -maxsize 1000000”. After motifs were identified, we ran tomtom (4.11.1) against transfac (1–2017) to identify known binding sites. GO enrichment analysis was performed using a hypergeometric test in R. GO terms were considered enriched if they had a BH-adjusted p-value < 0.05. Selected terms of interest are shown in the bar plot. Bars in the bar plot indicate percentage of genes in the list being tested having the term divided by the percentage of genes in the genome having the term. Peaks found by FAST-iCLIP in all three replicates were assigned to various features in the genome. Promoters were defined as upstream 150 bases from the TSS. “trans_stop” was defined as upstream and downstream 200 bases from the transcript start site.
Cord blood transduction
Cord blood transduction was conducted as described previously.11 Briefly, fresh CD34+ cord blood cells or flow-sorted CD34+ CD38− cells were prestimulated for 12–18 h in StemSpan medium (StemCell Technologies) supplemented with growth factors interleukin 6 (IL-6; 20 ng/ml, Peprotech), SCF (100 ng/ml, Peprotech), Flt3 ligand (FLT3-L; 100 ng/ml, Peprotech), and thrombopoietin (TPO; 20 ng/ml, Peprotech). Lentiviruses were then added in the same medium at a MOI of 50–200 for 24 h. Cells were then allowed 2 days recovery after transduction before in vitro or in vivo assays. Human YTHDF2 was knocked down by shRNA targeting 5′-AAGGACGTTCCCAATAGCCAA-3′ near the N terminus of the coding region as in a previous report.30 Scramble shRNA (seed sequence 5′-GCGCGATAGCGCTAATAAT-3′) was used as control.
Clonogenic progenitor assays
Flow-sorted GFP+ cord blood cells from 10-day cultured transduced cells (12,000 per ml) were resuspended in semi-solid methylcellulose medium (Methocult H4034; StemCell Technologies). Colony counts were carried out after 14 days of incubation.
hUBC HSPC culture
Two days after transduction, human cord blood CD34+ or CD34+ CD38− cells were collected, and the percentage that were GFP+ was determined by flow cytometry. To ensure that equal numbers of GFP+ cells were cultured before expansion, we added identically cultured GFP− cells to the ones with a higher percentage GFP+ to match the proportions of GFP+ cells between control and human YTHDF2 KD. Then cells were seeded at a density of 105 per ml in StemSpan medium (StemCell Technologies) supplemented with growth factors IL-6 (20 ng/ml), SCF (100 ng/ml), FLT3-L (100 ng/ml), TPO (20 ng/ml), and CHIR99021 (250 nM) (Stemgent).75
Human HSC xenotransplantation
For ex vivo expansion analysis of human cord blood HSCs, 105 sorted CD34+ CD38− cells were transduced with human YTHDF2 shRNA or control shRNA for 3 days and then analyzed for transduction efficiency (%GFP−/+) and stem cell markers. On day 10, cultured cells were collected for stem cell marker analysis. For the hUCB HSC primary LDA assay, CD34+ cells were enriched as described above and transduced with human YTHDF2 shRNA or control shRNA at a MOI of 50. Media were changed 24 h post infection. Equal numbers of GFP+ cells were sorted from control or YTHDF2 KD cells 3 days post infection and cultured overnight. Three doses of sorted GFP+ cells, 50, 20, and 10 thousand, were transplanted into sub-lethally irradiated (3.25 Gy) NSG mice. The cut-off for HSC engraftment was the exhibition of more than 1% human CD45+ GFP+ cells in a total of CD45+ cells in BM of primary transplantation recipients. For the hUCB HSC secondary LDA assay, BM cells were collected from primary recipients that had received the highest two doses and mixed together at 10 weeks post transplantation. Three doses of BM cells, 1.2 × 107, 8 × 106, and 4 × 106, were transplanted into sub-lethally irradiated (3.25 Gy) NSG mice, respectively. The cut-off for HSC engraftment was an exhibition of more than 0.2% human CD45+ GFP+ cells out of total CD45+ cells in BM of secondary transplantation recipients. HSC frequency was assessed using ELDA software.73 For all human cord blood xenotransplantation experiments, female NSG mice aged 6–8 weeks were used.
m6A-seq data analysis
Human and mouse m6A-seq data were aligned to the hg19 and mm10 transcriptome. In order to identify m6A peaks, the hg19 and mm10 transcriptome was divided into 25 nucleotide-wide tiles. The number of reads in the m6A immunoprecipitation (IP) and non-IP (control) sample was counted in each tile, and a p-value was calculated using Fisher’s exact test and adjusted for multiple testing. Tiles with significant enrichment of the m6A signal (adjust-Pval < = 0.05) were merged into bigger regions. Regions smaller than 100 bp were discarded, and regions over 200 bp were divided into 100 to 200 bp sub-regions; the m6A signal to control was calculated for each region; and regions with at least a twofold enrichment in all replicates were identified as m6A peaks. The distributions of m6A peaks and m6A marked genes were determined by overlapping all m6A peaks with the hg19 and mm10 RefGene annotation. m6A marked genes were identified by overlapping m6A peaks with hg19 RefGene. To filter for transcription factors, genes marked by m6A in all three samples were compared against the human transcription factor database (http://fantom.gsc.riken.jp/5/sstar/Browse_Transcription_Factors_hg19). GO term analysis was then performed using R package enrich GO. m6A marked human transcription factors were used as the searching list, and all the expressed genes were used as background. Hemopoiesis related biological process terms with significant enrichment were used to generate Fig. 4d.
RNA sequencing
CD34+ cells from human cord blood were transduced with control or human YTHDF2 KD lentivirus and GFP+ CD34+ sorted 10 days later. Three replicates of 12,000 GFP+ CD34+ cells were sorted for each group and used to extract total RNA. Four nanograms of high-quality total RNA was used for cDNA synthesis and library preparation using th the SMART-Seq v4 Ultra Low Input RNA Kit for Sequencing (Takara, 634891) and Nextera XT (Illumina, FC-131-1096) according to the manufacturers’ directions. The resulting short fragment libraries were checked for quality and quantity using an Agilent 2100 Bioanalyzer and Invitrogen Qubit Fluorometer. Equimolar libraries were pooled, requantified, and sequenced as 75 base pair single reads on a High Output flow cell on the Illumina NextSeq 500 instrument. Following sequencing, Illumina Primary Analysis version NextSeq RTA 2.4.11 and Secondary Analysis version bcl2fastq2 2.18 were applied to demultiplex reads for all libraries and generate FASTQ files.
For RNA-seq analysis, reads were aligned to UCSC genome hg38 with Tophat version 2.0.13 with default parameters, using Ensembl 87 gene models. Read counts were generated using HTSeq-count with -m intersection-nonempty. Reads were also aligned to ERCC control sequences and counts tabulated. A scaling factor was calculated based on the median of the ERCC counts for each sample and used for normalization. Differentially expressed genes were found using the edgeR package (3.18.1) in R (3.4.1). Differentially expressed genes were required to have a BH-adjusted p-value < 0.05 and a twofold change in expression.
RNA stability assay
A total of 15,000 sorted LT-, ST-HSCs, and MPPs were cultured in StemSpan SFEM medium (Stem Cell Technologies) supplemented with 10 μg/mL heparin (Sigma), 0.5 × penicillin/streptomycin (Sigma), 10 ng/mL rm SCF (Biovision, Inc.), and 20 ng/mL Tpo (Cell Sciences, Inc.)75 at 37 °C in an atmosphere containing 5% CO2 5% O2. Sorted cells were treated with 5 μM actinomycin D (Sigma) for inhibition of mRNA transcription. Cells were collected at 0 h or 4 h post treatment, and total RNA was extracted and used for RNA-seq.
m6A RNA methylation quantification
Mouse BM Lineage negative cells from wt and Ythdf2 KO mice were enriched using the mouse Lineage Cell Depletion Kit (Miltenyi Biotec), followed by total RNA extraction with TRIzol (Invitrogen). Quantification of m6A RNA methylation in Lin- cells was performed using the m6A RNA Methylation Quantification Kit (Abcam ab185912) following the manufacturer’s protocol. 200 ng total RNA were used per replicates for either group.
qPCR analysis
LSK cells (105) were sorted from wt and Ythdf2 KO mice. Total RNA was extracted with TRIzol (Invitrogen). cDNA synthesis was carried out using the High-Capacity RNA-to-cDNA™ Kit (Thermo) following the manufacturer’s protocol. qPCR primers used are listed in Supplementary information, Table S5.
Western blot and intracellular staining
To validate the KO or KD efficiency in the Ythdf2 KO mouse model or hUCB, 33,000 cKit+ cells or 120,000 GFP+ cells were sorted from BM or transfected hUCB samples, respectively. HeLa cells transduced to overexpress human YTHDF2 were used validate overexpression efficiency as shown in Supplementary information, Figure S8b. Immunoblotting was performed with anti-YTHDF2 rabbit polyclonal antibody (MBL, RN123PW) and β-actin mouse monoclonal antibody (NOVUS, NB600-501). The secondary antibodies used were IRDye 800CW Goat anti-Mouse IgG and IRDye 800CW Goat anti-Rabbit IgG antibodies (LI-COR). For intracellular staining, BM cells from wt and Ythdf2 KO mice were stained with HSC markers as above, then fixed using the Cytofix/Cytoperm kit (BD Biosciences) according to the manufacturer’s instructions. Fixed and permeabilized cells were immunostained with anti-YTHDF2 antibody (MBL RN123PW), anti-TAL1 antibody (Santa Cruz sc-393287), anti-GATA2 antibody (Santa Cruz sc-267), anti-RUNX1 antibody (Santa Cruz sc-365644), anti-STAT5 antibody (Santa Cruz sc-74442), and detected by Alexa-488 donkey anti-rabbit IgG antibody (Invitrogen).
Single-cell immunostaining
LSKs (10,000) from wt and Ythdf2 KO mice were sorted onto Poly-l-lysine coating slides, which were placed in a moist chamber and incubated at 4 °C for 30 mins to allow cells to settle onto the slides. Cells were fixed with chilled methanol at RT for 10 mins, blocked with universal blocking reagent (BioGenex) at RT for 30 min, and stained with mouse TAL1 antibody (Santa Cruz, SC393287) or mouse IgG control (Abcam) at 4 °C overnight. Cells were then stained with Alexa Fluor 488 donkey anti-mouse IgG (Thermo Fisher Scientific) at 4 °C for 30 min. Images were captured on a PerkinElmer Ultraview spinning disk system with a Yokagawa CS-X1 disk. Images were collected with a C9100-23 Hamamatsu EM-CCD using Velocity software (PerkinElmer). For Z-stacks, the step size was set at 400 nm. Staining intensity was quantified by ImageJ program.
FISH in conjugation with fluorescent immunostaining
Sorted LSKs were spun onto microscope glass slide (Fisher Scientific catalog number 12-544-4) using a Cytospin™ 4 Cytocentrifuge at 800 r.p.m. for 1 min with medium acceleration (Thermo Scientific, catalog number A78300003), followed by an immediate immersion into 4% PFA (diluted from 16% (wt/vol) aqueous solution, Electron Microscopy Sciences, catalog number 15710). Cells were fixed at RT (25 ± 2 °C) for 30 min. RNA in situ hybridization was performed using RNAscope multiplex fluorescent detection kit according to the manufacturer’s instructions (Advanced Cell Diagnostics) with a couple of modifications: antigen retrieval was unnecessary and digestion was performed with a 1:15 dilution of proteinase III solution for 10 mins at RT. RNAscope probes targeting mouse Tal1 and Gata2 were designed and produced by ACDbio. After the in situ hybridization was completed, slides were rinsed twice with phosphate buffered saline with Tween-20 (PBST) and directly processed with background blocking (Background buster solution, Innovex, catalog number NB306) and primary antibody incubation. Anti-YTHDF2 (MBL, 1: 500) and anti-DCP1a (Santa Cruz, SC100706, 1:200) antibodies were diluted with antibody diluent reagent buffer (Life Technologies, catalog number 003118) and incubated with the specimens at 4 °C overnight. Donkey anti- rabbit Alexa Fluor 488 (Invitrogen, 1:500) and donkey anti-mouse Alexa Fluor 633 (Invitrogen, 1:500) were used for protein target multiplexing.
Statistical analysis
Statistical analyses were carried out using Student’s t-test. The results are shown as the mean ± SEM.
Data availability
All sequencing data, including the m6A-seq, irCLIP-seq, and RNA-seq datasets, are available through the Gene Expression Omnibus under accession GSE107957.
Original Data Repository at http://www.stowers.org/research/publications/LIBPB-1248.
Change history
27 August 2018
In the initial published version of this article, there was an inadvertent omission from the Acknowledgements that this work was supported by Stowers Institute for Medical Research (SIMR-1004) and NIH National Cancer Institute grant to University of Kansas Cancer Center (P30 CA168524). This omission does not affect the description of the results or the conclusions of this work.
References
Li, L. & Clevers, H. Coexistence of quiescent and active adult stem cells in mammals. Science 327, 542–545 (2010).
Weissman, I. L. Stem cells: units of development, units of regeneration, and units in evolution. Cell 100, 157–168 (2000).
Walasek, M. A., van Os, R. & de Haan, G. Hematopoietic stem cell expansion: challenges and opportunities. Ann. N. Y. Acad. Sci. 1266, 138–150 (2012).
Sung, A. D. & Chao, N. J. Concise review: acute graft-versus-host disease: immunobiology, prevention, and treatment. Stem Cells Transl. Med. 2, 25–32 (2013).
Shlomchik, W. D. Graft-versus-host disease. Nat. Rev. Immunol. 7, 340–352 (2007).
Huang, X. et al. Activation of OCT4 enhances ex vivo expansion of human cord blood hematopoietic stem and progenitor cells by regulating HOXB4 expression. Leukemia 30, 144–153 (2016).
Boitano, A. E. et al. Aryl hydrocarbon receptor antagonists promote the expansion of human hematopoietic stem cells. Science 329, 1345–1348 (2010).
Fares, I. et al. Cord blood expansion. Pyrimidoindole derivatives are agonists of human hematopoietic stem cell self-renewal. Science (New Y., N. Y.) 345, 1509–1512 (2014).
Amsellem, S. et al. Ex vivo expansion of human hematopoietic stem cells by direct delivery of the HOXB4 homeoprotein. Nat. Med. 9, 1423–1427 (2003).
Antonchuk, J., Sauvageau, G. & Humphries, R. K. HOXB4-induced expansion of adult hematopoietic stem cells ex vivo. Cell 109, 39–45 (2002).
Rentas, S. et al. Musashi-2 attenuates AHR signalling to expand human haematopoietic stem cells. Nature 532, 508–511 (2016).
Himburg, H. A. et al. Pleiotrophin regulates the expansion and regeneration of hematopoietic stem cells. Nat. Med. 16, 475–482 (2010).
North, T. E. et al. Prostaglandin E2 regulates vertebrate haematopoietic stem cell homeostasis. Nature 447, 1007–1011 (2007).
Varnum-Finney, B. et al. Pluripotent, cytokine-dependent, hematopoietic stem cells are immortalized by constitutive Notch1 signaling. Nat. Med. 6, 1278–1281 (2000).
Chou, S., Flygare, J. & Lodish, H. F. Fetal hepatic progenitors support long-term expansion of hematopoietic stem cells. Exp. Hematol. 41, 479–490 e474 (2013).
Guo, B., Huang, X., Lee, M. R., Lee, S. A. & Broxmeyer, H. E. Antagonism of PPAR-gamma signaling expands human hematopoietic stem and progenitor cells by enhancing glycolysis. Nat. Med. 24, 360–367 (2018).
Zhao, B. S., Roundtree, I. A. & He, C. Post-transcriptional gene regulation by mRNA modifications. Nat. Rev. Mol. Cell Biol. 18, 31–42 (2017).
Roundtree, I. A., Evans, M. E., Pan, T. & He, C. Dynamic RNA Modifications in Gene Expression Regulation. Cell 169, 1187–1200 (2017).
Li, S. & Mason, C. E. The pivotal regulatory landscape of RNA modifications. Annu. Rev. Genom. Hum. Genet. 15, 127–150 (2014).
Batista, P. J. et al. m(6)A RNA modification controls cell fate transition in mammalian embryonic stem cells. Cell. Stem. Cell. 15, 707–719 (2014).
Geula, S. et al. Stem cells. m6A mRNA methylation facilitates resolution of naive pluripotency toward differentiation. Science 347, 1002–1006 (2015).
Yoon, K. J. et al. Temporal control of mammalian cortical neurogenesis by m6A methylation. Cell 171, 877–889 (2017).
Zhang, C. et al. m6A modulates haematopoietic stem and progenitor cell specification. Nature 549, 273–276 (2017).
Zhao, B. S. et al. m6A-dependent maternal mRNA clearance facilitates zebrafish maternal-to-zygotic transition. Nature 542, 475–478 (2017).
Lv, J. et al. Endothelial-specific m(6)A modulates mouse hematopoietic stem and progenitor cell development via Notch signaling. Cell Res. 28, 249–252 (2018).
Li, Z. et al. FTO plays an oncogenic role in acute myeloid leukemia as a N6-methyladenosine RNA demethylase. Cancer Cell. 31, 127–141 (2017).
Vu, L. P. et al. The N6-methyladenosine (m6A)-forming enzyme METTL3 controls myeloid differentiation of normal hematopoietic and leukemia cells. Nat. Med. 23, 1369–1376 (2017).
Weng, H. et al. METTL14 inhibits hematopoietic stem/progenitor differentiation and promotes leukemogenesis via mRNA m(6)A modification. Cell. Stem. Cell. 22, 191–205 e199 (2018).
Barbieri, I. et al. Promoter-bound METTL3 maintains myeloid leukaemia by m(6)A-dependent translation control. Nature 552, 126–131 (2017).
Wang, X. et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–120 (2014).
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3’ UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Schwartz, S. et al. High-resolution mapping reveals a conserved, widespread, dynamic mRNA methylation program in yeast meiosis. Cell 155, 1409–1421 (2013).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Wang, Z., Li, G., Tse, W. & Bunting, K. D. Conditional deletion of STAT5 in adult mouse hematopoietic stem cells causes loss of quiescence and permits efficient nonablative stem cell replacement. Blood 113, 4856–4865 (2009).
Ebina, W. & Rossi, D. J. Transcription factor-mediated reprogramming toward hematopoietic stem cells. EMBO J. 34, 694–709 (2015).
Orkin, S. H. & Zon, L. I. Hematopoiesis: an evolving paradigm for stem cell biology. Cell 132, 631–644 (2008).
de Pater, E. et al. Gata2 is required for HSC generation and survival. J. Exp. Med. 210, 2843–2850 (2013).
Hock, H. et al. Tel/Etv6 is an essential and selective regulator of adult hematopoietic stem cell survival. Genes & Dev. 18, 2336–2341 (2004).
Lim, K. C. et al. Conditional Gata2 inactivation results in HSC loss and lymphatic mispatterning. J. Clin. Invest. 122, 3705–3717 (2012).
Reynaud, D. et al. SCL/TAL1 expression level regulates human hematopoietic stem cell self-renewal and engraftment. Blood 106, 2318–2328 (2005).
Kato, Y. et al. Selective activation of STAT5 unveils its role in stem cell self-renewal in normal and leukemic hematopoiesis. J. Exp. Med. 202, 169–179 (2005).
Li, H. B. et al. m6A mRNA methylation controls T cell homeostasis by targeting the IL-7/STAT5/SOCS pathways. Nature 548, 338–342 (2017).
Pinto do, O. P., Kolterud, A. & Carlsson, L. Expression of the LIM-homeobox gene LH2 generates immortalized steel factor-dependent multipotent hematopoietic precursors. EMBO J. 17, 5744–5756 (1998).
Zarnegar, B. J. et al. irCLIP platform for efficient characterization of protein-RNA interactions. Nat. Methods 13, 489–492 (2016).
Sheth, U. & Parker, R. Decapping and decay of messenger RNA occur in cytoplasmic processing bodies. Science 300, 805–808 (2003).
Kedersha, N. & Anderson, P. Mammalian stress granules and processing bodies. Methods Enzymol. 431, 61–81 (2007).
Lacombe, J. et al. Scl regulates the quiescence and the long-term competence of hematopoietic stem cells. Blood 115, 792–803 (2010).
Galan-Caridad, J. M. et al. Zfx controls the self-renewal of embryonic and hematopoietic stem cells. Cell 129, 345–357 (2007).
Zheng, G. et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol. Cell 49, 18–29 (2013).
Zhou, J. et al. Dynamic m(6)A mRNA methylation directs translational control of heat shock response. Nature 526, 591–594 (2015).
Alarcon, C. R. et al. HNRNPA2B1 is a mediator of m(6)A-dependent nuclear RNA processing events. Cell 162, 1299–1308 (2015).
Zhang, S. et al. m6A demethylase ALKBH5 maintains tumorigenicity of glioblastoma stem-like cells by sustaining FOXM1 expression and cell proliferation program. Cancer Cell 31, 591–606 e596 (2017).
Lence, T. et al. m6A modulates neuronal functions and sex determination in Drosophila. Nature 540, 242–247 (2016).
Haussmann, I. U. et al. m6A potentiates Sxl alternative pre-mRNA splicing for robust Drosophila sex determination. Nature 540, 301–304 (2016).
Chen, T. et al. m(6)A RNA methylation is regulated by microRNAs and promotes reprogramming to pluripotency. Cell. Stem Cell. 16, 289–301 (2015).
Alarcon, C. R., Lee, H., Goodarzi, H., Halberg, N. & Tavazoie, S. F. N6-methyladenosine marks primary microRNAs for processing. Nature 519, 482–485 (2015).
Xiao, W. et al. Nuclear m(6)A Reader YTHDC1 Regulates mRNA Splicing. Mol. Cell 61, 507–519 (2016).
Wojtas, M. N. et al. Regulation of m6A transcripts by the 3’– > 5’ RNA helicase YTHDC2 is essential for a successful meiotic program in the mammalian germline. Mol. Cell 68, 374–387 e312 (2017).
Ivanova, I. et al. The RNA m6A reader YTHDF2 is essential for the post-transcriptional regulation of the maternal transcriptome and oocyte competence. Mol. Cell 67, 1059–1067 e1054 (2017).
Fustin, J. M. et al. RNA-methylation-dependent RNA processing controls the speed of the circadian clock. Cell 155, 793–806 (2013).
Slobodin, B. et al. Transcription impacts the efficiency of mRNA translation via co-transcriptional N6-adenosine methylation. Cell 169, 326–337 e312 (2017).
Schwartz, S. et al. Transcriptome-wide mapping reveals widespread dynamic-regulated pseudouridylation of ncRNA and mRNA. Cell 159, 148–162 (2014).
Pendleton, K. E. et al. The U6 snRNA m6A methyltransferase METTL16 regulates SAM synthetase intron retention. Cell 169, 824–835 e814 (2017).
Shi, H. et al. YTHDF3 facilitates translation and decay of N6-methyladenosine-modified RNA. Cell Res. 27, 315–328 (2017).
Huang, H. et al. Recognition of RNA N(6)-methyladenosine by IGF2BP proteins enhances mRNA stability and translation. Nat. Cell Biol. 20, 285–295 (2018).
Bertero, A. et al. The SMAD2/3 interactome reveals that TGFbeta controls m(6)A mRNA methylation in pluripotency. Nature 555, 256–259 (2018).
Liu, N. et al. N(6)-methyladenosine-dependent RNA structural switches regulate RNA-protein interactions. Nature 518, 560–564 (2015).
Joseph, C. et al. Deciphering hematopoietic stem cells in their niches: a critical appraisal of genetic models, lineage tracing, and imaging strategies. Cell. Stem. Cell. 13, 520–533 (2013).
Barbieri, I. et al. Promoter-bound METTL3 maintains myeloid leukaemia by m(6)A-dependent translation control. Nature 552, 126–131 (2017).
He, S., Nakada, D. & Morrison, S. J. Mechanisms of stem cell self-renewal. Annu. Rev. Cell. Dev. Biol. 25, 377–406 (2009).
Qian, P. et al. The Dlk1-Gtl2 locus preserves LT-HSC function by inhibiting the PI3K-mTOR pathway to restrict mitochondrial metabolism. Cell. Stem Cell. 18, 214–228 (2016).
He, X. C. et al. Homing and migration assays of hematopoietic stem/progenitor cells. Methods Mol. Biol. 1185, 279–284 (2014).
Hu, Y. & Smyth, G. K. ELDA: extreme limiting dilution analysis for comparing depleted and enriched populations in stem cell and other assays. J. Immunol. Methods 347, 70–78 (2009).
Purton, L. E. & Scadden, D. T. Limiting factors in murine hematopoietic stem cell assays. Cell. Stem Cell. 1, 263–270 (2007).
Perry, J. M. et al. Cooperation between both Wnt/{beta}-catenin and PTEN/PI3K/Akt signaling promotes primitive hematopoietic stem cell self-renewal and expansion. Genes Dev. 25, 1928–1942 (2011).
Simsek, D. et al. The mammalian Ribo-interactome reveals ribosome functional diversity and heterogeneity. Cell 169, 1051–1065 e1018 (2017).
Acknowledgements
We thank K. Tannen for proofreading and editing, and M. Hembree, H. Marshall, K. Zapien, D. Dukes, B. Lewis, A. Moran, M. Durnin, J. McCann, A. Box, J. Park, L. Holmes, J. Blank, and D. DeGraffenreid for technical support. We are grateful to X. Zhuang at University of Chicago for transferring the mice.
Author information
Authors and Affiliations
Contributions
Z.L. and P.Q. conceived the project, performed experiments, analyzed data, and wrote the manuscript. Z.L. conducted m6A-seq and irCLIP-seq. W.S. and J.Z. analyzed the m6A-seq and mRNA stability data. M.G., S.C., and H.L. analyzed irCLIP-seq and RNA-seq data. X.C.H. provided the training and performed the experiments. B.S., M.Q., Y.Z., and X.Z. generated the genetic mouse model. Z.Y., Y.W., J.P., F.T., K.Z., D.H., Y.H., C.Z., and H.X. performed experimental work. R.A., A.P., K.H., M.P., A.P., J.H., and T.P. provided technical assistance. S.H. and C.H. provided key suggestions and plasmids. L.L. supervised the study.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Rights and permissions
About this article
Cite this article
Li, Z., Qian, P., Shao, W. et al. Suppression of m6A reader Ythdf2 promotes hematopoietic stem cell expansion. Cell Res 28, 904–917 (2018). https://doi.org/10.1038/s41422-018-0072-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41422-018-0072-0
Keywords
This article is cited by
-
The role of the methyltransferase METTL3 in prostate cancer: a potential therapeutic target
BMC Cancer (2024)
-
Microplastics dampen the self-renewal of hematopoietic stem cells by disrupting the gut microbiota-hypoxanthine-Wnt axis
Cell Discovery (2024)
-
A Mettl16/m6A/mybl2b/Igf2bp1 axis ensures cell cycle progression of embryonic hematopoietic stem and progenitor cells
The EMBO Journal (2024)
-
Regulation of adult stem cell quiescence and its functions in the maintenance of tissue integrity
Nature Reviews Molecular Cell Biology (2023)
-
Transient regulation of RNA methylation in human hematopoietic stem cells promotes their homing and engraftment
Leukemia (2023)