Abstract
Maintenance of immunological homeostasis between tolerance and autoimmunity is essential for the prevention of human diseases ranging from autoimmune disease to cancer. Accumulating evidence suggests that p53 can mitigate phagocytosis-induced adjuvanticity thereby promoting immunological tolerance following programmed cell death. Here we identify Inhibitor of Apoptosis Stimulating p53 Protein (iASPP), a negative regulator of p53 transcriptional activity, as a regulator of immunological tolerance. iASPP-deficiency promoted lung adenocarcinoma and pancreatic cancer tumorigenesis, while iASPP-deficient mice were less susceptible to autoimmune disease. Immune responses to iASPP-deficient tumors exhibited hallmarks of immunosuppression, including activated regulatory T cells and exhausted CD8+ T cells. Interestingly, iASPP-deficient tumor cells and tumor-infiltrating myeloid cells, CD4+, and γδ T cells expressed elevated levels of PD-1H, a recently identified transcriptional target of p53 that promotes tolerogenic phagocytosis. Identification of an iASPP/p53 axis of immune homeostasis provides a therapeutic opportunity for both autoimmune disease and cancer.
Similar content being viewed by others
Introduction
One of the most fundamental functions of the immune system is the clearance of dead cells following normal tissue turnover, malignancy, injury, or infection. In this capacity, the immune system exhibits remarkable capability to discriminate whether to respond in an immunogenic or tolerogenic manner depending on the nature of cell death [1]. The daily homeostatic turnover of billions of cells rarely invokes an immune response, while the corpses of relatively few infected or malignant cells can elicit robust antigen-specific immune responses and immunological memory. Defects in this dichotomy can lead to human disease. For example, mistaken immunogenic phagocytosis in combination with epitope spreading contributes to autoimmune disease, while tumors can evade the immune system by promoting tolerogenic phagocytosis [2, 3]. Fundamentally, adaptive immunogenicity requires simultaneous antigenicity and adjuvanticity, and although cell death and subsequent phagocytosis almost universally promote antigen presentation, cell death can elicit varying adjuvanticity.
The nature of the immune response to cell death, whether immunogenic or tolerogenic, is relayed from the innate to adaptive immune system by manipulation of co-stimulation and cytokine-mediated adjuvanticity following phagocytosis [4]. Apoptosis or programmed cell death is typically immunologically silent while necrosis stimulates inflammation [5]. For example, phagocytosis of apoptotic cells can generate anergic cytotoxic T lymphocytes due to insufficient co-stimulation, stimulate immune checkpoint pathways, promote immunosuppressive cytokine release, and induce systemic antigenic tolerance [6,7,8,9]. In contrast, inflammatory cell death such as necrosis, pyroptosis, or ferroptosis can induce inflammation via recognition of pathogen-associated molecular patterns, damage-associated molecular patterns (DAMPs), inflammasome activation, release of pro-inflammatory cytokines, and licensing CD4+ T cell help of cytotoxic T cell development [1, 10,11,12]. The tissue context of the responding antigen-presenting cell (APC) might also influence the immunological consequences of cell death, as mature MHCIIhiCD80hi dendritic cells (DCs) are necessary to mount effective immune responses [13], while immature DCs are tolerogenic [14]. Ferroptosis of tumor neutrophil myeloid-derived suppressor cells can be immunosuppressive and promote tumor growth in vivo [15]. Therefore, the identity, location, activation status, and life history of the dying cell as well as those of the cell engulfing it all influence the nature of the immune response to cell death.
Physiologic apoptosis and unnatural cell death elicit fundamentally different immune responses, yet the pathways intrinsic to dying cells responsible for modulating immune homeostasis and phagocytic response remain poorly characterized. However, master regulators of cell death, such as p53, may play key roles in the establishment of an inherently immune activating or tolerizing transcriptional program [16, 17]. P53 potently induces apoptosis or cellular senescence following genotoxic stress by transactivating and trans-repressing hundreds of target genes, including genes involved in immune regulation [18,19,20]. Dominant negative and missense mutations in TP53 or its effectors Bim, p21, Gadd45a, and Fas are associated with an increasing number of autoimmune diseases [21,22,23,24,25,26,27,28,29,30,31,32], demonstrating the importance of p53 in maintaining immune tolerance. Expression of the p53 target Fas by apoptotic cells and subsequent Fas-Fas ligand (FasL) signaling in responding lymphocytes is necessary for APC release of the immunosuppressive cytokines TGF-β and IL-10, and maintenance of immune privilege and immune tolerance [33,34,35,36,37,38]. More recent studies have identified novel roles of p53 in regulation of both innate and adaptive immune responses with potential non-cell-autonomous influences on tumor immunology [39, 40], including upregulation of PD-L1 and PD-1H [17, 41] and suppression of pro-inflammatory cytokine secretion [42, 43]. Although recent evidence suggests that dying cells communicate with the immune system via p53, the mechanisms by which p53 and its regulators influence immunological tolerance remain largely unknown.
One of the most ancient regulators of p53 is the Ankyrin repeat domain, SH3 domain and Proline rich sequence containing Protein (ASPP) family of proteins [44]. Inhibitor of Apoptosis Stimulating p53 Protein (iASPP) is an ASPP family member that binds and inhibits the transcriptional activity of p53 [45] and its family members p63 and p73 [46]. iASPP was also identified as a binding partner and inhibitor of NF-κB subunit p65/RelA in cardiomyocytes and endothelium [47, 48]. Mice deficient for iASPP develop dilated cardiomyopathy with features of arrhythmogenic right ventricular cardiomyopathy (ARVC) with fibrosis and extensive immune infiltration [49]. Cardiomyocyte-specific and keratinocyte-specific iASPP-deficient mice develop ARVC or impaired cell adhesion and wound healing, respectively [50]. Although tissue-specific iASPP deletion largely phenocopies germline iASPP deletion, keratinocyte-specific iASPP-deficient mice have reduced cutaneous phenotype penetrance and severity, suggesting a possible non-cell autonomous role of iASPP. Disruption of immune homeostasis contributes to delayed wound healing [51,52,53,54] and increased inflammation is associated with more severe ARVC [55], suggesting that lymphocytes might provide this non-cell autonomous effect.
Given the recent implication of p53 in immune homeostasis, we hypothesized that iASPP regulates p53-mediated induction of immune tolerance by fundamentally altering the transcriptional profiles of dying cells, communicating the ultimate nature of cell death, whether physiologic or unnatural, to the responding immune system. We recently used genome-wide RNA-seq and p53 ChIP-seq analyses to discover iASPP regulation of a subset of p53 target genes involved in apoptosis and senescence [56]. Crystal structure analysis of the iASPP/p53 complex revealed that, unlike all other known cellular binding partners of p53, iASPP alters p53 target selectivity by binding to its L1 loop, which forms contacts with nucleotides defined by the iASPP/p53 sequence-specific signature. Here we demonstrate that perturbation of this iASPP/p53 complex in vivo disrupts the equilibrium between immunogenic versus tolerogenic cell death, with profound cell-intrinsic and non-cell-autonomous effects on the development of autoimmune disease and cancer.
Results
Experimental autoimmune encephalomyelitis is less severe in iASPP-deficient mice
To investigate the role of iASPP in immune homeostasis, we first characterized the peripheral immune system in iASPP−/− mice [46] in the absence of immune challenge. iASPP−/− mice had an increased proportion of CD4+FoxP3+ Treg cells in the colonic lamina propria, inguinal lymph node, and liver (Fig. S1A–C) and peripheral CD4+ T cell populations in iASPP−/− mice expressed elevated levels of PD-1 (Fig. S1D), suggesting that iASPP deficiency promotes peripheral T cell suppression and PD-1 expression. No increase in Treg frequency was observed in the thymus, suggesting that iASPP doest not affect the development of natural Treg (nTreg) cells. Interestingly, CD4+ and γδ T cells in the livers of aged iASPP−/− mice expressed elevated levels of PD-1H (Fig. S1E), an immunosuppressive transcriptional target of p53 [17]. Given the increased proportion of Treg cells observed in the periphery of iASPP−/− mice, we examined whether iASPP deficiency promotes Treg cell differentiation. Indeed, p53 induces Treg cell differentiation and FoxP3 transcription by directly binding its promoter [57, 58]. Naive CD4+CD44−CD62L+ (Tnaive) cells were purified and polarized toward Treg or effector CD4+ subpopulations in vitro using Th1, Th2, Th17, or Treg permissive conditions. iASPP deficiency decreased Tnaive polarization to all subpopulations in vitro (Fig. S1F), suggesting that iASPP might indiscriminately modulate CD4+ polarization and differentiation. These findings suggest that the increased frequency of peripheral Treg cells observed in iASPP−/− mice arise through peripheral mechanisms of immune tolerance rather than dysregulated CD4+ differentiation.
To determine functional consequences of iASPP-deficiency, we examined the proliferative and suppressive capacity of iASPP-deficient immune cells in vitro. iASPP−/− CD4+ T cells had a decrease (p = 0.029) in proliferation measured by CFSE dilution in response to anti-CD3 (Fig. 1A) while expressing greater PD-1 (p = 0.015, Fig. 1B) without any decrease in viability (Fig. 1C), suggesting that iASPP deficiency limits CD4+ T cell proliferation in response to polyclonal TCR stimulation. Due to the apparent increased frequency of Treg cells in iASPP−/− mice, we examined the suppressive activity of iASPP−/− Treg cells in vitro by co-culturing CD4+CD25− conventional T cells (Tconv) and CD4+CD25+ Treg cells purified from healthy WT or iASPP−/− mice. iASPP−/− Treg cells exhibited greater suppressive capacity measured by decreased CFSE dilution (Fig. 1D), suggesting that iASPP expression in Treg cells limits their suppressive potential. Together, these data suggest that cell-intrinsic iASPP expression may regulate CD4+ T cell proliferation and Treg cell immunosuppression but not development. Increased Treg cell frequency and CD4 T cell expression of PD-1H suggests that iASPP might regulate immune homeostasis through similar mechanisms as p53.
A Flow cytometry dot plots (left) and quantification (right) of CD4+ T cell division 72 h after anti-CD3 stimulation analyzed by CFSE dilution. B Flow cytometric quantification of PD-1 expression in CD4+ T cells in (A). MFI median fluorescent intensity. C Flow cytometry contour plots of live-dead stain of CD4+ T cells 72 h after anti-CD3 stimulation. D Flow cytometric quantification of suppressive activity of Treg cells examined by cellular division of responder Tconv cells as measured by CFSE dilution 72 h after co-culture.
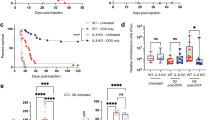
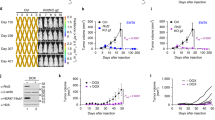
Given the influence of iASPP on tissue-resident Treg counts, we hypothesized that iASPP−/− mice would develop less severe autoimmune disease. To test this possibility, we utilized the myelin oligodendrocyte glycoprotein (MOG)-induced experimental autoimmune encephalomyelitis (EAE) model of T cell-mediated autoimmune demyelinating neuroinflammation (Fig. 2A). EAE was induced in WT (n = 7) or iASPP−/− (n = 10) mice and clinical severity was measured until mice reached a clinical score of 3 (paralysis of both hind limbs), at which point mice were euthanized. Both WT and iASPP−/− mice developed characteristic clinical features of EAE such as limp tail followed by limb paralysis and exhibited the expected relapsing-remitting disease course including initial attack and partial recovery after the peak phase of disease severity. However, iASPP−/− mice had delayed disease onset (p = 0.015), decreased duration of disease (p = 2.2 × 10–3), and decreased overall severity of EAE (p = 3.9 × 10–3) (Figs. 2B, S2A, B). Histological examination of CNS pathology 30 days after EAE induction revealed that iASPP−/− mice had less extensive demyelination measured by Luxol fast blue stain (Figs. 2C, S2C). iASPP−/− mice also exhibited a decreased number of infiltrating T cells in the cerebellum (Figs. 2C, S2D). Therefore, iASPP deficiency appears to decrease CNS autoimmune disease severity.
A Schematic of experimental autoimmune encephalomyelitis (EAE) model and experimental design. Mice were monitored until 30 days after immunization with MOG/CFA or until reaching primary endpoint of bilateral hind limb paralysis (clinical severity score = 3). MOG, 200 μg MOG35-55 peptide (MEVGWYRSPFSRVVHLYRNGK); CFA, Complete Freund’s Adjuvant supplemented with 400 μg H37Ra mycobacterium tuberculosis; PTx, 300 ng pertussis toxin. B Clinical course of MOG-induced EAE in WT (n = 7) or iASPP−/− (n = 10) mice. AUC, area under the curve. C Luxol fast blue (LFB), HE, and CD3 immunohistochemistry stains of corpus callosum, cerebellum, and spinal cord isolated from WT (n = 4) or iASPP−/− (n = 6) mice 30 days after immunization with MOG/CFA. Arrows indicate areas of demyelination. Scale, 1 mm (top) and 100 μm (bottom).
Given the decrease in autoimmune neuroinflammation observed in iASPP−/− mice, we analyzed potential sources of immunosuppression by performing flow cytometry on perfused whole brain hemispheres isolated from EAE mice 30 days after immunization. iASPP−/− mice with EAE had >6-fold decrease (p = 0.03) in mean numbers of CNS-infiltrating CD4+ T cells (Fig. S2E). Functionality of the endogenous autoimmune T cell response in EAE was examined by measuring splenocyte proliferation in response to challenge with immunogen. Splenocytes isolated from EAE mice at peak disease severity (clinical score = 3) were stimulated with MOG35-55 peptide and extent of proliferation was measured using CFSE dilution. In concordance with the observed clinical disease course and flow cytometry immunophenotyping of CNS-infiltrating lymphocytes, splenocytes from iASPP−/− mice with EAE were less responsive to in vitro immunogen challenge (Fig. S2F) and expressed increased amounts of PD-1H (Fig. S2G). Therefore, it appears that iASPP−/− mice with EAE have a less potent autoimmune antigen-specific T cell response, suggesting that iASPP predisposes mice to more severe CNS autoimmune disease by fostering CD4+ T cell-mediated neuroinflammation.
Pancreatic iASPP deficiency promotes an immunosuppressive tumor microenvironment
Given the stimulatory effect of iASPP on CD4+ T cell autoimmunity and that iASPP can have varying functions in different tissues [50], we next examined whether epithelial cell-specific disruption of the iASPP/p53 complex could influence immunological tolerance in a non-cell autonomous manner by studying consequences of iASPP deletion in oncogenic Kras driven pancreatic neoplasias. As iASPP modulates p53 target selectivity [56] and p53 is tumor suppressive, we expected iASPP deletion to delay tumor development. However, tumor suppressor genes such as p53 and tumor immunosurveillance represent independent safeguards against cancer development whose relative importance remains unclear [59,60,61,62]. Although KrasLSL−G12D/+;Trp53LSL−R172H/+;Pdx1-Cre (KPC) mice developed the most accelerated progression to PDAC, KrasLSL−G12D/+;iASPPfl/fl;Pdx1-Cre (KC;iASPPΔ8/Δ8) mice paradoxically had advanced pancreatic neoplasia development compared to KrasLSL−G12D/+;Pdx1-Cre (KC) mice (Fig. 3A). While most KC neoplasias exhibited non-invasive epithelial proliferation and minimal cellular abnormalities after 25 weeks, most KC;iASPPΔ8/Δ8 pancreata harbored sites of acinar to ductal metaplasia (ADM) and pancreatic intraepithelial neoplasia (PanIN) (Fig. 3B), indicative of more advanced disease. KC;iASPPΔ8/Δ8 pancreatic neoplasias had increased infiltration of Treg cells (Fig. 3C) and CD8 cells, but not CD4 cells (Fig. S3), suggesting that as in autoimmune disease, iASPP deficiency promotes Treg cell infiltration over conventional CD4 T cell infiltration in pancreatic neoplasia. Importantly, KC;iASPPΔ8/Δ8 mice had greater extents of intratumoral Treg cell infiltration than KC mice in early ADM lesions (Fig. 3D), demonstrating that increased presence of Treg cells is dependent on loss of iASPP and not simply on more advanced pancreatic cancer. While increased Treg cell infiltration was observed in KC;iASPPΔ8/Δ8 neoplasias and pre-cancerous lesions, sparse lymphocytic infiltration was observed in PDAC (Figs. 3D and S3), likely reflecting exclusion of lymphocytes from advanced pancreatic cancer [63]. Pancreatic CD4+ and CD8+ T cells exhibited a striking degree of exhaustion in KC;iASPPΔ8/Δ8 mice, including increased expression of PD-1 and Tim-3 (Fig. S3C), indicative of an exhausted T cell response in KC;iASPPΔ8/Δ8 mice. Together, these data suggest that advanced pancreatic neoplasia development observed in KC;iASPPΔ8/Δ8 mice is likely due to the emergence of an immunosuppressive tumor microenvironment that suppresses CD8+ anti-tumor cytotoxicity.
A Representative HE stains of pancreas isolated from 25-week-old KrasLSL−G12D/+;Pdx1-Cre (KC, n = 5), and KrasLSL−G12D/+;iASPPfl/fl;Pdx1-Cre (KC;iASPPΔ8/Δ8, n = 6), and KrasLSL−G12D/+;Trp53LSL−R172H/+;Pdx1-Cre (KPC, n = 5) mice. Higher magnification shown below. Scale, 200 μm (top) and 50 μm (bottom). B Proportion of reactive neoplasia, metaplasia, PanIN, versus PDAC of all pancreatic lesions in KC (n = 8, blue), KC;iASPPΔ8/Δ8 (n = 5, red), or KPC (n = 4, brown) mice. Data are sample proportion ± standard error of the sample proportion; Fisher’s exact test, p < 0.05 indicated on graph. C FoxP3 stains of pancreata isolated from KC (n = 5), KC;iASPPΔ8/Δ8 (n = 5), or KPC (n = 5) mice. Arrows indicate FoxP3+ cells. Scale, 100 μm. ADM acinar to ductal metaplasia, PanIN pancreatic intraepithelial neoplasia, PDAC pancreatic ductal adenocarcinoma. D Column scatter plot shows quantification of pancreatic neoplasia-infiltrating FoxP3+ Treg cells per area.
To test whether iASPP deficient oncogenic Kras-driven pancreatic cancer maintains an immunosuppressive microenvironment in the presence of a canonical inflammatory insult [64], we subjected KC;iASPPΔ8/Δ8 mice to caerulein-induced chronic pancreatitis (Fig. 4A). Chronic pancreatitis resulted in substantial fibrosis and increased infiltration of T cells into the pancreas, and deletion of iASPP exhibited an even greater degree of reactive stroma (Fig. 4B) and Treg infiltration (Fig. 4C) following caerulein insult. KC;iASPPΔ8/Δ8 pancreas-infiltrating DCs and macrophages expressed substantially increased amounts of MHCII and PD-1 (Fig. S4), and proportions of monocytic myeloid-derived suppressor cell (MDSC-M) and polymorphonuclear MDSC (MDSC-P) subpopulations were increased ~2-fold in KC;iASPPΔ8/Δ8 mice (Fig. 4D), indicative of the emergence of a potentially immunosuppressive population of APCs with iASPP deletion. Although KC;iASPPΔ8/Δ8 pancreas-infiltrating CD4+ T cells expressed increased CD44 (p = 5.5 × 10–3), they also had 2.3-fold increased expression of PD-1H (p = 5.3 × 10–3, Fig. 4E), suggesting that they might be subject to immunosuppression. Indeed, KC;iASPPΔ8/Δ8 pancreas-infiltrating CD4+ and CD8+ T cells secreted substantially less (p = 0.037) TNF-α and IFN-γ following chronic pancreatitis (Fig. 4E, F).
A Schematic of chronic pancreatitis model and experimental design. Six-week-old WT (n = 5), KrasLSL−G12D/+;Pdx1-Cre (KC, n = 17), or KrasLSL−G12D/+;iASPPfl/fl;Pdx1-Cre (KC;iASPPΔ8/Δ8 n = 16) mice received six weekly doses of caerulein. Mice were sacrificed 48 h after the final dose and pancreata were collected for analysis. B HE, CD3, and FoxP3 immunohistochemistry stains of pancreata following six weekly doses of caerulein. Scale, 250 μm. C Column scatter plot shows quantification of FoxP3+ Treg cells per neoplasia area. D Flow cytometry contour plot (left) and quantification (right) of gating strategy to identify MDSC-M, MDSC-P, monocytes, and neutrophils from KC (blue) or KC;iASPPΔ8/Δ8 (red) pancreata. E Flow cytometry plots (left) and quantification (right) of CD44, PD-1H, TNF-α, and IFN-γ expression in pancreata-infiltrating CD4+ cells from KC (blue) or KC;iASPPΔ8/Δ8 (red) pancreata. MFI median fluorescent intensity. F Flow cytometry histograms (left) and quantification (right) of TNF-α and IFN-γ expression in pancreata-infiltrating CD8+ cells from KC (blue) or KC;iASPPΔ8/Δ8 (red) pancreata. MFI median fluorescent intensity. G Flow cytometry contour plots (left) and quantification (right) of CD103, KLRG1, CD44 and CD62L expression in pancreata-infiltrating Treg cells from KC (blue) or KC;iASPPΔ8/Δ8 (red) pancreata. MFI median fluorescent intensity. H Flow cytometry histograms of CCR8, CD103, PD-1H, and CD44 expression in pancreata-infiltrating γδ T cells from KC (blue) or KC;iASPPΔ8/Δ8 (red) pancreata.
KC;iASPPΔ8/Δ8 pancreas-infiltrating Treg cells and γδ T cells (p = 0.043) expressed higher amounts of CD44, CD103, and PD-1 (Fig. 4G, H). CD103 is a hallmark of activated tumor-infiltrating Treg cells [65], but has not been previously described in γδ T cells. KC;iASPPΔ8/Δ8 pancreas-infiltrating γδ T cells expressed higher amounts of CCR8 (p = 2.1 × 10–3, Fig. 4H), whose expression strongly correlates with worse cancer prognosis [66, 67], and 41% of KC;iASPPΔ8/Δ8 pancreas-infiltrating γδ T cells expressed PD-1H (Fig. 4H). Flow cytometry performed on entire pancreata likely profiles both tumor-infiltrating lymphocytes and other components of the tumor microenvironment such as pancreatic lymph nodes and TLS. Indeed, tumor draining lymph nodes serve as unique reservoirs for anti-tumor immune responses [68, 69] and in pancreatic cancer contribute to an immunosuppressive tumor microenvironment [70, 71]. Deletion of iASPP in oncogenic Kras-driven pancreatic neoplasia elicited an immunosuppressive microenvironment that exhibited both canonical mechanisms of immunosuppression as well as previously undescribed Treg and γδ T cell phenotypes.
Oncogenic Kras driven lung adenocarcinoma is accelerated in iASPP-deficient lung tumors
Different tumor types employ vastly different mechanisms of immune escape and display varying responsiveness to immunotherapy [72, 73]. To determine whether iASPP-deficiency accelerates cancer development by attenuating anti-tumor immunity in a different cancer type, we employed the KrasLSL−G12D/+ model of lung adenocarcinoma (Fig. 5A). Intranasal delivery of adenovirus expressing Cre recombinase results in expression of oncogenic KrasG12D in the lung epithelium, as oncogenic KrasG12D can both promote tumorigenesis and the non-cell autonomous development of an inflammatory stroma via Myc, AP-1, and NF-κB [74,75,76]. In concordance with pancreatic neoplasia findings, 15 weeks after intranasal delivery of adenovirus expressing Cre recombinase, deletion of iASPP in KrasLSL−G12D/+;iASPPfl/fl (Ki) mice elicited >7-fold increased tumor burden (p = 0.0107, Fig. 5B–D) and more advanced histological grade (p = 4.0 × 10–10) of lung cancer compared to KrasLSL−G12D/+ (K) mice (Fig. 5B). The majority (96%) of oncogenic Kras driven tumors were graded as adenomas, whereas 50% of Ki tumors had progressed to lung adenocarcinoma 15 weeks after initiation (Fig. 5B). These histological observations indicate that loss of iASPP accelerates oncogenic Kras driven lung adenocarcinoma, despite being a canonical inhibitor of p53 [45].
A Schematic of initiation of lung adenocarcinoma in iASPPfl/fl (i, n = 2), KrasLSL−G12D/+ (K, n = 3), or KrasLSL−G12D/+;iASPPfl/fl (Ki, n = 3) mice with intranasal inhalation of adenoviral Cre (Ad-Cre). Lungs and mediastinal lymph node (mLN) were collected 15 weeks post-infection (pi). B Proportion of lung tumors classified by histological grade. P value computed by Exact Contingency test. C HE stains of lung lobes 15 weeks after tumor initiation. Scale, 5 mm (top) and 250 μm (bottom). D Quantification of tumor burden of lung tumors in (C). Tumor burden was calculated as total tumor area divided by total area of each lung lobe. E HE stains of lung lobes 15 weeks after tumor initiation. Arrows indicate tertiary lymphoid structures (TLS). Scale, 250 μm. F Quantification of numbers of tertiary lymphoid structures (TLS) present per tumor from mice 15 weeks after tumor initiation.
Tumor-associated lymphocytes were observed in oncogenic Kras-driven lung adenocarcinoma (Fig. 5E). However, most tumor-associated lymphocytes were retained in tertiary lymphoid structures (TLS) adjacent to lung tumors in Ki mice, while K mice had fewer TLSs present per tumor (Fig. 5F). TLSs are observed in human lung adenocarcinoma and correlate with positive prognosis [77,78,79], but have only been observed in mouse models of lung cancer that employ tumoral expression of strongly immunogenic antigens [80]. Therefore, Ki mice represent a distinct model of immunogenic lung adenocarcinoma, in which TLS surprisingly develop in mice with accelerated lung adenocarcinoma. Ki lung tumor-associated TLS contained elevated amounts of FoxP3+ Tregs (Fig. S5A), which might indirectly facilitate Treg activation, suggesting that TLS formed in Ki mice might serve as sites of immunosuppression [81]. Flow cytometry of Ki tumor-bearing lungs revealed that lung tumor-associated CD4+, CD8+, and Treg cells expressed increased PD-1, as did CD4+, CD8+, Treg, and myeloid cells in the tumor-draining mediastinal lymph node (Fig. S5B). Increased Treg infiltration and expression of T cell markers of exhaustion are indicative of an immunosuppressive tumor microenvironment. As Ki mice possess a wild type immune system, the iASPP/p53 complex likely promotes cancer immunosurveillance through juxtacrine or paracrine signals from transformed cancer cells to the tumor microenvironment. Together these data suggest that tumoral iASPP-deficiency dampens the development of anti-tumor T cell responses while facilitating Treg infiltration and activation, abating anti-tumor immune responses and accelerating lung cancer development.
iASPP is a regulator of cell death adjuvanticity
Given that loss of iASPP promoted an immunosuppressive pancreatic tumor microenvironment that accelerated tumor development in a non-cell autonomous manner, we next asked what tumor cell-specific molecular mechanisms might be responsible for this immunosuppression. RNA-sequencing (RNA-seq) of pancreatic tumor lysates and principal component analysis revealed that KC;iASPPΔ8/Δ8 pancreata bear closer resemblance to KC than KPC pancreata (Fig. S6A) and express greater Cd3, Cd4, Itgae, Pdcd1, Il10, Ccl5, Osm, and Cd274 (Fig. S6B), reflective of an immunosuppressive stromal reaction. Notably, this KC;iASPPΔ8/Δ8 -unique gene signature resembles the immunogenic subtype of human pancreatic cancer [82] for which there is currently no representative mouse model. KC;iASPPΔ8/Δ8 mice therefore represent a novel model of immunogenic pancreatic cancer. However, pancreatic tumor lysates include both transformed cancer cells and surrounding stroma. To identify the tumor cell-intrinsic factors responsible for promoting immunosuppression, we performed flow cytometry analysis of dissociated iASPP-deficient pancreata from KC or KC;iASPPΔ8/Δ8 mice with a R26LSL−EYFP/EYFP allele. Transformed KC;iASPPΔ8/Δ8 pancreatic cells expressed higher amounts of PD-1H and PD-L1 in vivo (Fig. 6A), suggesting that these co-inhibitory receptors might be some of the tumor-intrinsic factors responsible for mitigating anti-tumor immunity following iASPP loss. Together these data suggest that transformed KC;iASPPΔ8/Δ8 tumor cells might modulate the surrounding tumor immune microenvironment via PD-1H juxtacrine signaling [17], activating PD-1H expression by local immunosuppressive Treg and γδ T cells and suppressing functional anti-tumor adaptive immune responses.
A Flow cytometry histograms (left) and quantification (right) of PD-L1 and PD-1H expression by transformed KC (blue) or KC;iASPPΔ8/Δ8 (red) pancreatic tumor cells. MFI, median fluorescent intensity. B Differentially regulated genes in A549 cells following iASPP (PPP1R13L) knockdown. A gene is considered as differentially regulated when the mNET-seq and two ChIP-seq pol II signals in the gene body (TSS + 500 bp to poly(A) site) were >2-fold increased (upregulated, blue) or decreased (downregulated, red) in the knockdown cell line versus wild type cell line. The number of differentially expressed genes is indicated for each category. The genes indicated on the left and right sides are the 15 most downregulated or upregulated genes. C Ranked analysis of transcription factor binding sites in down (blue) or up (red)-regulated genes from the total pol II mNET-seq analysis in A549 cells following iASPP (PPP1R13L) knockdown. Transcription factor enrichment in up or down regulated genes was calculated using DAVID. D Motif analysis of indicated transcription factor binding sites in A549 cells following iASPP (PPP1R13L) knockdown. E Heatmap of log2FC in gene expression of indicated immune-related genes following iASPP (PPP1R13L) knockdown in A549 cells quantified by total pol II mNET-seq. The presence of a TP53 ChIP peak or TP53, AP-1, or NFκB 1/2 binding motif is indicated to the right for each gene. AP-1 and NFκB -1 motif presence was determined by GSEA and p53 motif presence was determined by MEME. F Network plot of gene ontology analysis of differentially expressed genes following iASPP (PPP1R13L) knockdown in A549 cells measured by total pol II mNET-seq. Tan circles represent gene sets and colored dots represent genes colored by log2FC.
Furthermore, knockout of the short isoform of PPP1R13L (gene name for iASPP) in A549 lung adenocarcinoma cells resulted in knockdown of full length iASPP as well as transcriptional changes consistent with immunomodulatory activity. First, we conducted total RNA polymerase II (pol II) ChIP-seq and mNET-seq to profile polymerase occupancy as a measure of active transcription for each gene following iASPP knockdown. iASPP knockdown resulted in increased polymerase occupancy at a Vsir (gene name for PD-1H) enhancer site with active chromatin marks (Fig. S6C), reaffirming that PD-1H might be an additional iASPP/p53-regulated mediator of immunological tolerance. Transcription factor motif analysis of pol II mNET-seq and ChIP-seq revealed that families of genes whose expression changed following iASPP knockdown contained TP53, AP-1, and/or NFκB1/2 motifs (Fig. 6B–D), suggesting that iASPP has transcription factor-specific modulatory ability. P53 binding was confirmed by P53 ChIP-seq (Fig. 6E). Gene set enrichment analysis confirmed that iASPP knockdown impacts P53, NFκB, and apoptosis-related genes (Figs. 6E, S6D), while gene network analysis provided insight into potential interactions between differentially-expressed genes (Fig. 6F). TNF and IL-17 signaling pathway activation is likely due to upstream AP-1 and NFκB signaling (Fig. 6E). These data suggest that iASPP-deficiency enacts broad genomic transcriptional changes by regulating p53 or other transcription factor target selectivity, increasing expression of both pro-inflammatory and immunosuppressive molecules normally expressed during physiologic cell death. Inflammatory molecule expression may facilitate tumorigenesis [83], while immunosuppressive molecules may antagonize cell corpse adjuvanticity and promote immunological tolerance.
Discussion
Here we identify iASPP as a novel regulator of immunological tolerance. iASPP can promote autoimmune responses in vivo by limiting Treg cell activation, while also enhancing cancer immunosurveillance by inhibiting T cell exhaustion and antagonizing Treg and γδ T cell-mediated immune tolerance. These immune mechanisms are maintained by distinct cell types in which iASPP has complementary functions, namely oncogene transformed cancer cells and CD4+ T cells. In this model, iASPP and p53 function as regulators of transcriptional profiles intrinsic to dying cells that influence their adjuvanticity, thereby communicating to the immune system whether physiologic or unnatural cell death has occurred (Fig. S7).
iASPP paradoxically hindered tumor growth and development in vivo, contradicting the widely-accepted presumption that iASPP is a constitutive oncogene [84]. Although iASPP might function as an oncogene once tumors have escaped immunosurveillance, these data suggest that iASPP expression in developing tumors is critical for the establishment of a functional anti-tumor immune response. iASPP-deficient tumors displayed several novel mechanisms of immune escape not previously associated with lung or pancreatic cancer. MDSC-Ms and DCs expressed increased PD-1, whose expression on tumor-associated macrophages limits anti-tumor immunity in syngeneic tumor models [85]. γδ T cells contribute to PDAC development via an immunosuppressive phenotype [86], and were observed in both greater numbers and activation in iASPP-deficient tumors. Although iASPP deletion promoted a tolerogenic tumor microenvironment in lung and pancreatic tumors, the cell-intrinsic oncogenic properties of iASPP might dominate in other cancer types with less significant immunosurveillance. Not only does iASPP exhibit differential localization and function depending on cell type [46, 49, 87, 88], but neoplastic cells also employ diverse mechanisms of immune escape depending on tissue context [89]. Although we focus on the effects of iASPP deletion on lymphocytes here, iASPP might exert non-cell autonomous functions on other cell types as well, such as cancer-associated fibroblasts in pancreatic neoplasias, or astrocytes and microglia in EAE. Further experiments in different mouse models of cancer are necessary to assess the universality of iASPP non-cell autonomous tumor-suppression.
Mechanistically, as iASPP was selectively deleted in neoplastic cells in mouse models of lung adenocarcinoma and pancreatic cancer, transformed cells themselves must modulate the activation of an otherwise wild-type immune system, likely through juxtacrine or paracrine signaling. We suggest that iASPP functions by inhibiting p53-mediated immunosuppression, particularly by modulating p53 transcriptional promotion of tolerance following physiologic cell death. Fas is one of the most significantly iASPP-regulated p53 target genes [56], and its expression on apoptotic cells promotes tolerogenic phagocytosis [35, 90]. Loss of function or dominant-negative mutations in Fas/FasL are associated with autoimmune disease [28, 91,92,93,94,95,96,97,98] and Fas overexpression facilitated tumorigenesis in vivo in a non-cell autonomous manner by promoting MDSC and Treg cell recruitment [99]. Interestingly, Treg cells express decreased FasL and are resistant to Fas-mediated apoptosis [100], potentially explaining their relative abundance we observed in vivo. PD-1H was another p53 target overexpressed in iASPP-deficient tumor cells as well as responding lymphocytes, suggesting that iASPP might inhibit p53 transcriptional activation of PD-1H and its subsequent juxtacrine signaling [17]. iASPP−/− mice exhibited significantly increased immune tolerance that largely phenocopied the consequences of treatment with a PD-1H agonist [101]. In addition to upregulation via p53 in the setting of programmed cell death [17, 102], Fas and PD-1H both promote immunological tolerance in a non-cell autonomous manner by acting as negative regulators of cell death-related adjuvanticity to responding phagocytes [16, 34, 90]. Without iASPP-mediated modulation of target selectivity, p53 might also promote tumorigenesis via tolerogenic or inflammatory DAMPs, such as PGE2 and IL-1α [103, 104], and general immunosuppression via senescence or tolerogenic molecules such as CD47, MFG-E8, or thrombospondin [105, 106]. Indeed, iASPP-deficient cells increased expression of inflammatory cytokines S100a8 and S100a9, which are also expressed by apoptotic cancer cells and can promote metastasis [107, 108]. Further investigation is necessary to determine if these potential cell-intrinsic mechanisms, in addition to Fas and PD-1H, are responsible for promoting the non-cell autonomous tolerance observed in vivo.
Although the data presented here suggest that iASPP promotes adjuvanticity by antagonizing p53-mediated tolerogenic phagocytosis, both iASPP and p53 have additional independent effects on the immune system. p53 functions in a positive feedback loop to drive type I IFN-mediated antiviral immunity [109], and can suppress STAT1 and NF-κB mediated inflammation [32]. p53-dependent activation of Batf3 in intratumoral monocytic MDSCs is essential for the development of anti-tumoral Ly6c+CD103+ DCs [110]. iASPP can inhibit NF-κB subunit RelA/p65-mediated inflammation and pro-proliferative functions [47, 111], even in the absence of p53 [48, 112], indicating that iASPP inhibition of RelA/p65 is functionally orthogonal to its inhibition of p53. RelA/p65 expression is essential for maintenance of Treg function, identity, and immune tolerance [113]. Therefore, iASPP inhibition of NF-κB mediated immune tolerance in Treg cells is a potential p53-independent mechanism of iASPP immunological influence, although iASPP likely also has Treg cell-extrinsic roles as tumor-specific deletion of iASPP promoted tolerogenic immune responses. iASPP regulates p63 target selectivity in keratinocytes with differential binding to promoters with AP-1 versus E2F motifs [114], suggesting that cross-talk between iASPP and other co-factors might be responsible for cell type-specific transcriptional regulation. Indeed, we identified AP-1, P53, E2F, and NF-κB binding motifs amongst inflammatory genes differentially expressed in iASPP deficient A549 lung cancer cells. Ultimately, the multifaceted roles of iASPP likely come together to influence the adaptive immune system, as chronic inflammation can both promote tumorigenesis and induce immunosuppression [83, 115,116,117].
This study identifies iASPP as a novel component of the maintenance of immune homeostasis between tolerance and autoimmunity. Loss of iASPP in cancer cells, CD4+ T cells, or with germline deletion elicited strikingly elevated immune tolerance. These data suggest that iASPP is a previously unidentified regulator of the immunological phenotypes of dying cells and thereby orchestrates the ensuing innate and adaptive immune responses to cell death. Although many signaling mediators such as DAMPs and ‘eat-me signals’ have been identified, the underlying pathways responsible for discriminating between physiologic apoptosis and unnatural cell death have remained previously uncharacterized. While this study demonstrates iASPP promotion of non-physiologic cell death adjuvanticity in the contexts of autoimmune disease and cancer, the normal physiologic role of iASPP in providing adjuvant co-stimulation remains unclear. Investigation of iASPP and its regulatory mediators provides a previously unappreciated opportunity to identify immunological therapeutic targets with implications for multiple diseases.
Materials and methods
Mice
iASPP−/− [46, 49], iASPPfl/fl [46], ASPP2fl/fl [118], KrasLSL−G12D [119], Pdx1-Cre [120], Trp53LSL−R172H/+ [121], Rosa26LSL−EYFP [122], and Rosa26CreER [123] mice have been previously described. iASPPfl/fl mice were back-crossed into a C57BL/6 background strain prior to being bred with Pdx1-Cre and Rosa26LSL−EYFP/EYFP mice (Jackson Laboratory) to generate iASPPfl/fl;Rosa26LSL−EYFP/EYFP (i) and iASPPfl/fl;Pdx1-Cre;Rosa26LSL−EYFP/EYFP (iC) mice. KrasLSL−G12D/+;Rosa26LSL−EYFP/EYFP (K), KrasLSL−G12D/+;Pdx1-Cre;Rosa26LSL−EYFP/EYFP (KC), KrasLSL−G12D/+;iASPPfl/fl;Rosa26LSL−EYFP/EYFP (Ki), and KrasLSL−G12D/+; iASPPfl/fl;Pdx1-Cre;Rosa26LSL−EYFP/EYFP (KC;iASPPΔ8/Δ8) mice were generated by crossing KrasLSL−G12D heterozygous mice with iC or Pdx1-Cre mice. KrasLSL−G12D/+;Trp53LSL−R172H/+ (KP) and KrasLSL−G12D/+;Trp53LSL−R172H/+;Pdx1-Cre;Rosa26LSL−EYFP/EYFP (KPC) mice were generated by crossing KC mice with Trp53LSL−R172H heterozygous mice. Mice heterozygous for deletion of exon 8 of Ppp1R13l (iASPPΔ8/+) were used to generate iASPP−/− mice and littermate wild type controls. The health of mice was monitored daily by veterinary staff at the Wellcome Trust Centre for Human Genetics (WTCHG) at the University of Oxford. No sample size calculation estimate or randomization were performed. Both male and female mice were used, mice were gender and age-matched within experiments where possible, and adult mice (>8 weeks old) were used for all experiments. Experimental and control mice were co-housed whenever appropriate. All mice were housed and all procedures were performed at the WTCHG at the University of Oxford. All animal studies described in this study were performed in accordance with guidelines provided by the University of Oxford Institutional Animal Care and Use Committee. Procedures were performed under the Home Office Animal Scientific Procedures Act 1986 guidelines (license PPL 30/3451 and PPL 30/3096).
PCR genotyping
After mice were weaned, DNA was isolated from ear clips by incubation in TDB (50 mM KCl, 10 mM Tris-HCl, 0.1% Triton-X100 in H2O) with 0.4 mg/mL Proteinase K (Qiagen) for 2 h at 55 °C, then 10 min at 95 °C. Pdx1-Cre and Rosa26CreER mice were genotyped using the following primers: Cre-F, 5′-CATTTGGGCCAGCTAAACAT-3′; Cre-B, 5′-ATTCTCCCACCGTCAGTACG-3′; Actin-F, 5′-GGTGTCATGGTAGGTATGGGT-3′; and Actin-B, 5′-CGCACAATCTCACGTTCAG-3′. iASPP2 fl/fl mice were genotyped using the following primers: FRANT9, 5′-GGGTAGGAAAAAGGGCTGAG-3′; FLP2, 5′-CCGAATTGGAGAAGTGAAGC-3′. KrasLSL−G12D/+ mice were genotyped using the following primers: Y116, 5′-TCCGAATTCAGTGACTACAGATG-3′; Y117, 5′-CTAGCCACCATGGCTTGAGT-3′; Y118, 5′-ATGTCTTTCCCCAGCACAGT-3′. Rosa26LSL−EYFP/EYFP mice were genotyped using the following primers: YFPc, 5′-AAAGTCGCTCTGAGTTGTTAT-3′ and YFPwt, 5′-GGAGCGGGAGAAATGGATATG-3′; or YFPc and YFPmut, 5′-AAGACCGCGAAGAGTTTGTC-3′. Trp53LSL−R172H/+ mice were genotyped using the following primers: p53u, 5′-CTTGGAGACATAGCCACACTG-3′ and p53wt, 5′-TTACACATCCAGCCTCTGTGG-3′; or p53u and p53mut, 5′-AGCTAGCCACCATGGCTTGAGTAAGTCTGCA-3′. iASPP−/− mice were genotyped using the following primers: I8-2, 5′- ACAGGCAGCTACTGGTATTC-3′ and E8-2, 5′-AGAGCAGCCTCAGAGCATGG-3′; or I8-2 and FLP2.
PCR reactions were performed on 0.5 μL of sample DNA using Green Taq DNA Polymerase (GenScript) according to manufacturer’s protocols. The Pdx1-Cre and Rosa26CreER allele yields a 308 bp product while the wild type allele yields a 753 bp product from primers Cre-F, Cre-B, Actin-F, and Actin-B. The ASPP2 fl/fl allele yields a 203 bp product while the wild type allele yields a 169 bp product from primers SDL1s and SDL2s. The iASPP fl/fl allele yields a 400 bp product while the wild type allele yields a 265 bp product from primers FRANT9 and FLP2. The KrasLSL−G12D allele yields a 327 bp product while the wild type allele yields a 450 bp product from primers Y116, Y117, and Y118. The Rosa26LSL−EYFP allele yields a 320 bp product from primers YFPc and YFPmut, while the wild type allele yields a 600 bp product from primers YFPc and YFPwt. The Trp53LSL−R172H allele yields a 270 bp product from primers p53u and p53mut, while the wild type allele yields a 166 bp product from primers p53u and p53wt. The iASPP−/− allele yields a 700 bp product from primers I8-2 and FLP2, while the wild type allele yields a 600 bp product from primers I8-2 and E8-2.
Preparation of mouse T cell populations
Spleens from 6–8-week-old mice were dissociated by mechanical dissociation using the plunger of a 2.5 mL syringe on a 70 μm cell strainer (Falcon). Red blood cells were lysed with ACK Lysis buffer (150 mM NH4Cl, 10 mM KHCO3, 0.1 mM EDTA in H2O, pH 7.4) for 2 min at RT. Following centrifugation cells were resuspended in RPMI 1640 (Gibco) supplemented with 1% heat-inactivated fetal calf serum (HI-FCS), 2 mM L-glutamine, 100 U/mL penicillin, and 100 μg/mL streptomyocin and counted using a hemocytometer. Splenocytes were then washed and resuspended in 0.5% BSA and 2 mM EDTA in PBS at 1 × 107 cells/mL for magnetic-activated cell sorting (MACS). CD4+ T cells were isolated by negative selection using a CD4+ T Cell Isolation Kit and LS columns (Miltenyi Bitoec), naive CD4+CD44−CD62L+ T cells were isolated using a Naive CD4+ T Cell Isolation Kit and LS columns (Miltenyi Biotec), and CD4+CD25− conventional T cells (Tconv) and CD4+CD25+ regulatory T cells (Treg) were isolated by sequential negative and positive selection using a CD4+CD25+ Regulatory T Cell Isolation Kit and LD and MS columns (Miltenyi Biotec) according to manufacturer’s protocols.
In vitro T cell activation
CD4+ T cells were washed and resuspended at 1 × 107 cells/mL in PBS and labeled with 5 μM CellTrace carboxyfluorescein succinimidyl ester (CFSE) Cell Proliferation Dye (Life Technologies) for 10 min at 37 °C. Staining was quenched with five volumes of HI-FCS and cells were resuspended at 5 × 105 cells/mL in RPMI 1640 supplemented with 10% HI-FCS, 10 mM HEPES (Life Technologies), 50 μM β-mercaptoethanol (Sigma), 2 mM L-glutamine, 100 U/mL penicillin, and 100 μg/mL streptomyocin with or without 40 ng/mL murine IL-2 (PeproTech), Cells were stimulated with indicated concentrations of plate-bound anti-CD3ε mAb (145-2C11, BioLegend) at 200 μL/well in a 96-well plate, and CFSE dilution was measured by flow cytometry after 72 h.
Treg suppression assay
CD4+CD25− Tconv cells were washed and resuspended at 1.5 × 106 cells/mL in PBS and labeled with 5 μM CFSE for 10 min at 37 °C. Staining was quenched with five volumes of HI-FCS and cells were resuspended at 5 × 105 cells/mL in RPMI 1640 supplemented with 10% HI-FCS, 10 mM HEPES (Life Technologies), 50 μM β-mercaptoethanol (Sigma), 1 mM sodium pyruvate (Life Technologies), 100 mM non-essential amino acids (Life Technologies), 2 mM L-glutamine, 100 U/mL penicillin, and 100 μg/mL streptomyocin. Control splenocytes were resuspended at 1 × 107 cells/mL and exposed to 30 Gy gamma irradiation and labeled with soluble 1 μg/mL anti-CD3ε mAb. CD4+CD25+ Treg cells were co-cultured at indicated ratios with 5 × 104 irradiated splenocytes and 2.5 × 104 Tconv cells/well in round-bottom 96-well plates (Corning). After 72 h cells were washed and CFSE dilution was measured by flow cytometry.
CD4 differentiation assay
Naive CD4+CD44−CD62L+ T cells were resuspended at 1 × 106 cells/mL in RPMI 1640 supplemented with 10% HI-FCS, 50 μM β-mercaptoethanol, 2 mM L-glutamine, 100 U/mL penicillin, and 100 μg/mL streptomyocin and stimulated with plate-bound 1 μg/mL anti-CD3ε mAb and 1 μg/mL anti-CD28 (37.51, BioLegend) at 200 μL/well in a 96-well plate. Cells were polarized as indicated using the following culture conditions: Th0, 5 μg/mL anti-IL-4 mAb (11B11, BioLegend) and 5 μg/mL anti-IFN-γ mAb (XMG1.2, BioLegend); Th1, 10 ng/mL murine IL-12 (PeproTech) and 5 μg/mL anti-IL-4 mAb; Th2, 10 ng/mL murine IL-4 (PeproTech) and 5 μg/mL anti-IFN-γ mAb; Th17, 5 μg/mL anti-IL-4 mAb, 5 μg/mL anti-IFN-γ mAb, 10 ng/mL human IL-23 (PeproTech), 1 ng/mL human TGF-β (PeproTech), and 5 ng/mL murine IL-6 (PeproTech); Treg, 1 ng/mL human TGF-β, 5 μg/mL anti-IL-4 mAb, and 10 μg/mL anti-IFN-γ mAb. After 72 hr cells were analyzed by intracellular flow cytometry for CD4+ Th subset polarization.
Experimental autoimmune encephalomyelitis
Experimental autoimmune encephalomyelitis (EAE) was induced in 8–10-week-old female mice by bilateral subcutaneous (s.c.) injection of a total of 200 μg MOG35-55 (MEVGWYRSPFSRVVHLYRNGK, Severn Biotech) emulsified 1:1 in complete Freund’s adjuvant supplemented with 400 μg H37Ra mycobacterium tuberculosis (CFA, Chondrex) in incomplete Freund’s adjuvant (Sigma). Immunization was boosted immediately and 48 h after s.c. injection with 300 ng pertussis toxin (PTx, Merck Chemicals) by i.v. injection. Mice were weighed and neurological impairment was measured daily in a genotype-blind fashion using the following clinical severity scoring system: 0, healthy; 0.5, partial tail limpness; 1, tail paralysis; 1.5, righting reflex weakness; 2, hind limb weakness; 2.5, paralysis of one hind limb; 3, paralysis of both hind limbs; 3.5 hind limb paralysis and paralysis of one fore limb; 4, quadriplegia; 5, moribund. When mice reached a score of 1, cages were changed to dry bedding and wet mash was placed on the cage floor. Under recommendation of the animal ethics committee, mice were humanely euthanized when they reached a clinical score greater than or equal to 3. Any mice exhibiting adverse signs related to EAE such as >20% weight loss, labored breathing, or inability to access food were humanely euthanized. Mice remaining alive 30 days after immunization were sacrificed and spleens, brains, spinal cords, iLNs, and blood were collected for analysis.
MOG35-55 stimulation assay
Splenocytes were washed and resuspended at 1 × 107 cells/mL in PBS and labeled with 5 μM CFSE for 10 min at 37 °C. Staining was quenched with five volumes of HI-FCS and cells were resuspended at 1 × 106 cells/mL in RPMI 1640 supplemented with 40 ng/mL murine recombinant IL-2, 10% HI-FCS, 2 mM L-glutamine, 100 U/mL penicillin, and 100 μg/mL streptomyocin. Splenocytes were then stimulated with indicated concentrations of MOG35-55 peptide for 72 h, and CFSE dilution was measured by flow cytometry.
Adenoviral infection
Autochthonous lung adenocarcinoma was initiated in 9–13 week old mice by intranasal infection with 2.5 × 107 PFU adenovirus as previously described [124, 125]. Briefly, Ad-CMV-iCre (Vector Biolabs) was resuspended in 50 μL/mouse Opti-MEM (Life Technologies) with 10 mM calcium chloride. Mice were anesthetized with isoflurane until respiration rate reached 2 respirations/sec and 25 μL of adenovirus suspension was administered dropwise to a single nostril. After 5 min of recovery, adenoviral administration was repeated on the same nostril. Mice were monitored daily and any mice exhibiting adverse signs related to tumor burden such as hunched posture, labored respiration, or >15% weight loss were humanely euthanized. All mice were sacrificed 15 weeks after tumor initiation and spleens, lungs, livers, and mediastinal lymph nodes (mLN) were collected for analysis.
Caerulein-induced chronic pancreatitis
Chronic pancreatitis was induced in 6–8-week-old mice by six weekly intraperitoneal (i.p.) injections of 250 μg/kg of caerulein (Sigma) in PBS. Approximately equal numbers of male and female mice were used in all experimental groups. Mice were monitored daily and any mice exhibiting adverse signs related to chronic pancreatitis or tumor burden such as hunched posture, piloerection, >15% weight loss, palpable abdominal masses, lethargy, diarrhea, rectal bleeding, ulcerated tumors, tumors interfering with locomotion, feeding, breathing, or vision, genital prolapse, or dermatitis were humanely euthanized. Mice were sacrificed 48 h following the final caerulein injection and spleens, pancreas, lungs, and livers were collected for analysis.
Tissue collection
Mice were sacrificed by cervical dislocation, transcardially perfused with 40 mL PBS and tissues were allocated for immunohistochemistry (IHC), immunofluorescence, flow cytometry, RNA isolation, in vitro assays, or tissue lysate preparation. In cases where tissues were only taken for IHC or immunofluorescence, mice were subsequently perfused with paraformaldehyde-lysine-periodate fixative (PLP; 50 mM phosphate buffer with 1% paraformaldehyde (Sigma), 0.1 M L-lysine buffer (pH 7.4), 0.2% NaIO4 (Sigma)). Tissue lysates were prepared by homogenizing organs in 50 μL/mg RPMI with 1% Triton X-100 (Sigma) with a rotor-stator tissue homogenizer for 15 s, followed by centrifugation at 5000 g for 10 min. RNA was prepared from snap-frozen tissues by grinding with mortar and pestle in liquid nitrogen, followed by tissue homogenization in 600 μL Buffer RLT (Qiagen) with β-mercaptoethanol and RNA isolation using the RNeasy Mini Kit according to manufacturer’s protocols.
Immunohistochemistry
Isolated tissues were individually perfused with PBS followed by PLP. Tissues were fixed overnight in PLP at 4 °C and stored in 70% ethanol until processing. PLP-fixed tissues were dehydrated overnight using a Shandon Excelsior Tissue Processor (Thermo), embedded in paraffin, sectioned at a thickness of 4 μm, and stained with haematoxylin and eosin using standard methods. For IHC, slides were dewaxed and antigen retrieval was performed by boiling in Tris-EDTA (pH 9) and 0.5% Tween-20 (Sigma) for 3 min at 120 °C in a pressure cooker. Slides were then cooled, washed with dH2O, and endogenous peroxidases were blocked with 3% hydrogen peroxide (Sigma) in methanol (Sigma) for 15 min at RT. Tissues were blocked with 5% normal goat (Vector Laboratories) or donkey serum (Sigma) in PBS, and stained with primary antibody overnight at 4 °C followed by biotinylated secondary anti-rabbit, anti-hamster, anti-rat, or anti-goat (Vector Laboratories) for 30 min at RT and Vectastain Elite ABC (Vector Laboratories) for 30 min at RT. Staining was visualized using ImmPACT DAB Peroxidase (Vector Laboratories) using manufacturer’s protocols, and slides were counterstained with haematoxylin for 4 s, dehydrated to Histo-Clear II (National Diagnostics), and coverslipped using VectaMount Permanent Mounting Medium (Vector Laboratories). The following primary antibodies were used for IHC: rabbit anti-CD3 (ab5690; 1:200, 4 min develop), rat anti-FoxP3 (FJK-16s, eBioscience; 1:50, 30 min develop). Images were obtained using a NanoZoomer S210 (Hamamatsu) and analyzed using NDP.view2 software, or using a Leica DM5500B and analyzed using Fiji.
Luxol fast blue staining was performed by incubating sections in 0.1% luxol fast blue (Fisher) in 95% ethanol with 0.5% glacial acetic acid overnight at 56 °C. Slides were then differentiated in 0.05% lithium carbonate (Scientific Laboratory Supplies Ltd) in dH2O for 30 s, then 70% ethanol for 30 s, and counterstained with 0.1% cresyl violet acetate (MP Biomedicals) in dH2O with glacial acetic acid.
Histological quantification
Grading of pancreatic neoplasias was performed in a blind fashion as described [126]. Briefly, pancreatic neoplasias were categorized as metaplasia (ADM) when exhibiting non-invasive epithelial proliferation and cellular abnormalities, PanIN when glandular epithelial proliferation was confined to pancreatic ducts, and pancreatic ductal adenocarcinoma (PDAC) when exhibiting ductal differentiation and penetration of the basement membrane. Lung tumor grading was performed in a blind fashion as previously described [125, 127, 128]. Briefly, lung adenomas were identified by prominent nucleoli, while lung adenocarcinomas were graded by extent of nuclear atypia, cellular pleomorphism, invasiveness, mitotic index, cellular heterogeneity, and desmoplasia.
IHC images were blinded using blindanalysis v1.0 [129]. Treg infiltration was assessed in a blind fashion by counting the number of FoxP3+ nuclei present in IHC images of tumors. Tumor area was measured in Fiji by outlining the perimeter of tumors and calculating the enclosed area. For analysis, average Treg infiltration was calculated by diving the total number of infiltrating FoxP3+ cells by the total tumor area across three or more fields of view per mouse. Tumor burden was measured per lung lobe in NDPview.2 by outlining the area of individual tumors and of individual lobes. CD3 infiltration into the CNS was assessed in a blind fashion by counting the number of CD3 cells present in in IHC images of brain and spinal cord. Average CD3 infiltration was calculated by averaging across three or more fields of view for each mouse. Demyelination was quantified by measuring Luxol fast blue intensity using the Mean Gray Value function in Fiji.
Flow cytometry
Lymph nodes were dissociated by mechanical dissociation between two frosted microscope slides (Fisher). Spleens, thymus, livers, and brains were dissociated by mechanical dissociation using the plunger of a 2.5 mL syringe on a 70 μm cell strainer (Falcon). Livers and brains were further processed by centrifugation in a Percoll (VWR) gradient at 700 g for 25 min with no brake, and lymphocytes were isolated from the 40–70% Percoll interface.
Lungs, pancreas, kidneys, and hearts were placed in 5 mL collagenase buffer (125 U/mL collagenase IV (Worthington), 40 U/mL DNaseI (Roche), 25 mM HEPES, and 0.5% HI-FCS in 1X HBSS) in C tubes (Miltenyi Biotec). Tissues were dissociated using program “m_imp_tumor_01” for lungs, “m_imp_tumor_04” for pancreas, “multi_E_01” for kidneys, or “m_heart_01” for hearts on a gentleMACS dissociator (Miltenyi Biotec), incubated at 37 °C for 40 min with gentle agitation, then dissociation was repeated using program “m_imp_tumor_01” for lungs, “m_imp_tumor_04” for pancreas, “multi_E_02” for kidneys, or “m_heart_02” for hearts. Following dissociation cells were filtered through a 100 μm cell strainer (Falcon).
Colons were cut longitudinally and washed in PBS, then cut into 5 mm pieces and incubated in 5 mM EDTA in PBS for 15 min at 37 °C. Washed tissues were then placed in 5 mL of RPMI 1640 supplemented with 5% HI-FCS, 20 mM HEPES, 100 U/mL penicillin, 100 μg/mL streptomyocin, and 100 U/mL collagenase VIII (Sigma) in C tubes. Tissues were dissociated using program “m_imp_intestine_01”, incubated for 60 min at 37 °C with gentle agitation, dissociated again using program “m_imp_intestine_01”, and filtered through a 100 μm cell strainer. Cells were then resuspended and centrifuged in a Percoll gradient at 700 g for 25 min with no brake, and lymphocytes were isolated from the 40 to 70% Percoll interface.
Red blood cells were lysed with ACK Lysis buffer for 2 min at RT and resuspended in RPMI 1640 supplemented with 1% HI-FCS, 2 mM L-glutamine, 100 U/mL penicillin, and 100 μg/mL streptomyocin, counted, and adjusted to 1 × 107 cells/mL. 3 × 103 to 1 × 107 cells were Fc receptor blocked with rat anti-CD16/CD32 (2.4G2, BD Biosciences; 1:100) for 5 min, and stained for 30 min on ice in round-bottom 96-well plates with Zombie Green Viability Dye (BioLegend; 1:1000) or Zombie NIR Viability Dye (BioLegend; 1:1000) and directly-conjugated antibodies in 0.5% HI-FCS and 0.05% sodium azide in PBS. Following staining cells were fixed with Fixation/Permeabilization Buffer (eBioscience) overnight at 4 °C. The following surface antibodies were used: PerCP-Cy5.5 anti-CCR8 (SA214G2; 1:100), APC-Cy7 anti-CD3 (17A2; 1:100), PE-Cy5 anti-CD4 (RM4-5; 1:300), BV421 anti-CD4 (GK1.5; 1:200), FITC anti-CD4 (RM4–5, eBioscience; 1:100), APC anti-CD8α (53–6.7; 1:200), BV510 anti-CD8α (53–6.7; 1:200), BV785 anti-CD8α (53–6.7; 1:200), BV510 anti-CD11b (M1/70; 1:200), PerCP-Cy5.5 anti-CD11b (M1/70; 1:100), BV421 anti-CD11c (N418; 1:200), PE-Cy7 anti-CD11c (N418; 1:250), BV650 anti-CD19 (6D5; 1:150), PE-Cy5 anti-CD24 (M1/69; 1:600), BV510 anti-CD25 (PC61; 1:100), BV785 anti-CD44 (IM7; 1:200), FITC anti-CD45 (30-F11; 1:300), PE/Dazzle 594 anti-CD49b (DX5; 1:100), Alexa Fluor 700 anti-CD62L (MEL-14; 1:150), PE-Cy7 anti-CD68 (FA-11; 1:250), BV605 anti-CD80 (16–10A1; 1:50), PE anti-CD80 (16–10A1; 1:100), PE anti-CD86 (GL-1; 1:100), PE-Cy7 anti-CD103 (2E7; 1:100), PE/Dazzle 594 anti-CTLA-4 (UC10–4B9; 1:100), BV421 anti-F4/80 (BM8; 1:100), BV711 anti-F4/80 (BM8; 1:100), BV421 anti-KLRG1 (2F1/KLRG1; 1:100), BV785 anti-Ly6C (HK1.4; 1:500), PerCP-Cy5.5 anti-Ly6G (1A8; 1:250), Alexa Fluor 700 anti-MHCII (M5/114.15.2; 1:250), BV711 anti-PD-1 (29 F.1A12; 1:100), APC anti-PD-1H (MH5A; 1:100), PE-Cy7 anti-PD-L1 (10 F.9G2; 1:250), PE anti-TCRγδ (UC7–13D5; 1:100), and PE-Cy7 anti-Tim-3 (RMT3–23; 1:100). For intracellular staining, cells were washed with Permeabilization Buffer (eBioscience) following fixation, incubated with directly-conjugated antibodies in Permeabilization Buffer for 30 min on ice, washed, and acquired directly. The following intracellular antibodies were used: PE-Cy5 anti-FoxP3 (FJK-16s, eBioscience; 1:100), PE anti-GATA3 (16E10A23; 1:50), Pacific Blue anti-GranzymeB (GB11; 1:50), PE/Dazzle 594 anti-Helios (22F6; 1:100), PerCP-Cy5.5 anti-IFN-γ (XMG1.2; 1:50), BV421 anti-IL17a (TC11–18H10.1; 1:50), BV650 anti-RORγt (Q31–378, BD Biosciences; 1:50), BV605 anti-T-bet (4B10; 1:50), and BV510 anti-TNF-α (MP6-XT22; 1:50). All antibodies were purchased from BioLegend unless otherwise indicated.
Following fixation, cells were read using 3–11 fluorophore flow cytometry on a LSRFortessa X-20 (BD Biosciences) with 405, 488, 561, and 633 lasers. Compensation matrices were determined for each individual acquisition date using unstained and single fluorophore-stained controls. Data was analyzed using FlowJo software (Tree Star) and cell numbers were calculated from tissue cell counts preceding staining.
Cell culture and PPP1R13L knockdown
A549 cells (ATCC) were cultured in DMEM (Corning) supplemented with L-glutamine (2 mM), penicillin (100 U/mL), streptomycin (100 μg/mL) and 10% fetal calf serum (FCS; Gibco) at 37 °C in a 5% CO2 humidified atmosphere. PPP1R13L knockdown was achieved by cloning a sgRNA targeting the PAM sequence TTGTATGCCCTGGAAGTTGTGG into PX459 (Addgene). A549 cells were transfected with this construct using Lipofectamine (Thermo) following the manufacturer’s protocol and selected with puromycin. Single colonies were expanded and CRISPR/Cas9 indel mutation was validating by sanger sequencing, confirming knockout of the short isoform of iASPP (sviASPP) and knockdown of iASPP.
ChIP-seq
ChIP-seq of total pol II and p53 were carried out as previously described [130] with 5 µg per IP of anti-pol II antibody (NBP2-32080, Novus Biologicals) and anti-p53 antibodies (9282S, Cell Signaling and sc-126, Santa Cruz). Preparation of ChIP-seq library and ChIP sequencing was prepared with the NEBNext Ultra II DNA Library Prep Kit for Illumina (NEB), according to the manufacturer’s instructions and conducted by the high throughput genomics team of the WTCHG, Oxford. A549 Input (DRR016939), H3K4me1 (DRR016935), and H3K27ac (DRR016938) ChIP-seq were obtained from [131]. Adapters were trimmed with Cutadapt in paired-end mode with the following parameters: -q 15, 10 --minimum-length 10 -A GATCGTCGGACTGTAGAACTCTGAAC -a TGGAATTCTCGGGTGCCAAGG. Obtained sequences were mapped to the human hg19 reference genome with Bowtie2. Properly paired and mapped reads were filtered with SAMtools. PCR duplicates were removed with Picard MarkDuplicates tool. Library-size normalized bedgraph files were created with Bedtools and trackhubs in the UCSC browser were generated with the UCSC bedGraphToBigWig tool. To obtain better peak calling results due to the difficulties of performing ChIP-seq of p53 in A549, for each condition, the ChIP-seq of both p53 antibodies were merged. Peak calling was called from merged biological replicates with MACS2 version 2.1.1.20160309 and the parameters: callpeak –f BAMPE -g 2.9e9 -B -q 0.01 –call-summits. Motif analysis was performed with the MEME Suite. The genomic locations of the AP-1 (JUN/JUNB/JUND/FOS/FOSB/FOSL1/FOSL2), NFκB1/2, and p53 transcription factors were obtained from the Gene Transcription Regulation Database (GTRD) [132]. All the metaclusters were retrieved for each transcription factor and filtered out any metaclusters that did not have support from four independent experiments and from the MACS peak caller. Each genomic location was then overlapped with gene TSS −/+ 10 kb to assign each metacluster to genes.
mNET-seq
mNET-seq was carried out as previously described [133] with minor changes. In brief, the chromatin fraction was isolated from 1 × 107 A549 cells. Chromatin was digested in 100 µL of MNase (40 units/mL) reaction buffer for 5 min at 37 °C in a thermomixer (1400 rpm). After addition of 10 µL EGTA (25 mM) to inactivate MNase, soluble digested chromatin was collected by 13,000 rpm centrifuge for 5 min. The supernatant was diluted with 400 µL of NET-2 buffer and added to 10 µg of total RNA pol II antibody (MABI0601, MBL International) conjugated with Dynabeads™ M-280 Sheep Anti-Mouse IgG. Immunoprecipitation was performed at 4 °C for 1 h. The beads were washed with 1 ml of NET-2 buffer six times and then with 100 µl of 1xPNKT (1xPNK buffer and 0.05 % Triton X-100) buffer once in the cold room. Washed beads were incubated in 50 µl PNK reaction mix (1xPNKT, 1 mM ATP and 0.05 U/ml T4 PNK 3′phosphatase minus (NEB) in Thermomixer (1400 rpm) at 37 °C for 6 min. After the reaction, the beads were washed with 1 ml of NET-2 buffer once and RNA was extracted with Trizol reagent. RNA was suspended in urea Dye (7 M Urea, 1xTBE, 0.1% BPB and 0.1% XC) and resolved on 6% TBU gel (Invitrogen) at 200 V for 5 min. To size select 30–160 nt RNAs, a gel fragment was cut between BPB and XC dye markers. 0.5 mL tube was prepared with 3–4 small holes made with 25 G needle and placed in a 1.5 mL tube. Gel fragments were placed in the layered tube and broken down by centrifugation at 12,000 rpm for 1 min. The small RNAs were eluted from gel using RNA elution buffer (1 M NaOAc and 1 mM EDTA) at 25 °C for 1 h in Thermomixer (900 rpm). Eluted RNA was purified with SpinX column (Coster) with 2 glass filters (Millipore) and the flow-through RNA was ethanol precipitated. RNA libraries were prepared according to manual of NEBNext small RNA library prep kit (NEB). 12–14 cycles of PCR were used to amplify the library. Deep sequencing (Hiseq4000, Illumina) was conducted by the high throughput genomics team of the WTCHG, Oxford.
mNET-seq data were processed as follows: adapters were trimmed with Cutadapt version 1.9.1 [134] in paired-end mode with the following parameters: -q 15, 10 --minimum-length 10 -A GATCGTCGGACTGTAGAACTCTGAAC -a TGGAATTCTCGGGTGCCAAGG. Trimmed reads were mapped to the human hg19 reference sequence with Tophat2 version 2.1.0 [135] and the parameters –g 1 –r 3000 –no-coverage-search. SAMtools version 1.3.1 [136] was used to retain only properly paired and mapped reads (-f 3). A custom python script [137] was used to obtain the 3′ nucleotide of the second read and the strandedness of the first read. Strand-specific bam files were generated with SAMtools. Library-size normalized bedgraph files were created with Bedtools version 2.23 [138] genomecov –bg –scale and trackhubs in the UCSC browser were generated with the UCSC bedGraphToBigWig tool.
Reads quantification and transcription factors binding sites
Total read base count for mNET-seq data were computed with samtools bedcov tool using strand-specific bam files and normalized to 100 million paired-end reads and to the region’s length. For ChIP-seq, total read base count were computed with samtools bedcov, normalized to 100 million paired-end reads, then the Input signal was subtracted to the IP signal and normalized to the region’s length. Only the regions with a positive signal in at least one sample were kept. For the samples having a signal ≤0 on the remaining regions, their values were put to the minimal value divided by two. The quantification is thus defined for the mNET-seq as:
and for ChIP-seq as:
The quantification regions is defined as follow: Gene body (GB) of protein-coding genes: TSS + 0.5 kb to TES. The lists of transcription factors binding sites associated with down-regulated or up-regulated genes were obtained with the DAVID Bioinformatics Resources [139, 140].
Gene set enrichment analysis
Gene set enrichment analysis was performed using fgsea (version 3.15). Genes were ranked according to log2FC and used to query GOBP signature libraries. Network plot was generated using clusterProfiler (version 1.0.12).
RNA-seq and principal component analysis
RNA was prepared from snap-frozen tissues by grinding with mortar and pestle in liquid nitrogen, followed by tissue homogenization in 600 μL Buffer RLT (Qiagen) with β-mercaptoethanol and RNA isolation using the RNeasy Mini Kit according to manufacturer’s protocols. Following mRNA isolation, libraries were prepared using a QuantSeq 3′ mRNA-seq Library Kit (Lexogen) and sequenced on a HiSeq 4000 (Illumina). FASTQ files were aligned using BWA-mem to the GRCm38.ERCC genome, counted using featureCounts, and annotated using the TxDb.Mmusculus.UCSC.mm10.knownGene database. Normalized counts were obtained using the VST normalization function of DeSeq v1.16.1 and used to generate expression scatter plots and volcano plots. Principal component analysis was performed using plotPCA. All RNA-Seq analyses were conducted in RStudio.
Statistical analyses
Statistical analysis was performed using unpaired two-tailed Student’s t tests where indicated. Where multiple comparisons were performed, statistical significance was determined using the two-stage linear step-up procedure of Benjamini, Krieger and Yekutieli with a false-discovery rate (Q) = 5% without assuming consistent standard deviations [141]. EAE clinical severity was analyzed using non-compartmental estimation of the area under the curve (AUC) using the log-trapezoidal rule. Median EAE duration and Treg infiltration was compared using the Mann–Whitney U test due to non-normal distribution. Disease-free survival and overall survival Kaplan–Meier estimations were compared using the log-rank Mantel-Cox test with one degree of freedom, and hazard ratios were calculated using the Mantel–Haenszel method [142]. MOG35-55 stimulation-induced T cell proliferation was analyzed using two-way ANOVA with between-subject factors of genotype and within-subject factors of peptide concentration. Where Student’s t tests and ANOVA were performed, normality was tested using the D’Agostino and Pearson test with p = 0.05. Variance was compared using an F test for unpaired t-tests and the Bartlett’s test for ANOVA. Fisher’s exact test was performed on contingency tables to determine statistical significance of differences in tumor grade between different genotypes. P values <0.05 were considered significant, except in cases where adjusted q values <0.05 were used to account for false discovery. For calculations of total cell counts, outliers were removed using robust nonlinear regression [143] with Q = 1%. All analyses were performed in Prism 7 (GraphPad Software). Data are presented as mean ± standard error of the mean (s.e.m.).
Data availability
References
Galluzzi L, Buqué A, Kepp O, Zitvogel L, Kroemer G. Immunogenic cell death in cancer and infectious disease. Nat Rev Immunol. 2017;17:97–111.
Fuchs Y, Steller H. Programmed cell death in animal development and disease. Cell 2011;147:742–58.
van Kempen TS, Wenink MH, Leijten EFA, Radstake TRDJ, Boes M. Perception of self: distinguishing autoimmunity from autoinflammation. Nat Rev Rheumatol. 2015;11:483–92.
Medzhitov R, Janeway CA. Decoding the patterns of self and nonself by the innate immune system. Science. 2002;296:298–300.
Thompson CB. Apoptosis in the pathogenesis and treatment of disease. Science 1995;267:1456–62.
Arandjelovic S, Ravichandran KS. Phagocytosis of apoptotic cells in homeostasis. Nat Immunol. 2015;16:907–17.
Blander JM. The many ways tissue phagocytes respond to dying cells. Immunol Rev. 2017;277:158–73.
Elliott MR, Ravichandran KS. Clearance of apoptotic cells: implications in health and disease. J Cell Biol. 2010;189:1059–70.
Garza KM, Chan SM, Suri R, Nguyen LT, Odermatt B, Schoenberger SP, et al. Role of Antigen-Presenting Cells in Mediating Tolerance and Autoimmunity. J Exp Med. 2000;191:2021–8.
Cao X. Self-regulation and cross-regulation of pattern-recognition receptor signalling in health and disease. Nat Rev Immunol. 2016;16:35–50.
Latz E, Xiao TS, Stutz A. Activation and regulation of the inflammasomes. Nat Rev Immunol. 2013;13:397–411.
Vénéreau E, Ceriotti C, Bianchi ME. DAMPs from Cell Death to New Life. Front Immunol. 2015;6:422.
Albert ML, Jegathesan M, Darnell RB. Dendritic cell maturation is required for the cross-tolerization of CD8+ T cells. Nat Immunol. 2001;2:1010–7.
Steinman RM, Turley S, Mellman I, Inaba K. The Induction of Tolerance by Dendritic Cells That Have Captured Apoptotic Cells. J Exp Med. 2000;191:411–6.
Kim R, Hashimoto A, Markosyan N, Tyurin VA, Tyurina YY, Kar G, et al. Ferroptosis of tumour neutrophils causes immune suppression in cancer. Nature 2022;612:338–46.
Muñoz-Fontela C, Mandinova A, Aaronson SA, Lee SW. Emerging roles of p53 and other tumour-suppressor genes in immune regulation. Nat Rev Immunol. 2016;16:741–50.
Yoon KW, Byun S, Kwon E, Hwang SY, Chu K, Hiraki M, et al. Control of signaling-mediated clearance of apoptotic cells by the tumor suppressor p53. Science 2015;349:1261669.
Mori T, Anazawa Y, Iiizumi M, Fukuda S, Nakamura Y, Arakawa H. Identification of the interferon regulatory factor 5 gene (IRF-5) as a direct target for p53. Oncogene 2002;21:2914–8.
Rivas C, Aaronson SA, Munoz-Fontela C. Dual Role of p53 in Innate Antiviral Immunity. Viruses 2010;2:298–313.
Watanabe M, Moon KD, Vacchio MS, Hathcock KS, Hodes RJ. Downmodulation of tumor suppressor p53 by T cell receptor signaling is critical for antigen-specific CD4(+) T cell responses. Immunity 2014;40:681–91.
Balomenos D, Martín-Caballero J, García MI, Prieto I, Flores JM, Serrano M, et al. The cell cycle inhibitor p21 controls T-cell proliferation and sex-linked lupus development. Nat Med. 2000;6:171–6.
Bouillet P, Metcalf D, Huang DC, Tarlinton DM, Kay TW, Köntgen F, et al. Proapoptotic Bcl-2 relative Bim required for certain apoptotic responses, leukocyte homeostasis, and to preclude autoimmunity. Science 1999;286:1735–8.
Hussain SP, Amstad P, Raja K, Ambs S, Nagashima M, Bennett WP, et al. Increased p53 mutation load in noncancerous colon tissue from ulcerative colitis: a cancer-prone chronic inflammatory disease. Cancer Res. 2000;60:3333–7.
Liu L, Tran E, Zhao Y, Huang Y, Flavell R, Lu B. Gadd45 beta and Gadd45 gamma are critical for regulating autoimmunity. J Exp Med. 2005;202:1341–7.
Okuda Y, Okuda M, Bernard CCA. Regulatory role of p53 in experimental autoimmune encephalomyelitis. J Neuroimmunol. 2003;135:29–37.
Salvador JM, Hollander MC, Nguyen AT, Kopp JB, Barisoni L, Moore JK, et al. Mice lacking the p53-effector gene Gadd45a develop a lupus-like syndrome. Immunity 2002;16:499–508.
Santiago-Raber ML, Lawson BR, Dummer W, Barnhouse M, Koundouris S, Wilson CB, et al. Role of cyclin kinase inhibitor p21 in systemic autoimmunity. J Immunol. 2001;167:4067–74.
Singer GG, Carrera AC, Marshak-Rothstein A, Martínez C, Abbas AK. Apoptosis, Fas and systemic autoimmunity: the MRL-lpr/lpr model. Curr Opin Immunol. 1994;6:913–20.
Takatori H, Kawashima H, Suzuki K, Nakajima H. Role of p53 in systemic autoimmune diseases. Crit Rev Immunol. 2014;34:509–16.
Tapinos NI, Polihronis M, Moutsopoulos HM. Lymphoma development in Sjögren’s syndrome: novel p53 mutations. Arthritis Rheum. 1999;42:1466–72.
Yamanishi Y, Boyle DL, Rosengren S, Green DR, Zvaifler NJ, Firestein GS. Regional analysis of p53 mutations in rheumatoid arthritis synovium. Proc Natl Acad Sci USA. 2002;99:10025–30.
Zheng SJ, Lamhamedi-Cherradi SE, Wang P, Xu L, Chen YH. Tumor suppressor p53 inhibits autoimmune inflammation and macrophage function. Diabetes 2005;54:1423–8.
Chen W, Frank ME, Jin W, Wahl SM. TGF-beta released by apoptotic T cells contributes to an immunosuppressive milieu. Immunity 2001;14:715–25.
Ferguson TA, Herndon J, Elzey B, Griffith TS, Schoenberger S, Green DR. Uptake of apoptotic antigen-coupled cells by lymphoid dendritic cells and cross-priming of CD8(+) T cells produce active immune unresponsiveness. J Immunol. 2002;168:5589–95.
Gao Y, Herndon JM, Zhang H, Griffith TS, Ferguson TA. Antiinflammatory effects of CD95 ligand (FasL)-induced apoptosis. J Exp Med. 1998;188:887–96.
Griffith TS, Brunner T, Fletcher SM, Green DR, Ferguson TA. Fas ligand-induced apoptosis as a mechanism of immune privilege. Science 1995;270:1189–92.
Griffith TS, Yu X, Herndon JM, Green DR, Ferguson TA. CD95-induced apoptosis of lymphocytes in an immune privileged site induces immunological tolerance. Immunity 1996;5:7–16.
Schwarz A, Grabbe S, Grosse-Heitmeyer K, Roters B, Riemann H, Luger TA, et al. Ultraviolet light-induced immune tolerance is mediated via the Fas/Fas-ligand system. J Immunol. 1998;160:4262–70.
Guo G, Cui Y. New perspective on targeting the tumor suppressor p53 pathway in the tumor microenvironment to enhance the efficacy of immunotherapy. J Immunother Cancer. 2015;3:9.
Menendez D, Shatz M, Resnick MA. Interactions between the tumor suppressor p53 and immune responses. Curr Opin Oncol. 2013;25:85–92.
Cortez MA, Ivan C, Valdecanas D, Wang X, Peltier HJ, Ye Y, et al. PDL1 Regulation by p53 via miR-34. J Natl Cancer Inst. 2016;108:djv303.
Komarova EA, Krivokrysenko V, Wang K, Neznanov N, Chernov MV, Komarov PG, et al. p53 is a suppressor of inflammatory response in mice. FASEB J. 2005;19:1030–2.
Lowe JM, Menendez D, Bushel PR, Shatz M, Kirk EL, Troester MA, et al. p53 and NF-κB coregulate proinflammatory gene responses in human macrophages. Cancer Res. 2014;74:2182–92.
Sullivan A, Lu X. ASPP: a new family of oncogenes and tumour suppressor genes. Br J Cancer. 2007;96:196–200.
Bergamaschi D, Samuels Y, O’Neil NJ, Trigiante G, Crook T, Hsieh JK, et al. iASPP oncoprotein is a key inhibitor of p53 conserved from worm to human. Nat Genet. 2003;33:162–7.
Notari M, Hu Y, Koch S, Lu M, Ratnayaka I, Zhong S, et al. Inhibitor of apoptosis-stimulating protein of p53 (iASPP) prevents senescence and is required for epithelial stratification. PNAS 2011;108:16645–50.
Herron BJ, Rao C, Liu S, Laprade L, Richardson JA, Olivieri E, et al. A mutation in NFkB interacting protein 1 results in cardiomyopathy and abnormal skin development in wa3 mice. Hum Mol Genet. 2005;14:667–77.
Yang JP, Hori M, Sanda T, Okamoto T. Identification of a Novel Inhibitor of Nuclear Factor-κB, RelA-associated Inhibitor. J Biol Chem. 1999;274:15662–70.
Notari M, Hu Y, Sutendra G, Dedeić Z, Lu M, Dupays L, et al. iASPP, a previously unidentified regulator of desmosomes, prevents arrhythmogenic right ventricular cardiomyopathy (ARVC)-induced sudden death. PNAS 2015;112:E973–81.
Dedeić Z, Sutendra G, Hu Y, Chung K, Slee EA, White MJ, et al. Cell autonomous role of iASPP deficiency in causing cardiocutaneous disorders. Cell Death Differ. 2018;25:1289–303.
Havran WL, Jameson JM. Epidermal T Cells and Wound Healing. J Immunol. 2010;184:5423–8.
Jameson J, Ugarte K, Chen N, Yachi P, Fuchs E, Boismenu R, et al. A role for skin gammadelta T cells in wound repair. Science 2002;296:747–9.
Nosbaum A, Prevel N, Truong HA, Mehta P, Ettinger M, Scharschmidt TC, et al. Cutting Edge: Regulatory T Cells Facilitate Cutaneous Wound Healing. J Immunol. 2016;196:2010–4.
Toulon A, Breton L, Taylor KR, Tenenhaus M, Bhavsar D, Lanigan C, et al. A role for human skin–resident T cells in wound healing. J Exp Med. 2009;206:743–50.
Campuzano O, Alcalde M, Iglesias A, Barahona-Dussault C, Sarquella-Brugada G, Benito B, et al. Arrhythmogenic right ventricular cardiomyopathy: severe structural alterations are associated with inflammation. J Clin Pathol. 2012;65:1077–83.
Chen S, Wu J, Zhong S, Li Y, Zhang P, Ma J, et al. iASPP mediates p53 selectivity through a modular mechanism fine-tuning DNA recognition. Proc Natl Acad Sci USA. 2019;116:17470–9.
Jung DJ, Jin DH, Hong SW, Kim JE, Shin JS, Kim D, et al. Foxp3 expression in p53-dependent DNA damage responses. J Biol Chem. 2010;285:7995–8002.
Kawashima H, Takatori H, Suzuki K, Iwata A, Yokota M, Suto A, et al. Tumor suppressor p53 inhibits systemic autoimmune diseases by inducing regulatory T cells. J Immunol. 2013;191:3614–23.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell 2011;144:646–74.
Schreiber RD, Old LJ, Smyth MJ. Cancer immunoediting: integrating immunity’s roles in cancer suppression and promotion. Science 2011;331:1565–70.
Weinberg RA. Coming full circle-from endless complexity to simplicity and back again. Cell 2014;157:267–71.
Zitvogel L, Tesniere A, Kroemer G. Cancer despite immunosurveillance: immunoselection and immunosubversion. Nat Rev Immunol. 2006;6:715–27.
Karamitopoulou E. Tumour microenvironment of pancreatic cancer: immune landscape is dictated by molecular and histopathological features. Br J Cancer. 2019;121:5–14.
Guerra C, Schuhmacher AJ, Cañamero M, Grippo PJ, Verdaguer L, Pérez-Gallego L, et al. Chronic pancreatitis is essential for induction of pancreatic ductal adenocarcinoma by K-Ras oncogenes in adult mice. Cancer Cell. 2007;11:291–302.
Anz D, Mueller W, Golic M, Kunz WG, Rapp M, Koelzer VH, et al. CD103 is a hallmark of tumor-infiltrating regulatory T cells. Int J Cancer. 2011;129:2417–26.
De Simone M, Arrigoni A, Rossetti G, Gruarin P, Ranzani V, Politano C, et al. Transcriptional Landscape of Human Tissue Lymphocytes Unveils Uniqueness of Tumor-Infiltrating T Regulatory Cells. Immunity 2016;45:1135–47.
Plitas G, Konopacki C, Wu K, Bos PD, Morrow M, Putintseva EV, et al. Regulatory T Cells Exhibit Distinct Features in Human Breast Cancer. Immunity 2016;45:1122–34.
Schenkel JM, Herbst RH, Canner D, Li A, Hillman M, Shanahan SL, et al. Conventional type I dendritic cells maintain a reservoir of proliferative tumor-antigen specific TCF-1+ CD8+ T cells in tumor-draining lymph nodes. Immunity 2021;54:2338–2353.e6.
Connolly KA, Kuchroo M, Venkat A, Khatun A, Wang J, William I, et al. A reservoir of stem-like CD8+ T cells in the tumor-draining lymph node preserves the ongoing antitumor immune response. Sci Immunol. 2021;6:eabg7836.
Spear S, Candido JB, McDermott JR, Ghirelli C, Maniati E, Beers SA, et al. Discrepancies in the Tumor Microenvironment of Spontaneous and Orthotopic Murine Models of Pancreatic Cancer Uncover a New Immunostimulatory Phenotype for B Cells. Front Immunol [Internet]. [cited 2022 Mar 31]. 2019;10. https://www.frontiersin.org/article/10.3389/fimmu.2019.00542
Torphy RJ, Schulick RD, Zhu Y. Understanding the immune landscape and tumor microenvironment of pancreatic cancer to improve immunotherapy. Mol Carcinogenesis. 2020;59:775–82.
Pardoll DM. The blockade of immune checkpoints in cancer immunotherapy. Nat Rev Cancer. 2012;12:252–64.
Zou W, Wolchok JD, Chen L. PD-L1 (B7-H1) and PD-1 pathway blockade for cancer therapy: Mechanisms, response biomarkers, and combinations. Sci Transl Med. 2016;8:328rv4.
Karin M. Inflammation and cancer: the long reach of Ras. Nat Med. 2005;11:20–1.
Kortlever RM, Sodir NM, Wilson CH, Burkhart DL, Pellegrinet L, Swigart LB, et al. Myc Cooperates with Ras by Programming Inflammation and Immune Suppression. Cell 2017;171:1301–1315.e14.
Meylan E, Dooley AL, Feldser DM, Shen L, Turk E, Ouyang C, et al. Requirement for NF-κB signalling in a mouse model of lung adenocarcinoma. Nature 2009;462:104–7.
Dieu-Nosjean MC, Antoine M, Danel C, Heudes D, Wislez M, Poulot V, et al. Long-Term Survival for Patients With Non–Small-Cell Lung Cancer With Intratumoral Lymphoid Structures. JCO. 2008;26:4410–7.
Germain C, Gnjatic S, Tamzalit F, Knockaert S, Remark R, Goc J, et al. Presence of B cells in tertiary lymphoid structures is associated with a protective immunity in patients with lung cancer. Am J Respir Crit Care Med. 2014;189:832–44.
Hiraoka N, Ino Y, Yamazaki-Itoh R. Tertiary Lymphoid Organs in Cancer Tissues. Front Immunol [Internet]. 2016;7. http://www.ncbi.nlm.nih.gov/pmc/articles/PMC4916185/.
Joshi NS, Akama-Garren EH, Lu Y, Lee DY, Chang GP, Li A, et al. Regulatory T Cells in Tumor-Associated Tertiary Lymphoid Structures Suppress Anti-tumor T Cell Responses. Immunity 2015;43:579–90.
de Chaisemartin L, Goc J, Damotte D, Validire P, Magdeleinat P, Alifano M, et al. Characterization of chemokines and adhesion molecules associated with T cell presence in tertiary lymphoid structures in human lung cancer. Cancer Res. 2011;71:6391–9.
Bailey P, Chang DK, Nones K, Johns AL, Patch AM, Gingras MC, et al. Genomic analyses identify molecular subtypes of pancreatic cancer. Nature 2016;531:47–52.
Greten FR, Grivennikov SI. Inflammation and Cancer: Triggers, Mechanisms, and Consequences. Immunity 2019;51:27–41.
Dong P, Ihira K, Hamada J, Watari H, Yamada T, Hosaka M, et al. Reactivating p53 functions by suppressing its novel inhibitor iASPP: a potential therapeutic opportunity in p53 wild-type tumors. Oncotarget 2015;6:19968–75.
Gordon SR, Maute RL, Dulken BW, Hutter G, George BM, McCracken MN, et al. PD-1 expression by tumour-associated macrophages inhibits phagocytosis and tumour immunity. Nature 2017;545:495–9.
Daley D, Zambirinis CP, Seifert L, Akkad N, Mohan N, Werba G, et al. γδ T Cells Support Pancreatic Oncogenesis by Restraining αβ T Cell Activation. Cell 2016;166:1485–1499.e15.
Lu M, Breyssens H, Salter V, Zhong S, Hu Y, Baer C, et al. Restoring p53 function in human melanoma cells by inhibiting MDM2 and cyclin B1/CDK1-phosphorylated nuclear iASPP. Cancer Cell. 2013;23:618–33.
Slee EA, Gillotin S, Bergamaschi D, Royer C, Llanos S, Ali S, et al. The N-terminus of a novel isoform of human iASPP is required for its cytoplasmic localization. Oncogene 2004;23:9007–16.
DuPage M, Jacks T. Genetically engineered mouse models of cancer reveal new insights about the antitumor immune response. Curr Opin Immunol. 2013;25:192–9.
Bosurgi L, Hughes LD, Rothlin CV, Ghosh S. Death begets a new beginning. Immunol Rev. 2017;280:8–25.
Chan O, Shlomchik MJ. A new role for B cells in systemic autoimmunity: B cells promote spontaneous T cell activation in MRL-lpr/lpr mice. J Immunol. 1998;160:51–9.
Chu JL, Drappa J, Parnassa A, Elkon KB. The defect in Fas mRNA expression in MRL/lpr mice is associated with insertion of the retrotransposon, ETn. J Exp Med. 1993;178:723–30.
Fields ML, Nish SA, Hondowicz BD, Metzgar MH, Wharton GN, Caton AJ, et al. The influence of effector T cells and Fas ligand on lupus-associated B cells. J Immunol. 2005;175:104–11.
Fisher GH, Rosenberg FJ, Straus SE, Dale JK, Middleton LA, Lin AY, et al. Dominant interfering Fas gene mutations impair apoptosis in a human autoimmune lymphoproliferative syndrome. Cell 1995;81:935–46.
Rieux-Laucat F, Le Deist F, Hivroz C, Roberts IA, Debatin KM, Fischer A, et al. Mutations in Fas associated with human lymphoproliferative syndrome and autoimmunity. Science 1995;268:1347–9.
Volpe E, Sambucci M, Battistini L, Borsellino G. Fas–Fas Ligand: Checkpoint of T Cell Functions in Multiple Sclerosis. Front Immunol [Internet]. Sep 27 [cited 2018 Jul 31]. 2016;7. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5037862/.
Watanabe-Fukunaga R, Brannan CI, Copeland NG, Jenkins NA, Nagata S. Lymphoproliferation disorder in mice explained by defects in Fas antigen that mediates apoptosis. Nature 1992;356:314–7.
Yamada A, Arakaki R, Saito M, Kudo Y, Ishimaru N. Dual Role of Fas/FasL-Mediated Signal in Peripheral Immune Tolerance. Front Immunol [Internet]. Apr 5 [cited 2018 Jul 30]. 2017;8. https://www.ncbi.nlm.nih.gov/pmc/articles/PMC5380675/.
Zhang Y, Liu Q, Zhang M, Yu Y, Liu X, Cao X. Fas Signal Promotes Lung Cancer Growth by Recruiting Myeloid-Derived Suppressor Cells via Cancer Cell-Derived PGE2. J Immunol. 2009;182:3801–8.
Singh N, Yamamoto M, Takami M, Seki Y, Takezaki M, Mellor AL, et al. CD4(+)CD25(+) regulatory T cells resist a novel form of CD28- and Fas-dependent p53-induced T cell apoptosis. J Immunol. 2010;184:94–104.
Flies DB, Han X, Higuchi T, Zheng L, Sun J, Ye JJ, et al. Coinhibitory receptor PD-1H preferentially suppresses CD4+ T cell–mediated immunity. J Clin Investig. 2014;124:1966–75.
Bennett M, Macdonald K, Chan SW, Luzio JP, Simari R, Weissberg P. Cell Surface Trafficking of Fas: A Rapid Mechanism of p53-Mediated Apoptosis. Science 1998;282:290–3.
Hangai S, Ao T, Kimura Y, Matsuki K, Kawamura T, Negishi H, et al. PGE2 induced in and released by dying cells functions as an inhibitory DAMP. PNAS 2016;113:3844–9.
Hou J, Greten TF, Xia Q. Immunosuppressive cell death in cancer. Nat Rev Immunol. 2017;17:nri.2017.46.
Garg AD, Agostinis P. Cell death and immunity in cancer: From danger signals to mimicry of pathogen defense responses. Immunol Rev. 2017;280:126–48.
Xue W, Zender L, Miething C, Dickins RA, Hernando E, Krizhanovsky V, et al. Senescence and tumour clearance is triggered by p53 restoration in murine liver carcinomas. Nature 2007;445:656–60.
Coffelt SB, Kersten K, Doornebal CW, Weiden J, Vrijland K, Hau CS, et al. IL-17-producing γδ T cells and neutrophils conspire to promote breast cancer metastasis. Nature 2015;522:345–8.
Hiratsuka S, Watanabe A, Aburatani H, Maru Y. Tumour-mediated upregulation of chemoattractants and recruitment of myeloid cells predetermines lung metastasis. Nat Cell Biol. 2006;8:1369–75.
Takaoka A, Hayakawa S, Yanai H, Stoiber D, Negishi H, Kikuchi H, et al. Integration of interferon-alpha/beta signalling to p53 responses in tumour suppression and antiviral defence. Nature 2003;424:516–23.
Sharma MD, Rodriguez PC, Koehn BH, Baban B, Cui Y, Guo G, et al. Activation of p53 in Immature Myeloid Precursor Cells Controls Differentiation into Ly6c+CD103+ Monocytic Antigen-Presenting Cells in Tumors. Immunity 2018;48:91–106.e6.
Falik-Zaccai TC, Barsheshet Y, Mandel H, Segev M, Lorber A, Gelberg S, et al. Sequence variation in PPP1R13L results in a novel form of cardio-cutaneous syndrome. EMBO Mol Med. 2017;9:319–36.
Laska MJ, Lowe SW, Zender L, Hearn S, Vogel U, Jensen UB, et al. Enforced expression of PPP1R13L increases tumorigenesis and invasion through p53-dependent and p53-independent mechanisms. Mol Carcinog. 2009;48:832–42.
Oh H, Grinberg-Bleyer Y, Liao W, Maloney D, Wang P, Wu Z, et al. An NF-κB Transcription-Factor-Dependent Lineage-Specific Transcriptional Program Promotes Regulatory T Cell Identity and Function. Immunity 2017;47:450–465.e5.
Moussawi K, Chung K, Carroll T, Osterburg C, Smirnov A, Lotz R, et al. Mutant Ras and inflammation-driven skin tumorigenesis is suppressed via a JNK-iASPP-AP1 axis. Cell Rep. 2022;41:111503.
Wang D, DuBois RN. Immunosuppression associated with chronic inflammation in the tumor microenvironment. Carcinogenesis 2015;36:1085–93.
Coussens LM, Werb Z. Inflammation and cancer. Nature 2002;420:860–7.
Grivennikov SI, Greten FR, Karin M. Immunity, Inflammation, and Cancer. Cell. 2010;140:883–99.
Sottocornola R, Royer C, Vives V, Tordella L, Zhong S, Wang Y, et al. ASPP2 binds Par-3 and controls the polarity and proliferation of neural progenitors during CNS development. Dev Cell. 2010;19:126–37.
Jackson EL, Willis N, Mercer K, Bronson RT, Crowley D, Montoya R, et al. Analysis of lung tumor initiation and progression using conditional expression of oncogenic K-ras. Genes Dev. 2001;15:3243–8.
Hingorani SR, Petricoin EF, Maitra A, Rajapakse V, King C, Jacobetz MA, et al. Preinvasive and invasive ductal pancreatic cancer and its early detection in the mouse. Cancer Cell. 2003;4:437–50.
Olive KP, Tuveson DA, Ruhe ZC, Yin B, Willis NA, Bronson RT, et al. Mutant p53 Gain of Function in Two Mouse Models of Li-Fraumeni Syndrome. Cell 2004;119:847–60.
Srinivas S, Watanabe T, Lin CS, William CM, Tanabe Y, Jessell TM, et al. Cre reporter strains produced by targeted insertion of EYFP and ECFP into the ROSA26 locus. BMC Dev Biol. 2001;1:4.
Vooijs M, Jonkers J, Berns A. A highly efficient ligand‐regulated Cre recombinase mouse line shows that LoxP recombination is position dependent. EMBO Rep. 2001;2:292–7.
DuPage M, Dooley AL, Jacks T. Conditional mouse lung cancer models using adenoviral or lentiviral delivery of Cre recombinase. Nat Protoc. 2009;4:1064–72.
Jackson EL, Olive KP, Tuveson DA, Bronson R, Crowley D, Brown M, et al. The Differential Effects of Mutant p53 Alleles on Advanced Murine Lung Cancer. Cancer Res. 2005;65:10280–8.
Hruban RH, Adsay NV, Albores-Saavedra J, Anver MR, Biankin AV, Boivin GP, et al. Pathology of genetically engineered mouse models of pancreatic exocrine cancer: consensus report and recommendations. Cancer Res. 2006;66:95–106.
DuPage M, Mazumdar C, Schmidt LM, Cheung AF, Jacks T. Expression of tumour-specific antigens underlies cancer immunoediting. Nature 2012;482:405–9.
Nikitin AY, Alcaraz A, Anver MR, Bronson RT, Cardiff RD, Dixon D, et al. Classification of proliferative pulmonary lesions of the mouse: recommendations of the mouse models of human cancers consortium. Cancer Res. 2004;64:2307–16.
Salter J. blindanalysis: v1.0 [Internet]. Zenodo; 2016. https://zenodo.org/record/44678#.WarFbdN9560.
Tellier M, Zaborowska J, Caizzi L, Mohammad E, Velychko T, Schwalb B, et al. CDK12 globally stimulates RNA polymerase II transcription elongation and carboxyl-terminal domain phosphorylation. Nucleic Acids Res. 2020;48:7712–27.
Suzuki A, Makinoshima H, Wakaguri H, Esumi H, Sugano S, Kohno T, et al. Aberrant transcriptional regulations in cancers: genome, transcriptome and epigenome analysis of lung adenocarcinoma cell lines. Nucleic Acids Res. 2014;42:13557–72.
Yevshin I, Sharipov R, Valeev T, Kel A, Kolpakov F. GTRD: a database of transcription factor binding sites identified by ChIP-seq experiments. Nucleic Acids Res. 2017;45:D61–7.
Nojima T, Gomes T, Carmo-Fonseca M, Proudfoot NJ. Mammalian NET-seq analysis defines nascent RNA profiles and associated RNA processing genome-wide. Nat Protoc. 2016;11:413–28.
Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011;17:10–2.
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013;14:R36.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009;25:2078–9.
Nojima T, Gomes T, Grosso ARF, Kimura H, Dye MJ, Dhir S, et al. Mammalian NET-Seq Reveals Genome-wide Nascent Transcription Coupled to RNA Processing. Cell 2015;161:526–40.
Quinlan AR, Hall IM. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics 2010;26:841–2.
Huang DW, Sherman BT, Lempicki RA. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat Protoc. 2009;4:44–57.
Huang DW, Sherman BT, Lempicki RA. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009;37:1–13.
Benjamini Y, Krieger AM, Yekutieli D. Adaptive linear step-up procedures that control the false discovery rate. Biometrika 2006;93:491–507.
Bernstein L, Anderson J, Pike MC. Estimation of the proportional hazard in two-treatment-group clinical trials. Biometrics 1981;37:513–9.
Motulsky HJ, Brown RE. Detecting outliers when fitting data with nonlinear regression – a new method based on robust nonlinear regression and the false discovery rate. BMC Bioinforma. 2006;7:123.
Acknowledgements
We would like to thank M. Shipman for microscopy assistance, J. Thompson for performing gamma irradiation, and S. McCuaig and F. Powrie for generously providing flow cytometry reagents and helpful advice. We thank the High-Throughput Genomics Group at the Wellcome Trust Centre for Human Genetics (funded by Wellcome Trust grant reference 203141/Z/16/Z) for the generation of the sequencing data.
Funding
This work is supported by the Ludwig Institute for Cancer Research and Rhodes Scholarships to EAG and TMC. EAG was supported by National Institutes of Health/National Institute of General Medical Sciences Grant T32GM007753 and by National Institutes of Health/National Institute of Allergy and Infectious Diseases Grants T32AI007529 and F30AI160909. SM, JZ, and MT were supported by grants WT106134AIA and WT210641/Z/18/Z from the Wellcome Trust to SM.
Author information
Authors and Affiliations
Contributions
Conceptualization, EAG, PM, XL; Methodology, EAG, PM, TC, MT; Software, EAG, TC, MT; Validation, EAG; Formal Analysis, EAG, PM, TC, MT, RG; Investigation, EAG, PM, TC, GS, LB, MT, RG, ES, JZ; Data Curation, EAG, PM, TC; Writing – Original Draft, EAG; Visualization, EAG, TC, MT; Supervision, FGS, SM, XL; Funding Acquisition, XL.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by Professor Gerry Melino
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Akama-Garren, E.H., Miller, P., Carroll, T.M. et al. Regulation of immunological tolerance by the p53-inhibitor iASPP. Cell Death Dis 14, 84 (2023). https://doi.org/10.1038/s41419-023-05567-9
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-023-05567-9
This article is cited by
-
p53 inhibitor iASPP is an unexpected suppressor of KRAS and inflammation-driven pancreatic cancer
Cell Death & Differentiation (2023)