Abstract
Circular RNAs (circRNAs) are a class of endogenous RNAs characterized by a covalent loop structure. In comparison to other types of RNAs, the abundance of circRNAs is relatively low but due to the circular configuration, their stability is very high. In addition, circRNAs display high degree of tissue specificity. The sponging activity of circRNAs toward microRNAs is the best-described mode of action of circRNAs. However, the ability of circRNAs to bind with specific proteins, as well as to encode short proteins, propose alternative functions. This review introduces the biogenesis of circRNAs and summarizes the roles played by circRNAs in human diseases. These include examples of their functional roles in several organ-specific cancers, such as head and neck and breast and lung cancers. In addition, we review potential functions of circRNAs in diabetes, cardiovascular, and neurodegenerative diseases. Recently, a growing number of studies have demonstrated involvement of circRNAs in a wide spectrum of signaling molecular pathways, but at the same time many different and controversial views on circRNAs role and function are emerging. We conclude by offering cellular homeostasis generated by networks comprising circular RNAs, other non-coding RNAs and RNA-binding proteins. Accordingly, it is predictable that circRNAs, due to their highly stable nature and remarkable tissue specificity, will emerge as reliable biomarkers of disease course and treatment efficacy.
Similar content being viewed by others
Facts
-
circRNAs are single-stranded circles of RNA, which form highly stable closed loops.
-
circRNAs can have different functions. Among these, the miRNA sponging is the best-characterized role.
-
circRNAs are widely expressed in human tissues and their expression is highly tissue-specific.
-
circRNAs are involved in many human diseases, including cancer and neurodegenerative disorders.
-
The biochemical characteristics of circRNAs, especially stability and tissue specificity, make them ideal biomarkers for clinical use.
Open questions
-
Are there yet unknown features controlling circRNA biogenesis?
-
Are there still undiscovered functional aspects and mechanisms of circRNAs?
-
How do circRNAs function in already well-characterized molecular pathways?
-
To what extent would deeper understanding and utilization of circRNAs help improve human health?
Introduction
Circular RNAs (circRNAs) are covalently closed circular RNA molecules recently reconsidered for their important roles in cancer and in other human diseases1,2,3. Since 2013, when Memczak et al.4 reported that circRNAs act as post-transcriptional regulators, additional circRNAs have been identified, implying an important regulatory potential for this class of molecules. It is currently broadly recognized that circRNAs have significant roles to play in cell proliferation5,6,7, motility and metastasis5,6,7,8,9, as well as in cell cycle progression10, angiogenesis11,12, and apoptosis13.
To exemplify the biomedical potential of circRNAs, we briefly review the biogenesis of circRNAs and then describe the role of some circRNAs in head and neck squamous cell carcinoma, breast and lung cancer. Among neurological disorders, we focus herein on Alzheimer disease. In addition, we report examples of circRNAs playing functional roles in cardiovascular diseases and in diabetes. Although the sponging mechanism of circRNAs toward microRNAs has emerged as the most common mechanism of action, additional modes of action have been proposed. CircRNAs can interact with proteins and some are translated into novel polypeptides or act as transcriptional regulators2,14,15,16,17,18,19,20,21. Presumably, as many circRNAs are being characterized, additional modes of action will be uncovered soon. CircRNAs are involved in many signaling pathways and some of these molecular pathways have been already characterized for their important roles in human diseases and they are subjects of clinical trials22,23. These characteristics, together with their presence in accessible body fluids, such as saliva, blood, and urine, make the circRNAs promising therapeutic targets and potential biomarkers for human diseases24,25,26.
Biogenesis and function of circRNAs
CircRNAs can derive from exons, introns, antisense, 5′ or 3′ untranslated and intergenic genomic regions27. Exonic circRNAs (ecircRNAs) represent the most abundant species and they are produced by a “back-splicing” mechanism. During the biogenesis process, a downstream 5′ splice site of an exon is joined to an upstream 3′ splice site of the same or another exon, involving single or multiple exons1,28,29,30,31,32. The molecules derived from this mechanism form a closed circular transcript and an alternatively spliced linear RNA with skipped exons31. Thus, the mechanism that generates circRNAs uses the canonical spliceosomal machinery31. As a consequence, transcription of circRNAs competes with canonical pre-mRNA splicing and affects the rate of canonical gene expression28.
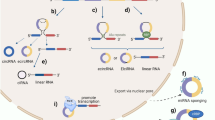
One of the best-described mechanisms explaining the biological function of circRNAs is the ability to effectively sponge microRNAs33 (Fig. 1). The circ-SRY33, CDR1as4,33,34, circ-ITCH35, circHIPK3 (ref. 36), circ_000984 (ref. 37), circ-TTBK2 (ref. 38), and circPVT1 (ref. 39) are examples of circRNAs that act as miRNA sponges (also called competing endogenous RNA; ceRNA). CircRNAs function as ceRNA via microRNA (miRNA) sequestration, by binding to miRNA response elements (MREs)4. Each circRNA can have many MREs on the miRNA target and the number of MREs is related to the length of the circRNAs themselves4.
CircRNAs can also interact with proteins (Fig. 1). Examples of circRNAs interacting with proteins include circ-PABPN14, circ-Foxo3 (ref. 10), and circ-Amotl1 (ref. 15). RNA-binding proteins (RBPs) specifically interact with RNA molecules to form ribonucleoprotein complexes14. The RBP HuR can bind circ-PABPN1 in human cervical carcinoma HeLa cells and it is responsible for the translation rate of the PABPN1 gene14 (Fig. 1). The RNA-binding protein quaking-5 (QKI-5) promotes circRNA biogenesis during epithelial-to-mesenchymal transition (EMT) through interaction with introns flanking the circRNA-forming exons16. Another protein that regulates circRNA biogenesis is the splicing factor called muscleblind (MBL). MBL promotes the formation of circMBL through interaction with introns flanking the circRNA itself28.
There are several examples of circRNA-protein interactions in the cancer context (Fig. 1). The tumor suppressor circ-Foxo3 interacts with CDK2 and p21 to form a ternary complex and inhibit cell cycle progression in cancer10. The oncogenic circRNA circ-Amotl1 promotes cell growth through an interaction with the proto-oncogene c-MYC. Circ-Amotl1 is able to increase the retention of nuclear c-MYC, promote c-myc stability, and up-regulating c-myc targets15.
There is also evidence showing that circRNAs can be translated into functional proteins (Fig. 1). Circ-ZNF609 is one of the first examples described of a circRNA that can be translated into a protein. Circ-ZNF609 is involved in the regulation of myoblast proliferation17.
The circular form of the SNF2 histone linker PHD RING helicase (SHPRH) gene, which encodes the protein SHPRH-146aa represents an additional example18. Both circ-SHPRH and SHPRH-146aa are highly expressed in normal human brains and their expression was found to be down-regulated in glioblastoma, suggesting a tumor suppressor role18. In a similar way, Zheng et al.19 identified circPPP1R12A, which is up-regulated in colon cancer (CC) and can be translated into a protein contributing to the rapid proliferation of CC cells via the Hippo-YAP pathway.
Finally, it is well accepted that intronic circRNAs (ciRNAs) and exon–intron circRNAs (ElciRNAs) can act as transcriptional regulators (Fig. 1)20,21. The intronic circRNA ci-ankrd52 is able to regulate its parental gene expression by modulating RNA polymerase II’s elongation activity21. Similarly, two ElciRNAs, circEIF3J and circPAIP2, are able to regulate the expression of their parental genes through a specific RNA–RNA interaction between U1 snRNA and the circRNA20. More recently Stoll et al.40 showed that the intronic circle ci-Ins2, located mainly in the nucleus of pancreatic β cells, is able to regulate insulin secretion through interaction with the TAR DNA-binding protein 43 kDa (TDP-43).
CircRNAs in head and neck squamous cell carcinoma
Head and neck cancers represent the sixth most common cancer worldwide41,42. This cancer usually initiates in the squamous cells that line the mucosal surfaces inside the head and neck and can arise from the mucosal surfaces of the oral cavity (OSCC), oropharynx (OPSCC), and larynx. Head and neck cancers can also begin in the salivary glands and in paranasal sinuses and nasal cavities41,42. We showed that the circRNA circPVT1 acts as an oncogene in head and neck squamous cell carcinoma (HNSCC)39. CircPVT1 expression is regulated through the mut-p53/YAP/TEAD complex binding its own promoter, which is independent from the host gene PVT1 promoter39. CircPVT1 is overexpressed in tumors compared to matched-non-tumoral tissues and its expression is particularly high in patients with TP53 mutations39. This is an example of a circRNA acting as an oncogene and modulating the expression of miR-497-5p and some of its targets, such as aurka, mki67, and bub1, all genes involved in the control of cell proliferation. This is in line with the known role of miR-497-5p as a tumor suppressor in several cancers39,43,44,45,46.
Using high-throughput sequencing and RT-qPCR, Li et al.47 showed that hsa_circ_0008309 is down-regulated in OSCC tissues relative to paired adjacent normal tissues (ANTs)47,48 and statistically correlated with pathological differentiation48. Intriguingly, bioinformatics analysis showed that hsa_circ_0008309 might function within a molecular network involving miR-1290, miR-136-5p, miR-382-5p and the ATXN1 gene, coding for the DNA-binding protein Ataxin-1 (refs. 47,48).
Xuan et al.8 analyzed the circRNA expression in a cohort of Laryngeal squamous cell carcinoma (LSCC) tissues. They found that two circRNAs, hsa_circRNA_100855 and hsa_circRNA_104912, were respectively up- and down-regulated in cancer tissues in comparison to the corresponding adjacent non-neoplastic tissues8. Patients with T3-4 stage, neck nodal metastasis, or advanced clinical stage had higher hsa_circRNA_100855 expression and a lower hsa_circRNA_104912 expression8,48.
CircHIPK3 is highly expressed in nasopharyngeal carcinoma (NPC)9. The silencing of circHIPK3 can reduce cell proliferation, migration, and invasion in vitro and it can repress tumor growth and metastasis in vivo9. CircHIPK3 functions in NPC by sponging the miR-4288, which in turn targets the E74-like ETS transcription factor 3 (ELF3)9. Studying the circHIPK3-miR-4288-ELF3 molecular pathway could instruct ways to identify new therapeutic strategies focused on this regulatory loop.
CircRNAs in breast cancer
Breast cancer is the most common cancer in females and it can be classified in three major cancer subtypes according to estrogen or progesterone receptor expression and ERBB2 gene amplification: hormone receptor positive/ERBB2 negative (HR+/ERBB2-), ERBB2 positive (ERBB2+), and triple-negative49.
Galasso et al. performed a pilot study in which they described one of the first panels of circRNAs expressed in breast cancer by analyzing RNA sequencing data from five paired breast cancer samples50. At the same time, Nair et al.51 developed an automated workflow called Circ-Seq to identify circRNAs in breast tumors and breast cancer cell lines. A recent work identified 235 differentially expressed circRNAs in breast cancer through high-throughput circRNA microarray analysis52. Among all the modulated circRNAs, circTADA2A-E6 (hsa_circ_0006220) and circTADA2A-E5/E6 (hsa_circ_0043278) were ranked in the top five down-regulated circRNAs by microarray analysis52. In particular, circTADA2A-E6 sponges miR-203a-3p and functions as a tumor suppressor by inhibiting cell proliferation, migration, and metastasis. The SOCS3 gene was predicted as a downstream target gene of the circTADA2A-E6/miR-203a-3p axis52, and a previous study reported that miR-203a-3p promotes cell proliferation by targeting SOCS3 in MCF-7 cells53. These results show that the circTADA2A-E6/miR-203a-3p/SOCS3 axis plays an important role in the inhibition of breast cancer progression.
CircRNA-000911 is another circRNA acting in breast cancer as a tumor suppressor54. Wang et al.54 showed that circRNA-000911 suppresses the proliferative, migratory, and invasive capacities of breast cancer cells by sponging miR-449a and releasing Notch1, a functional target of miR-449a. This mechanism includes the involvement of Ago2, an essential protein for circRNA sponge activity4,55. The consequence of circRNA-000911 down-regulation in breast cancer is the up-regulation of miR-449a and down-regulation of Notch1. Importantly, one downstream effector of Notch1 is the nuclear factor-kB (kB), which normally promotes breast cancer tumorigenesis and progression54.
The circRNA circEPSTI1 (hsa_ circRNA_000479) is up-regulated in breast cancer and it is a prognostic marker and mediator of triple-negative breast cancer (TNBC) progression56. CircEPSTI1 behaves as an oncogene promoting TNBC cell proliferation in vitro and in vivo, and it is able to sponge both miR-4753 and miR-6809 (ref. 56). BCL11A is a direct target gene of both miRNAs, and it is inhibited as a consequence of circEPSTI1 knockdown. It follows that the circEPSTI1-miR-4753/6809-BCL11A axis could be an interesting pathway to investigate in order to identify new therapeutic strategies for the treatment of TNBC56.
CircANKS1B is another circRNA up-regulated in TNBC and its expression is associated with both lymph node metastasis and advanced clinical stage57. CircANKS1B is able to sponge miR-148a-3p and miR-152-3p, thereby increases the expression of transcription factor USF1, which in turn up-regulates TGF-β1 expression57. The up-regulation of TGF-β1 results in activation of the TGF-β1/Smad signaling pathway, promoting epithelial-to-mesenchymal transition (EMT)57. The results suggest that circANKS1B is an interesting circRNAs to study in order to find alternative therapeutic strategies for inhibiting breast cancer metastasis.
CircRNAs in lung cancer
Lung cancer is one of the most common cancers in the world with 5-year survival rates varying from 92 to 0%, depending on disease stage at diagnosis58.
Circ-ITCH is generated from several exons of the ITCH E3 ubiquitin protein ligase (ITCH) and it shares the miR-7 and miR-214 binding sites with the three-prime untranslated regions (3′-UTR) of its parental gene ITCH59. Circ-ITCH plays an inhibitory role in lung cancer progression by sponging miR-7 and miR-214 and regulating the expression of ITCH59,60. ITCH negatively regulates canonical Wnt signaling by targeting the dishevelled-2 (Dvl2) protein61. In lung cancer the down-regulation of circ-ITCH brings to an increase of miR-7 and miR-214, thereby to a decrease of their target gene, ITCH. As a consequence, the Wnt/β-catenin pathway is enhanced, thereby promoting the development and progression of cancer59,60. Another circRNA that indirectly affects ITCH expression is hsa_circ_0043256. The circRNA hsa_circ_0043256 is able to sponge miR-1252, which binds the ITCH 3′-UTR62. Both circRNAs, circ-ITCH and hsa-circ_0043256, behave as tumor suppressors in lung cancer and their combined action could be used to design new strategies for the treatment of this malignancy.
In contrast to circ-ITCH, hsa_circ_0012673 is overexpressed in lung adenocarcinoma and promotes cell proliferation through the miR-22/ErbB3 pathway63. Hsa_circ_0012673 is able to sponge miR-22, which targets ERBB3/HER3, an important receptor tyrosine kinase in lung adenocarcinoma. ERBB3/HER3 is a member of the epidermal growth factor receptor (EGFR/ERBB) family64 and EGFR mutations were characterized for their important role in lung cancer65.
CircRNA in Alzheimer’s disease
Alzheimer’s disease (AD) is the most prevalent cause of dementia affecting millions of people worldwide66. AD is a progressive and neurodegenerative disorder characterized by widespread neuronal atrophy and two histopathological hallmarks: extracellular senile plaques consisting of amyloid-β peptides, and intracellular neurofibrillary tangles composed of abnormally hyperphosphorylated Tau protein66.
Dube et al.67 generated RNA-seq data from individuals with and without AD to quantify cortical circRNA expression. The results showed that there are significant associations between circRNA expression and AD diagnosis, clinical dementia severity, and neuropathological severity67. Interestingly, circRNA expression changes can be observed early on, in pre-symptomatic AD and in autosomal dominant AD67. The microtubule-associated Tau protein plays a central role in AD since it is responsible for amyloid-beta induced neuronal cell death68. The MAPT gene generates the Tau protein. Using a PCR screen of RNA from human brain tissues, Welden et al. showed that the MAPT locus generates circRNAs through a backsplicing mechanism, but the role of these circRNAs is still unclear69.
Similarly, CDR1as has been one of the first circRNAs that were characterized. It derives from the cerebellar degeneration-related protein 1 antisense transcript (CDR1AS) and contains over 70 conventional binding sites for miR-74,33,70. Down-regulation of CDR1as causes up-regulation of miR-7 and, consequently, negative regulation of its respective targets, such as ubiquitin protein ligase A (UBE2A)71,72,73. UBE2A is important for clearing amyloid peptides and it was found depleted in the AD brain71,72,73.
CircRNAs in cardiovascular diseases
RNA-Seq analysis of ribosome-depleted libraries from hearts of human, mouse, and rats origins, detected more than 9000 candidate circRNAs for each species74. A similar analysis listed more than 15,000 cardiac circRNAs in humans75. Although the study showed no statistically significant circRNA that was differentially expressed in diseased hearts compared to healthy hearts, other studies are needed to elucidate the role of circRNAs in cardiac diseases75. On the other hand, the analysis found significant differential expressed circRNAs during cardiomyocyte differentiation75.
Many of the identified cardiac circRNAs are yet uncharacterized in terms of their specific function. Nevertheless, the identification of cardiac circRNAs represents a potential strategy to use circRNAs as target molecules in the prevention and treatment of cardiovascular diseases.
The first circRNA described with a cardioprotective role was the heart-related circRNA, HRCR. This circRNA acts as a miR-223 sponge to inhibit cardiac hypertrophy and heart failure76. MiR-223 is able to suppress the expression level of its target, ARC, the apoptosis repressor with CARD domain protein. HRCR acts as an anti-hypertrophic molecule due to its sponging mechanism toward miR-223, which causes up-regulation of ARC76.
More recently, circFndc3b was identified as another circRNA involved in cardioprotection. CircFndc3b interacts with the RNA-binding protein Fused in Sarcoma (FUS) to regulate VEGF expression and signaling12. Acting on the FUS/VEGF-A axis, circFBDc3b is able to enhance angiogenesis and retard cardiomyocytes and endothelial cell apoptosis12.
Yet another circRNA, Cdr1as (ciRS-7), acts as a miR-7a sponge in myocardial cells77. It was shown that ciRS-7 induces apoptosis in myocardial infarction (MI) in mice by means of increasing caspase-3 activity. CiRS-7 is up-regulated in infarcted hearts, and it is able to inhibit the miR-7a mediated cardiomyocyte protection against MI injury acting as a miRNA sponge77,78. CiRS-7’s sponge mechanism toward miR-7a determines the up-regulation of two miR-7a targets, PARP and SP1. These proteins play pro-apoptotic roles during MI77.
MFACR (mitochondrial fission and apoptosis-related circRNA) regulates mitochondrial fission and apoptosis in the heart, while acting as a miRNA sponge for miR-652-3p79. MiR-652-3p down-regulates its target, MTP18, a nuclear-encoded mitochondrial membrane protein that contributes to mitochondrial fission in mammalian cells79,80. As a result, the MFACR-activated pathway instigates cardiomyocyte death through miR-652-3p-dependent up-regulation of MTP18 expression79.
Another circRNA involved in cardiomyocyte apoptotic events is circNCX1, which is generated from the sodium/calcium exchanger 1 (ncx1) gene81. circNCX1 acts as a miRNA sponge for miR-133a-3p, which is able to target the pro-apoptotic gene called Cell Death-Inducing p53-target Protein 1 (CDIP1). Importantly, miR-133a-3p plays a cardioprotective role and it is suppressed by the circNCX1 sponge mechanism81. This is an example of circRNAs that enhances damage following a MI event, primarily by promoting apoptosis of cardiomyocytes81.
CircRNAs in diabetes
Diabetes is a group of metabolic disorders all characterized by hyperglycemia, namely high levels of sugar in the blood. This condition is associated with various pathological states, such as cardiovascular disease, retinopathy, nephropathy, and neuropathy82.
A recent work showed the human pancreatic islets express thousands of circRNAs83. The circRNA Cdr1as is already known for its miR-7 sponging activity in embryonic zebrafish brains and in infarcted hearts4,34,84. Moreover, Cdr1as is able to affect miR-7 function in adult islet cells84. Xu et al.84 showed that miR-7 is highly expressed in islet cells, and its overexpression in transgenic mice β-cells causes diabetes due to impaired insulin secretion and β cell dedifferentiation. Cdr1as promotes insulin secretion by sponging miR-7 in islet cells84. Hence, the interaction between Cdr1as and miR-7 in insulin secretion may become a new therapeutic target for improving β cell function in diabetes84.
The circRNA circHIPK3 was found up-regulated in retinas and retinal endothelial cells of patients with diabetes. CircHIPK3 is able to regulate the retinal vascular endothelial function while sponging miR-30a-3p85. As a consequence of its action as miRNA sponge, circHIPK3 increases the expression of VEGFC, FZD4, and WNT2, leading to endothelial proliferation and vascular dysfunction85. It follows that circHIPK3 could serve as a valid target for diabetic retinopathy.
CircRNAs as potential disease biomarkers
Both prognostic biomarkers and markers that predict responses to a drug or other treatment modalities must bear high specificity for a given pathophysiological condition and a highly reproducible detection capacity. Accordingly, the renewed identification of circRNAs has opened a new potential strategy for diagnosis and for monitoring progression of different human diseases (Table 1). This is primarily due to the high tissue specificity of circRNAs, their relatively high stability in tissues and body fluids, as well as ease of detection using rather simple technologies, such as real-time PCR1,86,87. CircRNAs are highly abundant in blood25 and there are also evidences of circRNAs in urine samples26, for example to assist monitoring of patients who have undergone kidney transplantation, or for diagnosis of patients with bladder cancer26,88.
Importantly, several recent studies reported the presence of circRNAs in extracellular vesicles, mainly exosomes, which are targets for discovery of additional types of new biomarkers89,90,91. The abundance and diversity of circRNAs in human blood exosomes is already available in a database called exoRBase92. Likewise, circRNAs with diagnostic potential have been found in urine exosomes93,94. Additionally, another database, MiOncoCirc, was created based on sequencing of more than 2000 tumor samples, and many urine circRNAs were identified as possible biomarkers for prostate cancer95.
There are several papers that have shown correlations between expression of specific circRNAs and tumor grade, size, metastatic spread, and lymph node involvement. This is the case of hsa_circ_002059 and hsa_circ_0000190, which were found to be decreased in plasma of patients with gastric cancer96,97. Likewise, it has been reported that circFARSA is elevated in plasma of patients with non-small-cell lung cancer (NSCLC), in direct association with tumor cell aggressiveness in vitro98. The circRNA F-circEA-2a is another candidate biomarker in NSCLC. Generated from the prevalent fusion gene in lung cancer, EML4-ALK, circRNA F-circEA-2a appears elevated in plasma samples99.
A screening seeking differentially expressed circRNAs in plasma of patients with hepatocellular carcinoma related to the hepatitis B virus, reported elevated expression of hsa_circ_0027089 and classified it as a potential biomarker100. Additionally, an atlas of Blood-Based Biomarkers for Early Diagnosis of Cancers (BBcancer) has recently been established101. It includes data from 5000 samples across 15 different types of cancer101.
A recent study evaluated the presence of circRNAs in cerebrospinal fluid of patients with Alzheimer’s disease (AD) and found 112 up-regulated and 51 down-regulated circRNAs102. Some of these circRNAs were confirmed by real-time PCR, with circ-LPAR1, circ-AXL, and circ-GPHN elevated and circ-PCCA, circ-HAUS4, circ-KIF18B, and circ-TTC39C decreased in patients with AD102.
Another study showed that it is possible to differentiate patients with AD and healthy individuals by testing the expression of circRNAs in peripheral blood mononuclear cells (PMBCs)103. Hsa_circRNA_405619 and hsa_circRNA_000843 were shown to be elevated in PBMCs of patients with AD in comparison to healthy individuals, while hsa_circRNA_100861 and hsa_circRNA_102448 appear decreased in the same patients103.
Several published reports relate to circRNAs in different cardiovascular diseases104,105. The presence of hsa_circRNA_025016 in the plasma of patients is able to predict postoperative atrial fibrillation, while MICRA (myocardial infarction-associated circRNA) can help predicting left ventricular dysfunction in patients with acute myocardial infarction106,107. Similarly, in addition to being much more expressed than its linear form, an isoform of circANRIL has been shown to be elevated in whole blood of cardiac patients and playing an atheroprotective role, unlike its linear counterpart, which appears to play a proatherogenic role108.
CircRNAs implicated in molecular pathways disclose their potential use as therapeutic molecules
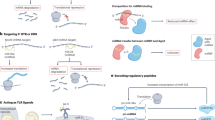
Although several circRNAs have been found to be either up- or down-regulated in human tissues, not always their specific role in molecular pathways has been elucidated. Specific circRNAs act in the Wnt signal transduction pathway: circRNA ITCH is active in lung cancer59 and cZNF292 is active in glioma109 (Fig. 2). Silencing cZNF292 blocked glioma cell cycle progression by means of inhibiting the Wnt/ß-catenin signaling pathway109.
Wnt signaling can be divided into β-catenin-dependent (or canonical) and β-catenin-independent (or non-canonical) signaling22,110,111. This pathway plays a critical role during embryonic development, including cell fate specification, cell proliferation, and cell migration. Moreover, the role of Wnt signaling has been well characterized in several diseases, such as cancer, diabetes, and cardiovascular disorders112,113,114. Accordingly, clinical trials that tested Wnt signaling drugs have shown promising outcomes, and circRNAs affecting the Wnt pathway might serve as targets for new therapies22,115.
CircANKS1B promotes the epithelial-to-mesenchymal transition (EMT) in triple-negative breast cancer (TNBC)57 (Fig. 3). EMT takes place in a diverse range of physiological and pathological conditions116. The molecular reprogramming occurring during EMT is orchestrated by a complex combination of factors, possibly including circRNAs. The biogenesis of numerous circRNAs is promoted during EMT transition by the RNA-binding protein quaking-5 (QKI-5)16. Recently, several clinical trials have been launched based on the current knowledge of EMT heterogeneity and plasticity117. The next challenge will be to include circRNAs as biomarkers or pharmacological targets in the protocols of new clinical trials addressing EMT.
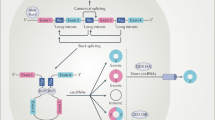
The circPVT1 and the circPPP1R12A act within the Hippo-YAP signaling pathway, respectively in head and neck squamous cell carcinoma and in colon cancer19,39 (Fig. 4). The Hippo pathway is recognized as an evolutionarily conserved signal transduction pathway that controls proliferation, organ size, and shape during development23. Moreover, the Hippo pathway is involved in multiple physiological processes, such as tissue growth, regeneration, and repair, maintaining the tissue homeostasis23. Hippo signaling plays an important role as a tumor suppressor in cancer and its deregulation is a key feature for cancer development, progression, and resistance to cancer treatment23,118,119. We showed that the mutant form of p53 (mut-p53) physically interacts with the transcriptional cofactor Yes-Associated Protein (YAP) in breast cancer120. YAP and TAZ are the main effectors of the Hippo pathway120. Hippo pathway inactivation determines the translocation to the nucleus of YAP and TAZ that regulate transcriptional activation in collaboration with mut-p53. In this context, Hippo effectors YAP and TAZ can act either as tumor suppressors, when located in the cytoplasm, or as oncogenes in the nucleus. In its wild type conformation, p53 works as a tumor suppressor regulating the cellular homeostasis23. At the same time the “p53 status”, wild type or mutant, can be considered a critical point in determining the tumor suppressor or oncogenic activity of the Hippo pathway23. Currently, there are several pathway modulators of the Hippo pathway that are subject of clinical development, such as the Verterpofin who inhibits the YAP–TEAD interaction121 or the PRIMA1-MET that restores the pro-apoptotic function of p53 with consequent activation of downstream target genes122. Importantly, we showed that YAP binds circPVT1 in head and neck squamous cell carcinoma. Moreover, we demonstrated that mut-p53 stabilizes the YAP/circPVT1 complex39. Thus, the current knowledge of circRNAs and their interaction with the Hippo pathway are expected to open new ways for the development of novel and more effective drugs.
The circPVT1 acts as oncogene in head and neck squamous cell carcinoma sponging the miR-497-5p and binding the complex YAP/mutp53. The circPPP1R12A can be translated into a protein and acts as oncogene in colon cancer where it is able to activated the proliferation of the tumor cells via theHippo-YAP pathway.
Divergent views of circRNA biogenesis and their mode of action
Despite the great interest that the circRNAs are raising in the scientific community, there are still some important questions regarding their biogenesis and function. The presence of repetitive inverted Alu elements flanking exons favors RNA circularization123. However, Zhang et al. demonstrated another mechanism not dependent on repetitive sequences for the generation of circRNAs, and occurring by the pairing between complementary sequences in introns flanking exons32. Another regulatory mechanism for circRNA biogenesis uses the ADAR protein, which is capable of modifying nucleotides in intronic repeat sequences29. Ivanov et al.29 demonstrated that ADAR antagonizes the expression of several circRNAs by editing intronic sites that flank exons and promote back-splicing. It is becoming clearer that introns are more important sequences for circRNAs biogenesis than initially anticipated, and that specific proteins might regulate backsplicing. Apparently, several different mechanisms control circRNA biogenesis, but it is yet unclear how do they work and which one, if any, is predominant over the others.
The unique circular configuration confers to cirRNAs not only resistance to digestion by ribonucleases, but it also translates to a longer half-life compared to the respective mRNAs124,125,126. As a consequence, circRNA levels are typically reduced in rapidly proliferating cells, such as in cancer cells. Thus, the association between lower circRNA levels and cancer could be due, in some cases, to a simple dilution effect mediated by cell division, as in colorectal cancer127. Hence, the meaning of circRNA expression levels should be carefully evaluated based on the specific cellular context. In addition, many circRNAs are sensitive to RNase R treatments, thus contradicting claims related to high stability of these molecules128.
Although the microRNA sponging mechanism is the better-described role for circRNAs, many circRNAs putatively act as sponges toward only a single, or very few miRNA targets129. Notably, there are some prerequisites that should be fulfilled in order to identify a circRNA as a putative ceRNA: the presence of multiple binding sites for the miRNA target, relatively high abundance of the circRNA, a miRNA target with fewer target genes, and a circRNA with a better affinity toward the miRNA than the mRNAs–miRNA affinity130. Moreover, a circRNA that triggers the degradation process of a target miRNA, and not only inhibits the interaction between miRNA and mRNA, might act as a better ceRNA candidate129.
The most common approach for assessing the sponging mechanism is the ectopic overexpression of binding sites for a specific miRNA. However, the result of this kind of experiment should be interpreted with caution since it could be biased by the introduction of sufficiently high numbers of binding sites able to inhibit the activity of the miRNA in question126. At the same time, one should also keep in mind that if a circRNA only binds with the cognate miRNA and inhibits its function without degrading it, the abundance of the miRNA would not be affected. Piwecka et al.131 have shown that the choice between either degradation or functional inhibition depends on whether the binding sites connecting circRNAs and miRNAs are completely complementary or they only partially match each other. It follows that a reliable in silico analysis of putative binding sites for circRNAs on miRNA targets is an essential requirement. While most studies have shown repression of miRNAs, Hansen et al.132 showed that the interaction between Cdr1 and miR-671 would actually lead to degradation of this circRNA through AGO2 rather than by the expected miRNA degradation mode132. Therefore, it is possible that additional circRNA–miRNA interactions regulate RNA circles.
With a few exceptions, the majority of circRNAs are expressed at low levels in both normal and cancer cells; hence they are unlikely to have only secondary roles in cellular physiology. However, the cascade of events that a single circRNA might unleash can potentially be of great importance from the clinical point of view, as we have shown for the circRNA circPVT1 (ref. 39). The roles of circRNAs must be carefully assessed in light of the various processes of their biogenesis and degradation, in addition to their broad capabilities for interacting with miRNAs and proteins.
Conclusions
The biochemical and molecular characteristics of circRNAs hold the promise that specific circles of RNA will be utilized in the future as disease biomarkers and pharmacological targets, thus opening new possibilities for early detection and treatment133,134,135. The large spectrum of mechanisms of action used by circRNAs makes the understanding of their role not only challenging but also promising in terms of resolving the complex molecular mechanisms activated in human disorders. Indeed, circRNAs can act as tumor suppressors or as oncogenes in oncology136,137. Likewise, circRNAs are involved in cardioprotection against heart failure, as well as mediate cardiomyocyte death in myocardial infarction12,76,77,78,79. Moreover, circRNAs are extensively expressed in the mammalian brain138,139. Networks of circRNAs, RNA-binding proteins and microRNAs play important roles in different human diseases, which reflects the complex regulatory potential of circRNAs. Hence, it is likely that the next few years will witness the discovery of more circRNAs and new modes of their action in human disorders.
References
Jeck, W. R. & Sharpless, N. E. Detecting and characterizing circular RNAs. Nat. Biotechnol. 32, 453–461 (2014).
Verduci, L., Strano, S., Yarden, Y. & Blandino, G. The circRNA-microRNA code: emerging implications for cancer diagnosis and treatment. Mol. Oncol. 13, 669–680 (2019).
Rong, D. et al. An emerging function of circRNA-miRNAs-mRNA axis in human diseases. Oncotarget 8, 73271–73281 (2017).
Memczak, S. et al. Circular RNAs are a large class of animal RNAs with regulatory potency. Nature 495, 333–338 (2013).
Hsiao, K. Y. et al. Noncoding effects of circular RNA CCDC66 promote colon cancer growth and metastasis. Cancer Res. 77, 2339–2350 (2017).
Han, D. et al. Circular RNA circMTO1 acts as the sponge of microRNA-9 to suppress hepatocellular carcinoma progression. Hepatology 66, 1151–1164 (2017).
Song, T. et al. CircRNA hsa_circRNA_101996 increases cervical cancer proliferation and invasion through activating TPX2 expression by restraining miR-8075. J. Cell Physiol. 234, 14296–14305 (2019).
Xuan, L. et al. Circular RNA: a novel biomarker for progressive laryngeal cancer. Am. J. Transl. Res. 8, 932–939 (2016).
Ke, Z., Xie, F., Zheng, C. & Chen, D. CircHIPK3 promotes proliferation and invasion in nasopharyngeal carcinoma by abrogating miR-4288-induced ELF3 inhibition. J. Cell Physiol. 234, 1699–1706 (2019).
Du, W. W. et al. Foxo3 circular RNA retards cell cycle progression via forming ternary complexes with p21 and CDK2. Nucleic Acids Res. 44, 2846–2858 (2016).
He, Q. et al. circ-SHKBP1 regulates the angiogenesis of U87 glioma-exposed endothelial cells through miR-544a/FOXP1 and miR-379/FOXP2 pathways. Mol. Ther. Nucleic Acids 10, 331–348 (2018).
Garikipati, V. N. S. et al. Circular RNA CircFndc3b modulates cardiac repair after myocardial infarction via FUS/VEGF-A axis. Nat. Commun. 10, 4317 (2019).
Geng, H. H. et al. The circular RNA Cdr1as promotes myocardial infarction by mediating the regulation of miR-7a on its target genes expression. PLoS ONE 11, e0151753 (2016).
Abdelmohsen, K. et al. Identification of HuR target circular RNAs uncovers suppression of PABPN1 translation by CircPABPN1. RNA Biol. 14, 361–369 (2017).
Yang, Q. et al. A circular RNA promotes tumorigenesis by inducing c-myc nuclear translocation. Cell Death Differ. 24, 1609–1620 (2017).
Conn, S. J. et al. The RNA binding protein quaking regulates formation of circRNAs. Cell 160, 1125–1134 (2015).
Legnini, I. et al. Circ-ZNF609 is a circular rna that can be translated and functions in myogenesis. Mol. Cell 66, 22–37 (2017).
Zhang, M. et al. A novel protein encoded by the circular form of the SHPRH gene suppresses glioma tumorigenesis. Oncogene 37, 1805–1814 (2018).
Zheng, X. et al. A novel protein encoded by a circular RNA circPPP1R12A promotes tumor pathogenesis and metastasis of colon cancer via Hippo-YAP signaling. Mol. Cancer 18, 47 (2019).
Li, Z. et al. Exon-intron circular RNAs regulate transcription in the nucleus. Nat. Struct. Mol. Biol. 22, 256–264 (2015).
Zhang, Y. et al. Circular intronic long non coding RNAs. Mol. Cell 51, 792–806 (2013).
Jung, Y. S. & Park, J. I. Wnt signaling in cancer: therapeutic targeting of Wnt signaling beyond β-catenin and the destruction complex. Exp. Mol. Med. 52, 183–191 (2020).
Ferraiuolo, M., Verduci, L., Blandino, G. & Strano, S. Mutant p53 protein and the hippo transducers YAP and TAZ: a critical oncogenic node in human cancers. Int. J. Mol. Sci. 18, 961 (2017).
Bahn, J. H. et al. The landscape of microRNA, Piwi-interacting RNA, and circular RNA in human saliva. Clin. Chem. 61, 221–230 (2015).
Memczak, S., Papavasileiou, P., Peters, O. & Rajewsky, N. Identification and characterization of circular RNAs as a new class of putative biomarkers in human blood. PLoS ONE 10, e0141214 (2015).
Jacky Lam, W. K. & Dennis, Lo. Y. M. Circular RNAs as urinary biomarkers. Clin. Chem. 65, 1196–1198 (2019).
Salzman, J., Gawad, C., Wang, P. L., Lacayo, N. & Brown, P. O. Circular RNAs are the predominant transcript isoform from hundreds of human genes in diverse cell types. PLoS ONE 7, e30733 (2012).
Ashwal-Fluss, R. et al. circRNA biogenesis competes with pre-mRNA splicing. Mol. Cell 56, 55–56 (2014).
Ivanov, A. et al. Analysis of intron sequences reveals hallmarks of circular RNA biogenesis in animals. Cell Rep. 10, 170–177 (2015).
Liang, D. & Wilusz, J. E. Short intronic repeat sequences facilitate circular RNA production. Genes Dev. 28, 2233–2247 (2014).
Starke, S. et al. Exon circularization requires canonical splice signals. Cell Rep. 10, 103–111 (2015).
Zhang, X. O. et al. Complementary sequence-mediated exon circularization. Cell 159, 134–147 (2014).
Hansen, T. B. et al. Natural RNA circles function as efficient microRNA sponges. Nature 495, 384–388 (2013).
Yu, L. et al. The circular RNA Cdr1as act as an oncogene in hepatocellular carcinoma through targeting miR-7 expression. PLoS ONE 11, e0158347 (2016).
Li, F. et al. Circular RNA ITCH has inhibitory effect on ESCC by suppressing the Wnt/b-catenin pathway. Oncotarget 6, 6001–6013 (2015).
Zheng, Q. et al. Huang, circular RNA profiling reveals an abundant circHIPK3 that regulates cell growth by sponging multiple miRNAs. Nat. Commun. 7, 11215 (2016).
Xu, X. W. et al. Circular RNA hsa_circ_000984 promotes colon cancer growth and metastasis by sponging miR-106b. Oncotarget 8, 91674–91683 (2017).
Zheng, J. et al. TTBK2 circular RNA promotes glioma malignancy by regulating miR-217/HNF1b/Derlin-1 pathway. J. Hematol. Oncol. 10, 52 (2017).
Verduci, L. et al. The oncogenic role of circPVT1 in head and neck squamous cell carcinoma is mediated through the mutant p53/YAP/TEAD transcription-competent complex. Genome Biol. 18, 237 (2017).
Stoll, L. et al. A circular RNA generated from an intron of the insulin gene controls insulin secretion. Nat. Comun. 11, 5611 (2020).
Leemans, C. R., Braakhuis, B. J. M. & Brakenhoff, R. H. The molecular biology of head and neck cancer. Nat. Rev. Cancer 11, 9–22 (2011).
O’Rorke, M. A. et al. Human papillomavirus related head and neck cancer survival: a systematic review and meta-analysis. Oral. Oncol. 48, 1191–1201 (2012).
Guo, S. T. et al. MicroRNA-497 targets insulin-like growth factor 1 receptor and has a tumour suppressive role in human colorectal cancer. Oncogene 32, 1910–1920 (2013).
Huang, C. et al. MiR-497 suppresses YAP1 and inhibits tumor growth in non-small cell lung cancer. Cell Physiol. Biochem. 37, 342–352 (2015).
Li, D. et al. Analysis of miR-195 and miR-497 expression, regulation and role in breast cancer. Clin. Cancer Res. 17, 1722–1730 (2011).
Wang, S. et al. The potent tumor suppressor miR-497 inhibits cancer phenotypes in nasopharyngeal carcinoma by targeting ANLN and HSPA4L. Oncotarget 6, 35893–35907 (2015).
Li, B. et al. Hsa_circ_0008309 may be a potential biomarker for oral squamous cell carcinoma. Dis. Markers 2018, 7496890 (2018).
Guo, Y. et al. Circular RNAs and their roles in head and neck cancers. Mol. Cancer 18, 44 (2019).
Waks, A. G. & Winer, E. P. Breast cancer treatment: a review. JAMA 321, 288–300 (2019).
Galasso, M. et al. Profiling of the predicted circular RNAs in ductal in situ and invasive breast cancer: a pilot study. Int. J. Genomics 2016, 4503840 (2016).
Nair, A. A. et al. Circular RNAs and their associations with breast cancer subtypes. Oncotarget 7, 80967–80979 (2016).
Xu, J. Z. et al. circTADA2As suppress breast cancer progression and metastasis via targeting miR-203a-3p/SOCS3 axis. Cell Death Dis. 10, 175 (2019).
Muhammad, N., Bhattacharya, S., Steele, R. & Ray, R. B. Anti-miR-203 suppresses ER-positive breast cancer growth and stemness by targeting SOCS3. Oncotarget 7, 58595–58605 (2016).
Wang, H., Xiao, Y., Wu, L. & Ma, D. Comprehensive circular RNA profiling reveals the regulatory role of the circRNA-000911/miR-449a pathway in breast carcinogenesis. Int. J. Oncol. 52, 743–754 (2018).
Du, W. W. et al. Identifying and characterizing circRNA-protein interaction. Theranostics 7, 4183–4191 (2017).
Chen, B. et al. circEPSTI1 as a prognostic marker and mediator of triple-negative breast cancer progression. Theranostics 8, 4003–4015 (2018).
Zeng, K. et al. The pro-metastasis effect of circANKS1B in breast cancer. Mol. Cancer 17, 160 (2018).
Neal R. D., Sun F., Emery J. D. & Callister M. E. Lung Cancer. BMJ 365, l1725 (2019).
Wan, L. et al. Circular RNA-ITCH suppresses lung cancer proliferation via inhibiting the Wnt/β-catenin pathway. Biomed. Res. Int. 2016, 1579490 (2016).
Chen, Y., Wei, S., Wang, X., Zhu, X. & Han, S. Progress in research on the role of circular RNAs in lung cancer. World J. Surg. Oncol. 16, 215 (2018).
Wei, W., Li, M., Wang, J., Nie, F. & Li, L. The E3 ubiquitin ligase ITCH negatively regulates canonical Wnt signaling by targeting dishevelled protein. Mol.Cell Biol. 32, 3903–3912 (2012).
Tian, F., Yu, C. T., Ye, W. D. & Wang, Q. Cinnamaldehyde induces cell apotosis mediated by a novel circular RNA hsa_circ_0043256 in non-small cell lung cancer. Biochem. Biophys. Res. Commun. 493, 1260–1266 (2017).
Wang, X. et al. Increased circular RNA hsa_circ_0012673 acts as a sponge of miR-22 to promote lung adenocarcinoma proliferation. Biochem. Biophys. Res. Commun. 496, 1069–1075 (2018).
Kiavue, N. et al. ERBB3 mutations in cancer: biological aspects, prevalence and therapeutics. Oncogene 39, 487–502 (2020).
Gazdar, A. F. Activating and resistance mutations of EGFR in non-small-cell lung cancer: role in clinical response to EGFR tyrosine kinase inhibitors. Oncogene 28, S24–S31 (2009).
Scheltens, P. et al. Alzheimer’s disease. Lancet 388, 505–517 (2016).
Dube, U. et al. An atlas of cortical circular RNA expression in Alzheimer disease brains demonstrates clinical and pathological associations. Nat. Neurosci. 22, 1903–1912 (2019).
Roberson, E. D. et al. Reducing endogenous tau ameliorates amyloid beta-induced deficits in an Alzheimer’s disease mouse model. Science 316, 750–754 (2007).
Welden, J. R., van Doorn, J., Nelson, P. T. & Stamm, S. The human MAPT locus generates circular RNAs. Biochim. Biophys. Acta Mol. Basis Dis. 1864, 2753–2760 (2018).
Zou, Y. et al. The role of circular RNA CDR1as/ciRS-7 in regulating tumor microenvironment: a pan-cancer analysis. Biomolecules 9, 429 (2019).
Lukiw, W. J. Variability in micro RNA (miRNA) abundance, speciation and complexity amongst different human populations and potential relevance to Alzheimer’s disease (AD). Front. Cell Neurosci. 7, 133 (2013).
Lonskaya, I. et al. Diminished parkin solubility and co-localization with intraneuronal amyloid-β are associated with autophagic defects in Alzheimer’s disease. J. Alzheimers Dis. 33, 231–247 (2013).
Lukiw, W. J. Circular RNA (circRNA) in Alzheimer’s disease (AD). Front. Genet. 4, 307 (2013).
Werfel, S. et al. Characterization of circular RNAs in human, mouse and rat hearts. J. Mol. Cell Cardiol. 98, 103–107 (2016).
Tan, W. L. W. et al. A landscape of circular RNA expression in the human heart. Cardiovasc. Res. 113, 298–309 (2017).
Wang, K. et al. A circular RNA protects the heart from pathological hypertrophy and heart failure by targeting miR-223. Eur. Heart J. 37, 2602–2611 (2016).
Geng, H. H. et al. The circular RNA Cdr1as promotes myocardial infarction by mediating the regulation of miR-7a on its target genes expression. PLoS ONE 11, e0151753 (2016).
Li, B. et al. MicroRNA-7a/b protects against cardiac myocyte injury in ischemia/reperfusion by targeting poly(ADP-ribose) polymerase. PLoS ONE 9, e90096 (2014).
Wang, K. et al. Circular RNA mediates cardiomyocyte death via miRNA-dependent upregulation of MTP18 expression. Cell Death Differ. 24, 1111–1120 (2017).
Tondera, D. et al. The mitochondrial protein mtp18 contributes to mitochondrial fission in mammalian cells. J. Cell Sci. 118, 3049–3059 (2005).
Li, M. et al. A circular transcript of ncx1 gene mediates ischemic myocardial injury by targeting miR-133a-3p. Theranostics 8, 5855–5869 (2018).
American Diabetes Association. Diagnosis and classification of diabetes mellitus. Diabetes Care 36, S67–S74 (2013).
Stoll, L. et al. Circular RNAs as novel regulators of β-cell functions in normal and disease conditions. Mol. Metab. 9, 69–83 (2018).
Xu, H., Guo, S., Li, W. & Yu, P. The circular RNA Cdr1as, via miR-7 and its targets, regulates insulin transcription and secretion in islet cells. Sci. Rep. 5, 12453 (2015).
Shan, K. et al. Circular noncoding RNA HIPK3 mediates retinal vascular dysfunction in diabetes mellitus. Circulation 136, 1629–1642 (2017).
Panda A. C. & Gorospe M. Detection and Analysis of Circular RNAs by RT-PCR. Bio Protoc. 8, e2775 (2018).
Salzman, J., Chen, R. E., Olsen, M. N., Wang, P. L. & Brown, P. O. Cell-type specific features of circular RNA expression. PLoS Genet. 9, e1003777 (2013).
Song, Z. et al. Identification of urinary Hsa_circ_0137439 as a potential biomarker and tumor regulator of bladder cancer. Neoplasma 67, 137–146 (2020).
Wang, Y. et al. Exosomal circRNAs: biogenesis, effect and application in human diseases. Mol. Cancer 18, 116 (2019).
Fanale, D., Taverna, S., Russo, A. & Bazan, V. Circular RNA in exosomes. Adv. Exp. Med. Biol. 1087, 109–117 (2018).
Li, Y. et al. Circular RNA is enriched and stable in exosomes: a promising biomarker for cancer diagnosis. Cell Res. 25, 981–984 (2015).
Li, S. et al. ExoRBase: a database of CircRNA, LncRNA and MRNA in human blood exosomes. Nucleic Acids Res. 46, D106–D112 (2018).
Ma, H. et al. Differential expression study of circular RNAs in exosomes from serum and urine in patients with idiopathic membranous nephropathy. Arch. Med. Sci. 15, 738–753 (2019).
Chen, X. et al. PRMT5 circular RNA promotes metastasis of urothelial carcinoma of the bladder through sponging MiR-30c to induce epithelial–mesenchymal transition. Clin. Cancer Res. 24, 6319–6330 (2018).
Vo, J. N. et al. The landscape of circular RNA in cancer. Cell 176, 869–881 (2019).
Chen, S., Li, T., Zhao, Q., Xiao, B. & Guo, J. Using circular hsa_circ_0000190 as a new biomarker in the diagnosis of gastric cancer. Clin. Chim. Acta 466, 167–171 (2017).
Li, P. et al. Using circular RNA as a novel type of biomarker in the screening of gastric cancer. Clin. Chim. Acta 444, 132–136 (2015).
Hang, D. et al. A novel plasma circular RNA CircFARSA is a potential biomarker for non-small cell lung cancer. Cancer Med. 7, 2783–2791 (2018).
Tan, S. et al. Circular RNA F-CircEA-2a derived from EML4-ALK fusion gene promotes cell migration and invasion in non-small cell lung cancer. Mol. Cancer 17, 138 (2018).
Zhu, K. et al. Plasma Hsa_circ_0027089 is a diagnostic biomarker for hepatitis B virus-related hepatocellular carcinoma. Carcinogenesis 41, 296–302 (2020).
Zuo, Z. et al. BBCancer: an expression atlas of blood-based biomarkers in the early diagnosis of cancers. Nucleic Acids Res. 48, D789–D796 (2020).
Li, Y. et al. Circular RNA expression profile of Alzheimer’s disease and its clinical significance as biomarkers for the disease risk and progression. Int. J. Biochem. Cell Biol. 123, 105747 (2020).
Li, Y. et al. Profiling of differentially expressed circular RNAs in peripheral blood mononuclear cells from Alzheimer’s disease patients. Metab. Brain Dis. 35, 201–213 (2020).
Lu, D. & Thum, T. RNA-based diagnostic and therapeutic strategies for cardiovascular disease. Nat. Rev. Cardiol. 16, 661–674 (2019).
Aufiero, S., Reckman, Y. J., Pinto, Y. M. & Creemers, E. E. Circular RNAs open a new chapter in cardiovascular biology. Nat. Rev. Cardiol. 16, 503–514 (2019).
Vausort, M. et al. Myocardial infarction-associated circular RNA predicting left ventricular dysfunction. J. Am. Coll. Cardiol. 68, 1247–1248 (2016).
Zhang, J. et al. Plasma circular RNAs, Hsa_circRNA_025016, predict postoperative atrial fibrillation after isolated off-pump coronary artery bypass grafting. J. Am. Heart Assoc. 7, e006642 (2018).
Holdt, L. M. et al. Circular non-coding RNA ANRIL modulates ribosomal RNA maturation and atherosclerosis in humans. Nat. Commun. 7, 12429 (2016).
Yang, P. et al. Silencing of cZNF292 circular RNA suppresses human glioma tube formation via the Wnt/β-catenin signaling pathway. Oncotarget 7, 63449–63455 (2016).
Grumolato, L. et al. Canonical and noncanonical Wnts use a common mechanism to activate completely unrelated coreceptors. Genes Dev. 24, 2517–2530 (2010).
Katoh, M. Canonical and non-canonical WNT signaling in cancer stem cells and their niches: cellular heterogeneity, omics reprogramming, targeted therapy and tumor plasticity (Review). Int. J. Oncol. 51, 1357–1369 (2017).
Bose, M., Almas, S. & Prabhakar, S. Wnt signaling and podocyte dysfunction in diabetic nephropathy. J. Investig. Med. 65, 1093–1101 (2017).
Gay, A. & Towler, D. A. Wnt signaling in cardiovascular disease: opportunities and challenges. Curr. Opin. Lipidol. 28, 387–396 (2017).
Wang, Z., Liu, C. H., Huang, S. & Chen, J. Wnt signaling in vascular eye diseases. Prog. Retin. Eye Res. 70, 110–133 (2019).
Harb, J., Lin, P. J. & Hao, J. Recent development of wnt signaling pathway inhibitors for cancer therapeutics. Curr. Oncol. Rep. 21, 12 (2019).
Yang, J. et al. Guidelines and definitions for research on ephitelial-mesenchymanl transition. Nat. Rev. Mol. Cell Biol. 21, 341–352 (2020).
Nieto, M. A., Huang, R. Y. J., Jackson, R. A. & Thiery, J. P. EMT: 2016. Cell 166, 21–45 (2016).
Sebio, A. & Lenz, H. J. Molecular pathways: hippo signaling, a critical tumor suppressor. Clin. Cancer Res. 21, 5002–5007 (2015).
Peng, Q. S. et al. circRNA_0000140 suppresses oral squamous cell carcinoma growth and metastasis by targeting miR-31 to inhibit Hippo signaling pathway. Cell Death Dis. 11, 112 (2020).
Di Agostino, S. et al. YAP enhances the pro-proliferative transcriptional activity of mutant p53 proteins. EMBO Rep. 17, 188–201 (2016).
Feng, J. et al. Verteporfin, a suppressor of YAP-TEAD complex, presents promising antitumor properties on ovarian cancer. Onco Targets Ther. 9, 5371–5381 (2016).
Sobhani, M., Abdi, J., Manujendra, S. N., Chen, C. & Chang, H. PRIMA-1Met induces apoptosis in Waldenström’s Macroglobulinemia cells independent of p53. Cancer Biol. Ther. 16, 799–806 (2015).
Jeck, W. R. et al. Circular RNAs are abundant, conserved, and associated with ALU repeats. RNA 19, 141–157 (2013).
Zhang, Y. et al. The biogenesis of nascent circular RNAs. Cell Rep. 15, 611–624 (2016).
Enuka, Y. et al. Circular RNAs are long-lived and display only minimal early alterations in response to a growth factor. Nucleic Acid Res. 44, 1370–1383 (2016).
Goodall, G. J. & Wickramasinghe, V. O. RNA in cancer. Nat. Rev. Cancer 21, 22–36 (2021).
Bachmayr-Heyda, A. et al. Correlation of circular RNA abundance with proliferation-exemplified with colorectal and ovarian cancer, idiopathic lung fibrosis, and normal human tissues. Sci. Rep. 5, 8057 (2015).
Szabo, L. & Salzman, J. Detecting circular RNAs: bioinformatic and experimental challenges. Nat. Rev. Genet. 17, 679–692 (2016).
Li, H. M., Ma, X. L. & Li, H. G. Intriguing circles: conflicts and controversies in circular RNA research. Wiley Interdiscip. Rev. RNA 10, e1538 (2019).
Bak, R. O. & Mikkelsen, J. G. miRNA sponges: soaking up miRNAs for regulation of gene expression. Wiley Interdiscip. Rev. RNA 5, 317–333 (2014).
Piwecka, M. et al. Loss of a mammalian circular RNA locus causes miRNA deregulation and affects brain function. Science 357, eaam8526 (2017).
Hansen, T. B. et al. miRNA-dependent gene silencing involving Ago2-mediated cleavage of a circular antisense RNA. EMBO J. 30, 4414–4422 (2011).
Costello, A., Lao, N. T., Barron, N. & Clynes, M. Reinventing the wheel: synthetic circular RNAs for mammalian cell engineering. Trends Biotechnol. 38, 217–230 (2020).
Ma, L. et al. Silencing of circRACGAP1 sensitizes gastric cancer cells to apatinib via modulating autophagy by targeting miR-3657 and ATG7. Cell Death Dis. 11, 169 (2020).
Xiong, D. D. et al. High throughput circRNA sequencing analysis reveals novel insights into the mechanism of nitidine chloride against hepatocellular carcinoma. Cell Death Dis. 10, 658 (2019).
Liu, Z. et al. CircRNA-5692 inhibits the progression of hepatocellular carcinoma by sponging miR-328-5p to enhance DAB2IP expression. Cell Death Dis. 10, 900 (2019).
Cao, L. et al. Circular RNA circRNF20 promotes breast cancer tumorigenesis and Warburg effect through miR-487a/HIF-1α/HK2. Cell Death Dis. 11, 145 (2020).
Rybak-Wolf, A. et al. Circular RNAs in the mammalian brain are highly abundant, conserved, and dynamically expressed. Mol. Cell 58, 870–875 (2015).
You, X. et al. Neural circular RNAs are derived from synaptic genes and regulated by development and plasticity. Nat. Neurosci. 18, 603–610 (2015).
Acknowledgements
We would like to thank the support of AIRC, Lazio Innova/Regione Lazio, MAECI Italy/USA bilateral grant program, and Alliance Against Cancer (ACC). Moreover, we would like to thank Professor Yosef Yarden’s laboratory for its kind collaboration. Y.Y. is the incumbent of the Harold and Zelda Goldenberg Professorial Chair.
Funding
This work was supported by AIRC IG 2017 ID 20613 and fondazione AIRC under 5 per mille 2019 - ID. 22759 program - GL Blandino Giovanni. Lazio Innova/Regione Lazio, MAECI Italy/USA bilateral grant program, and Alliance Against Cancer (ACC). YY laboratory was supported by the European Research Council, the Israel Science Foundation, the Israel Cancer Research Fund, and the Dr. Miriam and Sheldon G. Adelson Medical Research Foundation.
Author information
Authors and Affiliations
Contributions
Guarantors of the article: G.B. and Y.Y.; L.V. and G.B. identified the scope of this review article and wrote and revised the review. Y.Y. and E.T. revised the review and contributed to the writing. S.S. revised the review. L.V. conceived the original figures. E.T. revised the figures. All authors approved the final manuscript and agreed to be responsible for this review.
Corresponding authors
Ethics declarations
Ethics
This study did not require ethical approval.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by G. Melino
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Verduci, L., Tarcitano, E., Strano, S. et al. CircRNAs: role in human diseases and potential use as biomarkers. Cell Death Dis 12, 468 (2021). https://doi.org/10.1038/s41419-021-03743-3
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-021-03743-3
This article is cited by
-
Circular RNAs and cervical cancer: friends or foes? A landscape on circRNA-mediated regulation of key signaling pathways involved in the onset and progression of HPV-related cervical neoplasms
Cell Communication and Signaling (2024)
-
Circular RNAs and their roles in idiopathic pulmonary fibrosis
Respiratory Research (2024)
-
Non-coding RNAs in disease: from mechanisms to therapeutics
Nature Reviews Genetics (2024)
-
Paclitaxel-induced inhibition of NSCLC invasion and migration via RBFOX3-mediated circIGF1R biogenesis
Scientific Reports (2024)
-
Early-stage idiopathic Parkinson’s disease is associated with reduced circular RNA expression
npj Parkinson's Disease (2024)