Abstract
FOXA2 has been identified as an essential factor for pancreas development and emerging evidence supports an association between FOXA2 and diabetes. Although the role of FOXA2 during pancreatic development is well-studied in animal models, its role during human islet cell development remains unclear. Here, we generated induced pluripotent stem cells (iPSCs) from a patient with FOXA2 haploinsufficiency (FOXA2+/− iPSCs) followed by beta-cell differentiation to understand the role of FOXA2 during pancreatic beta-cell development. Our results showed that FOXA2 haploinsufficiency resulted in aberrant expression of genes essential for the differentiation and proper functioning of beta cells. At pancreatic progenitor (PP2) and endocrine progenitor (EPs) stages, transcriptome analysis showed downregulation in genes associated with pancreatic development and diabetes and upregulation in genes associated with nervous system development and WNT signaling pathway. Knockout of FOXA2 in control iPSCs (FOXA2−/− iPSCs) led to severe phenotypes in EPs and beta-cell stages. The expression of NGN3 and its downstream targets at EPs as well as INSUILIN and GLUCAGON at the beta-cell stage, were almost absent in the cells derived from FOXA2−/− iPSCs. These findings indicate that FOXA2 is crucial for human pancreatic endocrine development and its defect may lead to diabetes based on FOXA2 dosage.
Similar content being viewed by others
Introduction
During human development, early endodermal tissue becomes specified toward a pancreatic fate before evagination of pancreatic buds, populated with pancreatic progenitors. All adult pancreatic cells are originated from the same progenitors expressing a group of transcription factors (TFs), including PDX1, SOX9, FOXA2, NKX6.1, HNF6, and PTF1A1,2. Monogenic diabetes (MD) is caused by a mutation or defect in a single gene-regulating beta-cell development and/or function3. Several heterozygous mutations in the TFs expressed during pancreatic development are associated with a specific form of MD, known as MODY. However, homozygous mutations of the same TFs lead to neonatal diabetes, which can be associated with pancreatic hypoplasia/agenesis in some mutations4. This indicates that the onset and severity of the diabetes phenotype are correlated with the dosage of the TF expression during pancreatic development.
FOXA2 is expressed in several tissues and performs distinct functions as evident in the phenotypes of mouse models5. Foxa2 knockout mice die at an early embryonic stage and show developmental defects in the foregut (FG) and neural tube6,7,8. During pancreatic development, FOXA2 is expressed at early stages starting from the endoderm stage and its protein level is increased during the endocrine specification stage1,9, while the exocrine and ductal cells express a low level of FOXA29. Mouse studies showed that Foxa2 is important for islet development and beta-cell functionality6,10,11,12 and its specific deletion in beta-cells leads to hyperinsulinemic hypoglycemic phenotype13,14. In human, previous reports demonstrated that patients with heterozygous FOXA2 mutations develop hyperinsulinemia, hypoglycemia, hypopituitarism, endodermal organ defects, and craniofacial abnormalities15,16. Recent genomic studies found that type 2 diabetes (T2D) risk alleles are associated with FOXA2-bound enhancers in human17. Another recent study reported a patient with diabetes due to a heterozygous missense variant in FOXA2, indicating that FOXA2 defect can lead to MD18. FOXA2 is known to regulate the expression of multiple TFs controlling pancreatic endocrine cell fate and insulin secretion19,20.
Most of the information available on FOXA2 functions have been obtained from animal studies7,10,11,13,21, which do not fully reflect human phenotypes. Recent progress in the induced pluripotent stem cell (iPSC) technology allowed us to generate in vitro models to study human genetic diseases22. To our knowledge, there is only one recent study that used FOXA2 knockout hESCs (FOXA2−/− hESCs) to study FOXA2 during pancreatic progenitor differentiation23. To better understand the role of FOXA2 in beta cell development and diabetes progression, we established iPSCs from a patient with a heterozygous deletion in FOXA2 (FOXA2+/− iPSCs)24. We showed a number of key genes essential for pancreatic development were dysregulated in FOXA2+/− iPSC-derived pancreatic cells and the results obtained were further validated by generating CRISPR/Cas9-mediated FOXA2 knockout (FOXA2−/− iPSCs). Our results suggest that FOXA2 have a crucial role in pancreatic endocrine development in human.
Results
Derivation of iPSCs from a patient with FOXA2 heterozygous deletion
Recently, we reported the generation of an iPSC line from a patient with a heterozygous deletion in FOXA2 gene (QBRIi009-A; FOXA2+/− iPSC-C1)24 and from a healthy control (QBRIi001; Ctr1-iPSCs)25. In this study, we generated additional iPSC lines from another healthy control (Ctr2-iPSCs). All the established iPSC lines showed the expression of the pluripotency markers and showed the ability to differentiate into the three germ layers (Supplementary Figs. S1 and S2). Ctr-iPSC lines showed normal karyotype, while FOXA2+/− iPSC lines showed a translocation between the short arm of chromosome 6 and 20, karyotype: [46 XY, t (6; 20) (p11; p11)] and a deletion [46, XY, del (20) (p11.21p11.22)] as expected24 and as reported in the patient sample26 (Supplementary Fig. S1E). Three well-characterized clones from FOXA2+/− iPSCs were utilized for the subsequent experiments.
Effect of FOXA2 haploinsufficiency on definitive endoderm (DE) formation
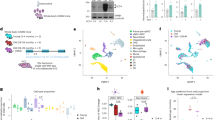
To investigate the effect of FOXA2 heterozygous deletion on pancreatic beta-cell development, Ctr-iPSCs and three FOXA2+/− iPSC lines were differentiated into all stages of beta cells as previously reported27,28,29 (Supplementary Fig. S3A). Since FOXA2 expression starts at the definitive endoderm (DE) stage, we examined the impact of FOXA2 haploinsufficiency on DE formation. The protein and mRNA levels of FOXA2 were significantly reduced in DE derived from the three FOXA2+/− iPSC lines in comparison to those derived from Ctr-iPSC lines (Fig. 1A–D), confirming FOXA2 haploinsufficiency. qPCR analysis showed a significant decrease in the expression of SOX17, the DE marker (Fig. 1B). Furthermore, immunostaining and flow cytometry analyses showed a marked reduction in the expression of the DE markers, SOX17 and FOXA2 and a significant increase in the expression of the pluripotency markers, OCT4 and SOX2 in the DE derived from the FOXA2+/− iPSC lines compared to those derived from Ctr-iPSCs (Fig. 1C, D). OCT4+ cells ranged from 35 to 51% in the DE derived from FOXA2+/− iPSCs compared to 5–9% in those derived from Ctr-iPSCs (Fig. 1D) indicating the retention of the pluripotency markers and a delay in DE formation as a result of FOXA2 deficiency.
A Western blotting of FOXA2 expression in the definitive endoderm (DE) derived from Ctr-iPSCs and FOXA2+/− iPSCs. B qPCR quantification of the expression of DE markers, FOXA2 and SOX17 in DE derived from Ctr-iPSCs and FOXA2+/− iPSCs (n = 3). C Representative immunofluorescent images showing the expression of SOX17 and OCT4 or FOXA2 and SOX2 in DE derived from Ctr-iPSCs and FOXA2+/− iPSCs. D Representative flow cytometry histograms and its quantification showing the expression of FOXA2, SOX17, and OCT4 in DE derived from Ctr-iPSCs and FOXA2+/− iPSCs. Data are represented as mean ± SD; *p < 0.05, **p < 0.01, ***p < 0.001.
FOXA2 deficiency leads to aberrant development of PDX1+/NKX6.1+ progenitors and endocrine progenitors
We next sought to assess the impact of FOXA2 deficiency on the differentiation of pancreatic progenitors (PP2) and endocrine progenitors (EPs). FOXA2 protein and mRNA levels were markedly reduced in PP2 derived from FOXA2+/− iPSCs (Fig. 2A, B), confirming the haploinsufficiency. We noticed that the efficiency of PP2 was negatively impacted as a result of FOXA2 reduction. The immunostaining and flow cytometry analyses showed a significant decrease in the expression of the key pancreatic progenitor TFs, PDX1 and NKX6.1 (above 35% reduction in PDX1 and NKX6.1) (Fig. 2C–E), further confirmed at mRNA levels using qPCR (Fig. 2F). The co-expression of PDX1 and NKX6.1 in the PP2 is known to be the origin of pancreatic beta cells27,28. Furthermore, we investigated the mRNA expression levels of other TFs, expressed at the PP2 stage and are important for beta-cell development, including SOX9, GATA6, ONECUT1 (HNF6), HNF1A, and PAX4 as well as the exocrine precursor markers, PTF1A and AMYLASE. Interestingly, all these pancreatic markers were significantly downregulated in PP2 derived from FOXA2+/− iPSC lines compared to those derived from Ctr-iPSCs (Fig. 2F), indicating the importance of FOXA2 in PP2.
Western blotting (A) and qPCR analysis (B) for FOXA2 protein and mRNA expression levels in iPSC-derived PP2. C Representative immunofluorescent images showing the expression of PDX1 and NKX6.1 in the PP2 derived from Ctr-iPSCs and FOXA2+/− iPSCs. D Flow cytometry histograms showing the expression of PDX1 and NKX6.1 in the PP2 derived from Ctr-iPSCs and FOXA2+/− iPSCs. E Quantification of the expression of PDX1 and NKX6.1 shown in D. F qPCR quantification for the expression of key PP2 markers, PDX1, NKX6.1, SOX9, HNF1A, GATA6, PAX4, ONECUT1 (HNF6), PTF1A, and AMYLASE in the generated PP2. Data are represented as mean ± SD; *p < 0.05, **p < 0.01, ***p < 0.001.
To investigate the effect of FOXA2 deficiency on EPs, the PP2 were further differentiated into EPs. The downregulation of FOXA2 was confirmed at the end of stage 5 (Fig. 3A, B). The immunostaining showed a decrease in the expression of the endocrine marker, chromogranin A (CHGA), as a result of FOXA2 deficiency (Fig. 3A). Also, the expression levels of NEUROGENIN 3 (NGN3), a crucial TF for endocrine specification, and its downstream targets, NEUROD1 and NKX2.2, as well as NKX6.1, were markedly reduced in the EPs derived from FOXA2+/− iPSCs compared to those generated from Ctr-iPSCs as indicated by immunostaining and qPCR (Fig. 3A, B). Furthermore, we examined the effect of FOXA2 deficiency on FOXA1 expression at PP2 and EPs. We found that FOXA2 deficiency resulted in a significant reduction in the FOXA1 mRNA levels (Supplementary Fig. S3B), suggesting a lack of a compensatory mechanism.
A Representative immunofluorescent images showing the expression of FOXA2, NGN3, CHGA, NEUROD1, and NKX6.1 in endocrine progenitors (EPs) derived from Ctr-iPSCs and FOXA2+/− iPSCs. B Real-time PCR analysis for the expression of the key EPs markers, FOXA2, NGN3, CHGA, NEUROD1, NKX6.1, and NKX2.2 in the EPs derived from Ctr-iPSCs and FOXA2+/− iPSCs. Data are represented as mean ± SD; *p < 0.05, **p < 0.01, ***p < 0.001.
Transcriptomic alterations in PP2 and EPs derived from FOXA2+/− iPSCs
To gain further insight into the gene regulatory network influenced by FOXA2 leading to progenitor development defects, we performed whole-transcriptome profiling using RNA-Seq analysis on PP2 and EPs. Our transcriptome analysis identified 610 downregulated (Log2 FC < −1.0, p < 0.05) and 427 upregulated (Log2 FC > 1.0, p < 0.05) differentially expressed genes (DEGs) in PP2 derived from FOXA2+/− iPSCs compared to those generated from Ctr-iPSCs (Fig. 4A). To investigate biological functions and pathways enriched in FOXA2-deficient PP2, GO and KEGG pathway enrichment analyses were performed. The enriched pathways of the downregulated DEGs were mainly associated with pancreatic development, liver development, glucose homeostasis, insulin secretion, MODY, T2D, and NOTCH signaling (Fig. 4B, C and Supplementary Table S4). However, the upregulated DEGs led to several enriched pathways, such as axon guidance, nervous system development, and WNT and BMP signaling (Fig. 4B, C and Supplementary Fig. S3C, Supplementary Table S5). Consistent with the qPCR results in Fig. 2F, the main PP2-specific TFs were significantly downregulated, such as PDX1, NKX6.1, FOXA2, SOX9, GATA6, PAX4, HNF6, and PTF1A (Figs. 2F, 4C and Supplementary Table S4). Furthermore, several genes involved in the endocrine specification and diabetes development were significantly downregulated, such as HES1, INSM1, HNF1A, NEUROD1, NGN3, NKX2.2, GIPR, MNX1, PROX1, TCF7L2, and HHEX (Fig. 4C and Supplementary Table S4). Consistent with pancreatic developmental defects, we noticed a significant reduction in the genes involved in the NOTCH signaling pathway, such as SOX9, NOTCH1, HES1, and HEY1, which have a key role in PP2 differentiation (Fig. 4C and Supplementary Table S4).
A Volcano plot plotting the log2 fold change (FC) and the adjusted p-value for all the transcripts. B Pathway enrichment analysis on the upregulated and downregulated differentially expressed genes (DEGs) in FOXA2+/− iPSC-derived pancreatic progenitors (PP2) using the DAVID function annotation tool. The enriched pathways were plotted against −log10 (p-value). C Heatmaps of pathway-associated DEGs in PP2 derived from FOXA2+/− iPSCs compared to those derived from Ctr-iPSCs. The downregulated genes were associated with pancreatic development, maturity-onset diabetes of the young (MODY), and NOTCH signaling pathways, while the upregulated genes were associated with the WNT signaling pathway and nervous system development. The relative value for each gene is depicted by color intensity, with red indicating upregulated and blue indicating downregulated genes. RNA-Seq results were validated using qPCR for the downregulated (D) and the upregulated genes (E). Data are represented as mean ± SD; *p < 0.05, **p < 0.01, ***p < 0.001.
To validate the RNA-Seq data, we examined the mRNA expression levels of selected DEGs. qPCR showed a significant reduction in NEUROD1, TCF7L2, INSR, NOTCH1, HES1, MNX1, ISL1, PROX1, PTPRN, ADRA2A, and CDX2 (Fig. 4D). Furthermore, the results showed a significant upregulation in BMP4, NOGGIN, MSX1, MSX2, DKK1, DLX2, DLX5, EMX2, ERG2, OTX1, OTX2, SOX2, and PAX6 (Fig. 4E). These results are consistent with the RNA-Seq data.
Our RNA-Seq analysis on EPs identified 643 downregulated (Log2 FC < −1.0, p < 0.05) and 564 upregulated (Log2 FC > 1.0, p < 0.05) DEGs in EPs derived from FOXA2+/− iPSCs compared to those generated from Ctr-iPSCs (Fig. 5A). The enriched pathways of the downregulated and upregulated DEGs were similar to those observed in PP2 (Fig. 5B). The downregulated DEGs were mainly associated with endocrine pancreas development, NOTCH signaling pathway, MODY, and insulin secretion and glucose homeostasis (Fig. 5B, C and Supplementary Table S6). However, the upregulated DEGs were mainly associated with neurogenesis and nervous system development, WNT pathway, and BMP signaling pathway (Fig. 5B, C and Supplementary Fig. S3D, Supplementary Table S7). In addition to the downregulated pancreatic endocrine TFs shown in Fig. 3B, we further validated selected upregulated DEGs using qPCR. Consistent with RNA-Seq data, the mRNA levels of DLX5, EMX2, OTX1, OTX2, BMP4, NOGGIN, and SIX4 were significantly upregulated in the EPs derived from FOXA2+/− iPSCs compared to those derived from Ctr-iPSCs (Fig. 5D).
A Volcano plot plotting the log2 fold change (FC) and the adjusted p-value for all the transcripts. B Pathway enrichment analysis on upregulated and downregulated differentially expressed genes (DEGs) in FOXA2+/− iPSC-derived EPs using DAVID function annotation tool. The enriched pathways were plotted against −log10 (p-value). C Heatmaps of pathway-associated DEGs in FOXA2+/− iPSC-derived EPs compared to those derived from Ctr-iPSCs. The downregulated genes were associated with pancreatic development, maturity-onset diabetes of the young (MODY), and NOTCH signaling pathways, while the upregulated genes were associated with the WNT signaling pathway and nervous system development. The relative value for each gene is depicted by color intensity, with red indicating upregulated and blue indicating downregulated genes. D RNA-Seq results were validated using qPCR for the upregulated genes associated with WNT signaling and nervous system development. Data are represented as mean ± SD; *p < 0.05, **p < 0.01, ***p < 0.001.
Effect of FOXA2 haploinsufficiency on beta-cell differentiation
In order to investigate the influence of FOXA2 haploinsufficiency on beta cells, the EPs were further differentiated into beta cells. Ctr-iPSCs and FOXA2+/− iPSCs were able to generate INSULIN (INS)+ cells. However, the expression levels of INS, NKX6.1, C-PEP, and GLUCAGON (GCG) were downregulated as examined by immunostaining and flow cytometry in stage 7 derived from FOXA2+/− iPSCs compared to those derived from Ctr-iPSCs (Fig. 6A–C). The expression of NKX6.1 in INS+ cells, one of the hallmarks of the functionality of beta cells, was also reduced. Furthermore, the mRNA levels of ABCC8, KCNJ11, NKX2.2, and PDX1, which are known to be regulated by FOXA219,20 were significantly downregulated due to FOXA2 haploinsufficiency (Fig. 6D). These results indicate that the number of beta and alpha cells are reduced as a result of FOXA2 haploinsufficiency.
Representative immunofluorescent images (A) and flow cytometry histograms (B) showing the expression of INSULIN (INS), NKX6.1, C-PEPTIDE (C-PEP), and GLUCAGON (GCG) in stage 7 derived from Ctr-iPSCs and FOXA2+/− iPSCs. C Flow cytometry quantification for the data shown in B. D qPCR quantification for the expression of the pancreatic markers, FOXA2, INS, GCG, ABCC8, KCNJ11, NKX6.1, NKX2.2, and PDX1 in beta cells derived from Ctr-iPSCs and FOXA2+/− iPSCs. GSIS assay: fold change of C-PEPTIDE secretion in iPSC-derived beta cells challenged with low (LG; 2.8 mM) and high glucose (HG; 20 mM) (E) or stimulated with or without 30 mM KCl in the presence of LG (F). Data are represented as mean ± SD; *p < 0.05, **p < 0.01, ***p < 0.001.
To test the functionality of the generated beta cells, we performed a GSIS assay. The ability of iPSC-derived beta-cell aggregates to secrete C-PEP in response to low (2.8 mM) and high (20 mM) glucose and KCl stimulation was measured. Beta cells derived from FOXA2+/− iPSCs and Ctr-iPSCs showed a significant increase in the secreted C-PEP in response to high glucose (FC > 3.0) and KCl stimulation compared to low glucose levels (Fig. 6E, F). We did not notice a significant difference in glucose responsiveness between the beta cells derived from FOXA2+/− iPSCs and those derived from Ctr-iPSCs (Fig. 6E). However, KCl stimulation showed a significant reduction in the C-PEP content in beta cells derived from FOXA2+/− iPSCs compared with those derived from Ctr-iPSCs (Fig. 6E). These results indicate that FOXA2-deficient beta cells are functional, but the total content of insulin is reduced as a result of a lower number of beta cells due to FOXA2 haploinsufficiency.
FOXA2 knockout in iPSCs impairs endocrine pancreatic development
To confirm the results obtained from the FOXA2+/− iPSCs and to exclude the possibility of cell line variations, we used CRISPR/Cas9 to KO the FOXA2 gene in the iPSCs (FOXA2−/− iPSCs) (Supplementary Fig. S4A–D). We generated FOXA2−/− iPSCs from Ctr1-iPSCs and Ctr2-iPSCs used in the current study and compared them with isogenic FOXA2+/+ wild type (WT) controls. Characterization of both FOXA2−/− iPSC lines showed that they were pluripotent and displayed normal karyotype (Supplementary Fig. S4E). Next, FOXA2−/− iPSC lines and their WT controls were differentiated into beta cells. Western blotting showed that the FOXA2 protein was completely lost in the DE and PP2 derived from FOXA2−/− iPSC lines (Fig. 7A, B). Consistent with the results obtained from FOXA2 haploinsufficiency, the expression of the key TFs, PDX1 and NKX61 were markedly reduced in PP2 derived from FOXA2−/− iPSC lines compared to WT iPSCs as examined by immunostaining (Fig. 7C). At the EPs stage, the immunostaining analysis showed that the depletion of FOXA2 resulted in the generation of a very low number of NGN3+/NKX2.2+ cells in comparison to WT controls (Fig. 7D). Furthermore, qPCR results showed a highly significant reduction in the expression of the main endocrine TFs, NGN3, NEUROD1, and NKX2.2 in the EPs derived from FOXA2−/− iPSC lines (Fig. 7E). Interestingly, immunostaining showed that only a small number of INS+ and GCG+ cells were detected in FOXA2−/− iPSC-derived beta cells compared to WT (Fig. 7F). Consistent with these findings, qPCR showed significant downregulation in the endocrine and beta-cell genes, NEUROD1, NKX2.2, INS, GCG, ABCC8, KCJ11, PDX1, and HHEX1 (Fig. 7G). Our results showed that the FOXA2−/− phenotypes were more severe than those obtained from FOXA2+/− iPSCs indicating that FOXA2 is required for endocrine pancreatic development.
A Western blotting of FOXA2 expression in the definitive endoderm (DE) derived from Ctr-iPSCs, FOXA2+/− iPSCs, and FOXA2−/− iPSCs. B Quantification of FOXA2 protein levels in DE and PP2 derived from Ctr-iPSCs, FOXA2+/− iPSCs, and FOXA2−/− iPSCs. Note the complete absence of FOXA2 expression in the DE derived from FOXA2−/− iPSCs. Representative immunofluorescence images showing the expression of PDX1 and NKX6.1 in PP2 (C) and NGN3 and NKX2.2 in EPs (D) derived from WT controls and FOXA2−/− iPSCs. E Real-time PCR analysis for the expression of FOXA2, NEUROD1, NKX2.2, and NGN3 in EPs derived from FOXA2−/− iPSCs and their WT controls. F Representative immunofluorescence images showing the expression of INS and GCG in stage 7 derived from FOXA2−/− iPSCs and Ctr1-iPSCs. G qPCR quantification for the expression of FOXA2, NEUROD1, NKX2.2, INS, GCG, ABCC8, KCNJ11, PDX1, and HHEX in stage 7 derived from FOXA2−/− iPSC line and its WT control, Ctr1-iPSCs. The data are presented as mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001.
Overexpression of FOXA2 in pancreatic progenitors reverses the phenotype associated with FOXA2 deficiency
In order to examine whether the phenotype observed due to FOXA2 deficiency can be reversed by re-introducing FOXA2 expression, we overexpressed FOXA2 at day 2 of stage 4 of differentiation (PP2 stage) derived from FOXA2+/− iPSCs and FOXA2−/− iPSCs (Fig. 8A). qPCR analysis at the end of stage 4 showed that FOXA2 OE resulted in a significant increase in the expression of the genes that were downregulated in FOXA2-deficient PP2 cells (Fig. 8B). Furthermore, FOXA2 OE led to a significant downregulation in the majority of the genes that were upregulated in FOXA2-deficient PP2 cells (Fig. 8C).
A Representative western blotting showing the overexpression of FOXA2 in the PP2 derived from FOXA2+/− iPSCs, and FOXA2−/− iPSCs, 48 h after transfection with FOXA2 plasmid or empty vector (EV). B Real-time PCR analysis for the expression of PDX1, NKX6.1, SOX9, GATA6, PAX4, PTF1A, AMYLASE, TCF7L2, INSR, NOTCH, HES1, MNX1, ISL1, PROX1, PTPRN1, ADRA2A, and CDX2 in PP2 derived from FOXA2+/− iPSCs and FOXA2−/− iPSCs 48 h after transfection. C Real-time PCR analysis for the expression of BMP4, NOG, MSX1, MSX2, DKK1, DLX2, DLX5, EMX2, ERG2, OTX1, OTX2, and RFX6 in PP2 derived from FOXA2+/− iPSCs and FOXA2−/− iPSCs 48 h after transfection. D Real-time PCR analysis for the expression of INS, GCG, SST, PDX1, NKX6.1, ABCC8, KCNJ11, HHEX, and NEUROD1 in stage 7 derived from FOXA2+/− iPSCs and FOXA2−/− iPSCs transfected with FOXA2 plasmid or EV at stage 4. E Representative immunofluorescence images showing the expression of INS and GCG in stage 7 derived from FOXA2+/− iPSCs and FOXA2−/− iPSCs transfected with FOXA2 plasmid or EV at stage 4. F C-PEPTIDE (C-PEP) secretion (% of content) after stimulation with or without 30 mM KCl in the presence of low glucose (LG) in FOXA2+/− iPSC-derived beta cells transfected with FOXA2 plasmid or EV at stage 4. The data are presented as mean ± SD. *p < 0.05, **p < 0.01, ***p < 0.001.
In order to assess the effect of the transient FOXA2 OE at the PP2 stage on iPSC differentiation towards pancreatic beta cells, we evaluated the key beta-cell markers and GSIS in stage 7 cells derived from both FOXA2+/− iPSCs and FOXA2−/− iPSCs after FOXA2 overexpression. At stage 7, we found that FOXA2 OE at PP2 resulted in a significant increase in the expression of the key pancreatic endocrine genes, including INS, GCG, SST, PDX1, NKX6.1, ABCC8, KCNJ11, HHEX, and NEUROD1 in comparison to those transfected with the empty vector (Fig. 8D). Consistent with these findings, the immunostaining results showed an increase in the number of INS+ and GCG+ cells derived from FOXA2+/− iPSCs and FOXA2−/− iPSCs after FOXA2 OE (Fig. 8E). Since the total content of insulin was reduced in beta cells derived from FOXA2+/− iPSCs (Fig. 6E), we examined the effect of FOXA2 OE on the C-PEP content. KCl stimulation showed a significant increase in the C-PEP content in beta cells derived from FOXA2+/− iPSCs overexpressed with FOXA2 compared with those transfected with the empty vector (Fig. 8E). These results indicate that FOXA2 OE can reverse the phenotype observed in the FOXA2-deficient pancreatic cells.
Effect of PDX1 overexpression on FOXA2-deficient pancreatic progenitors
Previous studies reported that FOXA2 controls the PDX1 gene during pancreatic beta-cell development in rodents and humans10,12,30. Therefore, we thought to examine the effect of PDX1 overexpression (PDX1 OE) at the PP2 stage on reversing the phenotype associated with FOXA2 deficiency (Supplementary Fig. S5A, B). As expected, PDX1 OE led to an increase in the mRNA expression levels of several key PP2 genes that were downregulated in FOXA2-deficient PP2 derived from FOXA2+/− iPSCs and FOXA2−/− iPSCs, such as NKX6.1, GATA6, PAX4, PTF1A, AMYLASE, TCF7L2, INSR, HES1, MNX1, PROX1, PTPRN1, and ADRA2A (Supplementary Fig. S5C, D). Furthermore, PDX1 OE led to a downregulation in the expression of some genes that were upregulated in the FOXA2-deficient PP2 derived from FOXA2+/− iPSCs and FOXA2−/− iPSCs, such as ERG2, OTX1, and OTX2 (Supplementary Fig. S5C, D). On the other hand, some genes were not significantly changed as a result of PDX1 OE, such as SOX9, ONECUT1, NOG, DKK1, DLX2, DLX5, and EMX2. Interestingly, PDX1 OE led to an upregulation of BMP4 and ERG2 (Supplementary Fig. S5C, D).
Inhibition of BMP and WNT signaling pathways in FOXA2-deficient pancreatic progenitor cells
Since FOXA2 deficiency led to a significant upregulation in the WNT and BMP signaling pathways, we thought to analyze whether inhibiting the BMP and WNT pathways are able to counteract the deleterious effects in the generation of PP2. We treated the cells with the WNT inhibitor, XAV939, for 4 days during stage 4 of differentiation (Supplementary Fig. S6A). We noticed that WNT inhibition led to a significant downregulation in the genes that were upregulated in the PP2 derived from FOXA2+/− iPSCs and FOXA2−/− iPSCs (Supplementary Fig. S6B). Also, WNT inhibition led to a significant increase in the expression of some genes that were downregulated in the PP2 derived from FOXA2+/− iPSCs and FOXA2−/− iPSCs, particularly NEUROD1, PAX4, TCF7L2, NOTCH, HES1, MNX1, PROX1, and PTPRN1 (Supplementary Fig. S6C). This indicates that inhibition of the WNT pathway, at least, partially reversed the phenotype noticed in the FOXA2-deficient PP2 stage (Supplementary Fig. S6A–C).
In the current protocol, we used BMP inhibitor, NOGGIN, during stages 2, 3, and 4 of differentiation. Therefore, to test the effect of BMP inhibition dose on the PP2 phenotype, we modified the protocol by treating stage 4 cells derived from FOXA2+/− iPSCs and FOXA2−/− iPSCs with two different concentrations of NOGGIN, 100 and 50 ng/mL (Supplementary Fig. S6D). The high concentration of NOGGIN (100 ng/mL) resulted in a significant reduction in the BMP pathway and other genes that were upregulated in the FOXA2-deficient PP2 in comparison to those treated with 50 ng/mL (Supplementary Fig. S6E). Although the high concentration of NOGGIN increased the expression of PP2 markers, these increases was not significant for most of the genes examined and showed some variations between PP2 derived from FOXA2+/− iPSCs and those derived from FOXA2−/− iPSCs (Supplementary Fig. S6F).
Discussion
The role of FOXA2 in pancreatic development has been studied in mouse models; however, the mechanism by which FOXA2 defects contribute to the pathogenesis of human diseases is still not well-known due to the lack of the appropriate model that can mimic human phenotypes. In the current study, we established a human iPSC model to gain insight into the role of FOXA2 during human pancreatic islet development and investigated the effect of FOXA2 haploinsufficiency on pancreatic beta-cell differentiation in vitro. Our results showed that FOXA2 is critical for the pancreatic endocrine specification. Taken together with the recent findings from other groups18,23, we suggest that FOXA2 defects may lead to monogenic diabetes with other abnormalities.
In the current study, FOXA2 haploinsufficiency led to a significant reduction in the expression of the key DE-specific TFs and retained the pluripotency markers in the iPSC-derived DE. This result disagrees with the recent findings obtained from FOXA2−/− hESCs, which showed that the DE and FG differentiation have not been affected by a complete loss of the FOXA2 gene23. However, our results are in agreement with a recent study that demonstrated that FOXA2 is a master regulator of endoderm formation and its absence in ESCs impairs the endoderm differentiation with improper activation of DE genes and sustained pluripotency markers31. Taken together, these data suggest a crucial role for FOXA2 during DE formation in human.
Our results obtained from the PP2 and EPs stages showed that FOXA2 haploinsufficiency led to a significant reduction in the expression of the monogenic diabetes-associated genes, such as PDX1, HNF4A, HNF1A, NEUROD1, NGN3, PAX4, ABCC8, KCNJ11, GATA6, and MNX1. In addition, several key pancreatic development-specific TFs were significantly downregulated in PP2 and EPs derived from FOXA2+/− iPSCs, such as NKX6.1, NKX6.2, SOX9, NKX2.2, ISL1, and GATA6, among others. Consistent with our findings, in mouse, Foxa2 has been found to control the expression of the majority of those TFs, including Pdx1, Nkx6.1, Nkx2.2, Hnf1a, Hnf4a, Kcnj11, and Abcc819,32. Furthermore, a recent study reported a significant reduction in the number of PDX1+ pancreatic progenitors derived from FOXA2−/− hESCs23. Given the important role of PDX1 in regulating the expression of several genes that are critical for pancreatic development and endocrine specification33,34 and the fact that PDX1 is the downstream target of FOXA230, our results suggest that the effect of FOXA2 on PDX1 at the PP2 stage is one of the key mechanisms by which FOXA2 control pancreatic endocrine development. In addition, at the PP2 stage, FOXA2 haploinsufficiency resulted in a significant reduction in the NOTCH signaling pathway, which is important for early pancreatic development by enhancing PP2 expansion. Notch1 is expressed in the mouse pancreatic progenitors and activates the downstream target genes, Hes1 and Hey135,36. Interestingly, we observed that NOTCH1, HES1, and HEY1 were significantly downregulated at the PP2 stage as a result of FOXA2 deficiency. Although early studies showed that Notch inactivation enhances Ngn3 and activates premature endocrine pancreatic differentiation37, recent studies reported that Notch is required for pancreatic endocrine specification by activating the expression of Nkx6.1 and Ngn3 activator, Sox938,39. Consistent with these findings, in addition to NOTCH pathway inhibition, we noticed downregulation of endocrine-specific TFs, such as SOX9, NKX6.1, NGN3, NKX2.2, NEUROD1, and PAX4 as a result of FOXA2 deficiency.
Furthermore, our results demonstrated that FOXA2 haploinsufficiency resulted in significant upregulation in the genes related to WNT and BMP signaling pathways at PP2 and EPs. Considering that WNT inhibition is crucial for endocrine development40 and BMP signaling inhibition is required for PDX1 induction in pancreatic progenitors41, we anticipate that increased both pathways are associated with the suppression of the pancreatic differentiation due to FOXA2 haploinsufficiency. Confirming this concept, we demonstrated that inhibition of WNT or BMP pathways during the PP2 stage partially rescued the phenotype associated with FOXA2 deficiency. The complete absence of FOXA2 had a more marked effect on beta-cell differentiation, particularly after the PP2 stage. FOXA2−/− iPSCs showed severe impairment in generating INS+ and GCG+ cells. These findings indicate that the severity of the phenotypes is associated with the dosage of FOXA2 expression. Transient FOXA2 OE in PP2 derived from FOXA2+/− iPSCs and FOXA2−/− iPSCs significantly upregulated the key pancreatic development-specific genes at PP2 and beta-cell stages and significantly downregulated the expression of WNT and BMP genes, indicating that the re-introduction of FOXA2 could reverse the phenotype associated with FOXA2 deficiency. Given the important roles of the downregulated and upregulated genes in controlling pancreatic endocrine development and beta-cell functionality, our findings indicate the important role of FOXA2 during pancreatic development and suggest that the patient with FOXA2 haploinsufficiency may develop diabetes later in life.
RNA-Seq and qPCR analyses showed that the top upregulated genes were associated with neurogenesis and nervous system development, which could be reversed after FOXA2 OE. This result is in agreement with the previous study that showed the activation of neuronal genes in the pancreatic islet of Foxa2-deficient mice11. Taken together, these findings suggest an important role for FOXA2 in suppressing neuronal fate during pancreatic differentiation to enhance the beta-cell program.
Although FOXA2 haploinsufficiency led to significant downregulation in the genes related to pancreatic endocrine development, it appears that the expression levels of those genes were still sufficient to generate functional, glucose-responsive beta cells at stage 7. In mice, Foxa2 haploinsufficiency is enough for normal islet development; however, the deletion of both Foxa1 and Foxa2 in the mouse pancreas leads to pancreatic hypoplasia as a result of a Pdx1 suppression10, indicating that Foxa1 can compensate for the absence of Foxa2 in rodents10. In this study, we found that FOXA1 expression was significantly downregulated in PP2 and EPs derived from FOXA2+/− iPSCs. Consistent with our findings, Lee et al. reported the reduction of FOXA1 in the DE and FG derived from FOXA2−/− hESCs23. These findings indicate that FOXA1 does not compensate for FOXA2 deficiency, at least, during the early stages of pancreatic development in human, highlighting the physiological differences between mouse and human.
In addition to the significant reduction of INS expression, GCG was significantly downregulated as a result of FOXA2 haploinsufficiency. Confirming the effect of FOXA2 deficiency on GCG expression, iPSCs lacking FOXA2 generated a very small number of GCG+ cells at stage 7. Previous mouse studies showed that Foxa2 activates the expression of Gcg and maintain the alpha cells by transcriptional regulation of MafB, Kir6.2, Isl1, and Nkx2.220,42,43. In agreement with these findings, we found that all the endocrine-specific TFs were significantly downregulated as a result of FOXA2 deficiency. These findings indicate that FOXA2 may also be involved in alpha cell development and regulates GCG expression in human pancreatic islet.
Whether the defect in FOXA2 leads to hypo-or hyperglycemia is still controversial. Recent reports have identified two young patients (2 and 5 years old) with hypoglycemic hyperinsulinemic phenotype due to two different FOXA2 mutations15,16. However, a more recent study identified a patient with diabetes due to another FOXA2 mutation18. Of note, these hypo-and hyperglycemic patients shared common features, such as growth hormone deficiency, hypopituitarism, and early hypoglycemia18. The hypoglycemic episode followed by hyperglycemia during childhood due to FOXA2 mutations is similar to some MODY gene mutations such as ABCC844, HNF1A45, and HNF4A46. The link between FOXA2 defect and diabetes development has been reported in another human genomic study, in which an association of T2D risk alleles with FOXA2-bound enhancers has been found17. Taken together, our work supports the concept suggested that FOXA2 defects may lead to hypoglycemia at the early stage of life followed by hyperglycemia during late childhood or at a later stage18. Stekelenburg et al18. explained the possible mechanism of an early hypoglycemic episode associated with FOXA2 defects as a result of growth hormone deficiency16 or the reduction in hepatic gluconeogenesis associated with the downregulation in the GCG expression18. Further clinical and experimental studies are needed to understand how FOXA2 defects influence glucose homeostasis and pancreatic islet functionality over time in human.
In conclusion, our current study provides the first human iPSC model for FOXA2 haploinsufficiency carrying the genetic information of the patient. Our results indicate that FOXA2 haploinsufficiency negatively impacted endocrine islet development by downregulating key TFs associated with islet cell development and upregulating genes associated with neuronal development and WNT and BMP pathways. Taken together with other recent reports17,18,23, our findings suggest that FOXA2 defects may lead to monogenic diabetes based on the gene dosage. However, it will be of interest to perform further studies to investigate how FOXA2 works with other TFs to main normal pancreatic endocrine development in human.
Materials and methods
Study approval
The protocols to obtain blood samples were approved by the Institutional Review Board (IRB) of Sidra Medicine (no. 1702007608) and QBRI/HBKU (no. 2018-002). Informed consent forms were signed by human subjects.
Generation of iPSCs and their differentiation into pancreatic beta cells
iPSCs were generated from a patient with FOXA2 heterozygous deletion (QBRIi009-A; FOXA2+/− iPSCs) and healthy controls (Ctr1-iPSCs and Ctr2-iPSCs) as we previously reported24,25. All lines were extensively characterized as previously reported24,25. The first four stages of the pancreatic differentiation were performed using our protocol27 and for further differentiation to beta cells, Rezania et al.28 protocol was used (Supplementary Fig. S3A and Supplementary Table S1).
For WNT signaling inhibition, the cells were treated with the WNT inhibitor, 5 µM XAV939 (Torics Bioscience, 3748) for 4 days during stage 4 and the control cells were treated with DMSO. For BMP inhibition, the cells were treated every day during stage 4 with 100 ng/mL NOGGIN, while the control cells were treated with 50 ng/mL of NOGGIN. The cells were collected at end of stage 4 to examine the effect of WNT and BMP inhibition on the dysregulated genes associated with FOXA2 deficiency.
Immunocytochemistry, flow cytometry, RT-PCR, and qPCR
Immunostaining and flow cytometry were performed as previously reported27,29. RT-PCR and qPCR were performed as previously reported25. (The antibody and primer details are listed in protocol was used in Supplementary Tables S2 and S3.)
Western blot analysis
Total protein was extracted using RIPA buffer and separated by SDS-PAGE. The PVDF membranes were blocked with 10% skim milk in 0.5% TBST, incubated with primary antibodies overnight at 4 °C, and incubated with secondary antibodies at room temperature (protocol was used Supplementary Table S2). The membranes were incubated in SuperSignal West Pico Chemiluminescent substrate (Pierce, Loughborough, UK) detection reagent (VWR) and visualized using the iBright™ CL 1000 Imaging System (Invitrogen).
RNA-sequencing analysis
The RNA-Sequencing (RNA-Seq) experiments and data analysis were performed as we previously reported25,47. For the analysis, the RNA-Seq data were pooled from two Ctr-iPSC lines and three FOXA2+/− iPSC lines of two independent experiments.
CRISPR/Cas9-mediated generation of FOXA2-KO iPSCs
We generated two FOXA2 iPSC lines from two control iPSC lines (Ctr1 and Ctr2). The cells were transfected with a plasmid expressing spCas9 and gRNA (Addgene, 79144) using Lipofectamine Stem Transfection Reagent following the manufacturer’s instructions (ThermoFisher Scientific) (Supplementary Fig. S4 and Supplementary Table S8).
Glucose-stimulated insulin secretion (GSIS)
Beta-cell aggregates were incubated with low (2.8 mM) and high (20 mM) glucose, which was repeated two times. In the end, the clusters were incubated in low glucose with 30 mM KCL to depolarize the cells and release their C-PEP (C-PEPTIDE) contents. The C-PEP concentration in the collected supernatants was measured using Human C-Peptide ELISA Kit (Abcam, ab178641). Each value of C-PEP level of low or high glucose condition was normalized to its own total content of C-PEP in each sample. The stimulation indices were determined through the ratio between the C-PEP level at high glucose concentration to its own basal level secretion at the two challenges.
Overexpression of FOXA2 and PDX1
At the end of day 2 of stage 4, the cells were dissociated with TrypLE and resuspended in the differentiation medium with 10 μM Y-27632. The resuspended cells were either transfected with FOXA2 plasmid (clone HsCD00330288 in pLenti6.2/v5-DEST; DNASU Plasmid Repository, Arizona State University, Tempe, AZ), PDX1 plasmid (pLenti-GIII-CMV-RFP-2A-Puro, LV259702, abm), or the empty vectors. Transfection was carried out using the Lipofectamine™ 3000 Transfection Reagent following the manufacturer’s instruction (ThermoFisher, L3000015). At 48 h post-transfection (end of stage 4), the cells were harvested for RNA and protein extraction. Some experiments were continued until the end of stage 7 (beta-cell stage) for further analyses.
Statistical analysis
At least three biological replicates were used in most of the experiments and statistical analysis was carried out using unpaired two-tailed Student’s t-test by Prism 8.
Data availability
The RNA-Seq data sets generated in the current study are available on the Zenodo repository at https://zenodo.org/record/4362011#.X98RVC0RrAI
Change history
09 April 2021
The OA funding note had to be added to fulfill the contractual requirement of the QNL Fully OA agreement.
References
Al-Khawaga, S. et al. Pathways governing development of stem cell-derived pancreatic beta cells: lessons from embryogenesis. Biol. Rev. Camb. Philos. Soc. 93, 364–389 (2018).
Pan, F. C. & Wright, C. Pancreas organogenesis: from bud to plexus to gland. Dev. Dyn. 240, 530–565 (2011).
Murphy, R., Ellard, S. & Hattersley, A. T. Clinical implications of a molecular genetic classification of monogenic beta-cell diabetes. Nat. Clin. Pr. Endocrinol. Metab. 4, 200–213 (2008).
De Franco, E. From biology to genes and back again: gene discovery for monogenic forms of beta-cell dysfunction in diabetes. J. Mol. Biol. 432, 1535–1550 (2020).
Friedman, J. R. & Kaestner, K. H. The Foxa family of transcription factors in development and metabolism. Cell Mol. Life Sci. 63, 2317–2328 (2006).
Kaestner, K. H., Hiemisch, H. & Schutz, G. Targeted disruption of the gene encoding hepatocyte nuclear factor 3gamma results in reduced transcription of hepatocyte-specific genes. Mol. Cell Biol. 18, 4245–4251 (1998).
Weinstein, D. C. et al. The winged-helix transcription factor HNF-3 beta is required for notochord development in the mouse embryo. Cell 78, 575–588 (1994).
Ang, S. L. & Rossant, J. HNF-3 beta is essential for node and notochord formation in mouse development. Cell 78, 561–574 (1994).
Willmann, S. J. et al. The global gene expression profile of the secondary transition during pancreatic development. Mech. Dev. 139, 51–64 (2016).
Gao, N. et al. Dynamic regulation of Pdx1 enhancers by Foxa1 and Foxa2 is essential for pancreas development. Genes Dev. 22, 3435–3448 (2008).
Gao, N. et al. Foxa1 and Foxa2 maintain the metabolic and secretory features of the mature beta-cell. Mol. Endocrinol. 24, 1594–1604 (2010).
Lee, C. S. et al. Foxa2 controls Pdx1 gene expression in pancreatic beta-cells in vivo. Diabetes 51, 2546–2551 (2002).
Sund, N. J. et al. Tissue-specific deletion of Foxa2 in pancreatic beta cells results in hyperinsulinemic hypoglycemia. Genes Dev. 15, 1706–1715 (2001).
Dines, J. N. et al. Expanding phenotype with severe midline brain anomalies and missense variant supports a causal role for FOXA2 in 20p11.2 deletion syndrome. Am. J. Med. Genet. A 179, 1783–1790 (2019).
Giri, D. et al. Novel FOXA2 mutation causes hyperinsulinism, hypopituitarism with craniofacial and endoderm-derived organ abnormalities. Hum. Mol. Genet. 26, 4315–4326 (2017).
Vajravelu, M. E. et al. Congenital hyperinsulinism and hypopituitarism attributable to a mutation in FOXA2. J. Clin. Endocrinol. Metab. 103, 1042–1047 (2018).
Gaulton, K. J. et al. Genetic fine mapping and genomic annotation defines causal mechanisms at type 2 diabetes susceptibility loci. Nat. Genet. 47, 1415–1425 (2015).
Stekelenburg, C. et al. Exome sequencing identifies a de novo FOXA2 variant in a patient with syndromic diabetes. Pediatr. Diabetes 20, 366–369 (2019).
Bastidas-Ponce, A. et al. Foxa2 and Pdx1 cooperatively regulate postnatal maturation of pancreatic beta-cells. Mol. Metab. 6, 524–534 (2017).
Heddad Masson, M. et al. Foxa1 and Foxa2 regulate alpha-cell differentiation, glucagon biosynthesis, and secretion. Endocrinology 155, 3781–3792 (2014).
Lantz, K. A. et al. Foxa2 regulates multiple pathways of insulin secretion. J. Clin. Invest. 114, 512–520 (2004).
Abdelalim, E. M. Modeling different types of diabetes using human pluripotent stem cells. Cell Mol. Life Sci. https://doi.org/10.1007/s00018-020-03710-9. (2020).
Lee, K. et al. FOXA2 is required for enhancer priming during pancreatic differentiation. Cell Rep. 28, 382–393.e387 (2019).
Elsayed, A. K. et al. Generation of a human induced pluripotent stem cell line (QBRIi009-A) from a patient with a heterozygous deletion of FOXA2. Stem Cell Res. 42, 101705 (2020).
Ali, G. et al. Keratinocytes derived from patient-specific induced pluripotent stem cells recapitulate the genetic signature of psoriasis disease. Stem Cells Dev. 29, 383–400 (2020).
Mohammed, I. et al. Haploinsufficiency of the FOXA2 associated with a complex clinical phenotype. Mol. Genet. Genomic Med 8, e1086 (2020).
Memon, B., Karam, M., Al-Khawaga, S. & Abdelalim, E. M. Enhanced differentiation of human pluripotent stem cells into pancreatic progenitors co-expressing PDX1 and NKX6.1. Stem Cell Res. Ther. 9, 15 (2018).
Rezania, A. et al. Reversal of diabetes with insulin-producing cells derived in vitro from human pluripotent stem cells. Nat. Biotechnol. 32, 1121–1133 (2014).
Aigha, I. I., Memon, B., Elsayed, A. K. & Abdelalim, E. M. Differentiation of human pluripotent stem cells into two distinct NKX6.1 populations of pancreatic progenitors. Stem Cell Res. Ther. 9, 83 (2018).
Ben-Shushan, E., Marshak, S., Shoshkes, M., Cerasi, E. & Melloul, D. A pancreatic beta -cell-specific enhancer in the human PDX-1 gene is regulated by hepatocyte nuclear factor 3beta (HNF-3beta), HNF-1alpha, and SPs transcription factors. J. Biol. Chem. 276, 17533–17540 (2001).
Cernilogar, F. M. et al. Pre-marked chromatin and transcription factor co-binding shape the pioneering activity of Foxa2. Nucleic Acids Res. 47, 9069–9086 (2019).
Wang, H., Gauthier, B. R., Hagenfeldt-Johansson, K. A., Iezzi, M. & Wollheim, C. B. Foxa2 (HNF3beta) controls multiple genes implicated in metabolism-secretion coupling of glucose-induced insulin release. J. Biol. Chem. 277, 17564–17570 (2002).
Gao, T. et al. Pdx1 maintains beta cell identity and function by repressing an alpha cell program. Cell Metab. 19, 259–271 (2014).
Bernardo, A. S. et al. Biphasic induction of Pdx1 in mouse and human embryonic stem cells can mimic development of pancreatic beta-cells. Stem Cells 27, 341–351 (2009).
Lammert, E., Brown, J. & Melton, D. A. Notch gene expression during pancreatic organogenesis. Mech. Dev. 94, 199–203 (2000).
Rooman, I. et al. Expression of the Notch signaling pathway and effect on exocrine cell proliferation in adult rat pancreas. Am. J. Pathol. 169, 1206–1214 (2006).
Li, X. Y., Zhai, W. J. & Teng, C. B. Notch signaling in pancreatic development. Int. J. Mol. Sci. 17, 48 (2015).
Afelik, S. et al. Notch-mediated patterning and cell fate allocation of pancreatic progenitor cells. Development 139, 1744–1753 (2012).
Shih, H. P. et al. A Notch-dependent molecular circuitry initiates pancreatic endocrine and ductal cell differentiation. Development 139, 2488–2499 (2012).
Sharon, N. et al. Wnt signaling separates the progenitor and endocrine compartments during pancreas development. Cell Rep. 27, 2281–2291 e2285 (2019).
Tan, M., Jiang, L., Li, Y. & Jiang, W. Dual inhibition of BMP and WNT signals promotes pancreatic differentiation from human pluripotent stem cells. Stem Cells Int. 2019, 5026793 (2019).
Gauthier, B. R. et al. Hepatic nuclear factor-3 (HNF-3 or Foxa2) regulates glucagon gene transcription by binding to the G1 and G2 promoter elements. Mol. Endocrinol. 16, 170–183 (2002).
Gosmain, Y., Avril, I., Mamin, A. & Philippe, J. Pax-6 and c-Maf functionally interact with the alpha-cell-specific DNA element G1 in vivo to promote glucagon gene expression. J. Biol. Chem. 282, 35024–35034 (2007).
Huopio, H. et al. Dominantly inherited hyperinsulinism caused by a mutation in the sulfonylurea receptor type 1. J. Clin. Invest. 106, 897–906 (2000).
Dusatkova, P. et al. HNF1A mutation presenting with fetal macrosomia and hypoglycemia in childhood prior to onset of overt diabetes. J. Pediatr. Endocrinol. Metab. 24, 187–189 (2011).
Kapoor, R. R. et al. Persistent hyperinsulinemic hypoglycemia and maturity-onset diabetes of the young due to heterozygous HNF4A mutations. Diabetes 57, 1659–1663 (2008).
Karam, M., Younis, I., Elareer, N. R., Nasser, S. & Abdelalim, E. M. Scalable generation of mesenchymal stem cells and adipocytes from human pluripotent stem cells. Cells 9, 710 (2020).
Acknowledgements
We thank Ms. Bushra Memon (QBRI/HBKU) for helping in iPSC characterization. Also, we thank Dr. Gerald Pfister (QBRI/HBKU) for technical assistance in cell sorting and the Genomic Core (QBRI/HBKU) for technical support in RNA-sequencing. This work was funded by grants from Qatar Biomedical Research Institute (QBRI)/HBKU (Grant No. IGP 2014009; IGP 2016001) and from Qatar National Research Fund (QNRF) (Grant No. NPRP9-283-3-056).
Funding
Open access funding provided by the Qatar National Library.
Author information
Authors and Affiliations
Contributions
E.M.A. conceived and designed the study and obtained research funding. A.K.E. performed the experiments. A.K.E. and E.M.A. analyzed and interpreted the data and wrote the manuscript. I.Y. analyzed the RNA-Seq data. G.A. participated in some experiments. K.H. contributed to sample and clinical data collection. All authors reviewed and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
The study has been approved by the appropriate institutional research ethics committee and has been performed in accordance with the ethical standards as laid down in the 1964 Declaration of Helsinki and its later amendments or comparable ethical standards.
Informed consent
Informed consent was obtained from all individual participants included in the study.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by A. Finazzi-Agrò
Supplementary information
41419_2021_3390_MOESM4_ESM.docx
Supplementary Table 4: Top downregulated genes in PP2 derived from FOXA2+/- iPSCs in comparison to those derived from Ctr-iPSCs
41419_2021_3390_MOESM5_ESM.docx
Supplementary Table 5: Top Upregulated genes in PP2 derived from FOXA2+/- iPSCs in comparison to those derived from Ctr-iPSCs
41419_2021_3390_MOESM6_ESM.docx
Supplementary Table 6: Top downregulated genes in the EPs derived from FOXA2+/- iPSCs in comparison to those derived from Ctr-iPSCs
41419_2021_3390_MOESM7_ESM.docx
Supplementary Table 7: Top upregulated genes in the EPs derived from FOXA2+/- iPSCs in comparison to those derived from Ctr-iPSCs
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Elsayed, A.K., Younis, I., Ali, G. et al. Aberrant development of pancreatic beta cells derived from human iPSCs with FOXA2 deficiency. Cell Death Dis 12, 103 (2021). https://doi.org/10.1038/s41419-021-03390-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-021-03390-8
This article is cited by
-
Chromosome 20p11.2 deletions cause congenital hyperinsulinism via the loss of FOXA2 or its regulatory elements
European Journal of Human Genetics (2024)
-
Genome-wide differential expression profiling of long non-coding RNAs in FOXA2 knockout iPSC-derived pancreatic cells
Cell Communication and Signaling (2023)
-
iPSC-Derived Pancreatic Progenitors Lacking FOXA2 Reveal Alterations in miRNA Expression Targeting Key Pancreatic Genes
Stem Cell Reviews and Reports (2023)
-
Loss of FOXA2 induces ER stress and hepatic steatosis and alters developmental gene expression in human iPSC-derived hepatocytes
Cell Death & Disease (2022)