Abstract
BRCA2 is crucial for repairing DNA double-strand breaks with high fidelity, and loss of BRCA2 increases the risks of developing breast and ovarian cancers. Herein, we show that BRCA2 is inactively mutated in 10% of gastric and 7% of colorectal adenocarcinomas, and that this inactivation is significantly correlated with microsatellite instability. Villin-driven Brca2 depletion promotes mouse gastrointestinal tumor formation when genome instability is increased. Whole-genome screening data showed that these BRCA2 monoallelic and biallelic mutant tumors were selectively inhibited by mitomycin C. Mechanistically, mitomycin C provoked double-strand breaks in cancer cells that often recruit wild-type BRCA2 for repair; the failure to repair double-strand breaks caused cell-cycle arrest at the S phase and p53-mediated cell apoptosis of BRCA2 monoallelic and biallelic mutant tumor cells. Our study unveils the role of BRCA2 loss in the development of gastrointestinal tumors and provides a potential therapeutic strategy to eliminate BRCA2 monoallelic and biallelic mutant tumors through mitomycin C.
Similar content being viewed by others
Introduction
Homologous recombination (HR) is an evolutionarily conserved process for repairing DNA double-strand breaks (DSBs) with high fidelity1. Upon DNA damage, HR-repair proteins such as BRCA2 directly bind with RAD51 through BRC motifs and promote RAD51 filament formation to recruit homologous templates to complete error-free DSB repair2,3,4. The employment of BRCA1, which lacks physical interaction with RAD51 to damaged chromatin, is also important for HR-mediated DSBs but is not fully understood5. When BRCA1 and/or BRCA2 is defective, nonhomologous end-joining (NHEJ) with no requirement for homologous template is an alternative process used to repair DSBs with potential genome instability, which accelerates tumor development6,7.
BRCA1 and BRCA2 are frequently mutated at both the germline and somatic levels in human breast, ovarian, and pancreatic cancers8,9. For example, BRCA2 is mutated in approximately 2.5% of breast cancers and 6% of ovarian cancers10,11. The inappropriate DSB repair caused by defective BRCA1/2 often leads to chromosomal dislocation, although this is not sufficient to drive tumor formation per se. However, simultaneous inactivation of Trp53 strongly promotes spontaneous tumor formation in the breast, ovarian, and pancreatic organs of mice12,13,14,15. This suggests that BRCA1/2 loss allows uncontrolled genetic mutations that drive tumorigenesis16.
Poly (ADP-ribose) polymerase (PARP) is a family of proteins that are responsible for repairing DNA single-strand breaks (SSBs). When BRCA1 and BRCA2 are fully depleted, PARP inhibition is synthetically lethal for these HR-defective tumors17,18. To date, PARP inhibitors have achieved remarkable clinical responses for treating BRCA1/2 deficient breast and ovarian tumors19,20,21. However, these HR-defective tumors quickly become resistant to PARP inhibitors as well as platinum analogues through secondary mutation of the BRCA1/2 gene even in a single allele22,23,24. Therefore, the need for potent therapeutic agents to treat BRCA1/2 monoallelic and biallelic mutant tumors is unmet.
Gastric and colorectal cancers are two major common cancer types worldwide, and the 5-year survival rates are ~30% and 60% for gastric and colorectal tumors, respectively25,26. To date, fluorouracil and platinum-based chemotherapy is still the first-line choice for treating gastrointestinal cancers, except for tumors with EGFR and HER2 expression27. Recently, the genetic landscapes of mutations in gastric and colorectal tumors have been well characterized and made known to the public28,29,30. By retrieving these data, we observed that BRCA2 is mutated in gastric and colorectal tumor samples at frequencies of 10% and 7%, respectively, but the role of defective BRCA1/2 in gastrointestinal tumorigenesis is still unknown5. To address this question, we established a Villin-driven Brca2 conditional knockout mouse to monitor gastrointestinal tumor formation, with or without the application of N-methyl-N-nitrosourea treatment or Trp53 deletion. Meanwhile, we identified mitomycin C as the most effective agent to eliminate BRCA2 monoallelic and biallelic mutant tumors by inhibiting BRCA2. Our study not only uncovers the role of BRCA2 mutation in gastrointestinal tumorigenesis, but also provides a feasible therapeutic agent to potently eliminate BRCA2 monoallelic and biallelic mutant tumors other than for those with p53 deficiency.
Results
BRCA2 is frequently mutated in both gastric and colorectal tumor samples and is associated with microsatellite instability
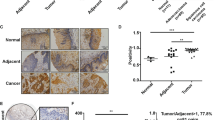
By retrieving the TCGA stomach cancer database28,31,32, we observed that genes belonging to the Fanconi anaemia (FA) pathway were frequently mutated in gastric adenocarcinoma33,34,35 (Fig. 1a). BRCA2, also termed FANCD1, ranked as the top candidate of 15 bona fide FA members at a frequency of 10% mutation (41/293), while BRCA1, another FA-like member, had an ~5% mutation rate (17/293). A similar pattern was also observed in colorectal cancer samples30, showing that BRCA1 and BRCA2 were mutated in ~4% (26/619) and 7% (59/619) of colorectal tumor samples, respectively (Fig. S1a). Thus, we selected the top candidate BRCA2 to perform more studies. Notably, BRCA2 mutations were associated with three features: (1) they were likely inactive (3 nonsense, 23 frameshift, and 74 missense mutations); (2) they were significantly associated with a microsatellite instability tumor subtype (p = 1.042E−11 in 293 gastric tumor samples and p = 7.39E−11 in 619 colorectal tumor samples by two-sided Fisher’s exact test), suggesting that frameshift mutations of BRCA2 gene (20/23 cases) might be caused by deregulation of the mismatch repair pathway (Supplemental Table 1)36; and (3) they were not significantly associated with TP53 mutations (p = 0.73 and p = 0.079 in gastric and colorectal tumors, respectively, by two-sided Fisher’s exact test). To examine the relationship between BRCA2 and patient survival, we further selected the TCGA provisional gastric and colorectal tumor databases to perform survival rate analysis, and the results showed that the low levels of BRCA2 mRNA expression were significantly correlated with the poor survival rates of both gastric and colorectal cancer patients (gastric: p = 0.04, n = 354; colon: p = 0.04, n = 597; Figs. 1b and S1b). Taken together, these data led us to assume that BRCA2 deficiency was pivotal for developing gastrointestinal tumors.
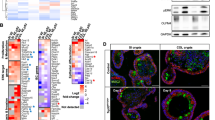
a Mutation status of FA- and FA-like members in 2014 TCGA stomach adenocarcinoma samples (n = 293). MSI, high microsatellite instability; GS, genomic stable; EBV, Epstein-Barr virus positivity; CIN, chromosomal instability. b Survival curves of BRCA2-low (n = 209) and BRCA2-high (n = 145) mRNA expression cases from provisional TCGA gastric cancer databases. *p < 0.05. c Brca2 staining in the stomach tissues of Brca2 wild-type and knockout mice. Scale bar, 10 μm. d Experimental design: Three-week-old Brca2 knockout mice or littermate wild-type mice were divided into four groups; MNU was administered to mice in two groups via drinking water at 120 mg/L on alternate weeks for a total exposure of 10 weeks, and vehicle was administered to the other two groups. e Disease incidence of Villin-Cre−; Brca2fl/fl mice (n = 10, WT group), Villin-Cre+; Brca2fl/fl (n = 12, KO group), Villn-Cre−; Brca2fl/fl with MNU (n = 16, WT plus MNU group), and Villin-Cre+; Brca2fl/fl with MNU (n = 18, KO plus MNU group). f Tumor incidence and location of all the tested mice in the four groups. g Kaplan–Meier survival curves of Brca2 wild-type (n = 10) and knockout mice (n = 12) without MNU treatment. h Kaplan–Meler survival curves of Brca2 wild-type (n = 8) and knockout mice (n = 6) with MNU treatment. **p < 0.01. i Gross morphology and H&E staining of stomach sections from 10-month-old MNU-treated Brca2 wild-type and knockout mice. The white arrow indicates the visible tumors of Brca2 knockout mice. Scale bar, 200 μm. j Disease incidence and location of Villin-Cre+; Brca2+/+; Trp53fl/+ mice (Group A, n = 5) and Villin-Cre+; Brca2fl/fl; Trp53fl/+ mice (Group B, n = 9). k Kaplan–Meier survival curves of Villin-Cre+; Brca2+/+; Trp53fl/+ and Villin-Cre+; Brca2fl/fl; Trp53fl/+ mice. *p < 0.05.
Villin-driven Brca2 deficiency induces tumorigenesis in gastrointestinal epithelial cells
To verify the role of BRCA2 deficiency in tumor development, we generated Villin-Cre+; Brca2fl/fl mice to achieve Brca2 deletion in gastrointestinal epithelial cells12,37. Brca2 was selectively depleted in gastrointestinal epithelial cells of Brca2 knockout mice at protein and genetic levels compared to that of Brca2 wild-type mice (Figs. 1c, S1c and S1d). When housed in specific pathogen-free conditions, Villin-Cre+; Brca2fl/fl (KO) mice had a very small chance of developing spontaneous intestinal tumor (1/12 mice) compared with the Villin-Cre-; Brca2fl/fl (WT) group (Fig. 1d–f), showing no difference in the median survival time between the two groups (505 vs 560.5 days, p > 0.05, Fig. 1g). These data suggest that Brca2 deficiency alone is not sufficient to promote gastrointestinal tumorigenesis.
Gastrointestinal tumor-associated risk factors such as Helicobacter pylori infection, nitrate and nitrite uptake often lead to genome instability38,39. Therefore, we selected N-methyl-N-nitrosourea (MNU), an alkylating agent widely accepted for inducing gastrointestinal tumorigenesis40, to test whether Brca2 deficiency can elicit gastrointestinal tumor formation when increasing genome instability (Fig. 1d). MNU treatment strongly provoked the formation of spontaneous and invasive gastric adenocarcinomas at both antrum and corpus sites in 9 out of 18 mice in the KO group (50%), and hyperplasia in the majority of the remaining mice (40%) (Fig. 1e, f, and i). Additionally, MNU-treated KO mice had a much lower survival rate than MNU-treat WT mice (p < 0.01, Fig. 1h). As a control, MNU-treated WT mice had no detectable tumors with normal and hyperplastic conditions observed in 70% and 30% of all tested mice, respectively (Fig. 1e, f, and i). These data suggest that Brca2 deficiency can boost gastrointestinal tumor formation when genomic DNA is damaged.
p53 is a key guardian of genome stability that prevents cells from entering the S phase with unrepaired DNA sequences41. When heterozygously inactivated, it robustly facilitates breast tumor formation in Brca2 conditional knockout mice12. Thus, we asked whether Brca2 loss can promote gastrointestinal tumorigenesis when Trp53 is heterozygously depleted by Villin-Cre expression (Fig. S1c and S1e). In the Villin-Cre+; Brca2fl/fl; Trp53fl/+ group (Group B), seven out of nine tested mice (78%) developed spontaneous and invasive adenocarcinomas at stomach and/or intestine sites, while no tumors detected in the Villin-Cre+; Brca2+/+; Trp53fl/+ group (Group A, 5/5 mice) (Fig. 1j). The H&E staining of tumor samples is shown in Fig. S1f. Mice in group B had a significantly lower survival rate than those in group A (p < 0.05, log-rank test, Fig. 1k).
Collectively, Brca2 deficiency driven by Villin-Cre can boost gastrointestinal tumor formation when genome instability is increased. In particular, Brca2 deficiency promotes gastric tumor formation when genome stability is selectively impaired in gastric epithelial cells.
Mitomycin C potently eliminates BRCA2 monoallelic and biallelic mutant tumor cells
In the literature, cancer cells with full BRCA2 deficiency are sensitive to DNA-damaging agents including mitomycin C (MMC)6, cisplatin42, melphalan43,44 and olaparib17,18,19,20,21. Therefore, we examined whether gastrointestinal cancer cell lines harboring BRCA2 mutations are sensitive to these DNA-damaging agents. The great majority of gastrointestinal tumor cell lines used in this study harbor BRCA1/2 heterozygous mutations; the exception is HGC-27, which harbors homozygous BRCA2p.F2058L mutation (Fig. S2a)45. Cell viability data demonstrated that olaparib, MMC and melphalan significantly decreased the viability of BRCA2 monoallelic and biallelic mutant gastrointestinal tumor cells at a concentration of 10 µM compared with that of BRCA2 wild-type tumor cells (Fig. S2b–S2e). Consistent with previous reports46, 1 µM olaparib treatment for 5 days significantly decreased about 50% of the viability of HGC-27 cells (Fig. S2f). Cisplatin achieved significant inhibition in HGC-27 cells when exposed for a longer time at 20 µM (Fig. S2g). Among all the tested chemicals, MMC exhibited the most potent inhibitory effects in all six BRCA2 monoallelic and biallelic mutant gastrointestinal cancer cell lines within 3 days at a concentration of 300 nM, surpassing the extent of inhibition by olaparib and other DNA-damaging agents (Fig. 2a). The BRCA2 mutations of SNU-1, SNU-5, and HGC-27 cell lines were validated by Sanger sequencing, showing two monoallelic mutations and one biallelic mutation of the BRCA2 gene in these three cell lines (Fig. S2h).
a After SNU-1, SNU-5 and HGC-27 cells were treated with the indicated drugs for 72 h, the cell viability was determined at OD570, with normalization to DMSO treatment. b Representative images of γ-H2AX and BRCA2 foci formation in BRCA2 wild-type and mutant cell lines after treated with 0.3 μM MMC for 18 h. Scale bar, 10 μm. c Representative images of γ-H2AX and RAD51 foci formation in BRCA2 wild-type and mutant cell lines after treatment with 0.3 μM MMC or vehicle for 18 h. Scale bar, 10 μm. d The quantification of γ-H2AX and BRCA2 foci formation in BRCA2 wild-type and mutant cell lines in the absence and presence of 0.3 μM MMC treatment for 18 h. e The quantification of γ-H2AX and RAD51 foci formation in BRCA2 wild-type and mutant cell lines after treatment with 0.3 μM MMC or vehicle for 18 h. f The flow cytometry data of SNU-216, SNU-1, SNU-5, and HGC-27 after treatment with 0.1% DMSO or 1 μM MMC for 24 h. g The cell-cycle phases of SNU-216, SNU-1, SNU-5, and HGC-27 were presented after treatment with 0.1% DMSO or 1 μM MMC for 24 h. h The relative S phases of SNU-216, SNU-1, SNU-5, and HGC-27 were calculated and statistical analyzed after treatment with 0.1% DMSO or 1 μM MMC for 24 h. i Single-cell electrophoresis in the indicated cell lines after treatment with 3 μM MMC or vehicle for 36 h. Scale bar, 10 μm. j Quantification of single-cell electrophoresis in indicated cell lines. k Immunoblotting analysis of γ-H2AX, PARP and cleaved-PARP in the indicated cell lines after treatment with 0.3 μM MMC for 18 h plus withdrawal for 24 h. Data represent the mean ± SD of 3–5 replicates, *p < 0.05, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Because BRCA2 recruits RAD51 to DBS sites for executing HR repair3, we asked whether BRCA2/RAD51 signaling is impaired in these BRCA2 monoallelic and biallelic mutant tumor cell lines. MMC treatment induces strong phosphorylation of histone H2AX at Ser 139 (γ-H2AX) (Fig. 2b, c, and k), which is a very early event of DNA damage responses that recruits DNA damage response proteins such as BRCA2 to DSB sites47. BRCA2 foci formation data showed that MMC provoked strong BRCA2 foci formation in the BRCA2 wild-type cell line, which overlapped well with γ-H2AX staining, but it had little or no foci formation in all three BRCA2 monoallelic and biallelic mutant gastrointestinal cancer cell lines (p < 0.0001, Fig. 2b and d). Consistently, RAD51 foci formation data revealed that MMC elicited strong RAD51 foci formation in BRCA2 wild-type cells, showing colocalization with γ-H2AX, but that it failed to do so in BRCA2 monoallelic and biallelic mutant cells (p < 0.0001, Fig. 2c and e). These data suggest that BRCA2/RAD51 signaling is functionally inactivated in BRCA2 monoallelic and biallelic mutant cancer cells upon MMC treatment.
MMC provokes irreversible DNA damage in BRCA2 monoallelic and biallelic mutant cancer cells
MMC, an antibiotic obtained from Streptomyces caespitosus48, causes numerous DSBs in cells by selectively crosslinking GC-rich DNA due to CpG-specific two-step alkylation49,50. When DNA replication is stalled, cells arrest at the G1/S transition to preserve enough time to execute DNA repair; failure to repair severe DNA damage activates apoptotic signaling to eradicate these cells, preventing the rise of DNA mutations and genome instability51. Cell-cycle data showed that MMC markedly arrested these BRCA2 monoallelic and biallelic mutant cell lines at the S phase after 24 h of treatment; in contrast, no significant cell arrest was observed in BRCA2 wild-type cells (Fig. 2f, g, and h). DNA comet tail movement data showed that compared with BRCA2 wild-type cells, severe DNA damage occurred in BRCA2 monoallelic and biallelic mutant cells after MMC treatment for 36 h (Fig. 2i and j).
Interestingly, γ-H2AX was sustained at high levels in all three BRCA2 monoallelic and biallelic mutant tumor cell lines even after MMC removal but returned to baseline phosphorylation status in wild-type SNU-216 cells during the same time frame (Fig. 2k). The irreversible DSB damage finally resulted in the apoptosis of three BRCA2 monoallelic and biallelic mutant tumor cell lines but not of BRCA2 wild-type cells (Fig. 2k). Based on these data, we conclude that MMC induces large amounts of DSBs in cells, which can be quickly removed by BRCA2 wild-type cells, and that the failure to correct these DSBs in BRCA2-defective tumors leads to cell apoptosis.
BRCA2 is a crucial therapeutic target of MMC
Next, we asked why BRCA2 monoallelic and biallelic mutant gastrointestinal tumor cells were hypersensitive to MMC. As shown in Fig. 3a, BRCA2 protein was expressed at lower levels in monoallelic and biallelic mutant tumor cell lines than in wild-type cell lines. Importantly, MMC significantly decreased BRCA2 expression at both the protein and mRNA levels in BRCA2 monoallelic and biallelic mutant cell lines but not in BRCA2 wild-type cell lines (Figs. 3b, S3a, S3b, and S3c)52,53,54. Olaparib and cisplatin had minor inhibitory effects on BRCA2 protein expression levels. These data suggest that BRCA2 inhibition is critical for achieving the therapeutic effects of MMC, which explains why MMC has better antitumor activity than other DNA-damaging reagents due to the inhibition of BRCA2 expression.
a The BRCA2 protein expression in BRCA2 wild-type and mutant gastrointestinal tumor cell lines was shown by immunoblotting. b The BRCA2 protein expression was determined in SNU-1 and HGC-27 cell lines by immunoblotting after treatment with the indicated drugs. c Working flowchart of genome-scale CRISPR-Cas9 knockout (GeCKO) screening. d CRISPR score analysis data showed that BRCA2 knockout selectively sensitized SNU-1 cells to MMC treatment, but did not affect the viability of SNU-1 cells when treated with vehicle. e The knockout efficiency of BRCA2 protein was determined by immunoblotting. f sgBRCA2 and sgControl SNU-1 cells were treated with 0.1 μM MMC or 0.1% DMSO for 72 h, respectively, and the cell viability was determined at OD570. g Primary tumor cells derived from tumor tissues of Villin-Cre+; Brca2fl/fl mice (#1305) were treated with 0.1 and 1 μM MMC or 0.1% DMSO for 72 h. h Cell viability, expression status of BRCA2 and foci formation in HGC-27 cells treated with MMC after introduction with wild-type BRCA2 or control vector. Scale bar, 10 μm. i The viability of SNU-1 and SNU-5 cells in the absence or presence of wild-type BRCA2 was determined when treatment with 0.1 μM MMC for 72 h, 0.1% DMSO was used as a control. j Tumor volume and survival rate of Rag2−/−;Il2r−/− mice harboring SNU-1 tumor xenografts. Mice were intraperitoneally treated with vehicle or 1 mg/kg MMC, once per week for total four weeks, n = 5. k Tumor volume and survival rate of nude mice with SNU-5 tumor xenografts. Mice were intraperitoneally treated with vehicle or 1 mg/kg MMC, once per week for total four weeks, n = 5. l Representative images of SNU-5 tumor-bearing nude mice after 3 weeks of vehicle or MMC treatment. m Tumor volume and survival rate of nude mice with HGC-27 tumor xenografts. Mice were intraperitoneally treated with vehicle or 1 mg/kg MMC, once per week for total 4 weeks, n = 5. n Representative images of HGC-27 tumor-bearing nude mice after three weeks of vehicle or MMC treatment. Data represent the mean ± SD, *p < 0.05, **p < 0.01, ***p < 0.01, ****p < 0.0001.
To verify this hypothesis, we applied genome-scale CRISPR-Cas9 knockout (GeCKO) screening to simultaneously knockout 19050 human genes in SNU-1 cells to explore whether BRCA2 monoallelic mutant tumor cells are selectively targeted by MMC55. The experimental procedure of GeCKO screening is shown in Fig. 3c and the Material and Methods56. At the endpoint of treatment, at least 2 × 107 cells in each group were harvested to achieve 300-fold coverage of the sgRNA library. After analysing the sequencing data with a MAGeCK algorithm57, we obtained a total of 18 gene candidates that significantly enhanced the MMC killing effects but did not reduce the cell viability of the control group when they were removed (ratio < 0.5, FDR < 0.05, with duplicated hits per screening, Supplemental Table 2). BRCA2 was the top 1 candidate selected in this screening (ratio = 0.3, FDR < 0.001, Fig. 3d). By contrast, BRCA1 showed no significant change during the screening (ratio = 1.68, p = 0.12), probably due to the inability to physically associate with RAD515. In addition, we performed a new round of GeCKO screening in BRCA2 wild-type SNU-216 cells, showing that FA genes, including FANCB, FANCF, FANCM, and FANCD2, were positively selected (Fig. S3d). This suggests that knockout of FA genes upstream of BRCA2 has minor effects when BRCA2 is largely impaired but that these genes are functionally important when BRCA2 is intact. Nevertheless, whole-genome screening data suggest that BRCA2 inhibition is critical for optimizing the therapeutic effects of MMC.
Next, we fully depleted BRCA2 protein expression in both alleles of SNU-1 cells using two different sgRNAs (Figs. 3e and S2i). As expected, these full BRCA2-deficient cells were more sensitive to MMC than sgControl cells (p < 0.001, Fig. 3f), suggesting that BRCA2 function was impaired but was still active in SNU-1 cells. To support our notion, primary tumor cells derived from MNU-treated Villin-Cre+; Brca2fl/fl murine spontaneous tumors were dose-dependently inhibited by MMC (p < 0.0001, Fig. 3g). HGC-27 cells were less sensitive to MMC when wild-type BRCA2 was re-expressed (p < 0.001), as evidenced by the fact that homozygous BRCA2p.F2058L alleles were primarily replaced by the wild-type BRCA2 allele (Fig. 3h). MMC treatment could induce the formation of BRCA2 and RAD51 foci when BRCA2 was reintroduced into HGC-27 cells but not in the control cells, indicating that the HR-repair ability was partially restored in the BRCA2-re-expressing cells (Fig. 3h). Similarly, both SNU-1 and SNU-5 cells were more insensitive to MMC when wild-type BRCA2 was reintroduced (Fig. 3i). Taken together, we conclude that BRCA2 is a target that could be selectively inhibited by MMC.
MMC eliminates BRCA2 monoallelic and biallelic mutant tumor xenografts in vivo
Next, we tested the in vivo efficacy of MMC in three BRCA2 monoallelic and biallelic mutant tumor xenograft models. Compared to vehicle treatment, MMC treatment potently inhibited the in vivo growth of heterozygous BRCA2 monoallelic mutant SNU-1 tumor xenografts in immune-deficient mice (p < 0.01, Fig. 3j top) and significantly improved the survival rate of the mice (p < 0.05, Fig. 3j bottom). Similar results were obtained in two other BRCA2 monoallelic and biallelic mutant tumor xenograft models (p < 0.01, Fig. 3k and m upper), and significantly improved survival rates for the MMC-treated mice compared with the control mice (p < 0.01, Fig. 3k and m bottom). Notably, MMC induced complete tumor regression in three out of five mice bearing HGC-27 tumor xenografts and five out of five mice bearing SNU-5 tumor xenografts (Fig. 3l and n).
p53 is responsible for MMC-induced apoptosis of BRCA2 monoallelic and biallelic mutant gastrointestinal tumors
We further investigated how MMC triggers the apoptosis of BRCA2 monoallelic and biallelic mutant cancer cells. Gene set enrichment analysis (GSEA)58 revealed that the gene signature of the p53 pathway was positively selected after comparing MMC with vehicle in SNU-1 cells (FDR < 0.0001, Fig. 4a and b). The mRNA expression levels of p53 transcriptionally regulated CDKN1A, PMAIP1, MDM2, and GADD45A genes were markedly increased after MMC treatment (Fig. 4a). Therefore, we asked whether p53 is responsible for MMC-induced cell apoptosis. To test this hypothesis, we depleted TP53 gene expression in SNU-1 cells using two independent sgRNAs and observed that TP53 knockout subclones were no longer sensitive to MMC compared with sgControl subclones (p < 0.0001, Fig. 4c). The p53 gene expression signature was also reversed by comparing MMC-treated sgControl and sgTP53 cells (FDR < 0.0001, Fig. 4d). The knockout effect of p53 protein in two subclones is presented in the upper panel of Fig. 4c. When wild-type p53 was reintroduced, these sgTP53 cells were re-sensitized to MMC (p < 0.0001, Fig. 4c), reflecting that it is not an off-target effect. Similar results were also observed in the BRCA2p.I2672fs mutant HCT116 colorectal cancer cell line (p < 0.0001, Fig. 4e). Importantly, TP53 knockout not only abolished the killing effects of MMC in vitro but also greatly reduced the therapeutic efficacy of MMC in vivo (p > 0.05, Fig. 4f). Primary cells derived from Villin-Cre+; Brca2fl/fl; Trp53fl/+ spontaneous tumor also became less sensitive to MMC than those derived from Villin-Cre+; Brca2fl/fl; Trp53+/+ (Fig. 4g).
a Nonbiased clustering of the top 20 up and downregulated genes by comparison MMC-treated to DMSO-treated SNU-1 cells. b Gene set enrichment analysis of the gene signature of the p53 pathway in SNU-1 cells treated with 3 μM MMC or 0.1% DMSO for 36 h. c SNU-1 sgTP53, sgControl and p53-putback cells were treated with 0.3 μM MMC or 0.1% DMSO for 72 h, and cell viability was determined at OD570. The knockout effect of p53 protein is shown in the upper panel. d Gene set enrichment analysis of the gene signature of the p53 pathway in SNU-1 (TP53 WT) and SNU-1(TP53 KO) cells treated with 3 μM MMC for 36 h. e HCT116 sgTP53 and sgControl cells were treated with 0.1 μM MMC or 0.1% DMSO for 72 h, and the cell viability was determined at OD570. The knockout effects of p53 protein are shown in the upper panel. f Tumor volume of SNU-1 (TP53 KO) tumor xenografts in nude mice. Mice were intraperitoneally treated with vehicle or 3 mg/kg MMC, once per week for a total of four weeks, n = 5. Representative images are shown on the right. g Cell viability of primary cancer cells derived from Villin-Cre+; Brca2fl/fl; Trp53+/+ mouse or Villin-Cre+; Brca2fl/fl; Trp53fl/+ mice treated with 1 μM MMC or 0.1% DMSO for 72 h. h RT-qPCR analysis of NOXA and PUMA mRNA levels in SNU-1 sgControl and sgTP53 cells when treated with 3 μM MMC or 0.1% DMSO for 24 h. i Immunoblotting analysis of PARP, cleaved-PARP, p53, p-p53(ser15), NOXA and PUMA of sgControl and sgTP53 SNU-1 cells. Cells were treated with 3 μM MMC or 0.1% DMSO for 36 h. j Cell viability of the SNU-1 NOXA knockout cell line, control cell line (sgControl), SNU-1 PUMA KD cell line and control cell line (Scramble) treated with 0.1 μM MMC or 0.1% DMSO for 72 h. Cell viability was determined at OD570. The knockout effects of NOXA and knockdown effects of PUMA are shown on the right. Data are presented as the mean ± SD of 3–5 replicates, **p < 0.01, ***p < 0.001, ****p < 0.0001.
Next, we asked which downstream gene is responsible for p53-mediated cell apoptosis. NOXA and PUMA are considered two key effectors of the p53 apoptosis pathway59. In this study, we observed that MMC significantly induced the mRNA and protein expression levels of both NOXA (encoded by the PMAIP1 gene) and PUMA, but it was diminished when TP53 was knocked out (Fig. 4h and i), suggesting that it is a p53-dependent event. Notably, individual inhibition of NOXA or PUMA significantly reduced the in vitro efficacy of MMC (both p < 0.01, Fig. 4j), indicating that NOXA and PUMA are required for p53-mediated apoptosis of BRCA2 monoallelic and biallelic mutant cells upon MMC treatment. As we previously mentioned, TP53 mutations were almost equally distributed in both BRCA2 wild-type and mutant gastrointestinal tumors, and it would be of value to examine the genetic mutations of the TP53 gene before applying MMC to patients with BRCA2 monoallelic and biallelic mutant tumors.
Discussion
It has been known for many years that BRCA2 deficiency increases the lifetime risk of developing breast, ovarian, and pancreatic cancers60,61,62,63. However, the role of BRCA2 mutants in gastrointestinal cancer is still largely unknown. In this study, we first showed that BRCA2 is mutated in ~7–10% of gastrointestinal tumors, and that it was significantly correlated with microsatellite instability. For example, 20 out of 23 frameshift mutations occurred at the polytract sites of the BRCA2 gene. Coincidentally, a recent study reported that the tumors of 2.3% of gastrointestinal cancer patients harboring deleterious alterations in BRCA2 were associated with high microsatellite instability64. Thus, it is a unique way for gastrointestinal tumors to develop BRCA2 mutations that is distinct from how these mutations occur in breast and ovarian cancer.
Second, we showed that Brca2 deficiency alone driven by Villin-Cre was not sufficient to promote gastrointestinal tumor formation but that it robustly elicited tumor formation when genome instability was increased. MNU treatment preferentially induced tumor in the stomach, while heterozygous depletion of Trp53 provoked cancer in both stomach and intestine. In addition, we did not observe tumor formation when Trp53 was heterozygously depleted alone in gastrointestinal epithelial cells. These data indicate that co-mutation of Brca2 and Trp53 may have a synergic effect towards inducing tumorigenesis.
BRCA2 deficiency can increase chromosomal instability when the error-free HR-repair system is impaired and eventually contributes to carcinogenesis6. p53 is a guardian of the genome for keeping the genome intact, and will prevent cells from entering the cell cycle with unrepaired DSBs65. When p53 is defective, BRCA2-mutant cells can quickly accumulate genomic instability and lead to tumorigenesis without inducing cell apoptosis. This may explain why other researchers cannot observe tumor formation when Brca2 was deleted alone in the small intestine66. At the same time, we observed that MMC treatment induced cell apoptosis in a p53-dependent manner. Thus, we conclude that deregulated p53 can efficiently facilitate the formation of BRCA2-mutant gastrointestinal cancer.
In agreement with previous reports6,43,44, our genome-wide knockout screen data revealed that full depletion of BRCA2 is critical for DNA-damaging agents to achieve the best therapeutic responses. For example, PARP inhibitors only kill the BRCA2 fully inactivated tumor cells17,18. Interestingly, we found that MMC was remarkably effective for eliminating BRCA2 monoallelic and biallelic mutant tumors by reducing BRCA2 mRNA and protein expression levels. The efficacy of MMC surpasses that of other DNA-damaging agents including cisplatin and olaparib. Thus, MMC offers a new therapeutic opportunity for patients harboring BRCA2 monoallelic and biallelic mutations when they do not respond to PARP inhibitors.
Materials and methods
Cell culture
The human gastric cancer cell lines were purchased from KCLB, RCB, and JCRB cell banks. RKO, HCT116 cell lines were purchased from the cell bank of Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences. HCT-15 was kindly provided by Dr. Zehong Miao, Shanghai Institute of Material Medica, Chinese Academy of Sciences. Cells were cultured in RPMI 1640, DMEM/F-12 or MEM medium with 10% fetal bovine serum and penicillin and streptomycin at 37 °C with 5% CO2. All the cell lines were recently authenticated by STR, keeping mycoplasma free.
Mice genotyping and spontaneous tumor models
All the animal experiments were performed according to the procedures approved by the Institutional Animal Care and Use Committee, Shanghai Institute of Nutrition and Health. The Brca2fl/fl mouse (01XB9, STOCK Brca2tm1Brn/Nci) and Trp53fl/fl mouse (01XC2, FVB.129P2-Trp53tm1Brn/Nci) were the kind gifts from Drs. Jos Jonkers and Anton Berns with signed material transfer agreements from NIH12. Villin-Cre mouse (T002376, B6/JNju-Tg(Pvillin-cre)I/Nju) was purchased from NRCMM (Nanjing, China). BALB/c mice were purchased from Shanghai SLAC Laboratory Animal Co. Ltd. To obtain Brca2 conditional knockout BALB/c mice in gastrointestinal epithelial cells, Brca2fl/fl mouse was first crossed with Villin-Cre mouse, and then the offspring were backcrossed onto BALB/c background for at least six generations before the studies. To get double knockout mice of Brca2 and Trp53, the Villin-Cre+; Brca2fl/fl mouse was crossed with Trp53fl/fl mouse and the offspring were backcrossed onto C57BL/6 background for at least three generations.
Mice were marked and genotyped about 10 days after born. Briefly, tail tip was boiled for 20 min in 200 µL of 0.05 M NaOH, then neutralized with 20 µL Tris-EDTA buffer (Tris 1 M, pH 8, EDTA 10 mM), serving as PCR template. The genotype of mouse was validated by the size of PCR products. For the loxP site in intron 10 of Brca2fl/fl allele, the PCR products were 376 bp for the floxed allele and 298 bp for the wild-type allele, respectively. For the loxP site in intron 11 of Brca2fl/fl allele, the PCR products were 529 bp for the floxed allele and 450 bp for the wild-type allele, respectively. Detection of the Brca2Δ11 allele yielded a 324-bp fragment. For the loxP site in intron 1 of Trp53fl/fl allele, the PCR products were 370 bp for the floxed allele and 288 bp for the wild-type allele, respectively. For the loxP site in intron 10 of Tp53fl/fl allele, the PCR products were 584 bp for the floxed allele and 431 bp for the wild-type allele, respectively. Detection of the of the Trp53Δ2–10 allele yielded a 612-bp fragment. Detection of the Cre yielded a 350-bp fragment. Primers for genotyping were listed in Supplementary Table 3.
For Brca2 knockout experiment, mice were separated into four groups with indicated numbers after genotyped, termed as Villin-Cre−; Brca2fl/fl mice without MNU, Villin-Cre+; Brca2fl/fl without MNU, Villin-Cre−; Brca2fl/fl with MNU and Villin-Cre+; Brca2fl/fl with MNU. MNU was added in drinking water at 120 mg/L on alternate weeks for a total exposure of 10 weeks. The drinking water was freshly prepared every three days in light-shielded bottles. For Brca2 and Trp53 double knockout experiment, mice were divided into two groups with indicated numbers.
Genome-scale CRISPR-Cas9 knockout screening
Lentivirus containing human GeCKOv2 library were produced by HEK293T and were precipitated by PEG buffer with a final concentration of 5% PEG8000 and 0.15 M NaCl. 2 × 108 SNU-1 cells were spin-infected at 37 °C with virus containing library sgRNAs at a MOI of 0.3. After spininfection, cells were selected with 2 µg/ml puromycin for another 7 days. After puromycin selection, cells were divided into three groups: 2 × 107 cells without further treatment were harvested at −80 °C termed as DMSODay0 group. The remaining cells were divided into another two groups, 2 × 107 cells receiving 0.1% DMSO for another 14 days termed as DMSODay14 group, and 2 × 108 cells treated with 0.3 µM MMC for another 14 days termed as MMCDay14 group. After experiment stopped, cells of all the three groups were harvested for extracting genomic DNA. After two rounds PCR amplification, PCR products were purified and sequenced by Illumina 2500 by CloudHealth (Shanghai, China). The sequencing data were analyzed by both MAGeCK and gene score analysis57. CRISPR gene score (CS) = average [log2 (MMCDay14 sgRNA abundance/ DMSODay0 sgRNA abundance)] or average [log2 (DMSODay14 sgRNA abundance/ DMSODay0 sgRNA abundance)]. PCR primers were shown in Supplemental Table 7.
Statistical analysis
All statistical analyses were done in GraphPad Prism 7 software. Data were represented as Means ± SD with at least three replicates. Log-rank (Mantel-Cox) test was used for calculating p values of survival curves, One-way ANOVA using Tukey’s comparison test to compare multiple groups. Two-way ANOVA with Sidak’s multiple comparisons test or Dunnett’s multiple comparisons test were used where appropriate. Two-sided Fisher’s exact test was used to determine the association between BRCA2 mutation and MSI or TP53 mutation.
Data availability
All the datasets are available from the corresponding author upon request. RNA sequencing data and GeCKO screening data have been deposited into NCBI database under accession number GSE132327 and GSE132431.
Code availability
All the codes are publically available according to the references.
Change history
01 October 2020
The original version of this Article was updated shortly after publication, because some of the Supplementary Information files were inadvertently omitted. The error has now been fixed and the Supplementry Information PDF is available to download from the HTML version of the Article.
References
Ceccaldi, R., Rondinelli, B. & D’Andrea, A. D. Repair pathway choices and consequences at the double-strand break. Trends Cell Biol. 26, 52–64 (2016).
Wong, A. K., Pero, R., Ormonde, P. A., Tavtigian, S. V. & Bartel, P. L. RAD51 interacts with the evolutionarily conserved BRC motifs in the human breast cancer susceptibility gene brca2. J. Biol. Chem. 272, 31941–31944 (1997).
Davies, A. A. et al. Role of BRCA2 in control of the RAD51 recombination and DNA repair protein. Mol. Cell 7, 273–282 (2001).
Patel, K. J. et al. Involvement of Brca2 in DNA repair. Mol. Cell 1, 347–357 (1998).
Roy, R., Chun, J. & Powell, S. N. BRCA1 and BRCA2: different roles in a common pathway of genome protection. Nat. Rev. Cancer 12, 68–78 (2011).
Yu, V. P. et al. Gross chromosomal rearrangements and genetic exchange between nonhomologous chromosomes following BRCA2 inactivation. Genes Dev. 14, 1400–1406 (2000).
Venkitaraman, A. R. Linking the cellular functions of BRCA genes to cancer pathogenesis and treatment. Annu. Rev. Pathol. 4, 461–487 (2009).
Antoniou, A. et al. Average risks of breast and ovarian cancer associated with BRCA1 or BRCA2 mutations detected in case Series unselected for family history: a combined analysis of 22 studies. Am. J. Hum. Genet. 72, 1117–1130 (2003).
Venkitaraman, A. R. Connecting Fanconi’s anaemia to breast cancer predisposition. Lancet 360, 1344–1345 (2002).
Tung, N. et al. Frequency of germline mutations in 25 cancer susceptibility genes in a sequential series of patients with breast cancer. J. Clin. Oncol. 34, 1460–1468 (2016).
Pennington, K. P. et al. Germline and somatic mutations in homologous recombination genes predict platinum response and survival in ovarian, fallopian tube, and peritoneal carcinomas. Clin. Cancer Res. 20, 764–775 (2014).
Jonkers, J. et al. Synergistic tumor suppressor activity of BRCA2 and p53 in a conditional mouse model for breast cancer. Nat. Genet. 29, 418–425 (2001).
Skoulidis, F. et al. Germline Brca2 heterozygosity promotes Kras(G12D) -driven carcinogenesis in a murine model of familial pancreatic cancer. Cancer Cell 18, 499–509 (2010).
Rowley, M. et al. Inactivation of Brca2 promotes Trp53-associated but inhibits KrasG12D-dependent pancreatic cancer development in mice. Gastroenterology 140(1303-1313), e1301–e1303 (2011).
Ludwig, T., Chapman, D. L., Papaioannou, V. E. & Efstratiadis, A. Targeted mutations of breast cancer susceptibility gene homologs in mice: lethal phenotypes of Brca1, Brca2, Brca1/Brca2, Brca1/p53, and Brca2/p53 nullizygous embryos. Genes Dev. 11, 1226–1241 (1997).
Kinzler, K. W. & Vogelstein, B. Cancer-susceptibility genes. Gatekeepers and caretakers. Nature 386(761), 763 (1997).
Bryant, H. E. et al. Specific killing of BRCA2-deficient tumours with inhibitors of poly(ADP-ribose) polymerase. Nature 434, 913–917 (2005).
Farmer, H. et al. Targeting the DNA repair defect in BRCA mutant cells as a therapeutic strategy. Nature 434, 917–921 (2005).
Tutt, A. et al. Oral poly(ADP-ribose) polymerase inhibitor olaparib in patients with BRCA1 or BRCA2 mutations and advanced breast cancer: a proof-of-concept trial. Lancet 376, 235–244 (2010).
Audeh, M. W. et al. Oral poly(ADP-ribose) polymerase inhibitor olaparib in patients with BRCA1 or BRCA2 mutations and recurrent ovarian cancer: a proof-of-concept trial. Lancet 376, 245–251 (2010).
Fong, P. C. et al. Inhibition of poly(ADP-ribose) polymerase in tumors from BRCA mutation carriers. N. Engl. J. Med. 361, 123–134 (2009).
Edwards, S. L. et al. Resistance to therapy caused by intragenic deletion in BRCA2. Nature 451, 1111–1115 (2008).
Lord, C. J. & Ashworth, A. Mechanisms of resistance to therapies targeting BRCA-mutant cancers. Nat. Med. 19, 1381–1388 (2013).
Sakai, W. et al. Secondary mutations as a mechanism of cisplatin resistance in BRCA2-mutated cancers. Nature 451, 1116–1120 (2008).
Bray, F. et al. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 68, 394–424 (2018).
Allemani, C. et al. Global surveillance of trends in cancer survival 2000-14 (CONCORD-3): analysis of individual records for 37 513 025 patients diagnosed with one of 18 cancers from 322 population-based registries in 71 countries. Lancet 391, 1023–1075 (2018).
Wadhwa, R. et al. Gastric cancer-molecular and clinical dimensions. Nat. Rev. Clin. Oncol. 10, 643–655 (2013).
The Cancer Genome Atlas Research Network. Comprehensive molecular characterization of gastric adenocarcinoma. Nature 513, 202–209 (2014).
Mortensen, D. L. et al. Effect of antigen binding affinity and effector function on the pharmacokinetics and pharmacodynamics of anti-IgE monoclonal antibodies. MAbs 4, 724–731 (2012).
Giannakis, M. et al. Genomic correlates of immune-cell infiltrates in colorectal carcinoma. Cell Rep. 15, 857–865 (2016).
Cerami, E. et al. The cBio cancer genomics portal: an open platform for exploring multidimensional cancer genomics data. Cancer Discov. 2, 401–404 (2012).
Gao, J. et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal 6, l1 (2013).
D’Andrea, A. D. & Grompe, M. The Fanconi anaemia/BRCA pathway. Nat. Rev. Cancer 3, 23–34 (2003).
Ceccaldi, R., Sarangi, P. & D’Andrea, A. D. The Fanconi anaemia pathway: new players and new functions. Nat. Rev. Mol. Cell Biol. 17, 337–349 (2016).
Howlett, N. G. et al. Biallelic inactivation of BRCA2 in Fanconi anemia. Science 297, 606–609 (2002).
Mori, Y. et al. Instabilotyping reveals unique mutational spectra in microsatellite-unstable gastric cancers. Cancer Res. 62, 3641–3645 (2002).
el Marjou, F. et al. Tissue-specific and inducible Cre-mediated recombination in the gut epithelium. Genesis 39, 186–193 (2004).
Suerbaum, S. & Josenhans, C. Helicobacter pylori evolution and phenotypic diversification in a changing host. Nat. Rev. Microbiol. 5, 441–452 (2007).
Joossens, J. V. et al. Dietary salt, nitrate and stomach cancer mortality in 24 countries. European Cancer Prevention (ECP) and the INTERSALT Cooperative Research Group. Int. J. Epidemiol. 25, 494–504 (1996).
Tatematsu, M. et al. Induction of adenocarcinomas in the glandular stomach of BALB/c mice treated with N-methyl-N-nitrosourea. Jpn J. Cancer Res. 83, 915–918 (1992).
Efeyan, A. & Serrano, M. p53: guardian of the genome and policeman of the oncogenes. Cell Cycle 6, 1006–1010 (2007).
Whicker, M. E., Lin, Z. P., Hanna, R., Sartorelli, A. C. & Ratner, E. S. MK-2206 sensitizes BRCA-deficient epithelial ovarian adenocarcinoma to cisplatin and olaparib. BMC Cancer 16, 550 (2016).
Osher, D. J., Kushner, Y. B., Arseneau, J. & Foulkes, W. D. Melphalan as a treatment for BRCA-related ovarian carcinoma: can you teach an old drug new tricks? J. Clin. Pathol. 64, 924–926 (2011).
Evers, B. et al. A high-throughput pharmaceutical screen identifies compounds with specific toxicity against BRCA2-deficient tumors. Clin. Cancer Res. 16, 99–108 (2010).
Barretina, J. et al. The Cancer Cell Line Encyclopedia enables predictive modelling of anticancer drug sensitivity. Nature 483, 603–607 (2012).
Tacconi, E. M. et al. Chlorambucil targets BRCA1/2-deficient tumours and counteracts PARP inhibitor resistance. EMBO Mol. Med 11, e9982 (2019).
Yuan, J., Adamski, R. & Chen, J. Focus on histone variant H2AX: to be or not to be. FEBS Lett. 584, 3717–3724 (2010).
Hata, T. et al. Mitomycin, a new antibiotic from Streptomyces. I. J. Antibiot. (Tokyo) 9, 141–146 (1956).
Tomasz, M. Mitomycin C: small, fast and deadly (but very selective). Chem. Biol. 2, 575–579 (1995).
Tomasz, M. et al. Isolation and structure of a covalent cross-link adduct between mitomycin C and DNA. Science 235, 1204–1208 (1987).
Bertoli, C., Skotheim, J. M. & de Bruin, R. A. Control of cell cycle transcription during G1 and S phases. Nat. Rev. Mol. Cell Biol. 14, 518–528 (2013).
Wu, K., Jiang, S. W. & Couch, F. J. p53 mediates repression of the BRCA2 promoter and down-regulation of BRCA2 mRNA and protein levels in response to DNA damage. J. Biol. Chem. 278, 15652–15660 (2003).
Andres, J. L. et al. Regulation of BRCA1 and BRCA2 expression in human breast cancer cells by DNA-damaging agents. Oncogene 16, 2229–2241 (1998).
Tan, S. L. W. et al. A class of environmental and endogenous toxins induces BRCA2 haploinsufficiency and genome instability. Cell 169, 1105–1118 (2017). e1115.
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Liu, T. et al. MYC predetermines the sensitivity of gastrointestinal cancer to antifolate drugs through regulating TYMS transcription. EBioMedicine 48, 289–300 (2019).
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014).
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005).
Villunger, A. et al. p53- and drug-induced apoptotic responses mediated by BH3-only proteins puma and noxa. Science 302, 1036–1038 (2003).
Breast Cancer Linkage Consortium. Cancer risks in BRCA2 mutation carriers. J. Natl Cancer Inst. 91, 1310–1316 (1999).
Teng, L. S., Zheng, Y. & Wang, H. H. BRCA1/2 associated hereditary breast cancer. J. Zhejiang Univ. Sci. B 9, 85–89 (2008).
Risch, H. A. et al. Population BRCA1 and BRCA2 mutation frequencies and cancer penetrances: a kin-cohort study in Ontario, Canada. J. Natl Cancer Inst. 98, 1694–1706 (2006).
Iqbal, J. et al. The incidence of pancreatic cancer in BRCA1 and BRCA2 mutation carriers. Br. J. Cancer 107, 2005–2009 (2012).
Parikh, A. R. et al. Analysis of DNA damage response gene alterations and tumor mutational burden across 17,486 tubular gastrointestinal carcinomas: implications for therapy. Oncologist 24, 1340–1347 (2019).
Chen, X., Ko, L. J., Jayaraman, L. & Prives, C. p53 levels, functional domains, and DNA damage determine the extent of the apoptotic response of tumor cells. Genes Dev. 10, 2438–2451 (1996).
Hay, T., Patrick, T., Winton, D., Sansom, O. J. & Clarke, A. R. Brca2 deficiency in the murine small intestine sensitizes to p53-dependent apoptosis and leads to the spontaneous deletion of stem cells. Oncogene 24, 3842–3846 (2005).
Acknowledgements
We thank the excellent service of institutional core facilities including Molecular Biology and Biochemistry, Cell Analysis, pathological analysis of biological samples, and laboratory animal technical platforms. We thank Dr. Ju Qiu for providing the immune-deficient mouse, and also thank Mr. Hang Dong and Xinlong Luo for their help with data analysis. This work was financially supported by grants from National Natural Science Foundation of China (81872890, 81322034, and 81372346), the Chinese Academy of Sciences (QYZDB-SSW-SMC034, XDA12020210), National Key R&D Program of China (2016YFC1302400).
Author information
Authors and Affiliations
Contributions
J.Y.L. conceived and supervised the study, wrote the manuscript. X.M.C. performed most of the experiments with data analysis, as well as conceived the study and wrote the manuscript. F.F.P. helped with the immunofluorescence staining, mouse genotyping and established some knockout cell lines. Y.J. and C.X.W. performed the bioinformatic analysis. T.T.L., H.W. helped with mouse genotyping. Y.M.H. and X.W. help with the in vivo tumor xenograft study. Y.F.Z. provided technical assistance. H.G.X. and X.Y.K. helped in design of the project and extensive discussions.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by A. Peschiaroli
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, X., Peng, F., Ji, Y. et al. Brca2 deficiency drives gastrointestinal tumor formation and is selectively inhibited by mitomycin C. Cell Death Dis 11, 812 (2020). https://doi.org/10.1038/s41419-020-03013-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-020-03013-8