Abstract
Glioblastoma multiform (GBM) is the most common brain tumor characterized by a dismal prognosis. GBM cancer stem cells (gCSC) or tumor-initiating cells are the cell population within the tumor-driving therapy resistance and recurrence. While temozolomide (TMZ), an alkylating agent, constitutes the first-line chemotherapeutic significantly improving survival in GBM patients, resistance against this compound commonly leads to GBM recurrence and treatment failure. Although the roles of protein-coding transcripts, proteins and microRNA in gCSC, and therapy resistance have been comprehensively investigated, very little is known about the role of long noncoding RNAs (lncRNAs) in this context. Using nonoverlapping, independent RNA sequencing and gene expression profiling datasets, we reveal that TP73-AS1 constitutes a clinically relevant lncRNA in GBM. Specifically, we demonstrate significant overexpression of TP73-AS1 in primary GBM samples, which is particularly increased in the gCSC. More importantly, we demonstrate that TP73-AS1 comprises a prognostic biomarker in glioma and in GBM with high expression identifying patients with particularly poor prognosis. Using CRISPRi to downregulate our candidate lncRNA in gCSC, we demonstrate that TP73-AS1 promotes TMZ resistance in gCSC and is linked to regulation of the expression of metabolism- related genes and ALDH1A1, a protein known to be expressed in cancer stem cell markers and protects gCSC from TMZ treatment. Taken together, our results reveal that high TP73-AS1 predicts poor prognosis in primary GBM cohorts and that this lncRNA promotes tumor aggressiveness and TMZ resistance in gCSC.
Similar content being viewed by others
Introduction
Glioblastoma multiform (GBM) is the most common primary tumor of the central nervous system (CNS) with a dismal outcome and a 5-year overall survival rate of <10%1. Despite multimodal therapeutic strategies encompassing surgical resection, radiation, and temozolomide (TMZ)-based chemotherapy2, GBM constitutes a major clinical challenge. This is due to its tendency to the infiltrative growth pattern and therapy resistance, both resulting in high recurrence rates, and eventually, therapeutic failure. A major advancement in deciphering GBM biology was the identification of glioblastoma multiform cancer stem cells (gCSC)3,4,5. These cells were shown to drive self-renewal, invasive GBM growth, and therapy resistance6,7. Therefore, numerous studies have focused on characterizing and targeting gCSC6,8.
To improve cure rates for GBM patients, a better understanding of the genetic and transcriptional events promoting tumor cell growth, survival, and drug resistance is urgently required9. While significant progress has been made in delineating the functions of protein-coding genes and microRNA in GBM biology, the functions of long noncoding RNAs (lncRNAs) in this disease are beginning to be elucidated. In one such study, a clinically relevant lncRNA, namely HIF1A-AS2, was found to promote stemness and tumorigenicity in mesenchymal gCSC10. However, the role and relevance of the vast majority of lncRNAs in GBM are currently unknown.
LncRNAs are RNA transcripts longer than 200 base pairs (bp) that do not code for proteins11. They are transcribed from genes with functional, well-conserved promoters12,13 that are bound by various transcription factors and that exhibit histone methylation patterns similar to those found in protein-coding genes14. The human genome harbors ~21,000 protein-coding genes and more than 50,000 lncRNA genes15, which are very poorly characterized to date. Nevertheless, in the past few years, lncRNAs have been found to play important roles in all aspects of cell biology, including stemness, immunity, development, regulation of gene expression, regulation of protein synthesis, and in various diseases16,17,18,19.
Recent studies suggest that the lncRNA TP73-AS1 may play a pivotal role in brain cancer biology. Specifically, DNA methylation of the TP73-AS1 promoter was reported to confer epigenetic downregulation of its expression in oligodendroglial tumors compared with the normal brain20. As part of a lncRNA-based signature, the expression of TP73-AS1 has been correlated with poor patient outcome in GBM21. Other studies suggested an association of high TP73-AS1 expression with low-grade glioma histology [25], while its forced overexpression resulted in reduced proliferation, as well as induction of apoptosis in conventional GBM cell lines [26]. Finally, hypermethylation and low expression of TP73-AS1 were found in GBM samples belonging to the less aggressive IDH and G-CIMP+ GBM subgroup22. Nevertheless, the clinical relevance or biological functions of TP73-AS1 in GBM, and in particular, in gCSC are currently unknown.
Here, we show that the lncRNA TP73-AS1 is clinically relevant in GBM, as high expression is associated with poor patient outcome in three independent, nonoverlapping primary GBM patient cohorts. Furthermore, TP73-AS1 downregulation leads to loss of ALDH1A1 expression and re-sensitizes gCSC to TMZ treatment. Together, our study underscores the importance of lncRNA-driven tumor biology in GBM and brings forth TP73-AS1 as a promising prognostic biomarker and a therapeutic target in this fatal disease.
Results
TP73-AS1 is a GBM-associated lncRNA
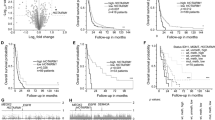
To assess whether TP73-AS1 is clinically relevant in GBM, we used GEPIA (http://gepia.cancer-pku.cn/index.html) where GBM expression data, obtained from the TCGA, are compared with normal brain tissue data, obtained from GTEx, in a standardized manner23. TP73-AS1 expression is significantly higher in primary GBM vs. normal brain tissue; however, it is lower in low-grade glioma (LGG) compared with normal tissue (Fig. 1a). Using R2, we analyzed the annotated FRENCH GBM cohort and found that the expression of TP73-AS1 is associated with the more aggressive gliomas as its expression is lower in tumors carrying an IDH1 mutation, as compared with tumors with wild-type (wt) IDH1 (Fig. 1b) and is higher in EGFR-amplified glioma tumors (Fig. 1b), both of which are more aggressive gliomas.
a TP73-AS1 expression in normal, LGG, and GBM tumor tissue. Normal tissue data were obtained from GTEx and tumor data from TCGA. Data were analyzed using GEPIA (gepia.cancer-pku.cn)23. b TP73-AS1 expression in tumor samples with or without IDH1 R132 mutation. Data were analyzed using R265. c Kaplan–Meier plots of patient outcome based on TP73-AS1 expression. Plots were generated using http://www.betastasis.com/ and R265. d Kaplan–Meier plots of patient outcome based on TP73-AS1 expression in GBM. Plots were generated using http://www.betastasis.com/. e Kaplan–Meier plots of patient outcome based on TP73-AS1 expression in glioma, stratified according to the annotated IDH mutation status and glioma subtype. Data, statistical evaluation, and visualization were obtained using the R2 website “R2: Genomics Analysis and Visualization Platform” (http://r2.amc.nl) and http://www.betastasis.com/
Importantly, we demonstrate that high expression levels of TP73-AS1 are associated with poor prognosis in three nonoverlapping, independent GBM cohorts (Fig. 1c). By analyzing TCGA survival and expression data, we found that high TP73-AS1 expression is correlated with poor patient outcome in the “mesenchymal” and “proneural” subtypes but did not correlate with survival in the “classical” or “neural” subtypes (Fig. 1d). In glioma, high expression of TP73-AS1 is significantly correlated with poor patient outcome in all tested cases, namely, IDH wt and mutant tumors, astrocytoma, and oligodendroglioma (Fig. 1e).
Collectively, these data suggest that TP73-AS1 comprises a clinically relevant lncRNA, potentially contributing to the aggressive tumor biology of GBM.
TP73-AS1 is linked to the stemness of gCSC
A key aspect of transcriptional regulation of stemness factors occurs at the epigenetic level. In particular, Suvà et al. compared the epigenetic landscapes, as determined by histone K27 acetylation (K27ac) between gCSC, differentiated gCSC (dCSC), and differentiated CSC that were reprogrammed back to gCSC using a set of their four reprogramming factors (iCSC)24. Our analysis of their data revealed that K27ac patterns within TP73-AS1 are similar to those of critical stemness factors in gCSC (Fig. 2a). Indeed, in gCSC, the TP73-AS1 promoter is strongly K27ac-positive, indicating active transcription, while almost no K27ac was observed in differentiated gCSC. Upon reprogramming of dCSC (iCSC), the amount of K27ac within the TP73-AS1 promoter increased again (Fig. 2a). These data indicate that the K27ac status of our candidate’s promoter is tightly linked to GBM cell “stemness” state, suggesting that TP73-AS1 may constitute an important factor in gCSC.
a Histone K27ac of the TP73-AS1 locus in gCSC (CSC), differentiated gCSC (dCSC), and differentiated gCSC that had been reprogrammed to gCSC (iCSC). The putative promoter region of TP73-AS1 is highlighted in gray. Data were obtained from graphs generated using http://www.broadinstitute.org/epigenomics/dataportal/clonePortals/Suva_Cell_2014.html. b The expression level of the TP73-AS1 in normal brain tissue versus neuronal stem cells and gCSC. Data, statistical evaluation, and visualization were obtained using the R2 website “R2: Genomics Analysis and Visualization Platform” (http://r2.amc.nl). c Efficient knockdown of the lncRNA TP73-AS1 in GBM tumor cells using CRISPRi. gCSC cells expressing a doxycycline-inducible dCAS9-KREB fusion protein and the indicated gRNAs targeting TP73-AS1 promoter region were generated. The indicated GBM tumor cell lines were engineered to express doxycyline-inducible dCAS9-KREB and gRNA targeting the TP73-AS1 promoter or scramble controls. Cells were induced with doxycycline for 10 days, after which the levels of TP73-AS1 were measured using qRT-PCR. *p < 0.05; n = 3. dCAS9-KREB expression was induced after 10 days, and subsequently, the percentage of cell death induction was determined using Annexin V and PI staining and was measured by flow cytometry. Representative images are shown. The graph represents average ± SD; n = 3
We interrogated published data established with patient-derived gCSC growing under adherent conditions4,25,26,27. We found that TP73-AS1 is highly expressed in gCSC and neuronal stem cells, as compared with the normal cortex (Fig. 2b). Due to the high expression of TP73-AS1 in gCSC, we subsequently used a gene silencing approach in these cells to study the functions of TP73-AS1.
In order to examine the role of TP73-AS1 in gCSC, we used two established gCSCs, G26 and G74,25. As the biological function of some lncRNAs is tightly linked to their transcription28,29,30, we performed CRISPR inhibition (CRISPRi) as a gene knockdown (kd) approach31,32. In CRISPRi, a guide RNA (gRNA) targeting the region close to the transcriptional start site of a specific gene is expressed in a cell. The gRNA binds to a DNAse dead mutant CAS9 (dCAS9) fused to the transcriptional inhibitor domain KREB and the complex silences transcription of the targeted gene. We used a lentivirus approach to generate a stable gCSC, G26, and G7, expressing a doxycycline-inducible dCAS9-KREB33 in combination with either one of the two gRNAs targeting TP73-AS1 (g#1 and g#2) or scramble controls (gSCR). Indeed, following doxycycline induction, qRT-PCR analysis showed significant reduction in TP73-AS1 expression in the gCSC models expressing either one of the two gRNAs as compared with controls (Fig. 2c).
To determine whether TP73-AS1 expression levels alter the stemness of gCSC, we assess the level of cellular differentiation upon TP73-AS1 depletion in gCSC. We depleted TP73-AS1 in gCSC for 10 days, and subsequently, we stained these cells for the established stemness (NESTIN) or differentiation markers (GFAP and TUBII)4 (Fig. 3a). Our results showed that TP73-AS1 depletion did not result in consistent changes in the expression of the tested markers in the two tested gCSC models (Fig. 3a). To functionally test the self-renewal, an important stemness trait of TP73-AS1-depleted gCSC, we performed a limiting dilution assay (LDA), which measures the ability of low cell numbers to form a sphere34. To this end, we induced kd of TP73-AS1 in gCSC for 10 days, plated cells in different dilutions in 96-well plates, and monitored sphere formation after 21 days (Fig. 3b). We found that TP73-AS1 kd reduced the ability to form a sphere in G7 cells as compared with control cells (Fig. 3b). Nevertheless, this was not the case in G26 gCSC (Fig. 3b). These data suggest that TP73-AS1 may act as a stemness factor in some gCSCs but not in all cases.
a gCSC cells expressing a doxycycline-inducible dCAS9-KREB fusion protein and the indicated gRNAs targeting TP73-AS1 promoter region were generated. dCAS9-KREB expression was induced for 10 days, after which the expression of the indicated stemness and differentiation markers was determined using immunofluorescence. Representative images are shown. Bar = 100 μM. b LDA assay measures the self-renewal capacity. dCAS9-KREB expression was induced for 10 days, after which the cells were plated in 96-well plates in the concentration of (1, 5, 10, and 20 cells per well). Spheres were counted manually 21 days after the induction. The estimate depicts the confidence interval for one per stem cell frequency. Data were analyzed using http://bioinf.wehi.edu.au/software/elda/
Co-expression analysis reveals that TP73-AS1 expression affects TP73 expression inconsistently in gCSCs
The lncRNA TP73-AS1 has an ~200-bp 3′ overlap with the 3′ untranslated region of an adjacent gene, TP73 (p73) (Fig. 2a). As a member of the p53 family of transcription factors35, p73 has been implicated in several biological categories, in particular in cancer36, brain development37, and importantly, in the stemness of neuronal progenitor cells38,39 and invasiveness of GBM tumor cells40. Because many lncRNAs are known to directly regulate the expression of adjacent genes28,29,30, we investigated whether TP73-AS1 regulates the expression of TP73. First, we tested whether silencing of TP73-AS1 affected the expression of TP73 in our gCSC models. However, we were not able to detect consistent changes in TP73 expression. While in G7 cells, TP73-AS1 kd led to a reduction of TP73 expression, but the levels of TP73 in G26 cells decreased upon silencing of TP73-AS1 using g#1, while it increased in g#2 cells (SUP Fig. 1a).
Next, we asked whether the expression of TP73-AS1 is correlated with that of TP73 in expression data obtained from various (627) human cell lines and two primary GBM tumor datasets. We found that the expression of these two genes is not correlated (SUP 1b). We therefore conclude that it is unlikely that TP73-AS1 directly regulates the expression of its neighboring geneTP73.
TP73-AS1 promotes gCSC resistance to TMZ
Treatment with TMZ is the most common therapeutic approach used for GBM patients; therefore, it is critical to better understand the mechanisms underlying TMZ resistance in gCSC41. Because the expression of TP73-AS1 is negatively correlated with patient survival, we asked whether TP73-AS1 is involved in TMZ resistance in gCSC.
Following the induction of TP73-AS1 depletion for 10 days, we treated both gCSC models with TMZ for 7 days, which was sufficient to observe the morphological changes suggestive of increased cell death in the TP73-AS1 kd cell culture (Fig. 4a). Subsequently, we quantified cell death induction in the TMZ-treated gCSC using annexin V/PI staining and FACS, and we found a significant increase in cell death in the TP73-AS1-depleted cells, as compared with control cells 5 days post treatment (Fig. 4b). Importantly, there was no difference in cell death between control and TP73-AS1 kd cells under control conditions or 2 days post TMZ treatment (SUP Fig. 2a, b) indicating that TP73-AS1 kd synergizes with TMZ to kill gCSC.
a Representative images of TP73-AS1 depleted and control gCSC cells treated with TMZ (500 µM) for 7 days. Images were recorded using a light microscope. b TP73-AS1 depleted and control gCSC models were treated with TMZ (500 µM) for 7 days, after which the viability was determined using Annexin V/PI staining and flow cytometry. Representative images are shown. Bar = 50 μM; *p < 0.05; n = 3; average ± SD. c TP73-AS1 depleted and control gCSC were treated with the indicated dose of TMZ for 7 days, after which viability was measured using crystal violet staining and a plate reader. Values represent the relative average OD; n ≥ 3; *p < 0.05 Student’s t test
To confirm these findings, we measured the viability of G7 and G26 gCSC treated with an escalating dose of TMZ for 5 days and found that the kd of TP73-AS1 in both gCSCs led to an increased sensitivity to TMZ (Fig. 4c).
We investigated the impact of TP73-AS1 on gCSC proliferation using a BrdU assay. We found that there was no significant difference between TP73-AS1 kd and control cells under resting conditions, and that proliferation was inhibited in all cases 2 days following TMZ treatment (SUP Fig. 3a). In addition, we measured reactive oxygen species (ROS) in TP73-AS1 depleted and control gCSC cells, using DCFDA (chloromethyl-2′,7′-dichlorofluorescein diacetate) and FACS, and found that in G7 cells, there were no differences in ROS levels in all tested conditions. In contrast, TP73-AS1 depletion resulted in increased ROS levels following 2 days of TMZ treatment and in non-treated G26 cells (SUP Fig. 3b).
These data indicate that TP73-AS1 contributes to TMZ resistance in gCSC and that the protective functions of TP73-AS1 are not linked to regulation of proliferation. Furthermore, TP73-AS1 protective functions in gCSC may be due to curbing ROS levels in a cell-line-specific manner.
TP73-AS1 interacts with multiple metabolic pathways in TMZ-treated gCSC
O-6-Methylguanine-DNA methyltransferase (MGMT) is known to repair DNA damage caused by alkylating agents, including TMZ, and it is regulated in GBM at the level of promotor methylation42, and MGMT methylation status is linked to the response to TMZ in patients. We asked what is the status of the promotor of MGMT in our gCSC using published data43,44,45 and found that the promoter is methylated in both cell lines (SUP Fig. 3c), suggesting that the increased sensitivity of the TP73-AS1 kd cells is not likely to be related to MGMT methylation.
To learn more about the cellular pathways interacting with TP73-AS1, we compared the transcriptome of TP73-AS1 kd vs. control G7 and G26 gCSC upon TMZ treatment and control conditions using RNAseq. According to our principal component analysis (PCA), we found that all three biological replicates from each experimental condition grouped together indicate that the major differences between samples are due to the biological perturbations caused by the knockdown of TP73-AS1 or TMZ treatment (SUP Fig. 4a).
Next, we asked which of the transcripts are affected by TP73-AS1 kd in each cell line upon TMZ treatment or in control conditions (SUP Table 1). To this end, we clustered the genes, which are significantly (DeSeq2 with a cutoff of the adjusted p value < 0.05) up- and downregulated upon TP73-AS1 kd. We found that there were profound transcriptional changes affecting over 2000 genes that were significantly changed in each experimental condition. For validation, we chose four transcripts, measured their expression using qRT-PCR, and found that their expression pattern matched the pattern found in the RNAseq data (Sup Fig. 5 and Fig. 6a).
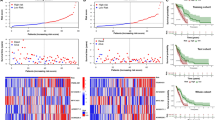
The TP73-AS1-dependent transcriptome perturbation indicates that TP73-AS1 comprises a major biological regulator in these cells (Fig. 5a). We therefore asked if there was a significant enrichment for particular biological pathways upon TP73-AS1 kd and found that there were hundreds of gene ontology (GO) pathways affected (Fig. 5b). In both G7 and G26 cells, TP73-AS1 kd led to an upregulation of genes enriched for GO categories related to neuronal development and function, in addition to other developmental pathways and cell–cell communication pathways (Fig. 5c and SUP Table 2).
a Heat map depicting transcripts whose expression is affected by TP73-AS1 kd. Red represents upregulation and blue represents downregulation. b Transcripts affected by TP73-AS1 kd in each cell line and conditions were analyzed to identify significantly affected GO categories. The number of GO categories identified in each cell line, condition, and direction of gene expression change (upregulation or downregulation by TP73-AS1 kd) are shown. c Adjusted p values (average ± SD) of GO categories related to the indicated biological functions are shown. Note that there were no common categories in the list of genes that were downregulated in the kd cells. See Table S1 for a complete list
Under TMZ treatment, genes belonging to neuronal differentiation and function were also upregulated in TP73-AS1 kd cells in both gCSC models (Fig. 5c and SUP Table 2). Interestingly, transcripts related to metabolism, mitochondria, and nucleotide metabolism in particular, were downregulated upon TP73-AS1 kd in both cell lines (Fig. 5c and Table 2). These findings may explain how TP73-AS1 kd leads to increased sensitivity to the alkylating agent TMZ in gCSC, as chemotherapy resistance is tightly linked to cellular metabolism46. Together, our transcriptomic analysis reveals that TP73-AS1 is a major regulator in gCSC by orchestrating multiple cellular pathways, including neuronal and metabolic pathways, providing protection against TMZ.
TP73-AS1 promotes the expression of ALDH1A1
One of the transcripts found to be downregulated in the two TP73-AS1 kd cell lines under TMZ treatment and control conditions was ALDH1A1 (Fig. 6a), which encodes the aldehyde dehydrogenase 1 family member A1 (ALDH1A1) protein. This protein family is known for their ability to do detoxification of endogenous and exogenous aldehyde substrates through NAD(P)+ oxidation47. Moreover, ALDH1A1 is a known marker of stem cells and cancer stem cells, and plays a crucial role in chemoresistance in several cancer types48,49 and specifically in GBM50.
a The indicated gCSC lines were treated with TMZ and the levels of ALDH1A1 were measured using qRT-PCR. The results matched the trends found in RNAseq experiments, as indicated by the RNAseq group each condition belongs to. b G7 cells were treated with TMZ (300 μM), DEAB (200 μM), or a combination of the two compounds for 5 days, after which cell death was measured using PI and FACS. Average values and SEM are shown. n = 3; *p < 0.05 ANOVA
To test if ALDH1A1 downregulation explains, at least in part, the protective functions of TP73-AS1 in gCSC, we treated gCSC with the TMZ (300 μM) in the presence or absence of the ALDH1 inhibitor, DEAB (200 μM), for 5 days, after which we measured cell death using PI and FACS (Fig. 6b). We found that DEAB increased the sensitivity of G7 cells to TMZ, confirming the protective functions of ALDH1 in gCSC and providing support to the model where TP73-AS1 enhances the resistance of gCSC to TMZ by promoting the expression of ALDH1A1.
Discussion
The biological role and the clinical relevance of TP73-AS1 in GBM are only emerging. Contradictory results were recently published, suggesting either a tumor promoting51,52 or suppressing53 function of TP73-AS1 in GBM. Specifically, a recent study demonstrated that overexpression of TP73-AS1 in U251 GBM tumor cells resulted in reduced proliferation and increased cell death53. Furthermore, the same study suggested that the expression of TP73-AS1 was higher in low-grade vs. high-grade gliomas. However, other studies showed that high expression of TP73-AS1 in peritumoral brain edema was correlated with poor patient outcome and that depletion of TP73-AS1 in GBM tumor cell lines results in reduced proliferation and migration52. To clarify the clinical relevance of the lncRNA TP73-AS1 in GBM, we interrogated three GBM patient cohorts and found that high TP73-AS1 expression was significantly correlated with poor patient outcome. This is in accordance with the finding that the expression of TP73-AS1 is high in GBM and low in LGG. Furthermore, TP73-AS1 is highly expressed in IDH1 wt vs. mutant tumors, the former being more aggressive than the latter.
To elucidate the biological functions of TP73-AS1, we utilized gCSC models, which are the most relevant in vitro models for studying GBM biology4. These gCSC models generate tumors upon orthotopic implantation into brains of immunocompromised mice, which most closely recapitulate the initial patient-derived tumor4,25. Using CRISPRi as a gene silencing approach and gCSC, we found that TP73-AS1 depletion does not affect neither proliferation nor gCSC survival. It may be that TP73-AS1 functions in a cell context-dependent manner and may promote or inhibit proliferation in standard GBM cell lines as opposed to gCSC.
Notably, the epigenetic profile of the TP73-AS1 promoter in differentiated, reprogrammed, and untreated gCSCs is similar to that found in the promoter of the established stemness factors24. Using a functional assay (LDA), we found a significant effect on self-renewal upon TP73-AS1 silencing in one of our tested gCSC, G7 but not in G26. These differences may be related to the type of tumor from which these cells were generated, the specific cell line biology, or compensatory mechanisms upon TP73-AS1 kd. Indeed, there are notable differences in key stemness factors expressed in these cells25. Nevertheless, we found that TP73-AS1 kd led to an upregulation of transcripts associated with neuronal differentiation in both models.
The current standard of care for GBM patients includes surgery, followed by radiation combined with chemotherapy54. Temozolomide is the major chemotherapeutic agent used for GBM patients. Hence, our finding that TP73-AS1 affected TMZ resistance in gCSC is of immediate clinical relevance. Specifically, we showed that TP73-AS1 silencing led to a significantly increased sensitivity to TMZ in two gCSC models. Despite an aggressive surgical approach and aggressive chemotherapy, the prognosis of GBM patients remains dismal54. Cancer stem cells are especially chemoresistant55,56 and are characterized by high expression of DNA repair proteins and efflux pumps57,58. Our data may explain the tight association existing between high TP73-AS1 expression and poor patient outcome in GBM and strongly suggest that TP73-AS1 exhibits a chemoprotective role in GBM tumors.
The mechanism by which TP73-AS1 protects gCSC from TMZ remains currently unknown. Because in some cases, lncRNAs function by regulating their gene neighbors28,29,30, we first tested whether TP73-AS1 may regulate the expression of its gene neighbor, TP73. We found that this is not the case, as there is no correlation in expression between the two genes in G26, in a collection of mammalian cell lines and in two independent GBM cohorts. However, it is possible that there is an indirect promotion of TP73 expression by TP73-AS1 in specific cellular contexts, as is the case in G7 cells. These data are consistent with the known functions of TP73 as a stemness factor in brain stem cells38,39 and the finding that in G7 gCSC, but not G26 gCSC, TP73-AS1 promotes self-renewal (Fig. 3b).
Using expression profiling, we found that TP73-AS1 kd and TMZ treatment resulted in reduced expression of transcripts related to several metabolic pathways, including mitochondria biology and nucleoside metabolism. While gCSCs can adapt to different nutrients found in their environment, they are highly dependent on the mitochondria for generating energy59. Furthermore, recent findings suggest that combining mitochondrial toxins with TMZ may be beneficial in eradicating gCSC60. Importantly, a recent CRISPR activation screen, aimed at identifying both protein-coding and lncRNA genes promoting drug resistance in acute myeloid leukemia tumor cells, found that lncRNA-protecting tumor cells from chemotherapy were often neighbors and regulators of metabolism-related genes, highlighting the importance of lncRNA regulation of the expression of metabolic genes in drug resistance46. It is therefore tempting to speculate that TP73-AS1 protects gCSC from TMZ by promoting mitochondrial and other metabolic pathways.
Our RNAseq data and validation showed that one of the transcripts that is downregulated under TMZ treatment and control conditions in the two TP73-AS1 kd gCSCs is ALDH1A1 (sup table 1 and sup Fig. 6), which encodes the aldehyde dehydrogenase 1 family member A1 (ALDH1A1) protein. ALDH1A1 is known to be a marker of cancer stem cells47,61 and importantly, is known to promote drug resistance in cancer62, cancer stem cells, and specifically in gCSC61,63. Using the ALDH1 inhibitor, DEAB, we confirmed that also in the case of the gCSC used in this study, ALDH1 inhibition leads to enhanced TMZ sensitivity, providing support to the premise that the increased TMZ sensitivity of TP73-AS1-depleted cells is linked to reduced ALDH1A1 expression. The mechanism of ALDH1A1-mediated therapy resistance is still not completely understood, but was proposed to be linked to reduction of oxidative stress caused by the chemotherapy62. We therefore propose that by promoting ALDH1A1 expression in gCSC, TP73-AS1 reduces their sensitivity to chemotherapy.
In conclusion, we uncovered that TP73-AS1 comprises a clinically relevant lncRNA conferring TMZ resistance in gCSC, and may therefore serve as an important predictive biomarker and a therapeutic target in a currently fatal disease.
Materials and methods
Cell culture
G26 and G7 were a kind gift from Steven Pollard. Cells were grown on plates coated with Laminin (Sigma-Aldrich, L2020), as detailed in ref. 4.
Immunofluorescence
Cells were cultured and differentiated in a 96-well plate covered with poly-L-ornithine hydrobromide (Sigma-Aldrich; P3655) and Laminin (10% in PBS). Cells were grown overnight, after which they were fixed with 4% paraformaldehyde (paraformaldehyde 16%, Alfa Aesar) in PBS for 30 min and permeabilized with permeabilization buffer (0.5% Triton X-100 (Sigma-Aldrich) in PBS). The cells were washed with washing solution (25% BSA (bovine serum albumin), Sigma-Aldrich) and 0.1% Tween-20 (Sigma-Aldrich, P9416; PBS). The cells were blocked for 1 h with 10% goat serum in wash buffer and subsequently in washing solution. The blocking buffer was removed and the cells were labeled with primary antibodies (anti-nestin 1:100; Abcam (10C2); anti-TUBIII 1:75 (Merck (MAB1637); anti-GFAP 1:100 (Santa Cruz (2E1)) diluted in blocking buffer and incubated overnight at 4 °C. The cells were incubated at room temperature (RT) for 30 min and then washed using washing solution. A secondary antibody (ENCO, Alexa 549) was added (1:200 in blocking solution) for 1 h at RT. The cells were washed with the washing solution and stained with DAPI for 1 min. The cells were washed three times with DDW. Images were taken using a fluorescent microscope.
Cell proliferation assessment with 5-bromo-2-deoxyuridine (BrdU)
The BrdU proliferation assay was performed using BrdU Cell Proliferation Assay Kit (Biovision Incorporated) following the manufacturer’s instructions. Cells were cultured in 96-well plates covered with poly-L-ornithine and laminin (10% in PBS) overnight, after which the cells were labeled with 5-bromo-2-deoxyuridine (BrdU) for 3 h and stained. Cells were visualized under a fluorescent microscope and the number of BrdU-positive cells and the total cell number were determined in each field.
RNA extraction, cDNA synthesis, and quantitative reverse transcription polymerase chain reaction (qRT-PCR)
The RNA from cultured cells was purified using the PureLink RNa Mini Kit (Thermo Fisher Scientific). From a starting amount of 100–200 ng of RNA, cDNA was generated using the cDNA synthesis kit (E6300S, BioLabs). Briefly, 20 µl of the cDNA synthesis reaction was subjected to the following conditions: 5 min at 25 °C, 60 min at 72 °C, and 5 min at 96 °C. cDNA is synthesized in vitro, from an mRNA template using the enzyme reverse transcriptase, resulting in single-stranded cDNA production. qPCR was carried out on the produced cDNA to allow for detection of mRNA expression levels in the cells of interest. The PCR conditions were as follows: initialization step at 95 °C for 10 min, 46 cycles of the denaturation step at 60 °C for 30 s, annealing step at 60 °C for 30 s, and elongation step at 72 °C for 1 s, followed by the final elongation at 40 °C for 10 min to ensure that any remaining single-stranded DNA is fully extended. Most of the forward and the reverse primers were designed to span different exons whenever possible. All primers were purchased from Integrated DNA Technologies (IDT). qRT-PCR was performed using an iCycler (Bio-Rad Laboratories) with a threshold cycle number determined with the use of iCycler software version. Reactions were performed in triplicate and threshold cycle numbers were averaged. The results were normalized to L32 and POL2.
RT-PCR primers used in this study
TP73-AS1: probe #29, primer1 ctccggacactgtgttttctc, primer2 tcttttaaggcggccatatc; P73: probe #60, primer1 cacgtttgagcacctctgg, primer2 cgcccaccacctcattatt; L32: probe #33, primer1 gcacactgactacagccttga, primer2 tacccaggtttggaggtgtg; ALDH1A1 #14: primer1 tttggtggattcaagatgtctg, primer2 cactgtgactgttttgacctctg; FNDC5 #8: primer1 taccaaaacaccccttctgg, primer2 tcttcctgtccgtggtgaat; ZNF536 #22: primer1 gagtcccagtcggtgagc, primer2 gctctcctcggtgacgttag; WNT5A #11: primer1 ttctggctccacttgttgct, primer2 gccaaagccactaggaagaac.
Plasmids
psPAX2 was a gift from Didier Trono (Addgene plasmid # 12260). pMD2.G was a gift from Didier Trono (Addgene plasmid # 12259). pHAGE TRE dCas9-KRAB was a gift from Rene Maehr and Scot Wolfe (Addgene plasmid # 50917). pLKO.1-puro U6 sgRNA BfuAI large stuffer was a gift from Scot Wolfe (Addgene plasmid # 52628). We used pLKO.1-puro U6 sgRNA BfuAI large stuffer to clone our gRNA sequences according to the instructions available on the addgene page. The sequences used were as follows:
scramble/control: ACCGCGCCAAACGTGCCCTGACGG; TP73-AS1 g#1: GCAGTCGGGGCTGACGGCGG; TP73-AS1 g#2: CCTAGATGGGAGCCGGGGAT.
Generation of stable cell lines for gene knockdown
Lentiviruses were generated using our standard protocol in 293T cells, as detailed in HACE1 reduces oxidative stress and mutant Huntingtin toxicity by promoting the NRF2 response64. Cells were grown in DMEM, 10% FBS, and transfected using CalFectin (Signagen) and plasmids at a ratio of 1:2:3 (PAX2; pMD2.G; transfer vector). Media was changed after 24 h and the viruses were collected after 48 h. The collected viruses were stored at −80 °C. G7 and G26 cells were grown to 80% confluence on 6-cm dishes, then infected with a vector ratio of 1:10 (cas9, gRNA), and left for 24 h in a 37 °C incubator. Following infection, the lentivirus-containing medium was removed and replaced by fresh medium and incubated at 37 °C with the selection antibiotic using puromycin (1 μg/ml) and G418 (1500 μg/ml).
Gene knockdown
The depletion of the TP73-AS1 was done using the CRISPRi method combined with viral infection. gRNA were designed to target the promoter region of the TP73-AS1 gene. Knockdown induction was done using doxycycline at a concentration of 2 µg/ml (Sigma-Aldrich, D3072). Gene knockdown was measured using qRT-PCR.
LDA-sphere formation assay
Cells are plated into a 96-well plate over a range of densities from high to low (number of cells/well = 1, 5, 10, and 20). The cells were allowed to grow for 21 days and form spheres. To determine the probability of sphere formation, the wells were scored for the presence or absence of sphere growth. The natural log fraction of the sphere-negative wells was plotted on a linear scale versus the density (cells/well). The frequency of sphere-forming cells in a particular cell type was determined using the ELDA web tool at http://bioinf.wehi.edu.au/software/elda.
TMZ treatment
The kd of TP73-AS1 was induced by addition of doxycycline for 10 days. The cells were plated in 12-well plates and TMZ (Sigma-Aldrich, T2577) or DMSO was added. Seven to ten days post treatment, the cells were washed and trichloroacetic acid (TCA) was added for an hour at 4 °C. The plates were washed and air dried. Crystal Violet (C0075, Sigma-Aldrich) was added for half an hour and additional rinsing was conducted. After dehydration, 10% acetic acid was added and the optical density was measured at 570 nm. The results were normalized to scramble gRNA.
FACS
Cell apoptosis was determined using flow cytometry. Cells were harvested 10 days post doxycycline treatment using Accutase (Biological Industries, Israel) and the cells were treated with TMZ for 7 days and harvested later. The cells were stained using an apoptosis detection kit (Miltenyi Biotec). The cells were resuspended in binding buffer, and were then stained with Annexin V FITC and propidium iodide (PI). The cells were analyzed using a flow cytometer (Sysmex) equipped with FCS Express software (De Novo Software) according to the manufacturer’s instructions. For the DEAB treatments, G7 cells were treated with 200 μM of DEAB (D86256, Sigma-Aldrich), an ALDH1A1 inhibitor, and with 300 μM of TMZ for 5 days, after which cell death was measured using PI staining and FACS.
ROS measurement
Cell quantitation of ROS was determined using flow cytometry. Cells were harvested 10 days post doxycycline treatment using Accutase (Biological Industries, Israel) and the cells were treated with TMZ for 2 days and harvested later. The cells were stained using a ROS detection kit (D6883, Sigma-Aldrich). The cells were treated with DCFDA for 1 h, and then harvested and washed with PBS. The cells were analyzed using a flow cytometer (Sysmex) equipped with FCS Express software (De Novo Software) according to the manufacturer’s instructions.
RNAseq
Libraries were prepared using QuantSeq 3′ mRNA-Seq Library Prep Kit FWD for Illumina (Lexogene) according to the manufacturer’s protocol. Samples were pooled and sequencing was performed using Nextseq5000 using all four lanes.
Thirty-three samples were sequenced in the present study. These are composed of three biological replicates of G7 cells expressing gRNA SCR (scrambled) and gRNA#1 in TMZ treated and untreated conditions and three biological replicates of G26 cells expressing gRNA SCR, gRNA#1, and gRNA#2 under TMZ treatment and non-treated conditions. Sequencing resulted in an average of 9,655,276 reads per sample with an average sequence length of 86 bp. Raw reads were then trimmed for sequencing adaptors, poly-A, as well as poor-quality bases using Trim galore (v0.4.2; https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/) and Cutadapt (v1.12.1; github.com/easybuilders/easybuild-easyconfigs/tree/master/easybuild/easyconfigs/c/cutadapt). After QC, an average of 9,561,457 reads per sample remained for downstream analysis with an average sequence length of 78 bp. Then mapping and gene-level read count estimation were performed using STAR (v2.5.3a) and RSEM (v1.2.31), respectively, against the human reference genome (GRCh38). The DeSeq2 R package was used to normalize gene counts and perform differential gene expression, followed by clustering and visualizations using the DeSeq2 VSD function (Variance Stabilizing Transformation). Clustering analysis was performed by employing a hierarchical clustering method (using Pearson correlation clustering and “ward.D2” agglomeration method), while the number of clusters were assessed using the “eclust” function from the “factoextra” R package. The “clusterprofiler” R package was used for Gene Ontology enrichment analysis, while gene annotation was retrieved from the “org.Hs.eg.db” R package. PCA was generated for the normalized count data using the plotPCA function from the DeSeq2 R package using the top 500 variable genes.
Cell viability measurements using crystal violet staining
Cells were grown in a medium containing antibiotic selection. KD was induced with doxycycline for 10 days, after which the cells were replated into 24-well plates in the presence or absence of TMZ (Sigma-Aldrich, T2577). One week post treatment, the cells were washed with trichloroacetic acid (TCA) for an hour at 4 °C. The plates were washed and air dried. Crystal Violet (C0075, Sigma-Aldrich) was added for half an hour and additional rinsing took place. Following dehydration, 10% acetic acid was added and the OD was measured at 590 nm. The results were normalized to scramble gRNA.
References
Reifenberger, G., Wirsching, H. G., Knobbe-Thomsen, C. B. & Weller, M. Advances in the molecular genetics of gliomas-implications for classification and therapy. Nat. Rev. Clin. Oncol. 14, 434–452 (2017).
Stupp, R. et al. Radiotherapy plus concomitant/nand adjuvant temozolomide for glioblastoma. N. Engl. J. Med. 352, 987–996 (2005).
Lathia, J. & Mack, S. Cancer stem cells in glioblastoma. Genes Dev. 29, 1203–1217 (2015).
Pollard, S. M. et al. Glioma stem cell lines expanded in adherent culture have tumor-specific phenotypes and are suitable for chemical and genetic screens. Cell Stem Cell 4, 568–580 (2009).
Stricker, S. H. et al. Widespread resetting of DNA methylation in glioblastoma-initiating cells suppresses malignant cellular behavior in a lineage-dependent manner. Genes Dev. 27, 654–669 (2013).
Chen, J. et al. A restricted cell population propagates glioblastoma growth after chemotherapy. Nature 488, 522–526 (2012).
Gabriely, G., Wheeler, M. A., Takenaka, M. C. & Quintana, F. J. Role of AHR and HIF-1α in glioblastoma metabolism. Trends Endocrinol. Metab. 28, 428–436 (2017).
Piccirillo, S. G. M. et al. Bone morphogenetic proteins inhibit the tumorigenic potential of human brain tumour-initiating cells. Nature 444, 761–765 (2006).
Rotblat. B. & Grunewald, T. G. P. Translating cancer-omics into function. Biol Cell 107, 1–2 (2015).
Mineo, M. et al. The long non-coding RNA HIF1A-AS2 facilitates the maintenance of mesenchymal glioblastoma stem-like cells in hypoxic niches. Cell Rep. 15, 2500–2509 (2016).
Sigova, A. A. et al. Divergent transcription of long noncoding RNA/mRNA gene pairs in embryonic stem cells. Proc. Natl Acad. Sci. USA 110, 2876–2881 (2012).
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009).
Necsulea, A. et al. The evolution of lncRNA repertoires and expression patterns in tetrapods. Nature 505, 635–640 (2014).
Kapranov, P. et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 316, 1484–1488 (2007).
Iyer, M. K. et al. The landscape of long noncoding RNAs in the human transcriptome. Nat. Genet. 47, 199–208 (2015).
Fatica, A. & Bozzoni, I. Long non-coding RNAs: new players in cell differentiation and development. Nat. Rev. Genet. 15, 7–21 (2014).
Li, W., Notani, D. & Rosenfeld, M. G. Enhancers as non-coding RNA transcription units: recent insights and future perspectives. Nat. Rev. Genet. 17, 207–223 (2016).
Wapinski, O. & Chang, H. Y. Long noncoding RNAs and human disease. Trends Cell Biol. 21, 354–361 (2011).
Schmitt, A. M. & Chang, H. Y. Long noncoding RNAs in cancer pathways. Cancer Cell 29, 452–463 (2016).
Pang, J. C.-S. et al. KIAA0495/PDAM is frequently downregulated in oligodendroglial tumors and its knockdown by siRNA induces cisplatin resistance in glioma cells. Brain Pathol. 20, 1021–1032 (2010).
Zhang, X.-Q. et al. A long non-coding RNA signature in glioblastoma multiforme predicts survival. Neurobiol. Dis. 58, 123–131 (2013).
Sturm, D. et al. Hotspot mutations in H3F3A and IDH1 define distinct epigenetic and biological subgroups of glioblastoma. Cancer Cell 22, 425–437 (2012).
Tang, Z. et al. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 45, W98–W102 (2017).
Suvà, M. L. et al. Reconstructing and reprogramming the tumor-propagating potential of glioblastoma stem-like cells. Cell 157, 580–594 (2014).
Bulstrode, H. et al. Elevated FOXG1 and SOX2 in glioblastoma enforces neural stem cell identity through transcriptional control of cell cycle and epigenetic regulators. Genes Dev. 31, 757–773 (2017).
Hubert, C. G. et al. Genome-wide RNAi screens in human brain tumor isolates reveal a novel viability requirement for PHF5A. Genes Dev. 27, 1032–1045 (2013).
Blake, S. M. et al. Inactivation of the ATMIN/ATM pathway protects against glioblastoma formation. Elife 5, 1–24 (2016).
Toiber, D., Leprivier, G. & Rotblat, B. Long noncoding RNA: noncoding and not coded. Cell Death Discov. 3, 16104 (2017).
Anderson, K. M. et al. Transcription of the non-coding RNA upperhand controls Hand2 expression and heart development. Nature 539, 433–436 (2016).
Engreitz, A. J. M. et al. Neighborhood regulation by lncRNA promoters, transcription, and splicing. Nature 539, 452–455 (2016).
Gilbert, L. A. et al. CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154, 442–451 (2013).
Gilbert, La et al. Resource genome-scale CRISPR-Mediated control of gene repression and activation. Cell 159, 647–661 (2014).
Kearns, Na et al. Cas9 effector-mediated regulation of transcription and differentiation in human pluripotent stem cells. Development 141, 219–223 (2014).
Lee, S. B. et al. An ID2-dependent mechanism for VHL inactivation in cancer. Nature 529, 172–177 (2016).
Dötsch, V., Bernassola, F., Coutandin, D., Candi, E. & Melino, G. p63 and p73, the ancestors of p53. Cold Spring Harb. Perspect. Biol. 2, a004887 (2010).
Flores, E. R. et al. Tumor predisposition in mice mutant for p63 and p73: evidence for broader tumor suppressor functions for the p53 family. Cancer Cell 7, 363–373 (2005).
Niklison-Chirou, M. V. et al. TAp73 knockout mice show morphological and functional nervous system defects associated with loss of p75 neurotrophin receptor. Proc. Natl Acad. Sci. USA 110, 18952–18957 (2013).
Agostini, M. et al. p73 regulates maintenance of neural stem cell. Biochem Biophys. Res Commun. 403, 13–17 (2010).
Fujitani, M. et al. TAp73 acts via the bHLH Hey2 to promote long-term maintenance of neural precursors. Curr. Biol. 20, 2058–2065 (2010).
Landré, V., Antonov, A., Knight, R. & Melino, G. p73 promotes glioblastoma cell invasion by directly activating POSTN (periostin) expression. Oncotarget 7, 11785–11802 (2016).
Beier, D., Schulz, J. B. & Beier, C. P. Chemoresistance of glioblastoma cancer stem cells--much more complex than expected. Mol. Cancer 10, 128 (2011).
Hegi, M. E. et al. MGMT gene silencing and benefit from temozolomide in glioblastoma. N. Engl. J. Med. 352, 997–1003 (2005).
Carén, H. et al. Glioblastoma stem cells respond to differentiation cues but fail to undergo commitment and terminal cell-cycle arrest. Stem Cell Rep. 5, 829–842 (2015).
Bady, P. et al. MGMT methylation analysis of glioblastoma on the Infinium methylation BeadChip identifies two distinct CpG regions associated with gene silencing and outcome, yielding a prediction model for comparisons across datasets, tumor grades, and CIMP-status. Acta Neuropathol. 124, 547–560 (2012).
Bady, P. mgmtstp27: DNA methylation of MGMT promoter. R package version 0.6-3.le (2015).
Bester, A. C. et al. An integrated genome-wide CRISPRa approach to functionalize lncRNAs in drug resistance. Cell 173, 649–652.e20 (2018).
Tomita, H., Tanaka, K., Tanaka, T. & Hara, A. Aldehyde dehydrogenase 1A1 in stem cells and cancer. Oncotarget 7, 11018–11032 (2016).
Januchowski, R. et al. Inhibition of ALDH1A1 activity decreases expression of drug transporters and reduces chemotherapy resistance in ovarian cancer cell lines. Int J. Biochem Cell Biol. 78, 248–259 (2016).
Croker, A. K. et al. Differential functional roles of ALDH1A1 and ALDH1A3 in mediating metastatic behavior and therapy resistance of human breast cancer cells. Int J. Mol. Sci. 18, 2039 (2017).
Schäfer, A. et al. Aldehyde dehydrogenase 1A1—a new mediator of resistance to temozolomide in glioblastoma. Neuro Oncol. 14, 1452–1464 (2012).
Xiao, S., Wang, R., Wu, X., Liu, W. & Ma, S. The long noncoding RNA TP73-AS1 interacted with miR-124 to modulate glioma growth by targeting inhibitor of apoptosis-stimulating protein of p53. DNA Cell Biol. 37, 117–125 (2018).
Zhang, R., Jin, H. & Lou, F. The long non-coding RNA TP73-AS1 interacted with miR-142 to modulate brain glioma growth through HMGB1/RAGE pathway. J. Cell Biochem 119, 3007–3016 (2017).
Zhang, J. et al. Function and mechanism of long non-coding RNA KIAA0495 in glioma. Int J. Clin. Exp. Pathol. 9, 12423–12430 (2016).
Stupp, R. et al. Effects of radiotherapy with concomitant and adjuvant temozolomide versus radiotherapy alone on survival in glioblastoma in a randomised phase III study: 5-year analysis of the EORTC-NCIC trial. Lancet Oncol. 10, 459–466 (2009).
Eyler, C. E. & Rich, J. N. Survival of the fittest: cancer stem cells in therapeutic resistance and angiogenesis. J. Clin. Oncol. 26, 2839–2845 (2008).
Pistollato, F. et al. Intratumoral hypoxic gradient drives stem cells distribution and MGMT expression in glioblastoma. Stem Cells 28, 851–862 (2010).
Dean, M., Fojo, T. & Bates, S. Tumour stem cells and drug resistance. Nat. Rev. Cancer 5, 275–284 (2005).
Gusyatiner, O. & Hegi, M. E. Glioma epigenetics: from subclassification to novel treatment options. Semin. Cancer Biol. 2017: 30258–4.
Vlashi, E. et al. Metabolic state of glioma stem cells and nontumorigenic cells. Proc. Natl Acad. Sci. USA 108, 16062–16067 (2011).
William, D., Walther, M., Schneider, B., Linnebacher, M. & Classen, C. F. Temozolomide-induced increase of tumorigenicity can be diminished by targeting of mitochondria in in vitro models of patient individual glioblastoma. PLoS ONE 13, e0191511 (2018).
Liu, P. et al. Cytotoxic effect of disulfiram/copper on human glioblastoma cell lines and ALDH-positive cancer-stem-like cells. Br. J. Cancer 107, 1488–1497 (2012).
Raha, D. et al. The cancer stem cell marker aldehyde dehydrogenase is required to maintain a drug-tolerant tumor cell subpopulation. Cancer Res. 74, 3579–3590 (2014).
Rasper, M. et al. Aldehyde dehydrogenase 1 positive glioblastoma cells show brain tumor stem cell capacity. Neuro Oncol. 12, 1024–1033 (2010).
Rotblat B. et al. Proc. Natl Acad. Sci. USA 111, 3032–7. (2014) Epub 2014 Feb 10. https://doi.org/10.1073/pnas.1314421111.
R2: Genomics analysis and visualization platform (http://r2.amc.nl).
Acknowledgements
This research was supported by the Israel Science Foundation (grant No. 1124/15) (BR). This research was supported by a grant from the Ministry of Science, Technology, and Space of the State of Israel and the German Cancer Research Center (DKFZ) (MR and BR). This research is funded by the Israeli Cancer Association (grant #20180012). We thank Steven Pollard for sharing with us GBM stem cells.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by M. Agostini
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Mazor, G., Levin, L., Picard, D. et al. The lncRNA TP73-AS1 is linked to aggressiveness in glioblastoma and promotes temozolomide resistance in glioblastoma cancer stem cells. Cell Death Dis 10, 246 (2019). https://doi.org/10.1038/s41419-019-1477-5
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41419-019-1477-5
This article is cited by
-
Identification of immune and Toll-like receptor signaling pathway related feature lncRNAs to construct diagnostic nomograms for acute ischemic stroke
Scientific Reports (2023)
-
METTL3-mediated m6A modification of LINC00839 maintains glioma stem cells and radiation resistance by activating Wnt/β-catenin signaling
Cell Death & Disease (2023)
-
METTL3 knockdown promotes temozolomide sensitivity of glioma stem cells via decreasing MGMT and APNG mRNA stability
Cell Death Discovery (2023)
-
Targeting Wnt/β-catenin-mediated upregulation of oncogenic NLGN3 suppresses cancer stem cells in glioblastoma
Cell Death & Disease (2023)
-
PSRC1 Regulated by DNA Methylation Is a Novel Target for LGG Immunotherapy
Journal of Molecular Neuroscience (2023)