Abstract
Mesenchymal stem cells (MSCs) can differentiate into endothelial cells; however, the mechanisms underlying this process in the tumor microenvironment (TME) remain elusive. This study shows that tumor necrosis factor alpha (TNF-α), a key cytokine present in the TME, promotes the endothelial differentiation of MSCs by inducing vascular endothelial growth factor receptor 2 (VEGFR2) gene expression. EGR1 is a member of the zinc-finger transcription factor family induced by TNF-α. Our findings indicate that EGR1 directly binds to the VEGFR2 promoter and transactivates VEGFR2 expression. We also demonstrate that EGR1 forms a complex with c-JUN activated by c-JUN N-terminal kinase (JNK) to promote VEGFR2 transcription and endothelial differentiation in MSCs in response to TNF-α stimulation. The shRNA-mediated silencing of EGR1 or c-JUN abrogates TNF-α-induced VEGFR2 transcription and the endothelial differentiation of MSCs. To further evaluated the role of EGR1 in the endothelial differentiation of BM-MSCs, we used a syngenic tumor implantation model. 4T1 mouse mammary tumor cells were injected subcutaneously into BALB/c mice with primary mBM-MSCs isolated from wild-type (Egr1+/+) or Egr1-null (Egr1−/−) mice. CD31-positive cells were predominantly observed at the border of the tumor in the 4T1 plus wild-type MSC group, while staining less in the 4T1 alone or 4T1 plus Egr1-null MSC group. Collectively, these findings demonstrate that the JNK-EGR1 signaling axis plays a crucial role in the TNF-α-induced endothelial differentiation of MSCs in the TME, which could be a potential therapeutic target for solid tumors vasculatures.
Similar content being viewed by others
Introduction
The vasculature in solid tumors is essential for tumor survival, progression, and metastasis, associated with poor prognosis. Under hypoxic conditions, cancer cells induce sprouting angiogenesis in neighboring blood vessels using various pro-angiogenic factors to meet their oxygen and nutrient demands and invade the newly formed vasculature [1, 2]. However, recent studies have reported that the tumor vasculature has morphological features different from the normal tissue vasculature [3]. In some cases, treatment targeting the angiogenesis of pre-existing vessels proves ineffective; this puzzling result cannot be explained well using the classical model [4]. Accordingly, there are ongoing efforts to identify other cell types in the tumor microenvironment (TME) that can differentiate into endothelial-like phenotypes [5, 6].
Mesenchymal stem cells (MSCs), a type of stromal cells present in the TME, are adult multipotent cells that can differentiate into various connective tissue-specific cells, such as osteoblasts, chondrocytes, and adipocytes [7]. Owing to this feature, MSCs play a central role in physiological wound healing and are of interest in regenerative therapy [8].
MSCs enhance the progression of various cancer types, including breast cancer, lung cancer, and colorectal cancer [9,10,11]. A well-known mechanism by which MSCs help promote tumorigenesis involves their differentiation into cancer-associated fibroblasts that secrete cytokines to modulate the TME [12]. These pro-oncogenic cytokines include angiogenic factors, such as vascular endothelial growth factor (VEGF), secreted in hypoxic TMEs [13]. Additionally, MSCs can directly differentiate into endothelial-like cells. In response to VEGF-A stimulation, mechanical shear, or co-culture with endothelial cells, MSCs acquire the ability to generate capillary-like structures on basement membrane extracts, expressing various endothelial markers, such as vascular endothelial growth factor receptor-2 (VEGFR2, also known as KDR and FLK1), VE-Cadherin, and CD31 (Ref. [14,15,16]). Based on the evidence from these studies, MSCs are now considered an alternative source of tumor vasculature [17]. However, the detailed molecular mechanisms underlying the endothelial differentiation of MSCs in the TME are yet to be elucidated.
Tumor necrosis factor alpha (TNF-α) is a pro-inflammatory cytokine produced by various immune and stromal cells [18]. It modulates the survival, proliferation, invasion, and metastasis of cancer cells in the TME [19]. In addition, it is often associated with the angiogenesis-related physiological roles of MSCs; for example, TNF-α can induce MSCs to migrate into ischemic tissues and secrete various chemokines, recruiting endothelial progenitor cells and promoting angiogenesis for repairing injured tissues [20]. TNF-α is also involved in the differentiation of MSCs into osteoblasts [21]. However, it remains unclear whether TNF-α directly induces the endothelial differentiation of MSCs.
This study aimed to investigate whether TNF-α directly induces the endothelial differentiation of MSCs and, if so, identify the TNF-α-regulated genes and signaling pathways associated with MSC differentiation.
Materials and methods
Reagents
Recombinant human TNF-α was purchased from PeproTech (London, UK) and reconstituted in 0.1% bovine serum albumin/PBS solution. U0126, SB20358, and SP600125 were purchased from Calbiochem (San Diego, CA, USA). Tan IIA was purchased from Sigma-Aldrich (St. Louis, MO, USA). U0126, SB20358, SP600125, and Tan IIA were dissolved in dimethyl sulfoxide.
Plasmids and transfection
The complete CDS fragments of human c-JUN and mouse Egr1 were cloned into a pcDNA3.1-V5-His vector (Thermo Fisher Scientific, Waltham, MA, USA) for a C-terminal V5–6xHis tag, or into a pCMV-Tag4B vector (Stratagene, La Jolla, CA, USA) for a C-terminal FLAG tag. For gene knockdown, shRNA templates were inserted into the pSilencer 2.1 U6 Neo vector (Thermo Fisher Scientific) using HindIII and BamHI restriction enzymes, according to the manufacturer’s instructions. The target sequences of each gene were as follows: shCT (control), 5′-ACT ACC GTT GTT ATA GGT GT-3′; shVEGFR2, 5′-GGC GGC ACG AAA TAT CCT CTT A-3′; shEGR1, 5′-GTT ACT ACC TCT TAT CCA T-3′; shJUN, 5′-TAG TAC TCC TTA AGA ACA CAA-3′. The plasmids were then transfected into cells using Lipofectamine 2000 (Thermo Fisher Scientific) according to the manufacturer’s instructions.
Cell culture
Human bone marrow-derived (BM)-MSCs were purchased from PromoCell (Heidelberg, Germany) and cultured in Mesenchymal Stem Cell Growth Medium 2 (MSCGM2, PromoCell). Human umbilical vein endothelial cells (HUVECs) were purchased from Lonza Bioscience (Switzerland) and cultured in EGM-2 (Lonza Bioscience) according to the manufacturer’s instructions. HEK293 (human embryonic kidney cell line) and 4T1 (mouse mammary carcinoma cell line) cells were obtained from the American Type Culture Collection (ATCC, Manassas, VA, USA) and cultured in DMEM supplemented with 10% FBS and 1% penicillin-streptomycin (BioWest, Kansas City, MO, USA).
Endothelial tube formation assay
BM-MSCs, HUVECs, or HEK293 cells (1 × 104 each) were suspended in 100 μL EGM-2 medium with or without 10 ng/mL TNF-α and then seeded onto 96-well plates coated with Corning Matrigel Growth Factor Reduced basement membrane matrix (Cat. 356230, Corning, NY, USA). After 12–24 h, the appearance of a capillary-like network structure was imaged using an EVOS FL Auto (Thermo Fisher Scientific) microscope system. The tube area was measured by quantifying the total tube length and nodes and expressed as the pixel per well (1024 × 768 pixels) using the Angiogenesis Analyzer in ImageJ software (http://image.bio.methods.free.fr/ImageJ/?Angiogenesis-Analyzer-for-ImageJ). Statistical data were obtained by independent tree experiments obtaining the tube length and number of nodes from the entire well.
Spheroid-based sprouting angiogenesis assay
Spheroid-based sprouting angiogenesis assay was performed according to the previous report [22]. BM-MSCs were cultured in MSCGM2. After 2–3 passages, the cells were resuspended at a density of 800 cells in 25 μL spheroid forming medium containing 80% MSCGM2 and 20% Methocel (ThermoFisher Scientific, Waltham, MA, USA) to assemble into a single spheroid. After 24 h, spheroids were embedded in a collagen hydrogel containing DMEM containing 0.5% FBS and 1 mg/mL rat tail collagen (Thermo Fischer Scientific). After the gels solidified, 10 ng/mL TNF-α were treated. After 24 h, the spheroid sprouting images were captured. For immunofluorescent staining, spheroids were fixed with 3.7% paraformaldehyde and permeabilized using 0.3% Triton X-100, followed by incubating with FITC-conjugated anti-CD44 antibody. F-actin was stained with Rhodamine Red X-conjugated Phalloidin (Jackson ImmunoResearch, West Grove, PA, USA). Images were acquired using Zeiss LSM900 confocal microscope (Carl Zeiss Microimaging Inc. Thornwood, NY, USA). Z-stack images were analyzed using ZEN Microscopy software (Zeiss AG, Oberkochen, Germany).
DNA microarray and gene set enrichment assay (GSEA)
MSCs were treated with or without 5 ng/mL TNF-α for 18 h. Total RNA was isolated using NucleoZOL (Macherey-Nagel, Düren, Germany) according to the manufacturer’s instructions. The DNA microarray experiment was performed using an Agilent human whole genome 4 × 44 K v2 microarray set (Agilent Technologies, Santa Clara, CA, USA). cDNA synthesis, DNA labeling, hybridization, and data analysis were performed by GenomicTree, Inc. (Daejeon, Republic of Korea, http://www.genomictree.com) with Agilent GeneSpring GX software. Gene Set Enrichment Analysis (GSEA) was performed using the Enrichr tool for screening enriched terms (available at https://maayanlab.cloud/Enrichr) [23] and another GSEA software (http://www.broadinstitute.org/gsea/index.jsp) provided by the joint project of UC San Diego (San Diego, CA, USA) and Broad Institute of MIT and Harvard University [24, 25].
Reverse transcription (RT)-PCR
The cultured cells were lysed in NucleoZOL (Macherey-Nagel, Germany), and total RNA was isolated according to the manufacturer’s instructions. Equal quantities (1 μg) of RNA samples were used for synthesizing cDNA with the iScript cDNA Synthesis Kit (Bio-Rad, Hercules, CA, USA). The cDNA samples were subjected to standard PCR using gene-specific primer pairs (Supplementary Table S1). The PCR conditions were as follows: 4 min at 95 °C for the initial denaturation, followed by 30 cycles of 30 s at 95 °C, 30 s at 60 °C, and 30 s at 72 °C. The amplicons were resolved using 2% agarose gel electrophoresis and visualized under UV light.
Immunoblotting
Total cell lysates were obtained using RIPA buffer (50 mM Tris-HCl (pH 7.4), 400 mM NaCl, 1% Triton X-100, 0.25% sodium deoxycholate, 1 mM EDTA, 1 mM Na3VO4, and 1 mM NaF). Next, 15–30 μg of the lysates were separated using SDS-PAGE in 7–10% gels and transferred to 0.45 µm nitrocellulose membranes. The membranes were blocked with 5% skim milk in Tris-Tween buffered saline (TTBS; 100 mM Tris-HCl (pH 7.5), 150 mM NaCl, 0.01% (v/v) Tween-20) and treated with primary antibodies overnight at 4 °C. Next, the membranes were treated with the appropriate secondary antibodies for 1 h and visualized using the WestPico PLUS Chemiluminescent Substrate (Thermo Fisher Scientific). The antibodies used for immunoblotting are listed in Supplementary Table S2.
Expression of VEGFR2 siRNA
MSCs cultured in 35-mm dishes were transfected with 100 pmol AccuTarget Predesigned siRNA against VEGFR2 (siVR2) (Bioneer Co, Daejeon, Republic of Korea) using Lipofectamine RNAiMAX (Invitrogen, Carlsbad, CA, USA) in 100 μL Opti-MEM (Thermo Fisher Scientific, Waltham, MA, USA). At 6 h post-transfection, the cells were serum-starved for 12 h and then treated with or without 10 ng/mL TNF-α. After 12 h, the cells were collected and subjected to immunoblotting or an endothelial tube formation assay. Silencing of VEGFR2 expression was confirmed using immunoblotting.
Immunofluorescent staining
BM-MSCs were cultured on coverslips, treated, and fixed using 3.7% paraformaldehyde. The fixed cells were permeabilized using 0.2% Triton X-100, blocked with 5% bovine serum albumin, and then incubated with the primary antibodies. The probed cells were visualized using the appropriate secondary antibodies conjugated to a fluorescent dye. The antibodies used in the immunofluorescent staining assay are listed in Supplementary Table S2. Cell nuclei were stained with 1 μg/mL Hoescht 33258. Images were acquired using the LSM900 laser-scanning confocal microscope system (Zeiss, Germany). For colocalization analysis, images were processed using ImageJ software and EzColocalization plugin (http://sites.imagej.net/EzColocalization/plugins/) as described previously [26]. The colocalization probability was expressed as a threshold overlap score (TOS) values (linear) ranging from 0 to 1 (0: no colocalization, 1: all pixels colocalization).
Fractionation of cytoplasmic, soluble nuclear, and chromatin-bound proteins
Soluble cytoplasmic and nuclear chromatin-bound proteins were fractionated as described previously [27], with minor modifications. Briefly, approximately 2 × 106 BM-MSCs were lysed in hypotonic lysis buffer (10 mM HEPES-NaOH (pH 7.5), 10 mM KCl, 1.5 mM MgCl2, 0.1 mM EGTA, and 0.4% Triton X-100), and the supernatants were collected for cytoplasmic protein fractionation. For fractionating soluble nuclear proteins, the pellets were resuspended in the L150 buffer (20 mM HEPES-NaOH (pH 7.5), 150 mM NaCl, 1.5 mM MgCl2, 0.1 mM EGTA, and 0.1% Triton X-100), and the supernatants were collected. For fractionating chromatin-bound proteins, the remaining pellets were digested using MNase for 15 min at 37 °C and completely lysed in RIPA buffer.
VEGFR2 promoter-reporter activity assay
Genomic sequence information of VEGFR2 (NM_002253) was retrieved from the UCSC Genome Browser and analyzed. The serial 5′-truncated fragments of the VEGFR2 promoter were amplified from human genomic DNA and cloned into the pGL4.17 reporter vector (Promega, Madison, WI, USA). For the −756/−21 sequence, the PCR products were amplified using the primers −756 F (5′-TTG ATG TCA CTC CCC AGG TGT-3′) and −21 R (5′-GAA ACT CTA GAG CGC GGA GG-3′), ligated into the T&A vector (RBC Bioscience, Taipei County, Taiwan), and subcloned into the pGL4.17 vector using EcoRV and BglII restriction enzymes. For the −456/−21 and −111/−21 sequences, the PCR products were amplified using the primer −456 F (5′-GCT GAG TAT CCG CTT CTC CC-3′) and −21R and ligated into the T&A vector, and then subcloned into the pGL4.17 vector by double-digestion with SacI-BglII (for −456/−21) or KpnI-BglII (for −111/−21). For the −69/−21 sequence, the PCR products were amplified using the primer −69F (5′-CCG CTC CGG CCC CGC-3′) and −21 R, ligated into the T&A vector, and subcloned into the pGL4.17 vector using KpnI and BglII. For generating mutant EBS (mtEGR1), the pVEGFR2-Luc(−69/−21) construct was subjected to site-directed mutagenesis (Enzynomics, Daejeon, Republic of Korea) with a primer pair (forward, 5′-GCC TCC GCG CTC TAG AGT T-3′; reverse, 5′-CGG GCC ATG CGG GGC G-3′). The pVEGFR2-Luc constructs were transfected into HEK293 cells, and the luciferase activities were measured using the Dual-Glo Lucferase assay system (Promega) according to the manufacturer’s instructions.
Electrophoretic mobility shift assay (EMSA)
Nuclear extracts of treated BM-MSCs were obtained and subjected to EMSA using the LightShift chemiluminescent EMSA kit (Thermo Fisher Scientific), as described previously [28]. Purified human EGR1 protein was purchased from Sigma-Aldrich (Cat. E5777). Biotin-labeled probes (5′-biotin-CAT GGC CCC GCC TCC GCG CTC TAGA-3′) were synthesized by Bioneer (Korea, Daejeon). For the competition assay, unlabeled (cold) probes with a consensus EGR1-binding sequence (5′-GGA TCC AGC GGG GGC GAG CGG GGG CGA-3′) or AP-1-binding sequence (5′-CGC TTG ATG ACT CAG CCG GAA-3′) were pre-incubated for 5 min before the binding reactions.
DNA affinity precipitation assay (DAPA)
The cultured cells were harvested, and the nuclear fractions were isolated. The nuclear extracts (100 μg) were diluted using a DNA binding buffer (10 mM HEPES-NaOH (pH 8.0), 50 mM KCl, 100 mM NaCl, 5 mM MgCl2, 0.1% NP-40, and 0.25 mM EDTA) and pre-incubated with 0.5 μg/mL of salmon sperm DNA at 4 °C. After 30 min, 4 μg of the biotinylated or unlabeled oligonucleotides used in EMSA were added, and the mixture was incubated for 2 h at 4 °C. Next, the DNA-protein complexes were pull-downed using 30 μl of streptavidin-conjugated agarose beads (Cat. 20353, Thermo Fisher Scientific). The beads were washed three times with the binding buffer and boiled in 2× Laemmli sample buffer. The supernatants were resolved using 8% PAGE and subjected to immunoblotting.
Co-immunoprecipitation assay
The nuclear extracts of HEK293 cells were diluted using a co-immunoprecipitation buffer (10 mM HEPES-NaOH (pH 8.0), 50 mM KCl, 100 mM NaCl, 5 mM MgCl2, 0.5% Triton X-100, and 0.25 mM EDTA), and 2 μg of antibodies was added. After overnight rotation at 4 °C, the antibody-antigen complexes were precipitated using 10 μL of Dynabead protein G (Cat. 10004D, Thermo Fisher Scientific), washed three times using a binding buffer, and eluted using 200 mM glycine solution (pH 2.5). The supernatants were analyzed by immunoblotting.
To immunoprecipitate endogenous EGR1 and p-c-JUN, nuclear extracts of MSCs treated with 10 ng/mL TNF-α for 50 min were prepared and diluted using immunoprecipitation buffer (20 mM HEPES-NaOH (pH 7.4), 50 mM KCl, 100 mM NaCl, 5 mM MgCl2, 0.1% Triton X-100, and 0.25 mM EDTA). Nuclear extracts (200 μg) were incubated with anti-phospho-c-JUN (Ser63) antibody (1:50, Cell Signaling Technology, Danvers, MA, USA) or normal IgG overnight at 4 °C with rotation. The antibody-antigen complexes were precipitated using 30 μL of Dynabead protein G (Thermo Fisher Scientific, Cat. 10004D), washed three times with binding buffer, and eluted using 1× SDS-PAGE loading buffer. The supernatants were analyzed by immunoblotting.
Chromatin immunoprecipitation (ChIP) assay
Approximately 4 × 106 BM-MSCs were cross-linked using 1% paraformaldehyde, and the ChIP assay was performed using the Magnetic ChIP Kit (Thermo Fisher Scientific) according to the manufacturer’s instructions. For each immunoprecipitation, 8 μg of antibodies was used. The ChIP samples were subjected to PCR analysis using locus-specific primer pairs. The primer sequences for the target site were 5′-TGG GGA GAT GTA AAT GGG CT-3′ (−159 F) and 5′-GAC CAC ACA TTG ACC GCT C-3′ (+41 R). For the negative control, off-target primer pair (forward, 5′-CCA CAA GAA GTC CAC ACA CG-3′ (−884F); reverse, 5′-CTT TTG TCC ATC GTG CTC CC-3′ (−673 R)) was used.
Animal study
Egr1-null mice on a C57BL/6 background were provided by Dr. Jeffrey Milbrandt (Washington University, St. Louis, MO, USA) [29]. Mouse BM-MSCs (mBM-MSCs) were isolated from wild-type and Egr1-null mice as reported previously [30] and cultured in MSC growth medium (10% FBS in DMEM/MSCGM2 (1:1)). Wild-type and Egr1-null genotypes were verified by PCR analysis of genomic DNA (forward primer, 5′-AAC CGG CCC AGC AAG ACA CC-3′; reverse primer, 5′-GGG CAC AGG GGA TGG GAA TG-3′). mBM-MSCs from passage 3 were labeled with CellTracker CM-Dil red fluorescence dye (Invitrogen) according to the manufacturer’s instructions. 4T1 mouse mammary tumor cells (5 × 105) and CM-Dil-labeled mMSCs (1 × 105) were co-injected subcutaneously into BALB/c mice (n = 5). After 5 days, the mice were sacrificed by exposure to CO2, and the tumor tissues were fixed in 3.7% paraformaldehyde overnight at 25 °C. After embedding in paraffin blocks, tumor tissues were cut into 5 μm sections. To observe DiI-labeled mMSCs and CD31-positive endothelial cells, tumor sections were assessed using fluorescent immunohistochemistry methods with an anti-CD31 antibody (Abcam, Cambridge, UK) and Alexa Fluor 488-conjugated anti-rabbit IgG secondary antibody and visualized under GFP and RFP fluorescence channel. The mouse experiments were carried out according to the guidelines for animal experiments and procedures of the Konkuk University Institutional Animal Care and Use Committee (IACUC) (approval number KU22041).
Statistical analysis
Data were analyzed using GraphPad Prism version 8.3.1 (GraphPad Software Inc., La Jolla, CA, USA), and expressed in terms of means ± SD. Statistical analysis was performed using a Student’s t-test or one-way analysis of variance, followed by Sidak’s or Dunnett’s multiple comparisons test. Differences with p < 0.05 were considered statistically significant.
Results
TNF-α induces the endothelial differentiation of BM-MSCs
To investigate whether TNF-α induces the endothelial differentiation of BM-MSCs, we performed a Matrigel-based tube formation assay [16]. Tube formation was analyzed by quantifying the total tube length and branch point (node) of the capillary-like networks using the Angiogenesis Analyzer program of the ImageJ software as pixel per total area. We found that cells initially attached to the Matrigel migrated toward each and then gradually formed endothelial capillary-like structures by 12–24 h (Fig. 1A). Notably, the capillary-like networks were more efficiently matured in the presence of TNF-α (Fig. 1B, C). TNF-α-stimulated BM-MSCs, but not non-endothelial HEK293 cells, were incorporated into endothelial capillary-like networks formed by HUVECs (Fig. 1D), suggesting that TNF-α-stimulated BM-MSCs exhibit the potential for endothelial differentiation. To confirm the potency of the endothelial differentiation of BM-MSCs, we performed a spheroid sprouting assay in collagen matrix-based three-dimensional (3-D) culture. The presence of TNF-α substantially increased the sprouting capacity of the spheroids (Supplementary Fig. S1A). Confocal microscopy immunofluorescence and 3-D image analysis revealed that sprouts stained with CD44 and F-actin formed a primary lumen that elongated along the sprout directions, as expected for new vessel formation (Supplementary Fig. S1B, C), suggesting that TNF-α induces endothelial cell differentiation of BM-MSCs, leading to the formation of capillary-like tubular structures under 3-D spheroid culture conditions.
A–C Representative images of capillary-like structures of TNF-α-stimulated (10 ng/mL) or unstimulated primary bone marrow-derived (BM)-MSCs on Matrigel. Images were obtained immediately (0 h) or 12 and 24 h after the cells adhered to Matrigel. The right panel shows images of the indicated areas at 24 h at a higher magnification. Dotted boxes represent ImageJ Analyzer images for quantifying total tube length and branch points (nodes). The tube length and node number per unit area of the structure were measured in (B) and (C), respectively. Error bars represent means±S.D (n = 3). **p < 0.01; ***p < 0.001; ns not significant, unpaired t-test. D Co-culture of PKH26-labeled human umbilical vein endothelial cells (HUVECs) (red fluorescence) alone or with GFP-expressing BM-MSCs or HEK293 cells on Matrigel in the presence of TNF-α. The lower panels show images of the indicated areas at a higher magnification. The incorporation efficiency of BM-MSCs or HEK293 cells into the capillary structure is quantified in the right panel. Error bars represent means ± S.D (n = 3). E Gene set enrichment analysis showed that vasculature development-related genes are enriched in TNF-α-stimulated BM-MSCs compared with that in unstimulated BM-MSCs. NES, normalized enrichment score; FDR, false discovery rate. F Serum-starved (0.5% FBS) BM-MSCs were treated with 10 ng/mL TNF-α for the indicated durations (0, 6, and 12 h), and total RNA was extracted. The mRNA levels of endothelial genes were measured using reverse transcription-polymerase chain reaction, as indicated. GAPDH was used as the internal control. G BM-MSCs were serum-starved (0.5% FBS) and treated with 10 ng/mL TNF-α for 12 h or left untreated. The cells were lysed, and the protein levels of indicated endothelial genes were measured by immunoblotting. GAPDH was used as the internal control.
To identify the gene sets that regulate the TNF-α-induced endothelial differentiation of MSCs, we examined the genes expressed by TNF-α in BM-MSCs using an Agilent human whole genome microarray (Supplementary Fig. S2A). Gene Ontology analysis using the Enrichr tool revealed that the angiogenesis gene set was highly expressed following TNF-α stimulation (Supplementary Fig. S2B, C). GSEA confirmed that a vasculature development-related gene set was highly enriched after TNF-α stimulation of MSCs (Fig. 1E). These data suggest that TNF-α induces endothelial differentiation of MSCs by upregulating endothelial inducer genes. Using RT-PCR and immunoblotting, we also observed that other endothelial marker proteins, including VCAM1 and VE-cadherin [31], were upregulated upon TNF-α stimulation (Fig. 1F, G).
VEGFR2 is required for the TNF-α-induced endothelial differentiation of BM-MSCs
VEGF-A stimulates the differentiation of MSCs into endothelial cells [16, 32]. We also observed that the length and node numbers of the capillary-like structure of differentiated BM-MSCs (assessed in the Matrigel-based tube formation assay) were enhanced when Endothelial Growth Medium-2 (EGM-2) was used compared to that observed when VEGF-A-free DMEM was used (Supplementary Fig. S3A–C). As assessed using RT-PCR, VEGFR2 mRNA expression increased considerably within 12 h of TNF-α stimulation, whereas VEGFR1 and VEGFR3 mRNA expression remained unaffected (Fig. 2A). Hence, we hypothesized that an increase in VEGFR2 expression might contribute to the TNF-α-induced differentiation of BM-MSCs into endothelial cells. To confirm this, we performed the siRNA-mediated knockdown of VEGFR2 expression in BM-MSCs. The expression of TNF-α-induced VCAM1 was substantially abrogated by siRNA-mediated knockdown of VEGFR2 expression (Fig. 2B). The Matrigel-based tube formation assay shows that TNF-α-induced endothelial-tube-like structure formation was significantly decreased upon transfection of siVR2 compared to that in control (siCT) (Fig. 2C). Pretreatment with neutralizing antibodies against VEGFR2 reduced the TNF-α-induced endothelial-tube-like structure formation (Fig. 2D). These data suggest that the TNF-α-induced endothelial differentiation of BM-MSCs requires VEGFR2 expression.
A Serum-starved (0.5% FBS) bone-marrow-derived (BM)-MSCs were treated or left untreated with TNF-α (10 ng/mL) for 12 h, and total RNA was extracted. The mRNA levels of VEGFR1, VEGFR2, and VEGFR3 were measured using reverse transcription-polymerase chain reaction, as indicated. B BM-MSCs were transfected with scrambled siRNA (siCT) or siVEGFR2 (siVR2 #1 and #2). After transfection, the siRNA-expressing cells were dissociated, serum-starved, and treated with TNF-α (10 ng/mL) for 12 h or left untreated. The VEGFR2 protein levels were measured by immunoblotting. C BM-MSCs transfecting control scrambled siRNA (siCT) or siVEGFR2 (siVR2) were seeded on Matrigel in the presence or absence of TNF-α (10 ng/mL). Images of the capillary-like structure of indicated BM-MSCs were obtained after 12 h. Dotted boxes represent ImageJ Analyzer images for quantifying total tube length and branch points (nodes). Scale bar, 1000 µm. The tube length and node number per unit area of the structure were measured with bar graphs. Error bars represent means ± S.D (n = 3). **p < 0.01; ***p < 0.001; ns not significant, unpaired t-test. D BM-MSCs were treated with anti-VEGFR2 antibody or normal immunoglobin (IgG) for 1 h, following which the Matrigel tube formation assay was performed in the presence or absence of TNF-α (10 ng/mL), as indicated. Images of the capillary-like structure of BM-MSCs under each treatment condition were obtained 12 h later. Scale bar, 1000 µm. The tube length and node number per unit area of the structure were measured with bar graphs. Error bars represent means±S.D. (n = 3). **p < 0.01, unpaired t-test.
EGR1 binds to and activates the VEGFR2 promoter in TNF-α-stimulated BM-MSCs
To determine the molecular mechanism underlying TNF-α-induced VEGFR2 expression, we generated four VEGFR2 promoter constructs with random deletions (Fig. 3A, top) and transfected them into HEK293 cells, followed by TNF-α treatment. Even the shortest −69/−21 construct responded to TNF-α stimulation in HEK293 cells (Fig. 3A, bottom). This indicated that the −69/−21 region is required for the complete activation of the VEGFR2 promoter under TNF-α stimulation.
A The upper panel shows the genomic map of the VEGFR2 promoter. The primers and restriction sites used for cloning are indicated. HEK293 cells were transfected with 0.1 µg of a series of 5′-deletion constructs of human VEGFR2 promoter-luciferase plasmids (pVEGFR2-Luc). Following transfection, the cells were dissociated, serum-starved, and treated with either the vehicle or 10 ng/mL TNF-α. After 12 h, the luciferase activities were measured. The upper panel shows the schematic locations of primers for the generation of pVEGFR2-Luc constructs. ‘+1’ indicates the transcription start site of the human VEGFR2 transcript (KDR; NM_002253). Error bars represent means ± S.D. (n = 3). *p < 0.05; ***p < 0.001; unpaired t-test. B The genomic sequence of the (−69/−21) region with a putative EBS. C The electromobility shift assay (EMSA) was performed to analyze the binding of EGR1 proteins to the putative EBS of the VEGFR2 promoter. The upper panel shows the sequence of the biotin-labeled probe. For EMSA, 50 mol of biotin-labeled EBS probes were incubated in the presence or absence of purified EGR1 proteins and resolved in a 6% acrylamide gel. For competition assays, 25, 50, and 100 molar excess quantities of unlabeled (cold) probes were pre-incubated with EGR1 proteins before binding reactions. The arrow indicates EGR1-binding probes. D HEK293 cells were transfected with indicated quantities of the pVEGFR2-Luc(−69/−21) plasmid, following which the luciferase assay was performed (upper panel). The residual lysates were subjected to immunoblotting for confirming EGR1 overexpression (bottom panel). Error bars represent means ± S.D. (n = 3). E The pVEGFR2-Luc(−69/−21) plasmids with wild-type (WT) or mutant EBS (mtEGR1) sequences were introduced into HEK293 cells. After serum-starvation, TNF-α was administered to the cells, and the luciferase activities were measured. Error bars represent means ± S.D. (n = 3). ***p < 0.001; ns not significant, unpaired t-test. F Serum-starved bone-marrow-derived (BM)-MSCs were cultured in the presence or absence of TNF-α for 90 min and cross-linked with formaldehyde. After cross-linking, chromatin immunoprecipitation assays were performed using normal rabbit IgG or anti-EGR1 antibodies. The input or ChIP samples were subjected to standard PCR with primers for the detection of the off-target region (Set B, −884/−673) or target EBS (Set A, −159/+41). DPC, DNA-protein complex. EBS, EGR1-binding site; TSS transcription start site, ChIP chromatin immunoprecipitation.
We analyzed the genome landscape of the VEGFR2 promoter using the UCSC genome browser to identify the transcription factors that regulate VEGFR2 transcription under TNF-α stimulation (Supplementary Fig. S4A). The −69/−21 region was layered on the H3K4Me3 marker and DNase I hypersensitive region. Additionally, the binding sites for the TATA-binding protein (TBP) and p300 co-activator were located in this region. We further analyzed the putative transcription factor-binding motifs in the −69/−21 region using the Genomatix MatInspector software. Seven transcription factors were predicted to bind to the −69/−21 region, of which EGR1 had the highest matrix similarity score (Fig. 3B, Supplementary Fig. S4B).
As EGR1 is upregulated by TNF-α in fibroblasts, keratinocytes, and MSCs [33,34,35], EGR1 may play a key role in the TNF-α-induced endothelial differentiation of MSCs by regulating VEGFR2 expression. To test this possibility, we first determined whether EGR1 binds to the VEGFR2 promoter using the EMSA. Purified EGR1 proteins formed a complex with a biotin-labeled deoxyoligonucleotide probe containing a putative EGR1-binding sequence (EBS; 5′-CAT GGC CCC GCC TCC GCG CTC TAG A-3′) in the −69/−21 region, which was dose-dependently inhibited by cold probes (Fig. 3C). Additionally, the promoter activity of the −69/−21 construct increased when the EGR1 expression vector was co-transfected into HEK293 cells (Fig. 3D). Next, to confirm whether the putative EBS motif is critical for activating promoter activity by TNF-α stimulation, we generated a mutant construct by substituting CGCC with GGCC, which significantly reduced the binding score of EGR1 (Supplementary Fig. S4C). This was followed by TNF-α treatment after the transfection of the mutant construct into HEK293 cells. The TNF-α-inducible promoter activity was suppressed in the mutant construct compared to that in the wild-type sequence (Fig. 3E). Finally, the binding of endogenous EGR1 to the VEGFR2 promoter was confirmed in a ChIP assay using BM-MSCs (Fig. 3F). The data suggest that EGR1 binds to the putative EBS in the −69/−21 region and enhances VEGFR2 promoter activity in response to the TNF-α stimulation of BM-MSCs.
EGR1 regulates TNF-α-induced VEGFR2 expression and the endothelial differentiation of BM-MSCs
To further confirm the role of EGR1 in regulating VEGFR2 expression, we generated a stable EGR1-knockdown cell line by transfecting EGR1 shRNA into BM-MSCs. The TNF-α-induced expression of VEGFR2 and VCAM-1 (an endothelial marker) was substantially suppressed in EGR1-knockdown cells (shEGR1) compared to that in control cells (shCT) (Fig. 4A). Furthermore, EGR1 silencing reduced TNF-α-induced endothelial tube-like structure formation (Fig. 4B) and the tube length (Fig. 4C). Therefore, EGR1 substantially contributes to TNF-α-induced endothelial differentiation of BM-MSCs by regulating VEGFR2 expression.
A BM-MSCs expressing control shRNA (shCT) or shEGR1 were serum-starved and cultured in the presence or absence of TNF-α for the indicating durations. The cell lysates were subjected to immunoblotting to measure the expression levels of EGR1, VEGFR2, and VCAM-1. B Capillary-like network of shCT or shEGR1-expressing BM-MSCs on Matrigel in the presence or absence of 10 ng/mL TNF-α. Scale bar, 1000 µm. Dotted boxes represent ImageJ Analyzer images for quantifying total tube length and branch points (nodes The tube length and node number per unit area of the structure were measured with bar graphs. Error bars represent means ± S.D. (n = 3). *p < 0.05; **p < 0.01; ***p < 0.001; ns, not significant, unpaired t-test.
The JNK and ERK pathways regulate the TNF-α-induced endothelial differentiation of BM-MSCs through EGR1 and VEGFR2 expression
TNF-α upregulated EGR1 expression in BM-MSCs within 30 min of its administration (Fig. 5A). Because EGR1 expression is induced by the MAPK family proteins in various cell types [34, 36, 37], we analyzed the activation of MAPK signaling pathways by TNF-α in BM-MSCs. ERK1/2, p38, and JNK1/2 MAPKs were rapidly phosphorylated within 15 min of TNF-α stimulation (Fig. 5B). The pharmacological inhibitors for MAPK kinase 1/2 (U0126), p38 (SB20358), and JNK1/2 (SP600126) were administered to determine the MAPK pathway involved in TNF-α-induced EGR1 and VEGFR2 expression in BM-MSCs. U0126 and SP600125 suppressed TNF-α-induced EGR1 expression, whereas SB203580 enhanced it (Fig. 5C). The effects of MAPK inhibitors on TNF-α-induced endothelial differentiation of BM-MSCs were further assessed using the Matrigel-based tube formation assay. Treatment with U0126 and SP600125 inhibited the TNF-α-induced capillary-like structure formation in BM-MSCs (Fig. 5D). Notably, U0126 (Fig. 5E) and SP600125 (Fig. 5F) dose-dependently inhibited TNF-α-induced VEGFR2 expression. These data suggest that both the ERK1/2 and JNK1/2 signaling pathways are involved in EGR1-mediated VEGFR2 expression upon TNF-α stimulation.
A, B Serum-starved bone-marrow-derived (BM)-MSCs were stimulated using TNF-α for the indicated durations, and total lysates were obtained. The lysates were subjected to immunoblotting with antibodies against EGR1 (A), p-ERK1/2 (T202/Y204), p-p38 (T180/Y182), or p-JNK1/2 (T183/Y185) (B). An anti-GAPDH antibody was used as the internal control. C Serum-starved BM-MSCs were pretreated with 1 μM U0126, 20 μM SB203580, or 25 μM SP600125 for 1 h and stimulated with TNF-α for 1 h. Cells were harvested and subjected to immunoblotting with antibodies against EGR1. An anti-GAPDH antibody was used as the internal control. D BM-MSCs were pretreated with U0126, SB203580, or SP600125 for 1 h, and then subjected to the Matrigel tube formation assay in the presence or absence of TNF-α, as indicated. The tube length and node number per unit area of the structure were measured with bar graphs. Error bars represent means ± S.D. (n = 3). **p < 0.01; ***p < 0.001, unpaired t-test. E, F Serum-starved BM-MSCs were pretreated with U0126 (E) and SP600125 at different concentrations (F) for 1 h, and then subjected to TNF-α stimulation for 12 h. Cells were harvested and subjected to immunoblotting with antibodies against VEGFR2. An anti-GAPDH antibody was used as the internal control.
c-JUN, a downstream component of JNK, cooperates with EGR1 to induce endothelial differentiation of BM-MSCs
JNK1/2 phosphorylates and activates the c-JUN protein, a member of the activator protein 1 (AP-1) family [38], which also reportedly regulates angiogenesis [39, 40]. In BM-MSCs, c-JUN was phosphorylated within 10 min of TNF-α stimulation (Fig. 6A). We further determined whether c-JUN activation also affects VEGFR2 induction. Tanshinone IIA (Tan IIA), an AP-1 inhibitor [41], was used to block AP-1 transcriptional activity in BM-MSCs before TNF-α treatment. Tan IIA suppressed VEGFR2 induction in a dose-dependent manner (Fig. 6B). Furthermore, TNF-α-induced VEGFR2 expression was attenuated by c-JUN knockdown (shJUN) compared to that in control (shCT) (Fig. 6C). Transfection of the c-JUN or EGR1 overexpression vector into BM-MSCs increased the formation of the capillary-like structure, whereas co-transfection of c-JUN and EGR1 synergistically promoted the formation of the capillary-like network (Fig. 6D, E). These data suggest that c-JUN and EGR1 cooperate to contribute to the endothelial differentiation of BM-MSCs. These data suggest that the cooperation between c-JUN and EGR1 contributes to the endothelial differentiation of BM-MSCs.
A Bone-marrow-derived (BM)-MSCs were serum-starved and stimulated with tumor necrosis factor alpha (TNF-α). After the indicated durations, immunoblotting was performed with antibodies against p-c-JUN (S63). Anti-c-JUN and GAPDH antibodies were used as internal controls. B Serum-starved BM-MSCs were pretreated with tanshinone IIA (Tan IIA) at different concentrations for 1 h, and then cultured in the absence or presence of TNF-α for 12 h. The vascular endothelial growth factor receptor 2 (VEGFR2) protein levels under each treatment condition were measured by immunoblotting. C BM-MSCs were transfected with the control shRNA (shCT) or shEGR1 vector. After 24 h, the cells were dissociated, serum-starved, and incubated with the vehicle or TNF-α for 12 h. The lysates were analyzed by immunoblotting using anti-c-JUN and anti-VEGFR2 antibodies. GAPDH was used as the internal control. D BM-MSCs were transfected with c-JUN- or Egr1-expressing vectors, as indicated. After transfection, the BM-MSCs were seeded on Matrigel for capillary-like tube formation. Middle panels, ImageJ Analyzer images for quantifying total tube length and nodes; Bottom panels, the higher magnification of the indicated areas. E The tube length and node number per unit area of the structure were measured with bar graphs. Error bars represent means ± S.D. (n = 3). *p < 0.05; **p < 0.01; ***p < 0.001; ns not significant, unpaired t-test.
c-JUN physically interacts with EGR1 on the EBS in the VEGFR2 promoter
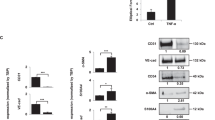
As c-JUN and EGR1 were found to stimulate TNF-α-induced VEGFR2 expression synergistically, we performed the ChIP assay to examine whether c-JUN binds to the −69/−21 region. Phosphorylated c-JUN bound to the −69/−21 region within 40 min of TNF-α-stimulation (Fig. 7A). Interestingly, no putative AP-1 homodimer/heterodimer binding motifs were identified around the −69/−21 region using the MatInspector or JASPAR software. EGR1 and AP-1 family proteins were shown to interact indirectly for regulating the tyrosine hydroxylase gene in PC12 cells [42]. Based on this finding and our observation, we hypothesized that c-JUN indirectly binds to the VEGFR2 promoter region by interacting with DNA-bound EGR1 protein. To test this, we first determined the localization of EGR1 and phosphorylated c-JUN. The subcellular fractionation experiment demonstrates that both EGR1 and p-c-JUN were detected in the chromatin-bound protein fractions upon TNF-α stimulation (Fig. 7B). Immunofluorescence staining also shows the probability of colocalization between EGR1 and p-c-JUN in the nucleus (Fig. 7C). To determine whether EGR1 and c-JUN interact directly, we performed an immunoprecipitation assay using nuclear extracts of HEK293 cells co-transfected with V5-tagged Egr1 and Flag-tagged c-JUN. c-JUN was precipitated by the anti-V5 antibody (Fig. 7D), and EGR1 was precipitated with an anti-FLAG antibody (Fig. 7E). Additionally, EGR1 co-immunoprecipitated with the anti-phospho-c-JUN antibody in TNF-α-treated MSCs (Fig. 7F). These data suggest that under TNF-α stimulation, EGR1 physically interacts with phosphorylated c-JUN in the nucleus
A Serum-starved bone-marrow-derived mesenchymal stem cells (BM-MSCs) were stimulated by TNF-α for 40 min. After chromatin cross-linking, the chromatin immunoprecipitation (ChIP) assay was performed using normal rabbit IgG or anti-p-c-JUN (S63) antibody. The input samples or ChIP samples were subjected to PCR with the indicated primer sets. B BM-MSCs were serum-starved, stimulated by TNF-α for 40 min, and subjected to subcellular fractionation. The fractionation samples were analyzed by immunoblotting with anti-EGR1 and anti-p-c-JUN (S63) antibodies. The GAPDH, lamin B, and histone H3 levels were used as internal controls for the cytosolic, soluble nuclear, and chromatin-bound protein fractions, respectively. Cyt, cytosolic fraction, S.N, soluble nuclear fraction, C.B.P, chromatin-bound proteins fraction. C Immunofluorescence staining of BM-MSCs using the indicated antibodies. Hoescht33258 was used for DNA staining. Confocal microscope images were processed using ImageJ software and EzColocalization plugin. Scale bar, 100 μm. The colocalization probability was expressed as a threshold overlap score (TOS) values (linear) ranging from 0 to 1 (0: no colocalization, 1: all pixels colocalization). Error bars represent means ± S.D. (n = 33 (0 min), 41 (20 min), 38 (50 min). **p < 0.01; ***p < 0.001, unpaired t-test. D, E The nuclear extracts of HEK293 cells expressing V5-tagged Egr1 and FLAG-tagged c-JUN were immunoprecipitated using an anti-V5 (α-V5) antibody (D) and an anti-FLAG (α-FLAG) antibody (E). F BM-MSC cells were stimulated by TNF-α for 40 min, and nuclear extracts were immunoprecipitated with a normal mouse IgG or p-c-JUN antibody. The eluted samples were subjected to immunoblotting using the indicated antibodies. G Nuclear extracts of BM-MSCs stimulated by the vehicle or TNF-α for 40 min were obtained, and treatment with the biotinylated probe was performed using the electromobility shift assay. For the competition assay, 50-fold molar excess unlabeled oligonucleotides (cold probe) were used, as indicated. DPC, DNA-protein complex. H Nuclear extracts of HEK293 cells expressing V5-tagged Egr1 or FLAG-tagged c-JUN were pull-downed by DNA-affinity purification assay (DAPA) with the biotinylated probes. I Serum-starved BM-MSCs were stimulated by TNF-α for 40 min, and nuclear extracts were isolated. The extracts were pull-downed by DAPA. Unlabeled oligonucleotide probes were used for the negative control. The elutes samples were subjected to immunoblotting using indicated antibodies. N.C. negative control.
To further determine the interaction of EGR1 and c-JUN on the VEFGR2 gene, we performed an EMSA using nuclear extracts with a biotin-labeled probe corresponding to the putative EBS in the VEGFR2 promoter −69/−21 region. The DNA-protein complex that increased upon TNF-α stimulation decreased upon the addition of the unlabeled consensus AP-1-binding sequence or EBS (Fig. 7G), suggesting that EGR1 and c-JUN form a complex on the EBS of VEGFR2. To evaluate the interaction between EGR1 and c-JUN in the VEGFR2 promoter, we performed a DAPA. Both EGR1 and c-JUN were detected in the pull-down of EBS oligonucleotides upon TNF-α stimulation (Fig. 7H). These data suggest that the EGR1:c-JUN complex could be formed on the EBS of the VEGFR2 promoter.
Egr1-deficient BM-MSCs impair tumor neovascularization
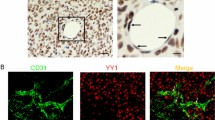
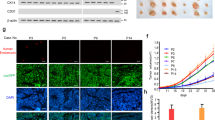
Finally, we evaluated the role of EGR1 in the endothelial differentiation of BM-MSCs and its contribution to tumor neovascularization using a syngenic tumor implantation model. 4T1 mouse mammary tumor cells were implanted subcutaneously into BALB/c mice with primary CM-Dil-labeled MSCs isolated from wild-type (Egr1+/+) or Egr1-null (Egr1/−) mice. Tumor neovascularization was assessed by immunostaining of the endothelial marker CD31. The two-color overlay shows that CD31-positive cells were predominantly observed at the border of the tumor in the 4T1 plus wild-type MSC group, while staining less in the 4T1 alone or 4T1 plus Egr1-null MSC group (Fig. 8). It can be seen that MSCs contribute to tumor neovascularization and that Egr1 plays a crucial role in the endothelial differentiation of MSC. In conclusion, the EGR1/VEGFR2 axis is functional in mediating the endothelial differentiation of BM-MSCs to promote tumor neovascularization.
A The BALB/c mice were implanted with 4T1 mouse mammary tumor cells alone, 4T1 cells plus mouse wild-type CM-Dil-labeled BM-MSCs (Dil-mMSC Egr1+/+), or 4T1 cells plus CM-Dil-labeled Egr1-null BM-MSCs (Dil-mMSC Egr1−/−). Tumor sections were stained with hematoxylin and eosin and immunostained with anti-CD31 antibody (green). Nuclear DNAs were stained with 1 µg/mL Hoechst 33258. Fluorescently labeled cells were examined using an EVOS FL fluorescence microscope (Advanced Microscopy Group, Bothell, WA, USA). Arrows indicate mMSC cells differentiated into endothelial cells. Scale bars, 100 μm or 50 μm.
Discussion
This study showed that TNF-α triggers endothelial differentiation of primary human BM-MSCs by upregulating VEGFR2 expression. In addition, we demonstrated that TNF-α-induced EGR1 physically interacts and forms a complex with c-JUN on the EBS in the core promoter region of the VEGFR2 to promote transcription of the VEGFR2.
The tumor vasculature was considered to be formed via sprouting angiogenesis from pre-existing blood vessels surrounding a tumor. However, recent findings show that the tumor vasculature has characteristics different from normal blood vessels, such as high tortuosity, leakiness, and low pericyte coverage [3, 4]. Therefore, the tumor vasculature is now thought to have an origin different from that of the normal vasculature. MSCs, which are adult multipotent cells that can differentiate into various connective tissues, can reportedly differentiate into endothelial-like cells. Although MSCs are now considered a non-endothelial source of the tumor vasculature, the effects of TME-related cytokines on the endothelial differentiation of MSCs are not well-studied.
TNF-α, a pro-inflammatory cytokine in the TME, stimulates MSCs to aid tumor angiogenesis [20]. Building on this prior evidence, we showed, using endothelial tube formation assays, gene set enrichment analysis, and immunoblotting assays, that TNF-α promotes the endothelial differentiation of BM-MSCs.
VEGF/VEGFR signaling is considered to play a critical role in the endothelial differentiation of MSCs since VEGF-A-containing media can stimulate the endothelial differentiation of MSCs [16, 32]. VEGF-A, secreted from hypoxic cancer cells but also by MSCs, is a key signal peptide that modulates tumor vascularization [43, 44]. Consistent with findings from previous studies, we observed that the capillary-like structure of BM-MSCs was well-formed in media containing VEGF-A, such as EGM-2. Although the expression of VEGFRs is required for the acceptance of autocrine or paracrine VEGF-A, the molecular mechanism underlying the upregulation of VEGFRs in MSCs in the TME remains unknown.
We observed that VEGFR2 expression was substantially increased in response to TNF-α-stimulation. Furthermore, VEGFR2 expression was necessary for the TNF-α-induced endothelial differentiation of BM-MSCs. Although VEGFR1 is the primary receptor for VEGF-A, it plays potentially inhibitory roles in endothelial proliferation during early development [45]. This dual role could explain why VEGFR1 is not expressed in TNF-α-stimulated BM-MSCs. BM-MSCs specifically express VEGFR2 in the presence of TNF-α, thereby efficiently binding VEGF-A and maximizing endothelial differentiation.
The TNF receptor (TNFR) family comprises TNFR1 and TNFR2. When TNF-α binds to trimeric TNFR, adaptor proteins such as TRADD are recruited, followed by the activation of various transcriptional factors, such as NF-κB, STAT, and EGR1, which regulate gene expression [46, 47]. Using promoter-reporter assays, we demonstrated that the region −69 to −21 bp upstream of the transcription start site represents the core region of the TNF-α-induced cis-regulatory element. In this region, the EBS motif with the highest score is present in the most conserved sequence close to −48 bp upstream of the transcription start site. We determined that EGR1 binds to the motif and functions as the primary regulator of VEGFR2 promoter activity.
EGR1 is a transcription factor with a three-tandem zinc finger domain that binds to GC-rich sequences and regulates gene expression in response to various cellular stimulations [48]. EGR1 participates in angiogenesis by regulating the expression of various pro-angiogenic factors in tumor cells [49, 50] and mediates MSC differentiation [51, 52]. EGR1 expression in BM-MSCs was regulated by the ERK and JNK pathways, two signaling pathways activated by TNF-α stimulation. Interestingly, only the JNK pathway is involved in VEGFR2 expression and the TNF-α-induced endothelial differentiation of MSCs.
JNK primarily activates the AP-1 transcription factor c-JUN by Ser63 phosphorylation [38]. phosphorylated c-JUN recruits interaction partners, such as c-FOS and FRA, to regulate the expression of several genes, including angiogenesis-related genes [53,54,55]. bZIP transcription factors form homodimers or heterodimers with other bZIP factors, mediated by the interaction among leucine-rich hydrophobic domains. However, recent studies have reported that they can also form complexes with other transcription factors without the bZIP domain. For example, the JUN:FOS heterodimer interacts with NFAT [56, 57], BACH1 interacts with NANOG, SOX2, and OCT [58], and PLAG1 interacts with USF2 [59]. Notably, c-JUN regulates tyrosine hydroxylase expression by forming a transcriptional complex indirectly with EGR1 [42]. In this study, using EMSA and pull-down assays, we showed that c-JUN co-localized with EGR1 on the VEGFR2 promoter. Furthermore, we confirmed that c-JUN could interact directly with EGR1 in the nucleus in vitro. Follow-up studies should be performed to obtain additional details on this mechanism; however, based on evidence from previous studies, the following could be hypothesized about the EGR1:c-JUN interaction. The consensus sequence TATA box was not detected in the VEGFR2 promoter. In a TATA-less gene, TBP can be recruited by other transcription factors to initiate transcription [60]. Meanwhile, it is known that c-JUN can recruit TBP [61]. ChIP-Seq data from the UCSC genome browser show that the EBS responsive to TNF-α lies in the TBP-binding region of the VEGFR2 promoter (Supplementary Fig. S4A). Based on these findings, we suggest that c-JUN interaction with EGR1 aids TBP recruitment and initiates transcription.
Additionally, gene regulation by chromatin remodeling may be involved in MSC differentiation. When MSCs were treated with the HDAC inhibitor Vorinostat (also known as suberoylanilide hydroxamic acid (SAHA)), VEGFR2 expression was activated without additional stimulation, suggesting that the VEGFR2 locus is present within a closed chromatin structure in an undifferentiated state (Supplementary Fig. S5A). The EGR1:c-JUN complex formed by TNF-α stimulation may also efficiently recruit co-activators that induce chromatin remodeling to the promoter region. Indeed, both EGR1 and c-JUN have been reported to interact with p300 acetyltransferases [62,63,64,65]. We also confirmed the presence of the c-JUN:p300 interaction (Supplementary Fig. S5B). Although the current study focused on VEGFR2 expression, it seems necessary to further confirm whether these putative complexes can modulate endothelial markers other than VEGFR2.
In summary, we demonstrated that TNF-α promotes the endothelial differentiation of BM-MSCs. Specifically, after TNF-α stimulation, EGR1 rapidly accumulates via the ERK and JNK pathways and forms a complex with c-JUN phosphorylated by JNK. The EGR1:c-JUN complex binds to the EBS in the VEGFR2 promoter, facilitating the endothelial differentiation of MSCs (Supplementary Fig. S6). Insight on the formation of this complex may provide a new perspective on the understanding of the tumor vasculature and indicate new antitumor targets.
Data availability
The data sets generated during the current study are available from the corresponding author upon reasonable request. The microarray data have been deposited in NCBI Gene Expression Omnibus (Accession number: GSE214834).
References
Forster JC, Harriss-phillips WM, Douglass MJ, Bezak E. A review of the development of tumor vasculature and its effects on the tumor microenvironment. Hypoxia. 2017;5:21–32.
Teleanu RI, Chircov C, Grumezescu AM, Teleanu DM. Tumor angiogenesis and anti-angiogenic strategies for cancer treatment. J Clin Med. 2020;9:84.
Ruoslahti E. Specialization of tumour vasculature. Nat Rev Cancer. 2002;2:83–90.
Lugano R, Ramachandran M, Dimberg A. Tumor angiogenesis: causes, consequences, challenges and opportunities. Cell Mol Life Sci. 2020;77:1745–70.
Rohde E, Malischnik C, Thaler D, Maierhofer T, Linkesch W, Lanzer G, et al. Blood monocytes mimic endothelial progenitor cells. Stem Cells. 2006;24:357–67.
Coukos G, Benencia F, Buckanovich RJ, Conejo-Garcia JR. The role of dendritic cell precursors in tumour vasculogenesis. Br J Cancer. 2005;92:1182–7.
Pittenger MF, Discher DE, Péault BM, Phinney DG, Hare JM, Caplan AI. Mesenchymal stem cell perspective: cell biology to clinical progress. NPJ Regen Med. 2019;4:22.
Satija NK, Singh VK, Verma YK, Gupta P, Sharma S, Afrin F, et al. Mesenchymal stem cell-based therapy: a new paradigm in regenerative medicine. J Cell Mol Med. 2009;13:4385–402.
Torsvik A, Bjerkvig R. Mesenchymal stem cell signaling in cancer progression. Cancer Treat Rev. 2013;39:180–8.
Hsu HS, Lin JH, Hsu TW, Su K, Wang CW, Yang KY, et al. Mesenchymal stem cells enhance lung cancer initiation through activation of IL-6/JAK2/STAT3 pathway. Lung Cancer. 2012;75:167–77.
Tsai KS, Yang SH, Lei YP, Tsai CC, Chen HW, Hsu CY, et al. Mesenchymal stem cells promote formation of colorectal tumors in mice. Gastroenterology. 2011;141:1046–56.
Mishra PJ, Mishra PJ, Humeniuk R, Medina DJ, Alexe G, Mesirov JP, et al. Carcinoma-associated fibroblast-like differentiation of human mesenchymal stem cells. Cancer Res. 2008;68:4331–9.
Yang KQ, Liu Y, Huang QH, Mo N, Zhang QY, Meng QG, et al. Bone marrow-derived mesenchymal stem cells induced by inflammatory cytokines produce angiogenetic factors and promote prostate cancer growth. BMC Cancer. 2017;17:1–10.
Yuan L, Sakamoto N, Song G, Sato M. High-level shear stress stimulates endothelial differentiation and VEGF secretion by human mesenchymal stem cells. Cell Mol Bioeng. 2013;6:220–9.
Lozito TP, Kuo CK, Taboas JM, Tuan RS. Human mesenchymal stem cells express vascular cell phenotypes upon interaction with endothelial cell matrix. J Cell Biochem. 2009;107:714–22.
Oswald J, Boxberger S, Jørgensen B, Bornhaeuser M, Ehninger G, Werner C. Mesenchymal stem cells can be differentiated into endothelial cells in vitro. Stem Cells. 2004;22:377–84.
Batlle R, Andrés E, Gonzalez L, Llonch E, Igea A, Gutierrez-Prat N, et al. Regulation of tumor angiogenesis and mesenchymal–endothelial transition by p38α through TGF-β and JNK signaling. Nat Commun. 2019;10:1–18.
Balkwill F. Tumour necrosis factor and cancer. Nat Rev Cancer. 2009;9:361–71.
Sethi G, Sung B, Aggarwal BB. TNF: a master switch for inflammation to cancer. Front Biosci. 2008;13:5094–107.
Kwon YW, Heo SC, Jeong GO, Yoon JW, Mo WM, Lee MJ, et al. Tumor necrosis factor-α-activated mesenchymal stem cells promote endothelial progenitor cell homing and angiogenesis. Biochim Biophys Acta Mol Basis Dis. 2013;1832:2136–44.
Huang H, Zhao N, Xu X, Xu Y, Li S, Zhang J, et al. Dose-specific effects of tumor necrosis factor alpha on osteogenic differentiation of mesenchymal stem cells. Cell Prolif. 2011;44:420–7.
Tetzlaff F, Fischer A. Human endothelial cell spheroid-based sprouting angiogenesis assay in collagen. Bio Protoc. 2018;8:e2995.
Xie Z, Bailey A, Kuleshov MV, Clarke DJB, Evangelista JE, Jenkins SL, et al. Gene set knowledge discovery with enrichr. Curr Protoc. 2021;1:e90.
Mootha VK, Lindgren CM, Eriksson K-F, Subramanian A, Sihag S, Lehar J, et al. PGC-1α-responsive genes involved in oxidative phosphorylation are coordinately downregulated in human diabetes. Nat Genet. 2003;34:267–73.
Subramanian A, Tamayo P, Mootha VK, Mukherjee S, Ebert BL, Gillette MA, et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc Natl Acad Sci USA. 2005;102:15545–50.
Stauffer W, Sheng H, Lim HN. EzColocalization: an ImageJ plugin for visualizing and measuring colocalization in cells and organisms. Sci Rep. 2018;8:15764.
Herrmann C, Avgousti D, Weitzman M. Differential salt fractionation of nuclei to analyze chromatin-associated proteins from cultured mammalian cells. Bio Protoc. 2017;7:1–13.
Yeo H, Ahn SS, Lee YH, Shin SY. Regulation of pro-opiomelanocortin (POMC) gene transcription by interleukin-31 via early growth response 1 (EGR-1) in HaCaT keratinocytes. Mol Biol Rep. 2020;47:5953–62.
Lee SL, Wang Y, Milbrandt J. Unimpaired macrophage differentiation and activation in mice lacking the zinc finger transcription factor NGFI-A (EGR1). Mol Cell Biol. 1996;16:4566–72.
Huang S, Xu L, Sun Y, Wu T, Wang K, Li G. An improved protocol for isolation and culture of mesenchymal stem cells from mouse bone marrow. J Orthop Transl. 2015;3:26–33.
Bouïs D, Hospers GAP, Meijer C, Molema G, Mulder NH. Endothelium in vitro: a review of human vascular endothelial cell lines for blood vessel-related research. Angiogenesis. 2001;4:91–102.
Janeczek Portalska K, Leferink A, Groen N, Fernandes H, Moroni L, van Blitterswijk C, et al. Endothelial differentiation of mesenchymal stromal cells. PLoS One. 2012;7:e46842.
Grimbacher B, Aicher WK, Peter HH, Eibel H. TNF-α induces the transcription factor Egr-1, pro-inflammatory cytokines and cell proliferation in human skin fibroblasts and synovial lining cells. Rheumatol Int. 1998;17:185–92.
Son SW, Min BW, Lim Y, Lee YH, Shin SY. Regulatory mechanism of TNFα autoregulation in HaCaT cells: The role of the transcription factor EGR-1. Biochem Biophys Res Commun. 2008;374:777–82.
Chaudhary LR, Cheng SL, Avioli LV. Induction of early growth response-1 gene by interleukin-1β and tumor necrosis factor-α in normal human bone marrow stromal and osteoblastic cells: regulation by a protein kinase C inhibitor. Mol Cell Biochem. 1996;156:69–77.
Wang B, Guo H, Yu H, Chen Y, Xu H, Zhao G. The role of the transcription factor EGR1 in cancer. Front Oncol. 2021;11:1–10.
Kim JH, Jung E, Choi J, Min DY, Lee YH, Shin SY. Leptin is a direct transcriptional target of EGR1 in human breast cancer cells. Mol Biol Rep. 2019;46:317–24.
Ip YT, Davis RJ. Signal transduction by the c-Jun N-terminal kinase (JNK) - From inflammation to development. Curr Opin Cell Biol. 1998;10:205–19.
Vleugel MM, Greijer AE, Bos R, van der Wall E, van Diest PJ. c-Jun activation is associated with proliferation and angiogenesis in invasive breast cancer. Hum Pathol. 2006;37:668–74.
Ma J, Zhang L, Han W, Shen T, Ma C, Liu Y, et al. Activation of JNK/c-Jun is required for the proliferation, survival, and angiogenesis induced by EET in pulmonary artery endothelial cells. J Lipid Res. 2012;53:1093–105.
Lee SP, Jun G, Yoon EJ, Park S, Yang CH. Inhibitory effect of methyl caffeate on Fos-Jun-DNA complex formation and suppression of cancer cell growth. Bull Korean Chem Soc. 2001;22:1131–5.
Nakashima A, Ota A, Sabban EL. Interactions between Egr1 and AP1 factors in regulation of tyrosine hydroxylase transcription. Brain Res Mol Brain Res. 2003;112:61–9.
Goel HL, Mercurio AM. VEGF targets the tumour cell. Nat Rev Cancer. 2013;13:871–82.
Beckermann BM, Kallifatidis G, Groth A, Frommhold D, Apel A, Mattern J, et al. VEGF expression by mesenchymal stem cells contributes to angiogenesis in pancreatic carcinoma. Br J Cancer. 2008;99:622–31.
Marcelo KL, Goldie LC, Hirschi KK. Regulation of endothelial cell differentiation and specification. Circ Res. 2013;112:1272–87.
Al-Lamki RS, Mayadas TN. TNF receptors: signaling pathways and contribution to renal dysfunction. Kidney Int. 2015;87:281–96.
Guo D, Dunbar JD, Yang CH, Pfeffer LM, Donner DB. Induction of Jak/STAT signaling by activation of the type 1 TNF receptor. J Immunol. 1998;160:2742–50.
Gashler A, Sukhatme VP. Early Growth Response Protein 1 (Egr-1): Prototype of a Zinc-finger family of transcription factors. Prog Nucleic Acid Res Mol Biol. 1995;50:191–224.
Li L, Ameri AH, Wang S, Jansson KH, Casey OM, Yang Q, et al. EGR1 regulates angiogenic and osteoclastogenic factors in prostate cancer and promotes metastasis. Oncogene. 2019;38:6241–55.
Wu SY, Rupaimoole R, Shen F, Pradeep S, Pecot CV, Ivan C, et al. A miR-192-EGR1-HOXB9 regulatory network controls the angiogenic switch in cancer. Nat Commun. 2016;7:1–14.
Seong Y, Moon J, Kim J. Egr1 mediated the neuronal differentiation induced by extremely low-frequency electromagnetic fields. Life Sci. 2014;102:16–27.
Guerquin MJ, Charvet B, Nourissat G, Havis E, Ronsin O, Bonnin MA, et al. Transcription factor EGR1 directs tendon differentiation and promotes tendon repair. J Clin Invest. 2013;123:3564–76.
Hess J, Angel P, Schorpp-Kistner M. AP-1 subunits: Quarrel and harmony among siblings. J Cell Sci. 2004;117:5965–73.
Vinson C, Myakishev M, Acharya A, Mir AA, Moll JR, Bonovich M. Classification of Human B-ZIP proteins based on dimerization properties. Mol Cell Biol. 2002;22:6321–35.
Hamik A, Wang B, Jain MK. Transcriptional regulators of angiogenesis. Arterioscler Thromb Vasc Biol. 2006;26:1936–47.
Morgunova E, Taipale J. Structural perspective of cooperative transcription factor binding. Curr Opin Struct Biol. 2017;47:1–8.
Chen L, Glover JN, Hogan PG, Rao A, Harrison SC. Structure of the DNA-binding domains from NFAT, Fos and Jun bound specifically to DNA. Nature. 1998;392:42–8.
Wei X, Guo J, Li Q, Jia Q, Jing Q, Li Y, et al. Bach1 regulates self-renewal and impedes mesendodermal differentiation of human embryonic stem cells. Sci Adv. 2019;5:eaau7887.
Belew MS, Bhatia S, Keyvani Chahi A, Rentas S, Draper JS, Hope KJ. PLAG1 and USF2 co-regulate expression of Musashi-2 in human hematopoietic stem and progenitor cells. Stem Cell Rep. 2018;10:1384–97.
Hernandez N. TBP, a universal eukaryotic transcription factor? Genes Dev. 1993;7:1291–308.
Franklin CC, McCulloch AV, Kraft AS. In vitro association between the Jun protein family and the general transcription factors, TBP and TFIIB. Biochem J. 1995;305:967–74.
Wang B, Chen J, Santiago FS, Janes M, Kavurma MM, Chong BH, et al. Phosphorylation and acetylation of histone H3 and autoregulation by early growth response 1 mediate interleukin 1β induction of early growth response 1 transcription. Arterioscler Thromb Vasc Biol. 2010;30:536–45.
Silverman ES, Du J, Williams AJ, Wadgaonkar R, Drazen JM, Collins T. cAMP-response-element-binding-protein-binding protein (CBP) and p300 are transcriptional co-activators of early growth response factor-1 (Egr-1). Biochem J. 1998;336:183–9.
Kim BK, Im JY, Han G, Lee WJ, Won KJ, Chung KS, et al. P300 cooperates with c-Jun and PARP-1 at the p300 binding site to activate RhoB transcription in NSC126188-mediated apoptosis. Biochim Biophys Acta Gene Regul Mech. 2014;1839:364–73.
Liu L, Guan H, Li Y, Ying Z, Wu J, Zhu X, et al. Astrocyte elevated gene 1 interacts with acetyltransferase p300 and c-Jun to promote tumor aggressiveness. Mol Cell Biol. 2017;37:e00456–16.
Acknowledgements
This paper was supported by the KU Research Professor Program of Konkuk University.
Funding
This work was supported by the National Research Foundation of Korea (NRF) grant funded by the Ministry of Science and ICT (2018R1A2B2004653 (YHL) and 2020R1A2C1005845 (SYS).
Author information
Authors and Affiliations
Contributions
Conceptualization, YHL and SYS; Methodology, EJ, SO, SSA, and HY; Investigation, EJ, SO, SSA, and HY; Formal Analysis, EJ, SSA, and HY; Writing – Original Draft, EJ; Writing – Review & Editing, YHL and SYS; Funding Acquisition, YHL and SYS; Project Administration and Supervision, SYS. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics statement
All animal experiments were approved by the Konkuk University Institutional Animal Care and Use Committee (IACUC) (approval number KU22041).
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by JP Medema
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jung, E., Ou, S., Ahn, S.S. et al. The JNK-EGR1 signaling axis promotes TNF-α-induced endothelial differentiation of human mesenchymal stem cells via VEGFR2 expression. Cell Death Differ 30, 356–368 (2023). https://doi.org/10.1038/s41418-022-01088-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-022-01088-8