Abstract
Autophagy targets cytoplasmic materials for degradation, and influences cell health. Alterations in Atg6/Beclin-1, a key regulator of autophagy, are associated with multiple diseases. While the role of Atg6 in autophagy regulation is heavily studied, the role of Atg6 in organism health and disease progression remains poorly understood. Here, we discover that loss of Atg6 in Drosophila results in various alterations to stress, metabolic and immune signaling pathways. We find that the increased levels of circulating blood cells and tumor-like masses in atg6 mutants vary depending on tissue-specific function of Atg6, with contributions from intestine and hematopoietic cells. These phenotypes are suppressed by decreased function of macrophage and inflammatory response receptors crq and drpr. Thus, these findings provide a basis for understanding how Atg6 systemically regulates cell health within multiple organs, and highlight the importance of Atg6 in inflammation to organismal health.
Similar content being viewed by others
Introduction
Macroautophagy (hereafter autophagy) is used by all cells to deliver cytoplasmic materials to lysosomes, and is induced by multiple stimuli, including nutrient deprivation, stress, and developmental signals [1]. Autophagy involves the formation of a double-membrane organelle, the autophagosome, and the fusion of autophagosomes with lysosomes leads to cargo degradation. Defects in this evolutionally conserved process are associated with various human disorders, including cancers, neurological disorders, and other diseases [2, 3]. Despite these implications, the mechanisms by which autophagy defects manifest as a decline in organismal health and diseases is not fully understood.
Elegant studies in yeast established the core autophagy machinery that is conserved from lower eukaryotes to humans. Many ATG genes were identified as part of this autophagy repertoire, which were subsequently shown to regulate various steps in the autophagy process, including induction, phagophore formation, autophagosome formation and termination by fusion with lysosomes [4,5,6]. One key autophagy protein is Atg6/Beclin-1, which forms a complex with the class III phosphatidylinositol 3-kinase, Vps34, Vps15 and Atg14 to form the Atg6-Vps34-Vps15-Atg14complex. This complex plays a vital role in autophagosome formation through Vps34 regulation of the formation of phosphatidylinositol 3-phosphate [7, 8]. This complex controls multiple vesicular trafficking pathways, including endocytosis, where it was originally identified as VPS30/ATG6 [9, 7, 10, 11].
Beclin-1 was initially thought to be an anti-apoptotic protein through its interaction with Bcl-2 [12]. Important work later discovered that this interaction with Bcl-2 serves to inhibit autophagy [13]. Meanwhile, several lines of evidence implicated Atg6/Beclin-1 as a critical mediator of health and disease. Beclin-1 is a tumor suppressor that is commonly deleted in breast cancer cell lines, and allelic loss of Beclin-1 in mice results in spontaneous carcinomas and lymphomas [14,15,16]. Patients with Alzheimer’s disease have decreased expression of Beclin-1, and heterozygous deletion of Beclin-1 in mice results in neurodegeneration and lysosomal dysregulation [17]. Conversely, mutations in the Bcl-2 binding domain of Beclin-1 in mice were shown to promote autophagy and enhance lifespan [18]. These studies illustrate the crucial role that Beclin-1/Atg6 plays in health and disease.
Atg6-deficient Drosophila display systemic health defects, similar to the phenotypes observed in Beclin-1 mice. For example, Atg6 loss-of-function Drosophila exhibit spontaneous formation of blood cell masses reminiscent of the lymphomas in Beclin-1 deficient mice [11, 14,15,16]. Alzheimer’s disease models in Drosophila demonstrated decreased Atg6 expression as flies aged [19]. Despite these observation in both mice and flies, the systemic influence of cellular and molecular changes in Beclin-1/Atg6 loss-of-function phenotypes remains poorly understood. Here we show that Atg6 loss-of-function Drosophila larvae possess drastic changes in stress and immune response, and metabolic pathways. Importantly, we demonstrate that hematopoietic and intestine cell lineages contribute to Atg6 mutant phenotypes, including organismal lethality. In addition, regulators of macrophage phagocytosis and inflammatory response are required for the blood-cell mass tumor-like phenotype observed in Atg6 mutants. Therefore, these studies reveal the important contribution of inflammation to the role of Atg6 in organismal health.
Results
Atg6 loss alters cell stress, metabolism, and ROS in cells and tissues
The complexity of atg6 and Beclin-1 mutant organismal phenotypes prompted us to consider genome wide changes in RNA levels in whole atg6 mutant Drosophila larvae. We used RNA sequencing (RNAseq) to compare changes in RNA levels between whole homozygous atg6(Δ) mutant larvae and age-matched w1118 control larvae (Table S1). Autophagy mitigates cell stress, and influences immunity and inflammation [3, 20, 21], processes that are associated with infection, wound healing, and cancer. Consistent with these roles for autophagy, we observed altered RNA levels of genes and pathways that have been associated with both cell stress and inflammation (Fig. 1A, B, Fig. S1A–D). Two Turandot-related genes, TotB and TotC that play a role in humoral stress response [22], were highly upregulated in atg6 mutants. Loss of Atg6 also resulted in upregulation of genes known to regulate immune response, such as BomS1, BomS2, BomS3, and Dso1 that are induced in response to bacterial and fungal infection [23, 24]. Two genes encoding enzymes important for metabolism and development, ugt36Bc and wunen-2 respectively [25, 26], were also highly upregulated in atg6 mutants. AICAR, an activator of AMPK that limits damage from sepsis [27,28,29,30], was also highly upregulated in intestine tissue.
A Bar chart displaying the genes related to stress response with the highest and lowest fold-changes in homozygous atg6 mutant larvae compared to w1118 control larvae. B A bar chart displaying the genes related to immunity and inflammation with the highest and lowest fold-changes in homozygous atg6 mutant larvae compared to w1118 control larvae. C atg6 (Δ)/(Δ) loss-of-function larval intestine cells (non-magenta) have increased GSTD1-GFP (green) compared to nearby atg6 (Δ/+) heterozygous control cells. D Quantification of mean GSTD1-GFP intensity in atg6 (Δ)/+ heterozygous control (n = 9) and atg6 (Δ)/atg6 (Δ) mutant (n = 9) intestine cells. E atg6 (Δ)/(Δ) loss-of-function larval intestine cells have increased TRE-GFP (green) compared to atg6 (Δ)/+ heterozygote control cells. F Quantification of the mean TRE-GFP intensity in atg6 (Δ)/(Δ) loss-of-function (n = 23) and atg6 (Δ)/+ heterozygote control (n = 27) larval intestine cells. Data in (D, F) were compared using two-tailed student t-test with error bars representing SEM. ***p < 0.001 and ****p < 0.0001. All experiments representative of at least 3 biological repeats.
To better understand the function of atg6 in cell stress, we investigated markers of stress signaling in mutant cells. We first analyzed the GSTD1 reporter, an oxidative response enzyme [31] that when combined with green fluorescent protein (GSTD1-GFP) serves as a reporter of oxidative stress [32, 33]. atg6(Δ) loss-of-function homozygous mutant larval intestine cells (non-magenta) exhibited higher amounts of GSTD1-GFP (green) compared to neighboring heterozygote control cells (magenta) (Fig. 1C, D), suggesting that atg6 mutant intestine cells harbor an activated oxidative stress response. The Jun N-terminal kinase (JNK) signaling pathway also plays an important role in Drosophila stress response [34], and the TRE-GFP reporter is a sensor of JNK activation [35]. As observed with GSTD1-GFP, homozygous atg6 (Δ) loss-of-function mutant intestine cells expressed higher amounts of TRE-GFP (green) than atg6 (Δ) /+ control intestine cells (Fig. 1E, F). Together, these results indicate that atg6 mutant cells display an upregulated stress response compared to controls.
Our analyses of RNA levels between whole age-matched w1118 control larvae and atg6(Δ) homozygous mutant larvae indicated significant changes in metabolism (Table S1, Fig. S1A–D), including genes that have been implicated in glucose and lipid metabolism (Fig. 2A). We next sought to directly investigate changes in metabolites in atg6 mutant tissues. Larval fat body, hemolymph, and intestines were isolated from homozygous atg6 (Δ) loss-of-function mutants and w1118 controls at the wandering larval stage and analyzed for changes in metabolites by liquid chromatography/mass spectrometry. In fat body, hemolymph, and intestines, Atg6-deficiency resulted in decreased amino acid metabolite levels compared to control tissues, while both Nα-acetyl glutamine and phosphoserine increased in hemolymph and intestines, but not fat body. Similarly, non-amino acid metabolite levels were reduced in Atg6-deficient fat body, while Atg6-deficient hemolymph and intestines exhibited both increases and decreases in non-amino acid metabolites (Fig. 2B, C, Fig. S2A–D, Fig. S3A–F, Table S2). Combined, these data indicate that atg6 is required for metabolic homeostasis in different tissues, and that these tissues possess distinct changes in metabolites.
A bar chart displaying the genes related to glucose and lipid metabolism with the highest and lowest fold-changes in homozygous atg6 mutant larvae compared to w1118 control larvae. B A bar chart displaying the 10 amino acids in the intestine with a p-value ≤ 0.1 and the highest fold change in homozygous atg6 mutant larvae compared with w1118 larvae. C A bar chart displaying the 10 non-amino acid metabolites in the intestine with a p-value ≤ 0.1 and the highest fold change in homozygous atg6 mutant larvae compared with w1118 larvae. D atg6 (Δ)/(Δ) loss-of-function larval intestine cells (non-magenta) have increased intestine-driven SREBP driven GFP (green) compared to nearby atg6 (Δ/+) heterozygous control cells. E Quantification of mean SREBP-GFP intensity in atg6 (Δ)/+ heterozygous control (n = 8) and atg6 (Δ)/atg6 (Δ) mutant (n = 8) intestine cells. F atg6 (Δ)/(Δ) loss-of-function larval intestine cells (non-magenta) have decreased Lipidtox puncta (green) staining compared to surrounding atg6 (Δ)/+ heterozygous control cells. G Quantification of Lipidtox puncta in atg6 (Δ)/ + heterozygous control (n = 14) and atg6 (Δ)/atg6 (Δ) mutant (n = 8) intestine cells. H atg6 (Δ)/(Δ) loss-of-function larval intestine cells (non-magenta) have decreased pS473 AKT puncta (green) compared nearby atg6 (Δ/+) heterozygous control cells. I Quantification of pS473 AKT puncta in atg6 (Δ)/+ heterozygous control (n = 9) and atg6 (Δ)/atg6 (Δ) mutant (n = 8) intestine cells. J Intestine lysates from atg6 (Δ)/(Δ) loss-of-function larvae (−/−) have increased phosphorylated AMPK (pAMPK) compared to heterozygous controls (+/−). Arrow indicates pAPMK-specific band. K Quantification of relative levels of pAMPK compared to Actin in atg6 (−/−) larval intestines compared to heterozygous controls (+/−). Representative of 4 independent experiments. Scale bars in D, F, H represent 40μm. Data in E, G, I, K were compared using two-tailed student t-test with error bars representing SEM. Abbreviations: **p < 0.01 and ****p < 0.0001. All experiments representative of at least 3 biological repeats.
Given these findings, we examined markers of lipid metabolism in atg6(Δ) homozygous mutant intestine cells. SREBP stimulates lipogenesis [36, 37], and we found that atg6 mutant intestine cells (non-magenta) possessed elevated levels of SREBP activity as demonstrated through a SREBP-dependent GFP reporter (green) [38] compared to neighboring heterozygous control cells (magenta) (Fig. 2D, E). We next analyzed atg6 mutant intestine cells for alteration in lipids by lipidtox staining. Consistent with altered SREBP reporter levels in atg6 mutant cells, lipidtox staining was significantly altered in cells with decreased atg6 function (Fig. 2F, G). In addition, levels of phosphorylated AKT, pAKT(S473) (green), were lower in homozygous atg6 mutant cells (non-magenta) compared to control cells (magenta) (Fig. 2H, I). Furthermore, atg6(Δ) mutant intestine lysates had elevated pAMPK levels compared to heterozygous atg6(Δ)/+ controls (Fig. 2J, K). Consistent with these results in individual mutant cells, the levels of GSTD1-GFP, SREBP activity, Lipidtox, and pAKT(S473) were different in whole animal atg6 (Δ) mutant intestines compared to heterozygous controls (Fig. S4A-D).
Changes in cellular metabolism and stress response could be linked to changes in the generation of reactive oxygen species (ROS). Freshly dissected atg6 loss-of-function mutant larval intestines and heterozygous controls were incubated with ROS markers. DHE and CellRox, markers that localize to the nucleus in the presence of high ROS stress, possessed increased nuclear localization in atg6 loss-of-function mutants compared to heterozygous controls (Fig. 3A–D). These results suggest that loss of Atg6 increased ROS production. To investigate mitochondrial ROS production, intestines were stained with MitoSox, a marker of mitochondrial ROS. An increase in mitochondrial MitoSox localization was detected in atg6 loss-of-function mutants compared to heterozygous controls (Fig. 3E, F).
A atg6 (Δ)/(Δ) loss-of-function larval intestine cells (bottom) have more nuclear localization of ROS-stress reporter DHE (red) than atg6 (Δ)/+ heterozygous control intestine cells (top). Nuclei were labeled with Hoescht (blue). B Quantification of DHE puncta that colocalized with Hoescht (nuclear-localizing) in atg6 (Δ)/(Δ) loss-of-function (n = 13) and atg6 (Δ)/+ heterozygous control (n = 12) larval intestine cells. C atg6 (Δ)/(Δ) loss-of-function larval intestine cells (bottom) have more nuclear localization of ROS-stress reporter CellRox-green (green) than atg6 (Δ)/+ heterozygous control intestine cells (top). Nuclei were labeled with Hoescht (blue). D Quantification of CellRox-green puncta that colocalized with Hoescht (nuclear-localizing) in atg6 (Δ)/(Δ) loss-of-function (n = 13) and atg6 (Δ)/+ heterozygous control (n = 12) larval intestine cells. E atg6 (Δ)/(Δ) loss-of-function larval intestine cells (bottom) have more mitochondrial ROS-stress reporter MitoSox puncta (red) than atg6 (Δ)/+ heterozygous control intestine cells (top). Mitochondria were labeled with Mito-Tracker green (green). Nuclei were labeled with Hoescht (blue). F Quantification of MitoSox puncta in atg6 (Δ)/(Δ) loss-of-function (n = 11) and atg6 (Δ)/+ heterozygous control (n = 10) larval intestine cells. Abbreviations: ****p < 0.0001. Scale bars in (A, C, E) represent 40μm. All p-values calculated from two-tailed student t test. All experiments representative of at least 3 biological repeats.
A failure to clear dysfunctional mitochondria through autophagy (mitophagy) could result in elevated production of mitochondrial ROS [39]. Therefore, atg6 mutants may possess increased mitochondrial ROS because of a decrease in mitophagy [40]. To test this possibility, we expressed the MitoQC construct in larval intestine cells and compared the differences in cells expressing atg6 RNAi and control luc RNAi. The MitoQC reporter construct utilizes mitochondria-localized tandem mCherry and GFP fluorescent proteins to label mitochondria outside of autolysosomes with both mCherry and GFP (yellow). By contrast, mitochondria inside autolysosomes are labeled only with red mCherry as the acidic environment of the autolysosome quenches the GFP signal [41, 42]. atg6 RNAi-expressing intestine cells had less mitolysosomes than luc RNAi-expressing control cells (Fig. S5A, B), consistent with the hypothesis that a defect in mitophagy could result in more ROS-producing mitochondria. Taken together, these results suggest a wide range of cellular changes that are associated with loss of Atg6, including altered metabolic, lipid, and ROS profiles.
Blood cell tumor-like masses are associated with the intestine of Atg6 mutants
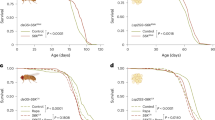
atg6 (Δ) mutant larvae possess tumor-like blood cell masses that accumulate around the intestine [11]. Unlike in mice [14,15,16], allelic loss of atg6 in Drosophila larvae does not result in either spontaneous tumor formation or alterations in animal survival (Fig. S6A–C). The close association of these blood cell masses with the intestines of most homozygous atg6 (Δ) mutant larvae, as well as significant changes in stress and immune signaling, prompted further investigation of this phenotype. atg6(Δ)/+ heterozygous control intestines have intact peripheral collagen as labeled by Vkg-GFP (green) (Fig. 4A). We visualized lamellocytes, specialized blood cells that participate in the immune response using msn9-mCherry [43]. Importantly, all control animals lack detectable msn9-mCherry (red) blood cell masses. By contrast, most homozygous atg6(Δ) mutant larval intestines have large msn9-mCherry labeled blood cell masses directly adjacent to areas of disrupted peripheral collagen on the intestine (Fig. 4A, B). These blood cell masses express Mmp1 (Fig. 4C), a proteinase that is involved in the breakdown of extracellular matrix that contributes to the invasiveness of tumor cells in Drosophila [44]. Transmission electron microscopy analyses of the blood cell mass and intestine interface shows cells that appear to be disrupting the continuity of the smooth muscle layer of the intestine (Fig. 4D, E). Together, these findings are consistent with blood cell masses in atg6 (Δ) mutants having tissue-invasive properties, which may be facilitated by expression of proteinases such as Mmp1.
A atg6 (Δ)/+ heterozygous control larvae (top) have intact intestinal architecture as shown by tubulin-driven Vkg-GFP (green) while atg6 (Δ)/(Δ) mutant larvae (bottom) have blood cell masses labeled by msn9-mCherry (red) that disrupt the intestine Vkg-GFP (green) architecture. B Quantification of the number of atg6 (Δ)/+ (n = 15) and atg6 (Δ)/(Δ) (n = 15) intestines that have blood masses that penetrate the intestine. Intestines were examined along the entire length. Intestines that had at least one msn9-mcherry labeled blood mass that completely bisected the intestine were considered intestines with mass attached. Fisher exact test was used to determine p-value. C msn9-mcherry (red) labeled blood cell masses that are present only in atg6 (Δ)/(Δ) co-localize with Mmp1 (green). D TEM section images of atg6 (Δ)/(Δ) blood cells [73] attached to intestine epithelium (Epi). The transition area between blood cells and intestine epithelium reveals disruption of the muscle layer (M). White arrows represent a region of muscle fiber separation due to a blood cell that extends toward the basement membrane of the intestine. Scale bar in the left TEM image represents 0.5 μm, middle TEM inset represents 10 nm, and right TEM inset represents 5 nm. E Number of blood cells associated with intestines that could be visualized by TEM in atg6 (Δ)/(Δ) (n = 134 TEM sections, from 6 processed intestines) and atg6 (Δ)/+ (n = 105 TEM sections, from 6 processed intestines) intestines. F Percentage of atg6 (Δ)/(Δ) larvae that died prematurely from developmental arrest compared to larvae that were able to reach adulthood (adult survival) expressing either GFP or ATG6-GFP driven by the NP1-GAL4 (intestines specific; n = 78 for GFP, n = 135 for ATG6-GFP), Dot-GAL4 (lymph gland and pericardial cell specific; n = 38 for GFP, n = 85 for ATG6-GFP), or Hml-GAL4 (blood cells; n = 124 for GFP, n = 78 for ATG6-GFP). G Percentage of larvae that died prematurely from developmental arrest in F that had blood cell masses upon observation with light microscope. Abbreviations: ****p < 0.0001 and ns, no significant differences between groups. Scale bars in (A, C) represent 40 μm. p-values in (A) calculated from two-tailed student t-test, p-values in (F, G) calculated from Fisher Exact test. All experiments representative of at least 3 biological repeats.
atg6 function in blood cells and intestines contributes to mutant lethality and blood cell tumors
To further characterize the role of different tissues on the atg6 mutant phenotype, we examined the influence of tissue-specific rescue of phenotypes by expression of Atg6 in mutant animal tissues. We reasoned that recruitment of blood cells to the intestine could be a result of either an increased intestinal stress response, dysregulated blood cell and inflammatory response, or both. To test these possibilities, we expressed Atg6 tagged with GFP (Atg6-GFP) using tissue-specific promoters. Both intestine- (NP1-GAL4) and hematopoietic lymph gland- (Dot-GAL4) specific Atg6-GFP expression partially rescued the developmental arrest phenotype in homozygous atg6 (Δ)/(Δ) mutant animals. However, rescue by expression of Atg6-GFP in the blood cells by an additional blood cell-specific promoter (Hml-GAL4) failed to suppress developmental arrest (Fig. 4F). In addition, we analyzed if tissue-specific expression of Atg6-GFP rescues mutant animal blood cell tumor-like mass formation. While expression of Atg6-GFP using the hematopoietic lineage promoters (Dot-GAL4 and Hml-GAL4) partially rescued blood cell mass formation, Atg6-GFP expression in the intestine (NP1-GAL4) failed to rescue the formation of blood cell masses (Fig. 4G). These results imply that Atg6-dependent mechanisms in both intestines and lymph glands promote survival beyond the late larval stage, while Atg6-expression in blood cells and lymph glands maintains blood cell homeostasis, preventing blood cell mass formation.
The Drosophila lymph gland is the Drosophila larval hematopoietic organ. In response to immune challenges and wound healing, larval lymph gland blood cell precursors proliferate, differentiate, and disseminate [45]. Therefore, we investigated the role of Atg6 in lymph glands because they appeared to play an important role in both lethality and blood cell mass atg6 mutant phenotypes. While lymph glands trended toward being enlarged in atg6 (Δ) mutants, this difference in lymph gland size was not statistically significant. However, Atg6-GFP expression in intestines also resulted in increased size of the atg6 (Δ) mutant lymph glands. Conversely, expression of Atg6-GFP in the lymph glands and blood cells of atg6 (Δ) mutants did not result in lymph gland enlargement (Fig. 5A–C). atg6 (Δ) mutant larvae possessed increased numbers of circulating blood cells compared to heterozygous controls [11]. Expression of Atg6-GFP in the hematopoietic system (Hml-GAL4 and Dot-GAL4) suppressed the atg6 (Δ) mutant increase in circulating blood cells. However, expression of Atg6-GFP in the intestine (NP1-Gal4) enhanced the increase in circulating blood cells (Fig. 5D). Together, these results demonstrate that Atg6 provides context-specific functions, and that the atg6 (Δ) mutant phenotypes can be suppressed depending on which organ retains Atg6 function.
A Representative fluorescent micrographs of larval Drosophila lymph glands from atg6 (Δ)/(Δ) (−/−) and atg6 (Δ)/(+) (−/+) animals expressing GFP in lymph gland and blood cells (Hml-Gal4). B Atg6 (Δ)/(Δ) animals in which Atg6-GFP was expressed in lymph gland and blood cells (Hml-Gal4), intestine (NP1-Gal4), and lymph gland and pericardial cells (Dot-Gal4). Bottom panels in (A, B) are color-thresholded to show the outlined area of the lymph gland. C-D Comparison of the average lymph gland area (C) or average number of circulating blood cells (D) per animal from heterozygous atg6 (Δ)/(+) and homozygous atg6 (Δ)/(Δ) mutant Drosophila larvae in addition to homozygous mutants in which Atg6-GFP is expressed under control of Hml-Gal4, NP1-Gal4, or Dot-Gal4. All scale bars represent 100 µm. Abbreviations: **p < 0.01, *** p < 0.001, and ****p < 0.0001; ns, no significant differences between groups. p-values in (C, D) calculated from one-way ANOVA analyses with Tukey post-hoc analysis. All experiments representative of 3 biological repeats.
Blood cell mass formation in atg6 mutants is mediated by macrophage receptors
Our data indicate that immune and inflammatory pathways might underlie the blood cell phenotypes in atg6 mutants. In addition, the Hml-GAL4 driver is expressed in most Drosophila hemocytes, which are the equivalent of mammalian neutrophils, macrophages, and dendritic cells [46], and Beclin-1 was shown to regulate neutrophil chemotaxis [47]. Therefore, we investigated if altered function of immune receptors influenced the atg6 (Δ) mutant blood cell phenotype. Crq and Drpr are two macrophage receptors that are involved in clearance of apoptotic cells [48, 49]. Larvae of atg6 (Δ) mutants possess blood cell aggregations that can be visualized as either black masses by brightfield microscopy or red msn9-mCherry masses by fluorescence microscopy, and no atg6 (Δ)/+ heterozygous control animals contain these masses (Fig. 6A). By contrast, we found that allelic loss of either crq or drpr resulted in partial suppression of blood cell mass formation in homozygous atg6 (Δ) mutant animals (Fig. 6A–C). These results suggest that the macrophage receptors Crq and Drpr facilitate blood cell aggregation in atg6 (Δ) mutant animals.
A atg6 (Δ/Δ) loss-of-function larvae (−/−) have enlarged blood cell masses (white arrows) visualized through brightfield (top panel) and lamellocyte marker msn9-mCherry expression (bottom panel) compared to heterozygous control atg6 (Δ/+) larvae (−/+). Allelic loss of either crq (+/−) or drpr (+/−) in an atg6 homozygous mutant genetic background results in suppression of the abnormal blood mass phenotype. msn9-mCherry blood cell masses are distinguishable from background msn9-mCherry expression in the mouth and cuticles, locations where these cells reside when immune and inflammatory responses are not active. B Quantification of the number of atg6 (+/−), atg6 (−/−), atg6 (−/−) crq (+/−), and atg6 (−/−) drpr (+/−) larvae with blood cell masses. Blood cell masses were defined as a dark mass greater than 0.1 mm2. C Quantification of the number of atg6 (+/−), atg6 (−/−), atg6 (−/−) crq (+/−), and atg6 (−/−) drpr (+/−) larvae with msn9-mCherry-positive masses greater than 1 mm2. Given background expression of mCherry along the outer cuticle and muscle layer, only mCherry-positive masses that were mobile and present medial to the cuticles were considered. Fisher Exact test p-values were calculated for (B, C). Abbreviations: ****p < 0.0001, **p < 0.01, *p < 0.05, and ns, no significant differences between groups. All experiments representative of 3 biological repeats.
Since expression of Atg6-GFP in blood cells rescued the blood cell mass formation phenotype in atg6 loss-of-function mutants, we next investigated if knockdown of crq and drpr using Dot-GAL4- and Hml-GAL4-driven expression of RNAi could suppress the atg6 blood cell mass phenotype. Knockdown of either crq or drpr in blood cells resulted in suppression of msn9-mCherry-positive (red) blood cell mass formation in atg6 loss-of-function mutants (Fig. 7A–C). Taken together, these results indicate that Crq and Drpr activity in lamellocytes is required for the blood cell mass phenotype in Atg6 deficient larvae.
A atg6 (−/−) larvae expressing Atg6-GFP or RNAi against either crq or drpr through the Dot-GAL4 (top) or Hml-GAL4 (bottom) drivers have less msn9-mCherry-positive blood cell masses compared to control atg6 (−/−) larvae expressing GFP. Quantification of the percentage of larvae with msn9-mCherry-positive blood cell masses in atg6 (−/−) larvae expressing GFP (Dot-GAL4, n = 38; Hml-GAL4 n = 51), Atg6-GFP (Dot-GAL4, n = 33; Hml-GAL4 n = 38), or RNAi targeting crq (Dot-GAL4, n = 35; Hml-GAL4 n = 34) or drpr (Dot-GAL4, n = 40; Hml-GAL4, n = 49) through the Dot-GAL4 (B) or Hml-GAL4 (C) promoters. Fisher Exact test p-values were calculated for (B, C). ****p < 0.0001 and ns, no significant differences between groups. All experiments representative of 3 biological repeats.
Discussion
Atg6/Beclin-1 has been implicated in the regulation of autophagy, endocytosis, apoptosis, and other cell processes [50, 51]. These diverse cellular roles suggest how Atg6/Beclin-1 loss may lead to health detriments and disease development [18, 52]. In mice, Beclin-1 loss enhanced tumorigenesis and invasive potential [15, 16]. Breast cancer cells deficient in Beclin-1 demonstrate enhanced tumorgenicity, potentially through modifications to cell membrane proteins like E-Cadherin [12, 53]. Like these mammalian models, Drosophila larvae lacking Atg6 possess cells with invasive properties in the form of blood cell masses. We demonstrate the complex and tissue-specific roles of Atg6 by showing Atg6 rescue specifically in either blood or intestine cells differentially modulates atg6 deficiency phenotypes. Importantly, we establish that the macrophage receptors Crq and Drpr contribute to atg6 mutant blood cell tumor-like aggregation, providing strong evidence of Atg6 function as a suppressor of inflammation.
Our studies of RNA levels in atg6 mutants revealed significant changes in multiple pathways. Stress response genes, including those in the Turandot and glutathione S-transferase family, were strongly upregulated with loss of atg6. These changes may reflect an increase in cellular stress, consistent with the metabolic roles of Atg6 [54]. Consistent with this possibility, a substantial difference in metabolite profiles were detected between atg6 mutants and controls in multiple tissue types. We detected an increase in specific metabolites in the atg6 mutant intestine and hemolymph, including phosphoserine and phosphoethanolamine. Interestingly, modulators of phosphoserine are known to affect cancer prognosis [55, 56]. In addition, previous work demonstrated that phosphoethanolamine accumulation protected breast cancer cells from glutamine deprivation and enhanced tumorigenicity [57]. RNAs of genes associated with ribosomes were also increased with loss of Atg6, and increased ribosome biogenesis is a hallmark of many cancer types [58, 59]. Metabolic changes may provide an enhanced environment for tumor cells to establish and spread, which may explain why atg6 mutants are so prone to developing invasive blood cell masses. It is tempting to speculate if haploinsufficient beclin-1 mice, which also share a predisposition for developing spontaneous tumors [14,15,16], have a similar metabolic profile that promotes tumorous growth. Future analyses of the metabolic changes observed in atg6 loss-of-function mutant Drosophila in haplo-deficient beclin-1 mice will help address this question.
Alterations in stress response pathways and imbalances in amino acid content can adversely affect the health of the cell, potentially explaining the increased ROS in atg6 loss-of-function mutant intestine cells. It is certainly possible that with loss of Atg6 function, cells are unable to clear unstable cargo and waste through autophagy, leading to increased ROS. The mitophagy deficiency in intestine cells with decreased Atg6 function is consistent with this hypothesis, as a failure to clear dysfunctional mitochondria could result in increased mitochondrial ROS. However, it is unknown if this principle applies to non-mitochondrial autophagic cargoes. Furthermore, we cannot rule out the possibility of a non-autophagy-related role of Atg6 in ROS production. Thus, these findings open new directions for how Atg6/Beclin-1 may function in tissue-specific manners to promote survival and stress-response during development, as well as suppress tumor formation.
Alterations in stress response markers, including GSTD1-GFP and TRE-GFP, in atg6 mutant intestine cells also support a role for Atg6 in suppression of stress. Furthermore, the upregulation of BomS1, BomS2, BomS3, Dso1, as well as genes involved in metabolism in atg6 mutants may be consistent with the role of Atg6 in inflammation and immunity [21]. Loss of Atg6 could also affect nutrient-sensing pathways because of the established role of autophagy in catabolism, which could explain decreased activation of AKT and increased levels of phosphorylated AMPK. AKT normally inhibits Atg6 activity in response to growth factors. The nutrient-deprived and sepsis-like characteristics of Atg6 loss-of-function mutants are counter-intuitive with AKT activity. Likewise, activation of AMPK serves to upregulate Atg6 activity, which would be expected to be increased if Atg6 activity is missing and the animal is deprived of nutrients [60]. Further investigation will determine if these phenotypes are due to either direct roles of Atg6 or if they reflect more global and indirect effects of Atg6 deficiency.
Previous work suggests that AKT activates SREBP [61]. Therefore, it is interesting that SREBP activity is increased in atg6 mutants when AKT activity is decreased. It is possible that loss of Atg6 uncouples this relationship by alteration of lipid content. While speculative, decreased lipid droplets in cells may stimulate SREBP activity, which through a negative feedback loop could inhibit AKT activity. Further studies, including investigation of the role of mTOR in these pathways, are necessary to elucidate the relationship between SREBP and AKT in the absence of Atg6.
Our characterization of the blood cell masses in atg6 mutant larvae demonstrates that Atg6-deficient blood cells display invasive phenotypes, including MMP1 positivity and disruption of intestine smooth muscle. Furthermore, atg6 mutant phenotypes, including developmental arrest, blood cell aggregation, lymph gland enlargement, and increased circulating blood cell numbers, were suppressed to varying degrees by expression of Atg6 in either intestine or blood cells. However, rescue expression in some tissue subsets failed to suppress mutant lethality or blood cell phenotypes, supporting a hypothesis that altered Atg6 function in a subset of tissues could drive tumorigenesis and disrupt systemic homeostasis. Future studies should evaluate how Atg6 loss in one tissue affects metabolites in tissues that retain Atg6 function.
Significantly, the blood cell immune receptors Crq and Drpr contributed to blood cell aggregation in atg6 mutants. It remains unclear if similar inflammatory mechanisms account for both the blood cell differences and tumors in Beclin-1 deficient mice [12, 53]. Mmp1 plays a role in cancer invasiveness, likely as a collagenase that facilitates tumor cell invasion through the basement membrane [62,63,64]. Although speculative, increased Mmp1 activity resulting from loss of Atg6/Beclin-1 and subsequent dysregulation of Crq and/or Drpr may explain the increase in invasive cell types in both Drosophila and mice. However, we cannot exclude that increased Mmp1 activity is simply an indirect byproduct of atg6 loss.
Interestingly, Beclin-1 provides a protective role against sepsis [65]. When compared to control animals, atg6 mutants display several molecular and metabolic hallmarks of sepsis. Upregulation of AICAR and AMP, as well as subsequent increase in pAMPK, is consistent with an increased response to septic shock [27,28,29]. As in human patients with sepsis, atg6 mutants display an acute inflammatory response, immune cell activation, and increased oxidative stress as evidenced by JNK activation, upregulation of immune response genes, and increased oxidative stress [66, 67]. Likewise, MMP1 activation is believed to contribute to sepsis development [68]. Taken together, these findings are consistent with a model whereby atg6 deficiency either results in increased susceptibility to sepsis or upregulates a sepsis-like response. These findings could explain the increased number of circulating blood cells, as increased circulating immune cells are characteristic of sepsis [66]. In addition, the fact that Atg6 expression specifically in immune cell lineages rescues multiple mutant phenotypes suggests that Atg6 regulates this process, at least in part, through its role in the immune system. However, we cannot exclude the possibility that loss of Atg6 contributes to this phenotype by enhancing blood cell proliferation through upregulation of cell proliferation regulators [69]. Additional studies of Atg6 function in sepsis development, along with other diseases including cancer, will be required to better understand how Beclin-1 suppresses these human conditions.
Materials and methods
Fly stocks
Flies were reared at 25 °C on standard cornmeal-molasses-agar media. For atg6(Δ) mutant cell clones, we crossed y w hsFlp;; FRT82B, Ubi-nlsGFP virgin females to FRT82B, atg6(Δ)/TM6B males. To obtain atg6(Δ) mutant cell clones combined with GSTD1-GFP and srebp-dependent GFP, UAS-GFP expression, we crossed y w hsFlp;; FRT82B, Ubi-mRFP virgin females to GSTD1-GFP, FRT82B, atg6(Δ)/TM6B and pP{GAL4-dSREBPg}, UAS-GFPFRT82B, atg6(Δ)/TM6B males, respectively. The following fly stocks used were from the Bloomington Drosophila Stock Center (BDSC): P{UAS-EGFP}34/TM3, Sb1 (5430), w1118; P{Hml-GAL4.G}6-4 (6396). The following fly stocks used for RNAi: TRiP.HMS01997 (crq RNAi) and drpr.dsRNA (Marc Freeman) (drpr RNAi). We thank the following researchers for mutant flies: pP{GAL4-dSREBPg}, UAS-GFP (Dirk Bohmann), TRE-GFP, (Dirk Bohmann) GSTD1-GFP (Dirk Bohmann), msn9-mCherry (Robert Schulz), Vkg-GFP (Andreas Bergmann). Additional fly stocks available in the Baehrecke lab include: atg6(Δ)/TM6B, UAS-atg6-GFP, Dot-GAL4, and Np1-GAL4.
Tissue-specific targeted metabolomics
Intestines and fatbody from female third instar larvae were dissected in cold water, centrifuged briefly, and snap-frozen in liquid nitrogen. 100ul of water was added to the samples and the samples were homogenized. Protein levels were measured using Bio-Rad Bradford assay and equal protein levels of each sample were used in subsequent steps. 80% methanol spiked with 1uM internal standard (MSK-A2-1.2, Cambridge Isotope Laboratories, Inc.) was added to the samples and vortexed at 4 °C. The samples were incubated at −20 °C for 1 h and centrifuged at 20,000 g for 5 min at 4 °C. The supernatant was transferred to a new tube vacuum dried for 2–3 h. Hemolymph was prepared by piercing third instar female larvae with tungsten needles and placing them in 0.5 ml Eppendorf tube contained within another 1.5 ml Eppendorf tube. The tubes were centrifuged 2X at 3000 g for 4 min at 4 °C. The hemolymph was collected and centrifuged at 3000 g for 3 min. Samples were transferred to a new tube and placed at 70 °C for 10 min and centrifuged at 13,000 g for 10 min. Samples were transferred to a new tube and 1 ml of 80% methanol containing 1uM internal standard (MSK-A2-1.2, Cambridge Isotope Laboratories, Inc.) was added to equal volumes/sample of hemolymph. Samples were then vacuum dried for 2–3 h. All samples were shipped on dry ice to the Metabolite Profiling Core Facility at the Whitehead Institute for Liquid Chromatography/Mass Spectrometry analyses using a Thermo Scientific Orbitrap mass spectrometer.
For statistical analysis and data interpretation, MetaboAnalyst 5.0 was used. For KEGG pathway analysis, the input data matrix, in csv format, contained all targeted metabolites and their concentrations. No normalization was performed and the reference metabolome used was the list of all targeted metabolites. Bar graphs were generated with Graphpad. For heatmap analysis, the input data in csv format contained targeted metabolites with a p-value <0.05. No normalization to the data was performed and autoscaling of features was performed. Euclidean distance measure and Ward clustering method was used.
RNA-Sequencing
RNA was extracted from third instar larvae using Qiagen RNEasy Kit. Next-Generation Sequencing was performed by Applied Biological Materials Inc., Canada. Quality check was performed by Qubit RNA Assay Kit and Agilent 2100 Bioanalyzer. PolyA mRNA was enriched and processed for cDNA using Illumina TruSeq Stranded mRNA library preparation kit. Paired-end sequencing was performed on the Illumina NextSeq. Illumina Casava 1.7 software was used for base calling. RNA seq reads were aligned against fruit fly reference genome dm6 using Hisat2 (version 2.1.0), FPKM values were determined by Cufflinks (v2.2.1) Differential expression was analyzed by limma with FPKM values as input. Downstream analyses were carried out using R and Bioconductor. All files were deposited into the Gene Expression Omnibus data repository under accession number GSE114790. KEGG pathway and Gene ontology analyses were performed by ShinyGO v0.741 (Brookings, SD, USA) using DE genes with an adjusted p-value <0.05.
Induction of mutant cell clones and quantification
Loss-of-function mutant cell clones in midgut were induced as previously described [70]. Adult flies were mated on low yeast food for 4 h at room temperature, and eggs were collected and heat shocked at 37 °C for 1 h. For 3rd instar larval midgut experiments, animals were dissected at 72 h. All puncta were quantitated as previously described [70].
Electron Microscopy
TEM was performed as previously described [71]. In brief, intestines were dissected in PBS 2 h after pupation, and then fixed in a solution of 2.5% glutaraldehyde and 2% paraformaldehyde in 0.1 M sodium cacodylate buffer, pH 7.4 (Electron Microscopy Sciences) overnight at 4 °C. Following osmium tetroxide treatment, samples were washed and stained en bloc in 1% aqueous uranyl acetate, dehydrated through a graded ethanol series, treated with propylene oxide, and infiltrated in SPI Pon Araldite for embedding. Ultrathin sections of the anterior region of the midgut were collected, and subsequently stained with uranyl acetate and lead citrate. Imaging was performed using a Phillips CM10 TEM.
ROS and Mitophagy Analysis
For labeling with immunofluorescent ROS markers, larvae were washed in 1X PBS and then dissected in cold Schneiders Media. Intestines were removed and immediately placed in respective ROS marker solutions dissolved in Schneiders media and Hoechst (Invitrogen #H3570, 200 ng/µL) for nuclear localization. For detection of ROS, CellRox Green Reagent (Invitrogen #C10444, 5 µM), DHE (Invitrogen # D11347, 30 µM), or MitoSox Red (Invitrogen # M36008, 5 µM) were used. Mitochondria were visualized in live tissue with MitoTracker Green FM (Invitrogen #M7514, 1 µM). Intestines were incubated in each of these solutions with gentle agitation, shielded from light before being briefly washed three times with cold Schneiders media. These intestines were then immediately mounted on coverslips with Schneider’s media for visualization for immunofluorescent microscopy. Only intestines visualized within 30 min of mounting were analyzed.
For mitophagy analysis by MitoQC, female flies expressing the MitoQC reporter were crossed with male flies with either the UAS-Luc RNAi (#31603) or UAS-Atg6 RNAi (#28060) transgenes. Intestines from wandering 3rd instar larvae progeny were dissected in cold PBS and immediately fixed in 4% paraformaldehyde for 30 min, and then incubated for 15 min in 0.1% PBSTx with Hoechst for nuclear visualization. These intestines were then mounted in Vectashield before visualization on immunofluorescent microscopy.
Immunolabeling and microscopy
Intestines were dissected in cold PBS before being placed in 4% paraformaldehyde solution for fixation at 4 °C overnight. Intestines were washed twice with PBS and then twice with PBS containing 0.1% Triton-X100 (PBSTx) before blocking in 5% normal goat serum for 90 min and incubation with primary antibody in 0.1% PBSTx overnight. Intestines were then stained with secondary antibody for 3 h before nuclei staining and mounting. For immunostaining, we used mouse monoclonal anti-MMP1 (1:1000, Developmental Hybridoma Bank, 3A6B4s) and rabbit polyclonal anti-phospho Drosophila AKT (Ser473) (1:1000, Cell Signaling, #4060P). Hoechst dye (Invitrogen #H3570, 10 ng/µL) was used to stain DNA. The following secondary antibodies (1:200): anti-mouse Alexa Fluor 488 (A-11029), anti-rabbit Alexa Fluor 488 (A-27034), anti-rabbit Alexa Fluor 546 (A-11035), anti-mouse Alexa Fluor 546 (A-11030), anti-guinea pig Alexa Fluor 546 (A-11074), anti-mouse Alexa Fluor 647 (A-S28181), anti-rabbit Alexa Fluor 647 (A-27040), anti-guinea pig Alexa Fluor 647 (A-21450), anti-rat Alexa 555 (A-48263) were used. All primary antibodies were incubated overnight and secondary antibodies were incubated for 2 h at room temperature. Tissues were mounted in VectaShield (Vector Laboratories, H-1200). For mCherry-Atg8a imaging, tissues were fixed in 4% paraformaldehyde for 20 min and stained with Hoescht before mounting. We imaged samples using a Zeiss LSM 700 confocal microscope equipped with a Plan-Apochromat 63×/1.40 Oil DIC M27 objective at room temperature (22 °C) using Zeiss Zen Software and a Nikon A1 HD25 confocal microscope equipped with a CFI Plan Apochromat 60×/1.4 Oil DIC objective at room temperature (22 °C) using NIS-Elements Viewer software. Images were deconvoluted using NIS-Elements Viewer software and processed using ImageJ.
We dissected lymph glands as described [72]. Briefly, we microdissected developmentally staged larvae from w1118 or atg6(Δ) animals in cold PBS by peeling away the larval cuticle from the posterior end, inverting the cuticle with minimal disruption of internal organs, and grasping the mouth hooks to separate the cuticle from the internal structures. Attached to the mouth hooks were several tissues, including the lymph gland and the ventral nerve cord. We removed any unneeded tissues except for the ventral nerve cord and lymph gland, using the ventral nerve cord as a handle for transferring the lymph gland to clean, ice cold PBS. We fixed the lymph glands in ice cold 4% paraformaldehyde for 30 min, washed 3 × 5 min in PBS, stained with DAPI to visualize nuclei, and mounted in Vectashield on slides with coverslips for immediate imaging. We imaged lymph glands at room temperature (22 °C) with a Zeiss AxioImager Z1 microscope equipped with a Plan-Apochromat 20×/0.8 M27 objective and an Axiocam MR3 camera using Axiovision Imaging software. Images were processed and lymph gland area was quantified using Image J (NIH, Bethesda, MD, USA).
We counted circulating bloods cells as described [72]. We collected developmentally staged larvae from w1118 or homozygous atg6(Δ) animals in dissecting dish wells filled with ice-cold PBS. For each larva, we placed 10 µl of 1x PBS in a microcentrifuge tube on ice and 10 µl 1x PBS on a clean dissecting pad under a stereo dissecting scope. After drying individual larvae on a Kim wipe, we transferred it to the PBS drop and carefully tore open the larval cuticle to expel circulating blood cells. Using a pipette, we transferred the cells to the microcentrifuge tube on ice. We counted cells within 30 min of collection using a hemocytometer and used Trypan blue to exclude dead cells.
Immunoblotting
For each sample, 5 whole larvae were lysed in 1x Laemli buffer and homogenized in solution using a plastic pestle for 30 s before being boiled at 99 °C for 6 min. Samples were run on 12% polyacrylamide gel, transferred onto 0.45 µm PVDF membranes (Millipore Sigma), and immuno-stained with antibodies using standard protocols. Primary antibodies used were rabbit anti-phospho AMPK (40H9) (1:1000), Cell Signaling, 2535T and mouse anti-Actin (1:1000, Proteintech). Full blots are shown (Fig. S7). No antibodies for total AMPK exist in Drosophila, so phospho-AMPK levels in lysates were normalized to Actin.
Statistical analysis
Image J (NIH, Bethesda, MD, USA) was used to quantify cell size, puncta, immunofluorescent intensity, co-localization, and protein level in western blot results. p-values were calculated using either a two-tailed unpaired t-test or one-way ANOVA followed by Dunnett’s or Bonferroni’s multiple comparisons from Graphpad Prism 5. No animals were excluded from statistical analyses, the experiments were not randomized, and the investigators were not blinded. All error bars are SEM.
Data availability
Please contact the Lead Contact, Dr. Eric Baehrecke, for additional information and requests for resources and reagents (eric.baehrecke@umassmed.edu).
References
Mizushima N, Komatsu M. Autophagy: renovation of cells and tissues. Cell 2011;147:728–41.
Mizushima N, Levine B. Autophagy in human diseases. N. Engl J Med. 2020;383:1564–76.
Klionsky DJ, Petroni G, Amaravadi RK, Baehrecke EH, Ballabio A, Boya P, et al. Autophagy in major human diseases. EMBO J. 2021;40:e108863.
Tsukada M, Ohsumi Y. Isolation and characterization of autophagy-defective mutants of Saccharomyces cerevisiae. FEBS Lett. 1993;333:169–74.
Yao Z, Delorme-Axford E, Backues SK, Klionsky DJ. Atg41/Icy2 regulates autophagosome formation. Autophagy 2015;11:2288–99.
Harding TM, Morano KA, Scott SV, Klionsky DJ. Isolation and characterization of yeast mutants in the cytoplasm to vacuole protein targeting pathway. J Cell Biol. 1995;131:591–602.
Funderburk SF, Wang QJ, Yue Z. The Beclin 1-VPS34 complex-at the crossroads of autophagy and beyond. Trends Cell Biol. 2010;20:355–62.
Burman C, Ktistakis NT. Regulation of autophagy by phosphatidylinositol 3-phosphate. FEBS Lett. 2010;584:1302–12.
Kruse KB, Brodsky JL, McCracken AA. Characterization of an ERAD gene as VPS30/ATG6 reveals two alternative and functionally distinct protein quality control pathways: one for soluble Z variant of human alpha-1 proteinase inhibitor (A1PiZ) and another for aggregates of A1PiZ. J Mol Cell Biol. 2006;17:203–12.
Kihara A, Kabeya Y, Ohsumi Y, Yoshimori T. Beclin-phosphatidylinositol 3-kinase complex functions at the trans-Golgi network. EMBO Rep. 2001;2:330–5.
Shravage BV, Hill JH, Powers CM, Wu L, Baehrecke EH. Atg6 is required for multiple vesicle trafficking pathways and hematopoiesis in Drosophila. Development 2013;140:1321–9.
Liang XH, Kleeman LK, Jiang HH, Gordon G, Goldman JE, Berry G, et al. Protection against fatal Sindbis virus encephalitis by beclin, a novel Bcl-2-interacting protein. J Virol. 1998;72:8586–96.
Pattingre S, Tassa A, Qu X, Garuti R, Liang XH, Mizushima N, et al. Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell 2005;122:927–39.
Aita VM, Liang XH, Murty VV, Pincus DL, Yu W, Cayanis E, et al. Cloning and genomic organization of beclin 1, a candidate tumor suppressor gene on chromosome 17q21. Genomics 1999;59:59–65.
Qu X, Yu J, Bhagat G, Furuya N, Hibshoosh H, Troxel A, et al. Promotion of tumorigenesis by heterozygous disruption of the beclin 1 autophagy gene. J Clin Invest. 2003;112:1809–20.
Yue Z, Jin S, Yang C, Levine AJ, Heintz N. Beclin 1. An autophagy gene essential for early embryonic development, is a haploinsufficient tumor suppressor. Proc Natl Acad Sci USA. 2003;100:15077–82.
Pickford F, Masliah E, Britschgi M, Lucin K, Narasimhan R, Jaeger PA, et al. The autophagy-related protein beclin 1 shows reduced expression in early Alzheimer disease and regulates amyloid beta accumulation in mice. J Clin Invest. 2008;118:2190–9.
Fernández ÁF, Sebti S, Wei Y, Zou Z, Shi M, McMillan KL, et al. Disruption of the beclin 1-BCL2 autophagy regulatory complex promotes longevity in mice. Nature 2018;558:136–40.
Haghi M, Masoudi R, Najibi SM. Distinctive alteration in the expression of autophagy genes in Drosophila models of amyloidopathy and tauopathy. Ups J Med Sci. 2020;125:265–73.
Levine B, Kroemer G. Autophagy in the pathogenesis of disease. Cell 2008;132:27–42.
Levine B, Mizushima N, Virgin HW. Autophagy in immunity and inflammation. Nature 2011;469:323–35.
Ekengren S, Hultmark D. A family of Turandot-related genes in the humoral stress response of Drosophila. Biochem Biophys Res Commun. 2001;284:998–1003.
Clemmons AW, Lindsay SA, Wasserman SA. An effector Peptide family required for Drosophila toll-mediated immunity. PLoS Pathog. 2015;11:e1004876.
Cohen LB, Lindsay SA, Xu Y, Lin SJH, Wasserman SA. The Daisho peptides mediate drosophila defense against a subset of filamentous Fungi. Front Immunol. 2020;11:9.
Ahn S-J, Marygold SJ. The UDP-Glycosyltransferase Family in Drosophila melanogaster: nomenclature update, gene expression and phylogenetic analysis. Front Physiol. 2021;12.
Hanyu-Nakamura K, Kobayashi S, Nakamura A. Germ cell-autonomous Wunen2 is required for germline development in Drosophila embryos. Development 2004;131:4545–53.
Kim J, Yang G, Kim Y, Kim J, Ha J. AMPK activators: mechanisms of action and physiological activities. Exp Mol Med. 2016;48:e224–e.
Huang J, Liu K, Zhu S, Xie M, Kang R, Cao L, et al. AMPK regulates immunometabolism in sepsis. Brain Behav Immun. 2018;72:89–100.
Mulchandani N, Yang W-L, Khan MM, Zhang F, Marambaud P, Nicastro J, et al. Stimulation of brain AMP-activated protein kinase attenuates inflammation and acute lung injury in sepsis. Mol Med. 2015;21:637–44.
Escobar DA, Botero-Quintero AM, Kautza BC, Luciano J, Loughran P, Darwiche S, et al. Adenosine monophosphate-activated protein kinase activation protects against sepsis-induced organ injury and inflammation. J Surg Res. 2015;194:262–72.
Sawicki R, Singh SP, Mondal AK, Benes H, Zimniak P. Cloning, expression and biochemical characterization of one Epsilon-class (GST-3) and ten Delta-class (GST-1) glutathione S-transferases from Drosophila melanogaster, and identification of additional nine members of the Epsilon class. Biochem J. 2003;370:661–9.
Landis G, Shen J, Tower J. Gene expression changes in response to aging compared to heat stress, oxidative stress and ionizing radiation in Drosophila melanogaster. Aging 2012;4:768–89.
Sykiotis GP, Bohmann D. Keap1/Nrf2 signaling regulates oxidative stress tolerance and lifespan in Drosophila. Dev Cell. 2008;14:76–85.
Davis RJ. Signal transduction by the JNK group of MAP kinases. Cell 2000;103:239–52.
Chatterjee N, Bohmann D. A versatile ΦC31 based reporter system for measuring AP-1 and Nrf2 signaling in Drosophila and in tissue culture. PLOS ONE. 2012;7:e34063.
Bertolio R, Napoletano F, Mano M, Maurer-Stroh S, Fantuz M, Zannini A, et al. Sterol regulatory element binding protein 1 couples mechanical cues and lipid metabolism. Nat Commun. 2019;10:1326.
Seegmiller AC, Dobrosotskaya I, Goldstein JL, Ho YK, Brown MS, Rawson RB. The SREBP pathway in Drosophila: regulation by palmitate, not sterols. Dev Cell. 2002;2:229–38.
Kunte AS, Matthews KA, Rawson RB. Fatty acid auxotrophy in Drosophila larvae lacking SREBP. Cell Metab. 2006;3:439–48.
Frank M, Duvezin-Caubet S, Koob S, Occhipinti A, Jagasia R, Petcherski A, et al. Mitophagy is triggered by mild oxidative stress in a mitochondrial fission dependent manner. Biochimica et Biophysica Acta. 2012;1823:2297–310.
Youle RJ, Narendra DP. Mechanisms of mitophagy. Nat Rev Mol Cell Biol. 2011;12:9–14.
McWilliams TG, Prescott AR, Allen GFG, Tamjar J, Munson MJ, Thomson C, et al. mito-QC illuminates mitophagy and mitochondrial architecture in vivo. J Cell Biol. 2016;214:333–45.
Lee JJ, Sanchez-Martinez A, Zarate AM, Benincá C, Mayor U, Clague MJ, et al. Basal mitophagy is widespread in Drosophila but minimally affected by loss of Pink1 or parkin. J Cell Biol. 2018;217:1613–22.
Tokusumi T, Sorrentino RP, Russell M, Ferrarese R, Govind S, Schulz RA. Characterization of a lamellocyte transcriptional enhancer located within the misshapen gene of Drosophila melanogaster. PLoS One. 2009;4:e6429.
Uhlirova M, Bohmann D. JNK- and Fos-regulated Mmp1 expression cooperates with Ras to induce invasive tumors in Drosophila. EMBO J. 2006;25:5294–304.
Letourneau M, Lapraz F, Sharma A, Vanzo N, Waltzer L, Crozatier M. Drosophila hematopoiesis under normal conditions and in response to immune stress. FEBS Lett. 2016;590:4034–51.
Williams MJ. Drosophila hemopoiesis and cellular immunity. J Immunol. 2007;178:4711–6.
Tan P, He L, Xing C, Mao J, Yu X, Zhu M, et al. Myeloid loss of Beclin 1 promotes PD-L1hi precursor B cell lymphoma development. J Clin Investig. 2020;129:5261–77.
Franc NC, Dimarcq JL, Lagueux M, Hoffmann J, Ezekowitz RA. Croquemort, a novel Drosophila hemocyte/macrophage receptor that recognizes apoptotic cells. Immunity 1996;4:431–43.
Manaka J, Kuraishi T, Shiratsuchi A, Nakai Y, Higashida H, Henson P, et al. Draper-mediated and phosphatidylserine-independent phagocytosis of apoptotic cells by Drosophila hemocytes/macrophages. J Biol Chem. 2004;279:48466–76.
Kang R, Zeh HJ, Lotze MT, Tang D. The Beclin 1 network regulates autophagy and apoptosis. Cell Death Differ. 2011;18:571–80.
Wirawan E, Lippens S, Vanden Berghe T, Romagnoli A, Fimia GM, Piacentini M, et al. Beclin1: a role in membrane dynamics and beyond. Autophagy 2012;8:6–17.
Jiang P, Mizushima N. Autophagy and human diseases. Cell Res. 2014;24:69–79.
Wijshake T, Zou Z, Chen B, Zhong L, Xiao G, Xie Y, et al. Tumor-suppressor function of Beclin 1 in breast cancer cells requires E-cadherin. Proc Natl Acad Sci USA. 2021;118:e2020478118.
Cao Y, Klionsky DJ. Physiological functions of Atg6/Beclin 1: a unique autophagy-related protein. Cell Res. 2007;17:839–49.
De Marchi T, Timmermans MA, Sieuwerts AM, Smid M, Look MP, Grebenchtchikov N, et al. Phosphoserine aminotransferase 1 is associated to poor outcome on tamoxifen therapy in recurrent breast cancer. Sci Rep. 2017;7:2099.
Park SM, Seo EH, Bae DH, Kim SS, Kim J, Lin W, et al. Phosphoserine phosphatase promotes lung cancer progression through the dephosphorylation of IRS-1 and a noncanonical L-serine-independent pathway. Mol Cells. 2019;42:604–16.
Osawa T, Shimamura T, Saito K, Hasegawa Y, Ishii N, Nishida M, et al. Phosphoethanolamine accumulation protects cancer cells under glutamine starvation through downregulation of PCYT2. Cell Rep. 2019;29:89–103.
Pelletier J, Thomas G, Volarević S. Ribosome biogenesis in cancer: new players and therapeutic avenues. Nat Rev Can. 2018;18:51–63.
Penzo M, Montanaro L, Treré D, Derenzini M. The ribosome biogenesis-cancer connection. Cells 2019;8:55.
Hill SM, Wrobel L, Rubinsztein DC. Post-translational modifications of Beclin 1 provide multiple strategies for autophagy regulation. Cell Death Differ. 2019;26:617–29.
Porstmann T, Santos CR, Griffiths B, Cully M, Wu M, Leevers S, et al. SREBP activity is regulated by mTORC1 and contributes to Akt-dependent cell growth. Cell Metab. 2008;8:224–36.
Iida J, McCarthy JB. Expression of collagenase-1 (MMP-1) promotes melanoma growth through the generation of active transforming growth factor-beta. Melanoma Res. 2007;17:205–13.
Liu H, Kato Y, Erzinger SA, Kiriakova GM, Qian Y, Palmieri D, et al. The role of MMP-1 in breast cancer growth and metastasis to the brain in a xenograft model. BMC Cancer. 2012;12:583.
Gabasa M, Radisky ES, Ikemori R, Bertolini G, Arshakyan M, Hockla A, et al. MMP1 drives tumor progression in large cell carcinoma of the lung through fibroblast senescence. Cancer Lett. 2021;507:1–12.
Sun Y, Yao X, Zhang QJ, Zhu M, Liu ZP, Ci B, et al. Beclin-1-dependent autophagy protects the heart during sepsis. Circulation 2018;138:2247–62.
Lewis AJ, Billiar TR, Rosengart MR. Biology and metabolism of sepsis: innate immunity, bioenergetics, and autophagy. Surg Infect. 2016;17:286–93.
Preau S, Vodovar D, Jung B, Lancel S, Zafrani L, Flatres A, et al. Energetic dysfunction in sepsis: a narrative review. Ann Intensive Care. 2021;11:104.
Vandenbroucke RE, Dejager L, Libert C. The first MMP in sepsis. EMBO Mol Med. 2011;3:367–9.
Ahn J-S, Ann E-J, Kim M-Y, Yoon J-H, Lee H-J, Jo E-H, et al. Autophagy negatively regulates tumor cell proliferation through phosphorylation dependent degradation of the Notch1 intracellular domain. Oncotarget 2016;7:79047–63.
Shen JL, Fortier TM, Wang R, Baehrecke EH. Vps13D functions in a Pink1-dependent and Parkin-independent mitophagy pathway. J Cell Biol. 2021;220:e202104073.
Shen JL, Fortier TM, Zhao YG, Wang R, Burmeister M, Baehrecke EH. Vmp1, Vps13D, and Marf/Mfn2 function in a conserved pathway to regulate mitochondria and ER contact in development and disease. Curr Biol. 2021;31:3028–39.
Reimels TA, Pfleger CM. Methods to Examine the Lymph Gland and Hemocytes in Drosophila Larvae. J Vis Exp. 2016;28:54544.
Tabchi S, Kassouf E, Rassy EE, Kourie HR, Martin J, Campeau M-P, et al. Management of stage III non–small cell lung cancer. Semin Oncol. 2017;44:163–77.
Acknowledgements
We thank the Baehrecke, Bohmann, Bergmann, and Freeman laboratories, the Bloomington Stock Center, the Developmental Hybridoma Bank, and the Electron Microscopy Core Facility at UMass Medical School for advice, flies, antibodies, cell lines, and technical support. We also thank Jun Yu and Julie Zhu from the UMass Bioinformatics Core for help with interpreting the RNA-seq data, and Caroline Lewis of the MIT Whitehead Institute Metabolomics Core for helping us with analyses and interpretation of metabolomics data. This work was supported by R01AG064892 to E.H.B. and F30CA239374 to J.L.S.
Author information
Authors and Affiliations
Contributions
JLS, JD, EA, and EHB designed experiments, JLS, JD, EA, and TMF performed experiments, JLS and EHB wrote the manuscript and all authors commented on it.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by M. Piacentini
Supplementary information
Rights and permissions
About this article
Cite this article
Shen, J.L., Doherty, J., Allen, E. et al. Atg6 promotes organismal health by suppression of cell stress and inflammation. Cell Death Differ 29, 2275–2287 (2022). https://doi.org/10.1038/s41418-022-01014-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-022-01014-y
This article is cited by
-
Macrophage polarization and metabolism in atherosclerosis
Cell Death & Disease (2023)