Abstract
Colorectal cancer (CRC) is the third leading cause of cancer-related deaths in the US. Understanding the mechanisms of CRC progression is essential to improve treatment. Mitochondria is the powerhouse for healthy cells. However, in tumor cells, less energy is produced by the mitochondria and metabolic reprogramming is an early hallmark of cancer. The metabolic differences between normal and cancer cells are being interrogated to uncover new therapeutic approaches. Mitochondria targeting PTEN-induced kinase 1 (PINK1) is a key regulator of mitophagy, the selective elimination of damaged mitochondria by autophagy. Defective mitophagy is increasingly associated with various diseases including CRC. However, a significant gap exists in our understanding of how PINK1-dependent mitophagy participates in the metabolic regulation of CRC. By mining Oncomine, we found that PINK1 expression was downregulated in human CRC tissues compared to normal colons. Moreover, disruption of PINK1 increased colon tumorigenesis in two colitis-associated CRC mouse models, suggesting that PINK1 functions as a tumor suppressor in CRC. PINK1 overexpression in murine colon tumor cells promoted mitophagy, decreased glycolysis and increased mitochondrial respiration potentially via activation of p53 signaling pathways. In contrast, PINK1 deletion decreased apoptosis, increased glycolysis, and reduced mitochondrial respiration and p53 signaling. Interestingly, PINK1 overexpression in vivo increased apoptotic cell death and suppressed colon tumor xenograft growth. Metabolomic analysis revealed that acetyl-CoA was significantly reduced in tumors with PINK1 overexpression, which was partly due to activation of the HIF-1α-pyruvate dehydrogenase (PDH) kinase 1 (PDHK1)-PDHE1α axis. Strikingly, treating mice with acetate increased acetyl-CoA levels and rescued PINK1-suppressed tumor growth. Importantly, PINK1 disruption simultaneously increased xenografted tumor growth and acetyl-CoA production. In conclusion, mitophagy protein PINK1 suppresses colon tumor growth by metabolic reprogramming and reducing acetyl-CoA production.
Similar content being viewed by others
Introduction
Colorectal cancer (CRC) is the third most common cancer as well as the third leading cause of cancer-related death in industrialized countries when data from men and women are combined [1]. Understanding the mechanisms of CRC development and progression is essential to improve treatment. Recently, we have found that intestine-specific overexpression of the novel mitochondrial ferrireductase six-transmembrane epithelial antigen of prostate 4 (STEAP4), which is highly increased in CRC, increases mitochondrial iron accumulation, oxidative stress and the susceptibility to colon tumors, while clinically used deferiprone chelates mitochondrial iron and reduces STEAP4-enhanced colon tumorigenesis in mice [2]. This study was the first to show that mitochondrial iron homeostasis is critical for colon carcinogenesis.
Generally, rapidly proliferating tumor cells have increased energy requirements in comparison to normal cells. Therefore, a more efficient energy source is needed for continual tumor growth. Though aerobic glycolysis, or the Warburg effect [3], has long since been accepted as an early hallmark of cancer cells [4], the latest cancer research confirms that mitochondrial respiration plays a critical role in tumorigenesis. Mitochondria act as the powerhouse for healthy cells as they consume nutrients such as glucose and generate most of the cell’s energy through oxidative phosphorylation [5]. However, in tumor cells, less energy is produced by the mitochondria and the nutrients are reorganized as building blocks to better support tumor cell growth [6]. The metabolic differences between normal and cancer cells are being interrogated to uncover potential new targeted therapeutic approaches [7]. Compared to cells from healthy colon tissues, mitochondrial number is increased in CRC cells, which leads to increased maximal respiratory rate [8]. However, metabolic reprogramming still occurs early at the adenoma stage and leads to induction of the Warburg effect in CRC [9].
Mitophagy, an essential mitochondrial quality control process in cells, removes damaged mitochondria through autophagic degradation [10, 11]. Defective mitophagy is implicated in various diseases, including neurodegenerative disorders and cancer [12]. One recent study showed that enhanced mitophagy caused by signal transducer and activator of transcription 3 inhibition in intestinal epithelial cells promotes anti-tumor immunity [13]. Another study showed that the expression level of mitochondria targeting PTEN-induced kinase 1 (PINK1), a critical mediator of mitophagy [14], was decreased in CRC compared to normal colorectal tissue and was regulated by MYC [9]. Moreover, PINK1 disruption promotes pancreatic tumorigenesis through increasing mitochondrial iron accumulation [15]. Furthermore, loss of PINK1 leads to increased mitochondrial reactive oxygen species (ROS), which activates hypoxia-inducible factor 1-alpha (HIF-1α) and reprograms glucose metabolism to sustain increased cell proliferation in mouse embryonic fibroblasts [16]. However, how PINK1 participates in metabolic reprogramming in CRC is not known.
In this study, we demonstrate that PINK1 acts as a tumor suppressor in CRC. Ablation of PINK1 increased colon tumorigenesis in two mouse models of colitis-associated CRC. In-depth analysis revealed that PINK1 is sufficient and required for suppressing tumor growth via metabolic reprogramming through activation of p53 signaling and HIF-1α/ pyruvate dehydrogenase (PDH) kinase 1 (PDHK1)/ PDHE1α pathways, which consequently impede acetyl-CoA production in tumor cells. Supplying tumors with acetyl-CoA from acetate rescued PINK1-suppressed tumor growth, demonstrating that acetyl-CoA serves as fuel for tumor growth.
Materials and methods
Animals
Animal studies were performed in accordance with the Institute of Laboratory Animal Resources guidelines, approved by the Institutional Animal Care and Use Committee (IACUC) at the University of New Mexico Health Sciences Center (Protocol# HSC-18-200699, 20-201060-HSC) and followed the National Institutes of Health guide for the care and use of laboratory animals (NIH Publications No. 8023, revised 1978). All mice were maintained in a standard cage on a 12-h light-dark cycle, in a temperature-controlled room and were allowed ad libitum access to a standard chow diet and water. Both male and female animals were used in this study. Pink1 knockout (Pink1−/−) mice were obtained from the Jackson laboratory (Stock No: 017946) and then were crossed with C57BL/6 mice to generate Pink1 heterozygous (Pink1+/−) mice, which were used as breeders. CDX2ERT2 ApcF/+ mice were generated as described previously [17], and were crossed with Pink1−/− mice to generate Pink1+/−; CDX2ERT2 ApcF/+ mice, which were then used as breeders. For the azoxymethane (AOM) /dextran sulfate sodium (DSS) induced colitis-associated CRC model, Pink1−/− and littermate control mice were intraperitoneally (i.p.) injected with AOM (10 mg/kg body weight) or saline at 6- to 8-week of age. It should be noted that no dysplasia was observed in mice receiving a single AOM dose, suggesting that the dose of carcinogen used was insufficient to induce colorectal tumors in the absence of inflammation. One day following the AOM injection mice received water with 1.5% DSS for 7 days (inflammatory phase). Then the mice were placed on regular drinking water for 14 days (recovery phase). Mice were subjected to two more inflammatory and recovery cycles to induce tumor formation and subsequently euthanized 2 weeks following the final recovery phase. For the colitis-associated CRC model under the Apc gene deletion setting, 6- to 8-week-old Pink1−/−; CDX2ERT2 ApcF/+ mice and littermate controls were treated with 100 mg/kg tamoxifen for 3 days by i.p. injection and then 7 days later received water with 1.5% DSS for 7 days, followed by regular drinking water for 28 days before euthanization. For the subcutaneous xenograft study, to test the role of PINK1 and metabolic rewiring under immune-intact conditions, murine MC38 cells with PINK1 overexpression or ablation were trypsinized and resuspended in sterile 1× phosphate-buffered saline (PBS). Cells were counted and diluted with 1× PBS to a concentration of 1 × 107 cells/mL, and then 100 μL containing 1 × 106 total cells were injected into the flanks of C57BL/6 mice. Two weeks later, mice were sacrificed and tumors were collected. For acetate treatment, mice were i.p. injected with 500 mg/kg sodium acetate weekly for three weeks. Proton linked monocarboxylate transporters (MCTs) and sodium coupled monocarboxylate transporters (SMCTs) are responsible for acetate transport across the cell membrane in mammals [18, 19]. Acetate then crosses the mitochondrial inner membrane through passive diffusion or via an unknown mitochondrial carrier [20, 21].
Real-time qPCR analysis
RNA was extracted using an IBI reagent Kit (IB47602, IBI Scientific, Dubuque, IA). qPCR was performed on a LightCycler 480 instrument (Roche Diagnostics, Indianapolis, IN). The mRNA level was measured by qPCR (the primers are listed in Supplementary Table 1). The gene levels were normalized by 18S. Expression level changes were calculated by comparing to the control.
Histology and immunofluorescence staining
Colon tissues were removed and washed with 1x PBS, and fixed with 10% formalin at room temperature overnight and embedded in paraffin. Tissue sections (5 μm) were deparaffinized in xylene and rehydrated in ethanol gradient. Histologic analysis was done on hematoxylin and eosin (H&E)-stained paraffin sections and microscopically analyzed by a gastrointestinal pathologist in a blinded manner. Immunofluorescence analysis was performed with primary antibodies for Ki-67 (12202T) and cleaved Caspase-3 (CC3, 9664) from Cell Signaling Technology (Danvers, MA) at a dilution of 1:100. Secondary antibodies were used at 1:200 for staining. 4′,6-diamidino-2-phenylindole (DAPI) was used for nuclei staining. The images were taken and analyzed under a fluorescence microscope (EVOS, Thermo Fisher Scientific).
Cell culture
Murine MC38 syngeneic CRC cells were a kind gift from Dr. Weiping Zou at University of Michigan and the authentication of this cell line was performed by the ATCC using mouse short tandem repeat profiling (137XV). These cells were maintained at 37 °C in 5% CO2 and cultured in Dulbecco’s modified Eagle medium supplemented with 10% fetal bovine serum and 1% penicillin and streptomycin (VWR, Radnor, PA). To establish MC38 cells with stable PINK1 overexpression, MC38 cells were transfected with a pLenti6-DEST PINK1-V5 WT plasmid obtained as a gift from Mark Cookson (Addgene Plasmid #13320) and then selected with blasticidin at 0.5–1 μg/mL. To establish MC38 cells with stable PINK1 knockout (sgPINK1) or control (sgEV), MC38 cells were infected with lentivirus targeting mouse PINK1 (CRISPRLV MMPD0000074951, CCAAGCCCGGCCCGCTGTT, 1 × 106 VP/mL, Lentivirus Backbone: (LV01) U6-gRNA: ef1a-puro-2A-Cas9-2A-tGFP, Sigma all-in-one lentivirus) or CRISPRLV NegativeControl1 (CGCGATAGCGCGAATATATT) and then selected with 1 μg/mL puromycin. pLenti6.3/V5-DEST TIGAR (HsCD00943600) and pLenti6.3/V5-DEST SCO2 (HsCD00938914) were obtained from DNASU Plasmid Repository (Arizona State University, Tempe, AZ), transfected into MC38 sgPINK1 and sgEV cells, and then selected with blasticidin at 0.5–1 μg/mL.
3-(4,5-dimethylthiazol-2-Yl)-2,5-diphenyltetrazolium bromide (MTT) cell proliferation assay
MC38 cells with PINK1 overexpression or disruption were seeded at a concentration of 5 × 104 cells/mL in 24-well plates. 125 μL 5 mg/mL MTT (Sigma, St Louis, MO) was added to each well and plates were incubated for 30 min. Dimethyl sulfoxide was added and absorbance was measured at 570 nm using a SpectraMax M2 Microplate Reader (Molecular Devices, Radnor, PA).
Cellular bioenergetic profile
MC38 cells were seeded at a total number of 4 × 104 cells/well 24 h prior to the beginning of the experiment. The extracellular acidification rate (ECAR) was monitored in a Seahorse XF24 instrument using the standard glycolysis stress test protocol of 3 min mix, 2 min wait, and 3 min measure. For the glycolysis stress test, cells were challenged with glucose (10 mM), oligomycin (1 µM), and 2-deoxyglucose (50 mM), allowing 3 measurements after each injection. For the oxygen consumption ratio (OCR) measurements, cells were stimulated with oligomycin (1 µM), FCCP (1 µM) and antimycin A (0.5 µM), also allowing 3 measurements after each injection. For the normalization of respiration to protein content, cells were lysed in radioimmunoprecipitation assay (RIPA) buffer and protein concentrations were measured using the Pierce BCA kit (5000006, Bio-Rad, Hercules, CA).
Western blot analysis
Tumor tissues were lysed with RIPA buffer and incubated on ice for 30 min. After incubation, cell extracts were centrifuged for 10 min at 4 °C. The supernatant was collected for protein concentration quantification using the Bradford assay. Equal amounts (30–50 μg) of protein were loaded for sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE), along with molecular weight markers. The gels were run for 1 h 30 min at 120 V. The proteins inside the gels were transferred onto nitrocellulose membranes for 1 h at 100 V using the wet transfer method as previously described [22]. The membranes were blocked with 3% milk for 1 h. Primary antibodies against p62 (sc-10117), p53 (sc-6243), PDHK1(sc-293160), Dynamin-related protein 1 (DRP1, sc-271583), TP53-induced glycolysis and apoptosis regulator (TIGAR, sc-166290), p21 (sc-6246), p38 (sc-7972), p-p38 (sc-166182), HIF1-α (sc-10790), succinate dehydrogenase complex flavoprotein subunit A (SDHA, sc-390381), SDHB (sc-271548), damage-regulated autophagy modulator (DRAM, sc-81713), translocase of outer mitochondrial membrane 20 (TOM20, sc-17764), actin (sc-8432) and glyceraldehyde 3-phosphate dehydrogenase (GAPDH, sc-47724) were from Santa Cruz Biotechnology (Dallas, TX). Primary antibodies against ATG16L1 (8089), LC3A/B (12741), LC3B (2775), p-P53 (9284), p-PDHK1 (3062T), Pyruvate dehydrogenase E1, alpha (PDHE1a, 3205T) and p-PDHE1a (31866) were from Cell Signaling Technology (Danvers, MA). V5 (14440-1-AP) synthesis of cytochrome c oxidase 2 (SCO2, 21223-1-AP), and p53 upregulated modulator of apoptosis (PUMA, 55120-1-AP) were from Proteintech (Rosemont, IL). Secondary antibodies against rabbit (7074S) or mouse IgG (7076S) were from Cell Signaling Technology (Danvers, MA).
Mitophagy assay with confocal imaging
We acquired a fluorescent mitophagy reporter plasmid pCLBW cox8-EGFP-mCherry as a gift from Dr. David Chan (Addgene, #78520) [23], and transiently transfected it into MC38 PINK1 overexpression cells and their parental controls using DreamFect™ Gold Transfection Reagent (DG81000, OZ Biosciences, San Diego, CA). We plated 2000 cells into each chamber of the 8-well Nunc™ Lab-Tek™ II Chambered Coverglass (155409, Thermo Scientific, Waltham, MA). GFP and mCherry fluorescent signals were visualized using a Leica TCS SP8 Confocal imaging system (Leica Microsystems Inc., Buffalo Grove, IL) and images were collected using the LAS X life science Microscope software platform.
ImageStream analysis
pCLBW cox8-EGFP-mCherry plasmid were transiently transfected into MC38 PINK1 overexpressing cells and their parental controls. After 24 h, cells were trypsinized to prepare single cell suspensions. Single cell images were acquired with ImageStreamxMark II (Amnis). Collected data were analyzed with IDEAS 5.0 software (Amnis). Cells were gated for single cells with the area and aspect ratio and for focused cells with the Gradient RMS feature.
Caspase-3/7 activity assay
We seeded 5000 MC38 PINK1 overexpressing and knockout cells and their parental controls into a white-wall 96-well plate. Background luminescence was measured using a BioTek Synergy HTX Multi-Mode Microplate Reader (BioTek Instruments, Inc., Winooski, VT) and subtracted from blank wells before seeding. Caspase-3/7 activity was measured by a Caspase-Glo3/7 Assay (Cat No: G8090, Promega) following the manufacturer’s protocol.
Metabolomic analysis
Targeted quantitative analysis was performed on 10 samples of tumors from MC38 with or without PINK1 overexpression at Human Metabolome America Technologies (HMT), Inc., (Boston, MA), using capillary electrophoresis mass spectrometry (Agilent CE-TOFMS system and Agilent CE-QqQMS) in the cation and anion analysis modes for analyzing cationic and anionic metabolites, respectively. A total of 116 metabolites (54 and 62 metabolites in the cation and anion mode, respectively) involved in glycolysis, pentose phosphate pathway, tricarboxylic acid (TCA) cycle, urea cycle, and polyamine, creatine, purine, glutathione, nicotinamide, choline, and amino acid metabolisms were annotated based on the HMT metabolite database. The samples were mixed with 750 μL of 50% acetonitrile in water (v/v) containing internal standards (20 μM) and homogenized (3500 rpm, 60 s × 5 times). The same amount of 50% acetonitrile in water (v/v) was then added and centrifuged. The supernatant (400 μL) was then filtrated through 5-kDa cut-off filter (ULTRAFREE-MC-PLHCC, Human Metabolome Technologies, Yamagata, Japan) to remove macromolecules. The filtrate was centrifugally concentrated and resuspended in 50 μL of ultrapure water immediately before the measurement.
Measurement of acetyl-CoA levels
The levels of acetyl-CoA in xenograft tumors were determined by an acetyl-CoA kit (K317-100, Biovision, Milpitas, CA) according to the manufacturer’s instructions. Briefly, tumor tissues were sonicated and the lysis was collected. Following the deproteinizing protocol, fluorescence was measured at excitation and emission wavelengths of 535 nm and 587 nm, respectively. The acetyl-CoA concentration in tissue samples were calculated based on the measured standard curves.
Statistical analysis
Data are expressed as mean ± SD with individual data points. We excluded any data point that was more than 2 standard deviation as an outlier. p values were calculated by independent two-sided t-test, paired t-test, one-way and two-way analysis of variance (ANOVA) as indicated. p < 0.05 was considered significant.
Results
PINK1 deficiency promotes tumor development in two colitis-associated CRC mouse models
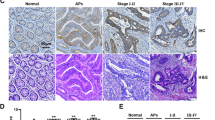
By mining the gene expression database Oncomine [24], we found that PINK1 expression is significantly downregulated in human colon tumors compared to adjacent normal colon tissues in The Cancer Genome Atlas (TCGA) cohort samples (Fig. 1A), which is consistent with a previous report [9]. To test the hypothesis that PINK1 is a colon tumor suppressor, we first confirmed that the gene expression of Pink1 was dose-dependently reduced in the colon tissues of wild-type (Pink1+/+), Pink1+/− and Pink1−/− mice by real-time qPCR analysis (Fig. 1B). We then treated these mice with a classical AOM/DSS protocol to induce colitis-associated CRC. Under macroscopy, we found that Pink1−/− mice developed more and bigger tumors and most of them were located in the proximal colon, whereas the tumors from Pink1+/− and Pink1+/+ mice were located in the distal colon (Fig. 1C). Quantitative analysis confirmed that Pink1−/− mice had significantly increased average tumor numbers and tumor burden than Pink1+/+ mice (Fig. 1D). Histological examination indicates that Pink1−/− and Pink1+/− mice had more severe inflammatory cell infiltration than Pink1+/+ mice (Fig. 1E). We also confirmed that the gene expression of Pink1 was dose-dependently reduced in both normal and tumor colon tissues of Pink1+/+, Pink1+/− and Pink1−/− mice (Fig. S1).
A PINK1 mRNA expression is decreased in human CRC patient tumor tissues (n = 102) compared to normal tissues (n = 19) from the TCGA database. B qPCR analysis in untreated colon tissues from Pink1−/− mice (n = 6), Pink1+/− mice (n = 5) and their wild-type littermates (n = 4). (C) Representative macroscopy images, (D) tumor number and tumor burden, and (E) H&E staining for Pink1−/− mice (n = 11), Pink1+/− mice (n = 7) and their wild-type littermates (n = 4) treated with an AOM/DSS colon tumor protocol (Bar = 400 μm). *p < 0.05, **p < 0.01, ***p < 0.001. Unpaired t-test. One-way ANOVA followed by Dunnett’s multiple comparisons test.
Though the AOM/DSS model is a widely accepted mouse model of colitis-associated CRC, recently it was criticized that this model may not reflect human colon tumorigenesis at the genomic level because some highly mutated genes in human CRC, such as APC, p53, KRAS and PIK3CA, were rarely identified in this model [25]. Thus, we also tested the effect of PINK1 deficiency on colon tumorigenesis using a recently developed CDX2ERT2 ApcF/+ mouse model [17]. We found that after treating with tamoxifen and DSS, Pink1−/− mice lost more body weight (Fig. S2A), had shorter colon length (Fig. S2B), and developed more and bigger tumors under macroscopy (Fig. 2A, B). The average tumor number (Fig. 2C), tumor burden (Fig. 2D), as well as the average tumor number at size of 1–2 mm were increased after PINK1 deletion (Fig. S2C). However, the average tumor size was not changed in Pink1−/− mice (Fig. S2D). Histological H&E staining showed that the percentage of high-grade dysplasia involved colon tissues was significantly increased in the Pink1−/− mice (Fig. 2E, F). Furthermore, adenoma was found in 2 out of 4 examined Pink1−/− mice, 1 out of 3 examined Pink1+/− mice, and 0 out of 3 examined Pink1+/+ mice, respectively. The cell proliferation marker Ki-67 was significantly increased in Pink1−/− mice compared with Pink1+/+ mice, whereas the cell apoptosis marker CC3 was not significantly changed (Fig. 2G–J). qPCR analysis confirmed again that the gene expression of Pink1 was significantly reduced in both normal and tumor colon tissues of Pink1−/− mice compared to Pink1+/+ mice (Fig. S2E). Interestingly, several proinflammatory cytokines including TNFα, IL-1β, IL-6 and IFN-β were significantly increased in Pink1−/− mice (Fig. S2F–I). Together, these data suggest that PINK1 is a tumor suppressor in CRC.
Representative images for (A) dissected colon tissues, (B) colon tumors under macroscope, (C) tumor number, (D) tumor burden, (E) H&E staining (Bar = 400 μm), (F) histological score, immunofluorescent staining and quantification of (G, H) Ki67 and (I, J) cleaved caspase 3 (CC3) (n = 3) for colon tumor tissues from Pink1−/−; CDX2ERT2 ApcF/+ mice (n = 10), Pink1+/−; CDX2ERT2 ApcF/+ mice (n = 7) and their littermates (n = 5) (Bar = 200 μm). *p < 0.05, **p < 0.01, NS not significant. One-way ANOVA followed by Dunnett’s multiple comparisons test.
PINK1 is critical for cell death in vitro
To further investigate the function of epithelial PINK1, we generated murine colon tumor MC38 cells with stable overexpression of human PINK1 and PINK1 knockout cells (sgPINK1). qPCR analysis confirmed that PINK1 was successfully overexpressed (Fig. 3A) or decreased (Fig. 3B). As PINK1 deficiency leads to increased tumor growth and it is a mitophagy regulating gene, we hypothesized that PINK1 might regulate cell proliferation and/or cell death. Firstly, we performed an MTT cell proliferation assay. Our results showed that PINK1 overexpression suppressed, whereas PINK1 deficiency increased cell growth in vitro at 48 h and 72 h (Fig. 3C, D). Western blot analysis demonstrated that several autophagy related genes including p62 and ATG16L, and the ratio of LC3II/LC3I were significantly increased by PINK1 overexpression, whereas PINK1 deletion cells exhibited significantly decreased ATG16L (Fig. 3E, F). We also validated that PINK1 itself and the vector tag V5, but not the mitochondrial fission protein DRP1, were increased by PINK1 overexpression (Fig. S3A). To test whether PINK1 overexpression leads to increased mitophagy, we transfected a fluorescent mitophagy reporter plasmid pCLBW cox8-EGFP-mCherry into MC38 PINK1 overexpressing cells and their parental controls. The cox8-EGFP-mCherry retroviral vector consists of the cox8 mitochondrial targeting sequence placed N-terminal to an EGFP-mCherry fusion [23]. Normal mitochondria are yellow, having both green and red fluorescence in the matrix, whereas mitochondria within acidic compartments show red-only fluorescence, due to the selective sensitivity of EGFP fluorescence to low pH [23]. We observed an increased ratio of red to green fluorescence signaling in MC38 PINK1 overexpressing cells compared to parental MC38 cells (Fig. 3G, H) under confocal microscopy. To efficiently quantify this change in more cells, we performed ImageStream analysis. Consistently, we found an increased ratio of red to green fluorescence signaling in MC38 PINK1 overexpressing cells due to decreased GFP signals (Fig. S3B, C). To examine the impact of PINK1 on apoptosis, we also measured caspase-3/7 activity in MC38 cells. Interestingly, PINK1 disruption significantly decreased caspase-3/7 activity, while PINK1 overexpression had no effect (Fig. S3D). Together, these data suggest that PINK1 may impact tumor cell growth by modulating cell death.
(A, B) qPCR analysis (n = 3), (C, D) MTT cell survival assay (n = 3), (E, F) western blot analysis (n = 3), (G) Confocal imaging (Bar = 6.2 μm) and (H) quantification of the fluorescence signals in murine MC38 CRC cells with (n = 5) or without stable PINK1 overexpression (n = 4). *p < 0.05, **p < 0.01, ***p < 0.001. Unpaired t-test. One-way ANOVA followed by Dunnett’s multiple comparisons test. 2-way ANOVA followed by Sidak’s multiple comparisons test.
PINK1 is important for metabolic homeostasis in vitro
As PINK1 is a mitochondrial protein and mitochondria are important for regulating cellular respiration, we further analyzed oxygen consumption and glycolysis after PINK1 overexpression and deletion using Seahorse analysis. Interestingly, we found that PINK1 overexpression in MC38 cells leads to increased mitochondrial respiration indicated by OCR (Fig. 4A), but decreased glycolysis indicated by ECAR (Fig. 4B). Specifically, the maximum respiration and spare respiration capacity were increased, whereas glycolysis and glycolytic capacity were decreased. To be noted, there was no change in basal ECAR, glycolytic reserve, basal respiration, ATP production and proton leak by PINK1 overexpression (Figs. S4A and S4B). In contrast, PINK1 knockout in MC38 cells leads to decreased mitochondrial respiration including basal respiration, maximum respiration, spare respiration capacity, ATP production and proton leak (Figs. 4C and S4C), and increased glycolysis including glycolytic capacity and glycolytic reserve (Figs. 4D and S4D). Activation of p53 has shown a similar metabolic effect and p53 is a direct target of PINK1 [26,27,28,29,30]. Thus, we examined the levels of p53, its active form phospho-p53 ser392 as well as p53 target proteins p21, TIGAR, SCO2, DRAM and PUMA after PINK1 overexpression or deletion. Western blot analysis showed that all of the above p53 related proteins were significantly increased by PINK1 overexpression but decreased by PINK1 deletion (Fig. 4E). Interestingly, the active form of a pro-survival protein p38, phospho-p38, was significantly decreased by PINK1 overexpression but increased in PINK1 knockout cells (Fig. 4E). Together, these data indicate that the metabolic changes and decreased cell survival after PINK1 overexpression may be at least partly due to the activation of p53 signaling pathways.
Seahorse XF (A) mitochondrial and (B) glycolysis stress test in MC38 cells with (n = 3) or without stable PINK1 overexpression (n = 5). Seahorse XF (C) mitochondrial and (D) glycolysis stress test in MC38 cells with (n = 5) or without stable PINK1 knockout (n = 5). (E) Western blot analysis in MC38 cells (n = 3) with PINK1 overexpression or ablation. *p < 0.05, **p < 0.01, ***p < 0.001. Unpaired t-test.
PINK1 overexpression suppresses tumor growth in vivo
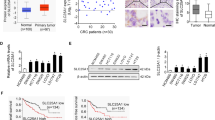
To study whether PINK1 overexpression can suppress tumor growth in vivo, we injected MC38 parental and PINK1 overexpressing cells into C57BL/6 mice. PINK1 overexpression significantly reduced MC38 xenograft tumor size and weight (Fig. 5A, B). By qPCR analysis, we confirmed that the PINK1 expression level was still higher in xenograft tumors derived from MC38 PINK1 overexpressing cells than parental cells (Fig. 5C). Furthermore, using immunofluorescence staining analysis, we found that PINK1 overexpression did not change the percentage of Ki67 positive cells (Fig. 5D), but increased the percentage of CC3 positive cells (Fig. 5E). Together, these data suggest that PINK1 overexpression leads to reduced tumor growth by promoting cell death in vivo.
(A) Representative images, (B) tumor weight (n = 10), (C) qPCR analysis (n = 5), immunofluorescent staining of (D) cell proliferation marker Ki67 (n = 3) and (E) apoptotic marker cleaved caspase 3 (CC3) (n = 3) for MC38 xenograft tumor tissues from C57BL/6 mice. **p < 0.01, NS not significant. Unpaired t test.
PINK1 drives metabolic reprogramming in xenograft tumor
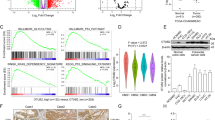
To further understand the impact of PINK1 on cellular metabolism in vivo, we performed an unbiased metabolomic analysis of xenograft tumors with or without PINK1 overexpression. We found the glycolytic product acetyl-CoA was decreased, whereas the TCA intermediates fumarate and malate were increased in tumor xenografts derived from MC38 cells with PINK1 overexpression (Fig. 6A, Supplementary Table 2). PDHE1α is the enzyme that can catalyze pyruvate to acetyl-CoA, whereas PDHK1 phosphorylates and inactivates PDHE1α (Fig. 6B). In addition, PDHK1 is a known target gene of HIF-1α [31]. The mitochondrial protein complex SDH is critical in transforming succinate to fumarate (Fig. 6B). Indeed, Western blot analysis confirmed that HIF-1α/PDHK1/PDHE1α, SDHA and SDHB were significantly increased by PINK1 overexpression, but decreased by PINK1 disruption in MC38 cells (Fig. 6C, D). Importantly, we also confirmed the activation of PDHK1 indicated by increased p-PDHK1 and the induction of mitophagy indicated by decreased mitochondrial protein TOM20 and LC3B accumulation in xenograft tumors derived from MC38 cells with PINK1 overexpression (Fig. S5). Interestingly, we found that the expression of PINK1 was correlated with SDHA, whereas the expression of SDHA was also correlated with p53 expression in colorectal adenoma tissues from the TCGA database (Fig. S6A, B). We then overexpressed SDHA in MC38 cells to test the effect of SDHA on colon tumorigenesis. qPCR analysis confirmed the successful overexpression of SDHA (Fig. S6C, D), but it didn’t change the tumor growth as indicated by xenograft tumor weight (Fig. S6E). Together, these data suggest that PINK1 overexpression drives metabolic reprogramming in tumors by activating p53, glycolytic and mitochondrial enzymes.
A Metabolite quantification by metabolomics for MC38 xenograft tumor tissues from C57BL/6 mice (n = 5). B Schematic diagram showing a simplified glycolysis process and TCA cycle. Blue arrow indicates downregulation, whereas red arrow indicates upregulation. Western blot analysis in MC38 cells (n = 3) with PINK1 (C) overexpression or (D) ablation. NS non-specific band. *p < 0.05, **p < 0.01, ***p < 0.001. Unpaired t-test.
Acetyl-CoA is a critical metabolite for driving colon tumor growth
It has been reported that acetyl-CoA is a critical determinant for tumor growth [32]. In addition to the production of acetyl-CoA from pyruvate oxidation through PDHE1α mediated glycolysis, acetyl-CoA can also be produced by acetyl-coenzyme A synthetase (ACSS) from acetate (Fig. 7A) [32,33,34]. Thus, we examined whether acetate treatment could increase acetyl-CoA levels in xenograft tumors and rescue the PINK1-mediated tumor suppression effect. Strikingly, PINK1 overexpression reproducibly inhibited tumor growth, whereas acetate treatment decreased tumor growth in MC38 parental cells but rescued PINK1-repressed tumor growth in mice (Fig. 7B, C). We also confirmed that the acetyl-CoA levels in tumors were decreased by PINK1 overexpression, whereas acetate treatment increased the acetyl-CoA levels in the tumor with PINK1 overexpression but not in the control group (Fig. 7D). Furthermore, PINK1 disruption significantly increased tumor growth and acetyl-CoA levels (Fig. 7E–G). To test whether p53 activation by PINK1 is responsible for the metabolic and tumor cell growth suppressing effects, we overexpressed a p53 target TIGAR or SCO2 in MC38 sgPINK1 cells and sgEV cells (Fig. S7A, B). TIGAR overexpression increased, whereas SCO2 overexpression decreased cell survival rate in both MC38 sgPINK1 and sgEV cells (Fig. S7C). This was further validated by a xenograft study (Fig. S7D). Together, our data suggest that mitophagy protein PINK1 suppresses colon tumor growth by activation of p53 signaling pathways and inhibiting the production of acetyl-CoA.
A Acetate can be catalyzed by ACSS directly into acetyl-CoA. (B) Representative images, (C) tumor weight (n = 8–10) and (D) acetyl-CoA levels (n = 3-4) for MC38 PINK1 overexpression xenograft tumors from C57BL/6 mice treated with or without 500 mg/kg sodium acetate weekly by i.p. injection for 3 weeks. (E) Representative images, (F) tumor weight and (G) acetyl-CoA levels for MC38 sgPINK1 xenograft tumors (n = 4). *p < 0.05, **p < 0.01 and ***p < 0.001. Unpaired and paired t-test. 2-way ANOVA followed by Sidak’s multiple comparisons test.
Discussion
CRC is the third most commonly diagnosed cancer worldwide and a leading cause of cancer-related deaths [1]. Mitochondria is a major intracellular organelle known for energy and ROS production via utilizing macronutrients. Thus, proper mitochondrial dynamics and quality control is required for normal mitochondrial functions. However, little is known about the exact role of mitochondrial quality control in the progression of CRC. In this paper, we demonstrate that mitophagy protein PINK1 suppresses tumor development by metabolic reprogramming. Mechanistically, PINK1 overexpression leads to the activation of p53 signaling pathways and the HIF-1α/PDHK1/ PDHE1α/acetyl-CoA axis (Fig. 8). Importantly, we demonstrate that acetyl-CoA is a limiting factor for CRC tumor growth.
PINK1 phosphorylates and activates p53, which induces its downstream target genes including TIGAR, SCO2, DRAM, PUMA and p21 [26,27,28,29]. TIGAR decreases glycolysis by dephosphorylating fructose-2,6-bisphosphate and modulates apoptosis [26, 27]. SCO2 is targeted to the inner membrane (IMM) of the mitochondria where it facilitates the assembly of the cytochrome c oxidase (COX) complex in the electron transport chain (ETC), and thus increases mitochondrial respiration and potential utilization of acetyl-CoA [28]. DRAM regulates autophagy and apoptosis [29]. PUMA is a known positive regulator for apoptosis [26]. Meanwhile, PINK1 can also induce HIF1α and its downstream target PDHK1, which phosphorylates PDHE1α and thus inhibits the conversion of pyruvate to acetyl-CoA. Exogenous acetate can be transported across the cell membrane via monocarboxylate transporters (MCTs) and sodium coupled monocarboxylate transporters (SMCTs) into the cell [18, 19]. It is then transported across the mitochondrial IMM through passive diffusion or via an unknown mitochondrial carrier into the mitochondrial matrix [20, 21] where it is catalyzed by ACSS2 to form acetyl-CoA and support tumor growth.
PINK1 is a negative regulator of growth and the Warburg effect in glioblastoma [35]. PINK1 overexpression suppressed the expression of HIF-1α and its downstream target PDHK1 in glioblastoma [35]. On the contrary, PINK1 deficiency sustains cell proliferation by upregulating the expression of HIF-1α and its downstream target PDHK1 in cultured Pink1−/− mouse embryonic fibroblasts and primary cortical neurons as well as in vivo [13]. PDHK1 activation is known to promote the Warburg effect by phosphorylation of PDHE1α, which impedes pyruvate binding and increases tumor development [36]. However, PINK1 overexpression upregulated expression of HIF-1α and PDHK1 despite reduced glycolysis in CRC. One possibility is that this reduced glycolysis is caused by p53 activation since p53 inhibits glycolysis by inducing TIGAR [26, 27]. In addition, p53 promotes mitochondrial respiration via induction of the protein SCO2 [28] and activates the gene expression of SDHA [37]. Consistent with this, we found that PINK1 overexpression increased mitochondrial respiration, whereas our metabolomics study showed that the fumarate levels were elevated after PINK1 overexpression. Thus, we propose that the upregulation of p53 and its downstream targets is responsible for the increased mitochondrial respiration by PINK1 overexpression. Indeed, PINK1 deficiency decreased p53 signaling and mitochondrial respiration, and increased glycolysis. TIGAR overexpression increased cell survival rate and xenograft tumor growth in both MC38 sgPINK1 and sgEV cells. This seems contradictory to our hypothesis that TIGAR overexpression inhibits glycolysis and suppresses PINK1 disruption-mediated tumor growth. However, expression of TIGAR may also protect cells from p53-induced apoptosis by reducing intracellular ROS [27]. In a mouse model of intestinal adenoma, TIGAR disruption decreased tumor burden and increased survival, while TIGAR expression is increased in human colon tumors indicating that TIGAR promotes cancer progression [38]. SCO2 overexpression was able to repress PINK1 depletion-enhanced tumor cell growth both in vitro and in vivo. This is consistent with our hypothesis that induction of oxidative phosphorylation may consume more acetyl-CoA and decrease colon tumor growth. Thus, SCO2 may be one key driver mediating the tumor suppressing effect of p53 and PINK1. However, the upregulation of SDHA may not be a major contributor to the tumor suppressive effect of PINK1 overexpression. SDHA overexpression in MC38 cells didn’t inhibit the xenograft tumor growth. Future studies will test the role of phosphomimetic p53 and SDHB in metabolic regulation and colon tumorigenesis.
We found when PINK1 is overexpressed in MC38 cells in vitro, mitophagy but not apoptosis was increased. This may have resulted from the simultaneous activation of anti-apoptosis proteins (p21, TIGAR and DRAM) and pro-apoptosis proteins (p53 and PUMA) [26, 27, 29, 39, 40]. However, when PINK1 was overexpressed in MC38 xenograft tumors in vivo, both mitophagy and apoptosis were increased. This may be due to the fact that nutrients are more restricted in the in vivo tumor environment compared to nutrient-rich in vitro cell culture conditions. In addition, we showed that autophagy markers and apoptosis were decreased in PINK1 knockout cells. Together, we believe PINK1 is a critical modulator of cell survival.
Our paper highlights the determinant role of acetyl-CoA in the growth of tumor xenografts. Acetyl-CoA is an important metabolic signal for growth and proliferation [41]. As mentioned above, acetate can provide an alternative source of acetyl-CoA in addition to pyruvate oxidation [42]. Interestingly, a recent paper showed that exogenous acetate supplementation suppresses intestinal stem cell loss in HNF4αγDKO mutant organoids but impairs the growth of wild-type organoids [43]. This is similar to our observation that acetate reduced tumor growth in MC38 parental cells but rescued PINK1-repressed tumor growth in mice. Under metabolically stressful conditions, ACSS2 promotes acetate utilization and tumor growth [44]. However, ACSS2 is downregulated in CRC tumor tissues compared with normal colonic mucosa and low expression of ACSS2 is an independent prognostic factor for poor 5-year progression-free survival [45]. This may be due to the fact that normal colonocytes use short chain fatty acids including acetate derived from intestinal microbiota fermentation as an energy source [45, 46]. Thus, future experiments are needed to examine the metabolic changes in colon tumor tissues derived from Pink1−/− mice compared to Pink1+/+ mice. Increased acetate percentage in short chain fatty acids was found in enema samples from CRC patients compared to normal subjects [47]. Thus, it would also be interesting to test whether decreasing acetate producing bacteria through modulation of gut microbiota would lead to decreased acetyl-CoA amounts and provide any benefits to CRC patients [48].
Mitochondrial damage can lead to broken mitochondrial membranes and subsequent mitochondrial DNA leakage, which can activate the NLRP3 inflammasome to release IL-1β and activate cGAS/STING to induce IFN-β [49]. Indeed, NLRP3 inflammasome activation has been found in cells from Pink1−/− mice under stressed conditions [50,51,52]. A recent landmark paper found that PINK1/Parkin alleviated the cGAS/STING pathway induced inflammation upon mitochondrial stress [53]. Considering that cytokines including IL-1β and IFN-β were increased in colon tumor tissues from our Pink1−/− mice, it would be important to investigate the involvement of the NLRP3 inflammasome and the cGAS/STING pathway in PINK1-suppressed colon tumorigenesis in the future.
In conclusion, we have demonstrated that mitophagy protein PINK1 is a tumor suppressor that modulates cellular metabolism and promotes cell death in CRC. Mechanistically, we showed that PINK1 suppresses colon tumorigenesis by metabolic reprogramming through p53 activation and reducing acetyl-CoA production. This study provides new insights that can be potentially used as therapeutic strategies for the treatment of CRC.
References
Siegel RL, Miller KD, Jemal A. Cancer statistics, 2020. CA Cancer J Clin. 2020;70:7–30.
Xue X, Bredell BX, Anderson ER, Martin A, Mays C, Nagao-Kitamoto H, et al. Quantitative proteomics identifies STEAP4 as a critical regulator of mitochondrial dysfunction linking inflammation and colon cancer. Proc Natl Acad Sci USA. 2017;114:E9608–E9617.
Liberti MV, Locasale JW. The Warburg effect: how does it benefit cancer cells? Trends Biochem Sci. 2016;41:211–8.
Ward PS, Thompson CB. Metabolic reprogramming: a cancer hallmark even warburg did not anticipate. Cancer Cell. 2012;21:297–308.
Spinelli JB, Haigis MC. The multifaceted contributions of mitochondria to cellular metabolism. Nat Cell Biol. 2018;20:745–54.
Ahn CS, Metallo CM. Mitochondria as biosynthetic factories for cancer proliferation. Cancer Metab. 2015;3:1.
Dang CV, Hamaker M, Sun P, Le A, Gao P. Therapeutic targeting of cancer cell metabolism. J Mol Med. 2011;89:205–12.
Kaldma A, Klepinin A, Chekulayev V, Mado K, Shevchuk I, Timohhina N, et al. An in situ study of bioenergetic properties of human colorectal cancer: the regulation of mitochondrial respiration and distribution of flux control among the components of ATP synthasome. Int J Biochem Cell Biol. 2014;55:171–86.
Satoh K, Yachida S, Sugimoto M, Oshima M, Nakagawa T, Akamoto S, et al. Global metabolic reprogramming of colorectal cancer occurs at adenoma stage and is induced by MYC. Proc Natl Acad Sci USA. 2017;114:E7697–E7706.
Shirihai OS, Song M, Dorn GW 2nd. How mitochondrial dynamism orchestrates mitophagy. Circ Res. 2015;116:1835–49.
Wang K, Klionsky DJ. Mitochondria removal by autophagy. Autophagy. 2011;7:297–300.
Um J-H, Yun J. Emerging role of mitophagy in human diseases and physiology. BMB Rep. 2017;50:299–307.
Ziegler PK, Bollrath J, Pallangyo CK, Matsutani T, Canli Ö, De Oliveira T, et al. Mitophagy in intestinal epithelial. Cells Triggers Adapt Immun Tumorigenesis Cell. 2018;174:88–101.e16.
Dagda RK, Cherra SJ 3rd, Kulich SM, Tandon A, Park D, Chu CT. Loss of PINK1 function promotes mitophagy through effects on oxidative stress and mitochondrial fission. J Biol Chem. 2009;284:13843–55.
Li C, Zhang Y, Cheng X, Yuan H, Zhu S, Liu J, et al. PINK1 and PARK2 suppress pancreatic tumorigenesis through control of mitochondrial iron-mediated immunometabolism. Dev Cell. 2018;46:441–455.e8.
Requejo-Aguilar R, Lopez-Fabuel I, Fernandez E, Martins LM, Almeida A, Bolaños JP. PINK1 deficiency sustains cell proliferation by reprogramming glucose metabolism through HIF1. Nat Commun. 2014;5:4514.
Xue X, Ramakrishnan SK, Weisz K, Triner D, Xie L, Attili D, et al. Iron uptake via DMT1 integrates cell cycle with JAK-STAT3 signaling to promote colorectal tumorigenesis. Cell Metab. 2016;24:447–61.
Halestrap AP, Price NT. The proton-linked monocarboxylate transporter (MCT) family: structure, function and regulation. Biochem J. 1999;343(Pt 2):281–99.
Ganapathy V, Thangaraju M, Gopal E, Martin PM, Itagaki S, Miyauchi S, et al. Sodium-coupled monocarboxylate transporters in normal tissues and in cancer. AAPS J. 2008;10:193–9.
Arduini A, Zammit V. Acetate transport into mitochondria does not require a carnitine shuttle mechanism. Magn Reson Med. 2017;77:11.
Rivière L, Moreau P, Allmann S, Hahn M, Biran M, Plazolles N, et al. Acetate produced in the mitochondrion is the essential precursor for lipid biosynthesis in procyclic trypanosomes. Proc Natl Acad Sci USA. 2009;106:12694–9.
Kim H, Yin K, Falcon DM, Xue X. The interaction of Hemin and Sestrin2 modulates oxidative stress and colon tumor growth. Toxicol Appl Pharm. 2019;374:77–85.
Rojansky R, Cha M-Y, Chan DC. Elimination of paternal mitochondria in mouse embryos occurs through autophagic degradation dependent on PARKIN and MUL1. Elife. 2016;5:e17896.
Rhodes DR, Yu J, Shanker K, Deshpande N, Varambally R, Ghosh D, et al. ONCOMINE: a cancer microarray database and integrated data-mining platform. Neoplasia. 2004;6:1–6.
Pan Q, Lou X, Zhang J, Zhu Y, Li F, Shan Q, et al. Genomic variants in mouse model induced by azoxymethane and dextran sodium sulfate improperly mimic human colorectal cancer. Sci Rep. 2017;7:25.
Green DR, Chipuk JE. p53 and metabolism: inside the TIGAR. Cell. 2006;126:30–32.
Bensaad K, Tsuruta A, Selak MA, Vidal MNC, Nakano K, Bartrons R, et al. TIGAR, a p53-inducible regulator of glycolysis and apoptosis. Cell. 2006;126:107–20.
Matoba S, Kang J-G, Patino WD, Wragg A, Boehm M, Gavrilova O, et al. p53 regulates mitochondrial Respiration. Science. 2006;312:1650–3.
Crighton D, Wilkinson S, O’Prey J, Syed N, Smith P, Harrison PR, et al. DRAM, a p53-induced modulator of autophagy, is critical for apoptosis. Cell. 2006;126:121–34.
Liu K, Lee J, Kim JY, Wang L, Tian Y, Chan ST, et al. Mitophagy controls the activities of tumor suppressor p53 to regulate hepatic cancer stem cells. Mol Cell. 2017;68:281–292.e5.
Kim J, Tchernyshyov I, Semenza GL, Dang CV. HIF-1-mediated expression of pyruvate dehydrogenase kinase: a metabolic switch required for cellular adaptation to hypoxia. Cell Metab. 2006;3:177–85.
Comerford SA, Huang Z, Du X, Wang Y, Cai L, Witkiewicz AK, et al. Acetate dependence of tumors. Cell. 2014;159:1591–602.
Frost G, Sleeth ML, Sahuri-Arisoylu M, Lizarbe B, Cerdan S, Brody L, et al. The short-chain fatty acid acetate reduces appetite via a central homeostatic mechanism. Nat Commun. 2014;5:3611.
Shubitowski TB, Poll BG, Natarajan N, Pluznick JL. Short-chain fatty acid delivery: assessing exogenous administration of the microbiome metabolite acetate in mice. Physiol Rep. 2019;7:e14005–e14005.
Agnihotri S, Golbourn B, Huang X, Remke M, Younger S, Cairns RA, et al. PINK1 is a negative regulator of growth and the warburg effect in glioblastoma. Cancer Res. 2016;76:4708–19.
Hitosugi T, Fan J, Chung T-W, Lythgoe K, Wang X, Xie J, et al. Tyrosine phosphorylation of mitochondrial pyruvate dehydrogenase kinase 1 is important for cancer metabolism. Mol Cell. 2011;44:864–77.
Sp N, Kang DY, Kim DH, Lee HG, Park Y-M, Kim IH, et al. Methylsulfonylmethane inhibits cortisol-induced stress through p53-mediated SDHA/HPRT1 expression in racehorse skeletal muscle cells: A primary step against exercise stress. Exp Ther Med. 2020;19:214–22.
Cheung EC, Athineos D, Lee P, Ridgway RA, Lambie W, Nixon C, et al. TIGAR is required for efficient intestinal regeneration and tumorigenesis. Dev Cell. 2013;25:463–77.
Gartel AL, Tyner AL. The role of the cyclin-dependent kinase inhibitor p21 in apoptosis. Mol Cancer Ther. 2002;1:639–49.
Weber JD, Zambetti GP. Renewing the debate over the p53 apoptotic response. Cell Death Differ. 2003;10:409–12.
Cai L, Sutter BM, Li B, Tu BP. Acetyl-CoA induces cell growth and proliferation by promoting the acetylation of histones at growth genes. Mol Cell. 2011;42:426–37.
Stine RR, Sakers AP, TeSlaa T, Kissig M, Stine ZE, Kwon CW, et al. PRDM16 maintains homeostasis of the intestinal epithelium by controlling region-specific metabolism. Cell Stem Cell. 2019;25:830–845.e8.
Chen L, Vasoya RP, Toke NH, Parthasarathy A, Luo S, Chiles E, et al. HNF4 regulates fatty acid oxidation and is required for renewal of intestinal stem cells in mice. Gastroenterology. 2020;158:985–999.e9.
Schug ZT, Peck B, Jones DT, Zhang Q, Grosskurth S, Alam IS, et al. Acetyl-CoA synthetase 2 promotes acetate utilization and maintains cancer cell growth under metabolic stress. Cancer Cell. 2015;27:57–71.
Bae JM, Kim JH, Oh HJ, Park HE, Lee TH, Cho N-Y, et al. Downregulation of acetyl-CoA synthetase 2 is a metabolic hallmark of tumor progression and aggressiveness in colorectal carcinoma. Mod Pathol. 2017;30:267–77.
Louis P, Hold GL, Flint HJ. The gut microbiota, bacterial metabolites and colorectal cancer. Nat Rev Microbiol. 2014;12:661–72.
Weaver GA, Krause JA, Miller TL, Wolin MJ. Short chain fatty acid distributions of enema samples from a sigmoidoscopy population: an association of high acetate and low butyrate ratios with adenomatous polyps and colon cancer. Gut. 1988;29:1539–43.
Walker AW, Duncan SH, McWilliam Leitch EC, Child MW, Flint HJ. pH and peptide supply can radically alter bacterial populations and short-chain fatty acid ratios within microbial communities from the human colon. Appl Environ Microbiol. 2005;71:3692–3700.
Youle RJ. Mitochondria—striking a balance between host and endosymbiont. Science. 2019;365:eaaw9855.
Zhang Y, Sauler M, Shinn AS, Gong H, Haslip M, Shan P, et al. Endothelial PINK1 mediates the protective effects of NLRP3 deficiency during lethal oxidant injury. J Immunol. 2014;192:5296–304.
Kang R, Zeng L, Xie Y, Yan Z, Zhou B, Cao L, et al. A novel PINK1- and PARK2-dependent protective neuroimmune pathway in lethal sepsis. Autophagy. 2016;12:2374–85.
Mouton-Liger F, Rosazza T, Sepulveda-Diaz J, Ieang A, Hassoun S-M, Claire E, et al. Parkin deficiency modulates NLRP3 inflammasome activation by attenuating an A20-dependent negative feedback loop. Glia. 2018;66:1736–51.
Sliter DA, Martinez J, Hao L, Chen X, Sun N, Fischer TD, et al. Parkin and PINK1 mitigate STING-induced inflammation. Nature. 2018;561:258–62.
Acknowledgements
This research was supported in part by the National Institutes of Health (P20 GM130422, K01DK114390), a Research Scholar Grant from the American Cancer Society (RSG-18-050-01-NEC), a Research Pilot Project Grant from University of New Mexico Environmental Health Signature Program and Superfund (P42 ES025589), a Shared Resources Pilot Project Award and a Research Program Support Pilot Project Award from UNM comprehensive cancer center (P30CA118100), a new investigator award from the Dedicated Health Research Funds at the University of New Mexico School of Medicine, and a Core Utilization Voucher Program award and a Metabolomic Studies Voucher Award from the Autophagy, Inflammation and Metabolism (AIM) Center of Biomedical Research Excellence (P20GM121176). We thank Dr. Jim Liu for providing us access to the SpectraMax M2 Microplate Reader, Dr. Laurie Hudson for providing us access to the microtome, Dr. Michael Paffett from the UNMCC Fluorescence Microscopy Core facility for helping with confocal imaging, Dr. Sharina P Desai from Scientific Cores off AIM center for helping with ImageStream analysis, and Dr. Jesse L Denson for proofreading the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by M. Oren
Supplementary information
Rights and permissions
About this article
Cite this article
Yin, K., Lee, J., Liu, Z. et al. Mitophagy protein PINK1 suppresses colon tumor growth by metabolic reprogramming via p53 activation and reducing acetyl-CoA production. Cell Death Differ 28, 2421–2435 (2021). https://doi.org/10.1038/s41418-021-00760-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-021-00760-9
This article is cited by
-
Soy isoflavones induces mitophagy to inhibit the progression of osteosarcoma by blocking the AKT/mTOR signaling pathway
Molecular Medicine (2024)
-
Hibiscus manihot L. flower extract induces anticancer activity through modulation of apoptosis and autophagy in A549 cells
Scientific Reports (2024)
-
Association of mitochondrial homeostasis and dynamic balance with malignant biological behaviors of gastrointestinal cancer
Journal of Translational Medicine (2023)
-
DNMT3A mutation promotes leukemia development through NAM-NAD metabolic reprogramming
Journal of Translational Medicine (2023)
-
METTL14-dependent maturation of pri-miR-17 regulates mitochondrial homeostasis and induces chemoresistance in colorectal cancer
Cell Death & Disease (2023)