Abstract
Increasing evidence has indicated that long noncoding RNAs (lncRNAs) play important roles in human diseases, including cancer; however, only a few of them have been experimentally validated and functionally annotated. Here, we identify a novel lncRNA that we term HITT (HIF-1α inhibitor at translation level). HITT is commonly decreased in multiple human cancers. Decreased HITT is associated with advanced stages of colon cancer. Restoration of the expression of HITT in cancer cells inhibits angiogenesis and tumor growth in vivo in an HIF-1α-dependent manner. Further study reveals that HITT inhibits HIF-1α expression, mainly by interfering with its translation. Mechanically, HITT titrates away YB-1 from the 5′-UTR of HIF-1α mRNA via a high-stringency YB-1-binding motif. The reverse correlation between HITT and HIF-1α expression is further validated in human colon cancer tissues. Moreover, HITT is one of the most altered lncRNAs upon the hypoxic switch and HITT downregulation is required for hypoxia-induced HIF-1α expression. We further demonstrate that HITT and HIF-1α form an autoregulatory feedback loop where HIF-1α destabilizes HITT by inducing MiR-205, which directly targets HITT for degradation. Together, these results expand our understanding of the cancer-associated functions of lncRNAs, highlighting the HITT–HIF-1α axis as constituting an additional layer of regulation of angiogenesis and tumor growth, with potential implications for therapeutic targeting.
Similar content being viewed by others
Introduction
Long noncoding RNAs (lncRNAs), defined as transcribed RNA molecules that are >200 nt in length, have attracted increasing attention [1, 2]. It has been shown that the central functions of lncRNAs encompass every level of the gene expression program involved in regulating multiple biological processes [3, 4], and aberrant lncRNA expression is associated with complex human diseases, such as cancer [5,6,7]. Further study of lncRNA functions is expected to revise our understanding of cancer biology and open up new opportunities for cancer treatment [8, 9]. However, the functions and regulation of the vast majority of lncRNAs currently remain completely unknown.
Hypoxia is a hallmark of the solid tumor microenvironment. Activation of the transcription factor hypoxia-inducible factor (HIF-1) enables cancer cells to adapt to the hypoxic environment via the transactivation of downstream target genes [10, 11]. Vascular endothelial growth factor (VEGF) is a well-established HIF-1 target that plays essential roles in promoting tumor angiogenesis [12, 13]. Once expressed upon HIF-1α activation, it stimulates the development of new blood vessels within tumors, providing nutrients and oxygen to facilitate tumor growth and metabolism, clearing metabolic waste and CO2 [13], and also providing a main route for tumor cells to metastasize to distant organs [14]. Increased expression of HIF-1 correlates with the promotion of angiogenesis, tumor metastasis, and poor prognosis in multiple types of solid tumors, including those of the colon and cervix [15, 16]. Thus, increasing our understanding of the regulatory mechanisms underlying HIF-1α expression and activation may lead to the development of new anticancer therapies that target HIF-1α [17, 18].
It is well established that HIF-1α is subjected to proteasome degradation under normal oxygen tension. Under hypoxia, HIF-1α escapes degradation and translocates to the nucleus, where it binds with the hypoxia response element (HRE) on promoters to initiate gene expression [19]. Although it is believed that such an increase in HIF-1α activity is mainly due to increased protein stability, recent evidence indicates that translational regulation of HIF-1α after the hypoxic switch may also play an important role. Galban et al. and Hui et al. have both shown that HIF-1α translation accounts for 40–50% of HIF-1α induction under hypoxia [20, 21]. It has been also reported that the activation of translational signals is correlated with elevated rates of HIF-1α translation and subsequent protein output [22,23,24]. However, although the regulation of HIF-1α degradation is well understood, much less is known about the regulation of HIF-1α biosynthesis.
Interestingly, evidence for the roles of lncRNAs in regulating HIF-1α activity is beginning to emerge. For example, an HIF-1α lncRNA can directly regulate the stability of HIF-1α mRNA and thus confers poor prognosis in lung cancer [25]. LncRNAs, such as LincRNA-p21 [26] and RERT [27], regulate HIF-1α indirectly by interfering with the proteasome degradation machinery that regulates HIF-1α. Without doubt, lncRNAs have added an additional layer of complexity to the HIF-1α-signaling network [28]; however, whether or not HIF-1α biosynthesis can be directly modulated by lncRNAs is still beyond our understanding. In particular, whether lncRNAs play any roles in HIF-1α’s regulation at the translational level remains unknown.
Chromosomal abnormalities are defining features of solid tumors. Frequently lost chromosome regions in cancer cells can unmask the locations of putative tumor suppressor genes. Notably, loss of 14q32 has been associated with the early onset and metastatic recurrence of colon cancer, as well as many other types of cancer [29,30,31], suggesting that important tumor suppressor genes might exist in this region. Among them, a previously uncharacterized lncRNA called linc00637 (we name this gene HITT: HIF-1α inhibitor at translation level) has attracted our attention because analysis of The Cancer Genome Atlas (TCGA) data sets reveals that this gene is significantly downregulated in multiple cancer types. We speculate that HITT may be one such 14q32 entity that possesses potential tumor suppressive activity and then selected it for further functional analysis.
Materials and methods
Cell lines and treatments
The human colorectal cancer (HCT116, HCT15, SW480, SW620, Colo205, and HT29), cervical cancer (HeLa, CASKi, SiHa, and C33A) cells, and HUVECs were cultured in RPMI-1640 medium (Gibco, Carlsbad, CA, USA) or Dulbecco’s modified Eagle’s medium supplemented with 10% (v/v) fetal bovine serum (Biological industries). All cells were grown at 37 °C in the humidified incubator with 5% CO2. For hypoxia treatment, cells were cultured in an atmosphere of 1% O2 at 37 °C in a hypoxia incubator for the indicated periods of time. All cell lines were authenticated and characterized by the supplier and were monitored regularly for their authenticity (Genetic Testing Biotechnology Corporation, Suzhou, China) and to be free of mycoplasma contamination.
Tissue samples
Human colorectal cancer tissues and their corresponding adjacent normal controls were collected from the Third Affiliated Hospital of Harbin Medical University in China. Written informed consent was obtained from all patients. The study has been approved by the Research Ethics Committee of Harbin Medical University, China. Specimens were collected and stored in liquid nitrogen immediately after surgery.
Xenografted tumor model in vivo
HITT expression was stably restored in HCT116 cells, by using empty vector as a control. The female nude mice between 4 and 5 weeks were purchased from Beijing HFK Bioscience Co., Ltd. The cells were paired and 1 × 107 paired cells were inoculated subcutaneously into either side of flank of the same female nude mouse. The tumor volumes were measured every week and calculated as length × width2 × 0.5. After 4 weeks, the mice were anesthetized and culled. The tumor was carefully removed, photographed, and weighed. All animal procedures were performed according to protocols approved by the Rules for Animal Experiments published by the Chinese Government (Beijing, China) and approved by the Research Ethics Committee of Harbin Medical University, China.
Small interfering RNA (siRNA) transfection
siRNA specifically targeting HITT, YB-1, HIF-1α, HIF-2α, or Ago2, and nonspecific si-scramble control were synthesized by GenePharma (Shanghai, China). All transfection experiments were conducted by using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA) following the manufacturer’s instructions. The siRNA sequences are listed in Supplementary Table S1.
RNA extraction and qRT-PCR
Total RNA was isolated using Trizol Reagent (Invitrogen, Carlsbad, CA, USA) following the manufacture’s protocol. qRT-PCR was performed in triplicate with an Applied Biosystems Prism 7500 Fast Sequence Detection System using TaqMan universal PCR master mix according to the manufacture’s protocol (Applied Biosystems Inc, Foster City, CA, USA). Gene expression levels relative to GAPDH, U6, or 18S rRNA were calculated by 2−ΔΔCT method. The primer sequences used in qRT-PCR are listed in Supplementary Table S2.
Western blot
Total proteins from cell lines or tissue samples were isolated with UREA buffer (8 M Urea, 1 M Thiourea, 0.5% CHAPS, 50 Mm DTT, and 24 Mm Spermine). Equal amount of proteins were separated at sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). After incubation with the indicated primary and secondary antibodies, signals were visualized by ECL. Membrane was then ready for scanning by Image studio system. Protein quantification was conducted by imageJ software. The antibodies were listed in Supplementary Table S3.
Immunohistochemistry
The tissue sample was subjected to antigen retrieval by boiling in 0.01 mol/L citrate buffer for 5 min (min). Slides were then incubated with anti-CD31 or anti-Ki-67 antibody at room temperature for 1 h. Detection was carried out by the REAL EnVision detection system (Dako) with diaminobenzidine peroxidase serving as chromogen. After that, the slides were briefly counterstained with hematoxylin before mounting. The images were captured under the same conditions and quantitative analysis using ImageJ software.
Tube formation assay
Each well of 24-well plates was coated with 200 μl of Matrigel (Corning, NY, USA) and polymerized for 1 h at 37 °C. HUVECs and 0.5 ml culture medium were seeded onto the solidified gel. After 8 h, tube formation was observed and captured with microscopy.
Luciferase assay
3× HRE sequence (GCATACGTGGGCGCATACGTGGGCGCATACGTGGGC) was cloned into PGL3-basic luciferase reporter plasmid, namely HRE-luciferase reporter. HIF-1α wild type (WT) 5′-UTR(1-256), 5′-UTR(1-138, mutant type, MT1), and 5′-UTR(139-256, MT2) were cloned into a PGL3 vector, respectively, with CMV promoter at upstream of the insert to generate reporter plasmids, namely HIF-1α-5′-UTR (1-256), HIF-1α-5′-UTR (1-138), and HIF-1α-5′-UTR (139-256), respectively. WT or MiR-205 binding site mutant (MT) HITT were cloned into PMIR-Reporter downstream of luciferase, namely WT HITT reporter and MT HITT reporter, respectively. After the indicated treatment, cells were harvested and subjected to an assay by using the Dual Luciferase Reporter Assay system (Promega, Madison, USA). The relative luciferase activities were normalized with the Renilla luciferase activities.
Chromatin immunoprecipitation (ChIP)
Chromatin immunoprecipitation (ChIP) assay was performed to estimate protein binding at specific promoter as described [32]. Briefly, 1 × 107 cells were collected in lysis buffer A and B (buffer A: 5 mM PIPES, 85 mM KCl, 0.5% NP40. buffer B: 1% SDS, 10 mM EDTA, 50 mM Tris-HCl.) supplemented with Protease Inhibitor Cocktail (MedChemExpress, Shanghai, China) and sonicated to generate chromatin samples with average fragment sizes of 100–500 bp. Cell lysates were treated with the indicated antibodies or IgG control at 4 °C overnight. Then, the supernatants were mixed with the blocked Protein A/D sepharose beads to collect the antibody–chromatin complexes. After washing four times, the immunoprecipitated DNA was eluted and purified for the subsequent qPCR analysis.
UV-cross-linking RNA-IP (CLIP)
RNP assay was performed to estimate physiological interaction between protein and RNA as described [32]. Briefly, 1 × 107 cells were washed and UV cross-linked at 400 mJ/cm2 and collected in lysis buffer supplemented with Protease Inhibitor Cocktail and RNase inhibitor (Thermo Fisher, Rockford, IL). After the cells were precleared with protein A/G sepharose beads, cell lysates were treated with indicated antibody or IgG control at 4 °C overnight. Then, the antibody–RNA complexes were collected by using the blocked Protein A/G sepharose beads. After four times washes in the presence of Protease Inhibitor Cocktail and RNase inhibitor, the immunoprecipitated RNA was eluted and Isolated for the subsequent reaction to reverse transcribed into first-strand cDNA and using for further qRT-PCR analysis for the target RNA of interest.
l-azidohomoalanine labeling to identify newly synthesized proteins
HeLa cells were washed and incubated for 30 min in methionine‐free medium to deplete intracellular methionine, followed by incubation with 50 μM l-azidohomoalanine (AHA, Invitrogen, Carlsbad, CA, USA) for 4 h under hypoxia. Cells were washed twice with PBS, lysis buffer were added, and then sonicated for two cycles of 30 s each and centrifuged cell at 13,000 g. Fifty micrograms of lysates were subjected to Click reactions according to the manufacturer’s instructions (Click-iT® Protein Reaction Buffer Kit; Invitrogen, Carlsbad, CA, USA). Total proteins from Click reactions were precipitated with methanol/chloroform and resolubilized in 50 mM Tris, pH 7.5, 0.01% SDS, and then incubated with 50 μl of Streptavidin coupled magnetic beads for 5 h at room temperature. Streptavidin coupled magnetic beads were incubated in 50 μl of 2× loading blue for 10 min at 100 °C and detected by western blots.
In vitro RNA pull-down assay
Biotin-labeled RNA were in vitro synthesized by Biotin RNA Labeling Mix (Roche, St Louis, MO, USA). After treatment with RNase-free DNase I, Biotin-labeled RNA was heated at 95 °C for 2 min and then cooled on ice for 3 min. Then secondary structure recovered Biotin-labeled RNA reacted with streptavidin agarose beads (Invitrogen, Carlsbad, CA, USA) overnight. The fresh HCT116 and HeLa lysates were collected and incubated with RNA-captured beads at 4 °C for 1 h. Beads were briefly washed for five times with wishing buffer (50 mM Tris-HCl (PH 7.4), 150 mM NaCl, 1 mM MgCl2, 0.05% NP40). The resulting beads were boiled at 95 °C for 5 min in SDS loading buffer and then subjected to western blot analysis. Than the samples were detected on 15% polyacrylamide TBE gel at 4 °C for 60 min at 200 V.
EMSA
In vitro synthesized RNA oligonucleotides were heated at 70 °C for 5 min and then cooled on ice for 2 min ready for EMSA. One picomole in vitro synthesized dsDNA and 20 pmol prepared ssRNA were incubated in 10 μl buffer that contain 1 μl 10× triplex forming buffer (100 mM Tris pH 7.5, 25 mM NaCl, and MgCl2) at 37 °C for 30 min. In control assay, triplex reaction was treated with either 5 U of RNase H or 20 mg/ml RNase A at 37 °C for 30 min.
Polysome fractionation
Cells were treated with 100 μg/ml cycloheximide (CHX) for 5 min and then lysed in polysome lysis buffer (15 mM Tris-HCL PH 7.5, 15 mM MgCl2, 0.3 M NaCl, 1% Triton X-100, 0.1 U/μl RNA inhibitor, 100 μg/ml CHX, 1 μg/ml Heparin, and 1× protease inhibitor cocktail). The sucrose gradient samples were obtained by centrifuge at 39,000 rpm for 2 h at 4 °C using SW40Ti rotor in a Beckman Coulter and monitored by using ultraviolet spectrophotometer at 254 nm. RNA in each sucrose gradient was collected and extracted in 3 volumes of Trizol, followed by qRT-PCR assay for the indicated genes.
ELISA
VEGF concentration of cell culture medium was measured by ELISA using the Quantikine human VEGF ELISA Kit according to the manufacturer’s instructions (R&D Systems, Minneapolis, MN, USA). The sample absorbance was determined at 450 and 540 nm. Each experiment was repeated three times.
Statistical analysis
Statistical analysis was done by GraphPad software, version 5. Data are presented as the means ± standard error of the means (SEM) or standard Deviation (SD). Student t test was applied to assess the statistical significance. Correlations were calculated according to Spearmen or Pearson correlation. p value < 0.05 were considered significant.
Results
LncRNA HITT is commonly reduced in human cancers
Rapid amplification of 3′- and 5′-complementary DNA ends (3′- and 5′-RACE) assays confirmed that HITT (linc00637, Fig. S1A) is a 2050-nt transcript possessing a polyadenylation site (Fig. S1B), with the expected size (~2 kb) as revealed by northern blot assay (Fig. S1C). HITT received a coding potential value of 0.387, similar to other well-characterized lncRNAs and much lower than protein-coding genes, as analyzed by Coding Potential Calculation (Fig. S1D). Comparable amounts of HITT were detected in both the nucleus and cytoplasm, using cytoplasmic GAPDH mRNA and the nuclear lncRNA MALAT1 [33] as controls (Fig. S1E).
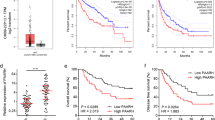
Remarkably, qRT-PCR results revealed that HITT was significantly downregulated in the colon cancer tissues analyzed relative to the paired normal controls (Fig. 1a). Absolute values of fold-change increased according to TNM stages (Fig. 1b). HITT levels were much lower in colon cancer cell lines than in pooled independent normal tissue controls (Fig. S2A). Furthermore, HITT levels were reduced in the cervical (n = 6) and renal cell carcinoma tissues (n = 17), and cervical cancer cell lines examined (Fig. S2B–C). Analysis of the TCGA database revealed that HITT is downregulated in multiple cancer types, such as colon cancer, thyroid cancer, and chromophobe renal cell carcinoma (Fig. S2D), and that decreased HITT is associated with advanced stages of bladder, breast, and liver cancers (Fig. S2E). Lower levels of HITT also predict poor clinical outcomes in lung and rectal cancers (Fig. S2F). These findings collectively indicate that HITT is commonly downregulated in human cancers.
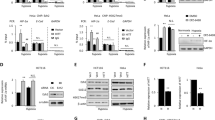
HITT prevents tumor growth in vivo by inhibiting angiogenesis in a HIF-1α dependent manner. The expression levels of HITT in human colorectal tumors (T) of different stages (B) relative to their paired adjacent normal controls (N) (n = 46, a, b) were determined by qRT-PCR. Tumor volumes at the indicated dates (c), as well as images (left) and tumor weights (right) at 4 weeks (d) of HCT116/vector (Vect.), HIC116/HITT, HCT116/(Vect. + si-HIF-1α), and HCT116/(HITT + si-HIF-1α) xenografts. The average values are present as bar graphs (means ± SD) (n = 6 for each group). Representative immunohistochemistry staining for Ki-67 (top) and CD31 (bottom) in the indicated xenografts (e), and the quantification of bar graphs (f). g HIF-1α and VEGF protein (upper panel) and VEGF mRNA (lower panel) levels were measured by WB and qRT-PCR, respectively, in HCT116 xenografts. Representative images are presented. h, i VEGF protein levels were measured by ELISA in clone stably transfected with HITT and vector controls after HIF-1α knockdown (KD) or ectopic HIF-1α expression under hypoxia. The KD efficiency of HIF-1α was confirmed by WB. j Representative images of HUVECs are presented (left) and quantification of HUVEC tube formation assays is presented in the bar graph (right). Data derived from three independent experiments are presented as mean ± SEM in the bar graph. Values in controls were normalized to 1 (a, f, g). *P < 0.05; **P < 0.01; ##P < 0.01, compared with hypoxia-treated controls (h–j). Vect. vector control, Ctl. control. Scale bar 50 μm. See also Fig. S1–S3
HITT suppresses angiogenesis and tumor growth in a HIF-1α-dependent manner
To understand its functional significance, single clones of HeLa and HCT116 cells stably expressing HITT at levels similar to those of normal controls were used for further analysis (Fig. S3A). However, no obvious differences in the anchorage-dependent or -independent growth capabilities was observed after restoration of HITT expression (Fig. S3B–C). Whereas HITT expression significantly reduced the tumor volumes and weights of HCT116 xenografts implanted in nude mice (Fig. 1c, d). These data suggest that HITT inhibits tumor growth, possibly by antagonizing detrimental microenvironmental factors.
Hypoxia is a principle microenvironment factor that initiates angiogenesis and promotes tumor growth. Given that HIF-1α is a master regulator of tumor angiogenesis, we examined whether the antitumor effect of HITT relies on the presence of HIF-1α protein. To this end, HIF-1α expression was inhibited using siRNA specifically targeted to HIF-1α. In line with previous reports, HIF-1α knockdown (KD) abolished HIF-1α expression and significantly reduced the tumor growth of HCT116 xenografts (2.5–3-fold, Fig. 1c, d). Importantly, the effect of HITT on tumor growth was completely abrogated by HIF-1α KD (Fig. 1c, d). Consistently, Ki-67 intensity and vascular density were reduced with HITT expression or HIF-1α KD, and no further reduction was observed upon their combination (Fig. 1e, f). VEGF, a known HIF-1α downstream pro-angiogenesis target [13], was inhibited by HITT (Fig. 1g), paralleling the decreased tumor vasculature (Fig. 1e, f) and HIF-1α protein levels (Figs. 1g and S3D).
The effect of HITT on HIF-1α-induced angiogenesis was verified using in vitro models. The conditioned medium from cells cultured under hypoxia significantly stimulated the branching abilities of HUVECs relative to the normoxia control (Fig. 1h–j). HITT transfection abrogated conditioned medium-induced HUVEC branching (Fig. 1j). HIF-1α KD also abrogated hypoxia-induced VEGF expression and HUVEC branching, and, interestingly, completely abolished the antiangiogenic effect of HITT (Fig. 1i). In contrast, HIF-1α overexpression rescued HITT-repressed HUVECs branching (Figs. 1h, j). Collectively, the findings indicate that HITT inhibits angiogenesis and tumor growth by interfering with HIF-1α-dependent signals.
HITT inhibits HIF-1α expression and activity
We next sought out to understand how HITT regulates HIF-1α signals. The results show that HITT markedly diminished hypoxia-induced HIF-1α expression by approximately five to sixfold (Fig. 2a, b), while HITT KD increased HIF-1α under normoxia (Fig. 2c–e). We also noticed that HITT consistently inhibited HIF-1α mRNA expression by approximately twofold (lower panel, Fig. 2b), while HITT KD promoted it (lower panel, Fig. 2e). However, the effect of HITT’s regulation of HIF-1α protein was much more dramatic than its effect on HIF-1α mRNA under the same conditions (Fig. 2b, e). These data indicate that HITT mainly regulates HIF-1α at protein levels. The inhibitory effect of HITT on HIF-1α is likely specific, as HITT overexpression or KD failed to change the mRNA or protein expression levels of its family member HIF-2α (Fig. S4A–B).
HITT inhibits HIF-1α expression and activity. HIF-1α protein was detected by WB of lysates from control and HITT stably transfected lines (a). Quantification of WB bands was conducted by ImageJ (b). HIF-1α mRNA relative to 18 s was detected by qRT-PCR (lower panel, b). The KD efficiency of HITT was confirmed by qRT-PCR (c). HIF-1α protein levels were measured by WB after HITT KD under normoxia in the presence or absence of proteasome inhibitor MG132 (d). Quantification of WB bands was conducted by ImageJ (e). HIF-1α mRNA relative to 18 s was detected by qRT-PCR (lower panel, e). Hypoxic response element (HRE) luciferase activity (f), and expression levels of VEGF and Angiogenin-1 (g), were measured by luciferase reporter assay or qRT-PCR, respectively, under the indicated conditions. Data derived from three independent experiments are presented as mean ± SEM in the bar graph. Values in controls were normalized to 1 (b, c, e–g). *P < 0.05; **P < 0.01; ##P < 0.01, compared with hypoxia-treated controls (b, f–g). Vect. vector control. See also Fig. S4
More importantly, the transcriptional activity of HIF-1α, as evidenced by HRE-dependent reporter activity, was markedly increased by hypoxia and this effect was abolished in a stable HITT-expressing cell line (Fig. 2f), while ectopic HIF-1α expression dampened the effect of HITT (Fig. 2f). In addition, the expression of well-established HIF-1α targets, such as VEGF and angiogenin-1, were increased by hypoxia (Fig. 2g), which was significantly blunted by HITT (Fig. 2g). These data collectively illustrate that HITT inhibits the transcriptional activity of HIF-1α, mainly by regulating its protein levels.
HITT inhibits HIF-1α translation
HITT is unlikely to affect HIF-1α protein stability. As shown, HIF-1α decayed at similar rates after cells were treated with the protein synthesis inhibitor CHX in a HITT-transfected stable line and control line (Fig. 3a). HITT similarly inhibited HIF-1α regardless of MG132 (Fig. 3b). We further compared the efficiency of HIF-1α protein synthesis by polysome fractionation. HITT expression did not change the polysome distribution profile (Fig. 3c), while polysome-associated HIF-1α mRNA was increased with hypoxia (Fig. 3c), which is in line with a previous report [21]. Importantly, ectopic HITT expression compromised hypoxia-induced association of HIF-1α mRNA with polysomes (Fig. 3c), while HITT KD increased their association (Fig. 3d). In contrast, the distribution of GAPDH mRNA remained unaltered, regardless of oxygen concentration or HITT levels (Fig. 3d). The role of HITT in regulating HIF-1α translation was further verified by a Click chemistry and AHA-labeled assay. As shown, HITT overexpression inhibited the newly synthesized HIF-1α protein (Fig. 3e), while HITT KD increased it under hypoxia (Fig. 3f). Newly synthesized α-tubulin served as a negative control, and was not changed by ectopic HITT expression or KD (Fig. 3e, f). Therefore, HITT inhibits HIF-1α mRNA translation.
HITT inhibits HIF-1α translation. a Representative images of HIF-1α protein levels detected by WB (left) and quantification of HIF-1α band intensities by ImageJ (right) for vector control and HITT ectopic expression HCT116 cells, in the presence of CHX, for the indicate time periods under hypoxia. HIF-1α levels in the CHX untreated group were normalized to 1. b HIF-1α protein levels were determined by WB after ectopic HITT expression in the presence or absence of MG132 in HCT116 and HeLa cells under hypoxia. c, d Polysomes in cytoplasmic extracts were fractionated through sucrose gradients, and mRNA levels of HIF-1α and GAPDH in HCT116 cells were measured by qRT-PCR in gradient fractions after HITT overexpression or KD. Representative data, as a percentage of total RNA of interest in the gradient from three independent experiments, are presented. e, f Affinity purification of biotinylated l-azidohomoalanine (AHA)-labeled acutely synthesized proteins from cell lysates. Purified HIF-1α and α-tubulin proteins were detected by WB
HITT acts by serving as a decoy of HIF-1α translational regulator YB-1
Recent evidence has shown that YB-1 promotes HIF-1α translation by directly interacting with its 5′-UTR [34, 35]. Although HITT had no obvious impact on YB-1 protein expression both in vitro and in vivo in HCT116 nude mice xenografts (Fig. 4a and Fig. S5A), eight YB-1-binding sequences were predicted in HITT via a high-stringency search using an online bioinformatics tool (Fig. 4b). YB-1 KD reduced HIF-1α levels and HITT lost its ability to affect HIF-1α expression in YB-1 KD cells (Fig. 4a). This brought us to ask whether HITT regulates HIF-1α transcription by competitively binding with YB-1.
HITT inhibits HIF-1α translation by competing with YB-1 for binding with HIF-1α 5′-UTR. a The expression levels of HIF-1α were measured by WB in HITT stable lines, with or without YB-1 KD, under hypoxia in HCT116 and HeLa cells. The KD efficiency of YB-1 was confirmed by WB. b Schematic of the positions of high-stringency YB-1 binding sites in HITT predicted by a bioinformatics online tool (http://rbpmap.technion.ac.il/). c Biotin-HITT pull-down using HeLa cell lysates showed that YB-1 bands were present in Biotin-HITT precipitates. YB-1-associated HITT (d) and HIF-1α 5′-UTR (e, f), as measured by qRT-PCR following CLIP YB-1 after HITT overexpression or KD, under normoxia or hypoxia, in HCT116 cells, as indicated in the figures. GAPDH mRNA and CLIP IgG were used as negative controls. g Schematic of the positions of high-stringency YB-1 binding sites in HITT (left). CLIP assay was used to detect the binding of YB-1 with full-length or fragmented HITT in 4T1 cells, a cell line with no endogenous HITT, after ectopic expression of the indicated HITT. CLIP IgG and 18 s were used as negative controls (lower panel). Similar expression levels of HITT were confirmed by qRT-PCR (upper panel). h Schematic of full-length and fragmented HITT, and the predicted YB-1 binding sites (upper panel). HIF-1α protein levels were measured by WB after transfection with full-length (1–2050) HITT and MT (1–1029) HITT (lower right panel). Similar expression levels of HITT were confirmed by qRT-PCR (lower left panel). i Schematic of positions of all potential YB-1 binding sites in HIF-1α 5′-UTR predicted by a bioinformatics online tool (http://rbpmap.technion.ac.il/)(Left). Luciferase reporter containing HIF-1α 5′-UTR (1–256) or two truncated mutants (1–138: M1 and 139–256: M2) was used to assess translational efficiency after HITT overexpression with or without YB-1 KD (right). Data derived from three independent experiments are presented as mean ± SEM in the bar graph (d–i). Values in controls were normalized to 1. **P < 0.01 (d–i); #P < 0.05, ##P < 0.01, compared with hypoxia-treated controls (d, e, and i). N.S. not significant (d–g). See also Fig. S5
As shown, YB-1 was co-precipitated with biotin-HITT in vitro (Fig. 4c). Both HITT and the HIF-1α 5′-UTR, but not GAPDH mRNA, were physically associated with YB-1, as revealed by CLIP assays (Figs. 4d, e and S5B). The enrichment of HITT was significantly decreased, while that of the 5′-UTR was increased in response to the hypoxic switch (Figs. 4d, e and S5B). Ectopic HITT expression restored the association of HITT with YB-1 under hypoxia and simultaneously abrogated the hypoxia-promoted association of the HIF-1α 5′-UTR with YB-1 (Figs. 4d, e and S5B). In contrast, HITT KD led to promotion of the association between YB-1 and the HIF-1α 5′-UTR (Fig. 4f). HITT containing the predicted YB-1-binding motif contributed to its associated with YB-1 (Fig. 4g). A HITT mutant that had lost the potential YB-1 binding sites completely lost its regulatory effect towards HIF-1α (Figs. 4h and S5C). These data together suggest that binding of YB-1 is required for HITT-mediated HIF-1α inhibition.
Furthermore, both YB-1 KD and ectopic HITT expression prevented hypoxia-induced WT HIF-1α-5′-UTR-luciferase reporter activity, while no further reduction was detected when they were introduced in combination (Figs. 4i and S5D). The truncated mutant reporters MT1 and MT2, were also induced by hypoxia but to a much lower extent (Figs. 4i and S5D). However, only MT2 responded to YB-1 KD and ectopic HITT expression, suggesting that sequences 2 and 3 in MT2 are the major YB-1 binding sites in HIF-1α-5′-UTR (Figs. 4i and S5D).
Altogether, HITT directly binds with YB-1 and prevents its association with HIF-1α 5′-UTR, leading to reduced HIF-1α translation.
HIF-1α lowers HITT’s stability via a feedback loop
To understand the physiological relevance of HITT in regulating HIF-1α under hypoxia, we examined the change in HITT expression under hypoxia. Interestingly, HITT was significantly reduced under hypoxia and the extent of the reduction was similar to that of the most dramatically altered lncRNAs reported previously [28] (Fig. 5a). In addition, HITT reduction was oxygen dose- and hypoxia duration-dependent (Fig. 5b, c). si-HIF-1α reduced HIF-1α levels and simultaneously restored the expression of HITT (Figs. 5d and S6A). In contrast, ectopic HIF-1α expression lowered HITT (Figs. 5e and S6B). HIF-2α did not show any obvious impact on the expression of HITT (Fig. S6C–D), suggesting that HITT is specifically regulated by HIF-1α via a feedback loop.
HIF-1α lowers HITT’s stability via negative feedback. a Hypoxia-responsive lncRNAs reported previously and HITT were measured by qRT-PCR under hypoxia relative to normoxia. HIF-1α expression was determined by WB, and HITT was measured by qRT-PCR in samples derived from cells subjected to dose- (b) and time-dependent (c) hypoxia treatment. The expression levels of HITHITT and HITT promoter luciferase reporter activity were detected by qRT-PCR and luciferase reporter assay, respectively, in HCT116 cells after HIF-1α KD (d) or overexpression (e) under hypoxia. The efficiency of HIF-1α overexpression and KD was confirmed by WB. f The half-lives of HITT and GAPDH mRNA were measured by qRT-PCR in the presences of ActD in HCT116 cells. Data derived from three independent experiments are represented as mean ± SEM in the bar graph. *P < 0.05, **P < 0.01, N.S. not significant. See also Fig. S6
Hypoxia/HIF-1α-induced HITT suppression is unlikely to occur at the transcriptional level. Firstly, no HRE motif was identified in HITT’s promoter. Secondly, neither ectopic HIF-1α expression nor KD had a significant effect on HITT promoter-driven luciferase activity (Figs. 5d, e and S6A–B). Nonetheless, the half-life of HITT, but not GAPDH mRNA, was decreased by hypoxia. HIF-1α KD blocked the effect of hypoxia (Figs. 5f and S6E). Consistently, comparable HITT levels were detected under normoxia and hypoxia in HIF-1α-deficient 786-O cells (Fig. S6F–G). These data demonstrate that HIF-1α governs HITT’s stability.
HIF-1α promotes HITT’s decay by MiR-205-mediated RNA degradation
We next sought to investigate the mechanisms underlying HIF-1α-mediated HITT destabilization. Two independent bioinformatics tools predicted that HITT is a candidate target of a number of microRNAs, of which MiR-205 was the most likely (Fig. 6a). In addition, among the microRNAs that indeed possess the ability to inhibit HITT expression as validated by qRT-PCR (Fig. S7A), only MiR-205 was induced over twofold by hypoxia (Fig. S7B). We also found that the expression of MiR-205 was dependent on the levels of HIF-1α, because hypoxia-induced MiR-205 was completely abolished by siRNA-mediated HIF-1α KD in both HCT116 and HeLa cells (Fig. 6b). Thus, it is possible that MiR-205 mediates HIF-1α-induced downstream events, such as HITT destabilization. Therefore, we further investigated the impacts of MiR-205 on HITT expression. As shown, a MiR-205 inhibitor prevented hypoxia-induced MiR-205 and also restored reduced levels of HITT under hypoxia (Figs. 6c and S7C), whereas the induction of MiR-205 under normoxia by transfecting MiR-205 mimics significantly reduced HITT (Fig. 6d). MiR-205 mimics also abolished HIF-1α KD-induced HITT recovery under hypoxia (Figs. 6e and S7D).
HIF-1α promotes HITT decay via MiR-205-mediated RNA degradation. a MiR-205 was predicted to be a potent microRNA that can bind HITT by two independent bioinformatics tools: http://www.mircode.org/index.php and http://annolnc.cbi.pku.edu.cn/index.jsp. b Expression levels of MiR-205 normalized to U6 were measured by qRT-PCR after HIF-1α KD under hypoxia. Relative expression levels of HITT were determined by qRT-PCR after transfection of MiR-205 inhibitors (Inh., c) or MiR-205 mimics (d) into HCT116 cells. MiR-205 levels normalized to U6 were measured by qRT-PCR. e The expression levels of HITT were determined by qRT-PCR in HCT116 cells after HIF-1α KD with or without MiR-205 inhibitor treatment. f–h Schematic of the hypothetical duplexes formed by interactions between the binding site in HITT (top), MiR-205 (middle), and mutated HITT (bottom) (f). The activities of wild type (WT) or MiR-205-binding defective mutant (MT) HITT luciferase reporters were detected in HCT116 cells after transfection of MiR-205 inhibitor (Inh., g) or mimics (h) under normoxia or hypoxia, as indicated in the figures. i Biotinylated precursor MiR-205 was transfected into HCT116 cells. MiR-205-associated endogenous HITT relative to total HITT was determined by qRT-PCR following CLIP biotin. CLIP GAPDH and nonbiotinylated MiR-205 were used as negative controls. j The expression levels of HITT were determined by qRT-PCR in HCT116 cells after Ago2 KD (right). KD efficiency was confirmed by WB (left). k Ago2-associated HITT levels relative to total HITT were determined by qRT-PCR following formaldehyde cross-linked IP of Ago2 in HCT116 cells under normoxia and hypoxia, with or without treatment with MiR-205 inhibitors (Inh.). GAPDH and IgG were used as negative controls. l Relative expression levels of HITT determined by qRT-PCR after treatment with MiR-205 mimics in control and Ago2 KD HCT116 cells. m HITT’s half-life was determined by qRT-PCR after treatment with MiR-205 inhibitor (Inh.) alone, Ago2 KD alone, or a combination of the two in the presences of RNA synthesis inhibitor ActD under hypoxia in HCT116 cells. Data derived from three independent experiments are presented as mean ± SEM in the bar graph. Values in controls were normalized to 1. *P < 0.05; **P < 0.01; #P < 0.05, compared with hypoxia treatment controls (c, e, g, k). N.S. not significant (e, g–i, k, l). See also Fig. S7
Furthermore, the activity of WT HITT luciferase reporter was decreased by hypoxia. MiR-205 inhibitor completely reversed the inhibitory effect of hypoxia on WT HITT luciferase activity (Figs. 6g and S7E). However, neither hypoxia nor the MiR-205 inhibitor influenced the luciferase activity of the MiR-205-binding defective MT reporter (Figs. 6g and S7E). In contrast, MiR-205 significantly reduced WT, but not MT, HITT luciferase activity (Figs. 6h and S7F). Furthermore, the direct association between MiR-205 and HITT was verified by pull-down biotinylated precursor MiR-205 followed by qRT-PCR analysis of HITT (Fig. 6i). Moreover, KD of Ago2, an important component of the RNA-induced silencing complex/MiR-205, produced a similar effect on HITT as MiR-205 inhibitors (Figs. 6j and S7G). RIP Ago2 showed that HITT was associated with Ago2 (Fig. 6k). Although the expression levels of Ago2 were not changed under hypoxia (Fig. S7H), levels of Ago2-associated HITT were increased (Fig. 6k), under which conditions HITT was destabilized. A MiR-205 inhibitor reduced the hypoxia-induced association between Ago2 and HITT (Fig. 6k), with MiR-205 losing its ability to affect HITT expression in Ago2 KD cells (Fig. 6l). Consistently, the MiR-205 inhibitor or Ago2 KD prolonged HITT’s half-life, while no further change was observed when they were combined (Fig. 6m).
Collectively, these data suggest a model where Ago2/miR-205 physiologically binds with HITT, facilitating MiR-205-mediated HITT destabilization.
HITT downregulation is correlated with high levels of HIF-1α in human colon cancer tissues
Having uncovered a novel HITT–HIF-1α axis, we verified the correlation between HIF-1α and HITT expression in human cancer tissues. The protein levels of HIF-1α and its target, VEGF, were examined in human colon cancer tissues and matched adjacent controls (n = 46). HIF-1α and VEGF protein levels were commonly increased in colon cancer tissues compared with their matched normal controls (Fig. 7a–c). Interestingly, reduced HITT levels were significantly correlated with increased HIF-1α protein and VEGF mRNA expression (Fig. 7d, e). These findings, combined with our in vitro and xenograft data, demonstrate that HITT regulates HIF-1α at the translational level, which results in reduced expression of HIF-1α and its targets.
The association of HITT with HIF-1α and VEGF levels in human colon cancer tissues. a Representative image of HIF-1α protein expression in human colon cancer tissue samples (T) and paired adjacent normal controls (N). The average fold-change of HIF-1α protein (b) and VEGF mRNA (c) levels normalized to 18S rRNA were detected by qRT-PCR and WB in human colorectal cancer tissue samples, (T) and paired adjacent normal controls (n = 46). Correlation analysis of HITT with HIF-1α protein (d) and VEGF mRNA (e). f Schematic diagram of the proposed HITT–HIF-1α feedback loop that controls hypoxia-induced angiogenesis and tumor growth. Under normoxia, HITT does not undergo HIF-1α/MiR-205-mediated destabilization. Thus, HITT acts as a molecular decoy for YB-1, such that it is associated with YB-1 via favorable binding sequences, titrating YB-1 away from HIF-1α 5′-UTR, resulting in the attenuation of HIF-1α protein translation. Under hypoxia, HITT is targeted by Ago2/MiR-205 for degradation in a HIF-1α-dependent manner, consequently diminishing the inhibitory effect of HITT on HIF-1α transcription, resulting in angiogenesis and tumor growth
Discussion
The discovery of the functional significance of protein-coding genes has led to great successes in the development of targeted therapies against cancer. Increasing evidence has suggested that nonprotein-coding genes account for the overwhelming majority of RNA transcripts and can drive important cancer phenotypes. Such discoveries have sparked great interest in the functions and regulation of lncRNAs [9, 36], with a view to improving the diagnosis and treatment of cancers.
HITT is one such as yet uncharacterized lncRNA in the human genome. We found that the expression of HITT was downregulated in multiple human tumors. Although it had no obvious impact on tumor growth in vitro, HITT elicited a marked suppressive effect in vivo by influencing angiogenesis. The key mechanism that we elucidated as accounting for this function of HITT is that it inhibits HIF-1α expression (Fig. 7f). The significance of our in vitro and animal experiment data is further strengthened by the fact that a reverse association between HITT and key components of this pathway, i.e. HIF-1α protein and VEGF transcripts, were detected in human colon cancer tissues. Of note, angiogenesis and increased HIF-1α levels are associated with poor prognosis in multiple cancers [11, 37]. In line with this notion, HITT levels were inversely correlated with advanced stages of colon cancers. These data encourage the future development of HITT-based cancer therapies for patients with increased HIF-1α, who are at high risk of metastasis [38], an outcome that results in ~90% of cancer-related deaths.
HIF-1α is a core transcription factor for the adaptive survival of cancer cells under hypoxia. The mechanisms underlying HIF-1α protein stability have been studied extensively and are also considered to be critical for the accumulation of HIF-1α under hypoxia [39]. Here, we have demonstrated that HITT is an important HIF-1α negative regulator. Intriguingly, the action of HITT is mainly fulfilled via its regulating of HIF-1α translation, but not its protein stability (Fig. 7f). This mechanism is distinct from lncRNA-mediated HIF-1α regulation reported previously, through which lncRNAs mainly effect the HIF-1α protein stabilization machinery [26] or sponging microRNAs target HIF-1α [40].
LncRNAs can act as ‘molecular decoys for RNA-binding proteins, which are themselves regulatory factors of gene expression [41, 42]. YB-1 is a well-characterized translational regulator. It has recently been reported to regulate HIF-1α translation. Like the typical YB-1 target SNAIL1 [43], HIF-1α 5′-UTR contains a complex stem-loop structure. Thereby, El-Naggar et al. have proposed that YB-1 binding may unwind such secondary structures, leading to the promotion of HIF-1α translation [34]. Our study reveals an additional level of regulation for YB-1-mediated HIF-1α translation via lncRNA. HITT prevents or interrupts the interaction of YB-1 with HIF-1α 5′-UTR to suppress HIF-1α expression. In fact, the lncRNAs CAR10 [44], TP53TG1 [45], and GAS5 [46] have been recently documented as regulators of YB-1 activity. These lncRNAs act by controlling YB-1’s localization or protein stability. In contrast to these lncRNAs, HITT inhibits YB-1 via a distinct mechanism by acting as a protein decoy. These findings combined suggest that lncRNAs may be important components of the regulatory machinery for modulating YB-1’s activity by diverse means. It will be also interesting to explore in the future whether HITT is involved in regulating multiple functions of YB-1 [47].
Given its importance in sensing hypoxic tension, it is not surprising that cells utilize feedback mechanisms to precisely control HIF-1α signals. In fact, a number of hypoxia-responsive lncRNAs contain HREs and subjected to direct regulation by HIF-1α [48]. However, the feedback regulation of HIF-1α by HITT, identified here, is unlikely to occur through a direct mechanism [49]. Our data suggest that MiR-205 is a primary microRNA to inhibit HITT under hypoxia. However, HITT may be regulated by additional microRNAs, as more microRNAs are predicted to undergo complementary base pairing with HITT. It will be interesting to explore the roles of these microRNAs on HITT expression under different physiological and pathological conditions in future studies. Furthermore, besides microRNAs, lncRNAs may also be regulated by typical transcription factors [26] or RNA-binding proteins [33]. The precise molecular mechanism of the downregulation of HITT requires further research.
Taken together, we have identified HITT, a novel lncRNA. Our data highlight the significance of the newly identified HITT–HIF-1α axis in regulating angiogenesis and tumor growth. Detailed dissection of the molecular mechanisms of HITT-mediated HIF-1α synthesis have provided insights into novel strategies to inhibit HIF-1α by restoring HITT activity.
References
Ulitsky I, Bartel DP. lincRNAs: genomics, evolution, and mechanisms. Cell. 2013;154:26–46.
Iyer MK, Niknafs YS, Malik R, Singhal U, Sahu A, Hosono Y, et al. The landscape of long noncoding RNAs in the human transcriptome. Nat Genet. 2015;47:199–208.
Wang KC, Chang HY. Molecular mechanisms of long noncoding RNAs. Mol Cell. 2011;43:904–14.
Guttman M, Rinn JL. Modular regulatory principles of large non-coding RNAs. Nature. 2012;482:339–46.
Huarte M. The emerging role of lncRNAs in cancer. Nat Med. 2015;21:1253–61.
Schmitt AM, Chang HY. Long noncoding RNAs in cancer pathways. Cancer Cell. 2016;29:452–63.
Prensner JR, Chinnaiyan AM. The emergence of lncRNAs in cancer biology. Cancer Discov. 2011;1:391–407.
Evans JR, Feng FY, Chinnaiyan AM. The bright side of dark matter: lncRNAs in cancer. J Clin Investig. 2016;126:2775–82.
Adams BD, Parsons C, Walker L, Zhang WC, Slack FJ. Targeting noncoding RNAs in disease. J Clin Investig. 2017;127:761–71.
Rankin EB, Giaccia AJ. Hypoxic control of metastasis. Science. 2016;352:175–80.
Wilson WR, Hay MP. Targeting hypoxia in cancer therapy. Nat Rev Cancer. 2011;11:393–410.
Ma L, Chen ZM, Li XY, Wang XJ, Shou JX, Fu XD. Nucleostemin and ASPP2 expression is correlated with pituitary adenoma proliferation. Oncol Lett. 2013;6:1313–8.
Jain RK. Antiangiogenesis strategies revisited: from starving tumors to alleviating hypoxia. Cancer Cell. 2014;26:605–22.
Potente M, Gerhardt H, Carmeliet P. Basic and therapeutic aspects of angiogenesis. Cell. 2011;146:873–87.
Birner P, Schindl M, Obermair A, Plank C, Breitenecker G, Oberhuber G. Overexpression of hypoxia-inducible factor 1alpha is a marker for an unfavorable prognosis in early-stage invasive cervical cancer. Cancer Res. 2000;60:4693–6.
Auyeung KK, Ko JK. Angiogenesis and oxidative stress in metastatic tumor progression: pathogenesis and novel therapeutic approach of colon cancer. Curr Pharm Des. 2017;23:3952–61.
Semenza GL. Hypoxia-inducible factors: mediators of cancer progression and targets for cancer therapy. Trends Pharm Sci. 2012;33:207–14.
Semenza GL. Targeting HIF-1 for cancer therapy. Nat Rev Cancer. 2003;3:721–32.
Dengler VL, Galbraith M, Espinosa JM. Transcriptional regulation by hypoxia inducible factors. Crit Rev Biochem Mol Biol. 2014;49:1–15.
Galban S, Kuwano Y, Pullmann R Jr, Martindale JL, Kim HH, Lal A, et al. RNA-binding proteins HuR and PTB promote the translation of hypoxia-inducible factor 1alpha. Mol Cell Biol. 2008;28:93–107.
Hui AS, Bauer AL, Striet JB, Schnell PO, Czyzyk-Krzeska MF. Calcium signaling stimulates translation of HIF-alpha during hypoxia. FASEB J. 2006;20:466–75.
Frede S, Stockmann C, Freitag P, Fandrey J. Bacterial lipopolysaccharide induces HIF-1 activation in human monocytes via p44/42 MAPK and NF-kappaB. Biochemical J. 2006;396:517–27.
Mi C, Ma J, Wang KS, Zuo HX, Wang Z, Li MY, et al. Imperatorin suppresses proliferation and angiogenesis of human colon cancer cell by targeting HIF-1alpha via the mTOR/p70S6K/4E-BP1 and MAPK pathways. J Ethnopharmacol. 2017;203:27–38.
Zhou C, Huang C, Wang J, Huang H, Li J, Xie Q, et al. LncRNA MEG3 downregulation mediated by DNMT3b contributes to nickel malignant transformation of human bronchial epithelial cells via modulating PHLPP1 transcription and HIF-1alpha translation. Oncogene. 2017;36:3878–89.
Singh L, Aldosary S, Saeedan AS, Ansari MN, Kaithwas G. Prolyl hydroxylase 2: a promising target to inhibit hypoxia-induced cellular metabolism in cancer cells. Drug Discov Today. 2018;23:1873–82.
Yang F, Zhang H, Mei Y, Wu M. Reciprocal regulation of HIF-1alpha and lincRNA-p21 modulates the Warburg effect. Mol cell. 2014;53:88–100.
Kapinova A, Kubatka P, Zubor P, Golubnitschaja O, Dankova Z, Uramova S, et al. The hypoxia-responsive long non-coding RNAs may impact on the tumor biology and subsequent management of breast cancer. Biomedicine Pharmacother. 2018;99:51–8.
Choudhry H, Harris AL. Advances in hypoxia-inducible factor biology. Cell Metab. 2018;27:281–98.
Taniwaki M, Matsuda F, Jauch A, Nishida K, Takashima T, Tagawa S, et al. Detection of 14q32 translocations in B-cell malignancies by in situ hybridization with yeast artificial chromosome clones containing the human IgH gene locus. Blood. 1994;83:2962–9.
Bando T, Kato Y, Ihara Y, Yamagishi F, Tsukada K, Isobe M. Loss of heterozygosity of 14q32 in colorectal carcinoma. Cancer Genet Cytogenet. 1999;111:161–5.
Dodd LE, Sengupta S, Chen IH, den Boon JA, Cheng YJ, Westra W, et al. Genes involved in DNA repair and nitrosamine metabolism and those located on chromosome 14q32 are dysregulated in nasopharyngeal carcinoma. Cancer Epidemiol Biomarkers Prev. 2006;15:2216–25.
Zhang Y, He Q, Hu Z, Feng Y, Fan L, Tang Z, et al. Long noncoding RNA LINP1 regulates repair of DNA double-strand breaks in triple-negative breast cancer. Nat Struct Mol Biol. 2016;23:522–30.
Huang J, Zhou N, Watabe K, Lu Z, Wu F, Xu M, et al. Long non-coding RNA UCA1 promotes breast tumor growth by suppression of p27 (Kip1). Cell Death Dis. 2014;5:e1008.
El-Naggar AM, Veinotte CJ, Cheng H, Grunewald TG, Negri GL, Somasekharan SP, et al. Translational activation of HIF1alpha by YB-1 promotes sarcoma metastasis. Cancer Cell. 2015;27:682–97.
Ivanova IG, Park CV, Yemm AI, Kenneth NS. PERK/eIF2alpha signaling inhibits HIF-induced gene expression during the unfolded protein response via YB1-dependent regulation of HIF1alpha translation. Nucleic Acids Res. 2018;46:3878–90.
Velagapudi SP, Cameron MD, Haga CL, Rosenberg LH, Lafitte M, Duckett DR, et al. Design of a small molecule against an oncogenic noncoding RNA. PNAS. 2016;113:5898–903.
Baba Y, Nosho K, Shima K, Irahara N, Chan AT, Meyerhardt JA, et al. HIF1A overexpression is associated with poor prognosis in a cohort of 731 colorectal cancers. Am J Pathol. 2010;176:2292–301.
Unwith S, Zhao H, Hennah L, Ma D. The potential role of HIF on tumour progression and dissemination. Int J cancer. 2015;136:2491–503.
Shenoy N, Pagliaro L. Sequential pathogenesis of metastatic VHL mutant clear cell renal cell carcinoma: putting it together with a translational perspective. Ann Oncol. 2016;27:1685–95.
Takahashi K, Yan IK, Haga H, Patel T. Modulation of hypoxia-signaling pathways by extracellular linc-RoR. J Cell Sci. 2014;127:1585–94.
Zhang J, Zhang Q. VHL and hypoxia signaling: beyond HIF in cancer. Biomedicines. 2018;6:1–13.
Yang J, Harris AL, Davidoff AM. Hypoxia and hormone-mediated pathways converge at the histone demethylase KDM4B in cancer. Int J Mol Sci. 2018;19:1–11.
Evdokimova V, Tognon C, Ng T, Ruzanov P, Melnyk N, Fink D, et al. Translational activation of snail1 and other developmentally regulated transcription factors by YB-1 promotes an epithelial-mesenchymal transition. Cancer Cell. 2009;15:402–15.
Cox TR, Erler JT, Rumney RMH. Established models and new paradigms for hypoxia-driven cancer-associated bone disease. Calcif Tissue Int. 2018;102:163–73.
Salem A, Asselin MC, Reymen B, Jackson A, Lambin P, West CML, et al. Targeting hypoxia to improve non-small cell lung cancer outcome. J Natl Cancer Inst. 2018;110:14–30.
Jahanban-Esfahlan R, de la Guardia M, Ahmadi D, Yousefi B. Modulating tumor hypoxia by nanomedicine for effective cancer therapy. J Cell Physiol. 2018;233:2019–31.
Lasham A, Print CG, Woolley AG, Dunn SE, Braithwaite AW. YB-1: oncoprotein, prognostic marker and therapeutic target? Biochemical J. 2013;449:11–23.
Choudhry H, Harris AL, McIntyre A. The tumour hypoxia induced non-coding transcriptome. Mol Asp Med. 2016;47-48:35–53.
Wang X, Yu M, Zhao K, He M, Ge W, Sun Y, et al. Upregulation of MiR-205 under hypoxia promotes epithelial-mesenchymal transition by targeting ASPP2. Cell Death Dis. 2016;7:e2517.
Acknowledgements
This work was funded by the National Natural Science Foundation of China (grant numbers 31301131, 31741084, and 31871389) and the Basic Science Foundation of Science and Technology Innovation Commission in Shenzhen (grant number JCYJ20170811154452255).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by I. Amelio
Rights and permissions
About this article
Cite this article
Wang, X., Li, L., Zhao, K. et al. A novel LncRNA HITT forms a regulatory loop with HIF-1α to modulate angiogenesis and tumor growth. Cell Death Differ 27, 1431–1446 (2020). https://doi.org/10.1038/s41418-019-0449-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-019-0449-8
This article is cited by
-
LncRNA ZNF674-AS1 drives cell growth and inhibits cisplatin-induced pyroptosis via up-regulating CA9 in neuroblastoma
Cell Death & Disease (2024)
-
Hypoxia and low-glucose environments co-induced HGDILnc1 promote glycolysis and angiogenesis
Cell Death Discovery (2024)
-
The strict regulation of HIF-1α by non-coding RNAs: new insight towards proliferation, metastasis, and therapeutic resistance strategies
Cancer and Metastasis Reviews (2024)
-
Lnc AC016727.1/BACH1/HIF-1 α signal loop promotes the progression of non-small cell lung cancer
Journal of Experimental & Clinical Cancer Research (2023)
-
LINC00665 activating Wnt3a/β-catenin signaling by bond with YBX1 promotes gastric cancer proliferation and metastasis
Cancer Gene Therapy (2023)