Abstract
TEAD1 (TEA domain transcription factor 1), a transcription factor known for the functional output of Hippo signaling, is important for tumorigenesis. However, the role of TEAD1 in the development of vascular smooth muscle cell (VSMC) is unknown. To investigate cell-specific role of Tead1, we generated cardiomyocyte (CMC) and VSMC-specific Tead1 knockout mice. We found CMC/VSMC-specific deletion of Tead1 led to embryonic lethality by E14.5 in mice due to hypoplastic cardiac and vascular walls, as a result of impaired CMC and VSMC proliferation. Whole transcriptome analysis revealed that deletion of Tead1 in CMCs/VSMCs downregulated expression of muscle contractile genes and key transcription factors including Pitx2c and myocardin. In vitro studies demonstrated that PITX2c and myocardin rescued TEAD1-dependent defects in VSMC differentiation. We further identified Pitx2c as a novel transcriptional target of TEAD1, and PITX2c exhibited functional synergy with myocardin by directly interacting with myocardin, leading to augment the differentiation of VSMC. In summary, our study reveals a critical role of Tead1 in cardiovascular development in mice, but also identifies a novel regulatory mechanism, whereby Tead1 functions upstream of the genetic regulatory hierarchy for establishing smooth muscle contractile phenotype.
Similar content being viewed by others
Introduction
Vasculogenesis is a critical step in cardiovascular development. In response to various extrinsic cues, vascular smooth muscle cell (VSMC) progenitors simultaneously undergo migration, proliferation, and differentiation to form the medial layers within the vascular wall [1]. During the process of development, VSMCs acquire a unique repertoire of specialized markers such as SM22α (TAGLN) [2], calponin (CNN1) [3], SM α-actin (ACTA2) [4], Hic-5 (TGFB1I1) [5], leiomodin 1 (LMOD1) [6], 130-kDa myosin light chain kinase (MYLK) [7], and SM myosin heavy chain (MYH11) [8] that collectively confer a differentiated phenotype. VSMCs are the major structural component of the vascular wall, but also are responsible for vasomotion through the abundantly expressed contractile proteins. Defects in the differentiation of VSMCs lead to congenital vascular malformations and this is seen in humans with genetic predispositions for vascular diseases. Therefore, identification of the factors responsible for VSMC differentiation is essential to a greater understanding of mechanisms underlying VSMC-driven vascular disease in humans [9].
The transcription factor serum response factor (SRF) plays a central role in regulating expression of smooth muscle-specific contractile genes through its binding to a specific DNA sequence, called CArG box [10]. CArG boxes can be found in the regulatory regions of almost every smooth muscle-specific gene [11]. The binding of SRF to the CArG box provides an initiating platform to recruit a variety of transcription cofactors that collectively regulate smooth muscle-specific gene expression [12]. Of the many cofactors that interact with SRF, PITX2, and myocardin (MYOCD) have been shown to play critical roles in VSMC differentiation [13,14,15]. However, the functional relationship between PITX2 and MYOCD in the regulation of smooth muscle-specific gene-expression remain elusive.
Initially identified in Drosophila melanogaster, the Hippo signaling pathway is crucial for organ size control and tumorigenesis [16,17,18]. The mammalian core components of the Hippo pathway include MST1/2 (mammalian Ste20-like kinase 1 or 2), LATS1/2 (large tumor suppressor 1 or 2), signaling down-stream effector YAP1 (yes-associated protein 1) or TAZ (transcriptional coactivator with PDZ-binding motif), and the YAP1/TAZ nuclear binding partners, TEAD1–4 (TEA domain transcription factor). Our previous studies have shown that the Hippo signaling effector, YAP1 not only plays a critical role in neointimal formation after arterial injury, but also is critical for cardiomyocyte (CMC) and VSMC proliferation during cardiovascular development in mice [19,20,21]. YAP1 or TAZ on its own cannot initiate transcription due to lack of a DNA binding domain, therefore the function of YAP1 or TAZ is dependent upon binding to many transcription factors including TEADs. Current dogma is that, the functional effects of YAP1 are mediated primarily through interaction with TEAD transcription factors, which bind to a consensus DNA sequence, 5′-CATTC-3′, known as the MCAT (muscle CAT) element [22,23,24,25]. However, a recent study suggested that TEAD itself is actively regulated by other factors, as evidenced by an analysis of YAP1 occupancy on DNA which overlapped with only a very small subset of TEAD binding sites [26]. Therefore, YAP1 and TEADs may play independent roles that may vary with cellular context and developmental stage. The TEAD family of proteins consist of four members, TEAD1, TEAD2, TEAD3, and TEAD4 [27, 28]. Among these TEAD family proteins, we recently reported that TEAD1 is the most abundant family member in VSMCs and plays a critical role in VSMC de-differentiation in vitro [29]. However, the specific role of TEAD1 in VSMC development in vivo has not been elucidated.
In this study, we found that loss of Tead1 in CMCs and VSMCs resulted in embryonic lethality at E14.5, due to impaired proliferation and differentiation of CMCs and VSMCs. Our study, for the first time, uncovers a critical role of the Tead1 in cardiovascular development in mice.
Materials and methods
Mouse breeding and genotyping
Tead1 flox (F) mice were generated as we previously described in details [30]. Tagln-Cre mice (B6.Cg-Tg(SM22α-Cre)1Her/J; Stock No: 017491) were purchased from the Jackson Laboratory [31]. Although TAGLN is well documented as a marker of smooth muscles, it is transiently expressed in CMCs between E8.5 and E12.5 in mice [2]. Thus, Cre recombinase directed by Tagln gene promoter will be expressed in CMCs in addition to VSMCs during embryogenesis [20]. The dual fluorescence reporter mTmG (membrane-targeted tandem dimer Tomato, mTomato or mT; membrane-targeted green fluorescent protein, mGFP or mG) mice (B6.129 (Cg)-Gt(ROSA)26Sortm4(ACTB-tdTomato,-EGFP)Luo/J; Stock No: 007676) were purchased from the Jackson Laboratory [32]. To generate mice in which the Cre transgene induces CMC/VSMC-specific KO of Tead1, we first generated Tagln-Cre+/TEAD1F/W mice, by crossing Tagln-Cre+ male mice with Tead1F/F female mice. Then Tagln-Cre+/TEAD1F/W male mice were bred with Tead1F/F/mTmG+/+ female mice to achieve CMC/VSMC-specific Tead1 deletion with green fluorescent protein (GFP) tag. This breeding strategy yielded 25% progeny with the homozygous CMC/VSMC-specific deletion (Tagln-Cre+/Tead1F/F/mTmG+/−) KO and 25% Tagln-Cre+/Tead1F/W/mTmG+/−, 25% Tagln-Cre−/TEAD1F/F/mTmG+/−, 25% Tagln-Cre−/TEAD1F/W/mTmG+/− littermate as controls. All mice used in this study were maintained on a C57BL/6 background. Embryonic day (E) 0.5 was defined as noon of the day when the vaginal plug was detected. Genotyping of these mice was performed as previously described and primer information is further provided in Online Table I [30, 31]. The use of experimental animals has been approved by the IACUC at Augusta University in accordance with NIH guidelines.
Sections, hematoxylin/eosin staining, immunofluorescence
Isolation of embryos, embedding in paraffin, sectioning and hematoxylin/eosin (HE) staining were performed following standard protocol as we previously described [5, 19, 20]. For immunofluorescence (IF), sections were deparaffinized and antigen retrieval was done by microwaving to heat at 98 °C for 10 min in citric acid buffer (10 mM, pH 6.0). After goat serum (10%, Invitrogen) blocking for 30 min, sections were then incubated with anti-ACTA2 (SM α-actin, Sigma, A2547, mouse, 1:100), anti-MYH11 (SM MHC, Biomedical Technologies Inc, BT562, rabbit, 1:30), anti-MYH1E (MF-20, obtained from Developmental Studies Hybridoma Bank, 1:200 dilution), anti-phospho-H3F3A, (H3 histone family member 3A, Ser10, Millipore, 06-570, rabbit, 1:50), anti-MKI67 (Thermo Scientific, RM-9106, rabbit, 1:30), and anti-GFP (Abcam, ab290, rabbit, 1:1000) antibodies. Subsequently, sections were stained with secondary antibody (488 nm anti-mouse secondary antibody, 568 nm anti-rabbit secondary antibody, 647 nm anti-mouse or rabbit secondary antibody, 1:250, Invitrogen) and diluted in blocking buffer for 1 h at room temperature. Sections were then immersed with mounting medium (ProLong Gold anti-fade reagent with DAPI, Invitrogen) to stain nuclei. Images were collected by using a confocal microscopy (LSM 780 upright, Zeiss) at the imaging core of Augusta University. The thickness of myocardium wall, arterial wall and arterial lumen area was measured by Image J software (NIH).
Optimal cutting temperature embedding, directly visualizing fluorescence in mTmG reporter mice and TEAD1 IF using tyramide signal amplification staining
Embryos were dissected from timed pregnant mice and fixed with 4% paraformaldehyde on ice for 30 min. After washing with ice-cold phosphate-buffered saline (PBS) twice for 10 min each time, embryos were then soaked in 30% sucrose/PBS on a shaker in 4 °C overnight until the embryos settled down. The processed embryos were subsequently embedded in optimal cutting temperature (OCT) and sectioned in 8-μm thickness. For directly visualizing fluorescence signal of mTmG reporter, OCT sections were air dried for 10–15 min then washed with PBS (3 times × 5 min/wash) to remove OCT. The slides were then briefly air dried and covered with Prolong DAPI mounting medium (Invitrogen) for counter-staining nuclei. GFP, Tomato, and DAPI signals were then collected by a confocal microscopy (LSM 780 upright, Zeiss) using 488, 594, 405-nm excitation channels, respectively. For IF staining of TEAD1 in embryos, a tyramide signal amplification (TSA) kit (TSA plus DIG, Perkin Elmer, cat. #: NEL748001KT) was used to amplify the TEAD1 signal. Briefly, after antigen retrieval as described above, sections were treated with H2O2 (30 μL 30% H2O2 diluted into 1 ml PBS) for 15 min and then incubated with blocking buffer for 30 min (2.5% horse serum + 0.1% Tween-20) at room temperature. Following immunostaining with other antibodies as described above, anti-TEAD1 antibody (Abcam, ab133533, rabbit, 1:500) was incubated with the sections overnight at 4 °C. After incubated with HRP labeled anti-rabbit secondary antibody (Vector Labs, cat. #: MP-7401) for 30 min at room temperature, TSA plus DIG Amplification working solution (1:50 dilution) was applied for 5 min at room temperature and further incubated with cy5 fluorescence-labeled anti-DIG antibody (TSA Plus Cyanine 5, NEL745E001KT, 1:250 dilution) for 30 min at room temperature. Finally, sections were mounted with Prolong Gold mounting medium with DAPI (Invitrogen) and images were acquired by a confocal microscopy (LSM 780 upright, Zeiss).
EdU incorporation assay
Before embryos were harvested from the timed pregnant mice, EdU (Invitrogen, A10044, 5mg/ml stock, 10 μl/g body weight of mouse) was peritoneally injected. Two hour later embryos were then embedded in OCT and sectioned. IF was performed as described above and EdU was detected using Click-iT EdU Imaging Kits (Invitrogen, cat. #: C10339) following the manufacturer’s protocol.
Preparation of primary aortic VSMCs from mouse embryos or adult mice
Mouse embryonic aortic VSMCs were prepared and cultured as described in our previous report [20]. Briefly, dorsal aortae from mouse E13.5 embryos were dissected and isolated. After removing periadventitial tissues under a dissecting fluorescence microscope, aortae were minced into small pieces by scissors and digested in 0.25% trypsin digestion buffer. The dispersed cells were further isolated by a flow cytometry cell sorter (Beckman Coulter MoFlo XDP) and then placed in 6-well plate containing 10% fetal bovine serum (FBS) Dulbecco's Modified Eagle Medium (DMEM) medium. After passaged 3 times, the cultured VSMCs were grown on coverslips overnight and then fixed with 4% paraformaldehyde. Following permeabilization with Triton X-100, the cells then were immersed in ProLong Gold mounting medium with DAPI (Invitrogen) to stain nuclei. Primary VSMCs were isolated from adult Tead1 floxed allele mice as previously reported [19, 21]. All embryonic and adult VSMCs used for further experiments were prior to passage 7. Embryonic or adult VSMCs from Tead1 floxed mice were infected with adenovirus expressing control GFP or Cre as described previously [21, 29, 33]. For TGF-β treatment experiments, embryonic VSMCs were grown to 80–90% confluence and infected with GFP or Cre for 24 h. Following overnight serum-starvation, cells were then treated with recombinant mouse TGF-β1 (10 ng/ml, transforming growth factor beta 1, R&D, cat. #: 7666-MB-005) for 48 h and harvested for Western blotting. Cells treated with vehicle served as controls.
Isolation of mouse embryonic fibroblasts
Mouse embryonic fibroblasts (MEFs) were prepared by a standard protocol. Briefly, E14.5 embryos were dissected and all inner organs, head, and limbs were removed. The remaining tissues were minced by scissors in 0.25% trypsin digestion buffer. Subsequently, the minced tissue was collected and incubated in 0.25% trypsin at 4 °C overnight and then cells were plated on 15-cm plates in 10% FBS/DMEM medium. After three passages, MEFs were plated for the luciferase assay or viral infections.
Next-generation sequencing and pathway enrichment analysis
GFP-positive heterozygous control and Tead1 KO cardiac and dorsal aorta tissues were dissected under a fluorescent stereoscope. Subsequently, the GFP-positive CMCs and VSMCs were dissociated by enzymatic digestion and collected by fluorescence activated cell sorting (FACS). Total RNA was then extracted from the purified GFP-positive cells using the RNeasy Micro Kit (Qiagen) and subjected to whole transcriptome RNA-seq analysis. Differential gene-expression analysis was performed at the Genome Technology Access Center at Washington University. Following amplification of the input RNA by Sigma SeqPlex RNA amplification kit, the library was prepared and single-end sequencing in 50-bp length was performed on a HiSeq 3000 system (Illumina). Adapter sequence and low-quality reads were removed using Trimmomatic 0.32 [34]. Pass-filtered reads were then mapped to Ensembl mouse reference genome mm10 using TopHat version 2.1.0 [35]. The genes annotated in Ensembl GRCm38 were quantified with HTSeq 0.6.1 [36] and only those with count ≥10 in at least 3 samples were considered as expressed genes and used for subsequent analysis. Differential expression analysis was performed with the R package edgeR (https://www.r-project.org) [37]. Cutoff values of fold change greater than 1.2 and P value less than 0.05 were considered significant between control and KO groups (3 embryos each group). Fragments per kilobase of transcript per million fragments mapped, which corrects for variations in contig length and read depth between samples, was calculated from raw counts for each sample (Online Table II) [38]. Volcano plots were generated using R program. DAVID was used to conduct GO (Gene Ontology) and KEGG (Kyoto Encyclopedia of Genes and Genomes) pathway analysis for up or downregulated genes (https://david.ncifcrf.gov/). Terms with P value <0.005 were considered significant. The chord plot to show the association between downregulated genes and enriched GO terms was generated graphically with GOplot [39]. Gene set enrichment analysis (GSEA) was performed using GSEA software (http://software.broadinstitute.org/gsea/index.jsp) and GO gene sets for mouse were generated with GO annotation obtained from Ensembl Biomart (http://useast.ensembl.org/biomart/martview/5bdb637f2ad9ce44c330fbda04c14ca3). The RNA-seq data generated in this study have been deposited in the Sequences Read Archive at the NCBI under accession number SRP154924.
Adenoviral construction and cell infection
Adenovirus encoding TEAD1, Myocd, or YAP1 was generated as described previously [5, 20, 29, 40]. To generate mouse Pitx2c adenovirus, the SuperScript III One-Step RT-PCR system (Invitrogen) was used to clone mouse Pitx2c cDNA using total RNA extracted from E12.5 moue embryonic cardiac and aortic tissues. Primers specific for mouse Pitx2c gene were used to synthesize and amplify the Pitx2c cDNA (primer sequences harboring KpnI and HindIII restriction enzyme sites were listed in the Online Table I). Subsequently, the Pitx2c cDNA was sub-cloned into an AdTrack shuttle vector and verified by Sanger sequencing (Genewiz). Transferring the Pitx2c expression cassettes from AdTrack vector into AdEasy viral backbone, viral packaging and cell infection was performed as we previously reported [19, 40].
Quantitative real time reverse transcription-PCR analysis
Total RNA from E12.5 or E13.5 embryonic aortic and heart tissue and VSMCs was isolated with TRIzol reagent (Invitrogen) as previously described [19, 20]. 0.5-1μg of RNA was utilized as a template for RT with random hexamer primers using the High Capacity RNA-to-cDNA Kit (Invitrogen). Real time polymerase chain reaction (RT-PCR) was performed with respective gene-specific primers as we previously reported or were otherwise listed in Online Table I [5, 19, 40]. All samples were amplified in duplicate and all experiments were repeated at least three independent times. Relative gene expression was converted using the 2−ΔΔCT method against the internal control house-keeping gene hypoxanthine phosphorribosyltransferase 1 (Hprt) where ΔΔCT = (CTexperimental gene − CTexperimental Hprt) − (CTcontrol gene − CTcontrol Hprt). The relative gene expression in control group was set to 1.
Co-immunoprecipitation assay
To determine if MYOCD, SRF and PITX2c can form a complex in VSMCs, mouse embryonic VSMCs were co-transduced with adenovirus encoding Myocd and Pitx2c for 72 h. Subsequently, nuclear protein was harvested for co-immunoprecipitation (Co-IP) assays using the Nuclear Complex Co-IP Kit (Active Motif) with anti-SRF antibody (Santa cruz, G-20, sc-335x, rabbit), anti-PITX2 antibody (Capra Science, PA-1020, rabbit), and anti-MYOCD antibody (Santa Cruz, H-300, sc-33766, rabbit) as described in our previous reports [41, 42]. Western blots were performed as described below.
GST pull-down assays
Primers harboring BamHI and NotI restriction enzyme sites specific for mouse Pitx2c cDNA (primer sequences were listed in the Online Table I) were used to amplify Pitx2c cDNA using AdTrack-Pitx2c plasmid as the template. Subsequently, the PCR product was subcloned into the pET28a vector (Novagen) that can express T7-tagged fusion protein. pET28 vector containing full-length mouse Myocd cDNA or pGEX4T vectors (express glutathione S-transferase (GST) fusion protein, Sigma-Aldrich) containing truncated Myocd cDNA were generated as we previously reported [43]. pGEX-4T vector encoding full-length SRF cDNA is a kind gift of Dr. B. Paul Herring [44]. Protein expression in bacteria and GST pull-down assays were performed as described in our previous reports [29, 41,42,43]. Briefly, bacterial expression vectors were transformed into Escherichia coli BL21-star (Stratagene) cells. After 1 h of induction with 0.4 mm isopropyl β-d-thiogalactoside, cultured bacteria were harvested by centrifugation, resuspended in PBS with protease inhibitors and then lysed by sonication. GST fusion protein lysates were clarified by centrifugation and incubated for 1 h with a 50% suspension of glutathione-agarose beads (Amersham Biosciences) in PBS. Beads bound with GST fusion proteins were resuspended and incubated for 2 h with 500 μl of T7-tagged proteins expressed by pET28 vector in a total volume of 1ml of binding buffer (PBS containing 1% Triton X-100, 1 mg/ml BSA + protease inhibitors). Beads were then washed 3 times in 1 ml of washing buffer (PBS containing 1% Triton X-100). Then bound proteins were eluted by heating at 95 °C for 5 min in sodium dodecyl sulfate (SDS) sample buffer. The eluted proteins were resolved on SDS-polyacrylamide gel electrophoresis (PAGE) and detected by Western blotting as described below.
Protein extraction and Western blotting
Protein lysate was extracted from embryonic tissues or cultured cells by radioimmunoprecipitation buffer (Fisher) as previously described [19, 21]. Following sonication and centrifugation, protein lysate was quantified by bicinchoninic acid assay and then loaded in a 6–12% SDS-PAGE gel at 3–10 μg per lane. Images were captured by ImageQuant LAS 4000 Imaging Station (GE) and band densities were quantified using the ImageQuant TL software (GE). Antibodies used in this study are: TUBA (α-tubulin, Cell Signaling, #2144, rabbit, 1:5000), ACTB (β-actin, Sigma, A5316, mouse, 1:5000), CNN1 (calponin, Abcam, ab46794, rabbit, 1:2500), CCNB1 (cyclin B1, Cell Signaling, #4138, rabbit,1:1000), CCND1 (cyclin D1, Cell Signaling, #2978, rabbit, 1:1000), CCND2 (cyclin D2, Cell Signaling, #3741, rabbit, 1:1000), CCNE1 (cyclin E1, Cell Signaling, #4129, mouse, 1:1000), Cre (Novagen, 69050-3, rabbit, 1:10000), GST (GE Healthcare, goat, 1:10000), (TNNI3 (cardiac troponin I type 3, Developmental Studies Hybridoma Bank, TI-1, 1:1000), TNNT2 (cardiac troponin T type 2, Developmental Studies Hybridoma Bank, RV-C2, mouse, 1:500), TGFB1I1 (Hic-5, BD, 611164, mouse 1:5000), HSP90 (Cell Signaling, #4874, rabbit, 1:1000), LMOD1 (leiomodin 1, Abcam, ab104858, rabbit, 1:2000), MYH1E (MF-20, Developmental Studies Hybridoma Bank, MF 20, mouse, 1:1000), MYLK (MLCK, Abcam, ab76092, rabbit, 1:2000), MYH7 (Santa Cruz, sc-53089, mouse, 1:1000), MYOCD (myocardin, Santa Cruz, H-300, sc-33766, rabbit,1:500), PCNA (Santa Cruz, sc-56, mouse,1:500), phospho-H3F3A, (H3 histone family member 3A, Ser10, Millipore, 06-570, rabbit, 1:1000), PITX2 (Capra Science, PA-1020, rabbit, 1:1000), TAGLN (SM22α, Abcam, ab10135, goat, 1:2000), ACTA2 (SM α-actin, Sigma, A2547, mouse,1:5000), MYH11 (SM MHC, Biomedical Technologies Inc, BT-562, rabbit, 1:1000), SRF (Santa Cruz, G-20, sc-335x, rabbit,1:1000), TEAD1 (Abcam, ab133533, rabbit, 1:1000), VCL (vinculin, Sigma, V4505, mouse,1:5000), and YAP1 (Sigma, WH0010413M1, mouse, 1:1000).
Luciferase reporter assays
Cloning of mouse Pitx2c gene promoter luciferase reporter and dual luciferase assay were performed as described in our recent reports [5, 20]. A 2541-bp fragment spanning mouse Pitx2c proximal promoter region (chr3:129,211,616–129,214,157) was amplified by PCR with primers harboring KpnI and XhoI restriction enzyme sites by using mouse genomic DNA as template (Promega) (primer sequences were listed in the Online Table I). The PCR product was first cloned into pSC-B blunt vector (Stratagene) then subcloned into pGL2B (Promega) luciferase reporter vector. A conserved MCAT element within Pitx2c gene promoter was identified by a sequence alignment among multiple species (http://genome.ucsc.edu). Mutation of the MCAT element in the Pitx2c promoter was carried out with QuikChange II XL Site-Directed Mutagenesis Kit (Stratagene). All plasmids were sequenced to verify the integrity of the insert. Mouse luciferase reporter plasmids for Lmod1 and mutant Lmod1 were generously provided by Dr. Joseph Miano at the University of Rochester [6]. Transfection was carried out with X-tremeGENE 9 transfection reagent (Roche) essentially following manufacturer’s protocol. The promoter activity was evaluated by measurement of the firefly luciferase activity relative to the internal control minimal TK promoter driven-renilla luciferase activity using the Dual Luciferase Assay System as described by the manufacturer (Promega). A minimum of four independent transfections was performed and all assays were duplicated.
Quantitative chromatin immunoprecipitation (qChIP) assays
Heart and aortic tissues were harvested from E13.5 embryos (total 80 embryos) and pooled together for fixing with formaldehyde. Embryonic VSMCs were isolated as described as above. A chromatin immunoprecipitation (ChIP) assay was performed by using anti-TEAD1 antibody (Abcam, ab133533) or control rabbit IgG antibody as described by the manufacturer (Active Motif) and in our previous reports [5, 41]. ChIP assays were repeated three times and each IP was duplicated. Equal amount of chromatin was used for IP with anti-TEAD1 antibody or IgG. The genomic DNA purified from the precipitated genomic DNA or from input were amplified by qPCR. Primers for quantitative evaluation of enrichment of the smooth muscle-specific gene promoter CArG region were listed in Online Table I. Data were expressed as relative binding by using the 2−ΔΔCT method against the IgG control samples (set to 1) where ΔΔCT = (CTIP TEAD1 − CTinput TEAD1) − (CTIP control IgG − CTinput control IgG).
Statistical analysis
All data are expressed as means ± SE of at least three independent experiments. Statistical analysis of the data was performed using one-way analysis of variance (ANOVA) followed by Bonferroni t test for data involving more than two groups, or unpaired two tailed t-test for data involving two groups only (GraphPad). Values of p ≤ 0.05 were considered statistically significant.
Results
Conditional deletion of Tead1 in CMCs/VSMCs in mice results in embryonic lethality at E14.5
To determine the specific role of Tead1 in CMC/VSMC development, Tead1 floxed allele mice (Tead1F/F) were crossed with transgenic mice expressing Cre recombinase under the control of the promoter of smooth muscle-specific gene Tagln (SM22α) [31]. First we assessed the specificity of Cre activity directed by Tagln gene promoter during mouse embryonic development, by intercross with the dual fluorescence reporter mouse, mTmG [32]. Because mGFP can only be switched on while mTomato is simultaneously switched off upon Cre-mediated excision, the mGFP signal can be utilized to permanently label Cre-expressing cells (Online Fig. I A). Directly visualizing the GFP signal and IF staining of GFP demonstrated that Tagln promoter used in the transgenic Cre mouse specifically directs recombinase activity only in CMCs and VSMCs at E13.5 mouse embryos (Online Fig. I B–D). Therefore, the Tagln-Cre mediated deletion of Tead1 occurred specifically in CMCs and medial layer VSMCs as anticipated (Online Fig. II).
Genotyping results from timed pregnant embryos demonstrated that offspring of male Tagln-Cre+/Tead1F/W mice crossed with female Tead1F/F/mTmG+/+ mice exhibited the expected 25% Mendelian ratio from E10.5 to E13.5. At E14.5, the numbers of viable mutant embryos were significantly reduced to ~ 5% and no viable KO mice detected at E15.5 to postnatal day 14 (P14) (Online Table III). These data suggest that specific deletion of Tead1 in CMCs/VSMCs leads to embryonic lethality in mice at E14.5.
Specific ablation of Tead1 in CMCs results in hypoplasia of the myocardial wall due to the impaired CMC proliferation
To determine the underlying cause of embryonic lethality resulting from Tead1 deficiency in CMCs/VSMCs, E12.5, E13.5, and E14.5 embryos were harvested to analyze the phenotype. Starting from E12.5 to E14.5, all of Tead1 mutant embryos (E12.5, n = 24/24; E13.5, n = 60/60; E14.5, n = 4/4, respectively) exhibited pericardial effusion or/and peripheral edema, suggesting a cardiovascular defect (Fig. 1a, Online Fig. III A and B). No gross abnormalities were found in Tead1 heterozygous embryos. Furthermore, compared to Tagln-Cre+/Tead1F/W/mTmG+/− heterozygous control embryos, Tagln-Cre+/Tead1F/F/mTmG+/− KO mutant hearts exhibited much weaker GFP fluorescence that marks the Cre expressing CMCs (Fig. 1b). Visualization of the GFP-tagged CMCs or HE staining of the sections demonstrated that Tead1 mutants displayed a marked thinning of the right and left ventricular wall with a reduced number of trabeculae at both E12.5 and E13.5 (Fig. 1c–f). Moreover, Tead1 mutants at E14.5 not only exhibited a hypoplastic myocardium wall with less trabeculae, but also displayed membranous ventricular septal defect and a double outlet right ventricle phenotype (Online Fig. III C). The thinning myocardial wall in the Tead1 deficient heart is most likely due to impaired cardiomyocyte proliferation as shown by markedly decreased expression of proliferative markers MKI67, phospho-H3F3A or EdU positive CMCs in both E12.5 and E13.5 mutant hearts (Online Figs. IV and V).
Conditional Tead1 KO mice display a severe hypoplastic cardiac phenotype. a Gross images of Tead1 selective CMC/VSMC KO embryos at E12.5 show pericardial effusion (red arrows) or/and peripheral edema (green arrows, far right panel). b The hearts of Tagln-Cre+/Tead1F/F/mTmG+/− KO mutants at E13.5 exhibit a wrinkled surface with blood retention in both ventricles (upper panel, arrows) and much weaker GFP expression which marks Cre-expressing CMCs (bottom panel, arrows), compared to Tagln-Cre+/Tead1F/W/mTmG+/− heterozygous control embryos. mTmG membrane targeted RFP tomato and membrane targeted EGFP. c Tagln-Cre+/Tead1F/W/mTmG+/− heterozygous control and Tagln-Cre+/Tead1F/F/mTmG+/− KO embryos at E12.5 were sectioned and directly visualized under a fluorescent microscope to assess GFP-tagged (Cre-positive) CMCs. RFP (tomato) marks Cre-negative cells. The boxed area of the right ventricle (RV) or left ventricle (LV, upper panel) is magnified and shown below the whole heart sections as indicated. Deletion of Tead1 in CMCs decreases the numbers of GFP positive CMCs, resulting in a thinning, compact myocardium (arrows) with less trabeculae in both cardiac ventricles. d, e HE staining of heart sections from control or Tead1 mutants at E12.5 (d) or E13.5 (e). The boxed area of the RV or LV within the whole-heart sections is magnified and shown below. Deletion of TEAD1 in CMCs led to severe hypoplasia of the myocardial wall (green arrows) with reduced trabeculae (back arrows) in both cardiac ventricles. f The thickness of the RV and LV myocardium wall at E12.5 (N = 4) and E13.5 (N = 9) as shown in “D–E” was quantified and presented relative to control (set to 1). *p < 0.05
Specific deletion of Tead1 in VSMCs leads to a dilated arterial lumen and thinning vascular wall due to compromised VSMC proliferation
In addition to the hypoplastic cardiac phenotype, loss of Tead1 in VSMCs resulted in significant thinning of vascular wall and enlargement of the vessel lumen area in both carotid artery and dorsal aorta (Fig. 2a–f) at E13.5. The hypoplastic vascular wall phenotype observed in Tead1 KO mutants resulted from the attenuated VSMC proliferation, as indicated by markedly decreased numbers of EdU and MKI67 positive VSMCs (Fig. 3a–f). Taken together, these data suggest that specific ablation of Tead1 in CMC/VSMCs leads to a severe hypoplasia cardiac and vascular wall phenotype secondary to impaired cellular proliferation. Therefore, the cardiovascular defects in mutant embryos are the primary cause of embryonic lethality.
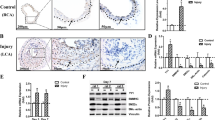
Conditional deletion of Tead1 in VSMCs leads to an atrophic arterial wall and dilated arterial lumen. At E13.5, Tagln-Cre+/Tead1F/W/mTmG+/− heterozygous control and Tagln-Cre+/TEAD1F/F/mTmG+/− mutant embryos were harvested and sectioned for the direct visualization of GFP-marked VSMCs (a, c) or for HE staining (b, d) of the left common carotid artery (LCA, a, b) and dorsal aorta (DA, c, d). Thickness of the media wall (e) and total lumen area (f) were quantified from the cross sections shown in panels (a) and (c). *p < 0.05
Tead1 KO embryos display proliferative defects in VSMCs. Pregnant dams carrying E13.5 control or Tead1 KO embryos were peritoneally injected with EdU to mark proliferating cells for 2 h. Embryos were then isolated, embedded and sectioned for immunostaining with antibodies for the smooth muscle differentiation marker MYH11 (green) together staining with the incorporated EdU (red) in LCA (a) or DA (b). Samples were counter-stained with DAPI (blue) to visualize nuclei. The percentage of EdU-positive VSMCs in the arterial wall of LCA or DA were measured and graphed as shown in panel (c). Similarly, sections were stained with antibodies for the smooth muscle differentiation marker ACTA2 (green) and the marker of cell proliferation, MKI67 (red) in LCA (d) or DA (e), respectively. The relative number of MKI67-positive VSMCs was measured and is shown in (f). N = 4 embryos per genotype. *p < 0.05
CMCs/VSMCs-specific deletion of Tead1 attenuates the expression of differentiation genes
To begin to define the molecular mechanism by which TEAD1 regulates cardiovascular development in mouse, we performed whole transcriptome screening by RNA-seq to determine the differentially expressed genes in CMCs/VSMCs between control and Tead1 KO embryos. We selectively isolated GFP positive hearts and aortic tissues of E13.5 Tagln-Cre+/Tead1F/W/mTmG+/− control or Tagln-Cre+/Tead1F/F/mTmG+/− KO embryos under a fluorescence stereoscope. After cell dispersal using enzymatic digestion, the live GFP positive cells were sorted using FACS and then total RNA was extracted for next-generation sequencing (Online Fig. VI A). Consistent with the proliferation defect, FACS revealed that the numbers of GFP positive cells in KO embryos were significantly less compared to control (Online Fig. VI A). Furthermore, RNA-seq data revealed that ablation of Tead1 in CMCs/VSMCs significantly downregulated 580 genes while upregulating 510 genes, respectively (Fig. 4a and Online Table II). Both GO enrichment analysis (Online Table IV and Online Fig. VI B) and KEGG pathway analysis (Online Table V and Online Fig. VI C) revealed the most significant pathway for downregulated genes is “muscle contraction” related to cardiomyocytes. GSEA revealed that, consistent with GO and KEGG pathway analysis, the gene set of “muscle contraction” is significantly downregulated by Tead1 deletion (Online Fig. VI D). Among these significantly downregulated genes, most of them are muscle structural proteins and key regulatory transcription factors including Myocd and Pitx2 that govern both VSMC and CMC differentiation gene programs [13, 15, 45, 46] (Fig. 4b). A careful examination of exon coverage in the mouse Pitx2 gene locus further revealed that the Pitx2 family gene Pitx2c is specifically expressed in E13.5 CMCs/VSMCs, but also downregulated in Tead1-deficient CMCs/VSMCs (Online Fig. VI E). These bioinformatics analyses collectively reveal a potential unrecognized role of TEAD1 in regulating muscle cell differentiation in vivo. qRT-PCR and Western blotting validated the RNA-seq data and revealed that both VSMC and CMC differentiation defects as indicated by the decreased expression of CMC and VSMC contractile genes, and that proliferation defects as shown by downregulation of cyclin family members in Tead1 mutants, respectively (Fig. 4c-e; Online Fig. VII A, B). Taken together, these data suggest that Tead1 is critical for the expression of genes controlling differentiation during VSMC and CMC development.
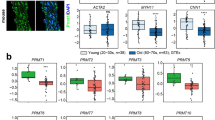
Deletion of Tead1 in embryonic VSMCs abolishes VSMC differentiation. a Volcano plot is to show differentially expressed genes between control and Tead1 KO CMCs/VSMCs (green: downregulated; red: upregulated) as assessed by RNA-seq. b Genes involved in CMC/VSMC differentiation that are downregulated in Tead1 KO cells are shown using a chord plot. MYOCD and PITX2, which govern both the VSMC and CMC contractile phenotype, are labeled in red. c At E13.5, hearts and aortae from control and Tead1 KOs were harvested for qRT-PCR analysis to validate changes in gene expression as revealed by RNA-seq analysis. N = 4–6 embryos per genotype. *p < 0.05. d Representative Western blot using protein lysates from the dorsal aorta of E13.5 control or TEAD1 KO embryos. e Densitometric analysis of protein expression in “D” normalized to the loading control HSP90 and relative to signals from control aortae (set to 1, red line). N = 4 embryos per genotype. *p < 0.05
Tead1 is critical for VSMC differentiation
Given that the role of the TEAD family and their co-factor YAP1 in regulating cell proliferation has been well-documented [19, 20, 24, 47], we next sought to establish a novel role of TEAD1 in VSMC differentiation. We first tested whether the VSMC differentiation defects observed in Tead1 mutant embryos are the primary consequence of the loss of TEAD1 in VSMCs. We isolated the GFP-tagged VSMCs from control and KO dorsal aorta by enzymatic digestion followed by FACS (Fig. 5a). Consistent with the findings in vivo (Fig. 4c-e), cultured embryonic VSMCs from Tead1 KO embryos displayed significantly lower expression levels of smooth muscle-contractile genes and the transcription factors, Pitx2c and Myocd without impacting its cofactor Yap1 expression (Fig. 5b). Restoration of TEAD1 expression through adenoviral-mediated gene transfer significantly rescued the VSMC differentiation defects, as shown by the elevated expression levels of PITX2c as well as smooth muscle-contractile genes (Fig. 5c, d). In contrast, over-expression of YAP1 exacerbated VSMC differentiation defects in Tead1 deficient VSMCs (Online Fig. VIII A, B). To further examine the obligatory role of TEAD1 in regulating VSMC differentiation, mouse embryonic VSMCs were prepared from E13.5 Tead1F/F embryonic aortae and transduced with control GFP or Cre recombinase adenovirus followed by treatment with TGF-β, a potent growth factor that promotes VSMC differentiation in vitro [48]. We found that deletion of Tead1, in the Tead1 floxed allele cells transduced with the Cre adenovirus, not only attenuated the expression of endogenous smooth muscle-specific genes, but also impaired TGF-β-induced VSMC differentiation (Fig. 5e, f). Luciferase reporter assays further demonstrated activity of reporter genes, driven by constructs encompassing the promoter regions with CArG boxes of Tgfb1i1 and Lmod1 genes was reduced in Tead1 deficient cells (Fig. 5g). Consistent with this finding, over-expression of Tead1 activated Tgfb1i1 and Lmod1 promoters (Fig. 5h), suggesting that the effects of TEAD1 on smooth muscle-specific gene expression occurs at the transcriptional level. In contrast, over-expression of the TEAD1 cofactor, YAP1 attenuated promoter activity of Tgfb1i1 and Lmod1 in embryonic VSMCs (Online Fig. VIII C).
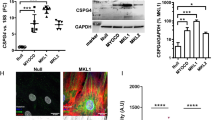
TEAD1 promotes embryonic VSMC differentiation. a. Using a flow cytometry, VSMCs were isolated from the dorsal aortae of E13.5 Tagln-Cre−/Tead1F/F/mTmG+/−, Tagln-Cre+/Tead1F/W/mTmG+/−, and Tagln-Cre+/Tead1F/F/mTmG+/− embryos and cultured to directly visualize GFP positive VSMCs. b Isolated VSMCs from the dorsal aorta of E13.5 control or Tead1 KO embryos were cultured and total RNA was extracted. qRT-PCR was performed to assess relative changes in gene expression as indicated. The level of gene expression in control VSMCs was set to 1 (red line). Data were obtained from cells cultured from 6 embryos per genotype. *p < 0.05. c Primary aortic VSMCs isolated from control or Tead1 mutant embryos were transduced with adenovirus encoding control GFP or TEAD1 and the expression of smooth muscle-specific genes was analyzed by Western blot. Restoration of TEAD1 expression in Tead1-null VSMCs rescues the differentiation defects seen in Tead1-deficient cells. d Densitometric analysis of protein expression in “C” normalized to the loading control VCL and relative to signals from control cells infected with GFP (set to 1, red line). N = 4. *p < 0.05, vs. Control + Ad-GFP group; #p < 0.05, vs. KO + Ad-GFP group. e Cultured primary VSMCs from E13.5 TEAD1F/F embryos were transduced with control GFP or Cre adenovirus and subsequently cells were treated with or without TGF-β, an established driver for VSMC differentiation. 48 h posttransduction, cells were harvested for analysis of smooth muscle-specific gene expression by Western blot. f Densitometric analysis of protein expression in “E” normalized to the loading control TUBA. *p < 0.05 and &p < 0.05, vs. cells infected with GFP and no TGF-β treatment (set to 1, red line); #p < 0.05, vs. cells infected with GFP and with TGF-β treatment. KO of endogenous TEAD1 expression impairs TGF-β induced smooth muscle-specific gene expression. g Luciferase reporters driven by the promoter regions of smooth muscle-specific genes, Tgfb1i1 or Lmod1 were transfected into control or Tead1 KO primary embryonic VSMCs and dual luciferase assays were carried out to determine the degree of promoter activity in transfected cells. Promoter activity in control cells was set to 1 (red line). N = 6. *P < 0.05. h Smooth muscle-specific luciferase reporters (Tgfb1i1 or Lmod1) were transfected into control embryonic VSMCs with or without co-transfection of a TEAD1 expression plasmid. At 36 h posttransfection, the effects of TEAD1 on promoter activity were determined using dual luciferase assays. Baseline promoter activity without TEAD1 co-transfection was set to 1 (red line). N = 3–6. *P < 0.05
In our previous study, we had shown that TEAD1 suppresses smooth muscle-specific gene expression in cultured adult VSMCs [29]. In contrast, data from embryonic VSMCs described above show that TEAD1 promotes smooth muscle-specific gene expression. To resolve this discrepancy, we isolated mature VSMCs from the dorsal aorta of adult Tead1F/F mice and transduced them with the Cre recombinase adenovirus to ablate Tead1. Data from these experiments showed that, in contrast to embryonic VSMCs, deletion of Tead1 in adult VSMCs significantly induced smooth muscle-contractile gene expression, suggesting an important temporal role of TEAD1 in regulating VSMC differentiation (Online Fig. IX A-E). Taken together, these data suggest that TEAD1 plays a distinct role from its cofactor YAP1 to promote VSMC differentiation in a time and cell-autonomous manner during vascular development.
Pitx2c is a direct transcriptional target of TEAD1 and promotes VSMC differentiation
Our data above demonstrated that deletion of Tead1 in embryonic VSMCs attenuates the expression of a large set of smooth muscle-contractile genes that do not contain obvious MCAT elements including Tgfb1i1 and Lmod1. Our RNA-seq data further revealed that ablation of Tead1 in embryonic VSMCs downregulated two crucial transcription factors, Myocd and Pitx2c that are critical for VSMC differentiation [14, 15]. Previous studies have shown that a conserved MCAT element within the Myocd gene enhancer governs MYOCD expression specifically in VSMCs [49]. Data from quantitative ChIP (qChIP) assays unveiled an approximate nine to tenfold enrichment of TEAD1 over the MCAT region of Myocd gene enhancer in E13.5 cardiovascular tissues (Online Fig. XA). Rescue experiments demonstrated that restoration of MYOCD expression in Tead1-defficient VSMCs induces expression of smooth muscle-contractile genes (Online Fig. XB, C).
Since the underlying mechanism by which TEAD1 regulates Myocd expression and the role of MYOCD in VSMC development is well studied [14, 49], we next sought to investigate the mechanism by which TEAD1 regulates the expression of the novel target Pitx2c. Analysis of the mouse Pitx2c gene revealed an evolutionally conserved TEAD binding MCAT element 5′-AGAATGTA-3′ in the proximal promoter region of the Pitx2c gene (Fig. 6a). qChIP assays experimentally validated that TEAD1 specifically binds to the putative MCAT element in vitro in embryonic VSMCs and in vivo in E13.5 heart and aortic tissues (Fig. 6b). To further test whether TEAD1 can activate the Pitx2c gene promoter activity through the MCAT element, we generated luciferase reporters that harbor the Pitx2c proximal gene promoter containing wild type (WT) or mutated MCAT elements and dual luciferase assays were performed. Data from the reporter assays showed that TEAD1 significantly trans-activates the WT promoter luciferase activity fivefold but not the MCAT mutant (Fig. 6c). Together, these data suggest that Pitx2c is a novel transcriptional target of TEAD1 which binds to the conserved MCAT element within the proximal promoter of Pitx2c gene. To establish a functional role of PITX2c in VSMC differentiation, rescue experiments revealed that ectopic expression of Pitx2c gene in Tead1-null embryonic VSMCs restores smooth muscle-specific gene expression level, suggesting that Pitx2c, at least in part, is an important downstream regulator of TEAD1 that promotes VSMC differentiation during vascular development (Fig. 6d, e). This concept is further supported by reporter assays which revealed that PITX2c trans-activates WT smooth muscle-specific gene reporters (Tgfb1i1 and Lmod1) but not CArG mutants (Fig. 6f, g), suggesting that the effects of PITX2c on expression of smooth muscle-specific genes occurs at the transcriptional level and is SRF/CArG-box dependent. Taken together, these data demonstrate that Pitx2c is a novel transcriptional target of TEAD1 and orchestrates the function of TEAD1 to induce VSMC differentiation.
Pitx2c is a direct transcriptional target of TEAD1 and promotes smooth muscle-specific gene activation in a CArG-dependent manner. a Schematic diagram of the mouse Pitx2c gene locus showing a conserved MCAT element within the proximal promoter region. A 2.5 kbp DNA fragment of the Pitx2c gene promoter containing a WT or mutated MCAT element (underscored) was cloned into the pGL2b luciferase vector. A set of primers spanning the MCAT element was used for qChIP assays in (b) as shown by arrows. b Chromatin was harvested from control embryonic VSMCs or heart and aortic tissues from E13.5 embryos and the chromatin was subjected to IP using either IgG or anti-TEAD1 antibodies. Immunoprecipitated DNA complexes were then analyzed by qPCR using the primer sets indicated in “A”. The degree of TEAD1 binding is expressed as relative to IgG control (set to 1). N = 3–4. *p < 0.05. c Pitx2c promoter-luciferase reporters (WT or MCAT mutant) were transfected into primary embryonic VSMCs in the absence or presence of TEAD1 expression plasmid and promoter activity was evaluated by a dual luciferase assay. Pitx2c promoter activity was normalized using the control vector (set to 1). N = 8. *p < 0.05. d Control or Tead1-null VSMCs were transduced with GFP or Pitx2c adenovirus and the expression of VSMC-specific genes analyzed by Western blot. e Densitometric analysis of protein expression in (d) normalized to the loading control VCL. *p < 0.05, relative to control cells infected with GFP (set to 1, red line); #p < 0.05, vs. KO cells infected with GFP. Over-expression of Pitx2c in Tead1-null VSMCs restores expression of VSMC-specific contractile proteins. f Tgfb1i1 or Lmod1 g gene promoter-luciferase reporters (WT or mutated CArG) were transfected into MEFs in the absence or presence of plasmids expressing Pitx2c and dual luciferase assays were performed. Baseline activity of the WT promoter in the absence of Pitx2c co-transfection was set to 1. N = 6. *P < 0.05
PITX2c synergistically promotes VSMC differentiation with MYOCD
Data described above suggest that TEAD1 directly targets two critical transcription factors, Pitx2c and Myocd to promote VSMC-specific gene expression. To further understand the functional relationship between PITX2c and MYOCD in VSMC differentiation, we sought to determine whether PITX2c and MYOCD can physically interact with each other in embryonic VSMCs. Due to lack of a validated antibody able to detect endogenous MYOCD in VSMCs [50], adenovirus expressing Myocd and Pitx2c were co-transduced into embryonic VSMCs and IP assays were performed. Western blotting of immune-precipitates revealed that PITX2c and MYOCD can form a ternary complex with SRF (Fig. 7a). Furthermore, GST pull-down assays confirmed the protein–protein interaction between SRF and PITX2c, SRF and MYOCD, respectively, but also the ternary complex formed by SRF, PITX2c, and MYOCD in vitro using bacterial expressed proteins (Fig. 7b, c). GST pull-down assays further defined the N-terminal MYOCD binds to PITX2c (Fig. 7d, e). More importantly, co-transduction of Pitx2c and Myocd into MEFs synergistically induced the expression of endogenous smooth muscle-genes to a greater extent than single transduction of Pitx2c or Myocd alone (Fig. 7f, g). Similarly, reporter assays further revealed that co-transfection of Pitx2c and Myocd synergistically trans-activates the Lmod1 gene promoter in a CArG-dependent manner (Fig. 7h). Collectively, these data suggest that both Pitx2c and Myocd are direct targets of TEAD1 and synergistically promote VSMC differentiation during vascular development through a physical and functional interaction (Fig. 7i).
PITX2c and MYOCD act synergistically to promote VSMC differentiation. a Adenovirus encoding Pitx2c and Myocd were co-transduced into embryonic VSMCs. PITX2c, MYOCD and endogenously expressed SRF were then immunoprecipitated using anti-PITX2, anti-MYOCD and anti-SRF antibodies, respectively. Proteins within immuno-complexes were determined by Western blot using the indicated antibodies. A species-matched IgG served as control. b Schematic diagram of mouse MYOCD domains demonstrating the MYOCD GST fusion proteins analyzed in GST pull-down assays and a summary of MYOCD domains mapped to interact with PITX2c. NTD N-terminal domain, ++ basic domain, Q polyQ domain, SAP, SAF-A/B, Acinus, and PIAS domain, LZ leucine zipper domain, TAD transcriptional activation domain, NT N-terminus, CT C-terminus, N1-4 N-terminal mutants 1–4. c Bacterial expressed full-length MYOCD or/and PITX2c was incubated with SRF-GST fusion protein or with GST control protein. Western blotting was performed to detect the SRF-GST bound MYOCD or/and PITX2c (upper panel). The lower panel indicates the expression of the GST or SRF-GST fusion proteins (arrows). The middle membrane within the dashed lines was cropped for detecting PITX2c. SRF was found to bind to both MYOCD and PITX2c. d Bacterial expressed PITX2c was incubated with NT- or CT-MYOCD-GST fusion protein or control GST protein (arrows). Western blotting was performed to detect the MYOCD-GST bound PITX2c (upper panel) and revealed that PITX2C binds to NT-MYOCD. The middle membrane within the dashed lines was cropped for detecting PITX2c. e Using truncation mutants of NT-MYOCD (N1-4, arrows) and CT-MYOCD GST fusion proteins, GST pull-down assay further showed N1–3 domains of NT-MYOCD specifically binds to PITX2c. f Adenovirus encoding GFP, Pitx2c, or/and Myocd were transduced into embryonic VSMCs and expression of endogenous VSMC-specific gene was determined by Western blot. g The band intensity shown in (f) was measured, normalized to its loading control HSP90 and plotted. N = 4. *P < 0.05, vs. Ad-GFP control group (set to 1, red line); #P < 0.05, vs. Ad-Pitx2c or Ad-MYOCD group. Co-expression of MYOCD and Pitx2c in VSMCs synergistically induces the expression of contractile proteins. h WT or mutated CArG Lmod1 gene promoter-luciferase reporters were transfected into MEFs with or without co-expression of Pitx2c and MYOCD expression plasmid. Dual luciferase reporter assays were then performed to determine the combinatorial effects between Pitx2c and MYOCD on Lmod1 gene promoter activity. The promoter activity of the WT reporter at baseline without Pitx2c or MYOCD transfection was set to 1. N = 4–6. *P < 0.05. i Schematic diagram depicting the regulatory mechanisms by which TEAD1 mediates VSMC-specific gene expression during VSMC development. TEAD1 functions upstream of the genetic regulatory hierarchy by directly regulating the critical transcription factors, PITX2c and MYOCD, both of which act not only physically but also synergistically to promote the expression of VSMC-specific genes that are reflective of differentiated VSMCs
Discussion
Our current study provides the first evidence that TEAD1 plays a crucial role in both proliferation and differentiation of VSMCs during vascular development, and that these effects are likely mediated through a mechanism independent of its cofactor, YAP1.
YAP1, as a prime binding factor to TEADs, is critical for VSMC proliferation both in vitro and in vivo [19, 20]. CMCs/VSMC-specific deletion of Yap1 as directed by Tagln-Cre transgene resulted in perinatal lethality due to proliferation defects in CMCs/VSMCs [20]. However, Yap1 deletion in embryonic VSMCs had no effect on smooth muscle-specific contractile gene expression, but instead significantly attenuated VSMC proliferation through the upregulation of cell cycle arrest genes without affecting cyclin D1 expression [20, 21]. In the current study, CMCs/VSMCs-specific deletion of Tead1 mediated by the same Tagln-Cre transgene resulted in embryonic lethality at E14.5, due to profound defects of both proliferation and differentiation in CMCs/VSMCs. The proliferation defects in Tead1-deficient VSMCs, however, are most likely due to the downregulation of cell cycle genes including cyclin D1, D2, D3, and E1 (Fig. 4c–e). Our studies support the concept that, although the primary function of both YAP1 and TEAD1 is to regulate cell proliferation during embryonic development, the underlying mechanisms are different, via regulating distinct target genes involved in cell cycle control. The earlier embryonic lethality of cardiovascular deletion of Tead1 in mice suggests that TEAD1 plays an earlier and more critical role in cardiovascular development as compared to YAP1. Our study further suggests that TEAD1 has functions that are different from its cofactor YAP1 in the regulation of CMC/VSMC differentiation and utilizes different regulatory mechanism for promoting VSMC proliferation during cardiovascular development. Given the embryonic lethality of Tead1 CMC/VSMC-specific KO mice, an inducible KO model is required to study the role of Tead1 in adult VSMCs in vivo as we recently reported [51].
A surprising finding of our study was that TEAD1 appears to have different and opposing roles in the regulation of VSMC differentiation in embryonic vs. adult VSMCs. In embryonic VSMCs, we found that deletion of Tead1 leads to defects in VSMC differentiation due to downregulation of the key transcription factors, Myocd and Pitx2c (Figs. 4 and 5). In contrast, we previously demonstrated that TEAD1, in cultured adult VSMCs, is a potent suppressor of VSMC contractile gene expression by disrupting the transcription factor complex between MYOCD and SRF [29]. This finding was confirmed in the current study by ablating Tead1 in VSMCs isolated from adult Tead1F/F mice followed by Cre recombinase virus infection (Online Fig. IX). We speculate that the disparate functions of TEAD1 in the deferent stages of VSMC development are most likely due to different levels of TEAD1 expression and perhaps more likely, different TEAD1 binding partners.
Our study further suggests that TEAD1 acts upstream of the genetic regulatory hierarchy by directly regulating the expression of multiple transcription factors that are critical for smooth and cardiac muscle differentiation. In addition to Pitx2c and Myocd, which were the focus of this study, additional transcription factors that are important for the development of cardiac and vascular myocytes were also downregulated in TEAD1 KO cardiac and aortic tissues, including Nkx2.5, Tbx3, and Prox1 [52,53,54]. In support of this transcription factor hierarchy, TEAD1 has also been shown to function as a “master” regulator of subordinate transcription factors that regulate the expansion of pancreatic progenitors [25].
Shang et al. reported that the expression of SMC differentiation markers was nearly abolished in vessels of Pitx2 KO mouse embryos but the vessel wall thickness in Pitx2 KO mouse appeared normal. This study suggested Pitx2c is critical for SMC differentiation but is dispensable for VSMC proliferation during vascular development [15]. Our current study identified Pitx2c as a novel TEAD1 target gene which works cooperatively with MYOCD through a direct physical interaction to promote smooth muscle-specific gene expression (Figs. 6 and 7). Previous study has shown that PITX2 and SRF can synergistically induce the expression of VSMC differentiation genes through an epigenetic mechanism whereby PITX2 induces the acetylation of histone H4 at the promoter regions of genes involved in VSMC differentiation [15]. Together, these studies further revealed the added complexity of tissue-specific gene regulation in addition to transcriptional mechanisms.
In summary, the current study not only revealed a critical role of the Hippo signaling factor, TEAD1, in regulating cardiovascular development in mice, but also uncovered novel regulatory hierarchy where TEAD1 acts upstream of the factors responsible for establishing differentiated cardiac and smooth muscle phenotypes, by directly regulating the expression of the critical transcription factors, PITX2c and MYOCD.
References
Hungerford JE, Little CD. Developmental biology of the vascular smooth muscle cell: building a multilayered vessel wall. J Vasc Res. 1999;36:2–27.
Li L, Miano JM, Cserjesi P, Olson EN. SM22 alpha, a marker of adult smooth muscle, is expressed in multiple myogenic lineages during embryogenesis. Circ Res. 1996;78:188–95.
Samaha FF, Ip HS, Morrisey EE, Seltzer J, Tang Z, Solway J, et al. Developmental pattern of expression and genomic organization of the calponin-h1 gene. A contractile smooth muscle cell marker. J Biol Chem. 1996;271:395–403.
Gabbiani G, Schmid E, Winter S, Chaponnier C, de Ckhastonay C, Vandekerckhove J, et al. Vascular smooth muscle cells differ from other smooth muscle cells: predominance of vimentin filaments and a specific alpha-type actin. Proc Natl Acad Sci USA. 1981;78:298–302.
Wang X, Hu G, Betts C, Harmon EY, Keller RS, Van De Water L, et al. Transforming growth factor-beta1-induced transcript 1 protein, a novel marker for smooth muscle contractile phenotype, is regulated by serum response factor/myocardin protein. J Biol Chem. 2011;286:41589–99.
Nanda V, Miano JM. Leiomodin 1, a new serum response factor-dependent target gene expressed preferentially in differentiated smooth muscle cells. J Biol Chem. 2012;287:2459–67.
Yin F, Hoggatt AM, Zhou J, Herring BP. 130-kDa smooth muscle myosin light chain kinase is transcribed from a CArG-dependent, internal promoter within the mouse mylk gene. Am J Physiol Cell Physiol. 2006;290:C1599–1609.
Miano JM, Cserjesi P, Ligon KL, Periasamy M, Olson EN. Smooth muscle myosin heavy chain exclusively marks the smooth muscle lineage during mouse embryogenesis. Circ Res. 1994;75:803–12.
Majesky MW. Developmental basis of vascular smooth muscle diversity. Arterioscler Thromb Vasc Biol. 2007;27:1248–58.
Miano JM. Serum response factor: toggling between disparate programs of gene expression. J Mol Cell Cardiol. 2003;35:577–93.
Sun Q, Chen G, Streb JW, Long X, Yang Y, Stoeckert CJ, et al. Defining the mammalian CArGome. Genome Res. 2006;16:197–207.
Owens GK, Kumar MS, Wamhoff BR. Molecular regulation of vascular smooth muscle cell differentiation in development and disease. Physiol Rev. 2004;84:767–801.
Wang D, Chang PS, Wang Z, Sutherland L, Richardson JA, Small E, et al. Activation of cardiac gene expression by myocardin, a transcriptional cofactor for serum response factor. Cell. 2001;105:851–62.
Miano JM. Myocardin in biology and disease. J Biomed Res. 2015;29:3–19.
Shang Y, Yoshida T, Amendt BA, Martin JF, Owens GK. Pitx2 is functionally important in the early stages of vascular smooth muscle cell differentiation. J Cell Biol. 2008;181:461–73.
Zhao B, Li L, Lei Q, Guan KL. The Hippo-YAP pathway in organ size control and tumorigenesis: an updated version. Genes Dev. 2010;24:862–74.
Pan D. The hippo signaling pathway in development and cancer. Dev Cell. 2010;19:491–505.
Dong J, Feldmann G, Huang J, Wu S, Zhang N, Comerford SA, et al. Elucidation of a universal size-control mechanism in Drosophila and mammals. Cell. 2007;130:1120–33.
Wang X, Hu G, Gao X, Wang Y, Zhang W, Harmon EY, et al. The induction of yes-associated protein expression after arterial injury is crucial for smooth muscle phenotypic modulation and neointima formation. Arterioscler Thromb Vasc Biol. 2012;32:2662–9.
Wang Y, Hu G, Liu F, Wang X, Wu M, Schwarz JJ, et al. Deletion of yes-associated protein (YAP) specifically in cardiac and vascular smooth muscle cells reveals a crucial role for YAP in mouse cardiovascular development. Circ Res. 2014;114:957–65.
Xu F, Ahmed AS, Kang X, Hu G, Liu F, Zhang W, et al. MicroRNA-15b/16 attenuates vascular neointima formation by promoting the contractile phenotype of vascular smooth muscle through targeting YAP. Arterioscler Thromb Vasc Biol. 2015;35:2145–52.
Stein C, Bardet AF, Roma G, Bergling S, Clay I, Ruchti A, et al. YAP1 exerts its transcriptional control via TEAD-mediated activation of enhancers. PLoS Genet. 2015;11:e1005465.
Zanconato F, Forcato M, Battilana G, Azzolin L, Quaranta E, Bodega B, et al. Genome-wide association between YAP/TAZ/TEAD and AP-1 at enhancers drives oncogenic growth. Nat Cell Biol. 2015;17:1218–27.
Zhao B, Ye X, Yu J, Li L, Li W, Li S, et al. TEAD mediates YAP-dependent gene induction and growth control. Genes Dev. 2008;22:1962–71.
Cebola I, Rodriguez-Segui SA, Cho CH, Bessa J, Rovira M, Luengo M, et al. TEAD and YAP regulate the enhancer network of human embryonic pancreatic progenitors. Nat Cell Biol. 2015;17:615–26.
Galli GG, Carrara M, Yuan WC, Valdes-Quezada C, Gurung B, Pepe-Mooney B, et al. YAP drives growth by controlling transcriptional pause release from dynamic enhancers. Mol Cell. 2015;60:328–37.
Yoshida T. MCAT elements and the TEF-1 family of transcription factors in muscle development and disease. Arterioscler Thromb Vasc Biol. 2008;28:8–17.
Lin KC, Park HW, Guan KL. Regulation of the Hippo pathway transcription factor TEAD. Trends Biochem Sci. 2017;42:862–72.
Liu F, Wang X, Hu G, Wang Y, Zhou J. The transcription factor TEAD1 represses smooth muscle-specific gene expression by abolishing myocardin function. J Biol Chem. 2014;289:3308–16.
Wen T, Yin Q, Yu L, Hu G, Liu J, Zhang W, et al. Characterization of mice carrying a conditional TEAD1 allele. Genesis 2017;55:10.1002.
Holtwick R, Gotthardt M, Skryabin B, Steinmetz M, Potthast R, Zetsche B, et al. Smooth muscle-selective deletion of guanylyl cyclase-A prevents the acute but not chronic effects of ANP on blood pressure. Proc Natl Acad Sci USA. 2002;99:7142–7.
Muzumdar MD, Tasic B, Miyamichi K, Li L, Luo L. A global double-fluorescent Cre reporter mouse. Genesis. 2007;45:593–605.
Chen M, Herring BP. Regulation of microRNAs by Brahma-related gene 1 (Brg1) in smooth muscle cells. J Biol Chem. 2013;288:6397–408.
Bolger AM, Lohse M, Usadel B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics. 2014;30:2114–20.
Kim D, Pertea G, Trapnell C, Pimentel H, Kelley R, Salzberg SL. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 2013;14:R36.
Anders S, Pyl PT, Huber W. HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics. 2015;31:166–9.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26:139–40.
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods. 2008;5:621–8.
Walter W, Sanchez-Cabo F, Ricote M. GOplot: an R package for visually combining expression data with functional analysis. Bioinformatics. 2015;31:2912–4.
Wang X, Hu G, Zhou J. Repression of versican expression by microRNA-143. J Biol Chem. 2010;285:23241–50.
Zhou J, Hu G, Wang X. Repression of smooth muscle differentiation by a novel high mobility group box-containing protein, HMG2L1. J Biol Chem. 2010;285:23177–85.
Hu G, Wang X, Saunders DN, Henderson M, Russell AJ, Herring BP, et al. Modulation of myocardin function by the ubiquitin E3 ligase UBR5. J Biol Chem. 2010;285:11800–9.
Zhou J, Blue EK, Hu G, Herring BP. Thymine DNA glycosylase represses myocardin-induced smooth muscle cell differentiation by competing with serum response factor for myocardin binding. J Biol Chem. 2008;283:35383–92.
Herring BP, Kriegel AM, Hoggatt AM. Identification of Barx2b, a serum response factor-associated homeodomain protein. J Biol Chem. 2001;276:14482–9.
Li S, Wang DZ, Wang Z, Richardson JA, Olson EN. The serum response factor coactivator myocardin is required for vascular smooth muscle development. Proc Natl Acad Sci USA. 2003;100:9366–70.
Franco D, Sedmera D, Lozano-Velasco E. Multiple roles of Pitx2 in cardiac development and disease. J Cardiovasc Dev Dis. 2017; 4:16.
Ota M, Sasaki H. Mammalian Tead proteins regulate cell proliferation and contact inhibition as transcriptional mediators of Hippo signaling. Development. 2008;135:4059–69.
Hirschi KK, Rohovsky SA, D'Amore PAPDGF. TGF-beta, and heterotypic cell-cell interactions mediate endothelial cell-induced recruitment of 10T1/2 cells and their differentiation to a smooth muscle fate. J Cell Biol. 1998;141:805–14.
Creemers EE, Sutherland LB, McAnally J, Richardson JA, Olson EN. Myocardin is a direct transcriptional target of Mef2, Tead and Foxo proteins during cardiovascular development. Development. 2006;133:4245–56.
Lyu Q, Dhagia V, Han Y, Guo B, Wines-Samuelson ME, Christie CK, et al. CRISPR-Cas9-mediated epitope tagging provides accurate and versatile assessment of myocardin-brief report. Arterioscler Thromb Vasc Biol. 2018;38:2184–90.
Osman I, He X, Liu J, Dong K, Wen T, Zhang F, et al. TEA domain transcription factor 1 (TEAD1) promotes smooth muscle cell proliferation through up-regulating SLC1A5-mediated glutamine uptake. Circ Res. 2019 (In press).
Lyons I, Parsons LM, Hartley L, Li R, Andrews JE, Robb L, et al. Myogenic and morphogenetic defects in the heart tubes of murine embryos lacking the homeo box gene Nkx2-5. Genes &. Genes Dev. 1995;9:1654–66.
Papaioannou VE. The T-box gene family: emerging roles in development, stem cells and cancer. Development. 2014;141:3819–33.
Risebro CA, Searles RG, Melville AA, Ehler E, Jina N, Shah S, et al. Prox1 maintains muscle structure and growth in the developing heart. Development. 2009;136:495–505.
Acknowledgements
We thank Dr. Nicole Winston for a critical reading of the paper. We also thank Dr. Guangyu Wu’s laboratory for their help to generate Pitx2c-pET28 plasmid. The work at the J.Z. laboratory is supported by a grant from the National Heart, Lung, and Blood Institute, NIH (HL132164). J.Z. is a recipient of Established Investigator Award (17EIA33460468) from the American Heart Association. K.D. and I.O. are supported by postdoctoral fellowships (19POST34450071 and 18POST34030400, respectively) from the American Heart Association.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by S. Kumar
Rights and permissions
About this article
Cite this article
Wen, T., Liu, J., He, X. et al. Transcription factor TEAD1 is essential for vascular development by promoting vascular smooth muscle differentiation. Cell Death Differ 26, 2790–2806 (2019). https://doi.org/10.1038/s41418-019-0335-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-019-0335-4
This article is cited by
-
SMYD2 regulates vascular smooth muscle cell phenotypic switching and intimal hyperplasia via interaction with myocardin
Cellular and Molecular Life Sciences (2023)
-
SRF: a seriously responsible factor in cardiac development and disease
Journal of Biomedical Science (2022)
-
Six Decades of History of Hypertension Research at the University of Toledo: Highlighting Pioneering Contributions in Biochemistry, Genetics, and Host-Microbiota Interactions
Current Hypertension Reports (2022)
-
TEAD1 protects against necroptosis in postmitotic cardiomyocytes through regulation of nuclear DNA-encoded mitochondrial genes
Cell Death & Differentiation (2021)
-
A reference map of murine cardiac transcription factor chromatin occupancy identifies dynamic and conserved enhancers
Nature Communications (2019)