Abstract
Somatic oncogenic mutation of BRAF coupled with inactivation of PTEN constitute a frequent combination of genomic alterations driving the development of human melanoma. Mice genetically engineered to conditionally express oncogenic BrafV600E and inactivate Pten in melanocytes following tamoxifen treatment rapidly develop melanoma. While early-stage melanomas comprised melanin-pigmented Mitf and Dct-expressing cells, expression of these and other melanocyte identity genes was lost in later stage tumours that showed histological and molecular characteristics of de-differentiated neural crest type cells. Melanocyte identity genes displayed loss of active chromatin marks and RNA polymerase II and gain of heterochromatin marks, indicating epigenetic reprogramming during tumour progression. Nevertheless, late-stage tumour cells grown in culture re-expressed Mitf, and melanocyte markers and Mitf together with Sox10 coregulated a large number of genes essential for their growth. In this melanoma model, somatic inactivation that the catalytic Brg1 (Smarca4) subunit of the SWI/SNF complex and the scaffolding Bptf subunit of the NuRF complex delayed tumour formation and deregulated large and overlapping gene expression programs essential for normal tumour cell growth. Moreover, we show that Brg1 and Bptf coregulated many genes together with Mitf and Sox10. Together these transcription factors and chromatin remodelling complexes orchestrate essential gene expression programs in mouse melanoma cells.
Similar content being viewed by others
Introduction
Melanoma is a highly aggressive skin cancer resulting from oncogenic transformation of melanocytes. Microphthalmia-associated transcription factor (MITF) and SOX10 are two critical transcription factors regulating gene expression programs essential for melanocyte and melanoma cell proliferation. We showed that MITF interacts with the PBAF chromatin remodelling complex, a member of the SWI/SNF family [1] in human melanoma cells [2,3,4]. Its catalytic subunit BRG1 is essential for the proliferation of MITF-high cells and immortalised Hermes 3A melanocytes. Both MITF and SOX10 [5] actively recruit BRG1 to chromatin to establish the epigenetic landscape of proliferative melanoma cells [4, 6]. In agreement with its potentially important role, BRG1 expression is frequently upregulated in human melanomas [7]. In mouse, somatic Brg1 inactivation in the melanocyte lineage leads to loss of developing melanoblasts and the resulting animals lack pigmentation [4, 8].
MITF also interacts with the NuRF chromatin remodelling complex and together they coregulate genes involved in the proliferation of melanoma cells [9]. The gene encoding BPTF, the scaffolding subunit of NuRF, is amplified in around 5–7% of human melanomas [10]. BPTF expression can be upregulated during tumour progression and elevated BPTF expression is associated with poor prognosis and acquisition of resistance to BRAF inhibitors [11]. However, in contrast to Brg1, Bptf is not required for mouse melanoblast development, but is required for generation of melanocytes from the adult melanocyte stem cell population [9]. These two chromatin remodelling complexes therefore play distinct but complementary roles in the establishment and renewal of the melanocyte lineage and in human melanoma.
Given the essential but differing roles of BRG1/PBAF and BPTF/NuRF in human melanoma cells in vitro and in the normal physiology of mouse melanocytes in vivo, we addressed their implication in melanoma in vivo using a genetically modified mouse model.
Results
Initiation and evolution of Braf/Pten melanoma tumours
We established a previously described model [12,13,14,15] where treatment with 4-hydroxy-Tamoxifen (4-OHT) induces expression of oncogenic BrafV600E and inactivates Pten selectively in the melanocyte lineage (Tyr:Cre-ERT2::BrafLSL-V600E/+::Ptenlox/lox). Adult mice were depilated and treated with a 5 μL drop of 5 mM 4-OHT leading to rapid appearance of small pigmented naevi and around 40 days later of prominent melanoma lesions (Fig. 1a). Fifteen days after 4-OHT treatment, histological examination revealed small naevus-like lesions with proliferating pigmented epithelioid melanocytes in the dermis surrounding the hair follicles (Fig. 1b) expressing Sox10, Mitf and Dct (Fig. 1c). As lesions progressed deeper into the dermis, a transition from pigmented epithelioid cells to a nonpigmented ovoid or spindle-shaped morphology was observed with some pigmented S100A4 or Dct-stained cells remaining close to the epidermis and more rarely deeper in the tumours (Figs. 1d, S1a and S2a). Later stage tumours were composed of bland spindle cells disposed haphazardly in a loose myxoid stroma and numerous regions showed a focal whorling of tumour cells (Fig. S1b, c). The invasive nature of these melanomas was highlighted by the appearance of pigmented and/or Sox10-labelled cells in lymph nodes and partially pigmented lung metastases (Fig. S1d, e). Cells invading the underlying dermal muscle by 26 days expressed invasive markers Zeb1 and Zeb2 (Fig. S2a, b). By 36 days however, Zeb1 and Zeb2 staining was weaker and more irregular with not all cells labelled (Fig. S2c).
Development of oncogenic Braf-driven mouse melanoma. a Photographs of the 4-OHT-treated area on the backs of mice at the indicated days after treatment. b H&E-stained skin sections 15 days after 4-OHT treatment. HF hair follicle, PM pigmented melanocytes, E epidermis, D dermis, AD adipocytes. Scale bar 100 µm. c Immunofluorescence staining of sections from mouse skin 15 days after 4-OHT treatment with the indicated antibodies. DAPI-stained nuclei are also shown. Scale bar 100 µm. d H&E-stained sections from tumours 29 days (upper panel) and 36 days (lower panel) after 4-OHT treatment. Scale bar 500 µm. e, f Immunofluorescence of sections from mouse skin with the indicated antibodies along with DAPI-stained nuclei. In (e), the upper, middle and lower panels show sections taken 15, 15 and 36 days after 4-OHT treatment, respectively. In (f), the upper and middle panels show sections taken 22 and the lower panel 36 days after 4-OHT treatment, respectively. T tumour. Scale bar 100 µm, S100A4 panel, 400 µm
At early stages, the majority of tumour cells stained with Mitf, Dct, S100A4, Sox10 (Fig. 1c, e) and Ki67 (Fig. 1f), whereas at later stages Mitf and Dct staining was progressively lost (Fig. 1e, f). Nonpigmented cells deep in larger tumours stained with KI67, whereas Dct staining was limited to the epidermal surface (Fig. S2d). Relative to their expression at day 22, expression of Mitf, Dct and Tyr was strongly diminished at later times, whereas expression of Sox10 increased (Fig. S3). While Mitf, Dct and Sox10 staining was restricted to tumour cells, Brg1 was present in cells of the tumour, the surrounding stroma and epidermis (Fig. 1e).
Hence, early melanoma cells express Mitf, and melanocyte markers, whereas later tumours invading deeper into the underlying dermis lost expression of Mitf and melanocyte genes, but maintained the SOX10 expression.
An epigenetic map of Braf/Pten melanoma tumours
We generated an epigenetic map of later stage tumours by ChIP-seq for acetylated lysine 27 of histone H3 (H3K27ac), marking active enhancers and promoters, trimethylated lysine 27 of histone H3 (H3K27me3), marking repressed heterochromatin, and RNA polymerase II (Pol II) (Fig. S4a, b). Around 2700 H3K27me3-marked genes with the absence of Pol II were identified (Fig. S4c) and strongly enriched in homeobox-containing and developmental genes involved in the differentiation of multiple ectodermal, mesodermal and endodermal tissues (Data set S1). These genes were stably repressed in the neural crest-derived tumour cells.
In contrast, the H3K27ac and Pol II data identified around 1500 highly transcribed genes with a ‘super-enhancer’ signature consisting of high levels of H3K27ac at wide regulatory elements around and within the coding region and high levels of transcribing Pol II [16,17,18,19,20] (Fig. 2a). Ontology analysis showed enrichment in transcription regulation, nucleosomes, angiogenesis, apoptosis and signalling pathways such as the cAMP response (Fig. 2a and Data set S1). Specifically, several clusters encoding core histones were identified along with transcription factors such as Mycn, Zeb2 and Snai2 or cyclins Ccnd1 and Ccnd3 (Fig. 2b and Data set S1). Core histones genes are short and intron-less and therefore may bias the analysis. Nevertheless, they were not detected in similar analyses of ChIP-seq data for ‘super-enhancer’ signature genes from mouse skeletal muscle or striatal neurons [21, 22]. Their presence in the signature here may rather reflect the active division of the mouse tumour cells.
An epigenetic map of oncogenic Braf-driven mouse melanoma. a Read density analyses of Pol II and H3K27ac of Ensembl annotated genes identifying those with a super-enhancer signature of high transcribing Pol II density and extended H3K27ac labelling. TSS transcription start site, TTS transcription termination site. Results of gene-annotation enrichment analysis of super-enhancer signature genes showing the enriched terms, the enrichment score (ES) and the modified Fisher exact p values. b, c UCSC genome browser view showing Pol II, H3K27ac and H3K27me3 profiles at the indicated loci
The Sox10 locus displayed a ‘super-enhancer’ signature, whereas, although extended regions with strong H3K27ac signals were seen throughout the Mitf locus, only a low level of Pol II was observed at the Mitf-A promoter with no Pol II signal at the Mitf-M promoter (Fig. 2b, c). No Pol II and low or no H3K27ac was seen at the Dct and Tyrp1 promoters (Fig. S5a, b). Moreover, these loci were marked by H3K27me3. At the Rab27a locus, Pol II and H3K27ac signals seen around the TSS corresponded to expression of the divergently transcribed Pigb, whereas Rab27a was selectively marked by H3K27me3 (Fig. S5c). At the Tyr locus, no Pol II was seen at the TSS, but H3K27ac and H3K27me3 were observed at the TSS and the upstream enhancer, suggesting this locus was in a transcriptionally silent, but ‘bivalent’ [23] state (Fig. S5d). Alternatively, tumours may comprise two populations of cells, with either H3K27ac or H3K27me3. Irrespective of the explanation, no Pol II was seen in keeping with their lost expression. In contrast, a ‘super-enhancer’ signature was seen at the Ngfr and Snai2 loci, two neural crest markers (Fig. S5e, f).
A transcriptional signature for human melanoma has been derived [24] comprising MITF and SOX10, but also TFAP2A, LEF1, DLX2, ALX1 PAX3 and GAS7. The Dlx2, Lef1 and Alx1 loci showed a striking absence of Pol II and H3K27ac and high levels of H3K27me3, whereas these loci were strongly marked by MITF, BRG1 and H3K27ac in human 501Mel melanoma cells (Fig. S6a, b). Only low levels of Pol II were seen at Pax3 and for Tfap2a a mixed signature of high levels of H3K27ac and H3K27me3 was seen. Only Gas7 showed high levels of H3K27ac and Pol II, a locus bound by SOX10 in 501Mel cells and in mouse melanocytes [25]. Persistent Gas7 expression may therefore reflect its activation by Sox10, whereas loss of Mitf expression may account for repression of the other genes.
Hence in this model, tumours initially expressed melanocyte markers, but then progressively adopted a de-differentiated state switching off expression of Mitf, melanocyte marker and human melanoma melanocyte signature target genes.
Brg1 and Bptf are required for normal melanoma development in vivo
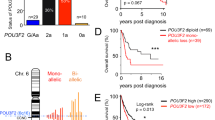
As mentioned in the introduction, BRG1 and BPTF are cofactors for MITF in human melanoma. We investigated their expression levels using TCGA datasets and recently published transcriptome data [26]. The SMARCA4 gene encoding BRG1 was altered in 8% of TCGA melanoma, 8 truncating mutations and 1 deep deletion (Fig. S7a). All cases with loss of function mutations were associated with either BRAF or NRAS mutations, suggesting that loss of SMARCA4 by itself is not a driver. ARID2 was the most affected subunit, altered in 15% of melanoma with frequent truncating mutations, 5 of which occurred in a ‘triple negative’ background. ARID1A and ARID1B were altered at a frequency similar to SMARCA4, but with numerous truncating mutations and deep deletions, several occurring in the triple-negative background. BPTF was altered in 11% of melanoma with no truncating mutations or deep deletions, but frequent amplifications of which 7 of 28 occurred in a triple-negative background.
Consistent with the increased BRG1 protein levels [9], SMARCA4 mRNA expression was upregulated in primary melanoma compared to benign naevi in the Badal data set (Fig. S7b). In contrast, BPTF expression was upregulated in metastatic versus primary melanoma (Fig. S7c). BPTF expression was also higher with NRAS mutations than with BRAF, NF1 or triple wild-type, whereas SMARCA4 was lower expressed with BRAF-mutation (Fig. S7d).
We addressed the requirement for Brg1 and Bptf in mouse melanoma in vivo. They and several subunits of their respective complexes were expressed at relatively constant levels throughout tumour development (Fig. S3). We crossed Tyr::Cre-ERT2:: BrafLSL-V600E/+::Ptenlox/lox animals with mice where the Bptf or Brg1 genes (Smarca4) were floxed such that they would be inactivated by the 4-OHT treatment and tumour formation was monitored.
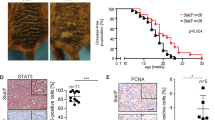
Compared to mice wild-type or heterozygous for Smarca4 and Bptf, inactivation of these genes strongly delayed tumour formation (Fig. 3a, b). Compared to large tumours in wild-type or heterozygous mice, much smaller lesions were observed on homozygous Smarca4 or Bptf mice. The effect was strongest for Bptf-floxed animals where only small pigmented foci were seen by day 44, whereas small tumours were seen by this time in the Smarca4-floxed mice. Comparison of the number of days required for the tumours to reach a 1 cm3 volume showed a strong delay in Bptf-floxed animals (median of 57 ± 6 days, compared to 34 ± 3 days for wild type) and a less marked delay (median of 46 ± 4 days) for Smarca4-floxed mice. Tumour formation was delayed from the earliest stages as the number of pigmented naevi seen at day 26 was dramatically reduced in the Smarca4- and Bptf-floxed animals (Fig. 3d, e).
Brg1 and Bptf are required for normal melanoma growth. a, b Photographs of the 4-OHT-treated area on the backs of Tyr:Cre-ERT2; BrafLSL-V600E/+; Ptenlox/lox mice at the indicated days (29, 38 or 44) after treatment. The genotypes of mice with respect to the Smarca4 and Bptf alleles are indicated to the left. c The number of days required for tumours to reach a volume of 1 cm3 on mice with the indicated genotypes are indicated. ***p < 0.001 with unpaired two-tailed Student’s t test. d Photographs of pigmented naevi developing on the Tam-treated areas on the backs of mice at day 26. The genotypes of mice are indicated to the left. e The numbers of pigmented naevi for each genotype are indicated. ***p < 0.001; **p < 0.005
Sections were prepared from the back skin or the tail of Smarca4-floxed mice 5 days after the 4-OHT application. In the hair follicle and tail epidermis, Dct-labelled melanocytes lacking Brg1 could be identified amongst surrounding Brg1-expressing keratinocytes (Fig. S8a, b). Nevertheless, a Dct-labelled melanocyte expressing Brg1 could be seen in the same follicle. Staining of another animal several days later also revealed a heterogeneous population where most Dct-labelled melanocytes emerging from the hair follicle expressed Brg1, while Brg1-negative cells were still observed (Fig. S8c). In the early-stage dermal tumours, strong and homogeneous Brg1 labelling was observed (Fig. S8d). Immunoblots of later stage tumour extracts revealed expression of Brg1 and other SWI/SNF subunits in tumours from animals with a Smarca4lox/lox genotype and expression of Bptf and other NuRF subunits in tumours from animals with a Bptflox/lox genotype (Fig. S9). Hence, tumours were formed from cells escaping full recombination of the floxed Smarca4 and Bptf alleles.
These results showed that 4-OHT treatment inactivated Brg1 in a many, but not all, melanocytes. As a result, naevi and tumour formation were both delayed as they arose from a smaller initial pool of cells.
Mitf and Sox10 regulate overlapping gene expression programs in mouse melanoma cells
Later stage tumours with wild-type or homozygous floxed Smarca4 or Bptf genotypes were excised, dissociated and the cells cultured in vitro. These in vitro cultured tumour cells expressed Sox10, and while Mitf was not expressed in later stage tumours from which the cells were isolated, immunostaining, RT-qPCR and RNA-seq showed its expression was reactivated after 24–48 h in vitro (Fig. 4a, b and Fig. 5). Cells from wild-type tumours were transfected with siRNAs directed against Sox10 or Mitf leading to pronounced morphological modifications (Fig. 4c) and a strong reduction in the number of viable cells (Fig. 4d).
Sox10 and Mitf are required for tumour cell growth in vitro. a Immunofluorescence staining of cultured mouse melanoma cells with the indicated antibodies and DAPI staining of nuclei. Scale bar 100 µm. 4-OHT treatment leads to loss of Brg1 staining in the cells with floxed Smarca4 alleles, but not in the other genetic backgrounds. b RT-qPCR of Mitf and Sox10 expression 48 h after transfection the indicated siRNAs. c Staining with Crystal Violet and immunolabelling with beta-tubulin and counting of cells 72 h after transfection. N = 3. *p < 0.05; **p < 0.01; ***p < 0.005
RNA-seq analyses of Mitf and Sox10-regulated gene expression. a, b Venn diagrams indicating the number of up- and downregulated genes after siMitf or siSox10 silencing compared to control siRNA. Each Venn shows the overlap between the two data sets using the indicated cut-off criteria. c, d Ontology analyses of regulated genes using GSEA or the DAVID Functional Enrichment tool. e Expression changes of selected cell cycle and mitosis genes after siMitf in mouse or human 501Mel melanoma cells
We performed RNA-seq from the siMitf and siSox10 silenced cells. Using standard criteria on triplicate samples (Log2 fold change >1; <−1 and p < 0.05) or more relaxed criteria (Log2 fold change >0.5; <−0.5 and p < 0.05), we found a large overlap between the regulatory programs of Mitf and Sox10 (Fig. 5a, b and Data set S2) consistent with the binding of MITF and SOX10 together at regulatory elements in human melanoma cells [4]. Ontology analyses showed pigmentation and UV-response genes were downregulated in both conditions along with cell membrane and extracellular proteins (Fig. 5c, d). Paradoxically, a large collection of cell cycle and mitosis genes positively coregulated by MITF and SOX10 in human melanoma cells [4, 27] appeared negatively regulated in mouse melanoma cells and were upregulated upon siMitf or siSox10 silencing (Fig. 5e). These cell cycle genes were observed only when more relaxed criteria were used showing they were not potently upregulated, but they were strongly represented in the 511 Mitf and Sox10 co-upregulated genes (Fig. 5b).
Brg1 and Bptf regulate extensive gene expression programs in mouse melanoma cells
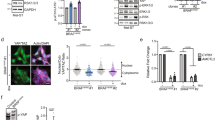
Cultured cells of appropriate genotypes were treated with 4-OHT to recombine residual floxed Smarca4 or Bptf alleles. Treatment of Smarca4lox/lox cells led to a loss not only of Brg1, but also of multiple other subunits of the PBAF complex and of Sox10 (Fig. 6a). Treatment of Bptflox/lox cells led to diminished Bptf and Smarca5 expression, but not of Smarca1 (Fig. 6b). 4-OHT treatment of wild-type tumour cells had little effect on their morphology, whereas treatment of cells from Smarca4lox/lox or Bptflox/lox tumours led to a major change in cell morphology, cytoskeleton reorganisation and reduced cytoplasmic volume (Fig. 6c). In addition, while 4-OHT treatment had little effect on the viability of wild-type cells, viability of Smarca4lox/lox or Bptflox/lox cells was strongly diminished (Fig. 6d) with a concomitant increase in apoptosis 72 h after treatment (Fig. 6e). After 10 days, almost no viable Smarca4lox/lox or Bptflox/lox tumour cells persisted.
4-OHT-induced inactivation of Brg1 and Bptf in cultured mouse melanoma cells. a, b Immunoblots from the indicated cells 48 h after treatment with 4-OHT or DMSO as control. The position of migration of molecular mass markers is indicated. c Bright field views and immunolabelling with beta-tubulin of cultured melanoma cells with the indicated genotypes 48 h after treatment with DMSO or 4-OHT. Scale bar 100 µm d Number of viable cells counted in each condition 48 h after the indicated treatments. N = 3. **p < 0.01. e FACS analyses to detect Annexin V-stained apoptotic cells 48 h after 4-OHT treatment
RNA-seq following 4-OHT-induced Brg1 or Bptf inactivation revealed de-regulation of more than 2000–3000 genes with a notable overlap of the programs controlled by each factor (Fig. 7a and Data set S2). Bptf inactivation deregulated genes involved in cell cycle and mitosis, innate immunity, secreted glycoproteins and oxidative phosphorylation (Fig. 7b). Brg1 inactivation also downregulated innate immunity, secreted glycoprotein genes and upregulated genes of the TP53 pathway (Fig. 7c). These data show that Brg1 and Bptf regulate extensive gene expression programs affecting multiple functional pathways in mouse melanoma cells.
RNA-seq analyses of Brg1- and Bptf-regulated gene expression. a Venn diagram indicating the number of up- and downregulated genes after Brg1 or Bptf inactivation compared to the 4-OHT-treated control cells. The Venn shows the overlap between the two data sets. b, c Ontology analyses of regulated genes using GSEA or the DAVID Functional Enrichment tool. d, e Venn diagrams indicating the overlap between the Mitf, Sox10, Brg1 and Bptf regulated genes highlighting the overlap between the data sets. The RF for the common up- and downregulated genes are indicated
A comparison of the Mitf, Sox10, Brg1 and Bptf regulated genes identified sets of coregulated genes (Fig. 7d, e). Using hypergeometric probability to calculate the representation factor (RF), we found in each case a highly significant overlap. More than 50% of genes downregulated by Mitf or Sox10 silencing were also down following Brg1 inactivation (Fig. 7d, e). This group of genes were enriched in signalling pathways such as Hippo, Wnt and MAP kinase, as well as melanogenesis (Data set S2). Brg1 and Bptf therefore coregulated a notable fraction of Mitf and Sox10 target genes. However, Brg1 and Bptf regulated additional genes and functional pathways, showing they acted as more general cofactors.
Discussion
Similarities and differences between human melanoma and mouse oncogenic BRAF-driven melanoma
The genetically modified mouse model described here was used to study the roles of CTNNB1, PIKC3A, DNMT3B and PDK1 in melanoma development [12, 13, 15, 26, 28]. At least two models have been independently developed [12, 14] using Cre-ERT2 transgenes with differing expression profiles. Despite this, the resulting tumours have not been fully characterised, particularly at the epigenetic level. At early stages, tumours comprised Sox10, Dct, and Mitf-expressing melanocytes, whereas cells in later tumours reaching deep into the dermis lost expression of melanocyte markers and gained that of invasion markers, such as Zeb1 and Zeb2, but with persistent expression of Sox10.
Primary dermal melanomas are a rare occurrence in humans. The focal whorling patterns and loose myxoid background seen in later stage mouse tumours also differentiate them from what is typically observed in human melanoma. However, using a Braf/Pten genetic model, Kohler et al. [29] performed cell lineage tracing to show that melanoma developed from mature melanocytes in the tail epidermis and pigmented Mitf-expressing bulb melanocytes analogous to what we observed here. Thus, transformed melanocytes originate from the epidermal compartment, but invasion of dermis is rapid occurring a few days after 4-OHT exposure. In the tail, transformed melanocytes clonally expanded in the epidermis, but underwent ‘de-differentiation’ with loss of Mitf and melanocyte markers upon invasion in the dermis [29]. In contrast, we found that transformed melanocytes from the hair follicles continue to express Mitf and melanocyte markers after invasion of the dermis and show ‘de-differentiation’ only later as they invade more deeply within the dermis.
Consistent with the loss of Mitf expression, no Pol II was detected at the Mitf-M isoform promoter. Moreover, Pol II was lost from other melanocyte lineage markers that were often organised into H3K27me3-marked heterochromatin. Thus, our observations are consistent with the idea of an epigenetic switch between early stages with expression of Mitf and melanocyte markers to a later de-differentiated state where these genes loose Pol II and often gain H3K27me3. De-differentiation takes place in tumour cells expressing Sox10 and Brg1, an axis normally required for melanocyte differentiation and activation of melanocyte marker genes [8]. The mechanisms responsible for bypassing this axis remain to be determined.
Transcriptome and epigenetic analyses of human melanomas showed the existence of MITF-high and MITF-low type cells [30, 31]. More recent classifications based on vulnerability to ferroptosis [32] and single-cell analyses of patient-derived xenografts [33] highlighted additional cell states, in particular neural crest-like cells expressing SOX10, but not MITF and ‘mesenchymal’ cells expressing neither MITF nor SOX10. The late-stage mouse tumour cells were reminiscent of a neural crest-like phenotype with the absence of melanocyte markers, but strong expression of Sox10 and Ngfr1. However, when cultured in vitro, they rapidly adopted a more differentiated phenotype with re-expression of Mitf and other melanocyte markers, suggesting that their expression was repressed in tumours by signals from the micro-environment. Previous reports showed that a proinflammatory micro-environment can induce a reversible de-differentiation and ‘mesenchymal-like’ phenotype through a mechanism involving c-JUN-mediated repression of MITF expression [34, 35]. Such dynamic transitions may be explained by the maintenance of the Mitf locus in an ‘open’ state retaining strong H3K27ac labelling even at the region around the Mitf-M promoter, facilitating re-recruitment of Pol II when the tumour cells were grown in the absence of repressive microenvironmental signals. In contrast, in human ‘mesenchymal’ cells exemplified by those profiled by Verfaillie et al. [31], the MITF and SOX10 loci displayed a more stable repressed state with low or no H3K27ac.
Contribution of Brg1 and Bptf to Mitf/Sox10 regulated gene expression in mouse melanoma
Here we show that Brg1 and Bptf are required for normal development of murine melanoma in vivo. Melanoma was strongly delayed by their inactivation, but Brg1 or Bptf-expressing tumours eventually developed from nonrecombined cells. This observation is in accordance with a previous dropout shRNA screen that identified BRG1 as an essential actor in human melanoma cells [36]. 4-OHT-induced Brg1 or Bptf inactivation in cultured tumour cells deregulated numerous genes affecting multiple functional pathways and was accompanied by apoptosis and loss of cell viability. These data highlight the essential roles of these chromatin remodelling factors in the regulation of gene expression in mouse melanoma cells.
In mouse melanoma cells, Mitf and Sox10 coregulated overlapping gene sets. A majority of Mitf-regulated genes were also regulated by Sox10, whereas Sox10 regulated an additional set of genes independently of Mitf. We noted a collection of cell cycle and mitosis genes whose expression was mildly induced upon Mitf/Sox10 silencing. Amongst these are numerous genes whose expression is normally activated by MITF and SOX10 in human melanoma cells. The basis for this apparent conversion of MITF/SOX10 from activators of these genes in human melanoma cells to repressors in in vitro cultured mouse melanoma cells remains to be established.
We reported that BRG1 and BPTF acted as cofactors coregulating subsets of MITF and SOX10 target genes in human melanoma [4, 9]. Here, we identified genes coregulated in mouse melanoma. This can in part be explained by reduced Sox10 expression after Brg1 inactivation. This role of Brg1 in activating Sox10 expression and acting as a potential cofactor for Mitf/Sox10 accounts for its essential role not only in melanoma, but more generally for melanocyte biology as previously shown [4, 8, 25, 37]. An essential role for BRG1 in melanocyte and melanoma biology is in line with the observation that it is rarely subject to loss-of-function mutations in melanoma and only in the background of BRAF or NRAS mutations. In contrast, ARID2, ARID1A and ARID1B showed more frequent loss of function even in the triple-negative background suggesting they may be tumour suppressors [38, 39]. SWI/SNF subunits therefore appear to make distinct contributions to melanoma, some acting as putative tumour suppressors and others such as BRG1 essential for tumour growth.
Methods
Mice and genotyping
All animal experiments were performed in accordance with the European and national guidelines and policies (2010/63/UE directive and French decree 2013-118) and with the approval of the National Ethics Committee. Mice bearing the Tyr:Cre-ERT2 transgene [40], BrafLSL-V600E/+ [41], and floxed alleles of Pten [38], Smarca4 [39] or Bptf [42] alleles were bred and kept on C57BL/6J background to generate experimental mice with the desired genotypes. Genotyping was carried out by PCR analysis of genomic tail, skin or tumour DNA with primers detailed in the respective publications.
Tumour induction and growth analysis
4-hydroxytamoxifen (4-OHT) was prepared freshly as a 25 mg/mL solution (65 mM) of 4-OHT (70% Z-isomer, Sigma) in dimethylsulfoxide (DMSO) and further diluted with 100% ethanol to a 1.9 mg/mL working solution (5 mM). Localised melanomas were induced by topical application of 5 µL of 4-OHT on the shaved back skin of 6−7-week-old mice for 2 consecutive days. For distal tail or ear inductions, 2 µL of 4-OHT was applied. Tumour growth was monitored twice weekly by digital photography of the skin, including a size reference. Tumour size was then analysed using ImageJ software. For Fig. 3e, only pigmented lesions with a minimal diameter of 1 mm were included for counting.
Primary cell culture and treatment
Primary melanoma cell cultures were derived from resected mouse tumours as described elsewhere [43] and cultured in Ham-F12 supplemented with 5% foetal calf serum. For in vitro Cre-ERT2-induced recombination, cultured melanoma cells were treated with 1 μM 4-OHT or vehicle (DMSO) for 48 h. The siRNA knockdown of MITF (SMARTpool L-047441-00, Dharmacon) and SOX10 (SMARTpool L-049957-01, Dharmacon) was performed with Lipofectamine RNAi max (Invitrogen, La Jolla, CA, USA) for 48 h following the manufacturer’s instructions. To assess cell viability, 50,000 cells were seeded into 24-well plates, treated and grown for 3 days, stained with Cristal Violet and counted under the microscope.
Immunofluorescence and histochemistry
Immunofluorescence staining of primary cells and formalin-fixed, paraffin-embedded skin and tumour samples was performed as described previously [4]. The following antibodies were used: goat anti-Dct (Santa Cruz Biotechnology, sc-10451), rabbit anti-Sox10 (Abcam, ab155279), rabbit anti-Brg1 (ab110641), rabbit anti-Mitf (Sigma, St Louis, MO), rabbit anti-ZEB1 (Santa Cruz, sc-25388), rabbit anti-ZEB2, rabbit anti-β-tubulin (Abcam, ab6046), Alexa 488 donkey anti-goat, and Alexa 555 donkey anti-rabbit (Invitrogen, Carlsbad, CA). For histology analysis, sections were stained with haematoxylin and eosin (H&E) following standard procedures. Immunostaining of melanoma cells in vitro was performed on cells after less than three passages and were seeded and grown on glass coverslips for 2 days prior to treatment with 4-OHT, DMSO or siRNAs.
Protein extraction and western blotting
Whole-cell extracts were prepared by the standard freeze-thaw technique using LSDB 500 buffer (500 mM KCl, 25 mM Tris at pH 7.9, 10% glycerol (v/v), 0.05% NP-40 (v/v), 1 mM dithiothreitol (DTT), and protease inhibitor cocktail) except for tumour pieces that were homogenised in RIPA (50 mM Tris pH 8.0, 150 mM sodium chloride, 0.5 mM Ethylenediaminetetraacetic acid (EDTA), 0.1% SDS, 0.5% sodium deoxycholate, 1% NP-40). Cell lysates were subjected to SDS–polyacrylamide gel electrophoresis (SDS-PAGE) and proteins were transferred onto a nitrocellulose membrane. Membranes were incubated with primary antibodies in 5% dry fat milk and 0.01% Tween-20 overnight at 4 °C. The membrane was then incubated with HRP-conjugated secondary antibody (Jackson ImmunoResearch) for 1 h at room temperature, and visualised using the ECL detection system (GE Healthcare).
Apoptosis and FACs analysis
Following treatments, primary cells were harvested and stained for annexin V using Annexin V-FITC Apoptosis Detection Kit (Sigma) and propidium iodide following the manufacturer instructions. Cells were analysed on an LSRII Fortessa (BD Biosciences) and data were analysed with Flowjo software (Tree Star).
Chromatin-immunoprecipitation and sequencing
For in vivo ChIP-seq, freshly resected mouse melanoma tumours were separated from the epidermis, cut into small pieces and homogenised by douncing in hypotonic buffer. Nuclei were isolated by centrifugation on a sucrose cushion (1.2 M sucrose, 60 mM KCl, 15 mM NaCl, 5 mM MgCl2, 0.1 mM EDTA, 15 mM Tris-HCl (pH 7.5), 0.5 mM DTT, 0.1 mM phenylmethylsulfonyl fluoride (PMSF), PIC). For Pol II ChIP, nuclei were further fixed in 0.4% PFA for 10 min and sonicated with a Covaris S220 as described [27]. Alternatively, H3K27ac and H3K27me3 ChIP experiments were performed on native Mnase-digested chromatin as described previously [4]. ChIP-seq libraries were prepared and sequenced as single-end 50-base reads, peak detection was performed using MACS [44] http://liulab.dfci.harvard.edu/MACS/). Datasets were normalised for the number of unique mapped reads for subsequent comparisons. Global clustering analysis and quantitative comparisons were performed using seqMINER [45] and R (http://www.r-project.org/). Gene ontology analyses were performed using the functional annotation clustering function of DAVID (http://david.abcc.ncifcrf.gov/).
RNA preparation, quantitative PCR and RNA-seq analysis
RNA isolation was performed according to standard procedure (Qiagen kit). qRT-PCR was carried out with SYBR Green I (Qiagen) and Multiscribe Reverse Transcriptase (Invitrogen) and monitored using a LightCycler 480 (Roche). GAPDH gene expression was used to normalise the results. Primer sequences for each cDNA were designed using Primer3 Software and are available upon request. RNA-seq was performed essentially as previously described. Gene ontology analyses were performed with the Gene Set Enrichment Analysis software GSEA v3.0 using the hallmark gene sets of the Molecular Signatures Database v6.2 and the functional annotation clustering and KDEGG pathway functions of DAVID (https://david.ncifcrf.gov/).
Analyses of public datasets
Analyses of mutations in the melanoma TCGA datasets were performed using the cBiopotal website http://www.cbioportal.org/. Melanoma (SKCM) RNAseqV2 data generated by the TCGA Research Network (http://cancergenome.nih.gov/), processed and normalised using RSEM were downloaded together with the patient clinical features (BRAF, NRAS, NF1 mutational status; Clark score of the lesion; primary tumour or metastatic origin) from cBioportal. Expression values of SMARCA4 and BPTF were extracted and compared between the groups indicated in the figures by Kruskal−Wallis test, assuming nonparametric distribution (***p < 0.001, **p < 0.01, *p < 0.05).
Analysis of SMARCA4 and BPTF expression in nevi and primary melanoma lesions was performed by extracting their normalised expression levels from RNA-seq data obtained from Badal et al. [46]. Two-tailed unpaired t test was used for statistical significance (***p < 0.001, **p < 0.01, *p < 0.05).
Data availability
All sequencing data in this paper have been submitted to the Geo database under the accession number. SuperSeries GSE129621.
References
Clapier CR, Iwasa J, Cairns BR, Peterson CL. Mechanisms of action and regulation of ATP-dependent chromatin-remodelling complexes. Nat Rev Mol Cell Biol. 2017;18:407–22.
Keenen B, Qi H, Saladi SV, Yeung M, de la Serna IL. Heterogeneous SWI/SNF chromatin remodeling complexes promote expression of microphthalmia-associated transcription factor target genes in melanoma. Oncogene. 2010;29:81–92.
de la Serna IL, Ohkawa Y, Higashi C, Dutta C, Osias J, Kommajosyula N, et al. The microphthalmia-associated transcription factor requires SWI/SNF enzymes to activate melanocyte-specific genes. J Biol Chem. 2006;281:20233–41.
Laurette P, Strub T, Koludrovic D, Keime C, Le Gras S, Seberg H, et al. Transcription factor MITF and remodeller BRG1 define chromatin organisation at regulatory elements in melanoma cells. eLife. 2015. https://doi.org/10.7554/eLife.06857.
Weider M, Kuspert M, Bischof M, Vogl MR, Hornig J, Loy K, et al. Chromatin-remodeling factor Brg1 is required for Schwann cell differentiation and myelination. Dev Cell. 2012;23:193–201.
Seberg HE, Van Otterloo E, Cornell RA. Beyond MITF: multiple transcription factors directly regulate the cellular phenotype in melanocytes and melanoma. Pigment Cell Melanoma Res. 2017;30:454–66.
Lin H, Wong RP, Martinka M, Li G. BRG1 expression is increased in human cutaneous melanoma. Br J Dermatol. 2010;163:502–10.
Marathe HG, Watkins-Chow DE, Weider M, Hoffmann A, Mehta G, Trivedi A, et al. BRG1 interacts with SOX10 to establish the melanocyte lineage and to promote differentiation. Nucleic Acids Res. 2017:45:6442–58. https://doi.org/10.1093/nar/gkx259.
Koludrovic D, Laurette P, Strub T, Keime C, Le Coz M, Coassolo S, et al. Chromatin-remodelling complex NURF is essential for differentiation of adult melanocyte stem cells. PLoS Genet. 2015;11:e1005555.
Cancer Genome Atlas N. Genomic classification of cutaneous melanoma. Cell. 2015;161:1681–96.
Dar AA, Nosrati M, Bezrookove V, de Semir D, Majid S, Thummala S, et al. The role of BPTF in melanoma progression and in response to BRAF-targeted therapy. J Natl Cancer Inst. 2015;107: pi: djv034. https://doi.org/10.1093/jnci/djv034.
Dankort D, Curley DP, Cartlidge RA, Nelson B, Karnezis AN, Damsky WE Jr., et al. Braf(V600E) cooperates with Pten loss to induce metastatic melanoma. Nat Genet. 2009;41:544–52.
Marsh Durban V, Deuker MM, Bosenberg MW, Phillips W, McMahon M. Differential AKT dependency displayed by mouse models of BRAFV600E-initiated melanoma. J Clin Invest. 2013;123:5104–18.
Dhomen N, Reis-Filho JS, da Rocha Dias S, Hayward R, Savage K, Delmas V, et al. Oncogenic Braf induces melanocyte senescence and melanoma in mice. Cancer Cell. 2009;15:294–303.
Micevic G, Muthusamy V, Damsky W, Theodosakis N, Liu X, Meeth K, et al. DNMT3b modulates melanoma growth by controlling levels of mTORC2 component RICTOR. Cell Rep. 2016;14:2180–92.
Whyte WA, Orlando DA, Hnisz D, Abraham BJ, Lin CY, Kagey MH, et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell. 2013;153:307–19.
Hnisz D, Abraham BJ, Lee TI, Lau A, Saint-Andre V, Sigova AA, et al. Super-enhancers in the control of cell identity and disease. Cell. 2013;155:934–47.
Pott S, Lieb JD. What are super-enhancers? Nat Genet. 2015;47:8–12.
Sengupta S, George RE. Super-enhancer-driven transcriptional dependencies in cancer. Trends Cancer. 2017;3:269–81.
Siersbaek R, Madsen JGS, Javierre BM, Nielsen R, Bagge EK, Cairns J, et al. Dynamic rewiring of promoter-anchored chromatin loops during adipocyte differentiation. Mol Cell. 2017;66:420–35 e425.
Joshi S, Davidson G, Le Gras S, Watanabe S, Braun T, Mengus G, et al. TEAD transcription factors are required for normal primary myoblast differentiation in vitro and muscle regeneration in vivo. PLoS Genet. 2017;13:e1006600.
Achour M, Le Gras S, Keime C, Parmentier F, Lejeune FX, Boutillier AL, et al. Neuronal identity genes regulated by super-enhancers are preferentially down-regulated in the striatum of Huntington’s disease mice. Hum Mol Genet. 2015;24:3481–96.
Bernstein BE, Mikkelsen TS, Xie X, Kamal M, Huebert DJ, Cuff J, et al. A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell. 2006;125:315–26.
Rambow F, Job B, Petit V, Gesbert F, Delmas V, Seberg H, et al. New functional signatures for understanding melanoma biology from tumor cell lineage-specific analysis. Cell Rep. 2015;13:840–53.
Fufa TD, Harris ML, Watkins-Chow DE, Levy D, Gorkin DU, Gildea DE, et al. Genomic analysis reveals distinct mechanisms and functional classes of SOX10-regulated genes in melanocytes. Hum Mol Genet. 2015;24:5433–50.
Damsky WE, Curley DP, Santhanakrishnan M, Rosenbaum LE, Platt JT, Gould Rothberg BE, et al. beta-catenin signaling controls metastasis in Braf-activated Pten-deficient melanomas. Cancer Cell. 2011;20:741–54.
Strub T, Giuliano S, Ye T, Bonet C, Keime C, Kobi D, et al. Essential role of microphthalmia transcription factor for DNA replication, mitosis and genomic stability in melanoma. Oncogene. 2011;30:2319–32.
Scortegagna M, Ruller C, Feng Y, Lazova R, Kluger H, Li JL, et al. Genetic inactivation or pharmacological inhibition of Pdk1 delays development and inhibits metastasis of Braf(V600E)::Pten(-/-) melanoma. Oncogene. 2014;33:4330–9.
Kohler C, Nittner D, Rambow F, Radaelli E, Stanchi F, Vandamme N. et al. Mouse cutaneous melanoma induced by mutant BRaf arises from expansion and dedifferentiation of mature pigmented melanocytes. Cell Stem Cell. 2017;21:679–93 e676.
Goodall J, Carreira S, Denat L, Kobi D, Davidson I, Nuciforo P, et al. Brn-2 represses microphthalmia-associated transcription factor expression and marks a distinct subpopulation of microphthalmia-associated transcription factor-negative melanoma cells. Cancer Res. 2008;68:7788–94.
Verfaillie A, Imrichova H, Atak ZK, Dewaele M, Rambow F, Hulselmans G, et al. Decoding the regulatory landscape of melanoma reveals TEADS as regulators of the invasive cell state. Nat Commun. 2015;6:6683.
Tsoi J, Robert L, Paraiso K, Galvan C, Sheu KM, Lay J. et al. Multi-stage differentiation defines melanoma subtypes with differential vulnerability to drug-induced iron-dependent oxidative stress. Cancer Cell. 2018;33:890–904 e895.
Rambow F, Rogiers A, Marin-Bejar O, Aibar S, Femel J, Dewaele M. et al. Toward minimal residual disease-directed therapy in melanoma. Cell. 2018;174:843–55 e819.
Landsberg J, Kohlmeyer J, Renn M, Bald T, Rogava M, Cron M, et al. Melanomas resist T-cell therapy through inflammation-induced reversible dedifferentiation. Nature. 2012;490:412–6.
Riesenberg S, Groetchen A, Siddaway R, Bald T, Reinhardt J, Smorra D, et al. MITF and c-Jun antagonism interconnects melanoma dedifferentiation with pro-inflammatory cytokine responsiveness and myeloid cell recruitment. Nat Commun. 2015;6:8755.
Bossi D, Cicalese A, Dellino GI, Luzi L, Riva L, D’Alesio C, et al. In vivo genetic screens of patient-derived tumors revealed unexpected frailty of the transformed phenotype. Cancer Disco. 2016;6:650–63.
Harris ML, Buac K, Shakhova O, Hakami RM, Wegner M, Sommer L, et al. A dual role for SOX10 in the maintenance of the postnatal melanocyte lineage and the differentiation of melanocyte stem cell progenitors. PLoS Genet. 2013;9:e1003644.
Suzuki A, Yamaguchi MT, Ohteki T, Sasaki T, Kaisho T, Kimura Y, et al. T cell-specific loss of Pten leads to defects in central and peripheral tolerance. Immunity. 2001;14:523–34.
Indra AK, Dupe V, Bornert JM, Messaddeq N, Yaniv M, Mark M, et al. Temporally controlled targeted somatic mutagenesis in embryonic surface ectoderm and fetal epidermal keratinocytes unveils two distinct developmental functions of BRG1 in limb morphogenesis and skin barrier formation. Development. 2005;132:4533–44.
Yajima I, Belloir E, Bourgeois Y, Kumasaka M, Delmas V, Larue L. Spatiotemporal gene control by the Cre-ERT2 system in melanocytes. Genesis. 2006;44:34–43.
Dhomen N, Dias SD, Hayward R, Ogilvie L, Hedley D, Delmas V, et al. Inducible expression of (V600E)Braf using tyrosinase-driven Cre recombinase results in embryonic lethality. Pigment Cell Melanoma Res. 2009;23:112–20.
Landry JW, Banerjee S, Taylor B, Aplan PD, Singer A, Wu C. Chromatin remodeling complex NURF regulates thymocyte maturation. Genes Dev. 2011;25:275–86.
Gallagher SJ, Luciani F, Berlin I, Rambow F, Gros G, Champeval D, et al. General strategy to analyse melanoma in mice. Pigment Cell Melanoma Res. 2011;24:987–8.
Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE, et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 2008;9:R137.
Ye T, Krebs AR, Choukrallah MA, Keime C, Plewniak F, Davidson I, et al. seqMINER: an integrated ChIP-seq data interpretation platform. Nucleic Acids Res. 2011;39:e35.
Badal B, Solovyov A, Di Cecilia S, Chan JM, Chang LW, Iqbal R, et al. Transcriptional dissection of melanoma identifies a high-risk subtype underlying TP53 family genes and epigenome deregulation. JCI Insight. 2017;2:pii: 92102. https://doi.org/10.1172/jci.insight.92102.
Acknowledgements
We thank R. Marais for the BrafLSL-V600E mice, D. Metzger for floxed Smarca4 and Pten mice, C. Wu for Bptf floxed mice, all the staff of the IGBMC high-throughput sequencing facility, a member of “France Génomique” consortium (ANR10-INBS-09-08). This work was supported by grants from the CNRS, the INSERM, Ligue National Contre le Cancer; Institut National du Cancer; ITMO-Cancer, Agence National de la Recherche, ANR10-Labex-0030-INRT. PL was supported by fellowships from the Ministère de l’enseignement supérieur et de la recherche and Fondation ARC pour la recherche sur le cancer. ID and LL are “équipes labellisées” of the Ligue Nationale contre le Cancer.
Author information
Authors and Affiliations
Contributions
PL and SC performed and analysed all of the wet lab experiments. PL and GM generated and maintained the mouse lines. PL and GD performed the bioinformatics analysis. IM performed the tail and tumour genotyping. PS performed histopathology analysis, LL provided mice, performed histology analysis and analysed the data, ML and WY provided samples from the lymph nodes and skin sections, ID, PL, SC and LL conceived experiments, analysed the data and wrote the paper.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Edited by S. Fulda
Rights and permissions
About this article
Cite this article
Laurette, P., Coassolo, S., Davidson, G. et al. Chromatin remodellers Brg1 and Bptf are required for normal gene expression and progression of oncogenic Braf-driven mouse melanoma. Cell Death Differ 27, 29–43 (2020). https://doi.org/10.1038/s41418-019-0333-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-019-0333-6
This article is cited by
-
Ssc-mir-221-3p regulates melanin production in Xiang pigs melanocytes by targeting the TYRP1 gene
BMC Genomics (2023)
-
PHF6 recruits BPTF to promote HIF-dependent pathway and progression in YAP-high breast cancer
Journal of Translational Medicine (2023)
-
The journey from melanocytes to melanoma
Nature Reviews Cancer (2023)
-
SMARCB1 regulates a TFCP2L1-MYC transcriptional switch promoting renal medullary carcinoma transformation and ferroptosis resistance
Nature Communications (2023)
-
Current understanding of epigenetics role in melanoma treatment and resistance
Cancer Cell International (2022)