Abstract
Necroptosis is a programmed form of necrotic cell death, which is tightly regulated by the necroptotic signaling pathway containing receptor-interacting protein (RIP)1, RIP3, and mixed-lineage kinase domain-like (MLKL) protein. In addition to the RIP1-RIP3-MLKL axis, other factors regulating necroptosis are still largely unknown. Here a cell-based small-molecule screening led to the finding that BET inhibitors protected cells from necroptosis in the TNFα/Smac-mimetic/Z-VAD-FMK (TSZ)-induced cell necroptosis model. Mechanistic studies revealed that BET inhibitors acted by downregulating MLKL expression. Further research demonstrated that BRD4, IRF1, P-TEFb, and RNA polymerase II formed a transcription complex to regulate the expression of MLKL, and BET inhibitors interfered with the transcription complex formation. In necroptosis-related disease model, the BET inhibitor JQ-1 showed promising therapeutic effects. Collectively, our studies establish, for the first time, BRD4 as a new epigenetic factor regulating necroptosis, and highlight the potential of BET inhibitors in the treatment of necroptosis-related diseases.
Similar content being viewed by others
Introduction
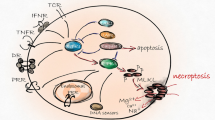
Programmed necrosis, termed necroptosis, has been implicated in many physiological and pathological processes, such as viral infection, inflammation, embryonic development, tissue injury, and cancer metastasis [1,2,3,4]. Necroptosis is regulated by the serine/threonine protein kinases receptor-interacting proteins 1 and 3, (RIP1 and RIP3) and the effector protein mixed-lineage kinase domain like (MLKL) [5, 6]. Necroptosis signaling can be triggered by various stimuli, among which tumor necrosis factor alpha (TNFα) is the most widely studied. TNFα binds to TNF receptor 1 (TNFR1), recruiting the adapter protein TNF-receptor-associated death domain (TRADD), which subsequently signals to RIP1 [7]. RIP1 associates with RIP3 to form a signaling complex called necrosome, which mediates necroptosis [8, 9]. RIP3 in the necrosome recruits and phosphorylates the pseudokinase MLKL [6]. The phosphorylated MLKLs form oligomers and translocate to the cytomembrane to execute necroptosis [10, 11].
Although the basic RIP1-RIP3-MLKL axis has been identified, other regulatory factors of necroptosis are largely unknown [12,13,14]. The discovery of new regulatory factors will not only enhance our understanding of the nature of cell necroptosis, but also provide evidence for the development of novel interventions against necroptosis-related diseases. In this study, we aimed to identify the novel regulatory factors of necroptosis. Instead of using traditional genetic manipulation methods, a small molecular strategy was adopted. Our results demonstrated, for the first time, the bromodomain protein 4 (BRD4) as a new epigenetic factor regulating necroptosis. In addition, our work also revealed the potential of bromodomain and extra-terminal domain (BET) inhibitors in the treatment of necroptosis-related diseases.
Results
Discovery of BET inhibitors capable of protecting cells from necroptosis
To identify new factors involved in the regulation of necroptosis, we adopted the classical TNFα/Smac-mimetic/Z-VAD-FMK (TSZ)-induced HT29 cell necroptosis model [6] to screen an in-house targeted small molecular library called T-Lib; this library collects compounds with their on-targets being known, and right now it contains 3600 agents. In the first round screening, TSZ and test compounds (3 µM) were added to the cultured HT29 cells at the same time; the screening scheme was called scheme I (Fig. 1a). From this screening, we found that eight compounds, including necrostatin-1, pazopanib, dabrafenib, DCC-2036, GSK2656157, GSK2606414, SKLB101126, and JNK-IN-8, significantly protected HT29 cells from TSZ-induced necroptosis (the survival rate > 60%; Fig. 1b). A target survey showed that necrostatin-1, pazopanib, dabrafenib, DCC-2036, GSK2656157, and GSK2606414 have been reported to be active against RIP1 or RIP3, either by an on-target or by an off-target effect [15,16,17,18,19]. Our further kinase profiling assays revealed that both SKLB101126 and JNK-IN-8 were also potent RIP1 inhibitors (Supplementary Fig. S1). These results implied that we did not find new molecules involved in the regulation of necroptosis in this screening.
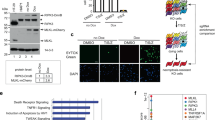
BET inhibitors are capable of protecting cells from necroptosis. a Two screening schemes (I and II). In scheme I, TSZ and test compounds (3 μM) were added at the same time. In scheme II, test compounds (3 μM) were added in advance. After a 72-h treatment, TSZ was added. b The compounds that protected cells from necroptosis in both the scheme I and II assays. Necroptosis of HT29 cells was induced by TNFα (10 ng/ml), Smac-mimetic (100 nM), and Z-VAD-FMK (20 μM). The cell survival rates were measured by CCK8 staining. Columns, mean (n = 3); bars, SD. ***P < 0.001. c The compounds that protected cells from necroptosis in the scheme II screening but not in scheme I screening. Columns, mean (n = 3); bars, SD. ***P < 0.001. d Dual staining with CytoCalcein™ Violet 450 (blue, living cell) and 7-AAD (red, necrotic cells) of HT29 cells treated with or without TSZ for 24 h. Columns, mean (n = 3); bars, SD. ***P < 0.001. Scale bar, 200μm. e Dose–response effect of JQ-1 on protecting cells from necroptosis. Columns, mean (n = 3); bars, SD. ***P < 0.001
We then modified the screening scheme. In the modified scheme (called scheme II; Fig. 1a), HT29 cells were pretreated with test compounds for three days and then TSZ was added to induce necroptosis. As expected, all the compounds that were active in the scheme I screening also showed comparable potency in the scheme II screening. Interestingly, five compounds (JQ-1, PFI-1, I-BET-151, I-BET-762, and AZD3514) were found to be active (the survival rate > 60%) in the scheme II screening but inactive in the scheme I screening (Fig. 1c). Among these compounds, JQ-1, PFI-1, I-BET-151, and I-BET-762 are known inhibitors of the BET proteins [20,21,22,23]; the BET proteins recognize acetylated chromatin through their bromodomains (BDs) and help in regulating gene expression. Also, AZD3514 is an inhibitor of androgen receptor and recently has been reported to be able to inhibit BET proteins due to an off-target effect [24]. Dual staining with CytoCalcein™ Violet 450 (for living cells) and 7-AAD (for necrotic cells) showed that JQ-1 inhibited TSZ-induced necroptosis and rescued cell survival (Fig. 1d). To substantiate this finding, dose–response experiments were then carried out using the five active compounds. As shown in Fig. 1e and Supplementary Fig. S2a-d, all the tested compounds displayed dose-dependent inhibitory effects on TSZ-induced necroptosis in HT29 cells. Similar effects were observed in U937 and L929 cell lines (Supplementary Fig. S2e, f), suggesting cell line-independent effects. To solidify the claim that BET inhibitors have a generalized anti-necroptotic effect, the influence of BET inhibitors on other necroptotic insults (besides TSZ) was assessed. Here the combination of Trail, Smac mimetic, and Z-VAD-FMK was used as an example. In these assays, all the BET inhibitors, including JQ-1, PFI-1, I-BET-151, and I-BET-762, exhibited dose-dependent inhibitory effects on necroptosis induced by Trail/Smac mimetic/Z-VAD-FMK in HT29 cells, indicating a generalized anti-necroptotic effect (Supplementary Fig. S2g). Here it is worth mentioning that there were totally 85 epigenetic inhibitors in our screening library, and except the BET inhibitors, the remaining agents, which are summarized in Supplementary Table S1, did not display any protective effect on TSZ-induced necroptosis.
Finally, to determine whether the 3-day treatment with BET inhibitors protects against TSZ-induced necroptosis by promoting cell proliferation and/or inhibiting cell apoptosis, we further examined the effects of the 3-day treatment with JQ-1 on the cell cycle, proliferation, and apoptosis of HT29 cells. The results showed that treatment with JQ-1 for 3 days did not promote but dose dependently inhibited cell proliferation (Supplementary Fig. S3). In line with this, treatment with JQ-1 for 3 days increased the cell population in the G1 phase of the cell cycle (Supplementary Fig. S4a) and slightly increased apoptosis (Supplementary Fig. S4b). These results indicate that the cell-protective effect of JQ-1 in the TSZ-induced necroptosis model was unlikely due to its promoting cell proliferation and/or inhibiting apoptosis. Of note, the considerable inhibitive effect of JQ-1 on cell proliferation at high concentrations may partly explain why the cell survival rate decreased also at high concentrations of JQ-1 (Fig. 1e).
Mechanism of BET inhibitors protecting cells from necroptosis
To explore the mechanism by which BET inhibitors protect cells from necroptosis, we examined the influence of BET inhibitors on the necroptotic signaling and key components of this pathway. Here, JQ-1 was selected as a representative of BET inhibitors. HT29 cells were pretreated with JQ-1 for 3 days and then exposed to TSZ. Co-immunoprecipitation (Co-IP) assays (Fig. 2a) showed that the exposure to TSZ, as expected, resulted in the association of TRADD, RIP1, RIP3, and MLKL, indicating the formation of a necrosome composed of RIP1, RIP3, and MLKL. JQ-1 treatment had no impact on the association among TRADD, RIP1, and RIP3, suggesting no influence on the priming of necroptotic signaling. Besides, JQ-1 treatment did not alter the protein levels of TRADD, RIP1, and RIP3, but obviously reduced the protein level of MLKL (Fig. 2a), implying that the BET inhibitor protects cells from TSZ-induced necroptosis possibly through the downregulation of MLKL. To this end, we further analyzed the MLKL protein levels in HT29 cells treated with different concentrations of JQ-1. As shown in Fig. 2b, the MLKL protein levels decreased upon JQ-1 treatment in a dose-dependent manner, which was inversely correlated with the protective effect against TSZ-induced necroptosis (Fig. 1e). Similar results were also obtained using the other BET inhibitors PFI-1, I-BET-151, I-BET-762, and AZD3514 (Supplementary Figs. S5 and S2a-d). We next examined the influence of the duration of JQ-1 treatment on cell protection and MLKL expression. For this, HT29 cells were pretreated with JQ-1 for 1–4 days, respectively, and then TSZ was added to induce necroptosis. As shown in Supplementary Fig. S6a, the cell-protective effect depended on the duration of JQ-1 treatment, and the 3-day treatment achieved the best protection. The MLKL protein level was inversely correlated with the duration of JQ-1 treatment (Supplementary Fig. S6b). Thus, our results demonstrated that BET inhibitors suppressed TSZ-induced necroptosis by reducing the protein level of MLKL.
BET inhibitors protect cells from necroptosis by suppressing the expression of MLKL. a Co-immunoprecipitation was conducted to detect the interactions among TRADD, RIP1, RIP3, and MLKL. HT29 cells were treated for 3 days with JQ-1 (0.3 µM), followed by a further treatment with TSZ for 6 h. b Protein levels of RIP1, RIP3, and MLKL in HT29 cells following a 3-day treatment with JQ-1 at indicated concentrations. c Cell viability was measured in HT29 cells after JQ-1 pretreatment at different concentrations with/without MG132 (200 nM) for 3 days and TSZ treatment for another 24 h. Columns, mean (n = 3); bars, SD. ***P < 0.001. d Protein level of MLKL was measured in HT29 cells treated with JQ-1 with/without MG132 (200 nM) for 3 days. e Cell viability was measured in HT29 cells after MG132 pretreatment at different concentrations for 3 days and TSZ treatment for another 24 h. f The mRNA level of MLKL in HT29 cells treated with JQ-1 for 24 h was measured by q-PCR. Columns, mean (n = 3); bars, SD. **P < 0.01; ***P < 0.001
To test whether the decreased protein level of MLKL is due to an increased protein degradation, MG132, a broad-spectrum proteasome inhibitor, was utilized. The results showed that the addition of MG132 did not influence HT29 cell survival (Fig. 2c) and MLKL protein level (Fig. 2d) compared with the vehicle (control) treatment. We further assessed the effect of MG132 alone (i.e., without TSZ) on MLKL protein level and HT29 cell survival. The results indicated that the proteasomal inhibitor did not independently influence MLKL expression (Fig. 2d) and HT29 cell survival at the concentrations we used (Fig. 2e). To examine whether the decreased MLKL protein level occurs at the mRNA level, q-PCR assays were performed. As shown in Fig. 2f, JQ-1 treatment resulted in a dose-dependent decrease in the mRNA level of MLKL. As a control, JQ-1 treatment did not obviously alter the mRNA levels of RIP1 and RIP3 (Fig. S6c). Similar results were seen in L929 cells (Supplementary Fig. S6d, e). These results indicate that JQ-1 dose dependently reduced the protein level of MLKL by downregulating the mRNA level of MLKL.
Identification of BRD4 involved in the regulation of MLKL expression
The BET family is composed of BRD2, BRD3, BRD4, and BRDT, and the BET inhibitors identified here are pan-BET inhibitors [20,21,22,23,24]. To pinpoint which BET family member is involved in the regulation of MLKL expression, we conducted gene knockdown assays. In these assays, BRD2, BRD3, and BRD4 were knocked down, respectively, in 293T cell line, and BRDT was excluded because it is a testis-specific bromodomain-containing protein [25]. We found that the knockdown of BRD4 rather than BRD2 or BRD3 decreased MLKL expression (Fig. 3a). Further q-PCR experiments showed that silencing BRD4 resulted in a decreased MLKL mRNA level (Fig. 3b). Similar results were observed in HT29 and HeLa cells (Fig. 3c). Overall, our results demonstrated that BRD4 rather than other BET family members was involved in the regulation of MLKL expression.
BRD4-regulated expression of MLKL. a BRD2, BRD3, or BRD4 shRNA was transfected into 293T cells, and protein levels of BRD2, BRD3, BRD4, and MLKL were detected by western blotting after 48 h. b BRD4 shRNA or control plasmid was transfected into 293T cells, and the mRNA levels of BRD4 and MLKL were measured by q-PCR after 36 h. Columns, mean (n = 3); bars, SD. **P < 0.01; ***P < 0.001. c HT29 and HeLa cells were infected with lentiviral shRNA to BRD4. Protein levels of BRD4 and MLKL were detected by western blotting
Mechanism behind BRD4 regulating MLKL expression
To unveil the mechanism by which BRD4 regulates MLKL expression, we first searched for all the transcription factors that might be involved in the regulation of MLKL gene expression from the SABiosciences’ DECODE database [26]. Then, for each of these transcription factor candidates, we further checked whether it had been reported to have connections with BRD4. We finally found a transcription factor IRF1, which might be involved in the regulation of MLKL gene expression and also have interactions with BRD4. To confirm whether IRF1 participates in BRD4-mediated MLKL expression, we cloned the human MLKL promoter (−1380bp upstream of the transcription starting site) to pGL3 vector and transfected it into 293T, followed by further transfecting the cells with IRF1 or BRD4. Expression of IRF1 or BRD4 alone was sufficient to increase the activity of promoter luciferase. Also, the co-expression of IRF1 and BRD4 further enhanced the luciferase activity (Fig. 4a), suggesting that IRF1 mediates MLKL expression. Moreover, RNA interference was employed. Of interest, knockdown of IRF1 decreased MLKL expression (Fig. 4b). The Co-IP assay also showed the association of BRD4 with IRF1. Furthermore, treatment with trichostatin A (TSA), a pan-HDAC inhibitor, enhanced the BRD4/IRF1 interaction (Fig. 4c), suggesting that BRD4 might bind to the acetylated IRF1, mediating the expression of MLKL.
BRD4 regulates MLKL expression through interacting with IRF1. a Shown is the enhancement of MLKL luciferase activity by the co-expression of IRF1 or/and BRD4 in 293T cells. Columns, mean (n = 3); bars, SD. **P < 0.01; ***P < 0.001. b HT29 and HeLa cells were infected with lentiviral IRF1 shRNA or the control shRNA. The protein level of MLKL was detected by western blotting. c Co-immunoprecipitation was conducted to detect the interactions between BRD4 and IRF1 in HT29 cells treated with/without TSA for 12 h. d 293T cells were co-transfected with FLAG-BRD4 (1-470) or FLAG-BRD4 (471–1362), together with HA-IRF1 by Lipofectamine 2000. After a 48-h transfection, the cells were harvested to conduct immunoprecipitation/immunoblotting with indicated antibodies. e 293T cells were co-transfected with FLAG-BRD4 (1–470) and HA-IRF1 by Lipofectamine 2000. After a 36-h transfection, the cells were treated with/without TSA for 12 h, followed by immunoprecipitation/immunoblotting with indicated antibodies. f 293T cells were co-transfected with FLAG-BD1 or FLAG-BD2, together with HA-IRF1 by Lipofectamine 2000. After a 48-h transfection, the cells were harvested to conduct immunoprecipitation/immunoblotting with indicated antibodies. g Co-immunoprecipitation was conducted to detect the interactions among BRD4, IRF1, RNA-POL II, and P-TEFb with or without JQ-1 treatment in HT29. h ChIP-qPCR was conducted to examine whether BRD4, IRF1, RNA-POL II, and P-TEFb bind to MLKL promoter. Columns, mean (n = 3); bars, SD. ***P < 0.001. i Schematically showing the mechanism by which BRD4 regulates MLKL expression
To determine which region of BRD4 associates with IRF1, we established a Flag-tagged N-terminal BRD4 (NTD, 1–470) construct, containing BD1 and BD2 domains, and a Flag-tagged C-terminal BRD4 (CTD, 471–1362) construct. 293T cells were co-transfected with the Flag-tagged NTD or CTD, together with HA-tagged IRF1. Co-IP assays showed that the NTD, but not the CTD of BRD4, associated with IRF1 (Fig. 4d). The interaction between the NTD and IRF1 was enhanced by TSA treatment (Fig. 4e), indicating again that the NTD associates with the acetylated IRF1. Subsequently, Flag-tagged BD1 (1–250) and BD2 (267–470) constructs were then generated and co-expressed in 293T cells with HA-tagged IRF1. As shown in Fig. 4f, an interaction between BD2 and IRF1 was detected. The results suggest that BRD4 forms a complex with IRF1 via its BD2 region.
It has been established that BRD4 participates in the regulation of gene transcription often through its interaction with the positive transcription elongation factor b (P-TEFb) and RNA polymerase II (RNA-POL II), which occupy the promoter of a gene [27]. We thus deduced that BRD4/IRF1/RNA-POL II/P-TEFb might form a transcriptional complex to regulate the expression of MLKL. To confirm this, co-IP assays were performed. The results did reveal the interactions among BRD4, IRF1, RNA-POL II, and P-TEFb (Fig. 4g). Furthermore, chromatin immunoprecipitation (ChIP)-qPCR assay also showed that the BRD4/IRF1/RNA-POL II/P-TEFb complex occupied the promoter of MLKL gene upstream of the transcription starting site (Fig. 4h). Collectively, all of these clearly demonstrated that BRD4 forms a transcriptional complex with IRF1, RNA-POL II, and P-TEFb, and regulates the expression of MLKL through occupying the promoter of MLKL gene. JQ-1 treatment disrupts the association between BRD4 and IRF1, hence interfering with the formation of the transcription complex of MLKL gene (Fig. 4g, h), which resulted in the downregulated expression of MLKL. The mechanism by which BRD4 regulates the MLKL expression is schematically summarized in Fig. 4i.
Activity of BET inhibitors in MLKL-independent cell death
Given that BRD4 regulates the expression of many genes [20, 22], one may argue that the protective effect of a prolonged treatment with BET inhibitors on inhibiting necroptosis might be a consequence of inducing cell cycle arrest, inhibiting cell proliferation, or inducing apoptosis, which could cause resistance to necroptosis. In this study, we did observe that treatment with JQ-1 for 3 days induced G1 cell cycle arrest, inhibited cell proliferation, and induced apoptosis (Supplementary Figs. S3 and S4). To address whether any of these cellular events contributes to inhibiting necroptosis, HT29 cells were treated for 3 days with a spectrum of small molecular compounds, which are known to arrest cell cycle, block cell proliferation, or induce apoptosis, followed by the necroptosis assay. Our results showed that none of these compounds were able to protect cells from necroptosis (Supplementary Fig. S7), suggesting that the protection against TSZ-induced necroptosis is not attributable to cell cycle arrest, proliferation inhibition, or apoptosis induction.
Since RIP1 not only participates in necroptosis, but also contributes to caspase-8-dependent apoptosis [28], we were curious about whether an inhibition of BRD4 protects against TNFα/Smac-mimetic (TS)-induced apoptosis. The results showed that BET inhibitors did not protect cells from RIP1-dependent apoptosis induced by TS; in contrast, high concentrations of BET inhibitors (>1 µM) led to a decrease in the cell survival rate (Fig. 5a). This might be partly related to the inhibition of cell proliferation by BET inhibitors at high concentrations (Supplementary Fig. S3a, b). In addition, dual staining with Hoechst33342 (for living cells) and Apopxin (for apoptotic cells) proved again that BET inhibitors did not significantly affect TS-induced apoptosis or rescue cell survival (Fig. S8).
BET inhibitors do not protect MLKL-independent cell death. a HT29 cells were pretreated with BET inhibitors at the indicated concentrations for 3 days. Then apoptosis was induced by TS [TNFα (10 ng/ml), Smac-mimetic (100 nM)]. In 24 h, cell viability was determined by CCK8 staining. Columns, mean (n = 3); bars, SD. ***P < 0.001. b HT29 cells were transfected with MLKL shRNA lentiviral particles or control in this experiment. In 48 h, apoptosis was induced by TS [TNFα (10 ng/ml), Smac-mimetic (100 nM)]. Cell viability was determined by CCK8 staining 24 h after TS treatment. Columns, mean (n = 3); bars, SD. ***P < 0.001. RNA interference efficiency was measured by western blotting. c HT1080 cells were pretreated with BET inhibitors for 3 days. Then ferroptosis was induced by erastin (10 μM) for 24 h. Cell viability was determined by CCK8 staining. d HT29 was treated with JQ-1 (0.3 μM) for 48 h. Then, JQ-1-treated HT29 cells were transfected with lentiviral particles of pCDH-MLKL-GFP or pCDH-RFP vector. In 48 h, TNFα/Smac mimetic/Z-VAD-FMK (TSZ) was added to induce necroptosis. Cell viability was measured by CCK8 staining. Columns, mean (n = 3); bars, SD. **P < 0.01; ***P < 0.001. e Protein levels of MLKL in HT29 cells changed following JQ-1 treatment and transfection with lentiviral particles of pCDH-MLKL-GFP or pCDH-RFP vector. Transfection efficiency was shown by GFP or RFP fluorescence. Scale bar, 200 μm
Next, we also wondered whether the downregulation of MLKL by BET inhibitors might predispose cells to RIP1-dependent apoptosis. For this, the TS-induced apoptosis assays were performed on the control and MLKL-silenced (shMLKL) HT29 cells. No difference in the number of apoptotic cells was found in the two assays (Fig. 5b), which is consistent with the other reports [29, 30]. All of these indicate that the downregulation of MLKL by BET inhibitors does not predispose cells to RIP1-dependent apoptosis. Furthermore, we tested whether BET inhibitors protect cells from MLKL-independent cell death. To this end, we studied the effect of JQ-1 on ferroptosis, another type of programmed cell death, which is MLKL independent [31, 32]. The results showed that JQ-1 had no impact on erastin-induced ferroptosis (Fig. 5c). This experiment, together with the above assay of TS-induced apoptosis, which is also MLKL-independent cell death, demonstrated that BET inhibitors could not protect cells from MLKL-independent cell death.
To substantiate the role of MLKL in the BET inhibitors’ protection against necroptosis, HT29 cells were treated with JQ-1 for 48 h and then transfected with lentiviral particles of pCDH-MLKL-GFP or pCDH-RFP vector. After transfection for 48 h, TSZ was added to induce necroptosis. As shown in Fig. 5d, the effect of JQ1 on inhibiting necroptosis could be totally bypassed in cells transfected with MLKL. Transfection efficiency was shown in Fig. 5e. The result further supports that BET inhibitors protect cells from TSZ-induced necroptosis by downregulating MLKL expression.
Evaluation of BET inhibitors in the treatment of necroptosis-related diseases
Because of the role of BRD4 in the regulation of necroptosis, we further explored the possible application of BET inhibitors in the treatment of necroptosis-related diseases. Here an experimental disease model was used: TNFα-driven shock model.
The TNFα-driven shock is a model of systemic inflammatory response syndrome (SIRS) and is a widely used necroptosis-related disease model [33, 34]. To examine the efficacy of BET inhibitors in this model, JQ-1 (20 mg/kg/d) or vehicle (control) was first intraperitoneally given to wild-type C57BL/6 mice, and three days later murine TNFα at a dose of 500 μg/kg was injected to the mice via the tail vein. The results showed that TNFα induced severe hypothermia and casualty in the control group (Fig. 6a). Pathological examination revealed severe liver, lung, and kidney injuries in the control group as well, which are consistent with the previous findings [33, 34]. Necrotic regions were detected in livers, and inflammatory lesions were detected in lungs and kidneys in the model group (Fig. 6b). Interestingly, JQ-1 treatment evidently ameliorated hypothermia and organic injuries, and increased the survival rate, compared with the vehicle treatment (Fig. 6a), indicating the therapeutic efficacy of JQ-1 in the SIRS model. Furthermore, immunohistochemical analysis showed that intraperitoneal injection of JQ-1 reduced MLKL expression in the lung and liver tissues (Supplementary Fig. S9).
JQ-1 ameliorates the systemic inflammatory response syndrome induced by TNFα. a Effects of JQ-1 on TNFα-induced shock-associated hypothermia and survival curve. b Representative images of HE staining of liver, lung, spleen, kidney, heart tissue sections; scale bar, 50 μm. Organs were harvested 6 h after the injection, then fixed in formalin, embedded in paraffin, and sliced
To further examine the role of MLKL in this model, the same experiment was carried out using MLKL knockout (KO) (MLKL-/-) mice. We found that TNFα led to an obviously ameliorated syndrome in the body temperature, casualty, and organic injury in MLKL-/- mice vs the wild-type counterpart. The degree of amelioration in MLKL-/- mice was slightly better than that of JQ-1 treatment in wild-type mice, which is understandable because JQ-1 could not entirely abrogate MLKL expression as the gene knockout did. Finally, the treatment with JQ-1 did not further enhance the ameliorative effect of MLKL-/- (Fig. 6a), confirming that JQ-1 executes its protection against necroptosis through downregulating MLKL expression.
Discussion
In this investigation, cell-based small molecule screening and subsequent mechanistic studies led to the finding that BRD4 is a new epigenetic regulator of necroptosis. We found that BRD4 interacted with acetylated IRF1, together with P-TEFb and RNA-POL II, to comprise a transcription complex. The formed complex BRD4/IRF1/P-TEFb/RNA-POL II occupied the promoter region of MLKL gene to regulate MLKL expression. BRD4 inhibitors interfered with the formation of the transcription complex of MLKL gene, leading to the downregulation of MLKL expression, thereby protecting cells from necroptosis.
BRDs are important epigenetic “reader” proteins that specifically recognize acetylated-lysine residues in nucleosomal histones, facilitating the recruitment of transcriptional proteins to chromatin [23]. Also, BRD4 is known to control the expression of genes critical for cell growth and contribute to the epigenetic memory of postmitotic cells by shaping transcriptional regulation across cell division [35]. Importantly, it has been demonstrated that BRD4 is required for the expression of c-Myc and other "tumor driving" oncogenes in hematologic cancers including multiple myeloma, acute myelogenous leukemia, and acute lymphoblastic leukemia [22, 23, 36, 37]. Therefore, BRD4 is thought as a promising target for cancer therapy. A number of BRD4 inhibitors have been developed, and some of them have entered clinical trials for treating various cancer types [38, 39]. Our finding in this study further expands the scope of application of BRD4 inhibitors to necroptosis-related diseases. Indeed, we have shown here that, in a necroptosis-related disease model, namely the TNFα-driven SIRS model, the BRD4 inhibitor JQ-1 displayed promising therapeutic effects.
BRD4 inhibitors have been studied in animal models for the treatment of autoimmune or inflammatory diseases, such as multiple sclerosis [40] and liver fibrosis [41]. Various mechanisms were suggested to be responsible for the therapeutic effect of BRD4 inhibitors. For example, in an experimental autoimmune encephalomyelitis mouse model, the BRD4 inhibitor JQ-1 was reported to be able to ameliorate experimental autoimmune encephalomyelitis severity by controlling TH17 differentiation [40]. In a CCl4-induced liver fibrosis mouse model, the BRD4 inhibitor JQ-1 showed a good therapeutic effect, which was found through a mechanism of abrogating cytokine-induced activation of hepatic stellate cells [41]. None of these studies addressed a possible mechanism linking BRD4 to necroptosis. Considering multiple sclerosis and liver fibrosis have been reported to be necroptosis-related diseases [42, 43], we postulate that inhibition of MLKL expression may also partly contribute to the therapeutic effects of BRD4 inhibitors in these models.
In conclusion, we, for the first time, report that the bromodomain protein BRD4 positively regulates necroptosis. Mechanistic studies indicate that BRD4 regulates necroptosis through modulating the expression of the necroptosis executor MLKL. This is a completely new epigenetic regulation mechanism of necroptosis. In addition, our work also firstly highlights the potential of BET inhibitors in the treatment of necroptosis-related diseases.
Materials and methods
Compounds
Compounds for screening were taken from an in-house targeted small molecular library (T-Lib) containing 3600 compounds with their on-targets being known. All compounds used in this study were of analytical purity.
Biochemical kinase binding assays
In vitro KINOMEscan kinase binding assays (DiscoverX) were employed to determine the binding affinity (Kd) of compounds.
Cell lines and cell culture conditions
HT29 and mouse L-cells NCTC 929 (L929) were obtained from the National Infrastructure of Cell Line Resource (China). U937 and 293T were obtained from American Type Culture Collection (ATCC). All cellular incubations were performed at 37 °C and 5% CO2 unless stated otherwise.
The EdU assay
The proliferation cells were assayed with a Fluro488-EdU DNA proliferation detection kit (KGA331; KeyGENEBioTECH) according to the manufacturer’s instructions.
Cell cycle and apoptosis assays
Flow cytometry was used for cell cycle and apoptosis assays. Cell Cycle Analysis Kit (KGA511; KeyGENEBioTECH) was used for cell cycle analysis and the Annexin V-FITC & PI Cell Apoptosis Analysis Kit (KGA107; KeyGENEBioTECH) was used for apoptosis assays. The data were analyzed using FLOWJO software and MODFIT software.
Co-immunoprecipitation and western blotting
Cells were directly lysed and incubated on ice for 15 min and then centrifuged. The supernatants were determined using the BCA Protein Assay Kit (P0010; Beyotime). Some supernatants were incubated with a certain antibody in a rotating way overnight at 4 °C, the rest were loaded with SDS-PAGE Sample Loading Buffer (5 × , P0015; Beyotime) and used as input. Overnight, Protein A (or G) Agarose Beads (11719408001; Roche) was added, and incubation was conducted for another 4–6 h. The antibody–protein bead complex was obtained and washed by centrifugation, followed by boiling with loading buffer and centrifugation. The supernatants were separated on 8–12% acrylamide gels by sodium dodecyl sulphate-polyacrylamide gel electrophoresis. Proteins were transferred to polyvinylidene difluoride membranes and analyzed by probing with anti-RIP1 antibody (1:1000, 610458; R&D), anti-RIP3 antibody (1:200, sc-135170; Santa Cruz), anti-TRADD antibody (1:500, sc-135170; Abcam), anti-MLKL antibody (1:1000, ab194699; Abcam), anti-P-MLKL(h) antibody (1:1000, ab187091; Abcam), anti-P-MLKL(m) antibody (1:500, ab196436; Abcam), anti-Ub antibody (1:200, sc-8017; Santa Cruz), anti-BRD4 antibody (1:1000, A301–985A50; Bethyl), anti-IRF1 antibody (1:500, ab26109; ABcam), anti-RNA-Polymerse II antibody (1:1000, 05–623; Millipore), anti-P-TEFb antibody (1:200, CDK9, sc-13130; Santa Cruz), anti-FLAG antibody (1:250, F1804; Sigma), and anti-HA antibody (1:1000, 66006-1-lg; Peprotech). Proteins were visualized by chemiluminescence.
Real-time q-PCR analysis
Total RNA in cells was extracted using RNAsimple total RNA kit (DP405; Tiangen) according to the manufacturer’s instructions. cDNA was generated using the iScriptTMcDNA Synthesis Kit (170–8891; Bio-Rad), and the RT-PCRs were performed using SsoAdvanced SYBR Green Supermix (172–5271; Bio-Rad).
Invitrogen Block-iT Pol II miR RNAi strategy
Oligonucleotides targeting human BRD2, BRD3, and BRD4 sequences compatible for use in cloning into Block-iT Pol II miR RNAi expression vectors were obtained using the online tool Block-iT RNAi Designer. The pcDNA 6.2-GW/EmGFP-miR-neg control plasmid contained an insert that can form a hairpin structure that is processed into mature miRNA, but was predicted not to target any known vertebrate gene.
Lentiviral shRNAs
BRD4 shRNA sequence used was 5′-AGCAAAGCCAAGGAACCTC-3′. The sequence was cloned into hU6-MCS-Ubiquitin-EGFP-IRES-puromycin vector. The BRD4 shRNA lentiviral particles were produced. IRF1 shRNA lentiviral particles (sc-35706-v) and MLKL shRNA lentiviral particles (sc-93430-v) were purchased from Santa Cruz.
cDNA constructs
The coding sequences for different regions of BRD4 (human) were cloned into pcDNA3.1 vector between the HindIII and XhoI sites. The coding sequence for a full-length IRF1 (human) was cloned into pcDNA3.1 vector between the HindIII and BamHI sites.
Chromatin immunoprecipitation
ChIP assay was performed using the EZ-Magna ChIPTM A/G Enzymatic Chromatin IP Kit (17–10086; Millipore), according to the manufacturer’s instructions.
Luciferase assay
Luciferase activity was measured with Luciferase assay system (E2920; Promega) according to the manufacturer’s instructions.
Mice
Parent MLKL-/- mice were kindly donated by Jiahuan Han’ lab (Xiamen University, China) and crossed with wild-type C57BL/6 mice to generate MLKL+/- mice. Then wild-type and MLKL-/- mice from the same background were obtained after MLKL+/- mice breeding.
TNFα-induced systemic inflammatory response syndrome model
JQ-1 (20 mg/kg/d) or vehicle (control) was first intraperitoneally given to wild-type C57BL/6 mice. Three days later, recombinant murine TNFα (AF-315–01A; PeproTech) was injected, at a dose of 500 μg/kg, to the mice via the tail vein. Body temperature was detected by an infrared thermometer. Partial mice were sacrificed 6 h after the injection, and their organs were collected, fixed in formalin, embedded in paraffin, and sectioned for histology. The rest of the mice were observed for mortality.
Hematoxylin and eosin staining
Mice were sacrificed and their organs were harvested 6 h after TNFα injection and then fixed in 10% buffered formalin for 24 h. The fixed tissues were then embedded into paraffin and sectioned at 5µm thickness with a microtome. Deparaffinized, rehydrated sections were stained with hematoxylin for 5 min, and then differentiated in 0.1% acid alcohol followed by bluing and counterstaining in 0.5% eosin Y solution and the final dehydration. The representative images were taken on a LEICA DM 2500 microscope with a LEICA DFC425 C camera. Image acquisition software used was LAS V3.7.
Immunohistochemical analysis
Paraffin-embedded organs were subjected to immunostaining with MLKL (66675-1-Ig; ProteinTech). The representative images were taken on a LEICA DM 2500 microscope with a LEICA DFC425 C camera. Image acquisition software used was LAS V3.7.
Animal care
All animals experiments were performed according to the guidelines of Institutional Animal Care and Use Committee of Sichuan University (Chengdu, Sichuan, China) and protocols were approved by the Institutional Animal Care and Use Committee of Sichuan University (Chengdu, Sichuan, China).
Statistical analysis
Data were expressed as mean ± standard deviation or mean ± standard error of the mean. For data analyses, one-way or two-way ANOVA test was performed using SPSS 16.0. In all comparisons, P < 0.05 was considered statistically significant.
References
Weinlich R, Oberst A, Beere HM.Green DR, Necroptosis in development, inflammation and disease. Nat Rev Mol Cell Biol. 2017;18:127–36.
Kaczmarek A, Vandenabeele P, Krysko DV. Necroptosis: the release of damage-associated molecular patterns and its physiological relevance. Immunity. 2013;38:209–23.
Berghe TV, Linkermann A, Jouan-Lanhouet S, Walczak H, Vandenabeele P. Regulated necrosis: the expanding network of non-apoptotic cell death pathways. Nat Rev Mol Cell Biol. 2014;15:135–47.
Ofengeim D, Yuan J. Regulation of RIP1 kinase signaling at the crossroads of inflammation and cell death. Nat Rev Mol Cell Biol. 2013;14:727–36.
Zhang D, Shao J, Lin J, Zhang N, Lu B, Lin S, et al. RIP3, an energy metabolism regulator that switches TNF-induced cell death from apoptosis to necrosis. Science. 2009;325:334–6.
Sun L, Wang H, Wang Z, He S, Chen S, Liao D, et al. Mixed lineage kinase domain-like protein mediates necrosis signaling downstream of RIP3 kinase. Cell. 2012;148:213–27.
Wang L, Du F, Wang X. TNF-α induces two distinct caspase-8 activation pathways. Cell. 2008;133:693–703.
Cho YS, Challa S, Moquin D, Genga R, Ray TD, Guildford M, et al. Phosphorylation-driven assembly of the RIP1-RIP3 complex regulates programmed necrosis and virus-induced inflammation. Cell. 2009;137:1112–23.
He S, Wang L, Miao L, Wang T, Du F, Zhao L, et al. Receptor interacting protein kinase-3 determines cellular necrotic response to TNF-alpha. Cell. 2009;137:1100–11.
Wang H, Sun L, Su J, Rizo J, Liu L, Wang L, et al. Mixed lineage kinase domain-like protein MLKL causes necrotic membrane disruption upon phosphorylation by RIP3. Mol Cell. 2014;54:133–46.
Chen X, Li W, Ren J, Huang D, He W, Song Y, et al. Translocation of mixed lineage kinase domain-like protein to plasma membrane leads to necrotic cell death. Cell Res. 2014;7:105–21.
Li D, Xu T, Cao Y, Wang H, Li L, Chen S, et al. A cytosolic heat shock protein 90 and cochaperone CDC37 complex is required for RIP3 activation during necroptosis. Proc Natl Acad Sci USA. 2015;112:5017–22.
Chen W, Wu J, Li L, Zhang Z, Ren J, Liang Y, et al. Ppm1b negatively regulates necroptosis through dephosphorylating Rip3. Nat Cell Biol. 2015;17:434–44.
Seo J, Lee EW, Sung H, Seong D, Dondelinger Y, Shin J, et al. CHIP controls necroptosis through ubiquitylation- and lysosome-dependent degradation of RIPK3. Nat Cell Biol. 2016;18:291–302.
Degterev A, Huang Z, Boyce M, Li Y, Jagtap P, Mizushima N, et al. Chemical inhibitor of nonapoptotic cell death with therapeutic potential for ischemic brain injury. Nat Chem Biol. 2005;1:112–9.
Fauster A, Rebsamen M, Huber KVM, Bigenzahn JW, Stukalov A, Lardeau CH, et al. A cellular screen identifies ponatinib and pazopanib as inhibitors of necroptosis. Cell Death Dis. 2015;6:e1767.
Li J, Feng M, Wang Y, Li X, Chen X, Su Y, et al. The B-RafV600E inhibitor dabrafenib selectively inhibits RIP3 and alleviates acetaminophen-induced liver injury. Cell Death Dis. 2014;5:e1278.
Najjar M, Suebsuwong C, Ray SS, Thapa RJ, Maki JL, Nogusa S, et al. Structure guided design of potent and selective ponatinib-based hybrid inhibitors for RIPK1. Cell Rep. 2015;10:1850–60.
Diego RR, Delvaeye T, Roelandt R, Nerinckx W, Augustyns K, Vandenabeele P, et al. When PERK inhibitors turn out to be new potent RIPK1 inhibitors: critical issues on the specificity and use of GSK2606414 and GSK2656157. Cell Death Differ. 2017;24:1100–10.
Filippakopoulos P, Qi J, Picaud S, Shen Y, Smith WB, Fedorov O, et al. Selective inhibition of BET bromodomains. Nature. 2010;468:1067–73.
Picaud S, Costa DD, Thanasopoulou A, Filippakopoulos P, Fish PV, Philpott M, et al. PFI-1, a highly selective protein interaction inhibitor, targeting BET bromodomains. Cancer Res. 2013;73:3336–46.
Dawson MA, Prinjha RK, Dittmann A, Giotopoulos G, Bantscheff M, Chan WI, et al. Inhibition of BET recruitment to chromatin as an effective treatment for MLL-fusion leukaemia. Nature. 2011;478:529–33.
Zhao Y, Yang C, Wang S. The making of I-BET762, a BET bromodomain inhibitor now in clinical development. J Med Chem. 2013;56:7498–7500.
Waring MJ, Chen H, Rabow AA, Walker G, Bobby R, Boiko S, et al. Potent and selective bivalent inhibitors of BET bromodomains. Nat Chem Biol. 2016;12:1097–104.
Pivot-Pajot C, Caron C, Govin J, Vion A, Rousseaux S, Khochbin S. Acetylation-dependent chromatin reorganization by BRDT, a testis-specific bromodomain-containing protein. Mol Cell Biol. 2003;23:5354–65.
Nakanome A, Brydun A, Matsumoto M, Ota K, Funayama R, Nakayama K, et al. Bach1 is critical for the transformation of mouse embryonic fibroblasts by RasV12 and maintains ERK signaling. Oncogene. 2013;32:3231–45.
Yang Z, Yik JHN, Chen R, He N, Jang MK, Ozato K, et al. Recruitment of P-TEFb for stimulation of transcriptional elongation by the bromodomain protein BRD4. Mol Cell. 2005;19:535–45.
Feokistova M, Geserick P, Kellert B, Dimitrova DP, Langlais C, Hupe M, et al. cIAPs block ripoptosome formation, a RIP1/Caspase-8 containing intracellular cell death complex differentially regulated by cFLIP isoforms. Mol Cell. 2011;43:323–5.
Zhao J, Jitkaew S, Cai Z, Choksi S, Li Q, Luo J, et al. Mixed lineage kinase domain-like is a key receptor interacting protein 3 downstream component of TNF-induced necrosis. Proc Natl Acad Sci USA. 2012;109:5322–7.
Wu J, Huang Z, Ren J, Zhang Z, He P, Li Y, et al. Mlkl knockout mice demonstrate the indispensable role of Mlkl in necroptosis. Cell Res. 2013;23:994–1006.
Dixon SJ, Lemberg KM, Lamprecht MR, Skouta R, Zaitsev EM, Gleason CE, et al. Ferroptosis: an iron-dependent form of nonapoptotic cell death. Cell. 2012;149:1060–72.
Dixon SJ, Stockwell BR. The role of iron and reactive oxygen species in cell death. Nat Chem Biol. 2014;10:9–17.
Duprez L, Takahashi N, Hauwermeiren FV, Vandendriessche B, Goossens V, Berghe TV, et al. RIP kinase-dependent necrosis drives lethal systemic inflammatory response syndrome. Immunity. 2011;35:908–18.
Newton K, Dugger DL, Maltzman A, Greve JM, Hedehus M, Martin-McNulty B, et al. RIPK3 deficiency or catalytically inactive RIPK1 provides greater benefit than MLKL deficiency in mouse models of inflammation and tissue injury. Cell Death Differ. 2016;23:1565–76.
Devaiah BN, Singer DS. Two faces of BRD4: mitotic bookmark and transcriptional lynchpin. Transcription. 2013;4:13–17.
Zuber J, Shi J, Wang E, Rappaport AR, Herrmann H, Sison EA, et al. RNAi screen identifies BRD4 as a therapeutic target in acute myeloid leukaemia. Nature. 2011;478:524–8.
Delmore JE, Issa GC, Lemieux ME, Rahl PN, Shi J, Jacobs HM, et al. BET bromodomain inhibition as a therapeutic strategy to target c-Myc. Cell. 2011;146:904–17.
Berthon C, Raffoux E, Thomas X, Vey N, Gomez-Roca C, Yee K, et al. Bromodomain inhibitor OTX015 in patients with acute leukaemia: a dose-escalation, phase 1 study. Lancet Haematol. 2016;3:e186–195.
Amorim S, Stathis A, Gleeson M, Lyengar S, Magarotto V, Leleu X, et al. Bromodomain inhibitor OTX015 in patients with lymphoma or multiple myeloma: a dose-escalation, open-label, pharmacokinetic, phase 1 study. Lancet Haematol. 2016;3:196–204.
Mele DA, Salmeron A, Ghosh S, Huang HR, Bryant BM, Lora JM. BET bromodomain inhibition suppresses TH17-mediated pathology. J Exp Med. 2013;210:2181–90.
Ding N, Hah N, Yu RT, Sherman MH, Benner C, Leblanc M, et al. BRD4 is a novel therapeutic target for liver fibrosis. Proc Natl Acad Sci USA. 2015;112:15713–8.
Ofengeim D, Ito Y, Najafov A, Zhang Y, Shan B, DeWitt JP, et al. (2015) Activation of necroptosis in multiple sclerosis. Cell Rep. 2015;10:1836–49.
Choi HS, Kang JW, Lee SM. Melatonin attenuates carbon tetrachloride-induced liver fibrosis via inhibition of necroptosis. Transl Res. 2015;166:292–303.
Acknowledgements
We are thankful to Dr. Jiahuan Han (Xiamen University, China) for generously providing MLKL-/- mice for this research. This work was supported by the National Natural Science Foundation of China (81473140, 81573349, 81773633, and 21772130), National Science and Technology Major Project (2018ZX09711002-014-002, 2018ZX09711002-011-019, and 2018ZX09711003-003-006), and 1.3.5 project for disciplines of excellence, West China Hospital, Sichuan University.
Author contributions
S.Y. and Y.W. conceived the project and designed the research. Y.X., L.L., L. Z., Y.C., and C.W. performed the experiments and analyzed the data. Q.Y., K.C., H.L., R.X., and Y.H. analyzed the data. S.Y., S.H., and Y.W. wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Xiong, Y., Li, L., Zhang, L. et al. The bromodomain protein BRD4 positively regulates necroptosis via modulating MLKL expression. Cell Death Differ 26, 1929–1941 (2019). https://doi.org/10.1038/s41418-018-0262-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-018-0262-9
This article is cited by
-
Regulation of programmed cell death by Brd4
Cell Death & Disease (2022)
-
Non-classical ferroptosis inhibition by a small molecule targeting PHB2
Nature Communications (2022)
-
Deficiency of PPP6C protects TNF-induced necroptosis through activation of TAK1
Cell Death & Disease (2022)
-
MLKL in cancer: more than a necroptosis regulator
Cell Death & Differentiation (2021)
-
The BET inhibitor attenuates the inflammatory response and cell migration in human microglial HMC3 cell line
Scientific Reports (2021)