Abstract
The tumor suppressor DAB2IP contributes to modulate the network of information established between cancer cells and tumor microenvironment. Epigenetic and post-transcriptional inactivation of this protein is commonly observed in multiple human malignancies, and can potentially favor progression of tumors driven by a variety of genetic mutations. Performing a high-throughput screening of a large collection of human microRNA mimics, we identified miR-149-3p as a negative post-transcriptional modulator of DAB2IP. By efficiently downregulating DAB2IP, this miRNA enhances cancer cell motility and invasiveness, facilitating activation of NF-kB signaling and promoting expression of pro-inflammatory and pro-angiogenic factors. In addition, we found that miR-149-3p secreted by prostate cancer cells induces DAB2IP downregulation in recipient vascular endothelial cells, stimulating their proliferation and motility, thus potentially remodeling the tumor microenvironment. Finally, we found that inhibition of endogenous miR-149-3p restores DAB2IP activity and efficiently reduces tumor growth and dissemination of malignant cells. These observations suggest that miR-149-3p can promote cancer progression via coordinated inhibition of DAB2IP in tumor cells and in stromal cells.
Similar content being viewed by others
Introduction
The tumor suppressor Disabled-2 Interacting Protein (DAB2IP), also known as ASK1 Interacting Protein (AIP1), is emerging as a pleiotropic modulator of cytoplasmic signaling. Through an interaction with signaling modulators and potential oncogenes, DAB2IP controls multiple pathways, including Ras/MAPK [1], apoptosis signal-regulating kinase 1 (ASK1)/c-Jun N-terminal kinase (JNK)/p38 MAPK [2], NF-κB [1], and phosphoinositide 3-kinase (PI3K)/AKT signaling [3], affecting cell behavior in response to a variety of extrinsic inputs. The loss-of-function of this modulator can therefore amplify the cancer cell’s response to multiple oncogenic signals, fostering tumor progression [4, 5].
A consistent body of literature reports epigenetic inactivation of DAB2IP by promoter methylation in malignant cells [6, 7]. Recently, other molecular mechanisms of DAB2IP inactivation in cancer have been identified. Multiple E3 ubiquitin-ligases can promote its turnover, while AKT kinase-mediated phosphorylation counteracts DAB2IP functions [4, 5]. In addition, cytoplasmic interaction with tumor-associated p53 mutants blocks DAB2IP inhibitory activity on NF-κB and on AKT, favoring respectively cancer cell dissemination in response to inflammatory stimuli [8], and insulin-induced proliferation and invasion of tumor cells [9]. Finally, due to the rather long 3′UTR sequence of its main transcripts, DAB2IP is a strong candidate for post-transcriptional regulation by microRNAs (miRNAs). Accordingly, some miRNAs have already been reported to target DAB2IP in various tumor cell models [10,11,12,13,14,15].
MicroRNAs are 20–22 nt small RNAs that control stability and translation of target mRNAs, and are involved in the regulation of virtually all biological processes [16]. In tumors, miRNAs impact every step of carcinogenesis, including metastasis formation and crosstalk with the tumor microenvironment [17]. Because of their ability to finely modulate key disease factors, miRNAs are also suitable candidates for tumor diagnosis as well as therapy [18, 19].
In the present study, we report the large-scale identification of miRNAs able to target and downregulate DAB2IP in tumor cells. We focus on miR-149-3p, and unveil its ability to sustain cell migration and tumor dissemination through cell-autonomous and cell non-autonomous targeting of DAB2IP.
Results
Identification of novel miRNAs targeting DAB2IP
Using an automated high-throughput screening approach, we tested a collection of 879 human miRNA mimics for their effect on a dual-Luciferase DAB2IP 3′UTR reporter in HeLa cells (see Materials and Methods for details). Empty (no miRNA), cel-miR-231, and cel-miR-67 were used as negative controls; hsa-miR-338-3p [13] and an siRNA targeting DAB2IP 3′UTR were used as positive controls. After two rounds of primary screening, and a secondary screen to expunge false positives, we obtained a list of 40 miRNAs able to downregulate the reporter (Fig. 1a); among them were miR-338-3p and miR-92b, already reported to target DAB2IP [12, 13], thus confirming the reliability of the assay. Interestingly, this approach identified miRNAs that are not detected by prediction algorithms, or have a very low probability score (Figure S1A).
Identification of microRNAs targeting DAB2IP by high-throughput functional screening. a The graph summarizes the list of top-scoring miRNAs identified in the screening. HeLa cells were reverse-transfected with miRNA mimics; after 24 h, cells were transfected with psiCHECK2 DAB2IP 3′UTR vector (primary screening) or with psiCHECK2 empty vector (secondary screening). Dual-Luciferase assay was performed 48 h later. The graph summarizes the results of two rounds of primary screen; false positives obtained with the secondary screen were expunged. The X axis reports Renilla Luciferase/Firefly Luciferase ratio for each miRNA, normalized to the average of negative controls. Empty (no miRNAs), cel-miR-231, and cel-miR-67 were used as negative controls, while miR-338-3p mimic and a DAB2IP siRNA targeting the 3′UTR were used as positive controls. miRNAs highlighted in red were considered for subsequent validation. b, c Identified miRNAs are able to reduce endogenous DAB2IP protein levels. b A representative western blot of PC3 and LNCaP cells transfected with indicated miRNA mimics for 48 h. DAB2IP siRNA (siDAB2IP) was used as a positive control. Actin was blotted as a loading control. c Heat map summarizing the effects of selected miRNA mimics on endogenous DAB2IP protein levels in the indicated cell lines. Cells were treated as in b. Protein bands were quantified by densitometry on autoradiography films (shown in Fig. 1b and Figure S1B) and normalized to siCtrl-transfected samples
Having a substantial number of hits, we focused on miRNAs with the highest efficiency of reporter inhibition; plus, we cherry-picked a few cancer-related miRNAs [20,21,22]. We screened selected hits by their effect on endogenous DAB2IP in multiple cancer cell lines, that differ in tissue origin, aggressiveness, and DAB2IP protein levels. Under these conditions, miR-149-3p, miR-555, miR-582-5p, and miR-92a induced strong reduction of DAB2IP in at least four out of seven cell lines (Fig. 1b, c and Figure S1B).
MiR-555 and miR-582-5p were excluded from further analysis because they are not detectable in our model cell lines (data not shown). Of the remaining hits, miR-92a belongs to the miR-17-92 cluster and its oncogenic properties are well established [20]; in contrast, the role of miR-149-3p in cancer is controversial [23,24,25,26]. Moreover, our screening also identified miR-149-5p (Fig. 1a, b), that results from processing of the same precursor as miR-149-3p, but has a different seed sequence (Figure S1C). Both miRNAs can be incorporated into RNA-Induced Silencing Complex (RISC) [27], and they share some common targets [27, 28]. We decided to focus on miR-149-3p and miR-149-5p in order to examine their biological functions related to DAB2IP inhibition.
miR-149-3p expression is increased in human malignancies
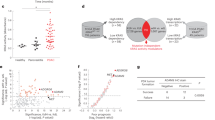
To explore the potential relevance of miR-149 in cancer, we analyzed miRNA expression data from The Cancer Genome Atlas (TCGA) consortium (Table S1). Among all available datasets, miR-149-3p is differentially expressed in bladder (BLCA), breast (BRCA), lung (LUSC), endometrial (UCEC), and prostate (PRAD) cancers. In the same datasets, DAB2IP mRNA displays an opposite trend (Fig. 2a).
A more focused analysis, on tumors matched with normal tissue from the same patient, revealed upregulation of miR-149-3p in bladder, breast, and lung cancers (Fig. 2b). In particular, scatter plots efficiently show that expression levels are truly different in normal vs. tumor samples.
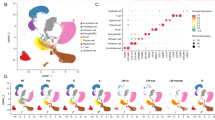
miR-149-3p expression in human cancers. a Differential expression analysis of miR-149-3p and DAB2IP in tumors compared to normal tissue across multiple TCGA datasets: Bladder Carcinoma (BLCA), Breast Carcinoma (BRCA), Lung Squamous Cell Carcinoma (LUSC), Uterine Corpus Endometrial Carcinoma (UCEC), and Prostate Adenocarcinoma (PRAD). Boxplots display levels of miR-149-3p calculated as log10 of its Counts Per Million (CPM), and levels of DAB2IP calculated as log10 of its Fragments per Kilobase of Transcript per Million (FPKM). b Paired differential expression analysis of miR-149-3p in tumor samples compared to matched normal tissue samples in TCGA Bladder Carcinoma (BLCA), Breast Carcinoma (BRCA), and Lung Squamous Cell Carcinoma (LUSC). Scatter plots display expression levels of miR-149-3p for paired tumor vs. matched-normal samples (log10 CPM). c Differential expression analysis of DAB2IP, miR-149-3p, and miR-149-5p in the GSE21032 prostate cancer dataset [30]. The table reports false discovery rate (q-values) after correction for multiple testing using Benjamini–Hochberg method
Conversely, the expression pattern of miR-149-5p is more variable and not always matches the expression of miR-149-3p. For instance, in prostate cancer (PRAD), differential expression is observed only for miR-149-3p (Table S1).
Since DAB2IP is a powerful inhibitor of cell migration and metastasis, especially in prostate cancer [29], we analyzed the GSE21032 prostate cancer study, which includes metastasis samples [30]. Notably, in this dataset miR-149-3p, but not miR-149-5p, shows augmented expression in primary tumors, with highest levels in metastasis (Fig. 2c). Concomitantly, DAB2IP expression is reduced both in tumors and metastases.
miR-149-3p increases cancer cell invasion
For functional studies, we first verified that miR-149-3p is expressed in our prostate and other cancer cell lines (Figure S2A). We next verified that transfection of a moderate amount of a miR-149-3p mimic (to reduce potential off-target effects) reproducibly induced a specific increase in mature cellular miR-149-3p levels, with corresponding downregulation of DAB2IP mRNA and protein (Figure S2B). We then transfected the miR-149-3p mimic or a miR-149-3p inhibitor in the metastatic androgen-independent prostate cancer cell line PC3, and performed transwell matrigel invasion assays. As shown in Fig. 3a, miR-149-3p overexpression increased the invasive properties of PC3 cells with similar, if not better, efficiency as DAB2IP depletion by siRNA.
Inhibition of endogenous miR-149-3p reduced the invasive capability of PC3, simultaneously increasing DAB2IP protein levels (Fig. 3a). Similar results were obtained in the androgen-sensitive prostate cancer cell line LNCaP (Fig. 3b). Importantly, results were recapitulated in breast and ovarian cancer cell lines, indicating that this effect is not prostate specific (Figure S2C).
miR-149-3p drives cell invasion, reducing DAB2IP protein. a, b miR-149-3p sustains prostate cancer cells invasion. PC3 (a) and LNCaP (b) cells were transfected with a control siRNA (siCtrl), DAB2IP siRNAs mix (siDAB2IP), miRNA mimic (miR-149-3p), a control hairpin inhibitor (inh Ctrl), or an inhibitor for miR-149-3p (inh miR-149-3p). After 48 h, invasion assays were performed. Cells were plated under low serum conditions; the lower chamber was filled with high serum medium. Graphs summarize migrated cells per area (mean ± SD; n = 3; ***P < 0.001; **P < 0.01; *P < 0.05). Western blots confirm increase or reduction of DAB2IP protein levels in transfected cells. Actin was blotted as a loading control. Representative images of migrated cells are also shown. Scale bar = 100 μm. c miR-149-3p functional depletion counteracts prostate cancer cells invasion. PC3 cells stably transduced with a lentivirus expressing a miR-149-3p sponge (TW3 S149-3p) were tested in invasion assays as in a (mean ± SD; n = 4; ***P < 0.001). Western blot for DAB2IP and GFP proteins confirmed the functionality of the sponge. Actin was blotted as a loading control. d miR-149-3p, but not miR-149-5p, promotes the invasion of non-transformed prostate epithelial cells. PWR-1E and RWPE-1 were transfected as indicated for 48 h. Invasion assays were performed as in a. Graph summarizes migrated cells per area (mean ± SD; n = 3; ***P 0.001; **P < 0.01; *P < 0.05; ns = not significant). DAB2IP levels were checked by western blot
For stable miR-149-3p inhibition, we cloned a specific antisense sponge construct (S149-3p) fused to the 3′ end of a green fluorescence protein (eGFP) expression cassette in the lentiviral vector Tween 3′UTR (TW3). RT-qPCR and western blot established the efficacy of the sponge in reducing miR-149-3p (and not miR-149-5p) levels, and augmenting DAB2IP protein (Figure S2D). PC3 cells expressing the sponge (TW3 S149-3p) showed marked reduction in matrigel invasion (Fig. 3c), confirming that inhibition of miR-149-3p reduces the invasive potential of these cells in tissue culture.
We also analyzed the effects of miR-149-5p in PC3 cells; interestingly, under conditions in which miR-149-3p strongly sustained invasion, no effects were observed upon miR-149-5p overexpression (Figure S2E). This result correlates with less efficient depletion of endogenous DAB2IP, and is in agreement with previous literature reporting tumor-suppressive functions for miR-149-5p [31, 32]. FACS analysis revealed that both miRNAs have negligible effects on the proliferation rate of transfected PC3 (Figure S2F), thus pointing to a major role in cell motility.
To further confirm this observation, we tested both miRNAs in non-transformed prostate epithelial cell lines RWPE-1 and PWR-1E: overexpression of miR-149-3p stimulated cell invasion and migration, while miR-149-5p had no detectable effects (Fig. 3d and S2G). These data suggest that the two miRNAs encoded by the miR-149 precursor, despite sharing the potential to target DAB2IP, can have different biological activities. In particular, they suggest a specific activity of miR-149-3p in sustaining tumor cells invasion.
miR-149-3p increases the invasive behavior of cancer cells through DAB2IP downregulation
To asses if DAB2IP downregulation is the main driver for invasion induced by miR-149-3p, we performed epistasis experiments in PC3 cells. Transfection of a miR-149-3p mimic, depletion of DAB2IP by siRNA, or co-transfection of both reagents induced a similar increase in cell invasion, supporting the notion that they act on the same molecular axis (Fig. 4a).
Functional interaction between miR-149-3p and DAB2IP in prostate cancer cells. a The effects of miR-149-3p and DAB2IP depletion are not additive. PC3 cells were transfected with the indicated reagents for 48 h. Invasion assays were performed as in Fig. 3a. Graph summarizes migrated cells per area (mean ± SD; n = 3; **P < 0.01; *P < 0.05). Western blot confirmed DAB2IP silencing. b DAB2IP overexpression blocks miR-149-3p-induced cell migration. PC3 cells were infected with retroviral construct expressing DAB2IP lacking 3′UTR sequences (pLPC DAB2IP), or control vector (pLPC empty). Subsequently, infected cells were transfected with miR-149-3p mimic or control siRNA for 48 h. Invasion assays were performed as in Fig. 3a. Graph summarizes migrated cells per area (mean ± SD; n = 3; **P < 0.01; *P < 0.05). A control western blot of endogenous and exogenous DAB2IP proteins is shown. c Schematic representation of three putative miR-149-3p target sites (seed sequence in red) on DAB2IP 3′UTR (NM_032552.2) as predicted by Target Scan and miRDB tools. Target Protector oligos were designed to cover site 1 (TP1), site 2 (TP2), and site 3 (TP3). d Target protectors increase endogenous DAB2IP protein levels. PC3 cells were transfected with target protectors (TP1, TP2, TP3), alone or in combination, or with a non-targeting control (TPc). DAB2IP protein levels were measured by western blot. Bottom: graph summarizes the fold increase over control (TPc transfected sample) measured by densitometry of autoradiography films (mean ± SD; n = 3). e Target protectors reduce prostate cancer cells invasion. PC3 cells were transfected with control siRNA (siCtrl) or miR-149-3p mimic, and target protectors mix (TPmix) or negative control (TPc) as indicated for 48 h. Invasion assays were performed as in Fig. 3a. Graph summarizes migrated cells per area (mean ± SD; n = 4; **P < 0.01). Immunoblotting was used to check endogenous DAB2IP protein levels. Actin was blotted as a loading control
We next tested whether overexpression of DAB2IP could rescue the effects of miR-149-3p. To this aim, we infected cells with a retrovirus encoding DAB2IP without the 3′UTR sequence; in line with a functional epistasis, miR-149-3p overexpression could not stimulate invasion of DAB2IP-overexpressing cells (Fig. 4b).
To prove that miR-149-3p-induced invasion depends specifically on its action on DAB2IP, we designed target protector oligos (TPs) for three putative miR-149-3p binding sites in the 3′UTR of human DAB2IP mRNA (Fig. 4c). Transfection of TPs significantly increased DAB2IP protein levels, confirming that endogenous miR-149-3p is active in PC3 cells (Fig. 4d). Results indicate that sites 1 and 2 are the primary targets of endogenous miR-149-3p in PC3, since protection of site 3 showed minimal impact. Very robust results were obtained by combined transfection of all three oligos (TPmix). Importantly, transfection of TPs significantly reduced both basal and miR-149-3p-induced cell motility (Fig. 4e), thus confirming that DAB2IP is the most relevant cellular target of miR-149-3p in mediating the invasive phenotype.
Inhibition of miR-149-3p limits tumor growth and dissemination in vivo
To analyze the role of miR-149-3p in sustaining tumor growth and dissemination in vivo, we initially used zebrafish (Danio rerio) xenotransplantation, a rapid yet reliable animal model for cancer growth [33]. We injected PC3 cells stably expressing the miR-149-3p sponge construct (TW3 S149-3p), or a control vector (TW3 empty), into the yolk sac of zebrafish embryos; expression of the miR-149-3p sponge significantly counteracted xenograft growth (Fig. 5a, b). Interestingly, growth of the tumor mass in animals injected with control cells correlated with increased expression of zebrafish angiogenesis markers (zVEGF A, zVEGFR-1, and zIL8), which was not observed in larvae injected with cells expressing the miR-149-3p sponge (Fig. 5c). These data not only support the notion that miR-149-3p fosters malignant cell growth in vivo, but also suggest an involvement of miR-149-3p in tumor crosstalk with the surrounding microenvironment.
To further study the role of miR-149-3p in vivo, we generated clones of PC3 stably expressing Luciferase, together with control (TW3 empty) or miR-149-3p sponge constructs (TW3 S149-3p). Cells were injected intravenously in NOD/SCID mice, and their growth and dissemination were evaluated at various times by bioluminescence assay (BLI). BLI signals were heavily reduced in mice injected with cells expressing the miR-149-3p sponge (Fig. 5d, e; P < 0.5; Anova test for repeated measures), indicating less efficient lung colonization.
miR-149-3p inhibition decreases growth rate and metastatic colonization of xenografted prostate cancer cells in vivo. a Representative images of the tumor mass formed by PC3 cells at 1 and 3 days post injection in zebrafish larvae. Left panels show a larva injected with PC3 cells expressing TW3 empty vector. Right panels show a larva injected with PC3 cells expressing the TW3 S149-3p sponge. Injected cells express eGFP (green), encoded by the TW3 vector, and are stained with DiI (red). Scale bar = 200 μm (dpf = day post fertilization; dpi = day post injection). b Fluorescence-based quantification of PC3 TW3 empty and PC3 TW3 S149-3p tumor masses at 1, 2, and 3 dpi. Data, normalized to 1 dpi, are expressed as mean ± SEM (N = 36). c mRNA levels of zebrafish VEGFA (zVEGFA), VEGFR-1 (zVEGFR-1) and IL8 (zIL-8), were evaluated by RT-qPCR at 1 and 3 dpi. All genes were normalized to zebrafish actin mRNA. Data are expressed as mean ± SD from two independent experiments (15 larvae per experiment; ***P < 0.001; **P < 0.01; *P < 0.05). d NOD/SCID mice (n = 18; 6/group) were injected i.v. with 0.5×106 cells PC3-PIP Luc transduced with TW3 empty vector, or with the miR-149-3p sponge construct TW3 (two different clones). Panels show two mice per group, as imaged by BLI at different time points. e Cumulative results of bioluminescence in regions of interest (ROI) in total body. Tumor growth was monitored as photon flux and quantified as photons * sec−1 * cm−2 * sr−1. Anova test for repeated measures was used for statistical analysis (TW3 empty vs. TW3 S149-3p clone B: P < 0.05. TW3 empty vs. TW3 S149-3p clone C: P < 0.05. TW3 S149-3p clone B vs. TW3 S149-3p clone C: P > 0.05)
miR-149-3p sustains an inflammatory and pro-angiogenic NF-kB-related gene expression program
To gain a molecular insight on the consequences of miR-149-3p-induced DAB2IP loss-of-function, we analyzed the transcriptional profile of PC3 transfected with miR-149-3p mimic or with DAB2IP-specific siRNAs. Not surprisingly, overexpression of miR-149-3p had a much broader impact on gene expression than DAB2IP knockdown, in line with its action on multiple cellular targets (Fig. 6a). Gene Set Enrichment Analysis (GSEA) revealed enrichment for hallmarks of inflammatory and immune response, apoptosis, and epithelial to mesenchymal transition (EMT) in genes upregulated by miR-149-3p (miR-149-3p_UP, 1337 genes) or DAB2IP siRNA (siDAB2IP_UP, 285 genes) (Table S2). One top enriched hallmarks in both conditions was “TNF signaling via NF-κB”, a pathway related to EMT and cell motility (Table S2 and Figure S3A).
To define the pathways and processes modulated by miR-149-3p through DAB2IP inhibition, we focused on genes upregulated both by DAB2IP depletion and by miR-149-3p overexpression (Common_UP, 112 genes): GSEA confirmed enrichment for genes involved in NF-κB signaling, inflammatory response, and EMT (Fig. 6a). The Common_UP gene set includes various inflammatory chemokines and cytokines, including CCL20 and CSF2 (GM-CSF), that modulate cell motility and crosstalk with the surrounding environment [34,35,36]. Of note, the Common_UP gene set includes IL8, an important inducer of cell motility, angiogenesis, and microenvironment remodeling [35, 37, 38].
We validated microarray results by performing RT-qPCR on a selection of Common_UP genes, confirming their upregulation upon transfection of a miR-149-3p mimic (Fig. 6b). Accordingly, a miR-149-3p inhibitor reduced basal expression levels of the same genes (Figure S3B).
miR-149-3p-mediated DAB2IP reduction reprograms gene expression promoting NF-κB activation. a–d PC3 cells were transfected with DAB2IP siRNAs, control siRNA, or miRNA-149-3p mimic for 48 h and analyzed by Illumina microarray. Genes displaying a variation greater than 1.5 fold were selected for gene set enrichment analysis (GSEA). a GSEA of common genes upregulated by miR-149-3p overexpression or DAB2IP silencing. Venn diagram shows the intersection of miR-149-3p_UP and siDAB2IP_UP gene sets (Common_UP). Graph summarizes the top enriched hallmarks in common upregulated genes (MSigDB = Molecular Signature Data Base). b Expression of selected genes in the Common_UP group was validated by RT-qPCR. Values are normalized to histone H3 and compared to siCtrl samples (mean ± SD; n = 4). A control western blot is shown on the right. c GSEA of common genes downregulated by miR-149-3p overexpression or DAB2IP silencing. Venn diagram shows the intersection of miR-149-3p_DW and siDAB2IP_DW gene sets (Common_DW). Graph summarizes the top enriched hallmarks in common downregulated genes. d Expression of selected genes in the Common_DW group was validated by RT-qPCR. Values are normalized to histone H3 and compared to siCtrl samples (mean ± SD; n = 3). e Overexpression of miR-149-3p or siRNA-mediated DAB2IP depletion increases nuclear localization of p65 RelA. PC3 cells were transfected as described in a and subjected to biochemical fractionation to purify nuclei from cytoplasm. Tubulin (cytoplasmic) and Lamin A/C (nuclear) were blotted as controls. DAB2IP depletion was checked by western blot of total samples. f Depletion of p65 RelA blocks miR-149-3p-induced cell invasion. PC3 cells were co-transfected as indicated. After 48 h, invasion assays were performed as in Fig. 3a (mean ± SD; n = 3; ***P < 0.001; *P < 0.05). Western blot confirmed DAB2IP and p65 RelA silencing. Actin was blotted as a loading control
We also analyzed genes downregulated by miR-149-3p (miR149-3p_DW, 1958 genes) or DAB2IP siRNA (siDAB2IP_DW, 1057 genes) (Table S2). Genes that are downregulated both by miR-149-3p and DAB2IP siRNA (Common_DW) displayed enrichment of hallmarks “E2F targets”, “TGF-β signaling”, and “apical junction” (Fig. 6c)—the latter suggesting a possible link with cell motility. We validated by RT-qPCR a set of Common_DW genes potentially involved in cell adhesion and actin rearrangement (Fig. 6d), confirming their regulation.
Therefore, miR-149-3p overexpression in PC3 dictates a pro-invasive transcriptional profile that comprises multiple secreted factors, suggesting a cell non-autonomous component in the invasive phenotype.
miR-149-3p sustains cell invasion through increased activation of NF-κB signaling
The hallmark most enriched in Common_UP genes is “TNF signaling via NF-κB” (Fig. 6a and Figure S3A), and validated miR-149-3p-induced genes are NF-κB targets (CCL20, IL8, CSF2, c-IAP2, and PTX3). We therefore tested if miR-149-3p could activate NF-κB. As reported in Fig. 6e, PC3 cells transfected with miR-149-3p had increased levels of nuclear p65 RelA—similar to cells depleted of DAB2IP by siRNA. We then tested the relevance of NF-κB activation for the invasive phenotype. Strikingly, knockdown of p65 totally abolished the increase in invasion triggered by miR-149-3p overexpression, or by DAB2IP depletion (Fig. 6f), confirming that NF-κB activity is required for this phenotype.
Inflammatory cytokines in the tumor microenvironment can activate NF-κB in cancer cells, affecting cancer progression and metastasis [39]. We therefore tested if miR-149-3p levels might affect the cell’s response to TNFα; we found that miR-149-3p overexpression enhanced TNF-dependent induction of pro-invasive inflammatory factors (Figure S3C), while transfection of a miR-149-3p inhibitor reduced TNF-dependent induction of CCL20 and IL8 (Figure S3D). Accordingly, miR-149-3p overexpression significantly enhanced TNF-induced cell motility (Figure S3E). We conclude that NF-κB is a key effector of invasion triggered by miR-149-3p in these cells—consistent with DAB2IP dysfunction.
miR-149-3p is secreted by cancer cells and downregulates DAB2IP in receiving cells
In a recent study, various miRNAs computationally predicted to target DAB2IP have been found in exosomes from PC3 [40]; notably, miR-149-3p is one of these (Figure S4A). By RT-qPCR, we readily confirmed the presence of miR-149-3p in the culture medium of PC3, with significantly lower levels when cells were infected with the sponge construct (Fig. 7a). We therefore hypothesized that secretion of miR-149-3p by cancer cells might reduce DAB2IP in neighboring non-transformed cells, potentially affecting their response to inflammatory and pro-invasive factors released by cancer cells, eventually creating a favorable condition for tumor growth. To test this, we treated non-transformed PWR-1E cells with PC3-conditioned medium. Notably, we observed a powerful increase in the invasive properties of these cells (Fig. 7b), together with augmented intracellular miR-149-3p levels (Figure S4B). In line with the action of miR-149-3p, we detected reduced DAB2IP mRNA and protein levels (Fig. 7c), and increased expression of miR-149-3p-related NF-κB target genes (Figure S4C).
miR-149-3p is secreted by PC3 and reduces DAB2IP levels in recipient non-transformed cells, shifting their behavior toward a pro-oncogenic phenotype. a miR-149-3p is secreted by PC3. Levels of miR-149-3p were measured by RT-qPCR in culture medium of PC3 transduced with control (TW3) or antisense miR-149-3p sponge construct (mean ± SD; n = 3; **P < 0.01). Values are normalized to a standard amount of cel-39 miRNA added to QIAzol prior to RNA extraction. b PC3-conditioned medium stimulates invasion of non-transformed prostate cells. PWR-1E were treated with control (Ctrl) or PC3-conditioned medium (CM_PC3), and subsequently seeded for matrigel invasion assays (see Methods for details). Graphs summarize migrated cells per area (mean ± SD; n = 3; ***P < 0.001). Representative images of migrated cells are shown. Scale bar = 100 μm. c PC3-conditioned medium reduces DAB2IP mRNA and protein levels in recipient cells. PWR-1E cells were treated as in b for 48 h. DAB2IP mRNA was quantified by RT-qPCR (mean ± SD; n = 3; ***P < 0.001). DAB2IP protein was detected by western blot with Actin as a loading control. d The effects of PC3-conditioned medium depend on secreted miR-149-3p. PWR-1E cells were treated with control medium (Ctrl), or with medium conditioned by PC3 transduced with empty vector (TW3) or miR-149-3p sponge construct. Invasion assays were performed as in b (mean ± SD; n = 3; *P < 0.05). DAB2IP protein levels were measured by western blot. e Expression of a miR-149-3p sponge protects recipient cells from the action of PC3-conditioned medium. PWR-1E cells stably transduced with control (TW3) or antisense miR-149-3p sponge construct were treated with control or PC3-conditioned medium. Invasion assays were performed as in b (mean ± SD; n = 3; *P < 0.05). DAB2IP and GFP protein levels were analyzed by western blot. f PC3-conditioned medium reduces DAB2IP mRNA and protein levels in vascular endothelial cells. HUVEC were treated with control or PC3-conditioned medium for 48 h. DAB2IP mRNA and protein were measured as in c (mean ± SD; n = 3; *P < 0.05). g PC3-conditioned medium stimulates migration and proliferation of vascular endothelial cells. HUVEC were treated as in f. Left graph: transwell migration assay (mean ± SE; n = 3; Mann–Whitney test, *P < 0.05). Right graph: expression of Ki67 proliferation marker as measured by fluorescent ELISA (mean ± SE; n = 3; Mann–Whitney test, **P < 0.01). Control western blots are shown below. h Proposed model for miR-149-3p-mediated cell autonomous and cell non-autonomous inhibition of DAB2IP
Medium conditioned by PC3 expressing the miR-149-3p sponge (TW3 S149-3p) had a much weaker effect on cell invasion and DAB2IP levels (Fig. 7d), implying that miR-149-3p is a key mediator of the observed phenotype. To confirm this, we transduced the miR-149-3p sponge in PWR-1E cells, thus making them resistant to the action of miR-149-3p (Figure S4D). Expression of the sponge counteracted DAB2IP reduction and matrigel invasion in cells treated with PC3-conditioned medium (Fig. 7e), strongly indicating that both phenotypes are mediated by secreted miR-149-3p.
miR-149-3p secreted by PC3 induces migration and proliferation of endothelial cell
Tissue-specific DAB2IP knockout in vascular endothelial cells was reported to potentiate tumor growth and metastasis in orthotopic breast cancer and subcutaneous melanoma mouse models, notably by favoring generation of a pro-metastatic niche [41]. We therefore asked whether secreted miR-149-3p could downregulate DAB2IP in endothelial cells. To this aim, we treated primary human umbilical vein endothelial cells (HUVEC) with PC3-conditioned medium; we observed reduction of DAB2IP protein and mRNA, with concurrent increase of intracellular miR-149-3p levels (Fig. 7f and Figure S4E,F). In line with the model, PC3-conditioned medium induced a significant increase in HUVEC migration and proliferation (Fig. 7g), as well as augmented expression of pro-angiogenic markers IL8, FGF2, and VEGFA (Figure S4G). We conclude that miR-149-3p secreted by the tumor cells can potentially reduce DAB2IP protein levels in adjacent endothelial cells, thereby reprograming their response to extrinsic inputs and facilitating remodeling of the tumor vasculature (Fig. 7h), with implications for cancer growth and dissemination.
Discussion
The tumor suppressor DAB2IP is an important modulator of multiple oncogenic signaling pathways, and is disabled in malignancies through multiple mechanisms [5]. Using high-throughput functional screening, we identified novel miRNAs able to target DAB2IP. Interestingly, we found a few miRNAs that are not predicted by available algorithms (e.g., miR-555 and miR-212) but can downregulate DAB2IP in cell lines; this corroborates the concept that sequence-based predictions need extensive experimental validation. On the other hand, all top hits could downregulate endogenous DAB2IP in at least one cell line, confirming that a reporter-based functional screen is a powerful tool for identifying target-specific miRNAs.
Among top hits, we uncovered known onco-miRNAs, such as miR-96 [42], miR-25, and miR-92a [20]. Although we have not investigated this aspect, it is possible that DAB2IP inhibition is part of the mechanisms through which these miRNAs promote proliferation, survival, and motility of cancer cells. For instance, miR-96 and miR-92a were reported to activate PI3K/AKT/mTOR signaling [20, 42], a phenotype potentially enhanced by DAB2IP loss [9].
For its robust effect on DAB2IP protein levels, we focused on miR-149-3p. Bioinformatic analysis of public cancer datasets revealed augmented expression of this miRNA in various human malignancies. Functional assays in tissue culture confirmed that miR-149-3p targets DAB2IP and stimulates invasion of prostate, breast, and ovarian cancer cell lines.
Intriguingly, also miR-149-5p, encoded by the same miRNA precursor, can target DAB2IP; nonetheless, miR-149-5p overexpression had no detectable effects on cell invasion. This may be partly explained by less efficient DAB2IP reduction, as suggested by some of our experiments. In addition, miR-149-5p has been described as a tumor-suppressive miRNA that can target pro-invasive proteins [31, 32], so its inhibition of oncogenic effectors may compensate the functional effects of DAB2IP reduction.
We analyzed the transcriptome of PC3 cells overexpressing either miR-149-3p or a specific DAB2IP siRNA. To our knowledge, this is the first study to examine the transcriptional program of prostate cancer cells silenced for DAB2IP. As expected, upon DAB2IP knockdown, we observed upregulation of genes involved in biological processes and pathways modulated by DAB2IP, such as NF-kB activation by TNF, unfolded protein response (UPR), EMT, apoptosis, K-Ras, p38MAP kinase, and STAT signaling [4, 5]. In addition, we detected enrichment for genes implicated in hypoxia, inflammation, and immune system regulation, suggesting a role for DAB2IP in additional molecular circuits.
In response to TNFα, DAB2IP limits the activation of NF-kB while promoting activation of ASK1/JNK kinases [1, 2]. Depletion of DAB2IP by miR-149-3p was therefore expected to stimulate NF-kB activity, as emerged clearly from this analysis and was experimentally confirmed in PC3. However, we cannot exclude that miR-149-3p action can potentiate additional signaling pathways controlled by DAB2IP, thus affecting the cell’s response to multiple extrinsic inputs.
Genes affected by miR-149-3p overexpression displayed enrichment for several additional biological processes, including glycolysis, response to estrogen, and heme metabolism, that are not enriched upon DAB2IP knockdown; these suggest a specific impact of this miRNA in additional cancer-related phenotypes.
By triggering NF-κB activation via DAB2IP inhibition, miR-149-3p enhances oncogenic responses to TNF and may thus increase aggressiveness of cancer cells in an inflammatory microenvironment [8]. By the same mechanism, miR-149-3p enhances transcription of various inflammatory factors, both in basal conditions and after TNF treatment; in particular CCL20, IL8, PTX3, and CSF2 that are inducers of metastasis, neo-angiogenesis, and remodeling of the tumor microenvironment, and often predict worst patient survival [34, 35, 38, 43]. It is therefore plausible that cancer cells overexpressing miR-149-3p not only increase their intrinsic motility, but also interact with stromal cells to reprogram the tumor microenvironment. In line with this concept, we observed that PC3 cells injected in Zebrafish embryos grow as a tumor and induce a pro-angiogenic gene expression signature in the host, and this effect is abolished by miR-149-3p inhibition.
Finally, we observed that miR-149-3p is secreted by PC3 cells and can reduce DAB2IP protein levels in receiving non-transformed cells. Although we cannot exclude that other secreted miRNAs, or proteins, contribute to DAB2IP downregulation in receiving cells, experiments with a specific inhibitor (sponge construct) demonstrated that miR-149-3p is a major mediator of this effect. Notably, Ji et al. recently reported that tissue-specific DAB2IP knockout in vascular endothelial cells accelerates neovascularization and promotes formation of a pre-metastatic niche, in mouse models of melanoma and breast cancer [41]. We found that PC3 cells can reduce DAB2IP in vascular endothelial cells by secreting miR-149-3p in the culture medium. This suggests a mechanism by which real-life tumors could locally establish a condition similar to that modeled by Ji and colleagues in transgenic mice [41]. In fact, PC3-conditioned medium triggered proliferation and motility of endothelial cells, together with increased IL8 and VEGF expression, supporting the notion of tumor-induced neo-angiogenesis. Intriguingly, we verified that human miR-149-3p can target murine DAB2IP, which contains six matching seed sequences in the 3′UTR (Figure S4H,I). It cannot therefore be excluded that part of the phenotype observed in NOD/SCID mice may be due to the action of secreted miR-149-3p.
In conclusion, our data suggest that the intracellular action of miR-149-3p enhances the invasive behavior of cancer cells and promotes expression of inflammatory and pro-angiogenic factors, while tumor-secreted miR-149-3p can act on stromal cells, establishing a supporting microenvironment that can potentially facilitate tumor growth (Fig. 7h). Future studies shall be aimed at testing this hypothesis in a broader context. If confirmed, the dual intracellular and extracellular role of miR-149-3p would make this miRNA an attractive candidate for targeted therapeutics, and a potentially interesting biomarker.
Materials and methods
Cell culture
The following human cell lines were used: HeLa, PC3, LNCaP, PWR-1E, RWPE-1, MCF10A, HBL100, SKOV-3, H1299, HCT116, HePG2, HUVEC, 293T, and 293GP. All cell lines were authenticated by STR genotyping, and tested for Mycoplasma weekly. For all experiments, cells were cultured for less than a month after thawing. HUVECs were isolated by collagenase treatment and cultured as previously described [44]. The study was approved by the institutional review board of Maternal-Children’s Hospital (IRCCS “Burlo Garofolo”, Trieste) and informed consent was obtained from all patients providing placental tissue specimens. All experiments were carried out within the third passage of cell culture.
For preparation of conditioned medium, cells were cultured for 30 h, then medium was collected, centrifuged, and filtered using 0.22 μm filters. Conditioned medium was used immediately or kept at 4 °C for up to 1 week.
Cells were treated with 10 ng/ml human recombinant TNFα (Invitrogen, PHC3015) in low serum (0.1% FBS) for 20 h. Cells were transfected with 50 nM siRNA oligonucleotides, 3 nM miRNAs mimics, 20 nM miRNAs hairpin inhibitors, and TPs, using Lipofectamine RNAiMax (Invitrogen), following manufacturer’s instructions.
Expression plasmids and viral constructs are described in Supplementary Information.
High-throughput screening
HeLa cells were transfected with MiRIDIAN miRNA mimics library (Dharmacon, GE Healthcare), as described previously [45], together with the reporter plasmid psiCHECK2 DAB2IP 3′UTR. Cells were then lysed in Glo Lysis Buffer (Promega); Firefly and Renilla activities were measured using Dual-Glo Luciferase Assay System (Promega) in a PerkinElmer Envision multimode plate reader. The screening was performed in duplicate at the ICGEB High-Throughput Screening Facility (http://www.icgeb.org/high-throughput-screening.html). Experimental details can be found in Supplementary Information.
Gene expression analysis
Total RNA was extracted with QIAzol (Qiagen). For mRNA expression analysis, 1 μg of total RNA was reverse-transcribed with QuantiTect Reverse Transcription (Qiagen). Analyzed genes were amplified using SsoAdvancedTMSYBR® Green Master Mix (Biorad) on a CFX96™ Real-Time PCR System (Biorad). For miRNAs expression analysis, 1 μg of RNA was retro-transcribed with miScript PCR System (Qiagen). MicroRNAs were amplified with miScript SYBR Green PCR kits (Qiagen), on a CFX96™ Real-Time PCR System (Biorad).
For microarray profiling, two biological replicates for each group (siCtrl, siDAB2IP, miR-149-3p) were hybrized on HumanHT-12-v4-BeadChip (Illumina). Primer sequences and microarray analysis are described in Supplementary Information.
Cell migration, invasion, and proliferation assays
Transwell migration and invasion assays were performed as described previously [8, 9]. In experiments with conditioned medium, cells were pre-incubated with Ctrl or PC3-conditioned medium for 24 h, and subsequently seeded in the presence of Ctrl or PC3-conditioned medium; the lower chamber of the transwell system was filled with high-serum medium.
HUVEC migration assays were performed as previously reported [46]. Cells were pre-incubated with Ctrl or PC3-conditioned medium for 24 h, and seeded in the upper compartment. The lower compartment was filled with Ctrl or PC3-conditioned medium.
For HUVEC proliferation assays, cells pre-incubated with Ctrl or PC3-conditioned medium for 24 h were seeded on a 2% gelatin-coated 96-well plate, and then stimulated with Ctrl or PC3-conditioned medium, for 18 h. Cells were then fixed and permeabilized with FIX & PERM kit (Società Italiana Chimici, Rome, Italy) and incubated with primary mAb (clone MIB-1) mouse anti-human anti Ki67, followed by FITC-conjugated goat anti-mouse IgG (Dako). After the staining, the number of Ki67-positive cells was measured using an Infinite 200 Pro plate reader (absorbance 485 nm, emission 535 nm) (TECAN, Italy).
Zebrafish xenografts
Zebrafish were raised and maintained as previously described [47]. Two days post fertilization (2 dpf), embryos were dechorionated; approximately 500 PC3 cells expressing TW3 empty or TW3 S149-3p vectors were microinjected into the yolk of each embryo. Embryos were maintained in E3 medium/PTU for 1 h at 28 °C, then kept at 34 °C to allow tumor cells survival and growth as previously described [33]. During all procedures under fluorescence microscope (Leica DM 2000), live animals, properly anesthetized, were positioned in 1.5% methylcellulose (Sigma Aldrich Co). The images were acquired in both green (GFP) and red signals (DiI staining) and then merged with the respective bright field image by using the Leica Application Suite X (LAS X) software. The evaluation of the tumor mass area was performed using the ImageJ software. For gene expression, total RNA was extracted using Trizol Reagent (Invitrogen, Life Technologies, Milan, Italy). One microgram of total RNA was reverse transcribed using M-MLV reverse transcriptase (Life Technologies). Gene expression was analyzed by RT-qPCR (Step One Plus, Applied Biosystems) using the SYBR Green system (Life Technologies). Primer sequences are listed in Supplementary Information.
Mouse xenografts
Six- to eight-week-old female immunodeficient NOD/SCID mice were inoculated intravenously (i.v.) with 0.5×106 PC3 cells transduced with TW3 empty vector, or with the miR-149-3p sponge construct TW3 S149-3p (two different clones). Before use, tumor sublines were transduced with a lentiviral vector encoding firefly luciferase, to subsequently track tumor growth metastases by BLI. BLI images were acquired at different time points using the IVIS Lumina II Imaging System (PerkinElmer, Zaventem, Belgium). Growth and spreading of the tumors were monitored weekly for up to 1 month.
Additional methods
Bioinformatic analysis of public datasets, protocols for protein studies, and more detailed procedures can be found in Supplementary Information.
Data availability
The full microarray dataset has been submitted to Gene Expression Omnibus (GSE104353).
References
Min J, Zaslavsky A, Fedele G, McLaughlin SK, Reczek EE, De Raedt T, et al. An oncogene-tumor suppressor cascade drives metastatic prostate cancer by coordinately activating Ras and nuclear factor-kappaB. Nat Med. 2010;16:286–94.
Zhang H, Zhang R, Luo Y, D’Alessio A, Pober JS, Min W. AIP1/DAB2IP, a novel member of the Ras-GAP family, transduces TRAF2-induced ASK1-JNK activation. J Biol Chem. 2004;279:44955–65.
Xie D, Gore C, Zhou J, Pong RC, Zhang H, Yu L, et al. DAB2IP coordinates both PI3K-Akt and ASK1 pathways for cell survival and apoptosis. Proc Natl Acad Sci USA. 2009;106:19878–83.
Liu L, Xu C, Hsieh JT, Gong J, Xie D. DAB2IP in cancer. Oncotarget. 2016;7:3766–76.
Bellazzo A, Di Minin G, Collavin L. Block one, unleash a hundred. Mechanisms of DAB2IP inactivation in cancer. Cell Death Differ. 2017;24:15–25.
Chen H, Tu SW, Hsieh JT. Down-regulation of human DAB2IP gene expression mediated by polycomb Ezh2 complex and histone deacetylase in prostate cancer. J Biol Chem. 2005;280:22437–44.
Dote H, Toyooka S, Tsukuda K, Yano M, Ouchida M, Doihara H, et al. Aberrant promoter methylation in human DAB2 interactive protein (hDAB2IP) gene in breast cancer. Clin Cancer Res. 2004;10:2082–9.
Di Minin G, Bellazzo A, Dal Ferro M, Chiaruttini G, Nuzzo S, Bicciato S, et al. Mutant p53 reprograms TNF signaling in cancer cells through interaction with the tumor suppressor DAB2IP. Mol Cell. 2014;56:617–29.
Valentino E, Bellazzo A, Di Minin G, Sicari D, Apollonio M, Scognamiglio G, et al. Mutant p53 potentiates the oncogenic effects of insulin by inhibiting the tumor suppressor DAB2IP. Proc Natl Acad Sci USA. 2017;114:7623–8.
Liao H, Xiao Y, Hu Y, Xiao Y, Yin Z, Liu L. microRNA-32 induces radioresistance by targeting DAB2IP and regulating autophagy in prostate cancer cells. Oncol Lett. 2015;10:2055–62.
Xu Y, He J, Wang Y, Zhu X, Pan Q, Xie Q, et al. miR-889 promotes proliferation of esophageal squamous cell carcinomas through DAB2IP. FEBS Lett. 2015;589:1127–35.
Huang J, Wang B, Hui K, Zeng J, Fan J, Wang X, et al. miR-92b targets DAB2IP to promote EMT in bladder cancer migration and invasion. Oncol Rep. 2016;36:1693–701.
Barik S. An intronic microRNA silences genes that are functionally antagonistic to its host gene. Nucleic Acids Res. 2008;36:5232–41.
Feng C, Sun P, Hu J, Feng H, Li M, Liu G, et al. miRNA-556-3p promotes human bladder cancer proliferation, migration and invasion by negatively regulating DAB2IP expression. Int J Oncol. 2017;50:2101–12.
Yun E-J, Zhou J, Lin C-J, Xu S, Santoyo J, Hernandez E, et al. The network of DAB2IP-miR-138 in regulating drug resistance of renal cell carcinoma associated with stem-like phenotypes. Oncotarget. 2017;8:66975–86.
Lin S, Gregory RI. MicroRNA biogenesis pathways in cancer. Nat Rev Cancer. 2015;15:321–33.
Bracken CP, Scott HS, Goodall GJ. A network-biology perspective of microRNA function and dysfunction in cancer. Nat Rev Genet. 2016;17:719–32.
Fogli S, Polini B, Carpi S, Pardini B, Naccarati A, Dubbini N, et al. Identification of plasma microRNAs as new potential biomarkers with high diagnostic power in human cutaneous melanoma. Tumour Biol. 2017;39:1010428317701646.
Rupaimoole R, Slack FJ. MicroRNA therapeutics: towards a new era for the management of cancer and other diseases. Nat Rev Drug Discov. 2017;16:203–22.
Li M, Guan X, Sun Y, Mi J, Shu X, Liu F, et al. miR-92a family and their target genes in tumorigenesis and metastasis. Exp Cell Res. 2014;323:1–6.
Yang Y, Gu X, Zhou M, Xiang J, Chen Z. Serum microRNAs: a new diagnostic method for colorectal cancer. Biomed Rep. 2013;1:495–8.
Li P, Sheng C, Huang L, Zhang H, Huang L, Cheng Z, et al. MiR-183/-96/-182 cluster is up-regulated in most breast cancers and increases cell proliferation and migration. Breast Cancer Res. 2014;16:473.
Lin RJ, Lin YC, Yu AL. miR-149* induces apoptosis by inhibiting Akt1 and E2F1 in human cancer cells. Mol Carcinog. 2010;49:719–27.
Jin L, Hu WL, Jiang CC, Wang JX, Han CC, Chu P, et al. MicroRNA-149*, a p53-responsive microRNA, functions as an oncogenic regulator in human melanoma. Proc Natl Acad Sci USA. 2011;108:15840–5.
Cao D, Jia Z, You L, Wu Y, Hou Z, Suo Y, et al. 18beta-glycyrrhetinic acid suppresses gastric cancer by activation of miR-149-3p-Wnt-1 signaling. Oncotarget. 2016;7:71960–73.
Fan SJ, Li HB, Cui G, Kong XL, Sun LL, Zhao YQ, et al. miRNA-149* promotes cell proliferation and suppresses apoptosis by mediating JunB in T-cell acute lymphoblastic leukemia. Leuk Res. 2016;41:62–70.
Okato A, Arai T, Yamada Y, Sugawara S, Koshizuka K, Fujimura L, et al. Dual strands of pre-miR-149 inhibit cancer cell migration and invasion through targeting FOXM1 in renal cell carcinoma. Int J Mol Sci. 2017;18:pii: E1969.
Chamorro-Jorganes A, Araldi E, Rotllan N, Cirera-Salinas D, Suarez Y. Autoregulation of glypican-1 by intronic microRNA-149 fine tunes the angiogenic response to FGF2 in human endothelial cells. J Cell Sci. 2014;127:1169–78.
Xie D, Gore C, Liu J, Pong RC, Mason R, Hao G, et al. Role of DAB2IP in modulating epithelial-to-mesenchymal transition and prostate cancer metastasis. Proc Natl Acad Sci USA. 2010;107:2485–90.
Taylor BS, Schultz N, Hieronymus H, Gopalan A, Xiao Y, Carver BS, et al. Integrative genomic profiling of human prostate cancer. Cancer Cell. 2010;18:11–22.
Bischoff A, Huck B, Keller B, Strotbek M, Schmid S, Boerries M, et al. miR149 functions as a tumor suppressor by controlling breast epithelial cell migration and invasion. Cancer Res. 2014;74:5256–65.
Chan SH, Huang WC, Chang JW, Chang KJ, Kuo WH, Wang MY, et al. MicroRNA-149 targets GIT1 to suppress integrin signaling and breast cancer metastasis. Oncogene. 2014;33:4496–507.
Jung DW, Oh ES, Park SH, Chang YT, Kim CH, Choi SY, et al. A novel zebrafish human tumor xenograft model validated for anti-cancer drug screening. Mol Biosyst. 2012;8:1930–9.
Park BK, Zhang H, Zeng Q, Dai J, Keller ET, Giordano T, et al. NF-kappaB in breast cancer cells promotes osteolytic bone metastasis by inducing osteoclastogenesis via GM-CSF. Nat Med. 2007;13:62–69.
Waugh DJ, Wilson C. The interleukin-8 pathway in cancer. Clin Cancer Res. 2008;14:6735–41.
Zeng W, Chang H, Ma M, Li Y. CCL20/CCR6 promotes the invasion and migration of thyroid cancer cells via NF-kappa B signaling-induced MMP-3 production. Exp Mol Pathol. 2014;97:184–90.
De Palma M, Biziato D, Petrova TV. Microenvironmental regulation of tumour angiogenesis. Nat Rev Cancer. 2017;17:457–74.
Matsuo Y, Ochi N, Sawai H, Yasuda A, Takahashi H, Funahashi H, et al. CXCL8/IL-8 and CXCL12/SDF-1alpha co-operatively promote invasiveness and angiogenesis in pancreatic cancer. Int J Cancer. 2009;124:853–61.
Karin M, Cao Y, Greten FR, Li ZW. NF-kappaB in cancer: from innocent bystander to major culprit. Nat Rev Cancer. 2002;2:301–10.
Hessvik NP, Phuyal S, Brech A, Sandvig K, Llorente A. Profiling of microRNAs in exosomes released from PC-3 prostate cancer cells. Biochim Biophys Acta. 2012;1819:1154–63.
Ji W, Li Y, He Y, Yin M, Zhou HJ, Boggon TJ, et al. AIP1 expression in tumor niche suppresses tumor progression and metastasis. Cancer Res. 2015;75:3492–504.
Siu MK, Tsai YC, Chang YS, Yin JJ, Suau F, Chen WY, et al. Transforming growth factor-beta promotes prostate bone metastasis through induction of microRNA-96 and activation of the mTOR pathway. Oncogene. 2015;34:4767–76.
Locatelli M, Ferrero S, Martinelli Boneschi F, Boiocchi L, Zavanone M, Maria Gaini S, et al. The long pentraxin PTX3 as a correlate of cancer-related inflammation and prognosis of malignancy in gliomas. J Neuroimmunol. 2013;260:99–106.
Tedesco F, Pausa M, Nardon E, Introna M, Mantovani A, Dobrina A. The cytolytically inactive terminal complement complex activates endothelial cells to express adhesion molecules and tissue factor procoagulant activity. J Exp Med. 1997;185:1619–27.
Mano M, Ippodrino R, Zentilin L, Zacchigna S, Giacca M. Genome-wide RNAi screening identifies host restriction factors critical for in vivo AAV transduction. Proc Natl Acad Sci USA. 2015;112:11276–81.
Bossi F, Tripodo C, Rizzi L, Bulla R, Agostinis C, Guarnotta C, et al. C1q as a unique player in angiogenesis with therapeutic implication in wound healing. Proc Natl Acad Sci USA. 2014;111:4209–14.
Tonon F, Zennaro C, Dapas B, Carraro M, Mariotti M, Grassi G. Rapid and cost-effective xenograft hepatocellular carcinoma model in Zebrafish for drug testing. Int J Pharm. 2016;515:583–91.
Acknowledgements
We thank Giada Pastore (LNCIB, Trieste) for assistance with tissue culture. We thank Andrea Lunardi (CIBIO, University of Trento) for sharing critical reagents. We thank Valeria Capaci, Marco Fantuz, and Lorenzo Bascetta (LNCIB, Trieste) for advice and discussion.
Funding
This work was funded by AIRC (Italian Association for Cancer Research) Investigator Grant (IG 14173) and Università di Trieste (FRA 2015) to LC, and AIRC Special Program Molecular Clinical Oncology “5 per mille” (Grant no. 10016) to GDS and AR. This study was also supported by NIH-NCI Grant R01CA200859 to LM. AB was supported by a “Guglielmina Lucatello e Gino Mazzega” postdoctoral fellowship from FIRC (Fondazione Italiana Ricerca sul Cancro). FT and CZ were supported by Fondazione la Nuova Speranza ONLUS - Lotta alla Glomerulosclerosi Focale, Rho (Milano).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Bellazzo, A., Di Minin, G., Valentino, E. et al. Cell-autonomous and cell non-autonomous downregulation of tumor suppressor DAB2IP by microRNA-149-3p promotes aggressiveness of cancer cells. Cell Death Differ 25, 1224–1238 (2018). https://doi.org/10.1038/s41418-018-0088-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-018-0088-5
This article is cited by
-
Disable 2, A Versatile Tissue Matrix Multifunctional Scaffold Protein with Multifaceted Signaling: Unveiling Role in Breast Cancer for Therapeutic Revolution
Cell Biochemistry and Biophysics (2024)
-
Advances in the understanding of circRNAs that influence viral replication in host cells
Medical Microbiology and Immunology (2024)
-
DDX17 induces epithelial-mesenchymal transition and metastasis through the miR-149-3p/CYBRD1 pathway in colorectal cancer
Cell Death & Disease (2023)
-
MiR-149-3p promotes the cisplatin resistance and EMT in ovarian cancer through downregulating TIMP2 and CDKN1A
Journal of Ovarian Research (2021)
-
Dichloroacetate restores colorectal cancer chemosensitivity through the p53/miR-149-3p/PDK2-mediated glucose metabolic pathway
Oncogene (2020)