Abstract
Cells undergoing Bax/Bak-mediated apoptosis exhibit signs of autophagy, but how it is activated and its significance is unknown. By directly activating Bax/Bak with BH3-only proteins or BH3 mimetic compounds, we demonstrate that mitochondrial damage correlated with a rapid increase in intracellular [AMP]/[ATP], phosphorylation of 5′ AMP-activated protein kinase (AMPK), and activation of unc-51 like autophagy activating kinase 1 (ULK1). Consequently, autophagic flux was triggered early in the apoptotic pathway, as activation of the apoptosome and caspases were not necessary for its induction. Bax/Bak-triggered autophagy resulted in the clearance of damaged mitochondria in an ATG5/7-dependent manner that did not require Parkin. Importantly, Bax/Bak-mediated autophagy inhibited the secretion of the pro-inflammatory cytokine interferon-β (IFN-β) produced in response to mitochondrial damage, but not another cytokine interleukin-6 (IL-6). These findings show that Bax/Bak stimulated autophagy is essential for ensuring immunological silence during apoptosis.
Similar content being viewed by others
Introduction
The autophagy machinery is used to recycle cytoplasmic material, and to protect cells from toxic protein aggregates, dysfunctional organelles and invading pathogens [1]. When energy levels are low, AMP-activated protein kinase (AMPK) is activated and stimulates autophagy by phosphorylating unc-51 like autophagy activating kinase 1 (ULK1) [2, 3]. ULK1 is also activated by nutrient starvation through the mechanistic/mammalian target of rapamycin complex 1 (mTORC1) [1, 3]. AMPK can also negatively regulate mTORC1 by phosphorylating Raptor to activate autophagy [4]. The activated ULK complex initiates the formation of a curved double membrane structure termed the phagophore, which is expanded via a core set of autophagy related (ATG) proteins, to form the completed autophagosome [1].
During canonical autophagy, members of the ATG8 family including microtubule-associated protein 1 light chain 3 beta (LC3B) associate with the growing phagophore and aid the elongation of autophagosomal membranes [5,6,7]. LC3B-I is cytoplasmic, but can associate with the phagophore when conjugated to the lipid phosphatidylethanolamine. This lipidated form of LC3B is termed LC3B-II. This lipidation is performed by a conjugation machinery that includes ATG5 and ATG7 [8], and is routinely used as a marker for autophagy [9]. LC3B-II is associated with both completed autophagosomes and also when this vesicles fuse with lysosomes to form autolysosomes, where encapsulated cytoplasmic material is degraded.
Two commonly used inhibitors of autolysosomal function are bafilomycin A1 (BafA1) and chloroquine (CQ), both of which inhibit the acidification of autolysosomes [1, 10]. Because many lysosomal hydrolases function at low pH, these compounds thereby inhibit degradation of the substrates of autophagy and autophagic flux.
Starvation-induced autophagy is predominantly a non-selective process that encapsulates cellular material within autophagosomal membranes. In contrast, selective forms of autophagy can degrade specific cargo. For instance, mitophagy degrades damaged mitochondria—the best-characterised version requires the E3 ubiquitin ligase Parkin and PTEN induced kinase 1 (PINK 1) [11,12,13,14].
In addition to stimulating autophagy, cellular stresses can also induce mitochondrial-mediated apoptosis. Bax and Bak are the key effector proteins of this form of programmed cell death (Fig. 1a) [15, 16]. Increased expression of BH3-only proteins, such as Bim, or addition of BH3 mimetic compounds such as ABT-737 and S63845, can either directly activate Bax/Bak or prevent pro-survival Bcl-2 family members (Bcl-2, Bcl-xL, Bcl-w, Mcl-1 and A1) from keeping Bax and Bak in check (Fig. 1a) [17, 18]. When activated, Bax/Bak oligomerize to induce mitochondrial outer membrane permeabilisation (MOMP), which stops energy production and releases cytochrome c into the cytoplasm. Cytochrome c can then bind to Apaf-1 and pro-caspase-9 to form the apoptosome, which activates the caspase cascade.
a Schematic of mitochondrial-mediated apoptosis. BH3 only proteins (e.g., Bim) and BH3 mimetic compounds sequester the pro-survival Bcl-2 family from Bax and Bak. Bax and Bak become activated, inducing mitochondrial outer membrane permiabilization, thus releasing cytochrome c. This activates the apoptosome and caspase 3 to induce rapid cell death. The pan-caspase inhibitor QVD-OPh (QVD) inhibits apoptotic caspase activity. b The pan caspase inhibitor QVD does not inhibit Bax/Bak-dependent LC3B lipidation. Mouse embryonic fibroblasts (MEFs) were treated with 1 μM ABT-737 and/or 10 μM QVD for 4 h prior to lysis and western blotting. c Wild type MEFs or those lacking Bax and Bak were infected with a doxycycline (Dox)-inducible Bims lentiviral construct. Cells were treated with 1 μg/ml Dox and/or 10 μM QVD for 4 h prior to lysis. d Apaf-1 is not required for LC3B lipidation following Bax/Bak activation. Wild-type and Apaf-1−/− MEFs previously bearing a Dox-inducible Bims lentiviral construct were treated with 1 μg/ml Dox and/or 10 μM QVD for 4 h. e MEFs lacking Apaf-1 were infected with construct expressing Cas9 and a guide RNA against Mcl-1, cloned to isolate knockouts, and subsequently treated for 4 h with 1 μM ABT-737 to induce Bax/Bak activation
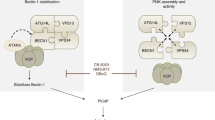
Two principal models have been proposed to explain how pro-survival Bcl-2 family members inhibit autophagy: [19] (1) by directing inhibiting of the autophagy protein Beclin 1 [20,21,22,23,24], and more recently, (2) by inhibiting Bax and Bak [25,26,27,28]. However, how and why Bax and Bak activation induces autophagy remained unclear. Here we show that, without requiring caspase activation by the apoptosome, Bax/Bak activation caused the rapid phosphorylation of AMPK and its substrate ULK1 and subsequent autophagic flux. Mitochondria were digested during Bax/Bak-induced autophagy, presumably to clear the damaged organelles. In addition, Bax/Bak-triggered autophagy reduced secretion of the pro-inflammatory cytokine interferon-β (IFN-β), thereby inhibiting a potential innate immune response to apoptotic cells.
Results
Bax/Bak activation induces autophagy independently of formation of the apoptosome
We previously demonstrated that inhibiting the pro-survival members of the Bcl-2 family not only leads to cell death but also activates LC3B lipidation and autophagic flux in a Bax/Bak-dependent manner (Fig. 1b) [25, 26]. However, the mechanism by which this occurs was unclear. To determine at what stage of the apoptotic pathway autophagy is induced (Fig. 1a), we activated Bax and Bak directly by the addition of the specific Bcl-2, Bcl-xL and Bcl-w inhibitor ABT-737 in mouse embryonic fibroblasts (MEFs) lacking Mcl-1, so that all the pro-survival proteins were inhibited (Fig. 1b) [18, 29]. We then tested whether inhibiting caspase activation with the pan-caspase inhibitor QVD-OPh would prevent LC3B lipidation (LC3B-I to LC3B-II). While QVD-OPh inhibited both the activating cleavage of caspase-3 and cell death after 4 h of treatment (Fig. 1b, Supplementary Fig. S1a), LC3B conversion from LC3B-I to LC3B-II remained unchanged (Fig. 1b). No further LC3B lipidation was seen in parallel experiments performed with cells lacking Bax and Bak (Fig. 1b) [25, 26]. Therefore, Bax/Bak activation can trigger LC3B conversion independently of caspase activity. LC3B lipidation was specific to mitochondrial-mediated apoptosis as induction of extrinsic apoptosis with TNF and the Smac mimetic Compound A did not stimulate LC3B-II formation (Supplementary Fig. S1b).
To complement the pharmacological approach, we activated Bax/Bak by over-expressing the BH3-only protein Bim using a doxycycline (Dox)-inducible vector. Again, caspase inhibition with QVD-OPh prevented cell death measured by propidium iodide (PI) uptake, but failed to prevent Bax/Bak from increasing LC3B-II formation (Fig. 1c, Supplementary Fig. S1c, d). To confirm that apoptotic caspases were not required for LC3B lipidation following Bax/Bak activation, we took a genetic approach and utilized cells lacking Apaf-1, a vital component of the apoptosome that activates caspase-9 and subsequently the downstream effector caspases, caspase-7 and caspase-3. As expected, Bim overexpression in Apaf-1−/− MEFs failed to induce caspase-3 cleavage, or PI uptake even after 16 h (Fig. 1d, Supplementary Fig. S1d). Despite this, LC3B-II lipidation still occurred within 4 h (Fig. 1d). We then deleted Mcl-1 using CRISPR/Cas9 technology from Apaf-1−/− MEFs and treated them with ABT-737. This also induced LC3B lipidation independently of PI uptake (Fig. 1e, Supplementary Fig. S1e). Therefore, induction of autophagy occurs early on in mitochondrial-mediated apoptosis pathway, independently of the apoptotic caspase cascade.
While LC3B lipidation is a useful marker of autophagy, we wanted to determine if the entire autophagy pathway was activated in an apoptosome-independent manner. We have previously shown that autophagic flux induced by both Bim and BH3 mimetics is strictly Bax/Bak-dependent in MEFs [25, 26]. To examine the requirement for apoptotic caspases, we used an mCherry-EGFP-LC3B marker, which reports on downstream autolysosome formation and acidification by a decrease in EGFP fluorescence (Fig. 2a). Under basal conditions, mCherry and EGFP fluorescence exhibited a linear correlation; however, when autophagy was stimulated by starving cells in amino-acid free Hank’s balanced salt solution (HBSS), EGFP fluorescence decreased as expected (Fig. 2b, c) [25]. When cells lacking Apaf-1 and Mcl-1 were treated with ABT-737 to induce Bax/Bak activation, EGFP fluorescence also decreased, while mCherry emission remained high (Fig. 2b, c), indicating that LC3B moved into autolysosomes. The drop in EGFP fluorescence could be prevented by the addition of the (auto)lysosomal inhibitor Bafilomycin A1 (BafA1) (Fig. 2b, c). Overexpression of Bim strongly induced autophagic flux, as measured by a drop of EGFP in cells lacking Apaf-1 (Fig. 2d, e). Therefore, autophagic flux induced by Bax/Bak activation occurs independently of the apoptosome and caspase activation.
a Schematic of the mCherry-EGFP-LC3B autophagic flux reporter. As LC3B is transported to the autolysosome via the autophagosome, EGFP fluorescence is inhibited due to a drop in pH. b Representative dot plots of Apaf-1−/− MEFs expressing a guide RNA against Mcl-1 and the mCherry-EGFP-LC3B autophagic flux reporter. Cells were treated overnight with 1 μM ABT-737, 0.1 μM bafilomycin A1 (BafA1), and/or starved by culturing them in HBSS prior to flow cytometry analysis. c Quantitation of (b) using two different clones of Apaf-1−/−ΔMcl-1 MEFs expressing the mCherry-EGFP-LC3B reporter (n = 7 experiments). d Representative dot plots of Apaf-1−/− MEFs expressing a Dox-inducible Bims and the mCherry-EGFP-LC3B autophagic flux reporter were treated overnight with 1 μg/mL Dox to express Bims, 0.1 μM bafilomycin A1 (BafA1), and/or starved by culturing them in HBSS. e Quantitation of (d) (n = 3 independent experiments). The mean is shown in all graphs; error bars represent the SEM
AMPK phosphorylates ULK1 following Bax/Bak activation
Because autophagy was stimulated independently of caspases, we sought to understand the pathway that links Bax/Bak activation to autophagic flux. To determine if the major regulators of autophagy were responsible, we treated cells lacking Mcl-1 with ABT-737 and immunoblotted for markers of mTORC1 activity (p-ULK1S757, p-S6S240/244 and p-4E-BP1S65) and AMPK activity (p-ULK1S317, p-ULK1S555 and p-AMPKT172) (Fig. 3a) [2, 3, 30]. Apaf-1−/− cells, in which the apoptosome cannot form, were used initially to ensure changes were not a consequence of apoptotic caspase activity.
a Schematic of the regulation of autophagy protein ULK1. AMPK activates ULK1 by phosphorylating both S317 and S555 on ULK1, while mTORC1 inhibits ULK1 activity by phosphorylating S757. b Apaf-1−/− MEFs with Mcl-1 deleted by CRISPR-Cas9 technology were treated with 1 μM ABT-737, 50 μM CCCP, or starved in HBSS for 4 h prior to lysis and western blotting. Identical aliquots of samples were run on four gels as follows: #1 (AMPK, p-ULK1S757), #2 (p-AMPKT172, p-ULK1S317, S6, β-actin), #3 (p-ULK1S555, 4E-BP1), #4 (p-S6S240/244, ULK1, p-4E-BP1S65, LC3B). c Apaf-1−/− MEFs were treated with 1 μg/mL Dox to express Bims, 50 μM CCCP, or starved in HBSS for 4 h prior to lysis and western blotting. Identical aliquots of samples were run on four gels as follows: #1 (LC3B, p-ULK1S757), #2 (p-AMPKT172, p-ULK1S317, 4E-BP, S6), #3 (p-4E-BP1S65, ULK1, AMPK), #4 (p-S6S240/244, p-ULK1S555, Bim, β-actin). See also Supplementary Fig. S2a. d AMPK activation is upstream of LC3B lipidation. Mcl-1−/− MEFs were infected with two different guide RNAs against ATG5 and selected for puromycin resistance. Pools were then treated with 1 μM ABT-737 for 4 h prior to western blotting. Identical amounts of sample were loaded on four different gels. e The intracellular ratio of AMP:ATP increases upon Bax/Bak-activation. MEFs were treated with either 1 μM ABT-737 or 1 μg/mL Dox to express Bims for 4 h and analysed by mass spectrometry. The mean of five to six experiments is shown; error bars represent the SEM
AMPK was robustly activated following Bax/Bak activation, and the autophagy protein ULK1 was phosphorylated by AMPK on both serine 317 and serine 555 (Fig. 3b). Similarly, Bim overexpression induced rapid ULK1 phosphorylation by AMPK (Fig. 3c). In contrast, the mTORC1 phosphorylation site on ULK1, serine 757, was unchanged by Bax/Bak activation, although other markers of mTORC1 activity, such as phosphorylation of S6 and 4E-BP1, were more variable (Fig. 3b, c, Supplementary Fig. S2a).
Interestingly, the activation of autophagy by Bax/Bak-activation had a distinct profile from that seen with either starvation-induced autophagy, or by treatment with carbonyl cyanide m-chlorophenyl hydrazone (CCCP), which induces mitophagy [11]. CCCP activated ULK1 by modulating both AMPK and mTORC1 signalling, while starvation in HBSS inhibited mTORC1 signalling under these experimental conditions (Fig. 3b, c). When ATG5 was deleted using CRISPR/Cas9 and two different guide RNAs, LC3B was no longer lipidated, but AMPK was still phosphorylated (Fig. 3d). This suggested that AMPK activation occurred either prior to or in parallel with formation of the apoptosome and caspase activation, and was upstream of autophagy induction.
However, when AMPK activity was inhibited by deletion of both AMPKβ1 and AMPKβ2, autophagic flux still occurred, indicating that AMPK is not the only regulator of Bax/Bak-stimulated autophagy (Supplementary Fig. S2b, c). Following Bax/Bak activation, mTORC1’s inhibitory phosphorylation of ULK1 was decreased in the AMPKβ 1/2−/− MEFs, but not in those with functional AMPK (Supplementary Fig. S2d, e). Therefore, AMPK appears to be the primary mechanism of the autophagy induction during Bax/Bak-activation, but mTORC1 can also contribute when AMPK is inactivated.
Because Bax/Bak activation disrupts mitochondrial membrane potential and therefore ATP production, we hypothesised that a drop in ATP levels was responsible for AMPK activation. We therefore measured total cellular [AMP]/[ATP] by mass spectrometry after 4 h treatment with ABT-737. The ratio of AMP to ATP increased approximately 4-fold compared to control cells, and occurred whether or not cells could activate caspases (Fig. 3e). A similar, but smaller, increase in [AMP]/[ATP] ratio was observed when Bim was induced with doxycycline (Fig. 3e). We suspect Bim caused a smaller change because induction of Bim required both transcription and translation prior to activation of Bax and Bak, whereas ABT-737 acted immediately.
Bax/Bak-activation induces the degradation of mitochondria by autophagy
PINK1 and Parkin are the best-characterised mediators of damage-induced mitophagy, a selective form of autophagy that degrades mitochondria [11, 12]. Because Bax/Bak activation damages the outer mitochondrial membrane, we hypothesised that this form of mitophagy would be stimulated. To investigate this possibility, we took advantage of the mitophagy marker mt-mKeima. This protein localises to the mitochondrial matrix and has a different excitation spectrum depending on pH, thereby indicating whether mitochondria have been delivered to the acidic lysosomal environment [31]. In a classic HeLa cell mitophagy model [14], treatment with the mitophagy inducers antimycin/oligomycin (A/O) caused mitochondria to be delivered to a more acidic environment in a Parkin-dependent manner, and the shift in excitation was repressed by the autolysosomal inhibitor BafA1, as expected (Supplementary Fig. S3a–c) [14, 31]. As with A/O treatment, Bax/Bak-activation caused by a combination of ABT-737 and the Mcl-1 inhibitor S63845 increased the acidic environment of mitochondria (Fig. 4a, b, Supplementary Fig. S3a–c). This shift was inhibited by the lysosomal inhibitor BafA1. However, unlike A/O-induced mitophagy, mitochondrial localisation to the lysosome did not require Parkin over-expression (Fig. 4a, b), although Parkin could increase the fluorescence shift (Supplementary Fig. S3a). Deletion of ATG5 inhibited both LC3B lipidation and mitochondrial acidification (Fig. 4a, b, Supplementary Fig. S3c), suggesting that some mitochondria were being trafficked to the lysosome in an autophagy-dependent manner.
a Bax/Bak activation stimulates mitochondria to move to the lysosome in ATG5-dependent but Parkin-independent manner. HeLa cells infected to over-express the mitophagy marker mt-mKeima where further infected to overexpress Parkin and/or to delete ATG5. Cells were treated with BH3 mimetics ABT-737 (1 μM) and S63845 (1 μM) to induce Bax/Bak activation in the presence of 10 μM QVD-OPh, and 0.1 μM BafA1 to inhibit autophagy for 16 h prior to analysis by flow cytometry (n = 3-4 independent experiments). b Representative flow cytometry dot plots and gating from a of HeLa cells with or without ATG5 expressing the mt-mKeima marker but without Parkin over-expression. c Apaf-1−/−ΔMcl-1 MEFs expressing the mt-mKeima mitophagy marker with or without over-expressed Parkin were treated for 8 h with mitophagy inducers 4 μM antimycin and 10 μM oligomycin (A/O) or 1 μM ABT-737 to induce Bax/Bak activation in the presence or absence of 0.1 μM BafA1 (n = 3 independent experiments). d Mcl-1−/− MEFs, with or without ATG7, were infected with the mt-mKeima marker. Cells were treated for 8 h with 1 μM ABT-737, 0.1 μM BafA1, and 10 μM QVD-OPh prior to analysis via flow cytometry (n = 3 independent experiments, using two independent guide RNAs—ATG7.113 and ATG7.160). e Mitochondria localise with LC3B puncta following Bax/Bak activation. Mcl-1−/− MEFs expressing GFP-LC3B and mCherry-Bak transmembrane domain (α9) to label mitochondria were treated with 1 μM ABT-737 and 10 μM QVD-OPh for 6 h prior to staining with DAPI and imaging. Representative of three independent experiments. Scale bar equals 10 μm. Also see Supplemental Fig. S4f. f Total mitochondrial load decreases after Bax/Bak activation. Mcl-1−/− MEFs were treated with 1 μM ABT-737, 0.1 μM BafA1, and 10 μM QVD-OPh overnight. Thirty minutes prior to harvesting, cells were treated with 0.1 μM of MitoTracker® Green FM and 0.1 μM MitoTracker® Deep Red FM and analysed at 488 nm and 633 nm, respectively. (n = 3 independent experiments) Graphs represent the mean, while error bars show the SEM in all graphs
Similar results were observed in fibroblasts. A/O treatment caused a BafA1-inhibitable mt-mKeima excitation shift and required Parkin (Fig. 4c, Supplementary Fig. S4a). While ABT-737 treatment was similarly suppressed by either BafA1 treatment or deletion of ATG7, it was independent of Parkin 1 (Fig. 4c, d, Supplementary Fig. S4a–c). Bim overexpression also induced Parkin-independent but ATG5- and ATG7-dependent mitochondrial localisation into lysosomes (Supplementary Fig. S4d, e).
Consistent with mt-mKeima data, mitochondria localised with GFP-LC3B puncta that formed following ABT-737 treatment (Fig. 4e, Supplementary Fig. S4f). To confirm mitochondria were being degraded, we activated Bax/Bak and measured mitochondrial load and membrane potential with MitoTracker® Green FM and Deep Red FM, respectively (Fig. 4f, Supplementary Fig. S4g). ABT-737 reduced both the mitochondrial membrane potential and mitochondrial load, the latter of which could be prevented by BafA1. Therefore, activation of Bax/Bak leads to mitochondria being degraded by autophagy.
Bax/Bak-activated autophagy suppresses type I interferon secretion
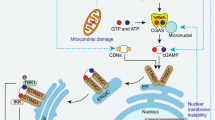
Bax/Bak-activation induces the pro-inflammatory cytokine IFN-β by releasing mitochondrial (mt)DNA and activating the cGAS/STING cytoplasmic DNA-sensing pathway (Fig. 5a). Apoptotic caspases suppress this pathway [32,33,34]. Because Bax/Bak-activation also induces autophagy and degrades mitochondria, we hypothesised that autophagy induced during apoptosis may also aid in limiting IFN-β secretion.
a Bax/Bak-activation activates the cGAS/STING pathway to induce IFN-β production, but the cytokine is degraded by apoptotic caspases. b Both autophagy genes and caspases inhibit IFN-β secretion following Bax/Bak-activation. Mcl-1−/− MEFs, those with CRISPR/Cas-9 deleted ATG5 or ATG7 (gRNA ATG5.299, ATG.405, ATG7.113 or ATG7.160), and Bax−/−Bak−/− MEFs were treated with 1 μM ABT-737 and 10 μM QVD-OPh as indicated. After 16 h, supernatants were harvested and IFN-β was measured by ELISA (n = 10–11 independent experiments). c IFN-β secretion after autophagy inhibition requires mitochondrial (mt) DNA. MEFs lacking Mcl-1 and either ATG5 or ATG7 were depleted of mtDNA (ρ0) and subsequently treated as in b (n = 4). d IL-6 secretion measured by ELISA in supernatants from b. e Autophagy inhibition does not increase Infb1 mRNA expression following ABT-737 treatment. Mcl-1−/− MEFs, and those with deleted ATG5 or ATG7, were treated for 4 h with 1 μM ABT-737 and/or 10 μM QVD-OPh prior to RNA purification. After cDNA synthesis, the fold change of ABT-737 treatment to untreated of Ifnb1/Gapdh was calculated (n = 3–13, using four different gRNAs independently: ATG5.299, ATG5.405, ATG7.113, and ATG7.160). f IFN-β is produced to similar extents irrespective of a functional canonical autophagy system. Cells were treated with 1 μM ABT-737 and/or 10 μM QVD-OPh for 4 h prior to lysis and immunoblotting. Representative of 3 experiments. g Apaf-1−/−ΔMcl-1 MEFs were treated with 1 μM ABT-737 for 4 h and subjected to sequential centrifugation as illustrated (left; adapted from ref. [38]). Samples were normalised to contain equal amount of total protein (30 μg), except for fractions 1–10 where the entire sample was loaded. Representative of three independent experiments. All graphs represent the mean and SEM
Addition of the caspase inhibitor QVD-OPh increased the levels of IFN-β in the supernatant of Mcl-1−/− MEFS induced to undergo apoptosis by treatment with ABT-737 (Fig. 5b), as expected [32]. When canonical autophagy was also inhibited by deletion of either ATG5 or ATG7 in addition to caspase inhibition (using cells from Fig. 3d and Supplementary Fig. S4c, respectively), IFN-β secretion increased five to sixfold (Fig. 5b). Indeed, following Bax/Bak activation, deletion of autophagy genes alone enhanced IFN-β secretion to a similar extent as caspase inhibition (Fig. 5b), suggesting the caspase and autophagy pathways suppress IFN-β secretion to similar extents.
To determine if IFN-β secretion depended on mtDNA for its production when autophagy was inhibited, we made mtDNA-depleted (ρ0) cells. Secretion of the cytokine was almost completely inhibited in ρ0 cells with or without caspase inhibition (Fig. 5c). Since mtDNA drives IFN-β secretion, we investigated if a difference in mtDNA levels was causing greater cytokine secretion in ATG5- or ATG7-deleted compared to parental cells; however, this was not the case (Supplementary Fig. S5a). In addition, ABT-737 induced cell death similarly in all cell lines (Supplementary Fig. S5b). Not all cytokine secretion is regulated by canonical autophagy during Bax/Bak-mediated apoptosis, however, as secretion of IL-6 was not greater in ATG5- or ATG7-deleted cells than parental ones (Fig. 5d).
As autophagy can reduce Infb1 mRNA in models of viral infection [35,36,37], we measured Infb1 mRNA levels following ABT-737 treatment. Surprisingly, inhibition of autophagy by deletion of ATG5 or ATG7 did not increase Infb1 mRNA levels (Fig. 5e), unlike caspase inhibition with QVD-OPh [32]. Consistent with this result, intercellular IFN-β protein levels were also similar between lines with deleted autophagy genes (Fig. 5f). Interestingly, chemical inhibition of autophagy by BafA1 did not produce as striking effects as deletion of ATG5 or ATG7 on IFN-β secretion (Fig. 5b, Supplementary Fig. S5c). These data suggest that ATG5 and ATG7 affect cytokine secretion rather than regulating its production, at least during the first few hours.
If this hypothesis is true, one would expect to find IFN-β associated with autophagosomes. Following sequential centrifugation to separate different organelles (Fig. 5g) [38], the majority of IFN-β and LC3B-II were both present in fraction 5 following the OptiPrep gradient. On the other hand, lysosomal (LAMP1), plasma membrane/endosome (TNFR1), nuclear (Histone H3), ER-Golgi (Calnexin), and mitochondrial (TIM44) markers, were also present in other fractions where IFN-β was absent, indicating that IFN-β were most likely associated with autophagosomes. Taken all together, our data suggest that autophagosomes sequester IFN-β to help keep apoptosis an immunologically silent.
Discussion
Programmed cell death can be used as a defensive strategy to limit replication of intracellular parasites such as viruses, and this may be why cell death mechanisms initially evolved. This is consistent with observations that cell death mechanisms can generate pro-inflammatory signals that aid in the immune response against infection. However, when apoptosis is used during development or to maintain homoeostasis of cell number, inflammation is undesirable. Here we describe a novel mechanism by which apoptotic cells prepare for death so that unwanted inflammation can be avoided. By inducing autophagy via AMPK (and under some circumstances through mTORC1 inhibition), apoptotic cells reduced secretion of the inflammatory cytokine IFN-β and degraded mitochondria (Fig. 6). We were surprised to find that Parkin was not essential for the degradation of mitochondria following Bax/Bak-activation. We believe that mitochondria are most likely non-selectively sequestered within autophagosomes, because even though alternative mitophagy pathways do occur, such as those employing NIX, BNIP3 or FUNDC1, these take place during reticulocyte maturation or hypoxia [39, 40]. Our results are consistent with previous findings showing that over-expression of Puma (a BH3 only protein) and Bax localise with LC3 on the mitochondria, and leads to the degradation of the mitochondrial proteins in a Bax/Bak-dependent manner [28].
Autophagic degradation of mitochondria following limited Bax/Bak activation could, under some circumstances, protect cells from unnecessary cell death [28, 41]. In laboratory settings, Bax/Bak are often activated by very strong stimuli that are sufficient to damage the vast majority of mitochondria. Conversely, minimal stresses might lead to moderate Bax/Bak activation and damage of only a few mitochondria, leading to subsequent ‘minority MOMP’ [42]. Cells can sometimes survive minority MOMP and it is possible Bax/Bak-stimulated autophagy may play a role in achieving this. Indeed, genetic abrogation of autophagy causes an accumulation of damaged mitochondria and increased sensitivity to cell death in erythroid and lymphoid cells [43, 44]. As the apoptosis machinery regulates the hematopoietic stem cell (HSC) compartment and autophagy is required for its maintenance [45, 46], it is possible that Bax/Bak-induced autophagy plays a role in haematopoiesis.
By degrading mitochondria, Bax/Bak-mediated autophagy could decrease the mtDNA available to produce an innate immune response. While this is the simplest explanation for how Bax/Bak-mediated autophagy limits IFN-β secretion, we suspect cGAS/STING activation occurs too quickly for degradation of mitochondria (and mtDNA) by autophagy to inhibit this early step effectively. Our data suggest that Bax/Bak-stimulated autophagy causes the cytokine to become sequestered. This is a distinct mechanism from that activated during viral infection. For instance, ATG9a deletion inhibited transcription of cGAS/STING-induced Infb1 following treatment with poly(dA-dT) dsDNA [35]. Similarly, Infb1 mRNA levels were increased in ATG5 or ATG7 knockouts upon exposure to viral dsRNA via the RIG-1 pathway [36, 37]. It remains to be determined if autophagosomes capture IFN-β by non-selective macroautophagy or selective autophagy.
Lastly, it is intriguing to speculate that the effect of Bax/Bak-induced autophagy on the immune response could be different in specialised cell types. Toll-like receptor (TLR)9, which recognises unmethylated CpG dinucleotides found in both bacterial and mitochondrial DNA, resides in endosomes (a type of vacuole that can fuse with autolysosomes) of B lymphocytes, monocytes, natural killer cells, and plasmacytoid dendritic cells [47]. Therefore it is possible that autophagy does not inhibit the IFN response during apoptosis of TLR9-expressing immune cells, but may even enhance it. It has been suggested that mtDNA-activated TLR9 may actually enhance autophagic flux [48, 49]. Consistent with this, TLR3 has been shown to sample apoptotic cells via endosomes after transfection with poly I:C, a model of viral infection [47, 50]. Our results suggest that even outside the immune system the links between cell death and inflammatory responses remain, but where an immune response is not needed, such as when cell death is used for development or to maintain homoeostasis of cell number, both autophagy and apoptosome-activated caspases suppress cytokine production and unwanted inflammation.
Materials and methods
Compounds
ABT-737 was purchased from Sellick, and QVD-OPh was from MP Biomedicals. Doxycycline, bafilomycin A1, carbonyl cyanide 3-chlorophenylhydrazone (CCCP), oligomycin A, and antimycin A were purchased from Sigma. The Mcl-1 inhibitor S63845 was synthesised as specified in US Patent Application No. 20150175623 (example 185) [17]. Compound A was a gift from TetraLogic Pharmaceuticals, while Fc-hTNF is made in house.
Cell lines and constructs
MEFs were immortalised with SV40 large T antigen and grown in Dulbecco’s modified Eagle’s medium supplemented with 8% foetal calf serum. Apaf-1−/−, Mcl-1−/−, AMPKβ 1/2−/−, and Bax−/−Bak−/− and wild-type MEFs have been previously described [25, 51,52,53]. The autophagic flux marker pBABE-puro/mCherry-EGFP-LC3B as described in [54], was obtained from AddGene (#22418). Parkin from pEGFP-parkin WT (AddGene #45875) was subcloned into pMX-IRES-GFP, and infected MEFs and ATG5-deleted HeLas [14] were sorted for GFP expression following infection. HeLas expressing pBMN-YFP-Parkin (AddGene #59416) and the retroviral pCHAC-mt-mKeima (AddGene #72342) have previously been described [14, 55].
Additional genes were deleted from MEFs by CRISPR/Cas9 technology using lentiCRISPR v2 (Addgene # 52961) and inserting guide RNA into BsmBI sites [56]. Cells were infected with viral vectors and selected with puromycin. For ATG5 and ATG7 deleted lines, pools of cells were used. For Mcl-1-deleted lines, cells were cloned after selection to insure sufficient Mcl-1 reduction. Guide sequences for MEFs are as follows:
Mcl-1.2 (GGGCGGCGGTTCTCCGGCAG) [57],
ATG5.299 (AAGATGTGCTTCGAGATGTG),
ATG5.405 (AAGAGTCAGCTATTTGACGT),
ATG7.113 (TTTAATAGTGCCCTGGACGT),
ATG7.160 (GAAGTTGAACGAGTACCGCC).
ATG5-deleted HeLa cells were made and described previously [14].
ATP and AMP concentrations
MEFs were seeded two days prior and treated for 4 h with doxycycline or ABT-737. Cells were washed in cold PBS and lysed in 0.5 M perchloric acid, clarified, and neutralised with 2.3 M KHCO3 prior to measurement by liquid chromatography mass spectrometry as described previously [58].
Western blots
Cells were treated as indicated prior to lysis in 20 mM Tris pH 7.5, 135 mM NaCl, 1.5 mM EDTA, 10% glycerol, 1% Triton X-100 including protease inhibitors (Roche, Dee Why, NSW, Australia) [25]. Equal amounts of protein were separated on 10% NUPAGE® Bis Tris gels (Invitrogen), transferred to nitrocellulose, and blotted with the antibodies raised against Mcl-1 (19C4; WEHI), Bim (3C5; WEHI), Bax (49F9-13-13; WEHI), Bak (Sigma), Parkin (sc-32282; Santa Cruz), LAMP1 (1D4B; Santa Cruz), TNFR1 (ab19139; Abcam), Calnexin (ab22595; Abcam), Apaf-1 (18H2; WEHI), TIMM44 (HPA043052; Sigma) and β-actin (AC-15; Sigma). The following antibodies were purchased from Cell Signalling: ATG5 (D5F5U), ATG7 (#2631) LC3B (D11), IFN-β (D2J1D), SQSTM1 (#5114), p-ULK1S317 (D2B6Y), p-ULK1S555 (D1H4), p-ULK1S757 (D7O6U), ULK1 (D8H5), p-S6S240/244 (D68F8), S6 (5G10), p-4E-BP1S65 (#9451), 4E-BP1 (53H11), cleaved caspase 3 (D175), Histone H3 (#9715), p-AMPKαT172 (40H9), AMPKβ (#4150) and AMPKα (#2532).
Autophagic flux assays were performed as previously described [9, 25, 26]. Briefly, cells were infected with pBABE-puro/mCherry-EGFP-LC3B and sorted prior to 16 h treatment with indicated treatments and processed for flow cytometry.
Mitophagy assays
Cells infected with pCHAC-mt-mKeima were treated as indicated to induce Bax/Bak activation for 16 h prior to analysis by flow cytometry [14, 31]. Mitochondrial load and potential were assayed with 0.1 μM of MitoTracker® Green FM and 0.1 μM MitoTracker® Deep Red FM, respectively, as per the manufacturer’s instructions. For confocal imaging, MEFs previously infected with the pBabe puro/EGFP-LC3 [59] and pMIH/mCherry-Bak-α9GGCK [60] were seeded in eight-well chamber μ-slides (Ibidi) and treated as indicated. Cells were subsequently fixed with 4% PFA in PBS for 10 min, washed with PBS, and stained with 4′,6-diamidine-2′-phenylindole dihydrochloride (DAPI; ThermoFisher) for 10 min. Images were obtained using a Zeiss LSM 780 microscope; with Z-stacks obtained using a 63x oil objective and acquired with ZEN software (Zeiss). Images were merged using FIJI software.
ELISAs
MEFs lacking Mcl-1 (20 000 cells) were seeded in 24-well plates. The next day, cells in 0.5 mL were treated as indicated for 16 h. After spinning plates to pellet cells, 0.25 ml supernatant were collected and stored at −80 °C. Supernatant samples were thawed at room temperature and IFN-β ELISAs were performed with 50 μL sample using the VeriKine-HSTM Mouse IFN Beta Serum ELISA Kit (PBL Assay Science) as per the manufacturer’s instructions. IL-6 ELISAs were performed on the same supernatents using the Mouse IL-6 ELISA Ready-SET-Go!® kit (eBioscience).
Rho (ρ0) cell production
Mitochondrial (mt)DNA-depleted cells were produced as previously reported [32]. Briefly, cells were cultured in 100 ng/ml ethidium bromide in media supplemented with 50 μg/ml uridine and 100 μg/ml sodium pyruvate for 7-8 days prior to seeding for experiments. Twice as many ethidium bromide-treated cells were seeded compared to controls as depleted cells had a greatly reduced division rate. Mitochondrial DNA depletion was confirmed by qPCR as previously described [32].
Autophagosome purification
Autophagosome enrichment was performed essentially as described [38]. Briefly, 10 15-cm dishes of Apaf-1−/−ΔMcl-1 MEFs were treated with 1 μM ABT-737 to induce Bax/Bak activation for 4 h. Cells were scraped, washed in ice cold PBS and homogenised through a 27-G needle in H Buffer [20 mM HEPES-KOH pH 7.2, 400 mM sucrose, 1 mM EDTA, protease inhibitors]. Lysate was sequentially centrifuged at 1000×g (10 min), and then at 3000×g (10 min), 25,000×g (20 min), and 100,000×g (30 min) at 4 °C in a TLA100.3 rotor (Beckman). 25,000×g pellet was resuspended in 0.75 ml 1.25 M sucrose buffer and overlayed with 0.5 mL 1.1 M and 0.25 M sucrose buffer (in 10 mM Tris-HCl pH 7.4). After centrifugation at 120,000 xg for 2 h at 4 °C in a TLS 55 rotor (Beckman), the pellet and light fraction, which is visible by eye, were collected. The light fraction was diluted in 110 mM Tris-HCl pH 7.4 to dilute sucrose and the pellet was collected following a 100,000×g spin for 1 h (SW40 Ti; Beckman). The pellet was resuspended in a 2.5 ml 19% OptiPrep (D1556; SIGMA) for a step gradient containing 1.25 ml 22.5%, 2.5 ml 19% (sample), 2.25 ml 16%, 2.25 ml 12%, 2.5 ml 8%, 1.25 ml 5% and 0.5 ml 0% OptiPrep each in buffer O (20 mM Tricine-KOH, pH 7.4, 250 mM sucrose, 1 mM EDTA). After centrifugation at 150,000×g for 3 h in a SW40 Ti rotor, 1.25 mL fractions were taken from the top and diluted in B88 buffer (20 mM HEPES-KOH, pH 7.2, 250 mM sorbitol, 150 mM potassium acetate, 5 mM magnesium acetate) to pellet samples at 100,000×g for 1 h (SW40 Ti). All samples were resuspended in loading buffer.
References
Klionsky DJ, Abdelmohsen K, Abe A, Abedin MJ, Abeliovich H, Acevedo Arozena A, et al. Guidelines for the use and interpretation of assays for monitoring autophagy (3rd edition. Autophagy. 2016;12:1–222.
Egan DF, Shackelford DB, Mihaylova MM, Gelino S, Kohnz RA, Mair W, et al. Phosphorylation of ULK1 (hATG1) by AMP-activated protein kinase connects energy sensing to mitophagy. Science. 2011;331:456–61.
Kim J, Kundu M, Viollet B, Guan KL. AMPK and mTOR regulate autophagy through direct phosphorylation of Ulk1. Nat Cell Biol. 2011;13:132–41.
Gwinn DM, Shackelford DB, Egan DF, Mihaylova MM, Mery A, Vasquez DS, et al. AMPK phosphorylation of raptor mediates a metabolic checkpoint. Mol Cell. 2008;30:214–26.
Nguyen TN, Padman BS, Usher J, Oorschot V, Ramm G, Lazarou M. Atg8 family LC3/GABARAP proteins are crucial for autophagosome-lysosome fusion but not autophagosome formation during PINK1/Parkin mitophagy and starvation. J Cell Biol. 2016;215:857–74.
Abeliovich H, Dunn WA Jr., Kim J, Klionsky DJ. Dissection of autophagosome biogenesis into distinct nucleation and expansion steps. J Cell Biol. 2000;151:1025–34.
Nakatogawa H, Ichimura Y, Ohsumi Y. Atg8, a ubiquitin-like protein required for autophagosome formation, mediates membrane tethering and hemifusion. Cell. 2007;130:165–78.
Mizushima N, Noda T, Yoshimori T, Tanaka Y, Ishii T, George MD, et al. A protein conjugation system essential for autophagy. Nature. 1998;395:395–8.
Mizushima N, Yoshimori T, Levine B. Methods in mammalian autophagy research. Cell. 2010;140:313–26.
Klionsky DJ, Elazar Z, Seglen PO, Rubinsztein DC. Does bafilomycin A1 block the fusion of autophagosomes with lysosomes? Autophagy. 2008;4:849–50.
Narendra D, Tanaka A, Suen DF, Youle RJ. Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J Cell Biol. 2008;183:795–803.
Narendra DP, Jin SM, Tanaka A, Suen DF, Gautier CA, Shen J, et al. PINK1 is selectively stabilized on impaired mitochondria to activate Parkin. PLoS Biol. 2010;8:e1000298.
Vives-Bauza C, Zhou C, Huang Y, Cui M, de Vries RL, Kim J, et al. PINK1-dependent recruitment of Parkin to mitochondria in mitophagy. Proc Natl Acad Sci USA. 2010;107:378–83.
Lazarou M, Sliter DA, Kane LA, Sarraf SA, Wang C, Burman JL, et al. The ubiquitin kinase PINK1 recruits autophagy receptors to induce mitophagy. Nature. 2015;524:309–14.
Lalaoui N, Lindqvist LM, Sandow JJ, Ekert PG. The molecular relationships between apoptosis, autophagy and necroptosis. Semin Cell Dev Biol. 2015;39:63–9.
Wei MC, Zong WX, Cheng EH, Lindsten T, Panoutsakopoulou V, Ross AJ, et al. Proapoptotic BAX and BAK: a requisite gateway to mitochondrial dysfunction and death. Science. 2001;292:727–30.
Kotschy A, Szlavik Z, Murray J, Davidson J, Maragno AL, Le Toumelin-Braizat G, et al. The MCL1 inhibitor S63845 is tolerable and effective in diverse cancer models. Nature. 2016;538:477–82.
van Delft MF, Wei AH, Mason KD, Vandenberg CJ, Chen L, Czabotar PE, et al. The BH3 mimetic ABT-737 targets selective Bcl-2 proteins and efficiently induces apoptosis via Bak/Bax if Mcl-1 is neutralized. Cancer Cell. 2006;10:389–99.
Lindqvist LM, Simon AK, Baehrecke EH. Current questions and possible controversies in autophagy. Cell Death Discov. 2015;1. 10.1038/cddiscovery.2015.36.
Pattingre S, Tassa A, Qu X, Garuti R, Liang XH, Mizushima N, et al. Bcl-2 antiapoptotic proteins inhibit Beclin 1-dependent autophagy. Cell. 2005;122:927–39.
Pedro JM, Wei Y, Sica V, Maiuri MC, Zou Z, Kroemer G, et al. BAX and BAK1 are dispensable for ABT-737-induced dissociation of the BCL2-BECN1 complex and autophagy. Autophagy. 2015;11:452–9.
Wei Y, Pattingre S, Sinha S, Bassik M, Levine B. JNK1-mediated phosphorylation of Bcl-2 regulates starvation-induced autophagy. Mol Cell. 2008;30:678–88.
Maiuri MC, Criollo A, Tasdemir E, Vicencio JM, Tajeddine N, Hickman JA, et al. BH3-only proteins and BH3 mimetics induce autophagy by competitively disrupting the interaction between Beclin 1 and Bcl-2/Bcl-X(L). Autophagy. 2007;3:374–6.
Liang XH, Kleeman LK, Jiang HH, Gordon G, Goldman JE, Berry G, et al. Protection against fatal Sindbis virus encephalitis by beclin, a novel Bcl-2-interacting protein. J Virol. 1998;72:8586–96.
Lindqvist LM, Heinlein M, Huang DC, Vaux DL. Prosurvival Bcl-2 family members affect autophagy only indirectly, by inhibiting Bax and Bak. Proc Natl Acad Sci USA. 2014;111:8512–7.
Reljic B, Conos S, Lee EF, Garnier JM, Dong L, Lessene G, et al. BAX-BAK1-independent LC3B lipidation by BH3 mimetics is unrelated to BH3 mimetic activity and has only minimal effects on autophagic flux. Autophagy. 2016;12:1083–93.
Lindqvist LM, Vaux DL. BCL2 and related prosurvival proteins require BAK1 and BAX to affect autophagy. Autophagy. 2014;10:1474–5.
Yee KS, Wilkinson S, James J, Ryan KM, Vousden KH. PUMA- and Bax-induced autophagy contributes to apoptosis. Cell Death Differ. 2009;16:1135–45.
Oltersdorf T, Elmore SW, Shoemaker AR, Armstrong RC, Augeri DJ, Belli BA, et al. An inhibitor of Bcl-2 family proteins induces regression of solid tumours. Nature. 2005;435:677–81.
Laplante M, Sabatini DM. mTOR signaling in growth control and disease. Cell 2012;149:274–93.
Katayama H, Kogure T, Mizushima N, Yoshimori T, Miyawaki A. A sensitive and quantitative technique for detecting autophagic events based on lysosomal delivery. Chem Biol. 2011;18:1042–52.
White MJ, McArthur K, Metcalf D, Lane RM, Cambier JC, Herold MJ, et al. Apoptotic Caspases Suppress mtDNA-Induced STING-Mediated Type I IFN Production. Cell. 2014;159:1549–62.
Rongvaux A, Jackson R, Harman CC, Li T, West AP, de Zoete MR, et al. Apoptotic caspases prevent the induction of type I interferons by mitochondrial DNA. Cell. 2014;159:1563–77.
Cai X, Chiu YH, Chen ZJ. The cGAS-cGAMP-STING pathway of cytosolic DNA sensing and signaling. Mol Cell. 2014;54:289–96.
Saitoh T, Fujita N, Hayashi T, Takahara K, Satoh T, Lee H, et al. Atg9a controls dsDNA-driven dynamic translocation of STING and the innate immune response. Proc Natl Acad Sci USA. 2009;106:20842–6.
Jounai N, Takeshita F, Kobiyama K, Sawano A, Miyawaki A, Xin KQ, et al. The Atg5 Atg12 conjugate associates with innate antiviral immune responses. Proc Natl Acad Sci USA. 2007;104:14050–5.
Tal MC, Sasai M, Lee HK, Yordy B, Shadel GS, Iwasaki A. Absence of autophagy results in reactive oxygen species-dependent amplification of RLR signaling. Proc Natl Acad Sci USA. 2009;106:2770–5.
Ge L, Melville D, Zhang M, Schekman R. The ER-Golgi intermediate compartment is a key membrane source for the LC3 lipidation step of autophagosome biogenesis. Elife. 2013;2:e00947.
Bernardini JP, Lazarou M, Dewson G. Parkin and mitophagy in cancer. Oncogene. 2016;36:1315–27.
Rogov V, Dotsch V, Johansen T, Kirkin V. Interactions between autophagy receptors and ubiquitin-like proteins form the molecular basis for selective autophagy. Mol Cell. 2014;53:167–78.
Colell A, Ricci JE, Tait S, Milasta S, Maurer U, Bouchier-Hayes L, et al. GAPDH and autophagy preserve survival after apoptotic cytochrome c release in the absence of caspase activation. Cell. 2007;129:983–97.
Ichim G, Lopez J, Ahmed SU, Muthalagu N, Giampazolias E, Delgado ME, et al. Limited mitochondrial permeabilization causes DNA damage and genomic instability in the absence of cell death. Mol Cell. 2015;57:860–72.
Mortensen M, Ferguson DJ, Edelmann M, Kessler B, Morten KJ, Komatsu M, et al. Loss of autophagy in erythroid cells leads to defective removal of mitochondria and severe anemia in vivo. Proc Natl Acad Sci USA. 2010;107:832–7.
Pua HH, Guo J, Komatsu M, He YW. Autophagy is essential for mitochondrial clearance in mature T lymphocytes. J Immunol. 2009;182:4046–55.
Mortensen M, Soilleux EJ, Djordjevic G, Tripp R, Lutteropp M, Sadighi-Akha E, et al. The autophagy protein Atg7 is essential for hematopoietic stem cell maintenance. J Exp Med. 2011;208:455–67.
Orelio C, Dzierzak E. Bcl-2 expression and apoptosis in the regulation of hematopoietic stem cells. Leuk Lymphoma. 2007;48:16–24.
Stetson DB, Medzhitov R. Type I interferons in host defense. Immunity. 2006;25:373–81.
De Leo MG, Staiano L, Vicinanza M, Luciani A, Carissimo A, Mutarelli M, et al. Autophagosome-lysosome fusion triggers a lysosomal response mediated by TLR9 and controlled by OCRL. Nat Cell Biol. 2016;18:839–50.
Lindqvist LM. Can the cargo control the car? Mitochondrial DNA as a stimulator of TLR9-mediated autophagosome-lysosome fusion. Cell Death Differ. 2016;23:1737–8.
Schulz O, Diebold SS, Chen M, Naslund TI, Nolte MA, Alexopoulou L, et al. Toll-like receptor 3 promotes cross-priming to virus-infected cells. Nature. 2005;433:887–92.
Willis SN, Chen L, Dewson G, Wei A, Naik E, Fletcher JI, et al. Proapoptotic Bak is sequestered by Mcl-1 and Bcl-xL, but not Bcl-2, until displaced by BH3-only proteins. Genes Dev. 2005;19:1294–305.
Marsden VS, O’Connor L, O’Reilly LA, Silke J, Metcalf D, Ekert PG, et al. Apoptosis initiated by Bcl-2-regulated caspase activation independently of the cytochrome c/Apaf-1/caspase-9 apoptosome. Nature. 2002;419:634–7.
Dite TA, Ling NXY, Scott JW, Hoque A, Galic S, Parker BL, et al. The autophagy initiator ULK1 sensitizes AMPK to allosteric drugs. Nat Commun. 2017;8:571.
N’Diaye EN, Kajihara KK, Hsieh I, Morisaki H, Debnath J, Brown EJ. PLIC proteins or ubiquilins regulate autophagy-dependent cell survival during nutrient starvation. EMBO Rep. 2009;10:173–9.
Yamano K, Fogel AI, Wang C, van der Bliek AM, Youle RJ. Mitochondrial Rab GAPs govern autophagosome biogenesis during mitophagy. Elife. 2014;3:e01612.
Sanjana NE, Shalem O, Zhang F. Improved vectors and genome-wide libraries for CRISPR screening. Nat Methods. 2014;11:783–4.
Aubrey BJ, Kelly GL, Kueh AJ, Brennan MS, O’Connor L, Milla L, et al. An inducible lentiviral guide RNA platform enables the identification of tumor-essential genes and tumor-promoting mutations in vivo. Cell Rep. 2015;10:1422–32.
Scott JW, Galic S, Graham KL, Foitzik R, Ling NX, Dite TA, et al. Inhibition of AMP-activated protein kinase at the allosteric drug-binding site promotes islet insulin release. Chem Biol. 2015;22:705–11.
Fung C, Lock R, Gao S, Salas E, Debnath J. Induction of autophagy during extracellular matrix detachment promotes cell survival. Mol Biol Cell. 2008;19:797–806.
Iyer S, Bell F, Westphal D, Anwari K, Gulbis J, Smith BJ, et al. Bak apoptotic pores involve a flexible C-terminal region and juxtaposition of the C-terminal transmembrane domains. Cell Death Differ. 2015;22:1665–75.
Acknowledgements
We thank J. Bernadini, S. Iyer, I. Tan, and G. Dewson (WEHI) for helpful discussions and technical assistance. S63845 was a kind gift from C. Burns (WEHI). Funding for this project was provided by NHMRC Program Grants 461221 (D.L.V.) and 1016647 (B.T.K.) and Project Grant 1106471 (M.L.). L.M.L. was supported by an NHMRC Peter Doherty Early Career Fellowship 1035502, and D.L.V. and B.T.K. held NHMRC Fellowships (1020136 and 1063008, respectively). J.S.O. (130100988) and M.L. (1601100063) are ARC Future Fellows and K.M. is supported by an Australian Federal Government Postgraduate Award. This work was made possible through Independent Research Institutes Infrastructure Support Scheme Grant (361646) from the Australian National Health and Medical Research Council and a Victorian State Government Operational Infrastructure Support Grant.
Author contributions
L.M.L., D.F., K.M. and T.A.D. performed the experiments; L.M.L., D.F., K.M., J.S.O. and B.T.K. designed the experiments; and L.M.L., K.M., M.L. and B.T.K. analysed the data. L.M.L., B.T.K. and D.L.V. wrote the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
D.L.V. used to be on the scientific advisory board for TetraLogic Pharmaceuticals.
Additional information
Edited by S. Kumar
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Lindqvist, L.M., Frank, D., McArthur, K. et al. Autophagy induced during apoptosis degrades mitochondria and inhibits type I interferon secretion. Cell Death Differ 25, 784–796 (2018). https://doi.org/10.1038/s41418-017-0017-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41418-017-0017-z
This article is cited by
-
Exposure of the inner mitochondrial membrane triggers apoptotic mitophagy
Cell Death & Differentiation (2024)
-
Mechanism and role of mitophagy in the development of severe infection
Cell Death Discovery (2024)
-
Mitochondrial outer membrane integrity regulates a ubiquitin-dependent and NF-κB-mediated inflammatory response
The EMBO Journal (2024)
-
Molecular mechanisms of mitochondrial DNA release and activation of the cGAS-STING pathway
Experimental & Molecular Medicine (2023)
-
Mitochondrial control of inflammation
Nature Reviews Immunology (2023)