Abstract
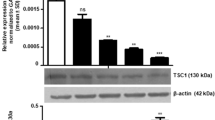
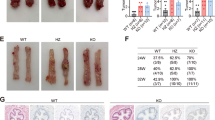
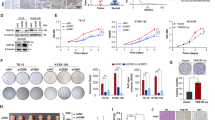
Mutations in ARID2 and TP53 genes are found to be implicated in the tobacco related tumorigeneses. However, the effect of loss of ARID2 in the TP53 mutated background in tobacco related cancer including oral cancer has not been investigated yet. Hence, in this study we knockdown ARID2 using shRNA mediated knockdown strategy in TP53 mutated oral squamous cell carcinoma (OSCC) cell line and studied its tumorigenic role. Our study revealed that suppression of ARID2 in TP53 mutated oral cancer cells increases cell motility and invasion, induces drastic morphological changes and leads to a marked increase in the expression levels of cytokeratins, and integrins, CK8, CK18 and β4-Integrin, markers of cell migration/invasion in oral cancer. ARID2 suppression also showed early onset and increased tumorigenicity in-vivo. Interestingly, transcriptome profiling revealed differentially expressed genes associated with migration and invasion in oral cancer cells including AKR1C2, NCAM2, NOS1, ADAM23 and genes of S100A family in ARID2 knockdown TP53 mutated oral cancer cells. Pathway analysis of differentially regulated genes identified “cancer pathways” and “PI3K/AKT Pathway” to be significantly dysregulated upon suppression of ARID2 in TP53 mutated OSCC cells. Notably, decreased ARID2 expression and increased CK8, CK18 expression leads to poor prognosis in Head and Neck cancer (HNSC) patients as revealed by Pan-Cancer TCGA data analysis. To conclude, our study is the first to demonstrate tumor suppressor role of ARID2 in TP53 mutated background indicating their cooperative role in OSCC, and also highlights its prognostic implications suggesting ARID2 as an important therapeutic target in OSCC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
Most data have been incorporated into the article and its online supplementary material. Remaining data are available on reasonable request to corresponding author.
References
Shain AH, Pollack JR. The spectrum of SWI/SNF mutations, ubiquitous in human cancers. PLoS One. 2013;8:e55119.
Cohet N, Stewart KM, Mudhasani R, Asirvatham AJ, Mallappa C, Imbalzano KM, et al. SWI/SNF chromatin remodeling enzyme ATPases promote cell proliferation in normal mammary epithelial cells. J Cell Physiol. 2010;223:667–78.
Imbalzano AN, Kwon H, Green MR, Kingston RE. Facilitated binding of TATA-binding protein to nucleosomal DNA. Nature 1994;370:481–5.
Kwon H, Imbalzano AN, Khavari PA, Kingston RE, Green MR. Nucleosome disruption and enhancement of activator binding by a human SW1/SNF complex. Nature 1994;370:477–81.
Yan Z, Cui K, Murray DM, Ling C, Xue Y, Gerstein A, et al. PBAF chromatin-remodeling complex requires a novel specificity subunit, BAF200, to regulate expression of selective interferon-responsive genes. Genes Dev. 2005;19:1662–7.
Olivier M, Hollstein M, Hainaut P. TP53 mutations in human cancers: origins, consequences, and clinical use. Cold Spring Harb Perspect Biol. 2010;2:a001008.
Loesch R, Chenane L, Colnot S. ARID2 chromatin remodeler in hepatocellular carcinoma. Cells 2020;9:2152.
Manceau G, Letouzé E, Guichard C, Didelot A, Cazes A, Corté H, et al. Recurrent inactivating mutations of ARID2 in non-small cell lung carcinoma. Int J Cancer. 2013;132:2217–21.
Maitra A, Biswas N, Amin K, Kowtal P, Kumar S, Das S, et al. Mutational landscape of gingivo-buccal oral squamous cell carcinoma reveals new recurrently-mutated genes and molecular subgroups. Nat Commun. 2013;4:2873.
Fadlullah MZ, Chiang IK, Dionne KR, Yee PS, Gan CP, Sam KK, et al. Genetically-defined novel oral squamous cell carcinoma cell lines for the development of molecular therapies. Oncotarget 2016;7:27802–18.
Pansare K, Gardi N, Kamat S, Dange P, Previn R, Gera P, et al. Establishment and genomic characterization of gingivobuccal carcinoma cell lines with smokeless tobacco associated genetic alterations and oncogenic PIK3CA mutation. Sci Rep. 2019;9:8272.
Silwal-Pandit L, Vollan HK, Chin SF, Rueda OM, McKinney S, Osako T, et al. TP53 mutation spectrum in breast cancer is subtype specific and has distinct prognostic relevance. Clin Cancer Res. 2014;20:3569–80.
Gong F, Fahy D, Liu H, Wang W, Smerdon MJ. Role of the mammalian SWI/SNF chromatin remodeling complex in the cellular response to UV damage. Cell Cycle. 2008;7:1067–74.
Wu RC, Wang TL, Shih IM. The emerging roles of ARID1A in tumor suppression. Cancer Biol Ther. 2014;15:655–64.
Cajuso T, Hänninen UA, Kondelin J, Gylfe AE, Tanskanen T, Katainen R, et al. Exome sequencing reveals frequent inactivating mutations in ARID1A, ARID1B, ARID2, and ARID4A in microsatellite unstable colorectal cancer. Int J Cancer. 2014;135:611–23.
Kadoch C, Hargreaves DC, Hodges C, et al. Proteomic and bioinformatic analysis of mammalian swi/snf complexes identifies extensive roles in human malignancy. Nat Genet. 2013;45:592–601.
Duan Y, Tian L, Gao Q, Liang L, Zhang W, Yang Y, et al. Chromatin remodeling gene ARID2 targets cyclin D1 and cyclin E1 to suppress hepatoma cell progression. Oncotarget 2016;7:45863–75.
Jiang H, Cao HJ, Ma N, Bao WD, Wang JJ, Chen TW, et al. Chromatin remodeling factor ARID2 suppresses hepatocellular carcinoma metastasis via DNMT1-Snail axis. Proc Natl Acad Sci USA. 2020;117:4770–80.
Zhang L, Wang W, Li X, He S, Jianfeng Y, Wang X, et al. MicroRNA-155 promotes tumor growth of human hepatocellular carcinoma by targeting ARID2. Int J Oncol. 2016;48:2425–34.
Liu X, Zhang H, Zhou P, Yu Y, Zhang H, Chen L, et al. CREB1 acts via the miR-922/ARID2 axis to enhance malignant behaviour of liver cancer cells. Oncol Rep. 2021;45:79.
Zhou C, Zhang Y, Yan R, Huang L, Mellor AL, Yang Y, et al. Exosome-derived miR-142-5p remodels lymphatic vessels and induces IDO to promote immune privilege in the tumour microenvironment. Cell Death Differ. 2021;28:715–29.
Wu M, Duan Q, Liu X, Zhang P, Fu Y, Zhang Z, et al. MiR-155-5p promotes oral cancer progression by targeting chromatin remodeling gene ARID2. Biomed Pharmacother. 2020;122:109696.
Moreno T, Monterde B, González-Silva L, Betancor-Fernández I, Revilla C, Agraz-Doblas A, et al. ARID2 deficiency promotes tumor progression and is associated with higher sensitivity to chemotherapy in lung cancer. Oncogene 2021;40:2923–35.
Fukumoto T, Lin J, Fatkhutdinov N, Liu P, Somasundaram R, Herlyn M, et al. ARID2 deficiency correlates with the response to immune checkpoint blockade in melanoma. J Invest Dermatol. 2020;S0022-202X:32396–4.
Goh AY, Ramlogan-Steel CA, Jenkins KS, Steel JC, Layton CJ. Presence and prevalence of UV related genetic mutations in uveal melanoma: similarities with cutaneous melanoma. Neoplasma 2020;67:958–71.
Williams EA, Shah N, Montesion M, Sharaf R, Pavlick DC, Sokol ES, et al. Melanomas with activating RAF1 fusions: clinical, histopathologic, and molecular profiles. Mod Pathol. 2020;33:1466–74.
Saqcena M, Leandro-Garcia LJ, Maag JLV, Tchekmedyian V, Krishnamoorthy GP, Tamarapu. SWI/SNF complex mutations promote thyroid tumor progression and insensitivity to redifferentiation therapies. Cancer Disco. 2020;11:1158–75.
Oba A, Shimada S, Akiyama Y, Nishikawaji T, Mogushi K, Ito H, et al. ARID2 modulates DNA damage response in human hepatocellular carcinoma cells. J Hepatol. 2017;66:942–51.
Pan D, Kobayashi A, Jiang P, Ferrari de Andrade L, Tay RE, Luoma AM, et al. A major chromatin regulator determines resistance of tumor cells to T cell-mediated killing. Sci. 2018;359:770–5.
Fuchs E, Cleveland DW. A structural scaffolding of intermediate filaments in health and disease. Sci. 1998;279:514–9.
Paramio JM, Jorcano JL. Beyond structure: do intermediate filaments modulate cell signalling? Bioessays 2002;24:836–44.
Moll R, Divo M, Langbein L. The human keratins: biology and pathology. Histochem Cell Biol. 2008;129:705–33.
Vaidya MM, Borges AM, Pradhan SA, Bhisey AN. Cytokeratin expression in squamous cell carcinomas of the tongue and alveolar mucosa. Eur J Cancer B Oral Oncol. 1996;32B:333–6.
Borradori L, Sonnenberg A. Hemidesmosomes: roles in adhesion, signalling and human diseases. Curr Opin Cell Biol 1996;8:647–56.
Yang Y, Dowling J, Yu QC, Kouklis P, Cleveland DW, Fuchs E. An essential cytoskeletal linker protein connecting actin microfilaments to intermediate filaments. Cell 1996;86:655–65.
Giancotti FG. Targeting integrin beta4 for cancer and anti-angiogenic therapy. Trends Pharmacol Sci 2007;28:506–11.
Alam H, Kundu ST, Dalal SN, Vaidya MM. Loss of keratins 8 and 18 leads to alterations in α6β4-integrin-mediated signalling and decreased neoplastic progression in an oral-tumour-derived cell line. J Cell Sci. 2011;124:2096–106.
Kippenberger S, Hofmann M, Zöller N, Thaçi D, Müller J, Kaufmann R, et al. A. Ligation of beta4 integrins activates PKB/Akt and ERK1/2 by distinct pathways-relevance of the keratin filament. Biochem Biophys Acta. 2010;1803:940–50.
Simbolo M, Vicentini C, Ruzzenente A, Brunelli M, Conci S, Fassan M, et al. Genetic alterations analysis in prognostic stratified groups identified TP53 and ARID1A as poor clinical performance markers in intrahepatic cholangiocarcinoma. Sci Rep. 2018;8:7119.
Zhang J, Hou S, You Z, Li G, Xu S, Li X, et al. Expression and prognostic values of ARID family members in breast cancer. Aging (Albany NY). 2021;13:5621–37.
Afrem M, CrăiŢoiu Ş, Hîncu M, Manolea H, Nicolae V, CrăiŢoiu M. Study of CK18 and GDF5 immunoexpression in oral squamous cell carcinoma and their prognostic value. Rom J Morphol Embryol. 2016;57:167–72.
Nagata M, Noman A, Suzuki K, Kurita H, Ohnishi M, Ohyama T, et al. ITGA3 and ITGB4 expression biomarkers estimate the risks of locoregional and hematogenous dissemination of oral squamous cell carcinoma. BMC Cancer. 2013;13:410.
Choudhari SK, Chaudhary M, Bagde S, Gadbail AR, Joshi V. Nitric oxide and cancer: a review. World J Surg Oncol. 2013;11:118.
Raab JR, Resnick S, Magnuson T. Genome-wide transcriptional regulation mediated by biochemically distinct SWI/SNF complexes. PLoS Genet. 2015;11:e1005748.
Pfister NT, Fomin V, Regunath K, Zhou JY, Zhou W, Silwal-Pandit L, et al. Mutant p53 cooperates with the SWI/SNF chromatin remodeling complex to regulate VEGFR2 in breast cancer cells. Genes Dev. 2015;29:1298–315.
Song H, Hollstein M, Xu Y. p53 gain-of-function cancer mutants induce genetic instability by inactivating ATM. Nat Cell Biol. 2007;9:573–80.
Acknowledgements
We would like to thank Genomics & NGS facility, electron microscopy facility, animal House, biorepository, ACTREC, Dr. Pradnya Kowtal and members of Sarin Lab. We are also grateful to TCGA/ULACAN databases to make data available free for analysis. We also thank NIBMG, and grant support from Department of Biotechnology, Govt. of India (Grant Number (BT/MB/01/ICGC/2009). We are grateful to Dr. T. Mahmoudi, Netherland for providing ARID2 antibody and Dr. Sorab Dalal, ACTREC, Mumbai for providing pLKO-PURO EGFP vector. The study has been supported by grant from Department of Biotechnology, Govt. of India (Grant Number (BT/MB/01/ICGC/2009) under the International Cancer Genome Consortium (ICGC) project.
Author information
Authors and Affiliations
Contributions
PS conceptualized the idea, conducted cloning and molecular biology experiments, TCGA data analysis, designed and wrote the manuscript. PD performed cell culture and all functional assays and helped in animal experiments. PC supervised BSM in conducting all animal experiments. NG analyzed bioinformatics data. RS conceived the project, supervised all the experiments and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Shukla, P., Dange, P., Mohanty, B.S. et al. ARID2 suppression promotes tumor progression and upregulates cytokeratin 8, 18 and β-4 integrin expression in TP53-mutated tobacco-related oral cancer and has prognostic implications. Cancer Gene Ther 29, 1908–1917 (2022). https://doi.org/10.1038/s41417-022-00505-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41417-022-00505-x