Abstract
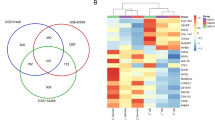
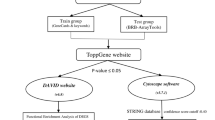
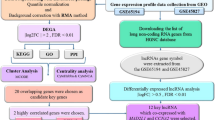
Triple-negative breast cancer (TNBC) has a high degree of malignancy, lack of effective diagnosis and treatment, and poor prognosis. Bioinformatics methods are used to screen the hub genes and signal pathways involved in the progress of TNBC to provide reliable biomarkers for the diagnosis and treatment of TNBC. Download the raw data of four TNBC-related datasets from the Gene Expression Omnibus (GEO) database and use them for bioinformatics analysis. GEO2R tool was used to analyze and identify differentially expressed (DE) mRNAs. DAVID database was used to carry out gene ontology (GO) analysis and Kyoto Encyclopedia of Genes and Genome Pathways (KEGG) signal pathway enrichment analysis for DE mRNAs. STRING database and Cytoscape were used to build DE mRNAs protein-protein interaction (PPI) network diagram and visualize PPI network, respectively. Through cytoHubba, cBioPortal database, Kaplan-Meier mapper database, Gene Expression Profiling Interactive Analysis (GEPIA) Database, UALCAN Database, The Cancer Genome Atlas (TCGA) database, Tumor Immunity Estimation Resource identify hub genes. Perform qRT-PCR, Human Protein Atlas analysis, mutation analysis, survival analysis, clinical-pathological characteristics, and infiltrating immune cell analysis. 22 DE mRNAs were identified from the four datasets, including 16 upregulated DE mRNAs and six downregulated DE mRNAs. Enrichment analysis of the KEGG showed that DE mRNAs were principally enriched in pathways in cancer, mismatch repair, cell cycle, platinum drug resistance, breast cancer. Six hub genes were screened based on the PPI network diagram of DE mRNAs. Survival analysis found that TOP2A, CCNA2, PCNA, MSH2, CDK6 are related to the prognosis of TNBC. In addition, mutations, clinical indicators, and immune infiltration analysis show that these five hub genes play an important role in the progress of TNBC and immune monitoring. Compared with MCF-10A, MCF-7, and SKBR-3 cells, TOP2A, PCNA, MSH2, and CDK6 were significantly upregulated in MDA-MB-321 cells. Compared with normal, luminal, and Her-2 positive tissues, CCNA2, MSH2, and CDK6 were significantly upregulated in TNBC. Through comparative analysis of GEO datasets related to colorectal cancer and lung adenocarcinoma, it was determined that these five hub genes were unique differentially expressed genes of TNBC. At last, the hub genes related to the progression, prognosis, and immunity of TNBC have been successfully screened. They are indeed specific to TNBC as prognostic features. They can be used as potential markers for the prognosis of TNBC and provide potential therapeutic targets.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout










Similar content being viewed by others
Data availability
Additional data pertaining to this work can be obtained from the corresponding author upon request.
Change history
01 July 2022
A Correction to this paper has been published: https://doi.org/10.1038/s41417-022-00498-7
References
Goldhirsch A, Winer EP, Coates AS, Gelber RD, Piccart-Gebhart M, Thürlimann B, et al. Personalizing the treatment of women with early breast cancer: highlights of the St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2013. Ann Oncol. 2013;24:2206–23.
Yin L, Duan JJ, Bian XW, Yu SC. Triple-negative breast cancer molecular subtyping and treatment progress. Breast Cancer Res. 2020;22:61.
Sung H, Ferlay J, Siegel RL, Laversanne M, Soerjomataram I, Jemal A, et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2021;71:209–49.
Mills MN, Yang GQ, Oliver DE, Liveringhouse CL, Ahmed KA, Orman AG, et al. Histologic heterogeneity of triple negative breast cancer: A National Cancer Centre Database analysis. Eur J Cancer. 2018;98:48–58.
Lehmann BD, Bauer JA, Chen X, Sanders ME, Chakravarthy AB, Shyr Y, et al. Identification of human triple-negative breast cancer subtypes and preclinical models for selection of targeted therapies. J Clin Invest. 2011;121:2750–67.
Dong P, Yu B, Pan L, Tian X, Liu F. Identification of key genes and pathways in triple-negative breast cancer by integrated bioinformatics analysis. Biomed Res Int. 2018;2018:2760918.
Zhong G, Lou W, Shen Q, Yu K, Zheng Y. Identification of key genes as potential biomarkers for triple‑negative breast cancer using integrating genomics analysis. Mol Med Rep. 2020;21:557–66.
Shah SP, Roth A, Goya R, Oloumi A, Ha G, Zhao Y, et al. The clonal and mutational evolution spectrum of primary triple-negative breast cancers. Nature. 2012;486:395–9.
Bianchini G, Balko JM, Mayer IA, Sanders ME, Gianni L. Triple-negative breast cancer: challenges and opportunities of a heterogeneous disease. Nat Rev Clin Oncol. 2016;13:674–90.
Kamps R, Brandão RD, Bosch BJ, Paulussen AD, Xanthoulea S, Blok MJ, et al. Next-generation sequencing in oncology: genetic diagnosis, risk prediction and cancer classification. Int J Mol Sci 2017;18:308.
Morgan AJ, Giannoudis A, Palmieri C. The genomic landscape of breast cancer brain metastases: a systematic review. Lancet Oncol. 2021;22:e7–e17.
Vagia E, Mahalingam D, Cristofanilli M. The landscape of targeted therapies in TNBC. Cancers (Basel) 2020;12:916.
Maire V, Baldeyron C, Richardson M, Tesson B, Vincent-Salomon A, Gravier E, et al. TTK/hMPS1 is an attractive therapeutic target for triple-negative breast cancer. PLoS ONE. 2013;8:e63712.
Wang L, Shen X, Xie B, Ma Z, Chen X, Cao F. Transcriptional profiling of differentially expressed long non-coding RNAs in breast cancer. Genom Data. 2015;6:214–6.
Cascione L, Gasparini P, Lovat F, Carasi S, Pulvirenti A, Ferro A, et al. Integrated microRNA and mRNA signatures associated with survival in triple negative breast cancer. PLoS ONE. 2013;8:e55910.
Komatsu M, Yoshimaru T, Matsuo T, Kiyotani K, Miyoshi Y, Tanahashi T, et al. Molecular features of triple negative breast cancer cells by genome-wide gene expression profiling analysis. Int J Oncol. 2013;42:478–506.
Gene Ontology Consortium. The Gene Ontology resource: enriching a GOld mine. Nucleic Acids Res 2021;49: D325-D334.
Kanehisa M, Sato Y, Kawashima M, Furumichi M, Tanabe M. KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res. 2016;44:D457–62.
Chin CH, Chen SH, Wu HH, Ho CW, Ko MT, Lin CY. cytoHubba: identifying hub objects and sub-networks from complex interactome. BMC Syst Biol. 2014;8:S11.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO, et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal. 2013;6:pl1.
Györffy B, Lanczky A, Eklund AC, Denkert C, Budczies J, Li Q, et al. An online survival analysis tool to rapidly assess the effect of 22,277 genes on breast cancer prognosis using microarray data of 1,809 patients. Breast Cancer Res Treat. 2010;123:725–31.
Lánczky A, Nagy Á, Bottai G, Munkácsy G, Szabó A, Santarpia L, et al. miRpower: a web-tool to validate survival-associated miRNAs utilizing expression data from 2178 breast cancer patients. Breast Cancer Res Treat. 2016;160:439–46.
Tang Z, Li C, Kang B, Gao G, Li C, Zhang Z. GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Res. 2017;45:W98–w102.
Chandrashekar DS, Bashel B, Balasubramanya SAH, Creighton CJ, Ponce-Rodriguez I, Chakravarthi B, et al. UALCAN: a portal for facilitating tumor subgroup gene expression and survival analyses. Neoplasia. 2017;19:649–58.
Uhlén M, Fagerberg L, Hallström BM, Lindskog C, Oksvold P, Mardinoglu A, et al. Proteomics. Tissue-based map of the human proteome. Science. 2015;347:1260419.
Uhlen M, Zhang C, Lee S, Sjöstedt E, Fagerberg L, Bidkhori G, et al. A pathology atlas of the human cancer transcriptome. Science 2017;357:eaan2507.
Li T, Fan J, Wang B, Traugh N, Chen Q, Liu JS, et al. TIMER: a web server for comprehensive analysis of tumor-infiltrating immune cells. Cancer Res. 2017;77:e108–e110.
Li B, Severson E, Pignon JC, Zhao H, Li T, Novak J, et al. Comprehensive analyses of tumor immunity: implications for cancer immunotherapy. Genome Biol. 2016;17:174.
Canela A, Maman Y, Huang SN, Wutz G, Tang W, Zagnoli-Vieira G, et al. Topoisomerase II-induced chromosome breakage and translocation is determined by chromosome architecture and transcriptional activity. Mol Cell. 2019;75:252–e8.
Seoane JA, Kirkland JG, Caswell-Jin JL, Crabtree GR, Curtis C. Chromatin regulators mediate anthracycline sensitivity in breast cancer. Nat Med. 2019;25:1721–7.
Kawale AS, Povirk LF. Tyrosyl-DNA phosphodiesterases: rescuing the genome from the risks of relaxation. Nucleic Acids Res. 2018;46:520–37.
Lu Y, Su F, Yang H, Xiao Y, Zhang X, Su H, et al. E2F1 transcriptionally regulates CCNA2 expression to promote triple negative breast cancer tumorigenicity. Cancer Biomark. 2022;33:57–70.
Guo Y, Gabola M, Lattanzio R, Paul C, Pinet V, Tang R, et al. Cyclin A2 maintains colon homeostasis and is a prognostic factor in colorectal cancer. J Clin Invest 2021;131:e131517.
Yu YL, Chou RH, Liang JH, Chang WJ, Su KJ, Tseng YJ, et al. Targeting the EGFR/PCNA signaling suppresses tumor growth of triple-negative breast cancer cells with cell-penetrating PCNA peptides. PLoS ONE. 2013;8:e61362.
Cardano M, Tribioli C, Prosperi E. Targeting proliferating cell nuclear antigen (PCNA) as an effective strategy to inhibit tumor cell proliferation. Curr Cancer Drug Targets. 2020;20:240–52.
Aebi S, Fink D, Gordon R, Kim HK, Zheng H, Fink JL, et al. Resistance to cytotoxic drugs in DNA mismatch repair-deficient cells. Clin Cancer Res. 1997;3:1763–7.
Dasgupta H, Islam S, Alam N, Roy A, Roychoudhury S, Panda CK. Hypomethylation of mismatch repair genes MLH1 and MSH2 is associated with chemotolerance of breast carcinoma: clinical significance. J Surg Oncol. 2019;119:88–100.
Jiang YZ, Liu Y, Xiao Y, Hu X, Jiang L, Zuo WJ, et al. Molecular subtyping and genomic profiling expand precision medicine in refractory metastatic triple-negative breast cancer: the FUTURE trial. Cell Res. 2021;31:178–86.
Bellayr IH, Marklein RA, Lo Surdo JL, Bauer SR, Puri RK. Identification of predictive gene markers for multipotent stromal cell proliferation. Stem Cells Dev. 2016;25:861–73.
Le DT, Uram JN, Wang H, Bartlett BR, Kemberling H, Eyring AD, et al. PD-1 blockade in tumors with mismatch-repair deficiency. New Engl J Med. 2015;372:2509–20.
Fremd C, Hlevnjak M, Zapatka M, Zoernig I, Halama N, Fejzibegovic N, et al. Mismatch repair deficiency drives durable complete remission by targeting programmed death receptor 1 in a metastatic luminal breast cancer patient. Breast Care (Basel). 2019;14:53–59.
Mills AM, Dill EA, Moskaluk CA, Dziegielewski J, Bullock TN, Dillon PM. The relationship between mismatch repair deficiency and PD-L1 expression in breast carcinoma. Am J Surg Pathol. 2018;42:183–91.
Slamon DJ, Neven P, Chia S, Fasching PA, De Laurentiis M, Im SA, et al. Phase III randomized study of ribociclib and fulvestrant in hormone receptor-positive, human epidermal growth factor receptor 2-negative advanced breast cancer: MONALEESA-3. J Clin Oncol. 2018;36:2465–72.
Hsu YH, Yao J, Chan LC, Wu TJ, Hsu JL, Fang YF, et al. Definition of PKC-α, CDK6, and MET as therapeutic targets in triple-negative breast cancer. Cancer Res. 2014;74:4822–35.
Dickler MN, Tolaney SM, Rugo HS, Cortés J, Diéras V, Patt D, et al. MONARCH 1, a phase II study of abemaciclib, a CDK4 and CDK6 inhibitor, as a single agent, in patients with refractory hr(+)/her2(-) metastatic breast cancer. Clin Cancer Res. 2017;23:5218–24.
Dieci MV, Miglietta F, Guarneri V. Immune infiltrates in breast cancer: recent updates and clinical implications. Cells 2021;10:223.
Author information
Authors and Affiliations
Contributions
JM, ZF and LR conceived and designed experiments. JM performed experiments. CC and SL carried out data analysis. DW, JJ and PH, DW participated in the preparation of reagents/materials/analysis tools. JM wrote the manuscript. ZF and LR supervised the manuscript. All data were generated in-house, and no paper mill was used. All authors agree to be accountable for all aspects of this work including the integrity and accuracy of the data presented.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Ma, J., Chen, C., Liu, S. et al. Identification of a five genes prognosis signature for triple-negative breast cancer using multi-omics methods and bioinformatics analysis. Cancer Gene Ther 29, 1578–1589 (2022). https://doi.org/10.1038/s41417-022-00473-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41417-022-00473-2
This article is cited by
-
The crucial prognostic signaling pathways of pancreatic ductal adenocarcinoma were identified by single-cell and bulk RNA sequencing data
Human Genetics (2024)
-
Characterization of protein-based risk signature to predict prognosis and evaluate the tumor immune environment in breast cancer
Breast Cancer (2023)