Abstract
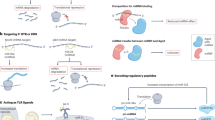
Gastric cancer is the third leading cause of cancer-related death worldwide, with relapse and metastasis being major contributors to the mortality. Circular RNAs (circRNAs) have been at the center of several researches and some circRNAs have been indicated to be involved in gastric cancer as sponges. Nevertheless, the mechanism underlying the function of circRNA remains largely unclear. Therefore, this study was conducted with the main objective of screening the associated circRNA in gastric cancer and exploring its mechanism. Expression of hsa_circRNA_0009172 was validated in gastric cancer tissues and cell lines after the correlation between hsa_circRNA_0009172 and prognosis was determined. Moreover, the binding site between miR-485-3p and hsa_circRNA_0009172 or NTRK3 was verified using dual luciferase assay and RNA pull down. Function-gain and -loss experiments were performed for the purpose of detecting the effect of hsa_circRNA_0009172 in vivo and in vitro as well as its mechanism with microRNA (miRNA)-485-3p and NTRK3 in gastric cancer. The hsa_circRNA_0009172 expression was downregulated in gastric cancer tissues and cell lines, indicating a positive association with patient prognosis. Functionally, hsa_circ_0009172 overexpression inhibited proliferative, invasive and migrative potential of gastric cancer cells as well as epithelial-mesenchymal transition (EMT)-related proteins by sponging miR-485-3p to inhibit NTRK3, while miR-485-3p overexpression could reverse the inhibitory effect of hsa_circ_0009172 on gastric cancer. Furthermore, either up-regulation of hsa_circ_0009172 or down-regulation of miR-485-3p led to the suppression of xenograft tumor growth in nude mice. In conclusion, hsa_circ_0009172 serves as a tumor suppressor in gastric cancer by targeting miR-485-3p/NTRK3 axis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
Data availability
The datasets generated/analyzed during the current study are available.
References
Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018;68:394–424.
Thrumurthy SG, Chaudry MA, Chau I, Allum W. Does surgery have a role in managing incurable gastric cancer? Nat Rev Clin Oncol. 2015;12:676–82.
Van Cutsem E, Sagaert X, Topal B, Haustermans K, Prenen H. Gastric cancer. Lancet. 2016;388:2654–64.
Dongre A, Weinberg RA. New insights into the mechanisms of epithelial-mesenchymal transition and implications for cancer. Nat Rev Mol Cell Biol. 2019;20:69–84.
Huang L, Wu RL, Xu AM. Epithelial-mesenchymal transition in gastric cancer. Am J Transl Res. 2015;7:2141–58.
Kristensen LS, Andersen MS, Stagsted LVW, Ebbesen KK, Hansen TB, Kjems J. The biogenesis, biology and characterization of circular RNAs. Nat Rev Genet. 2019;20:675–91.
Shan C, Zhang Y, Hao X, Gao J, Chen X, Wang K. Biogenesis, functions and clinical significance of circRNAs in gastric cancer. Mol Cancer. 2019;18:136.
Shang BQ, Li ML, Quan HY, Hou PF, Li ZW, Chu SF, et al. Functional roles of circular RNAs during epithelial-to-mesenchymal transition. Mol Cancer. 2019;18:138.
Tian M, Chen R, Li T, Xiao B. Reduced expression of circRNA hsa_circ_0003159 in gastric cancer and its clinical significance. J Clin Lab Anal. 2018;32:e22281.
Zhang Y, Liu H, Li W, Yu J, Li J, Shen Z, et al. CircRNA_100269 is downregulated in gastric cancer and suppresses tumor cell growth by targeting miR-630. Aging (Albany NY). 2017;9:1585–94.
Zhang J, Liu H, Hou L, Wang G, Zhang R, Huang Y, et al. Circular RNA_LARP4 inhibits cell proliferation and invasion of gastric cancer by sponging miR-424-5p and regulating LATS1 expression. Mol Cancer. 2017;16:151.
Lamouille S, Subramanyam D, Blelloch R, Derynck R. Regulation of epithelial-mesenchymal and mesenchymal-epithelial transitions by microRNAs. Curr Opin Cell Biol. 2013;25:200–7.
Song S, Ajani JA. The role of microRNAs in cancers of the upper gastrointestinal tract. Nat Rev Gastroenterol Hepatol. 2013;10:109–18.
Lou C, Xiao M, Cheng S, Lu X, Jia S, Ren Y, et al. MiR-485-3p and miR-485-5p suppress breast cancer cell metastasis by inhibiting PGC-1alpha expression. Cell Death Dis. 2016;7:e2159.
Muinos-Gimeno M, Guidi M, Kagerbauer B, Martin-Santos R, Navines R, Alonso P, et al. Allele variants in functional MicroRNA target sites of the neurotrophin-3 receptor gene (NTRK3) as susceptibility factors for anxiety disorders. Hum Mutat. 2009;30:1062–71.
Wood LD, Calhoun ES, Silliman N, Ptak J, Szabo S, Powell SM, et al. Somatic mutations of GUCY2F, EPHA3, and NTRK3 in human cancers. Hum Mutat. 2006;27:1060–1.
Shi E, Chmielecki J, Tang CM, Wang K, Heinrich MC, Kang G, et al. FGFR1 and NTRK3 actionable alterations in “Wild-Type” gastrointestinal stromal tumors. J Transl Med. 2016;14:339.
Bu JY, Lv WZ, Liao YF, Xiao XY, Lv BJ. Long non-coding RNA LINC00978 promotes cell proliferation and tumorigenesis via regulating microRNA-497/NTRK3 axis in gastric cancer. Int J Biol Macromol. 2019;123:1106–14.
Luo Y, Kaz AM, Kanngurn S, Welsch P, Morris SM, Wang J, et al. NTRK3 is a potential tumor suppressor gene commonly inactivated by epigenetic mechanisms in colorectal cancer. PLoS Genet. 2013;9:e1003552.
Xiong D, Sheng Y, Ding S, Chen J, Tan X, Zeng T, et al. LINC00052 regulates the expression of NTRK3 by miR-128 and miR-485-3p to strengthen HCC cells invasion and migration. Oncotarget. 2016;7:47593–608.
Padmanabhan N, Ushijima T, Tan P. How to stomach an epigenetic insult: the gastric cancer epigenome. Nat Rev Gastroenterol Hepatol. 2017;14:467–78.
Bang YJ, Van Cutsem E, Feyereislova A, Chung HC, Shen L, Sawaki A, et al. Trastuzumab in combination with chemotherapy versus chemotherapy alone for treatment of HER2-positive advanced gastric or gastro-oesophageal junction cancer (ToGA): a phase 3, open-label, randomised controlled trial. Lancet. 2010;376:687–97.
Tang W, Fu K, Sun H, Rong D, Wang H, Cao H. CircRNA microarray profiling identifies a novel circulating biomarker for detection of gastric cancer. Mol Cancer. 2018;17:137.
Yin Y, Long J, He Q, Li Y, Liao Y, He P, et al. Emerging roles of circRNA in formation and progression of cancer. J Cancer. 2019;10:5015–21.
Vo JN, Cieslik M, Zhang Y, Shukla S, Xiao L, Zhang Y, et al. The Landscape of Circular RNA in. Cancer Cell. 2019;176:869–81 e13.
Fang X, Wen J, Sun M, Yuan Y, Xu Q. CircRNAs and its relationship with gastric cancer. J Cancer. 2019;10:6105–13.
Peng Z, Wang CX, Fang EH, Wang GB, Tong Q. Role of epithelial-mesenchymal transition in gastric cancer initiation and progression. World J Gastroenterol. 2014;20:5403–10.
Naeli P, Pourhanifeh MH, Karimzadeh MR, Shabaninejad Z, Movahedpour A, Tarrahimofrad H, et al. Circular RNAs and gastrointestinal cancers: Epigenetic regulators with a prognostic and therapeutic role. Crit Rev Oncol Hematol. 2020;145:102854.
Liu H, Liu Y, Bian Z, Zhang J, Zhang R, Chen X, et al. Circular RNA YAP1 inhibits the proliferation and invasion of gastric cancer cells by regulating the miR-367-5p/p27 (Kip1) axis. Mol Cancer. 2018;17:151.
Deng G, Mou T, He J, Chen D, Lv D, Liu H, et al. Circular RNA circRHOBTB3 acts as a sponge for miR-654-3p inhibiting gastric cancer growth. J Exp Clin Cancer Res. 2020;39:1.
Wang J, Zhang H, Zhou X, Wang T, Zhang J, Zhu W, et al. Five serum-based miRNAs were identified as potential diagnostic biomarkers in gastric cardia adenocarcinoma. Cancer Biomark. 2018;23:193–203.
Dai D, Tan Y, Guo L, Tang A, Zhao Y. Identification of exosomal miRNA biomarkers for diagnosis of papillary thyroid cancer by small RNA sequencing. Eur J Endocrinol. 2020;182:111–21.
Cocco E, Scaltriti M, Drilon A. NTRK fusion-positive cancers and TRK inhibitor therapy. Nat Rev Clin Oncol. 2018;15:731–47.
Wada R, Arai H, Kure S, Peng WX, Naito Z. “Wild type” GIST: Clinicopathological features and clinical practice. Pathol Int. 2016;66:431–7.
Kubo T, Kuroda Y, Shimizu H, Kokubu A, Okada N, Hosoda F, et al. Resequencing and copy number analysis of the human tyrosine kinase gene family in poorly differentiated gastric cancer. Carcinogenesis. 2009;30:1857–64.
Acknowledgements
We would like to give our sincere appreciation to the reviewers for their helpful comments on this article.
Author information
Authors and Affiliations
Contributions
Hao Wang, Nan Wang, Xiaoli Zheng, Lei Wu, Chengcheng Fan, Xue Li, Ke Ye, and Suxia Han designed the study. Hao Wang, Nan Wang, and Xiaoli Zheng collated the data, carried out data analyses, and produced the initial draft of the manuscript. Lei Wu, Chengcheng Fan, Xue Li, Ke Ye, and Suxia Han contributed to drafting the manuscript. All authors have read and approved the final submitted manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Wang, H., Wang, N., Zheng, X. et al. Circular RNA hsa_circ_0009172 suppresses gastric cancer by regulation of microRNA-485-3p-mediated NTRK3. Cancer Gene Ther 28, 1312–1324 (2021). https://doi.org/10.1038/s41417-020-00280-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41417-020-00280-7