Abstract
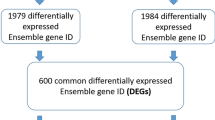
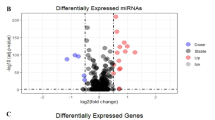
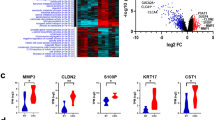
Doublecortin-like kinase 1 (DCLK1) is a cancer stem cell marker for the colorectal cancer (CRC). It plays critical roles in the oncogenesis, progression, and metastasis of CRC. DCLK1 can be an intriguing therapeutic target for CRC treatment. However, the molecular mechanism of how DCLK1 functions is unclear currently. In our research, we aim to apply RNA-Sequencing (RNA-Seq) technology, a high throughput massively Next-generation sequencing approach, to monitor transcriptome changes due to DCLK1 overexpression in the CRC cells. In order to achieve our goal, RNA from quadruplicate samples from two clones of isogenic DCLK1 stable overexpression cells and the parental wild-type HCT116 cells was sent for RNA-Seq on the Illumina NextSeq500 platform. Differentially expressed (DE) genes were evaluated by t-test (P < 0.05 and fold-change ± 1.5 or greater) using two methods: (1) FWER; and (2) Benjamani and Hochberg FDR (false discovery rate) which corrects for multiple comparisons. Gene networks and functional analysis were evaluated using Ingenuity Pathways Analysis (IPA). We identified 1463 DE genes common for both DCLK1 overexpression clone A and clone B cells. IPA results indicated that 72 canonical pathways were significantly modified by DCLK1 overexpression (P < 0.05), among which nine out of the top ten pathways are involved in the cell cycle regulation, indicating that DCLK1 might play its tumorigenesis role via activation of pathways facilitating cell proliferation, repression of pathways inhibiting cells proliferation, and function against pathways facilitating cell apoptosis. Cell cycle analysis results confirmed the IPA findings, which demonstrated that DCLK1 overexpression cells had much less G0/G1 cells but much more S and G2/M cells (P < 0.05). In conclusion, DCLK1 overexpression significantly modified transcriptome profile of CRC cancer cells. Control of the cell cycle regulation might be one of the critical mechanism for DCLK1 function. Our findings provide more direct evidence for the development of DCLK1 as a therapeutic target for CRC treatment, and will be of great benefit for the discovery of novel therapeutic target within the DCLK1 molecular network for the treatment of colorectal cancer patients.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
De Dosso S, Sessa C, Saletti P. Adjuvant therapy for colon cancer: present and perspectives. Cancer Treat Rev. 2009;35:160–6.
Yamakage K, Omori Y, Piccoli C, Yamasaki H. Growth control of 3T3 fibroblast cell lines established from connexin 43-deficient mice. Mol Carcinog. 1998;23:121–8.
Omori Y, Suzuki M, Ozaki K, Harada Y, Nakamura Y, Takahashi E, et al. Expression and chromosomal localization of KIAA0369, a putative kinase structurally related to doublecortin. J Hum Genet. 1998;43:169–77.
May R, Riehl TE, Hunt C, Sureban SM, Anant S, Houchen CW. Identification of a novel putative gastrointestinal stem cell and adenoma stem cell marker, doublecortin and CaM kinase-like-1, following radiation injury and in adenomatous polyposis coli/multiple intestinal neoplasia mice. Stem Cells. 2008;26:630–7.
Nakanishi Y, Seno H, Fukuoka A, Ueo T, Yamaga Y, Maruno T, et al. Dclk1 distinguishes between tumor and normal stem cells in the intestine. Nat Genet. 2013;45:98–103.
Gagliardi G, Goswami M, Passera R, Bellows CF. DCLK1 immunoreactivity in colorectal neoplasia. Clin Exp Gastroenterol. 2012;5:35–42.
Arruda-Olson AM, Bursi F, Gerber Y, May RH, Roger VL, Pellikka PA. Three-dimensional echocardiography for evaluating left ventricular function in patients with ST elevation myocardial infarction: a pilot study. Mayo Clin Proc. 2008;83:372–3.
Chandrakesan P, Weygant N, May R, Qu D, Chinthalapally HR, Sureban SM, et al. DCLK1 facilitates intestinal tumor growth via enhancing pluripotency and epithelial mesenchymal transition. Oncotarget. 2014;5:9269–80.
Westphalen CB, Asfaha S, Hayakawa Y, Takemoto Y, Lukin DJ, Nuber AH, et al. Long-lived intestinal tuft cells serve as colon cancer-initiating cells. J Clin Investig. 2014;124:1283–95.
Kantara C, O’Connell MR, Luthra G, Gajjar A, Sarkar S, Ullrich RL, et al. Methods for detecting circulating cancer stem cells (CCSCs) as a novel approach for diagnosis of colon cancer relapse/metastasis. Lab Investig. 2015;95:100–12.
Mirzaei A, Tavoosidana G, Modarressi MH, Rad AA, Fazeli MS, Shirkoohi R, et al. Upregulation of circulating cancer stem cell marker, DCLK1 but not Lgr5, in chemoradiotherapy-treated colorectal cancer patients. Tumour Biol. 2015;36:4801–10.
Sakaguchi M, Hisamori S, Oshima N, Sato F, Shimono Y, Sakai Y. miR-137 regulates the tumorigenicity of colon cancer stem cells through the inhibition of DCLK1. Mol Cancer Res. 2016;14:354–62.
Vega KJ, May R, Sureban SM, Lightfoot SA, Qu D, Reed A, et al. Identification of the putative intestinal stem cell marker doublecortin and CaM kinase-like-1 in Barrett’s esophagus and esophageal adenocarcinoma. J Gastroenterol Hepatol. 2012;27:773–80.
Whorton J, Sureban SM, May R, Qu D, Lightfoot SA, Madhoun M, et al. DCLK1 is detectable in plasma of patients with Barrett’s esophagus and esophageal adenocarcinoma. Dig Dis Sci. 2015;60:509–13.
Sureban SM, May R, Lightfoot SA, Hoskins AB, Lerner M, Brackett DJ, et al. DCAMKL-1 regulates epithelial-mesenchymal transition in human pancreatic cells through a miR-200a-dependent mechanism. Cancer Res. 2011;71:2328–38.
Bailey JM, Alsina J, Rasheed ZA, McAllister FM, Fu YY, Plentz R, et al. DCLK1 marks a morphologically distinct subpopulation of cells with stem cell properties in preinvasive pancreatic cancer. Gastroenterology. 2014;146:245–56.
Rao CV, Mohammed A. New insights into pancreatic cancer stem cells. World J Stem Cells. 2015;7:547–55.
Qu D, Johnson J, Chandrakesan P, Weygant N, May R, Aiello N, et al. Doublecortin-like kinase 1 is elevated serologically in pancreatic ductal adenocarcinoma and widely expressed on circulating tumor cells. PLoS ONE. 2015;10:e0118933.
Ali N, Allam H, Bader T, May R, Basalingappa KM, Berry WL, et al. Fluvastatin interferes with hepatitis C virus replication via microtubule bundling and a doublecortin-like kinase-mediated mechanism. PLoS ONE. 2013;8:e80304.
Sureban SM, Madhoun MF, May R, Qu D, Ali N, Fazili J, et al. Plasma DCLK1 is a marker of hepatocellular carcinoma (HCC): Targeting DCLK1 prevents HCC tumor xenograft growth via a microRNA-dependent mechanism. Oncotarget. 2015;6:37200–15.
Wang W, Zhang H, Wang L, Zhang S, Tang M. miR-613 inhibits the growth and invasiveness of human hepatocellular carcinoma via targeting DCLK1. Biochem Biophys Res Commun. 2016;473:987–92.
Sureban SM, May R, Mondalek FG, Qu D, Ponnurangam S, Pantazis P, et al. Nanoparticle-based delivery of siDCAMKL-1 increases microRNA-144 and inhibits colorectal cancer tumor growth via a Notch-1 dependent mechanism. J Nanobiotechnol. 2011;9:40.
Sureban SM, May R, Qu D, Weygant N, Chandrakesan P, Ali N, et al. DCLK1 regulates pluripotency and angiogenic factors via microRNA-dependent mechanisms in pancreatic cancer. PLoS ONE. 2013;8:e73940.
Deng H, Qianqian G, Ting J, Aimin Y. miR-539 enhances chemosensitivity to cisplatin in non-small cell lung cancer by targeting DCLK1. Biomed Pharmacother. 2018;106:1072–81.
Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods. 2001;25:402–8.
Li L, Jones K, Mei H. Doublecotin-like kinase 1 increases chemoresistance of colorectal cancer cells through the anti-apoptosis pathway. J Stem Cell Res Ther. 2019;9:1–7.
Weygant N, Qu D, Berry WL, May R, Chandrakesan P, Owen DB, et al. Small molecule kinase inhibitor LRRK2-IN-1 demonstrates potent activity against colorectal and pancreatic cancer through inhibition of doublecortin-like kinase 1. Mol Cancer. 2014;13:103.
Wenzel ES, Singh ATK. Cell-cycle checkpoints and aneuploidy on the path to cancer. In vivo. 2018;32:1–5.
Whittaker SR, Mallinger A, Workman P, Clarke PA. Inhibitors of cyclin-dependent kinases as cancer therapeutics. Pharm Ther. 2017;173:83–105.
Jin MH, Oh DY. ATM in DNA repair in cancer. Pharmacol Ther 2019;107391.
Zhang P, Wei Y, Wang L, Debeb BG, Yuan Y, Zhang J, et al. ATM-mediated stabilization of ZEB1 promotes DNA damage response and radioresistance through CHK1. Nat Cell Biol. 2014;16:864–75.
Han T, Lin J, Wang Y, Fan Q, Sun H, Tao Y, et al. Forkhead box D1 promotes proliferation and suppresses apoptosis via regulating polo-like kinase 2 in colorectal cancer. Biomed Pharmacother. 2018;103:1369–75.
Xie Y, Liu Y, Li Q, Chen J. Polo-like kinase 2 promotes chemoresistance and predicts limited survival benefit from adjuvant chemotherapy in colorectal cancer. Int J Oncol. 2018;52:1401–14.
Kolluri SK, Jin UH, Safe S. Role of the aryl hydrocarbon receptor in carcinogenesis and potential as an anti-cancer drug target. Arch Toxicol. 2017;91:2497–513.
Xue P, Fu J, Zhou Y. The aryl hydrocarbon receptor and tumor immunity. Front Immunol. 2018;9:286.
Foster JS, Henley DC, Bukovsky A, Seth P, Wimalasena J. Multifaceted regulation of cell cycle progression by estrogen: regulation of Cdk inhibitors and Cdc25A independent of cyclin D1-Cdk4 function. Mol Cell Biol. 2001;21:794–810.
Chandrakesan P, May R, Weygant N, Qu D, Berry WL, Sureban SM, et al. Intestinal tuft cells regulate the ATM mediated DNA Damage response via Dclk1 dependent mechanism for crypt restitution following radiation injury. Sci Rep. 2016;6:37667.
Yin L, Chang C, Xu C. G2/M checkpoint plays a vital role at the early stage of HCC by analysis of key pathways and genes. Oncotarget. 2017;8:76305–17.
Qiu Z, Oleinick NL, Zhang J. ATR/CHK1 inhibitors and cancer therapy. Radiother Oncol. 2018;126:450–64.
Kastan MB, Bartek J. Cell-cycle checkpoints and cancer. Nature. 2004;432:316–23.
Acknowledgements
This work was supported by the Mississippi INBRE, funded by an Institutional Development Award (IDeA) from the National Institute of General Medical Sciences of the National Institutes of Health under grant number P20GM103476. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of General Medical Sciences or the National Institutes of Health. The work performed through the UMMC Molecular and Genomics Facility is supported, in part, by funds from the NIGMS, including Mississippi INBRE (P20GM103476), Center for Psychiatric Neuroscience (CPN)-COBRE (P30GM103328), Obesity, Cardiorenal, and Metabolic Diseases-COBRE (P20GM104357), and Mississippi Center of Excellence in Perinatal Research (MS-CEPR)-COBRE (P20GM121334). The content of the paper is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health. Special acknowledgments to Dr Jonathan Lindner at University of Southern Mississippi for his great help with the FACS, Dr Sweta Khanal and Dr Alex Flynt at University of Southern Mississippi for his great help with the pathway heatmap establishment.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Co-first authors: Lianna Li, Hao Mei
Supplementary information
Rights and permissions
About this article
Cite this article
Li, L., Mei, H. & Commey, A.N.A. Application of RNA-sequencing to identify transcriptome modification by DCLK1 in colorectal cancer cells. Cancer Gene Ther 27, 691–701 (2020). https://doi.org/10.1038/s41417-019-0144-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41417-019-0144-4