Abstract
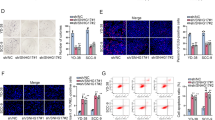
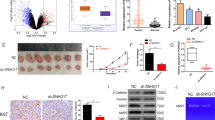
Small nucleolar RNA host gene 16 (SNHG16) has been documented to be involved in the pathogenesis of human cancers. Here, we elucidated the biological roles and regulatory mechanism of SNHG16 in the pathogenesis of oral squamous cell carcinoma (OSCC). In this paper, we found that c-Myc and SNHG16 were overexpressed in OSCC tissues and cell lines compared with normal tissues and normal human oral keratinocytes cells. There was a notable positive correlation between SNHG16 and c-Myc expression in OSCC tissues. c-Myc silencing by either shRNA c-Myc or by 10058-F4 (c-Myc inhibitor) resulted in a dose-dependent reduction in SNHG16 levels in CAL-27 and TSCCA cells; conversely, upregulation of c-Myc by pcDNA c-Myc markedly increased SNHG16 expression. Depletion of SNHG16 in CAL-27 cells strikingly inhibited cell proliferation, migration and invasion, as indicated by downregulation of proliferating cell nuclear antigen (PCNA), matrix metalloproteinase (MMP)-2 and MMP-9. Moreover, depletion of SNHG16 induced cell apoptosis and inhibited epithelial-to-mesenchymal transition as indicated by induction of cleaved caspase-3 and epithelial cadherin (E-cadherin) along with reduction of N-cadherin and Snail. Intriguingly, c-Myc knockdown led to the similar functional effects as that of SNHG16 knockdown in TSCCA cells. However, these changes caused by c-Myc knockdown were abrogated by SNHG16 overexpression. Knockdown of SNHG16 conspicuously repressed tumor growth in nude mice. Similarly, silencing of c-Myc markedly inhibited tumor growth and reduced SNHG16 expression in nude mice. Moreover, overexpression of SNHG16 blocked the inhibitory effect of c-Myc silencing on tumor growth in vivo. Thus, we conclude that c-Myc-induced upregulation of SNHG16 enhances progression and carcinogenesis in OSCC.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
GBD 2013 Mortality and Causes of Death Collaborators. Global, regional, and national age-sex specific all-cause and cause-specific mortality for 240 causes of death, 1990-2013: a systematic analysis for the Global Burden of Disease Study 2013. Lancet. 2015; 385:117–71.
Choi S, Myers JN. Molecular pathogenesis of oral squamous cell carcinoma: implications for therapy. J Dent Res. 2008;87:14–32.
Ernani V, Saba NF. Oral cavity cancer: risk factors, pathology, and management. Oncology. 2015;89:187–95.
Marsh D, Suchak K, Moutasim KA, Vallath S, Hopper C, Jerjes W, et al. Stromal features are predictive of disease mortality in oral cancer patients. J Pathol. 2011;223:470–81.
Gearhart J, Pashos EE, Prasad MK. Pluripotency redux--advances in stem-cell research. N Engl J Med. 2007;357:1469–72.
Dang CV. MYC, metabolism, cell growth, and tumorigenesis. Cold Spring Harb Perspect Med. 2013;3:a014217.
Dang CV, O’Donnell KA, Zeller KI, Nguyen T, Osthus RC, Li F. The c-Myc target gene network. Semin Cancer Biol. 2006;16:253–64.
Eberhardy SR, Farnham PJ. Myc recruits P-TEFb to mediate the final step in the transcriptional activation of the cad promoter. J Biol Chem. 2002;277:40156–62.
Cascon A, Robledo M. MAX and MYC: a heritable breakup. Cancer Res. 2012;72:3119–24.
Wiese KE, Walz S, von Eyss B, Wolf E, Athineos D, Sansom O, et al. The role of MIZ-1 in MYC-dependent tumorigenesis. Cold Spring Harb Perspect Med. 2013;3:a014290.
Kuzyk A, Mai S. c-MYC-induced genomic instability. Cold Spring Harb Perspect Med. 2014;4:a014373.
Horiuchi D, Kusdra L, Huskey NE, Chandriani S, Lenburg ME, Gonzalez-Angulo AM, et al. MYC pathway activation in triple-negative breast cancer is synthetic lethal with CDK inhibition. J Exp Med. 2012;209:679–96.
Eischen CM, Packham G, Nip J, Fee BE, Hiebert SW, Zambetti GP, et al. Bcl-2 is an apoptotic target suppressed by both c-Myc and E2F-1. Oncogene. 2001;20:6983–93.
Dang CV. MYC on the path to cancer. Cell. 2012;149:22–35.
Gabay M, Li Y, Felsher DW. MYC activation is a hallmark of cancer initiation and maintenance. Cold Spring Harb Perspect Med. 2014;4:a014241.
Hutter S, Bolin S, Weishaupt H, Swartling FJ. Modeling and targeting MYC genes in childhood brain tumors. Genes. 2017;8:107.
Bergmann JH, Spector DL. Long non-coding RNAs: modulators of nuclear structure and function. Curr Opin Cell Biol. 2014;26:10–18.
Wahlestedt C. Targeting long non-coding RNA to therapeutically upregulate gene expression. Nat Rev Drug Discov. 2013;12:433–46.
Yang T, Jin X, Lan J, Wang W. Long non-coding RNA SNHG16 has tumor suppressing effect in acute lymphoblastic leukemia by inverse interaction on hsa-miR-124-3p. IUBMB Life. 2018;71:134–142.
Zhao W, Fu H, Zhang S, Sun S, Liu Y. LncRNA SNHG16 drives proliferation, migration, and invasion of hemangioma endothelial cell through modulation of miR-520d-3p/STAT3 axis. Cancer Med. 2018;7:3311–20.
Yu M, Ohira M, Li Y, Niizuma H, Oo ML, Zhu Y, et al. High expression of ncRAN, a novel non-coding RNA mapped to chromosome 17q25.1, is associated with poor prognosis in neuroblastoma. Int J Oncol. 2009;34:931–8.
Christensen LL, True K, Hamilton MP, Nielsen MM, Damas ND, Damgaard CK. et al. SNHG16 is regulated by the Wnt pathway in colorectal cancer and affects genes involved in lipid metabolism. Mol Oncol. 2016;10:1266–82.
Qi P, Xu MD, Ni SJ, Shen XH, Wei P, Huang D, et al. Down-regulation of ncRAN, a long non-coding RNA, contributes to colorectal cancer cell migration and invasion and predicts poor overall survival for colorectal cancer patients. Mol Carcinog. 2015;54:742–50.
Lu YF, Cai XL, Li ZZ, Lv J, Xiang YA, Chen JJ, et al. LncRNA SNHG16 functions as an oncogene by sponging MiR-4518 and up-regulating PRMT5 expression in glioma. Cell Physiol Biochem. 2018;45:1975–85.
Cao X, Xu J, Yue D. LncRNA-SNHG16 predicts poor prognosis and promotes tumor proliferation through epigenetically silencing p21 in bladder cancer. Cancer Gene Ther. 2018;25:10–17.
Liao TT, Yang MH. Revisiting epithelial‐mesenchymal transition in cancer metastasis: the connection between epithelial plasticity and stemness. Mol Oncol. 2017;11:792–804.
Jeanes A, Gottardi CJ, Yap AS. Cadherins and cancer: how does cadherin dysfunction promote tumor progression? Oncogene. 2008;27:6920–9.
Wheelock MJ, Shintani Y, Maeda M, Fukumoto Y, Johnson KR. Cadherin switching. J Cell Sci. 2008;121:727–35.
Villarejo A, Cortes-Cabrera A, Molina-Ortiz P, Portillo F, Cano A. Differential role of Snail1 and Snail2 zinc fingers in E-cadherin repression and epithelial to mesenchymal transition. J Biol Chem. 2014;289:930–41.
Winkle M, van den Berg A, Tayari M, Sietzema J, Terpstra M, Kortman G, et al. Long noncoding RNAs as a novel component of the Myc transcriptional network. FASEB J. 2015;29:2338–46.
Kim T, Cui R, Jeon Y-J, Fadda P, Alder H, Croce CM. MYC-repressed long noncoding RNAs antagonize MYC-induced cell proliferation and cell cycle progression. Oncotarget. 2015;6:18780–9.
Doose G, Haake A, Bernhart SH, López C, Duggimpudi S, Wojciech F, et al. MINCR is a MYC-induced lncRNA able to modulate MYC’s transcriptional network in Burkitt lymphoma cells. Proc Natl Acad Sci USA. 2015;112:E5261–E5270.
Yang F, Xue X, Bi J, Zheng L, Zhi K, Gu Y, et al. Long noncoding RNA CCAT1, which could be activated by c-Myc, promotes the progression of gastric carcinoma. J Cancer Res Clin Oncol. 2013;139:437–45.
Cao L, Zhang P, Li J, Wu M. LAST, a c-Myc-inducible long noncoding RNA, cooperates with CNBP to promote CCND1 mRNA stability in human cells. eLife. 2017;6:e30433
Kim T, Jeon YJ, Cui R, Lee JH, Peng Y, Kim SH, et al. Role of MYC-regulated long noncoding RNAs in cell cycle regulation and tumorigenesis. J Natl Cancer Inst. 2015;107:dju505.
Wang O, Yang F, Liu Y, Lv L, Ma R, Chen C, et al. C-MYC-induced upregulation of lncRNA SNHG12 regulates cell proliferation, apoptosis and migration in triple-negative breast cancer. Am J Transl Res. 2017;9:533–45.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Li, S., Zhang, S. & Chen, J. c-Myc induced upregulation of long non-coding RNA SNHG16 enhances progression and carcinogenesis in oral squamous cell carcinoma. Cancer Gene Ther 26, 400–410 (2019). https://doi.org/10.1038/s41417-018-0072-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41417-018-0072-8
This article is cited by
-
Tumor reversion: a dream or a reality
Biomarker Research (2021)
-
Coding the noncoding: 2 years of advances in the field of microRNAs and long noncoding RNAs
Cancer Gene Therapy (2021)
-
LncRNA-SNHG16 promotes proliferation and migration of acute myeloid leukemia cells via PTEN/PI3K/AKT axis through suppressing CELF2 protein
Journal of Biosciences (2021)
-
LncRNAs as key players in the MYC pathways
Genome Instability & Disease (2021)
-
LncRNA SNHG16 regulates trophoblast functions by the miR-218-5p/LASP1 axis
Journal of Molecular Histology (2021)