Abstract
Background
Targeted RNA sequencing (RNA-seq) from FFPE specimens is used clinically in cancer for its ability to estimate gene expression and to detect fusions. Using a cohort of NSCLC patients, we sought to determine whether targeted RNA-seq could be used to measure tumour mutational burden (TMB) and the expression of immune-cell-restricted genes from FFPE specimens and whether these could predict response to immune checkpoint blockade.
Methods
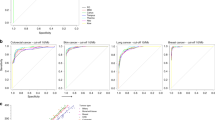
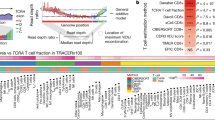
Using The Cancer Genome Atlas LUAD dataset, we developed a method for determining TMB from tumour-only RNA-seq and showed a correlation with DNA sequencing derived TMB calculated from tumour/normal sample pairs (Spearman correlation = 0.79, 95% CI [0.73, 0.83]. We applied this method to targeted sequencing data from our patient cohort and validated these results against TMB estimates obtained using an orthogonal assay (Spearman correlation = 0.49, 95% CI [0.24, 0.68]).
Results
We observed that the RNA measure of TMB was significantly higher in responders to immune blockade treatment (P = 0.028) and that it was predictive of response (AUC = 0.640 with 95% CI [0.493, 0.786]). By contrast, the expression of immune-cell-restricted genes was uncorrelated with patient outcome.
Conclusion
TMB calculated from targeted RNA sequencing has a similar diagnostic ability to TMB generated from targeted DNA sequencing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Data availability
The datasets generated and/or analysed during the current study are available from the corresponding author on reasonable request.
Code availability
R code that implements the variant filtering strategies, performs differential expression analysis and generates all figures contained in this manuscript, along with all relevant data are available on Figshare at https://doi.org/10.6084/m9.figshare.21332646.
References
Paez JG, Janne PA, Lee JC, Tracy S, Greulich H, Gabriel S, et al. EGFR mutations in lung cancer: correlation with clinical response to gefitinib therapy. Science. 2004;304:1497–500.
Recondo G, Facchinetti F, Olaussen KA, Besse B, Friboulet L. Making the first move in EGFR-driven or ALK-driven NSCLC: first-generation or next-generation TKI? Nat Rev Clin Oncol. 2018;15:694–708.
Paik PK, Drilon A, Fan PD, Yu H, Rekhtman N, Ginsberg MS, et al. Response to MET inhibitors in patients with stage IV lung adenocarcinomas harboring MET mutations causing exon 14 skipping. Cancer Discov. 2015;5:842–9.
Drilon A, Clark JW, Weiss J, Ou SI, Camidge DR, Solomon BJ, et al. Antitumor activity of crizotinib in lung cancers harboring a MET exon 14 alteration. Nat Med. 2020;26:47–51.
Drilon A, Hu ZI, Lai GGY, Tan DSW. Targeting RET-driven cancers: lessons from evolving preclinical and clinical landscapes. Nat Rev Clin Oncol. 2018;15:151–67.
Takeuchi K, Soda M, Togashi Y, Suzuki R, Sakata S, Hatano S, et al. RET, ROS1 and ALK fusions in lung cancer. Nat Med. 2012;18:378–81.
Remon J, Pignataro D, Novello S, Passiglia F. Current treatment and future challenges in ROS1- and ALK-rearranged advanced non-small cell lung cancer. Cancer Treat Rev. 2021;95:102178.
Majewski IJ, Mittempergher L, Davidson NM, Bosma A, Willems SM, Horlings HM, et al. Identification of recurrent FGFR3 fusion genes in lung cancer through kinome-centred RNA sequencing. J Pathol. 2013;230:270–6.
Reguart N, Teixido C, Gimenez-Capitan A, Pare L, Galvan P, Viteri S, et al. Identification of ALK, ROS1, and RET fusions by a multiplexed mRNA-based assay in formalin-fixed, paraffin-embedded samples from advanced non-small-cell lung cancer patients. Clin Chem. 2017;63:751–60.
McLeer-Florin A, Duruisseaux M, Pinsolle J, Dubourd S, Mondet J, Phillips Houlbracq M, et al. ALK fusion variants detection by targeted RNA-next generation sequencing and clinical responses to crizotinib in ALK-positive non-small cell lung cancer. Lung cancer. 2018;116:15–24.
Reck M, Rodriguez-Abreu D, Robinson AG, Hui R, Csoszi T, Fulop A, et al. Pembrolizumab versus chemotherapy for PD-L1-positive non-small-cell lung cancer. N Engl J Med. 2016;375:1823–33.
Hendry S, Byrne DJ, Wright GM, Young RJ, Sturrock S, Cooper WA, et al. Comparison of four PD-L1 immunohistochemical assays in lung cancer. J Thorac Oncol. 2018;13:367–76.
Cooper WA, Russell PA, Cherian M, Duhig EE, Godbolt D, Jessup PJ, et al. Intra- and interobserver reproducibility assessment of PD-L1 biomarker in non-small cell lung cancer. Clin Cancer Res. 2017;23:4569–77.
Butter R, t Hart NA, Hooijer GKJ, Monkhorst K, Speel EJ, Theunissen P, et al. Multicentre study on the consistency of PD-L1 immunohistochemistry as predictive test for immunotherapy in non-small cell lung cancer. J Clin Pathol. 2020;73:423–30.
Yarchoan M, Hopkins A, Jaffee EM. Tumor mutational burden and response rate to PD-1 inhibition. N Engl J Med. 2017;377:2500–1.
Marabelle A, Fakih M, Lopez J, Shah M, Shapira-Frommer R, Nakagawa K, et al. Association of tumour mutational burden with outcomes in patients with advanced solid tumours treated with pembrolizumab: prospective biomarker analysis of the multicohort, open-label, phase 2 KEYNOTE-158 study. Lancet Oncol. 2020;21:1353–65.
Ready N, Hellmann MD, Awad MM, Otterson GA, Gutierrez M, Gainor JF, et al. First-line nivolumab plus ipilimumab in advanced non-small-cell lung cancer (CheckMate 568): outcomes by programmed death ligand 1 and tumor mutational burden as biomarkers. J Clin Oncol. 2019;37:992–1000.
Hellmann MD, Paz-Ares L, Bernabe Caro R, Zurawski B, Kim SW, Carcereny Costa E, et al. Nivolumab plus ipilimumab in advanced non-small-cell lung cancer. N Engl J Med. 2019;381:2020–31.
Stenzinger A, Endris V, Budczies J, Merkelbach-Bruse S, Kazdal D, Dietmaier W, et al. Harmonization and standardization of panel-based tumor mutational burden measurement: real-world results and recommendations of the quality in pathology study. J Thorac Oncol. 2020;15:1177–89.
Strickler JH, Hanks BA, Khasraw M. Tumor mutational burden as a predictor of immunotherapy response: is more always better? Clin Cancer Res. 2021;27:1236–41.
Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011;17:3.
Kim D, Langmead B, Salzberg SL. HISAT: a fast spliced aligner with low memory requirements. Nat Methods. 2015;12:357–60.
Van der Auwera GA, Carneiro MO, Hartl C, Poplin R, Del Angel G, Levy-Moonshine A, et al. From FastQ data to high confidence variant calls: the Genome Analysis Toolkit best practices pipeline. Current protocols in bioinformatics/editoral board. Andreas D Baxevanis [et al.] 2013;43:11 0 1–0 33.
DiGuardo MA, Davila JI, Jackson RA, Nair AA, Fadra N, Minn KT, et al. RNA-seq reveals differences in expressed tumor mutation burden in colorectal and endometrial cancers with and without defective DNA-mismatch repair. J Mol Diagn. 2021;23:555–64.
Jessen E, Liu Y, Davila J, Kocher JP, Wang C. Determining mutational burden and signature using RNA-seq from tumor-only samples. BMC Med Genomics. 2021;14:65.
Zhao C, Jiang T, Hyun Ju J, Zhang S, Tao J, Fu Y, et al. TruSight oncology 500: enabling comprehensive genomic profiling and biomarker reporting with targeted sequencing. bioRxiv. [Preprint] 2020. https://www.biorxiv.org/content/10.1101/2020.10.21.349100v1.
Genomes Project C, Auton A, Brooks LD, Durbin RM, Garrison EP, Kang HM, et al. A global reference for human genetic variation. Nature. 2015;526:68–74.
Wang C, Davila JI, Baheti S, Bhagwate AV, Wang X, Kocher JP, et al. RVboost: RNA-seq variants prioritization using a boosting method. Bioinformatics. 2014;30:3414–6.
Neums L, Suenaga S, Beyerlein P, Anders S, Koestler D, Mariani A, et al. VaDiR: an integrated approach to variant detection in RNA. Gigascience. 2018;7:gix122.
Karczewski KJ, Francioli LC, Tiao G, Cummings BB, Alfoldi J, Wang Q, et al. The mutational constraint spectrum quantified from variation in 141,456 humans. Nature. 2020;581:434–43.
Chang H, Sasson A, Srinivasan S, Golhar R, Greenawalt DM, Geese WJ, et al. Bioinformatic methods and bridging of assay results for reliable tumor mutational burden assessment in non-small-cell lung cancer. Mol diagnosis Ther. 2019;23:507–20.
Merino DM, McShane LM, Fabrizio D, Funari V, Chen SJ, White JR, et al. Establishing guidelines to harmonize tumor mutational burden (TMB): in silico assessment of variation in TMB quantification across diagnostic platforms: phase I of the Friends of Cancer Research TMB Harmonization Project. J Immunother Cancer. 2020;8:e000147.
Fancello L, Gandini S, Pelicci PG, Mazzarella L. Tumor mutational burden quantification from targeted gene panels: major advancements and challenges. J Immunother Cancer. 2019;7:183.
Risso D, Schwartz K, Sherlock G, Dudoit S. GC-content normalization for RNA-Seq data. BMC Bioinforma. 2011;12:480.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26:139–40.
Team RC. R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2020.
Robin X, Turck N, Hainard A, Tiberti N, Lisacek F, Sanchez JC, et al. pROC: an open-source package for R and S+ to analyze and compare ROC curves. BMC Bioinforma. 2011;12:77.
Schloerke B, Cook D, Larmarange J, Briatte F, Marbach M, Thoen E, et al. GGally: Extension to ‘ggplot2’. 2021; R package version; 2.
Simko TWaV. R package “corrplot”: visualization of a correlation matrix. 2017. https://bmjopenrespres.bmj.com/content/4/1/e000250#ref-22andelsewhere.
Galvano A, Gristina V, Malapelle U, Pisapia P, Pepe F, Barraco N, et al. The prognostic impact of tumor mutational burden (TMB) in the first-line management of advanced non-oncogene addicted non-small-cell lung cancer (NSCLC): a systematic review and meta-analysis of randomized controlled trials. ESMO Open. 2021;6:100124.
Mosele F, Remon J, Mateo J, Westphalen CB, Barlesi F, Lolkema MP, et al. Recommendations for the use of next-generation sequencing (NGS) for patients with metastatic cancers: a report from the ESMO Precision Medicine Working Group. Annal Oncol. 2020;31:1491–505.
Cohen D, Hondelink LM, Solleveld-Westerink N, Uljee SM, Ruano D, Cleton-Jansen AM, et al. Optimizing mutation and fusion detection in NSCLC by sequential DNA and RNA sequencing. J Thorac Oncol. 2020;15:1000–14.
Pestinger V, Smith M, Sillo T, Findlay JM, Laes JF, Martin G, et al. Use of an integrated pan-cancer oncology enrichment next-generation sequencing assay to measure tumour mutational burden and detect clinically actionable variants. Mol diagnosis Ther. 2020;24:339–49.
Sorokin M, Gorelyshev A, Efimov V, Zotova E, Zolotovskaia M, Rabushko E, et al. RNA sequencing data for FFPE tumor blocks can be used for robust estimation of tumor mutation burden in individual biosamples. Front Oncol. 2021;11:732644.
Alborelli I, Leonards K, Rothschild SI, Leuenberger LP, Savic Prince S, Mertz KD, et al. Tumor mutational burden assessed by targeted NGS predicts clinical benefit from immune checkpoint inhibitors in non-small cell lung cancer. J Pathol. 2020;250:19–29.
Benayed R, Offin M, Mullaney K, Sukhadia P, Rios K, Desmeules P, et al. High yield of RNA sequencing for targetable kinase fusions in lung adenocarcinomas with no mitogenic driver alteration detected by DNA sequencing and low tumor mutation burden. Clin Cancer Res. 2019;25:4712–22.
Li MM, Datto M, Duncavage EJ, Kulkarni S, Lindeman NI, Roy S, et al. Standards and guidelines for the interpretation and reporting of sequence variants in cancer: a Joint Consensus Recommendation of the Association for Molecular Pathology, American Society of Clinical Oncology, and College of American Pathologists. J Mol Diagn: JMD. 2017;19:4–23.
Linette GP, Carreno BM. Tumor-infiltrating lymphocytes in the checkpoint inhibitor era. Curr Hematol Malig Rep. 2019;14:286–91.
Acknowledgements
We thank all the patients who participated in this study. We acknowledge the cooperation of the Melbourne Health Shared Pathology Service. We thank Ann Officer and Marliese Alexander for the study coordination. We thank Dr Michael Christie for the pathology review. Some results shown here are based on data generated and made publicly available by the TCGA Research Network (http://cancergenome.nih.gov/).
Funding
This work was funded by Bristol Myers Squibb.
Author information
Authors and Affiliations
Contributions
JFM and APF designed the work, played important roles in interpreting the results, and drafted and revised the manuscript. RL and DC played important roles in interpreting the results and revising the manuscript. TG acquired the laboratory data and drafted the manuscript. TM acquired the clinical data. JLL, TJ, BS and SBF played important roles in interpreting the results and revising the manuscript. All authors approved the final version of this manuscript and are accountable for all aspects of the work.
Corresponding author
Ethics declarations
Competing interests
SF is on advisory boards for AstraZeneca, Pfizer, Merck, Bayer, GSK, Roche, Janssen, Novartis, and Thermo Fisher. Honoraria are paid to Peter MacCallum Cancer Centre. BS is on advisory boards/receives honoraria for Roche/Genentech, Pfizer, Novartis, AstraZeneca, Merck, Bristol Myers Squibb, Amgen, BeiGene, Janssen and Lilly.
Ethics approval and consent to participate
All patients gave written informed consent for their tissue samples and medical history to be used in this research. This research was approved by the Peter MacCallum Cancer Centre Human Research Ethics Committee (HREC ref HREC/17/PMCC/42). This study was performed in accordance with the Declaration of Helsinki.
Consent to publish
All patients gave consent for the publication of anonymised information.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Markham, J.F., Fellowes, A.P., Green, T. et al. Predicting response to immune checkpoint blockade in NSCLC with tumour-only RNA-seq. Br J Cancer 128, 1148–1154 (2023). https://doi.org/10.1038/s41416-022-02105-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-022-02105-w