Abstract
Background
The interleukin (IL)-36 cytokines are a sub-family of the IL-1 family which are becoming increasingly implicated in the pathogenesis of inflammatory diseases and malignancies. Initial studies of IL-36 signalling in tumorigenesis identified an immune-mediated anti-tumorigenic function for these cytokines. However, more recent studies have shown IL-36 cytokines also contribute to the pathogenesis of lung and colorectal cancer (CRC).
Methods
The aim of this study was to investigate IL-36 expression in CRC using transcriptomic datasets and software such as several R packages, Cytoscape, GEO2R and AnalyzeR. Validation of results was completed by qRT-PCR on both cell lines and a patient cohort. Cellular proliferation was assessed by flow cytometry and resazurin reduction.
Results
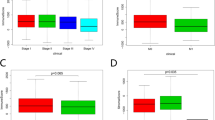
We demonstrate that IL-36 gene expression increases with CRC development. Decreased tumoral IL-36 receptor expression was shown to be associated with improved patient outcome. Our differential gene expression analysis revealed a novel role for the IL-36/IL-17/IL-23 axis, with these findings validated using patient-derived samples and cell lines. IL-36γ, together with either IL-17a or IL-22, was able to synergistically induce different genes involved in the IL-17/IL-23 axis in CRC cells and additively induce colon cancer cell proliferation.
Conclusions
Collectively, this data support a pro-tumorigenic role for IL-36 signalling in colon cancer, with the IL-17/IL-23 axis influential in IL-36-mediated colon tumorigenesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Data availability
The datasets analysed during the current study are available publically, as outlined in Table 1. The data underlying this article will be shared on a reasonable request to the corresponding author.
References
Xi Y, Xu P. Global colorectal cancer burden in 2020 and projections to 2040. Transl Oncol. 2021;14:101174.
Loomans-Kropp HA, Umar A. Increasing incidence of colorectal cancer in young adults. J Cancer Epidemiol. 2019;2019:9841295.
Xie Y-H, Chen Y-X, Fang J-Y. Comprehensive review of targeted therapy for colorectal cancer. Signal Transduct Target Ther. 2020;5:22.
Zaborowski AM, Winter DC, Lynch L. The therapeutic and prognostic implications of immunobiology in colorectal cancer: a review. Br J Cancer. 2021;125:1341–9.
Carlsen L, Huntington KE, El-Deiry WS. Immunotherapy for colorectal cancer: mechanisms and predictive biomarkers. Cancers. 2022;14:1028.
Hanahan D, Weinberg, Robert A. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74.
Van Der Kraak L, Gros P, Beauchemin N. Colitis-associated colon cancer: is it in your genes? World J Gastroenterol. 2015;21:11688–99.
Müller MF, Ibrahim AE, Arends MJ. Molecular pathological classification of colorectal cancer. Virchows Arch: Int J Pathol. 2016;469:125–34.
Friis S, Riis AH, Erichsen R, Baron JA, Sørensen HT. Low-dose aspirin or nonsteroidal anti-inflammatory drug use and colorectal cancer risk: a population-based, case-control study. Ann Intern Med. 2015;163:347–55.
Ciardiello D, Vitiello PP, Cardone C, Martini G, Troiani T, Martinelli E, et al. Immunotherapy of colorectal cancer: challenges for therapeutic efficacy. Cancer Treat Rev. 2019;76:22–32.
Byrne J, Baker K, Houston A, Brint E. IL-36 cytokines in inflammatory and malignant diseases: not the new kid on the block anymore. Cell Mol Life Sci: CMLS. 2021;78:6215–27.
Ridker PM, MacFadyen JG, Thuren T, Everett BM, Libby P, Glynn RJ. Effect of interleukin-1β inhibition with canakinumab on incident lung cancer in patients with atherosclerosis: exploratory results from a randomised, double-blind, placebo-controlled trial. Lancet. 2017;390:1833–42.
Bent R, Moll L, Grabbe S, Bros M. Interleukin-1 beta-A friend or foe in malignancies? Int J Mol Sci. 2018;19:2155.
Uppala R, Tsoi LC, Harms PW, Wang B, Billi AC, Maverakis E, et al. “Autoinflammatory psoriasis”—genetics and biology of pustular psoriasis. Cell Mol Immunol. 2021;18:307–17.
Gabay C, Towne JE. Regulation and function of interleukin-36 cytokines in homeostasis and pathological conditions. J Leukoc Biol. 2015;97:645–52.
Russell SE, Horan RM, Stefanska AM, Carey A, Leon G, Aguilera M, et al. IL-36α expression is elevated in ulcerative colitis and promotes colonic inflammation. Mucosal Immunol. 2016;9:1193–204.
Leon G, Hussey S, Walsh PT. The diverse roles of the IL-36 family in gastrointestinal inflammation and resolution. Inflamm Bowel Dis. 2021;27:440–50.
McCombs JE, Kolls JK. Walking down the “IL”: the newfound marriage between IL-36 and chronic obstructive pulmonary disease. Am J Respir Cell Mol Biol. 2021;64:153–4.
Baker KJ, Houston A, Brint E. IL-1 family members in cancer; two sides to every story. Front Immunol. 2019;10:1197.
Li D, Huang Y, Yu Z, Zhang J, Hu C, Bai Y, et al. IL-36β promotes anti-tumor effects in CD8 + T cells by downregulating micro-RNA let-7c-5p. Annal Transl Med. 2021;9:1734.
Wang X, Zhao X, Feng C, Weinstein A, Xia R, Wen W, et al. IL-36γ transforms the tumor microenvironment and promotes type 1 lymphocyte-mediated antitumor immune responses. Cancer Cell. 2015;28:296–306.
Zhao X, Chen X, Shen X, Tang P, Chen C, Zhu Q, et al. IL-36β promotes CD8(+) T cell activation and antitumor immune responses by activating mTORC1. Front Immunol. 2019;10:1803.
Weinstein AM, Chen L, Brzana EA, Patil PR, Taylor JL, Fabian KL, et al. Tbet and IL-36γ cooperate in therapeutic DC-mediated promotion of ectopic lymphoid organogenesis in the tumor microenvironment. Oncoimmunology. 2017;6:e1322238.
Weinstein AM, Giraldo NA, Petitprez F, Julie C, Lacroix L, Peschaud F, et al. Association of IL-36gamma with tertiary lymphoid structures and inflammatory immune infiltrates in human colorectal cancer. Cancer Immunol, immunotherapy: CII. 2019;68:109–20.
Mao D, Hu C, Zhang J, Feng C, Zhang Z, Wang J, et al. Long noncoding RNA GM16343 promotes IL-36β to regulate tumor microenvironment by CD8(+)T cells. Technol Cancer Res Treat. 2019;18:1533033819883633.
Huang L, Zhang H, Zhao D, Hu H, Lu Z. Interleukin-38 suppresses cell migration and proliferation and promotes apoptosis of colorectal cancer cell through negatively regulating extracellular signal-regulated kinases signaling. J Interferon Cytokine Res. 2021;41:375–84.
Baker K, O’Donnell C, Bendix M, Keogh S, Byrne J, O’Riordain M, et al. IL-36 signalling enhances a pro-tumorigenic phenotype in colon cancer cells with cancer cell growth restricted by administration of the IL-36R antagonist. Oncogene. 2022;41:2672–84.
Yang W, Dong H-P, Wang P, Xu Z-G, Xian J, Chen J, et al. IL-36γ and IL-36Ra reciprocally regulate colon inflammation and tumorigenesis by modulating the cell–matrix adhesion network and Wnt signaling. Adv Sci. 2022;9:2103035.
Li T, Fu J, Zeng Z, Cohen D, Li J, Chen Q, et al. TIMER2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Res. 2020;48:W509–w14.
Barrett T, Wilhite SE, Ledoux P, Evangelista C, Kim IF, Tomashevsky M, et al. NCBI GEO: archive for functional genomics data sets—update. Nucleic Acids Res. 2013;41:D991–D5.
Szklarczyk D, Gable AL, Nastou KC, Lyon D, Kirsch R, Pyysalo S, et al. The STRING database in 2021: customizable protein-protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 2021;49:D605–d12.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13:2498–504.
Bader GD, Hogue CWV. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinforma. 2003;4:2.
Guinney J, Dienstmann R, Wang X, de Reyniès A, Schlicker A, Soneson C, et al. The consensus molecular subtypes of colorectal cancer. Nat Med. 2015;21:1350–6.
Miller HE, Bishop AJR. Correlation AnalyzeR: functional predictions from gene co-expression correlations. BMC Bioinforma. 2021;22:206.
Yang W, Dong HP, Wang P, Xu ZG, Xian J, Chen J, et al. IL-36γ and IL-36Ra reciprocally regulate colon inflammation and tumorigenesis by modulating the cell-matrix adhesion network and Wnt signaling. Adv Sci. 2022;9:e2103035.
Rohr M, Beardsley J, Nakkina SP, Zhu X, Aljabban J, Hadley D, et al. A merged microarray meta-dataset for transcriptionally profiling colorectal neoplasm formation and progression. Sci Data. 2021;8:214.
Wong R. Proximal tumors are associated with greater mortality in colon cancer. J Gen Intern Med. 2010;25:1157–63.
le Rolle A-F, Chiu TK, Fara M, Shia J, Zeng Z, Weiser MR, et al. The prognostic significance of CXCL1 hypersecretion by human colorectal cancer epithelia and myofibroblasts. J Transl Med. 2015;13:199
Wang D, Sun H, Wei J, Cen B, DuBois RN. CXCL1 is critical for premetastatic niche formation and metastasis in colorectal cancer. Cancer Res. 2017;77:3655–65.
Scheibe K, Backert I, Wirtz S, Hueber A, Schett G, Vieth M, et al. IL-36R signalling activates intestinal epithelial cells and fibroblasts and promotes mucosal healing in vivo. Gut. 2017;66:823–38.
Eyerich S, Traidl-Hoffmann C, Behrendt H, Cavani A, Schmidt-Weber CB, Ring J, et al. Novel key cytokines in allergy: IL-17, IL-22. Allergol Sel. 2017;1:71–6.
Hu M, Tong Y, Fang H, Tang J, Liu L, Hu Y, et al. IL36 indicating good prognosis in human hepatocellular carcinoma. J Cancer. 2020;11:6248–55.
Pan Q-Z, Pan K, Zhao J-J, Chen J-G, Li J-J, Lv L, et al. Decreased expression of interleukin-36α correlates with poor prognosis in hepatocellular carcinoma. Cancer Immunol Immunother. 2013;62:1675–85.
Chang L, Guo R, Yuan Z. IL-36α suppresses proliferation of ovarian cancer cells. Tumor Biol. 2017;39:1010428317706918.
Chen F, Qu M, Zhang F, Tan Z, Xia Q, Hambly BD, et al. IL-36 s in the colorectal cancer: is interleukin 36 good or bad for the development of colorectal cancer? BMC Cancer. 2020;20:92.
Chelvanambi M, Weinstein AM, Storkus WJ. IL-36 signaling in the tumor microenvironment. Adv Exp Med Biol. 2020;1240:95–110.
Manzanares-Meza LD, Valle-Rios R, Medina-Contreras O. Interleukin-1 receptor-like 2: one receptor, three agonists, and many implications. J Interferon Cytokine Res. 2022;42:49–61.
Kovach MA, Che K, Brundin B, Andersson A, Asgeirsdottir H, Padra M, et al. IL-36 cytokines promote inflammation in the lungs of long-term smokers. Am J Respir Cell Mol Biol. 2021;64:173–82.
Yuan Z-C, Xu W-D, Liu X-Y, Liu X-Y, Huang A-F, Su L-C. Biology of IL-36 signaling and its role in systemic inflammatory diseases. Front Immunol. 2019;10:2532. -
Chi HH, Hua KF, Lin YC, Chu CL, Hsieh CY, Hsu YJ, et al. IL-36 signaling facilitates activation of the NLRP3 inflammasome and IL-23/IL-17 axis in renal inflammation and fibrosis. J Am Soc Nephrology: JASN. 2017;28:2022–37.
Razi S, Baradaran Noveiry B, Keshavarz-Fathi M, Rezaei N. IL-17 and colorectal cancer: From carcinogenesis to treatment. Cytokine. 2019;116:7–12.
Henry CM, Sullivan GP, Clancy DM, Afonina IS, Kulms D, Martin SJ. Neutrophil-derived proteases escalate inflammation through activation of IL-36 family cytokines. Cell Rep. 2016;14:708–22.
Ngo VL, Abo H, Maxim E, Harusato A, Geem D, Medina-Contreras O, et al. A cytokine network involving IL-36γ, IL-23, and IL-22 promotes antimicrobial defense and recovery from intestinal barrier damage. Proc Natl Acad Sci USA. 2018;115:E5076–E85.
Mercurio L, Failla CM, Capriotti L, Scarponi C, Facchiano F, Morelli M, et al. Interleukin (IL)-17/IL-36 axis participates to the crosstalk between endothelial cells and keratinocytes during inflammatory skin responses. PLoS ONE. 2020;15:e0222969.
Maxwell JR, Zhang Y, Brown WA, Smith CL, Byrne FR, Fiorino M, et al. Differential roles for interleukin-23 and interleukin-17 in intestinal immunoregulation. Immunity. 2015;43:739–50.
Straus DS. TNFα and IL-17 cooperatively stimulate glucose metabolism and growth factor production in human colorectal cancer cells. Mol Cancer. 2013;12:78.
Müller A, Hennig A, Lorscheid S, Grondona P, Schulze-Osthoff K, Hailfinger S, et al. IκBζ is a key transcriptional regulator of IL-36-driven psoriasis-related gene expression in keratinocytes. Proc Natl Acad Sci USA. 2018;115:10088–93.
Santiago-Sánchez GS, Pita-Grisanti V, Quiñones-Díaz B, Gumpper K, Cruz-Monserrate Z, Vivas-Mejía PE. Biological functions and therapeutic potential of lipocalin 2 in cancer. Int J Mol Sci. 2020;21:4365.
Kim SL, Shin MW, Seo SY, Kim SW. Lipocalin 2 potentially contributes to tumorigenesis from colitis via IL-6/STAT3/NF-κB signaling pathway. Biosci Rep. 2022;42:BSR20212418.
Kim SL, Lee ST, Min IS, Park YR, Lee JH, Kim DG, et al. Lipocalin 2 negatively regulates cell proliferation and epithelial to mesenchymal transition through changing metabolic gene expression in colorectal cancer. Cancer Sci. 2017;108:2176–86.
Funding
This work was funded by a grant from the Government of Ireland Postgraduate Scholarship Scheme GOIPG/2018/2974.
Author information
Authors and Affiliations
Contributions
KJB conceived and designed work, performed both transcriptomic analysis and experimental work and was involved in the writing of the manuscript. EB and AH conceived and designed work, undertook the data analysis and writing of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
The study was approved by the University College Cork Clinical Research Ethics Committee of the Cork Teaching Hospitals (ECM (3) P 3 September 2013). All samples were obtained during surgery at the Mercy University Hospital Cork following informed consent.
Consent to publish
Not applicable.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Baker, K.J., Brint, E. & Houston, A. Transcriptomic and functional analyses reveal a tumour-promoting role for the IL-36 receptor in colon cancer and crosstalk between IL-36 signalling and the IL-17/ IL-23 axis. Br J Cancer 128, 735–747 (2023). https://doi.org/10.1038/s41416-022-02083-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-022-02083-z
This article is cited by
-
Imbalance of Circulating Follicular Regulatory and Follicular Helper T Cell Subpopulations Is Associated with Disease Progression and Serum CYFRA 21-1 Levels in Patients with Non-small Cell Lung Cancer
Current Medical Science (2024)
-
Bioinformatics Identification of Therapeutic Gene Targets for Gastric Cancer
Advances in Therapy (2023)