Abstract
Epigenetic mechanisms play instrumental roles in gene regulation during embryonic development and disease progression. However, it is challenging to non-invasively monitor the dynamics of epigenomes and related gene regulation at inaccessible human tissues, such as tumours, fetuses and transplanted organs. Circulating cell-free DNA (cfDNA) in peripheral blood provides a promising opportunity to non-invasively monitor the genomes from these inaccessible tissues. The fragmentation patterns of plasma cfDNA are unevenly distributed in the genome and reflect the in vivo gene-regulation status across multiple molecular layers, such as nucleosome positioning and gene expression. In this review, we revisited the computational and experimental approaches that have been recently developed to measure the cfDNA fragmentomics across different resolutions comprehensively. Moreover, cfDNA in peripheral blood is released following cell death, after apoptosis or necrosis, mainly from haematopoietic cells in healthy people and diseased tissues in patients. Several cfDNA-fragmentomics approaches showed the potential to identify the tissues-of-origin in cfDNA from cancer patients and healthy individuals. Overall, these studies paved the road for cfDNA fragmentomics to non-invasively monitor the in vivo gene-regulatory dynamics in both peripheral immune cells and diseased tissues.
Similar content being viewed by others
Main
Multidimensional epigenetic layers play instrumental roles in mammalian gene regulation [1,2,3]. During cancer initiation and progression, embryonic development and organ transplantation, epigenomes vary across different time points and impact gene expression and biological function [4,5,6,7,8,9]. In animal models, the variations of the epigenome and transcriptome across different time points can be characterised through high-throughput sequencing technologies, such as ATAC-seq and RNA-seq, by sacrificing the animals with the same genetic background [10,11,12]. In cultured human cells still alive, the epigenome dynamics can be visualised by fluorescence live imaging [13,14,15]. However, in human subjects still alive, especially for the inaccessible human tissues, such as solid tumours, fetuses during pregnancy and transplanted organs, how to non-invasively monitor the dynamics of their epigenomes and transcriptomes is still largely unexplored.
Circulating cell-free DNA (cfDNA) in the peripheral blood offers a promising and non-invasive approach to monitoring the genome dynamics from the inaccessible human tissues [16,17,18]. CfDNA molecules from the tumours, foetus and transplanted organs can be separated out based on the allelic status of mutations or single-nucleotide polymorphisms (SNPs) within fragments measured through deep whole-genome sequencing (WGS). Thus, part of or the whole genome in these inaccessible tissues can be reconstructed across different time points [19,20,21]. However, most of the current cfDNA WGS studies only focus on the genetic aberrations in the cfDNA, such as SNPs or copy number variations (CNVs) [22,23,24], but not epigenomes. Since the cfDNA fragments are outside the cells and highly fragmented, it is challenging to directly characterise the epigenomes, except for covalent DNA modifications and very few other cases [25,26,27,28,29,30,31]. Interestingly, the fragmentation patterns of cfDNA are not uniform across the genome [32, 33]. Several studies suggested a tight correlation between cfDNA-fragmentation patterns and in vivo gene-regulatory maps within the cells, such as nucleosome positioning and gene expression [32,33,34]. These correlations raise the possibility of inferring back the epigenome and transcriptome within the cells from the cfDNA-fragmentation patterns. In this review, we will first go through the major milestones of cfDNA-fragmentation studies in the pre-omics era, which inspired many studies in the later-omics era. Further, we will summarise the state-of-art high-throughput methods, mostly computational, on the measurement of cfDNA fragmentation (Table 1). Finally, we will go through the approaches for the tissue-of-origin analysis using cfDNA fragmentation.
The pre-omics era of cfDNA-fragmentation studies
Early in 1948, the presence of cell-free nucleic acids in the circulation was first described by Mandel and Metais in human blood from healthy individuals and patients with different diseases [35]. In the beginning, circulating cell-free DNA (cfDNA) was demonstrated to be predominantly double-stranded [36]. Later in 1973, single-stranded cfDNA fragments were also characterised in the serum from systemic lupus erythematosus (SLE) patients [37]. The variations of strandness and sizes of cfDNA fragments altered the clearance kinetics and mechanism of DNA in the circulation [38]. Stroun et al., in 1987, showed that cfDNA from cancer patients was double-stranded primarily with the size of 0.5–21 kb and revealed that cfDNA from cancer patients is smaller than genomic DNA with fragmented status [39]. Later, several studies suggested that cfDNA is mainly released from apoptotic or necrotic cells [40, 41], which is further proved by multiple subsequent studies [42, 43]. In 2002, Lui et al. first demonstrated that the cfDNA from healthy individuals mostly comes from the haemopoietic cells by estimating the percentage of cfDNA from the Y chromosome in the plasma from sex-mismatched bone marrow transplantation patients [44]. In 2003, Deligezer et al. suggested a significant correlation between the fragmentation status in cfDNA and DNA methylation, as well as the nucleosomes, at the 1st exon of the p16 gene in lymphoma patients [45]. Further, the study of variations in cfDNA-fragment sizes was expanded to other fields, such as prenatal test [46].
Before the development of next-generation sequencing (NGS) technology, quantitative PCR (qPCR) was applied to study the fragmentation in cfDNA. By using qPCR, Diehl et al. discovered that mutant sequences are enriched in small DNA fragments (<180 bp) but not in long fragments [40, 47, 48]. Further, by optimised qPCR and xenografted model, Mouliere et al. first indicated that cancer-derived cfDNA showed higher fragmentation as compared with healthy controls highlighting higher fragmentation as a hallmark of cancer cfDNA (circulating tumour DNA, ctDNA) [49]. Sanchez et al. systematically compared the distribution of cfDNA-fragment sizes with cfDNA WGS in both double-stranded and single-stranded cfDNA by qPCR and discussed the association of mononucleosomes and chromatosomes with cfDNA [50].
Moreover, Meddeb et al. quantified the influence of several pre-analytical and demographic parameters on the overall variations of fragmentation in nuclear and mitochondrial circulating cfDNA [51]. Recently, droplet digital PCR (ddPCR) was combined with bisulfite treated by Shemer et al. to accurately quantify the tissues-of-origin in cfDNA [52]. Other approaches, such as sophisticated capillary electrophoresis, electron microscopy, Raman microscope, 3D laser-scanning confocal microscope and atomic force microscopy, were also utilised to study the cfDNA fragments besides NGS [53,54,55]. However, due to the limitation of technologies, cfDNA-fragmentation studies at the pre-omic era were mostly limited to the summary statistics or limited loci in the genome. The correlation between genome-wide fragmentation patterns in cfDNA and gene regulation within the cells is still not explored.
The cfDNA-fragmentomics era and gene regulation
Thanks to the advances in NGS, it is possible to measure the fragmentation patterns in millions of cfDNA molecules across different genomic locations in a high-precision and high-throughput way. Due to the overall short-fragment sizes in cfDNA, WGS library construction on cfDNA does not require the mechanic sonication step, which was widely used for the traditional WGS library construction on genomic DNA and will significantly affect the measurement of fragmentation patterns in cfDNA. Moreover, instead of a single summary-statistic score or characteristics from a limited number of loci, researchers can accurately estimate the fragment size, ends and strandness at every single cfDNA molecule, as well as the fragment coverage at each base of the reference genome [16, 56,57,58,59,60,61] [16, 56,57,58,59]. In 2008, Fan et al. identified the well-positioned nucleosomal patterns around TSS in plasma cfDNA but not in genomic DNA control by utilising the fragment coverages estimated from NGS [62]. In 2010, Fan et al. further demonstrated that the fragment size originated from the foetal side is smaller than those originated from the maternal side through paired-end NGS [63]. The concept of ‘cfDNA fragmentomics’ was first introduced by Ivanov et al. in 2015 [32]. Since then, in addition to the study of the fragment coverages and sizes, several innovative computational and experimental approaches have been developed to comprehensively measure the cfDNA-fragmentation patterns in plasma across different resolutions, including large-scale fragmentation patterns at megabase level (DELFI) [64], large-scale co-fragmentation patterns (FREE-C) [65], fragment coverage near transcription-start sites (TSS) [34], cfDNA-accessibility score near the transcription factor-binding sites (TFBS) [66], orientation-aware cfDNA fragmentation (OCF) [67], windowed protection score (WPS) [33], cfDNA-fragmentation hotspots [68], inference of DNA methylation from cfDNA-fragmentation patterns [69], the preferred-ended position of cfDNA [70, 71], the end-motif frequency and motif-diversity score (MDS) [72], jagged end [73, 74] and patterns outside the chromosomes [75,76,77,78]. Here, we will go through these state-of-art approaches applied at cfDNA fragmentation from a large-scale genomic bin to a single fragment.
Large-scale fragmentation patterns (DELFI) and chromatin organisations [64]
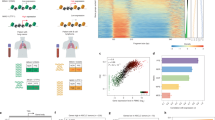
“DNA evaluation of fragments for early interception” (DELFI) was developed to detect genome-wide fragmentation abnormality of cfDNA by ~1X low-coverage whole-genome sequencing (WGS). DELFI evaluated the fragment coverage, size and other summary statistics within 100-kb non-overlapping bins and aggregated them into the 5-megabase (Mb) non-overlapping window, which will bring more than 20,000 reads per window at 1–2X genome coverage. They estimated the ratio between short cfDNA fragments (100–150 bp) and long cfDNA fragments (151–220 bp) within each window and found the increased aberrations at cancer patients, not at healthy individuals. They further utilised these summary statistics for the diagnosis of early-stage cancer. The fragment coverage among these statistics in the window was the most critical classification feature in their stochastic gradient-boosting model. Regarding the correlation with gene regulation, they utilised the window-protection score (WPS) [33] approach to infer the nucleosomal positioning in both cfDNA- and DNase-digested genomic DNA (gDNA) from lymphocytes. The median distance of the nucleosome from cfDNA showed a high correlation with that from DNase-digested gDNA and intermediated correlation with the chromatin compartment characterised by Hi-C from lymphocytes. These results indicated the important role of both nucleosome occupancy and high-ordered chromatin organisations for the cfDNA fragmentomics at cancers.
Large-scale co-fragmentation patterns (FREE-C) and 3D genome [65]
‘FRagmentation Evaluation of Epigenetics from CfDNA sequencing’ (FREE-C) was developed to evaluate the co-fragmentation patterns between pairs of large-scale genomic bins in the low-coverage WGS. They hypothesised that the cfDNA molecules released from two genomic regions would show similar fragmentation patterns at large scale if the two regions are spatially close to each other inside the cells, further informing the 3D genome status. There are two different approaches in FREE-C: multi-sample FREE-C and single-sample FREE-C. (1). In multi-sample FREE-C, they divided the chromosomes into 500-kilobase (kb) non-overlapping bins and calculated a ‘normalised fragmentation score’ based on fragment size at each bin for each individual. They then calculated the Pearson correlation coefficient of the fragmentation scores between each pair of bins in the same chromosome across all individuals. (2). Single-sample FREE-C calculated the fragment-size distributions of cfDNA in each genomic bin at a single cfDNA WGS sample. Further, FREE-C characterised the distance of the fragment-size distribution between each pair of genomic bins in the same chromosome using the Kolmogorov–Smirnov test. After obtaining the two-dimensional distance-correlation matrix between bins in each chromosome by multi-sample FREE-C and single-sample FREE-C, they found a high similarity with the correlation matrix characterised from Hi-C data in white blood cells (WBCs). The first eigenvector of the matrix from cfDNA in both approaches showed high similarity with the A/B compartments characterised from Hi-C data in WBCs. The fragment sizes of cfDNA are affected by multiple epigenetic backgrounds, such as DNA methylation and histone modifications [32, 79]. Thus, they performed the multivariate analysis and confirmed that the 3D genome signal is the major contributor to the co-fragmentation patterns in large-scale rather than sequence-composition bias or other large-scale epigenetic signals, such as CpG methylation and H3K4me1. This study has not been peer-reviewed and is supported by commercial entities.
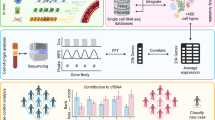
Fragment coverage near TSS and gene expression [34]
In 2016, Ulz et al. developed a computational approach to summarise the fragment coverages at two focal regions near TSS: 2000-bp region centred on the TSS (2K-TSS coverage) and −150 bp to +50 bp with respect to the TSS (NDR coverage). They normalised the fragment coverage for both regions with the relative copy number to exclude the potential effect of copy number alterations (CNAs) often observed in cancer. Further, Ulz et al. utilised these two features by a support-vector machine model for the binary classification of the housekeeping genes and constitutively not-expressed genes and achieved high accuracy in the cross-validation. Moreover, the model was applied in the cancer patients to predict the gene’s binary expression status or even several isoforms inside the tumour, suggesting a promising opportunity to non-invasively investigate the in vivo gene expression status at inaccessible tissues. The earlier study by Ivanov et al. [32] also revealed the associations between gene expression and nucleosome-fragmentation patterns near promoters but with whole-exome sequencing, which limited its capability to predict the gene expression status with high accuracy.
The cfDNA-accessibility score and the binding affinity of the transcription factors [66]
In 2019, Ulz et al. developed an accessibility score to estimate the overall binding affinity of the transcription factor (TF) by cfDNA WGS. They summarised the cfDNA-fragment coverages near each transcription factor-binding site (TFBS). The gDNA-accessibility pattern near TFBS was different in proximal and distal regions to the TSS. They found the low-frequency pattern of cfDNA-fragment coverage at proximal TFBS and the high-frequency pattern of cfDNA-fragment coverage at distal TFBS. They utilised Savitzky–Golay filters to suppress effects on the coverage not contributed by nucleosome positioning and remove the local biases. Further, the high-frequency signals were ranked and finally utilised as the accessibility score. Ulz’s cfDNA-accessibility score around the cell-type-specific TFs showed similar overall trends as that at gDNA accessibility measured by ATAC-seq between cancer and healthy. Thus, they concluded that the cfDNA-accessibility score indicated the global in vivo binding affinity of TFBS in each cfDNA sample. Further, Ulz et al. demonstrated the ability of the cfDNA- accessibility score to distinguish the subtypes of late-stage prostate cancer by aggregated low-coverage cfDNA WGS. Finally, the accessibility score across 504 TFs showed good performance in identifying the early-stage colorectal-cancer samples even with the low tumour fraction.
OCF and open-chromatin regions [67]
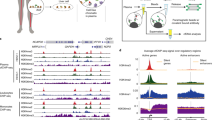
Sun et al. developed a computational approach to quantify the cfDNA-fragmentation patterns near the tissue-specific open-chromatin regions identified in previous epigenomic profiling by DNase-seq. First, they identified the tissue-specific open-chromatin regions from different publicly available DNA- accessibility data. Second, around the centre of these open-chromatin regions (±1000 bp), they calculated the coverage of upstream (U) fragment ends (the end mapped to the reference genome with a smaller coordinate), and the coverage of downstream (D) fragment ends (the end mapped to the reference genome with larger coordinate), respectively. Since the U and D ends showed peaks at −60 and +60 bp from the centre of the open-chromatin regions, respectively, they calculated the OCF value by quantifying the differences of coverages of the U and D ends in 20-bp windows around these two peaks for each tissue-specific open-chromatin regions. Naturally, the OCF value can be utilised to rank the relative contributions of different cell types to the circulating cfDNA and further application for the diagnosis of different pathological and physiological conditions.
WPS, single-stranded cfDNA, nucleosomes and other epigenetic backgrounds [33]
In the traditional WGS library-preparation approach, damaged and short double-stranded DNA molecules were poorly recovered. Thus, Snyder et al. developed a single-stranded cfDNA library- preparation method, which was adapted from studies of damaged ancient DNA [80]. Several follow-up studies on single-stranded cfDNA further revealed that most cfDNA is highly nicked, which might be subjected to continuous nuclease activity in the bloodstream [50, 67, 81]. Snyder and his colleagues also developed a computational approach, named WPS, to reconstruct the footprint of nucleosome positioning and transcription factor binding through deep WGS. Basically, WPS was calculated by the number of DNA fragments completely spanning a genomic window minus the number of fragments with an endpoint within that same window. They applied a peak-calling algorithm on the genome-wide WPS signals and identified many nucleosome-occupied regions, which showed similar genomic distributions as the nucleosome maps previously identified within the blood cells. The long-fraction WPS (L-WPS) was calculated by long fragments (120–180 bp) with a larger window (120 bp) and short-fraction WPS (S-WPS) was estimated by short fragments (35–80 bp) with a smaller window (16 bp). While L-WPS captured the signal from nucleosome occupancy, S-WPS represented the potential of TF binding, such as CTCF. Further, the nucleosomal spacing identified by WPS showed correlations with the A/B compartments identified by Hi-C, open-chromatin regions and gene expression specifically from haematopoietic cells, which enable its potential to identify the tissues-of-origin from cfDNA.
CfDNA-fragmentation hotspots, open-chromatin regions and transposons [68]
Cell-free dna fragmentation (CRAG) was developed to de novo characterise the genome-wide cfDNA- fragmentation hotspots in cfDNA WGS. Many studies suggested that local nucleosome structure reduces the fragmentation process, indicating the potential enrichment of cfDNA-fragmentation hotspots (lower coverage and smaller size) at the open-chromatin regions. Basically, they utilised a 200-bp sliding window to scan the genome. The fragment coverage was weighted by the ratio of average fragment sizes in the window versus that in the whole chromosome, named integrated fragmentation score (IFS). They further applied a negative binomial model to test if the windows showed significantly lower IFS than the local (5-kb and 10-kb centre to the window) and global background (whole chromosome). In cfDNA from healthy, cfDNA-fragmentation hotspots were enriched in gene-regulatory elements, including promoters, haematopoietic-specific enhancers and 3′ end of transposons. In cfDNA from early-stage cancers, IFS showed aberrant patterns at hotspots near microsatellites, CTCF and promoters of genes enriched in immune processes from peripheral immune cells. They also applied the IFS signals from hotspots for the diagnosis of multiple early-stage cancers with high accuracy. This study has not been peer-reviewed.
Inference of DNA methylation from cfDNA-fragmentation patterns [69]
The cfDNA-fragment sizes are significantly different at methylated and unmethylated fragments [45, 79]. Liu et al. developed a machine-learning approach to infer the base-pair resolution DNA methylation level by the fragmentation patterns from cfDNA WGS [69]. Fragment size, coverage and the distance of each CpG to the fragment end in each fragment were utilised as features for a non-homogeneous hidden Markov model. The emission probability was estimated by adding a Bayesian factor of cfDNA methylation level from healthy peripheral blood mononuclear cells to the multivariate Gaussian mixture model. In cfDNA WGBS, with the ground truth of the binary CpG methylation status in each CpG at each fragment, they achieved about 0.92 area under the curve in the balance-sampled CpG sites at CpG-rich regions (>=10 CpGs in each fragment). They also achieved a high correlation in a 1-kb window between the prediction from cfDNA WGS by the matched cfDNA WGBS from the same tube of blood in both healthy and multiple cancer patients with deep and shallow sequencing. Using hundreds of WGBS datasets from different tumour and normal cells as the reference map, they deconvoluted cfDNA’s tissue-of-origin status by inferred DNA methylation level at ultra-low-pass WGS from thousands of breast or prostate cancer samples and healthy individuals. The tissue-of-origin status in cancer patients showed a high concordance with confirmed metastasis tissues from physicians and correlation with some clinical metadata. This study has not been peer-reviewed.
The preferred-ended position of cfDNA [70, 71]
Taking advantage of PCR-free ultra-deep WGS, Chan et al. first identified the presence of foetus-associated cell-free DNA preferred ends at plasma from pregnant women [70]. They scanned the genome to check if certain locations had a significantly increased probability of being an ending position of plasma DNA fragments using a Poisson probability function. The ratio between foetal-specific preferred end and maternal-specific preferred end showed a high correlation with the foetal fraction estimated by the fraction of reads mapped to the chrY. Moreover, the fragments with these foetus-specific preferred ends showed similar fragment-size distributions as those overlapped with foetus-specific alleles. These observations disappeared at fragments that ended more than just 5 bp away from the foetal-specific preferred ends, which suggested the high specificity of these preferred-end positions from the foetus. Later, Jiang et al. discovered the liver-associated preferred ends at patients who received liver transplants and tumour-associated preferred DNA ends at cancer patients with a similar strategy [71]. In the follow-up study from the same group, Sun et al. suggested that the preferred-end positions were highly correlated with the nucleosomal structure [82]. They found that foetal preferred end sites were generally located in the nucleosome cores, while the maternal ones were located in the linker regions, which explained the relative shortness of foetal DNA in maternal plasma.
The end-motif frequency and MDS [72]
Jiang et al. measured the end-motif frequency by calculating the frequency of the first 4-nucleotide (i.e. 4-mer) sequence on each 5′-fragment end of plasma DNA after alignment to the reference genome. The comparison of end-motif frequency revealed the significant differences between hepatocellular carcinoma (HCC) patients and healthy at some of the motifs across all the 256 combinations. For example, the CCCA motif showed significant reductions at the cfDNA from cancer patients compared with the non-malignant controls. Jiang et al. further utilised the normalised Shannon entropy to summarise the variations of the motif frequency into a single MDS score for each cfDNA WGS sample. A higher MDS value suggested a higher variety of plasma DNA molecules with different end motifs, while a lower MDS value indicated fewer varieties of plasma DNA end motifs. They found a higher MDS value in HCC patients than the non-malignant controls and utilised it as a marker for the diagnosis of HCC. At the same dataset, the MDS value showed a better classification power than other fragmentomic metrics, such as fragment size or OCF value developed in the same group. More importantly, by utilising the DNASE1L3-deleted model system in mice, they discovered that the DNASE1L3 might have a major role in generating the CC motif fragments predominant in cfDNA. In the human cancer data from the TCGA research network, they observed that the expression levels of DNASE1L3 across multiple cancers were generally downregulated. In several other additional literature, homozygous loss-of-function mutations of DNASE1L3 in both mouse model and human were found to significantly impact the end-motif frequency [83, 84]. These findings finally open the door to dissect the molecular mechanism behind cfDNA fragmentomics.
Jagged end [73, 74]
The traditional WGS library-preparation step usually contains the DNA end-repair steps before the generation of NGS libraries, which erases the protruding-end (i.e. jagged-end) information. Early in 2008, Suzuki et al. identified the 5′ jagged ends in plasma DNA but not in gDNA by using the radioactive end-labelling procedure with the enzyme Klenow fragment [73]. CpG dinucleotides outside the CpG island in the human genome are usually methylated. In the library-preparation step of bisulfite sequencing, unmethylated cytosine is usually used for the end-repair step. Therefore, the uneven methylation status of CpG between the original and newly synthesised strand at the 3′ end may reflect the length of the jagged end. By utilising the publicly available cfDNA and gDNA WGBS data, Jiang et al. showed the proof-of-concept results of the genome-wide presence of jagged end at plasma DNA but not at sonicated genomic DNA from blood cells [74]. They developed a jag index (JI-U, Jagged Index-Unmethylated) in read 2 to measure the jaggedness of the fragments. The JI-U in cfDNA fragments between 130 and 160 bp showed a high performance in diagnosing the HCC in their previous cfDNA WGBS cohort. CpG dinucleotides are underrepresented in the human genome, which will affect the accuracy of measuring the length of the jagged end. Since non-CpG cytosine is usually unmethylated and widely distributed in the human genome, they further developed CC-tag technology by using methylated cytosine instead of unmethylated cytosine for the end-repair step. Therefore, in the context of CpH, they can utilise the JI-M (Jagged Index-Methylated) to characterise the jaggedness with higher accuracy. Finally, they observed the increases of jaggedness in mice with the deletion of DNASE1L3 (DNASE1L3−/−) compared with wild-type mice, whereas the decreases of jaggedness in mice with deletion of DNASE1 (DNASE1−/−), further revealed the possible biological mechanism behind jaggedness.
CfDNA-fragmentation patterns at eccDNA [76,77,78]
Zhu et al. and Kumar et al. published the articles almost the same time about the genome-wide identifications of extrachromosomal circular DNA (eccDNA) at the cfDNA from human blood [76, 77]. To enrich eccDNA from cfDNA, Zhu et al. utilised the ATP-dependent DNase to selectively digest linear DNA and then applied the multiple-displacement-amplification (MDA) method to amplify the remaining circular DNA preferentially. Kumar et al. utilised the proteinase and exonucleases to remove linear DNA and subjected to rolling-circle amplification to increase the yield of circular DNA. Very recently, Sin et al. utilised exonuclease V to digest the background linear DNA and follow-up with MspI digestion or Tn5 tagmentation to build the NGS library [78]. These results suggested that ecc-cfDNA showed predominate peaks at dinucleosomal fragment size and highly enriched in exons, 3′UTR and CpG island, as well as the DNase-hypersensitive sites, H3K4Me1 and H3K27Ac marks. The ecc-cfDNA was also identified at cfDNA from cancer patients with longer fragment size than that from healthy controls, which suggested it as a potential biomarker for the disease diagnosis.
Overall, different approaches emerged rapidly to measure the cfDNA fragmentomics and even benefit other non-fragmentation-based measurements of cfDNA. For example, the characteristics of small fragment sizes from ctDNA or foetus have been utilised to enrich the ctDNA or cell-free foetal DNA (cffDNA) and thus increase the performance of cancer diagnosis and NIPT by CNVs [85,86,87,88]. Moreover, the shorter fragment size in ctDNA was utilised to filter out the possible clonal haematopoiesis of indeterminate-potential (CHIP)-associated genetic variants and improve the classification performance for the genetic-based approaches [89, 90]. The non-random fragmentation and fragment-size information have recently been incorporated into the mutational calling algorithm specifically designed for the cfDNA [91].
Identification of tissues-of-origin by cfDNA fragmentomics
Different measurements of cfDNA fragmentomics described above are highly correlated with local and global epigenetic backgrounds, the patterns of which are known to be highly cell-type specific [92,93,94]. Therefore, it is possible to evaluate the tissues-of-origin by cfDNA fragmentomics.
DELFI, coverage near TFBS and cfDNA-fragmentation hotspots showed the potential to distinguish different cancer types in a supervised manner of machine learning but without providing the most relevant cell types contributed to cfDNA [64, 66, 68]. Preferred-end position, ended motif frequency and jagged end showed the potential for the estimation of the most relevant cell types but only demonstrated their correlations with the foetal sources in pregnant women, transplanted tissue source in organ- transplantation patients and tumour sources in cancer patients [70,71,72, 74].
Snyder et al. performed the fast Fourier transformation (FFT) on WPS signals in the first 10 kb of gene bodies [33]. Then they evaluated the correlations of the intensity of FFT signal against 76 gene expression datasets of human cell lines and primary tissues and found out that the most negatively correlated cell lines are haematopoietic lineages in healthy individuals. The intensity of FFT signals from the late-stage cancer patients showed the most positive correlations with cell lines from non-haematopoietic lineages. However, this WPS-based approach does not give the relative contributions from each cell type.
Sun et al. utilised OCF value around tissue-specific open-chromatin regions to solve this problem [67]. In healthy subjects, they observed the positive OCF values on open-chromatin regions from T cells and liver, and near or below zero on other tissue-specific open-chromatin regions. At cfDNA from pregnant women, liver transplantation and HCC patients, lung-cancer patients and colorectal-cancer patients, they observed the elevated OCF values on the placenta, liver, lungs or small intestine-specific open- chromatin regions, respectively. More importantly, the rank of the OCF value seems to provide the relative contributions from each cell type.
Liu et al. tried to estimate the absolute fractions of cell types that contributed to cfDNA by using A/B compartments inferred from FREE-C, which were represented as the linear combinations of 65 datasets from 18 tissues/cell types (Hi-C, H3K4me1 or WGBS data) [65]. However, the reference panel here is arbitrary and not completed yet, especially for the Hi-C data from different cell types. The inferred DNA methylation level from cfDNA seems to take advantage of the rich reference panel from DNA methylation and current well-established methylation-deconvolution methods to obtain the absolute contribution value from different cell types [69]. However, the inference accuracy of the methylation level in this approach is still not high enough for a robust and sensitive estimation of tissues-of-origin.
Overall, unlike the tissue-of-origin studies by cfDNA methylation in different pathological conditions [28, 95,96,97,98,99,100,101,102,103,104,105], the current tissue-of-origin analysis methods by cfDNA fragmentomics are still in their infant stage. An accurate and robust computational approach is still needed. Importantly, circulating cfDNA fragmentomics data from cancer patients contain the fragmentation information at DNA released from multiple cell types. A well-designed gold-standard system is needed for the complete benchmark and evaluation of the current tissue-of-origin analysis methods, including those based on cfDNA fragmentomics and methylation. Moreover, this is especially crucial in oncology for the fragmentation analysis in diagnosis and screening of cancers, given the presence of cfDNA deriving from a mixture of malignant, tumour-microenvironment and normal cells.
Discussion
In summary, the cfDNA fragmentomics showed the potential to characterise different molecular layers, especially epigenetics, in gene regulation within the cells. Therefore, circulating cfDNA in the peripheral blood offers a promising non-invasive approach to monitor the dynamics of genetics and epigenetics information together from multiple cell types in a single WGS experiment. CfDNA in peripheral blood is mainly from haematopoietic cells in healthy people and additional diseased tissues in patients [28, 44, 95]. Thus, cfDNA fragmentomics may inform the initiation and progression of complex diseases such as early-stage cancers and autoimmune diseases that are associated with genetic and epigenetic aberrations in both primarily affected tissues as well as multiple immune cells [106,107,108]. Moreover, cfDNA-fragmentomics study in cancer does not suffer the problem of CHIP-associated genetic variations, which are not specific to cancer but a normal phenotype of ageing [109,110,111].
Beyond the circulating blood, cfDNA also existed in many other bodily fluids, such as urine and cerebrospinal fluid (CSF) [112,113,114,115,116]. The cfDNA fragmentomics showed significantly different patterns at these bodily fluids. The comprehensive characterisation of cfDNA fragmentation and the related biological mechanisms at these bodily fluids are still largely unexplored. More interestingly, how the cfDNA fragmentomics evolves in the context of evolution across different species beyond the mice is still not known.
Circulating cfDNA showed different fragmentation patterns when released from cellular apoptosis or necrosis. Neutrophil extracellular DNA traps (NETs) were identified in 2004 [117]. It has been shown that NETosis may lead to the release of DNA without the cell lysing upon certain conditions [56, 118,119,120]. How does NETosis affect the fragmentation patterns, especially with the presence of different pathogens, will be an interesting question to explore.
Currently, some urgent computational and experimental-related questions need to be solved for cfDNA fragmentomics. For example, the cfDNA yield and fragment sizes varied significantly with different pre-analytical steps, for instance, the cfDNA-extraction kits [121]. The fragmentation patterns in plasma cfDNA will also be severely affected by the contamination of genomic DNA from white blood cells, which could be due to the delay between blood draw and plasma preparation, storage time, plasma centrifugation and preparation approach, and different storage tubes [122, 123]. How to compare the cfDNA fragmentomics at samples generated from different protocols is a big challenge. Moreover, the fragment coverage and sizes measured by NGS are known to be affected by PCR amplification, G + C% content, mappability and k-mer composition [33, 124, 125]. Several computational approaches have been proposed and applied to correct or normalise these technical artefacts for the coverage and sizes [33, 64, 124,125,126]. However, extensive benchmark and comparison is still needed to find the optimal approach for different fragmentomic analysis at different resolutions. For instance, whether or not these technical factors will affect the other fragmentomic approaches, such as the preferred end and end motif, is still not well understood. Further, the batch effect is the major hurdle for applying cfDNA fragmentomics in disease diagnosis and prognosis [127]. Research groups began to evaluate the importance of PCR-amplification bias for the application of DELFI on their follow-up cancer screening study [126]. The performance of most approaches mentioned in this review was evaluated by cross-validation from the same dataset. Whether or not the trained machine-learning model in one dataset can still be applied to other independent datasets is a challenging problem, especially for the multiple different measurements across different genomic resolutions. Moreover, a lot of fragmentation-based approaches were developed recently and need more follow-up studies from different labs to replicate their power in the prospective clinical studies. Although with some recent progress, such as cfDNApipe [128], the lack of well-documented and well-maintained open-source bioinformatic packages for cfDNA fragmentomics is still a hurdle for the field [57]. Finally, a limited number of fragmentomic features were integrated together to represent the overall fragmentation level, such as WPS to integrate fragment coverage and end [33]. However, unlike the integration-method development in single-cell multi-omics field, it is urgently needed in the cfDNA field that how to integrate multiple cfDNA fragmentomic features in the same dataset, as well as other cfDNA signals [22, 25, 26, 129, 130], and therefore boost the power for the disease diagnosis. Several recent efforts on the multi-modality integration across different resolutions of fragmentomics and with other measurements began to show their power on the cancer diagnosis, such as LIQUORICE and others [85, 89, 90, 126, 131].
In 2020, the American College of Obstetricians and Gynecologists’ (ACOG) guidelines recommended the NIPT for all pregnancies, regardless of risk, which could eventually generate millions of cfDNA WGS every year in the United States. Moreover, large-scale cfDNA WGS datasets have already been generated for cancer detection and many other purposes [132]. However, most cfDNA WGS datasets are deposited in the controlled-access repositories for the purpose of protecting patients’ genotype information, which is not needed for the cfDNA fragmentomic analysis. Due to the enormous commercial interests behind the cfDNA, the access of raw cfDNA NGS data in these repositories usually requires data-transfer agreements that may take several months of negotiations between the two organisations’ legal departments. Some initial efforts, such as FinaleDB, have been made recently to establish the cfDNA-fragmentomics database [133,134,135]. However, more community efforts are still needed to collect and uniformly process the comprehensive publicly available cfDNA datasets together with their rich metadata. A centralised and uniformly processed cfDNA-fragmentomics database, similar to the ENCODE [136], will finally benefit this community in the long run.
In the end, a large number of cfDNA WGS and their associated clinical metadata from individuals at different physiological conditions will eventually allow us to characterise the baseline of the cfDNA fragmentomics and their variations in the population level at both healthy and pathological conditions.
Data availability
Not applicable.
References
Thurman RE, Rynes E, Humbert R, Vierstra J, Maurano MT, Haugen E, et al. The accessible chromatin landscape of the human genome. Nature. 2012;489:75–82. https://doi.org/10.1038/nature11232.
Jones PA. Functions of DNA methylation: islands, start sites, gene bodies and beyond. Nat Rev Genet. 2012;13:484–92. https://doi.org/10.1038/nrg3230.
Li Y, Hu M, Shen Y. Gene regulation in the 3D genome. Hum Mol Genet. 2018;27:R228–33. https://doi.org/10.1093/hmg/ddy164.
Jones PA, Baylin SB. The epigenomics of cancer. Cell. 2007;128:683–92. https://doi.org/10.1016/j.cell.2007.01.029.
Corces MR, Granja JM, Shams S, Louie BH, Seoane JA, Zhou W, et al. The chromatin accessibility landscape of primary human cancers. Science 2018;362. https://doi.org/10.1126/science.aav1898.
Xu Q, Xie W. Epigenome in early mammalian development: inheritance, reprogramming and establishment. Trends Cell Biol. 2018;28:237–53. https://doi.org/10.1016/j.tcb.2017.10.008.
Zheng H, Xie W. The role of 3D genome organization in development and cell differentiation. Nat Rev Mol Cell Biol. 2019;20:535–50. https://doi.org/10.1038/s41580-019-0132-4.
Hug CB, Vaquerizas JM. The birth of the 3D Genome during early embryonic development. Trends Genet. 2018;34:903–14. https://doi.org/10.1016/j.tig.2018.09.002.
Mas VR, Le TH, Maluf DG. Epigenetics in kidney transplantation: current evidence, predictions, and future research directions. Transplantation. 2016;100:23–38. https://doi.org/10.1097/TP.0000000000000878.
Wu J, Huang B, Chen H, Yin Q, Liu Y, Xiang Y, et al. The landscape of accessible chromatin in mammalian preimplantation embryos. Nature. 2016;534:652–7. https://doi.org/10.1038/nature18606.
Cardoso-Moreira M, Halbert J, Valloton D, Velten B, Chen C, Shao Y, et al. Gene expression across mammalian organ development. Nature. 2019;571:505–9. https://doi.org/10.1038/s41586-019-1338-5.
Ke Y, Xu Y, Chen X, Feng S, Liu Z, Sun Y, et al. 3D Chromatin structures of mature gametes and structural reprogramming during mammalian embryogenesis. Cell. 2017;170:367–81.e20. https://doi.org/10.1016/j.cell.2017.06.029.
Cui C, Shu W, Li P. Fluorescence in situ hybridization: cell-based genetic diagnostic and research applications. Front Cell Dev Biol. 2016;4:89 https://doi.org/10.3389/fcell.2016.00089.
Bintu B, Mateo LJ, Su J-H, Sinnott-Armstrong NA, Parker M, Kinrot S, et al. Super-resolution chromatin tracing reveals domains and cooperative interactions in single cells. Science 2018;362. https://doi.org/10.1126/science.aau1783.
Barth R, Bystricky K, Shaban HA. Coupling chromatin structure and dynamics by live super-resolution imaging. Sci Adv 2020;6. https://doi.org/10.1126/sciadv.aaz2196.
Heitzer E, Haque IS, Roberts CES, Speicher MR. Current and future perspectives of liquid biopsies in genomics-driven oncology. Nat Rev Genet. 2019;20:71–88. https://doi.org/10.1038/s41576-018-0071-5.
Norwitz ER, Levy B. Noninvasive prenatal testing: the future is now. Rev Obstet Gynecol. 2013;6:48–62. https://www.ncbi.nlm.nih.gov/pubmed/24466384.
Knight SR, Thorne A, Lo Faro ML. Donor-specific Cell-free DNA as a biomarker in solid organ transplantation. a systematic review. Transplantation. 2019;103:273–83. https://doi.org/10.1097/TP.0000000000002482.
Kitzman JO, Snyder MW, Ventura M, Lewis AP, Qiu R, Simmons LE, et al. Noninvasive whole-genome sequencing of a human fetus. Sci Transl Med. 2012;4:137ra76 https://doi.org/10.1126/scitranslmed.3004323.
Fan HC, Gu W, Wang J, Blumenfeld YJ, El-Sayed YY, Quake SR. Non-invasive prenatal measurement of the fetal genome. Nature. 2012;487:320–4. https://doi.org/10.1038/nature11251.
Keller L, Belloum Y, Wikman H, Pantel K. Clinical relevance of blood-based ctDNA analysis: mutation detection and beyond. Br J Cancer. 2021;124:345–58. https://doi.org/10.1038/s41416-020-01047-5.
Adalsteinsson VA, Ha G, Freeman SS, Choudhury AD, Stover DG, Parsons HA, et al. Scalable whole-exome sequencing of cell-free DNA reveals high concordance with metastatic tumors. Nat Commun. 2017;8:1324 https://doi.org/10.1038/s41467-017-00965-y.
Zhang J, Li J, Saucier JB, Feng Y, Jiang Y, Sinson J, et al. Non-invasive prenatal sequencing for multiple Mendelian monogenic disorders using circulating cell-free fetal DNA. Nat Med. 2019;25:439–47. https://doi.org/10.1038/s41591-018-0334-x.
Peters D, Chu T, Yatsenko SA, Hendrix N, Hogge WA, Surti U, et al. Noninvasive prenatal diagnosis of a fetal microdeletion syndrome. N Engl J Med. 2011;365:1847–8. https://doi.org/10.1056/NEJMc1106975.
Erger F, Nörling D, Borchert D, Leenen E, Habbig S, Wiesener MS, et al. cfNOMe—a single assay for comprehensive epigenetic analyses of cell-free DNA. Genome Medicine. 2020;12. https://doi.org/10.1186/s13073-020-00750-5.
Sadeh R, Sharkia I, Fialkoff G, Rahat A, Gutin J, Chappleboim A, et al. ChIP-seq of plasma cell-free nucleosomes identifies gene expression programs of the cells of origin. Nat Biotechnol. 2021; https://doi.org/10.1038/s41587-020-00775-6.
Lun FMF, Chiu RWK, Sun K, Leung TY, Jiang P, Chan KCA, et al. Noninvasive prenatal methylomic analysis by genomewide bisulfite sequencing of maternal plasma DNA. Clin Chem. 2013;59:1583–94. https://doi.org/10.1373/clinchem.2013.212274.
Sun K, Jiang P, Chan KCA, Wong J, Cheng YKY, Liang RHS, et al. Plasma DNA tissue mapping by genome-wide methylation sequencing for noninvasive prenatal, cancer, and transplantation assessments. Proc Natl Acad Sci USA. 2015;112:E5503–12. https://doi.org/10.1073/pnas.1508736112.
Kang S, Li Q, Chen Q, Zhou Y, Park S, Lee G, et al. CancerLocator: non-invasive cancer diagnosis and tissue-of-origin prediction using methylation profiles of cell-free DNA. Genome Biol. 2017;18:53 https://doi.org/10.1186/s13059-017-1191-5.
Li W, Zhou XJ. Methylation extends the reach of liquid biopsy in cancer detection. Nat Rev Clin Oncol 2020;17:655–6. https://doi.org/10.1038/s41571-020-0420-0.
Li W, Li Q, Kang S, Same M, Zhou Y, Sun C, et al. CancerDetector: ultrasensitive and non-invasive cancer detection at the resolution of individual reads using cell-free DNA methylation sequencing data. Nucleic Acids Res. 2018;46:e89 https://doi.org/10.1093/nar/gky423.
Ivanov M, Baranova A, Butler T, Spellman P, Mileyko V. Non-random fragmentation patterns in circulating cell-free DNA reflect epigenetic regulation. BMC Genomics. 2015;16(Suppl 13):S1 https://doi.org/10.1186/1471-2164-16-S13-S1.
Snyder MW, Kircher M, Hill AJ, Daza RM, Shendure J. Cell-free DNA comprises an in vivo nucleosome footprint that Informs its tissues-of-origin. Cell. 2016;164:57–68. https://doi.org/10.1016/j.cell.2015.11.050.
Ulz P, Thallinger GG, Auer M, Graf R, Kashofer K, Jahn SW, et al. Inferring expressed genes by whole-genome sequencing of plasma DNA. Nat Genet. 2016;48:1273–8. https://doi.org/10.1038/ng.3648.
Mandel P, Metais P. Nuclear acids in human blood plasma. C R Seances Soc Biol Fil. 1948;142:241–3. https://www.ncbi.nlm.nih.gov/pubmed/18875018.https://www.ncbi.nlm.nih.gov/pubmed/18875018.
Tan EM, Schur PH, Carr RI, Kunkel HG. Deoxybonucleic acid (DNA) and antibodies to DNA in the serum of patients with systemic lupus erythematosus. J Clin Invest. 1966;45:1732–40. https://doi.org/10.1172/JCI105479.
Koffler D, Agnello V, Winchester R, Kunkel HG. The occurrence of single-stranded DNA in the serum of patients with systemic lupus erythematosus and other diseases. J Clin Invest. 1973;52:198–204. https://doi.org/10.1172/JCI107165.
Emlen W, Mannik M. Effect of DNA size and strandedness on the in vivo clearance and organ localization of DNA. Clin Exp Immunol. 1984;56:185–92. https://www.ncbi.nlm.nih.gov/pubmed/6609032.
Stroun M, Anker P, Lyautey J, Lederrey C, Maurice PA. Isolation and characterization of DNA from the plasma of cancer patients. Eur J Cancer Clin Oncol. 1987;23:707–12. https://doi.org/10.1016/0277-5379(87)90266-5.
Jahr S, Hentze H, Englisch S, Hardt D, Fackelmayer FO, Hesch RD, et al. DNA fragments in the blood plasma of cancer patients: quantitations and evidence for their origin from apoptotic and necrotic cells. Cancer Res. 2001;61:1659–65. https://www.ncbi.nlm.nih.gov/pubmed/11245480.
Anker P, Mulcahy H, Chen XQ, Stroun M. Detection of circulating tumour DNA in the blood (plasma/serum) of cancer patients. Cancer Metastasis Rev. 1999;18:65–73. https://doi.org/10.1023/a:1006260319913.
Lo YMD, Chan KCA, Sun H, Chen EZ, Jiang P, Lun FMF, et al. Maternal plasma DNA sequencing reveals the genome-wide genetic and mutational profile of the fetus. Sci Transl Med. 2010;2:61ra91 https://doi.org/10.1126/scitranslmed.3001720.
Zheng YWL, Chan KCA, Sun H, Jiang P, Su X, Chen EZ, et al. Nonhematopoietically derived DNA is shorter than hematopoietically derived DNA in plasma: a transplantation model. Clin Chem. 2012;58:549–58. https://doi.org/10.1373/clinchem.2011.169318.
Lui YYN, Chik K-W, Chiu RWK, Ho C-Y, Lam CWK, Lo YMD. Predominant hematopoietic origin of cell-free DNA in plasma and serum after sex-mismatched bone marrow transplantation. Clin Chem. 2002;48:421–7. https://www.ncbi.nlm.nih.gov/pubmed/11861434.
Deligezer U, Yaman F, Erten N, Dalay N. Frequent copresence of methylated DNA and fragmented nucleosomal DNA in plasma of lymphoma patients. Clin Chim Acta. 2003;335:89–94. https://doi.org/10.1016/s0009-8981(03)00279-1.
Chan KCA, Zhang J, Hui ABY, Wong N, Lau TK, Leung TN, et al. Size distributions of maternal and fetal DNA in maternal plasma. Clin Chem. 2004;50:88–92. https://doi.org/10.1373/clinchem.2003.024893.
Diehl F, Li M, Dressman D, He Y, Shen D, Szabo S, et al. Detection and quantification of mutations in the plasma of patients with colorectal tumors. Proc Natl Acad Sci USA. 2005;102:16368–73. https://doi.org/10.1073/pnas.0507904102.
Giacona MB, Ruben GC, Iczkowski KA, Roos TB, Porter DM, Sorenson GD. Cell-free DNA in human blood plasma: length measurements in patients with pancreatic cancer and healthy controls. Pancreas. 1998;17:89–97. https://doi.org/10.1097/00006676-199807000-00012.
Mouliere F, Robert B, Arnau Peyrotte E, Del Rio M, Ychou M, Molina F, et al. High fragmentation characterizes tumour-derived circulating DNA. PLoS ONE. 2011;6:e23418 https://doi.org/10.1371/journal.pone.0023418.
Sanchez C, Snyder MW, Tanos R, Shendure J, Thierry AR. New insights into structural features and optimal detection of circulating tumor DNA determined by single-strand DNA analysis. NPJ Genom Med. 2018;3:31 https://doi.org/10.1038/s41525-018-0069-0
Meddeb R, Dache ZAA, Thezenas S, Otandault A, Tanos R, Pastor B, et al. Quantifying circulating cell-free DNA in humans. Sci Rep. 2019;9:5220 https://doi.org/10.1038/s41598-019-41593-4.
Shemer R, Magenheim J, Dor Y. Digital droplet PCR for monitoring tissue-specific cell death using DNA methylation patterns of circulating cell-free DNA. Curr Protoc Mol Biol. 2019;127:e90 https://doi.org/10.1002/cpmb.90.
Andriamanampisoa C-L, Bancaud A, Boutonnet-Rodat A, Didelot A, Fabre J, Fina F, et al. BIABooster: online DNA concentration and size profiling with a limit of detection of 10 fg/μL and application to high-sensitivity characterization of circulating cell-free DNA. Anal Chem. 2018;90:3766–74. https://doi.org/10.1021/acs.analchem.7b04034.
Ito H, Hasegawa K, Hasegawa Y, Nishimaki T, Hosomichi K, Kimura S, et al. Silver nanoscale hexagonal column chips for detecting cell-free DNA and circulating nucleosomes in cancer patients. Sci Rep. 2015;5:10455 https://doi.org/10.1038/srep10455.
Pang D, Thierry AR, Dritschilo A. DNA studies using atomic force microscopy: capabilities for measurement of short DNA fragments. Front Mol Biosci. 2015;2:1 https://doi.org/10.3389/fmolb.2015.00001.
Lo YMD, Han DSC, Jiang P, Chiu RWK. Epigenetics, fragmentomics, and topology of cell-free DNA in liquid biopsies. Science 2021;372. https://doi.org/10.1126/science.aaw3616.
Chiu RWK, Heitzer E, Lo YMD, Mouliere F, Tsui DWY, Cell-Free DNA. Fragmentomics: the New ‘Omics’ on the Block. Clin Chem. 2020;66:1480–4. https://doi.org/10.1093/clinchem/hvaa258.
Heitzer E, Auinger L, Speicher MR. Cell-free DNA and apoptosis: how dead cells inform about the living. Trends Mol Med. 2020;26:519–28. https://doi.org/10.1016/j.molmed.2020.01.012.
Heitzer E, Perakis S, Geigl JB, Speicher MR. The potential of liquid biopsies for the early detection of cancer. NPJ Precis Oncol. 2017;1:36 https://doi.org/10.1038/s41698-017-0039-5.
Underhill HR, Kitzman JO, Hellwig S, Welker NC, Daza R, Baker DN, et al. Fragment length of circulating tumor DNA. PLoS Genet. 2016;12:e1006162 https://doi.org/10.1371/journal.pgen.1006162.
Chandrananda D, Thorne NP, Bahlo M. High-resolution characterization of sequence signatures due to non-random cleavage of cell-free DNA. BMC Med Genomics. 2015;8:29 https://doi.org/10.1186/s12920-015-0107-z.
Fan HC, Blumenfeld YJ, Chitkara U, Hudgins L, Quake SR. Noninvasive diagnosis of fetal aneuploidy by shotgun sequencing DNA from maternal blood. Proc Natl Acad Sci USA. 2008;105:16266–71. https://doi.org/10.1073/pnas.0808319105.
Fan HC, Blumenfeld YJ, Chitkara U, Hudgins L, Quake SR. Analysis of the size distributions of fetal and maternal cell-free DNA by paired-end sequencing. Clin Chem. 2010;56:1279–86. https://doi.org/10.1373/clinchem.2010.144188.
Cristiano S, Leal A, Phallen J, Fiksel J, Adleff V, Bruhm DC, et al. Genome-wide cell-free DNA fragmentation in patients with cancer. Nature. 2019;570:385–9. https://doi.org/10.1038/s41586-019-1272-6
Liu Y, Liu T-Y, Weinberg DE, White BW, De La Torre CJ, Tan CL, et al. Spatial co-fragmentation pattern of cell-free DNA recapitulates in vivo chromatin organization and identifies tissues-of-origin. bioRxiv. 2019;564773. Preprint at https://doi.org/10.1101/564773.
Ulz P, Perakis S, Zhou Q, Moser T, Belic J, Lazzeri I, et al. Inference of transcription factor binding from cell-free DNA enables tumor subtype prediction and early detection. Nat Commun. 2019;10:4666 https://doi.org/10.1038/s41467-019-12714-4.
Sun K, Jiang P, Cheng SH, Cheng THT, Wong J, Wong VWS, et al. Orientation-aware plasma cell-free DNA fragmentation analysis in open chromatin regions informs tissue of origin. Genome Res. 2019;29:418–27. https://doi.org/10.1101/gr.242719.118.
Zhou X, Liu Y. De novo characterization of cell-free DNA fragmentation hotspots boosts the power for early detection and localization of multi-cancer. biorxiv. 2020;2020.07.16.201350. Preprint at https://doi.org/10.1101/2020.07.16.201350.
Liu Y, Reed S, Choudhury AD, Parsons HA, Stover DG, Ha G, et al. Abstract 5689: Identify tissue-of-origin in cancer cfDNA by whole genome sequencing. Cancer Res. 2017;77:5689–5689. https://doi.org/10.1158/1538-7445.AM2017-5689.
Chan KCA, Jiang P, Sun K, Cheng YKY, Tong YK, Cheng SH, et al. Second generation noninvasive fetal genome analysis reveals de novo mutations, single-base parental inheritance, and preferred DNA ends. Proc Natl Acad Sci USA. 2016;113:E8159–68. https://doi.org/10.1073/pnas.1615800113.
Jiang P, Sun K, Tong YK, Cheng SH, Cheng THT, Heung MMS, et al. Preferred end coordinates and somatic variants as signatures of circulating tumor DNA associated with hepatocellular carcinoma. Proc Natl Acad Sci USA. 2018;115:E10925–33. https://doi.org/10.1073/pnas.1814616115.
Jiang P, Sun K, Peng W, Cheng SH, Ni M, Yeung PC, et al. Plasma DNA end-motif profiling as a fragmentomic marker in cancer, pregnancy, and transplantation. Cancer Disco. 2020;10:664–73. https://doi.org/10.1158/2159-8290.CD-19-0622.
Suzuki N, Kamataki A, Yamaki J, Homma Y. Characterization of circulating DNA in healthy human plasma. Clin Chim Acta. 2008;387:55–8. https://doi.org/10.1016/j.cca.2007.09.001.
Jiang P, Xie T, Ding SC, Zhou Z, Cheng SH, Chan RWY, et al. Detection and characterization of jagged ends of double-stranded DNA in plasma. Genome Res. 2020;30:1144–53. https://doi.org/10.1101/gr.261396.120.
An Q, Hu Y, Li Q, Chen X, Huang J, Pellegrini M, et al. The size of cell-free mitochondrial DNA in blood is inversely correlated with tumor burden in cancer patients. Precis Clin Med. 2019;2:131–9. https://doi.org/10.1093/pcmedi/pbz014.
Zhu J, Zhang F, Du M, Zhang P, Fu S, Wang L. Molecular characterization of cell-free eccDNAs in human plasma. Sci Rep. 2017;7:10968 https://doi.org/10.1038/s41598-017-11368-w.
Kumar P, Dillon LW, Shibata Y, Jazaeri AA, Jones DR, Dutta A. Normal and cancerous tissues release extrachromosomal circular DNA (eccDNA) into the circulation. Mol Cancer Res. 2017;15:1197–205. https://doi.org/10.1158/1541-7786.MCR-17-0095.
Sin STK, Jiang P, Deng J, Ji L, Cheng SH, Dutta A, et al. Identification and characterization of extrachromosomal circular DNA in maternal plasma. Proc Natl Acad Sci USA. 2020;117:1658–65. https://doi.org/10.1073/pnas.1914949117.
Jensen TJ, Kim SK, Zhu Z, Chin C, Gebhard C, Lu T, et al. Whole genome bisulfite sequencing of cell-free DNA and its cellular contributors uncovers placenta hypomethylated domains. Genome Biol. 2015;16:78 https://doi.org/10.1186/s13059-015-0645-x.
Gansauge M-T, Meyer M. Single-stranded DNA library preparation for the sequencing of ancient or damaged DNA. Nat Protoc. 2013;8:737–48. https://doi.org/10.1038/nprot.2013.038.
Sanchez C, Roch B, Mazard T, Blache P, Dache ZAA, Pastor B, et al. Circulating nuclear DNA structural features, origins, and complete size profile revealed by fragmentomics. JCI Insight 2021;6. https://doi.org/10.1172/jci.insight.144561.
Sun K, Jiang P, Wong AIC, Cheng YKY, Cheng SH, Zhang H, et al. Size-tagged preferred ends in maternal plasma DNA shed light on the production mechanism and show utility in noninvasive prenatal testing. Proc Natl Acad Sci USA. 2018;115:E5106–14. https://doi.org/10.1073/pnas.1804134115.
Serpas L, Chan RWY, Jiang P, Ni M, Sun K, Rashidfarrokhi A, et al. Dnase1l3 deletion causes aberrations in length and end-motif frequencies in plasma DNA. Proc Natl Acad Sci USA. 2019;116:641–9. https://doi.org/10.1073/pnas.1815031116.
Chan RWY, Serpas L, Ni M, Volpi S, Hiraki LT, Tam L-S, et al. Plasma DNA profile associated with DNASE1L3 gene mutations: clinical observations, relationships to nuclease substrate preference, and in vivo correction. Am J Hum Genet. 2020;107:882–94. https://doi.org/10.1016/j.ajhg.2020.09.006.
Mouliere F, Chandrananda D, Piskorz AM, Moore EK, Morris J, Ahlborn LB, et al. Enhanced detection of circulating tumor DNA by fragment size analysis. Sci Transl Med. 2018;10. https://doi.org/10.1126/scitranslmed.aat4921.
Yang Q, Du Z, Song Y, Gao S, Yu S, Zhu H, et al. Size-selective separation and overall-amplification of cell-free fetal DNA fragments using PCR-based enrichment. Sci Rep. 2017;7:40936 https://doi.org/10.1038/srep40936.
Liang B, Li H, He Q, Li H, Kong L, Xuan L, et al. Enrichment of the fetal fraction in non-invasive prenatal screening reduces maternal background interference. Sci Rep. 2018;8:17675 https://doi.org/10.1038/s41598-018-35738-0.
Marass F, Stephens D, Ptashkin R, Zehir A, Berger MF, Solit DB, et al. Fragment size analysis may distinguish clonal hematopoiesis from tumor-derived mutations in cell-free DNA. Clin Chem. 2020;66:616–8. https://doi.org/10.1093/clinchem/hvaa026.
Wan JCM, Heider K, Gale D, Murphy S, Fisher E, Mouliere F, et al. ctDNA monitoring using patient-specific sequencing and integration of variant reads. Sci Transl Med. 2020;12. https://doi.org/10.1126/scitranslmed.aaz8084.
Chabon JJ, Hamilton EG, Kurtz DM, Esfahani MS, Moding EJ, Stehr H, et al. Integrating genomic features for non-invasive early lung cancer detection. Nature. 2020;580:245–51. https://doi.org/10.1038/s41586-020-2140-0.
Li S, Noor ZS, Zeng W, Stackpole ML, Ni X, Zhou Y, et al. Sensitive detection of tumor mutations from blood and its application to immunotherapy prognosis. Nat Commun. 2021;12:4172.
Roadmap Epigenomics Consortium KundajeA, Meuleman W, Ernst J, Bilenky M, Yen A, et al. Integrative analysis of 111 reference human epigenomes. Nature. 2015;518:317–30. https://doi.org/10.1038/nature14248.
Stunnenberg HG, International Human Epigenome Consortium, Hirst M. The International Human Epigenome Consortium: A Blueprint for Scientific Collaboration and Discovery. Cell. 2016;167:1145–9. https://doi.org/10.1016/j.cell.2016.11.007.
Stricker SH, Köferle A, Beck S. From profiles to function in epigenomics. Nat Rev Genet. 2017;18:51–66. https://doi.org/10.1038/nrg.2016.138.
Moss J, Magenheim J, Neiman D, Zemmour H, Loyfer N, Korach A, et al. Comprehensive human cell-type methylation atlas reveals origins of circulating cell-free DNA in health and disease. Nat Commun. 2018;9:5068 https://doi.org/10.1038/s41467-018-07466-6.
Shen SY, Singhania R, Fehringer G, Chakravarthy A, Roehrl MHA, Chadwick D, et al. Sensitive tumour detection and classification using plasma cell-free DNA methylomes. Nature. 2018;563:579–83. https://doi.org/10.1038/s41586-018-0703-0.
Lehmann-Werman R, Neiman D, Zemmour H, Moss J, Magenheim J, Vaknin-Dembinsky A, et al. Identification of tissue-specific cell death using methylation patterns of circulating DNA. Proc Natl Acad Sci USA. 2016;113:E1826–34. https://doi.org/10.1073/pnas.1519286113.
Gala-Lopez BL, Neiman D, Kin T, O’Gorman D, Pepper AR, Malcolm AJ, et al. Beta cell death by cell-free DNA and outcome after clinical islet transplantation. Transplantation. 2018;102:978–85. https://doi.org/10.1097/TP.0000000000002083.
Akirav EM, Lebastchi J, Galvan EM, Henegariu O, Akirav M, Ablamunits V, et al. Detection of β cell death in diabetes using differentially methylated circulating DNA. Proc Natl Acad Sci USA. 2011;108:19018–23. https://doi.org/10.1073/pnas.1111008108.
Lebastchi J, Deng S, Lebastchi AH, Beshar I, Gitelman S, Willi S, et al. Immune therapy and β-cell death in type 1. Diabetes Diabetes. 2013;62:1676–80. https://doi.org/10.2337/db12-1207.
Lam WKJ, Gai W, Sun K, Wong RSM, Chan RWY, Jiang P, et al. DNA of erythroid origin is present in human plasma and informs the types of anemia. Clin Chem. 2017;63:1614–23. https://doi.org/10.1373/clinchem.2017.272401.
Gai W, Ji L, Jacky Lam WK, Sun K, Jiang P, Chan AWH, et al. Liver- and Colon-Specific DNA Methylation Markers in Plasma for Investigation of Colorectal Cancers with or without Liver Metastases. Clin Chem. 2018;64:1239–49. https://doi.org/10.1373/clinchem.2018.290304.
Zemmour H, Planer D, Magenheim J, Moss J, Neiman D, Gilon D, et al. Non-invasive detection of human cardiomyocyte death using methylation patterns of circulating DNA. Nat Commun. 2018;9:1443 https://doi.org/10.1038/s41467-018-03961-y.
Lehmann-Werman R, Magenheim J, Moss J, Neiman D, Abraham O, Piyanzin S, et al. Monitoring liver damage using hepatocyte-specific methylation markers in cell-free circulating DNA. JCI Insight. 2018;3. https://doi.org/10.1172/jci.insight.120687.
Liu MC, Oxnard GR, Klein EA, Swanton C, Seiden MV, CCGA Consortium. Sensitive and specific multi-cancer detection and localization using methylation signatures in cell-free DNA. Ann Oncol. 2020;31:745–59. https://doi.org/10.1016/j.annonc.2020.02.011.
Hanahan D, Weinberg RA. Hallmarks of cancer: the next generation. Cell. 2011;144:646–74. https://doi.org/10.1016/j.cell.2011.02.013.
Gjoneska E, Pfenning AR, Mathys H, Quon G, Kundaje A, Tsai L-H, et al. Conserved epigenomic signals in mice and humans reveal immune basis of Alzheimer’s disease. Nature. 2015;518:365–9. https://doi.org/10.1038/nature14252.
Nowling TK, Gilkeson GS. Mechanisms of tissue injury in lupus nephritis. Arthritis Res Ther. 2011;13:250 https://doi.org/10.1186/ar3528.
Abbosh C, Swanton C, Birkbak NJ. Clonal haematopoiesis: a source of biological noise in cell-free DNA analyses. Ann Oncol 2019;30:358–9. https://doi.org/10.1093/annonc/mdy552.
Genovese G, Kähler AK, Handsaker RE, Lindberg J, Rose SA, Bakhoum SF, et al. Clonal hematopoiesis and blood-cancer risk inferred from blood DNA sequence. N Engl J Med. 2014;371:2477–87. https://doi.org/10.1056/NEJMoa1409405.
Hu Y, Ulrich BC, Supplee J, Kuang Y, Lizotte PH, Feeney NB, et al. False-positive plasma genotyping due to clonal hematopoiesis. Clin Cancer Res. 2018;24:4437–43. https://doi.org/10.1158/1078-0432.CCR-18-0143.
Koide K, Sekizawa A, Iwasaki M, Matsuoka R, Honma S, Farina A, et al. Fragmentation of cell-free fetal DNA in plasma and urine of pregnant women. Prenat Diagn. 2005;25:604–7. https://doi.org/10.1002/pd.1213.
Cheng THT, Jiang P, Tam JCW, Sun X, Lee W-S, Yu SCY, et al. Genomewide bisulfite sequencing reveals the origin and time-dependent fragmentation of urinary cfDNA. Clin Biochem. 2017;50:496–501. https://doi.org/10.1016/j.clinbiochem.2017.02.017.
Burnham P, Dadhania D, Heyang M, Chen F, Westblade LF, Suthanthiran M, et al. Urinary cell-free DNA is a versatile analyte for monitoring infections of the urinary tract. Nat Commun. 2018;9:2412 https://doi.org/10.1038/s41467-018-04745-0.
Markus H, Zhao J, Contente-Cuomo T, Stephens MD, Raupach E, Odenheimer-Bergman A, et al. Analysis of recurrently protected genomic regions in cell-free DNA found in urine. Sci Transl Med. 2021;13. https://doi.org/10.1126/scitranslmed.aaz3088.
Mouliere F, Mair R, Chandrananda D, Marass F, Smith CG, Su J, et al. Detection of cell-free DNA fragmentation and copy number alterations in cerebrospinal fluid from glioma patients. EMBO Mol Med. 2018;10. https://doi.org/10.15252/emmm.201809323.
Brinkmann V, Reichard U, Goosmann C, Fauler B, Uhlemann Y, Weiss DS, et al. Neutrophil extracellular traps kill bacteria. Science. 2004;303:1532–5. https://doi.org/10.1126/science.1092385.
Yipp BG, Kubes P. NETosis: how vital is it? Blood. 2013;122:2784–94. https://doi.org/10.1182/blood-2013-04-457671.
Thierry AR, El Messaoudi S, Gahan PB, Anker P, Stroun M. Origins, structures, and functions of circulating DNA in oncology. Cancer Metastasis Rev. 2016;35:347–76. https://doi.org/10.1007/s10555-016-9629-x.
Jiménez-Alcázar M, Rangaswamy C, Panda R, Bitterling J, Simsek YJ, Long AT, et al. Host DNases prevent vascular occlusion by neutrophil extracellular traps. Science. 2017;358:1202–6. https://doi.org/10.1126/science.aam8897.
Markus H, Contente-Cuomo T, Farooq M, Liang WS, Borad MJ, Sivakumar S, et al. Evaluation of pre-analytical factors affecting plasma DNA analysis. Sci Rep. 2018;8:7375 https://doi.org/10.1038/s41598-018-25810-0.
El Messaoudi S, Rolet F, Mouliere F, Thierry AR. Circulating cell free DNA: preanalytical considerations. Clin Chim Acta. 2013;424:222–30. https://doi.org/10.1016/j.cca.2013.05.022.
Meddeb R, Pisareva E, Thierry AR. Guidelines for the preanalytical conditions for analyzing circulating cell-free DNA. Clin Chem. 2019;65:623–33. https://doi.org/10.1373/clinchem.2018.298323.
Benjamini Y, Speed TP. Summarizing and correcting the GC content bias in high-throughput sequencing. Nucleic Acids Res. 2012;40:e72 https://doi.org/10.1093/nar/gks001.
Chandrananda D, Thorne NP, Ganesamoorthy D, Bruno DL, Benjamini Y, Speed TP, et al. Investigating and correcting plasma DNA sequencing coverage bias to enhance aneuploidy discovery. PLoS ONE. 2014;9:e86993 https://doi.org/10.1371/journal.pone.0086993.
Mathios D, Johansen JS, Cristiano S, Medina JE, Phallen J, Larsen KR, et al. Detection and characterization of lung cancer using cell-free DNA fragmentomes. Nat Commun. 2021;12:5060 https://doi.org/10.1038/s41467-021-24994-w.
Wan N, Weinberg D, Liu T-Y, Niehaus K, Ariazi EA, Delubac D, et al. Machine learning enables detection of early-stage colorectal cancer by whole-genome sequencing of plasma cell-free DNA. BMC Cancer. 2019;19:832 https://doi.org/10.1186/s12885-019-6003-8.
Zhang W, Wei L, Huang J, Zhong B, Li J, Xu H, et al. cfDNApipe: a comprehensive quality control and analysis pipeline for cell-free DNA high-throughput sequencing data. Bioinformatics. 2021; https://doi.org/10.1093/bioinformatics/btab413.
Kinde I, Wu J, Papadopoulos N, Kinzler KW, Vogelstein B. Detection and quantification of rare mutations with massively parallel sequencing. Proc Natl Acad Sci USA. 2011;108:9530–5. https://doi.org/10.1073/pnas.1105422108.
Newman AM, Bratman SV, To J, Wynne JF, Eclov NCW, Modlin LA, et al. An ultrasensitive method for quantitating circulating tumor DNA with broad patient coverage. Nat Med. 2014;20:548–54. https://doi.org/10.1038/nm.3519.
Peneder P, Stütz AM, Surdez D, Krumbholz M, Semper S, Chicard M, et al. Multimodal analysis of cell-free DNA whole-genome sequencing for pediatric cancers with low mutational burden. Nat Commun. 2021;12:3230 https://doi.org/10.1038/s41467-021-23445-w.
Liu S, Huang S, Chen F, Zhao L, Yuan Y, Francis SS, et al. Genomic analyses from non-invasive prenatal testing reveal genetic associations, patterns of viral infections, and chinese population history. Cell. 2018;175:347–59.e14. https://doi.org/10.1016/j.cell.2018.08.016.
Yu F, Li K, Li S, Liu J, Zhang Y, Zhou M, et al. CFEA: a cell-free epigenome atlas in human diseases. Nucleic Acids Res. 2020;48:D40–4. https://doi.org/10.1093/nar/gkz715.
Zheng H, Zhu MS, Liu Y. FinaleDB: a browser and database of cell-free DNA fragmentation patterns. Bioinformatics. 2020; https://doi.org/10.1093/bioinformatics/btaa999.
Teif VB, Clarkson CT Nucleosome Positioning. In: Ranganathan S, Gribskov M, Nakai K, Schönbach C, eds. Encyclopedia of Bioinformatics and Computational Biology. Oxford: Academic Press 2019. 308–17. https://doi.org/10.1016/B978-0-12-809633-8.20242-2.
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature. 2012;489:57–74. https://doi.org/10.1038/nature11247.
Acknowledgements
We thank Dr. Li Wang and all the four reviewers for the critical reviews of this article.
Funding
Y.L. is supported by his startup grant, trustee award, and innovation fund from Cincinnati Children’s Hospital Medical Center, CCTST-mentored pilot translational award from the University of Cincinnati and MOMI Ideas Fund from Bill & Melinda Gates Foundation.
Author information
Authors and Affiliations
Contributions
Y.L. contributed to the design and writing of this review.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
Not applicable.
Consent to publish
Not applicable.
Competing interests
YL has an approved patent (“Methods for genome characterization”, US Patent App. 16/323,158, filed by MIT and Broad Institute of MIT and Harvard) and three provisional patents filed (“METHODS AND SYSTEMS FOR INFERRING CHROMOSOMAL STRUCTURE” and “METHODS AND SYSTEMS FOR ANALYZING MICROBIOTA” filed by Freenome Inc. “Cell-free DNA fragmentation pattern for cancer early detection” filed by Cincinnati Children’s Hospital Medical Center). YL owns stocks from Freenome Inc.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liu, Y. At the dawn: cell-free DNA fragmentomics and gene regulation. Br J Cancer 126, 379–390 (2022). https://doi.org/10.1038/s41416-021-01635-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-021-01635-z
This article is cited by
-
FinaleMe: Predicting DNA methylation by the fragmentation patterns of plasma cell-free DNA
Nature Communications (2024)
-
Cell type signatures in cell-free DNA fragmentation profiles reveal disease biology
Nature Communications (2024)
-
Circulating cell-free DNA fragmentation is a stepwise and conserved process linked to apoptosis
BMC Biology (2023)
-
Liquid biopsy for brain metastases and leptomeningeal disease in patients with breast cancer
npj Breast Cancer (2023)
-
Cancer signature ensemble integrating cfDNA methylation, copy number, and fragmentation facilitates multi-cancer early detection
Experimental & Molecular Medicine (2023)