Abstract
Background
Activating fusions of the NTRK1, NTRK2 and NTRK3 genes are drivers of carcinogenesis and proliferation across a broad range of tumour types in both adult and paediatric patients. Recently, the FDA granted tumour-agnostic approvals of TRK inhibitors, larotrectinib and entrectinib, based on significant and durable responses in multiple primary tumour types. Unfortunately, testing rates in clinical practice remain quite low. Adding plasma next-generation sequencing of circulating tumour DNA (ctDNA) to tissue-based testing increases the detection rate of oncogenic drivers and demonstrates high concordance with tissue genotyping. However, the clinical potential of ctDNA analysis to identify NTRK fusion-positive tumours has been largely unexplored.
Methods
We retrospectively reviewed a ctDNA database in advanced stage solid tumours for NTRK1 fusions.
Results
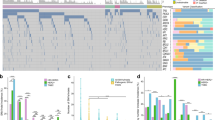
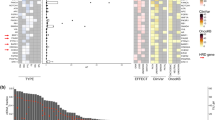
NTRK1 fusion events, with nine unique fusion partners, were identified in 37 patients. Of the cases for which clinical data were available, 44% had tissue testing for NTRK1 fusions; the NTRK1 fusion detected by ctDNA was confirmed in tissue in 88% of cases. Here, we report for the first time that minimally-invasive plasma NGS can detect NTRK fusions with a high positive predictive value.
Conclusion
Plasma ctDNA represents a rapid, non-invasive screening method for this rare genomic target that may improve identification of patients who can benefit from TRK-targeted therapy and potentially identify subsequent on- and off-target resistance mechanisms.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 24 print issues and online access
$259.00 per year
only $10.79 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Cocco E, Scaltriti M, Drilon A. NTRK fusion-positive cancers and TRK inhibitor therapy. Nat Rev Clin Oncol. 2018;15:731–47. https://doi.org/10.1038/s41571-018-0113-0.
Drilon A, Laetsch TW, Kummar S, DuBois SG, Lassen UN, Demetri GD, et al. Efficacy of larotrectinib in TRK fusion-positive cancers in adults and children. N Engl J Med. 2018;378:731–9. https://doi.org/10.1056/NEJMoa1714448.
Rolfo C. NTRK gene fusions: a rough diamond ready to sparkle. Lancet Oncol. 2020;21:472–4. https://doi.org/10.1016/S1470-2045(20)30026-7.
Russo A, Lopes AR, Scilla K, Mehra R, Adamo V, Oliveira J, et al. NTRK and NRG1 gene fusions in advanced non-small cell lung cancer (NSCLC). Precis Can Med. 2020;3:14.
Heitzer E, Haque IS, Roberts CES, Speicher MR. Current and future perspectives of liquid biopsies in genomics-driven oncology. Nat Rev Genet. 2019;20:71–88. https://doi.org/10.1038/s41576-018-0071-5.
Rolfo C, Cardona AF, Cristofanilli M, Paz-Ares L, Diaz Mochon JJ, Duran I, et al. Challenges and opportunities of cfDNA analysis implementation in clinical practice: perspective of the International Society of Liquid Biopsy (ISLB). Crit Rev Oncol Hematol. 2020;151:102978. https://doi.org/10.1016/j.critrevonc.2020.102978.
Odegaard JI, Vincent JJ, Mortimer S, Vowles JV, Ulrich BC, Banks KC, et al. Validation of a plasma-based comprehensive cancer genotyping assay utilizing orthogonal tissue- and plasma-based methodologies. Clin Cancer Res. 2018;24:3539–49. https://doi.org/10.1158/1078-0432.CCR-17-3831.
Clark TA, Chung JH, Kennedy M, Hughes JD, Chennagiri N, Lieber DS, et al. Analytical validation of a hybrid capture-based next-generation sequencing clinical assay for genomic profiling of cell-free circulating tumor DNA. J Mol Diagn. 2018;20:686–702. https://doi.org/10.1016/j.jmoldx.2018.05.004.
Westphalen CB, Krebs MG, Le Tourneau C, Sokol ES, Maund SL, Wilson TR, et al. Genomic context of NTRK1/2/3 fusion-positive tumours from a large real-world population. NPJ Precis Oncol. 2021;5:69. https://doi.org/10.1038/s41698-021-00206-y.
Marchiò C, Scaltriti M, Ladanyi M, Iafrate AJ, Bibeau F, Dietel M, et al. ESMO recommendations on the standard methods to detect NTRK fusions in daily practice and clinical research. Ann Oncol. 2019;30:1417–27. https://doi.org/10.1093/annonc/mdz204.
Russo A, De Miguel Perez D, Gunasekaran M, Scilla K, Lapidus R, Cooper B, et al. Liquid biopsy tracking of lung tumor evolutions over time. Expert Rev Mol Diagn. 2019;19:1099–108. https://doi.org/10.1080/14737159.2020.1680287.
Gerratana L, Zhang Q, Shah AN, Davis AA, Zhang Y, Wehbe F, et al. Performance of a novel Next Generation Sequencing circulating tumor DNA (ctDNA) platform for the evaluation of samples from patients with metastatic breast cancer (MBC). Crit Rev Oncol Hematol. 2020;145:102856. https://doi.org/10.1016/j.critrevonc.2019.102856.
Rolfo C, Mack PC, Scagliotti GV, Baas P, Barlesi F, Bivona TG, et al. Liquid biopsy for advanced non-small cell lung cancer (NSCLC): a statement paper from the IASLC. J Thorac Oncol. 2018;13:1248–68. https://doi.org/10.1016/j.jtho.2018.05.030.
Leighl NB, Page RD, Raymond VM, Daniel DB, Divers SG, Reckamp KL, et al. Clinical utility of comprehensive cell-free DNA analysis to identify genomic biomarkers in patients with newly diagnosed metastatic non-small cell lung cancer. Clin Cancer Res. 2019;25:4691–700. https://doi.org/10.1158/1078-0432.CCR-19-0624.
Remon J, Lacroix L, Jovelet C, Caramella C, Howarth K, Plagnol V, et al. Real-world utility of an amplicon-based next-generation sequencing liquid biopsy for broad molecular profiling in patients with advanced non-small-cell lung cancer. JCO Precis Oncol. 2019;3: https://doi.org/10.1200/PO.18.00211.
Mack PC, Banks KC, Espenschied CR, Burich RA, Zill OA, Lee CE, et al. Spectrum of driver mutations and clinical impact of circulating tumor DNA analysis in non-small cell lung cancer: Analysis of over 8000 cases. Cancer. 2020;126:3219–28. https://doi.org/10.1002/cncr.32876.
Strickler JH, Loree JM, Ahronian LG, Parikh AR, Niedzwiecki D, Pereira AAL, et al. Genomic landscape of cell-free DNA in patients with colorectal cancer. Cancer Disco. 2018;8:164–73. https://doi.org/10.1158/2159-8290.CD-17-1009.
Rich TA, Reckamp KL, Chae YK, Doebele RC, Iams WT, Oh M, et al. Analysis of cell-free DNA from 32,989 advanced cancers reveals novel co-occurring activating RET alterations and oncogenic signaling pathway aberrations. Clin Cancer Res. 2019;25:5832–42. https://doi.org/10.1158/1078-0432.CCR-18-4049.
Rolfo C, Mack P, Scagliotti GV, Aggarwal C, Arcila ME, Barlesi F, et al. Liquid biopsy for advanced non-small cell lung cancer: a consensus statement from The International Association for the Study of Lung Cancer (IASLC). J Thorac Oncol. 2021; https://doi.org/10.1016/j.jtho.2021.06.017.
Xia H, Xue X, Ding H, Ou Q, Wu X, Nagasaka M, et al. Evidence of NTRK1 fusion as resistance mechanism to EGFR TKI in EGFR+ NSCLC: results from a large-scale survey of NTRK1 fusions in chinese patients with lung cancer. Clin Lung Cancer. 2020;21:247–54. https://doi.org/10.1016/j.cllc.2019.09.004.
Piotrowska Z, Isozaki H, Lennerz JK, Gainor JF, Lennes IT, Zhu VW, et al. Landscape of acquired resistance to osimertinib in EGFR-mutant NSCLC and clinical validation of combined EGFR and RET inhibition with osimertinib and BLU-667 for acquired RET fusion. Cancer Disco. 2018;8:1529–39. https://doi.org/10.1158/2159-8290.CD-18-1022.
Cocco E, Lee JE, Kannan S, Schram AM, Won HH, Shifman S, et al. TRK xDFG Mutations trigger a sensitivity switch from type I to II kinase inhibitors. Cancer Discov. 2020, https://doi.org/10.1158/2159-8290.CD-20-0571.
Doebele RC. Acquired Resistance Is Oncogene and Drug Agnostic. Cancer Cell. 2019;36:347–9. https://doi.org/10.1016/j.ccell.2019.09.011.
Cocco E, Schram AM, Kulick A, Misale S, Won HH, Yaeger R, et al. Resistance to TRK inhibition mediated by convergent MAPK pathway activation. Nat Med. 2019;25:1422–7. https://doi.org/10.1038/s41591-019-0542-z.
Acknowledgements
Not applicable.
Author information
Authors and Affiliations
Contributions
CR, LK, KP and DRG conceived the work; LK and KP acquired data; AD, JJL, AMS, LK and KP provided detailed cases information and materials; CR, AR, LK, KP and DRG drafted the manuscript and analysed the data; all the co-authors revised the manuscript. All the co-authors approved the final version.
Corresponding author
Ethics declarations
Ethics approval and consent to participate
This research was conducted in accordance with the Declaration of Helsinki. Institutional Review Board approval (Advarra IRB Pro00034566/CR00218935) waived the need for individual informed consent for use of deidentified aggregate data for research purposes.
Consent to publish
Not applicable.
Competing interests
Dr. Rolfo reports grants for Lung Cancer Research Foundation-Pfizer Grant 2019 NHI U54 grant (Project co-leader); He has received personal fees for attending advisory board with Inivata, ArcherDx, EMD Serono, BMS, Novartis, Boston Pharmaceuticals, Pfizer, Mirati, and Eisai; fee for speaking bureau: MSD, Astra Zeneca, Roche, and GuardantHealth; Participation in Safety Monitoring Board for EMD Serono Non-financial conflict included research collaboration: GuardantHealth. Dr. Drilon reports Honoraria/Advisory Boards from Ignyta/Genentech/Roche, Loxo/Bayer/Lilly, Takeda/Ariad/Millenium, TP Therapeutics, AstraZeneca, Pfizer, Blueprint Medicines, Helsinn, Beigene, BergenBio, Hengrui Therapeutics, Exelixis, Tyra Biosciences, Verastem, MORE Health, Abbvie, 14ner/Elevation Oncology, Remedica Ltd., ArcherDX, Monopteros, Novartis, EMD Serono, Melendi, Liberum, Repare RX; Associated Research to Institution from Pfizer, Exelixis, GlaxoSmithKlein, Teva, Taiho, PharmaMar; Royalties from Wolters Kluwer; Other from Merck, Puma, Merus, Boehringer Ingelheim. Dr. Russo reports consultancy for Astra Zeneca, MSD, and Novartis outside the submitted work. Dr. Stinchcombe reports personal fees from Takeda, AstraZeneca, Genentech/Roche, Foundation Medicine, Pfizer, EMD Serono, Novartis, Daiichi Sankyo, Lilly, Medtronic, Puma Biotechnology, Janssen Oncology, Regeneron, non-financial support from Genentech/Roche, Blueprint Medicines, AstraZeneca, Takeda, Advaxis, Regeneron, outside the submitted work. Dr. Vidula reports institutional research funding from Radius, Merck, Daehwa, Pfizer (and travel), and Novartis; advisory board from AbbVie; Dr. Price is an employee and stock holder of Guardant Health.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Rolfo, C., Drilon, A., Hong, D. et al. NTRK1 Fusions identified by non-invasive plasma next-generation sequencing (NGS) across 9 cancer types. Br J Cancer 126, 514–520 (2022). https://doi.org/10.1038/s41416-021-01536-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-021-01536-1
This article is cited by
-
Circulating tumor DNA validity and potential uses in metastatic breast cancer
npj Breast Cancer (2024)
-
Network approach in liquidomics landscape
Journal of Experimental & Clinical Cancer Research (2023)
-
Prevalence and clinico-genomic characteristics of patients with TRK fusion cancer in China
npj Precision Oncology (2023)