Abstract
Background
Delayed detection of tumours contributes to poor prognosis in patients with gastric cancer (GC). The invasive nature of endoscopy and the absence of an effective serum markers highlight the need to develop novel, noninvasive biomarkers.
Methods
We performed biomarker discovery and validation to identify candidate genes in three gene expression data sets. After validating the gene panel in clinical tissues, we translated the gene panel into serum samples by performing training and validation in 89 samples from GC patients and 54 from healthy donors in two independent cohorts.
Results
We identified a nine-gene panel in the discovery phase, with subsequent validation in tissue specimens. Using a serum training cohort, we developed a 5-gene risk prediction formulae for the diagnosis of GC; bootstrapped analysis exhibited an AUC of 0.896. We validated this 5-gene biomarker panel using an independent serum cohort, yielding an AUC of 0.947. This biomarker panel successfully identified GC, regardless of tumour histology. Notably, biomarker performance for detection of stage 1 and 2 GC displayed an AUC of 0.928 and 0.980 in both serum cohorts.
Conclusions
We identified a novel 5-gene biomarker panel for noninvasive diagnosis of GC, which might serve as a potential diagnostic tool for early detection.
Similar content being viewed by others
Background
Gastric cancer (GC) is the second leading cause of cancer-related deaths worldwide [1]. In the United States, almost one-third of patients with GC are diagnosed at an advanced stage with distant metastases, and the 5-year survival rate is <30% [2]. Several factors, including tumour heterogeneity, limited treatment options, and insufficient response to conventional therapy, contribute to the poor prognosis of this disease. Further, the lack of a clinically available, noninvasive screening modality remains one of the most critical barriers for improving disease outcomes in this malignancy. Unfortunately, serum tumour markers (e.g. CEA, CA 19-9, and CA 72-4) are not useful for detecting GC due to their low sensitivity and specificity [3]. Instead, endoscopy is currently recommended as the standard for screening and diagnosing GC. However, endoscopy is a relatively expensive and invasive procedure with rare but serious complications; these limitations have hindered its widespread use for screening in Western countries [4, 5]. Intriguingly, in a few East Asian countries where endoscopy is available at an affordable cost and nationwide endoscopic surveillance programmes have been established, the proportion of patients diagnosed at stage 1 with excellent prognosis is gradually increasing, underscoring the importance of early-stage detection of GC to improve prognosis [6, 7]. This evidence suggests that a cost-effective and safely accessible screening modality is warranted to improve overall survival in patients with GC.
There have been continuous efforts for developing noninvasive biomarkers for cancer diagnosis and treatment. Cancer cells are characterised by rapid growth, invasion, and metastasis enabled by a rich blood supply, which leads to a continuous release of tumour cells into the bloodstream. Likewise, cancer-related nucleic acids are shed in the blood vessels via several mechanisms: apoptosis of cancer cells [8, 9], tumour necrosis [10], and active secretion of RNA- or DNA-containing vesicles by tumours [11, 12]. In this regard, blood-based biomarkers can reflect real-time biologic characteristics of tumours and have been recognised as emerging indicators for diagnosing cancer, detecting recurrence [13,14,15], or monitoring treatment response in several malignancies [16, 17]. Collectively, this highlights the importance of developing a blood-based biomarker for diagnosis of GC patients.
Recent advances in RNA sequencing technologies have enabled molecular characterisation of several malignancies and more precise biomarker discovery. Accordingly, over the past decade, several studies have analysed clinical specimens from patients with GC to identify circulating noncoding RNAs, primarily microRNAs and long noncoding RNAs, as diagnostic biomarkers [18,19,20]. While findings from some of these studies have been promising, limitations, including noncomprehensive biomarker discovery and validation approaches, a lack of clinical cohorts for independent validation, and unsatisfactory performance in detecting early-stage (stage 1 and 2) tumours, have restricted their translation into clinical practice. With regards to cell-free messenger RNA (mRNA), although they provide both genetic and functional information, the concerns with their stability in systemic circulation have discouraged their exploration and underestimated their biomarker significance. Consequently, to date, little is known about the ability of circulating mRNAs to diagnose GC.
Two main histologic types (diffuse and intestinal) are recognised in GC [21]. Because diffuse-type GC is associated with low sensitivity to standard treatment and poor prognosis, early detection is even more crucial for improving prognosis of this disease [22, 23]. Because intestinal-type tumours arise from premalignant lesions including intestinal metaplasia and adenoma, the co-existence of these lesions might hamper the effective discovery of malignancy-specific biomarkers.
To address these limitations, we conducted a systematic genome-wide expression profiling in tissue specimens from patients with diffuse-type GC, followed by rigorous bioinformatic approaches to identify a panel of genes that can allow discrimination of GC from noncancerous tissues. We next validated this gene panel using tissue samples in a clinical cohort of GC patients and successfully confirmed their ability to distinguish GC from normal. Subsequently, we translated our tissue-based gene panel into a serum-based assay, and assessed its diagnostic performance by analysing two independent cohorts consisting of GC patients and healthy donors (HDs). Towards this end, by using a comprehensive biomarker discovery and validation approach, we identified a 5-gene biomarker panel that enables early detection of GC, which may be clinically important for improving the prognosis of patients suffering from this malignancy.
Methods
This study was performed in accordance with STARD (STAndards for the Reporting of Diagnostic accuracy studies) guidelines [24].
Biomarker discovery and in silico validation in genome-wide transcriptomic expression profiling data sets
For the biomarker discovery phase of our study, we analysed genome-wide expression profiling data from three publicly available data sets (GSE29272, GSE62254, and GSE66222). Data were downloaded from the Gene Expression Omnibus database. Among them, GSE62254 and GSE66222 contain gene expression data of patient-matched GC and adjacent normal tissue, respectively. The GSE29272 and GSE62254/66222 cohorts included 134 and 300 patients with GC, respectively. To avoid the potential influence of precancerous lesions in the discovery phase, which frequently accompany intestinal-type GC, we selected only the cases with diffuse-type histology and those with available paired tissues for biomarker discovery; a total of 50 and 54 cases from each data set were included in the analysis.
Clinical cohorts for biomarker validation
Patients who received gastrectomy for biopsy-proven primary diffuse-type GC between 2008 and 2015 at Ajou University, Suwon, Korea were enrolled as a tissue validation cohort. Patients who had received neoadjuvant chemotherapy and those who developed tumours in the remnant stomach following previous partial gastrectomy were excluded. Paired cancer and adjacent normal tissue from the remaining 82 patients were included in tissue analysis, and their clinicopathologic characteristics are shown in Table S1.
For serum translation of the biomarkers, two independent cohorts consisting of 89 patients with GC and 54 HDs were enrolled. This included 54 patients who received surgical treatment between 2012 and 2015 at the Ajou University, Suwon, Korea and a control group of 31 healthy volunteers from the University of Ulsan and Asan Medical Centre, Seoul, Korea and 35 patients who received surgery between 2017 and 2018 at the Ajou University, Suwon, Korea and 23 HDs from Mie University, Mie, Japan. All HDs were enrolled after confirming that they had no history of malignancy. Clinicopathological characteristics of the two cohorts are described in Table S2.
Cancer tissues were obtained from representative malignant lesions; normal tissues were procured from the location furthest from the lesion in a resected stomach specimen during the operation and were assessed by pathologic examination to evaluate neoplastic purity or the presence of malignant cells. Tissues were snap-frozen in liquid nitrogen and stored at −80 °C. Blood was collected before surgery in cancer patients and centrifuged at 1600 × g for 10 min at 4 °C. The clear upper layer comprising of cell-free serum was extracted and stored at −80 °C. Blood samples from HDs were treated in the same manner.
The histological type was determined according to Lauren’s classification [21]. The tumour stage was determined according to the Seventh edition of the American Joint Committee on Cancer [25].
All procedures were conducted in accordance with the Helsinki Declaration. A written informed consent was obtained from all participants. This study was approved by the Institutional Review Boards of all participating institutions.
RNA extraction and gene expression analysis
Total RNA was isolated from 200 µl of serum and snap-frozen surgical tissues using the RNeasy Mini Kit (Qiagen, Hilden, Germany). Real-time quantitative reverse transcription polymerase chain reaction (RT-qPCR) was performed using the SensiFAST™ probe Lo-ROX Kit (Bioline, London, UK) and the QuantStudio 6 Flex Real-Time PCR System (Applied Biosystems, Foster City, CA). Gene expression levels were evaluated using the Applied Biosystems QuantStudio 6 Flex Real-Time PCR System Software. The relative abundance of target genes was assessed and corrected to the expression level of beta-actin as an internal control using the 2−ΔCt method; ΔCt refers to the difference of Ct values between the gene of interest and beta-actin. Values were further transformed into the log2 form. PCR primers used in the current study are described in Table S3.
Statistical analysis
Statistical analyses were performed using R version 3.6.3 and MedCalc Statistical Software version 19.2 (MedCalc Software Ltd, Ostend, Belgium). Wilcoxon signed-rank test was used to compare gene expression levels between cancer and normal tissue in the discovery phase. In the discovery and in silico validation phase, the performance of the risk model consisted of candidate genes assessed through single-step-based logistic regression analysis with group dichotomisation based on the median expression value of each gene. Then random forest classification with tenfold cross-validation, a machine learning technique, was used to assess the performance of candidate gene biomarkers. In the clinical phase, a gene-based risk score formulae were built by single-step based logistic regression, and dichotomised groups were defined according to Youden’s index for cancer diagnosis. Performance was evaluated using receiver operating characteristic (ROC) curves and area under the curve (AUC) values. Bootstrap aggregating, a machine learning ensemble meta-algorithm, was used and repeated 1000 times to improve the stability and accuracy of diagnostic performance in serum training and validation phases. An optimism-adjusted AUC was computed. A paired t test was used to compare gene expression levels between serum samples collected before and after surgery. Pearson correlation analysis was employed to measure the association of gene expression from cancer tissue with serum in patients with GC. The threshold for statistical significance was set at P < 0.05.
Results
Genome-wide transcriptomic expression profiling identifies a nine-gene panel for discriminating GC from noncancerous tissue
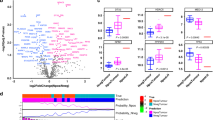
The overall workflow of this study is illustrated in Fig. S1. We performed a systematic, comprehensive, and unbiased biomarker discovery effort by analysing genome-wide transcriptomic expression profiling data from three publicly available data sets of patients with GC. The GSE29272 data set was selected for the initial biomarker discovery and expression data from paired cancer and adjacent normal tissues from 50 patients were analysed. We identified a panel of 12 genes that were differentially expressed between GC and noncancerous tissues (P < 0.05; absolute fold change >2; expression level >6; Pearson’s correlation <0.5). With this panel, we compared the expression pattern of 12 genes in validation data sets (GSE62254/66222) and excluded two genes that showed discrepancy in gene expression pattern between data sets. In addition, one gene located within the sex chromosome was also excluded from the panel. Accordingly, we finalised a panel of nine genes: ALDOB, MSMB, TNFRSF17, HBB, KRT7, CEACAM6, UBD, PLA2G2A, and ISG15 (Table S4). With nine genes, we developed a logistic regression model to distinguish GC from normal tissue, which exhibited an AUC of 0.978 (95% confidence interval (CI): 0.960–1.000, P < 0.001). We used a heatmap to visualise disparities in the expression of the candidate genes between GC and normal tissue (Fig. 1a). Analysis using a volcano plot revealed that the nine genes derived from the discovery step were highly significant in discriminating GC tissues and displayed marked expression change (Fig. 1b).
a A heatmap of nine genes derived from the GSE29272 discovery cohort demonstrates a disparity in expression between patient-matched tumour and normal tissues. b A volcano plot illustrates that the nine genes are accompanied by remarkable expression change and a significant discriminant value for GC tissues. c A heatmap of the nine-gene panel in the GSE62254 and GSE66222 validation cohorts exhibit consistent disparities in expression between patient-matched tumour and normal tissues compared to the discovery phase.
Next, we validated the accuracy of the 9-gene panel in GSE62254 and GSE66222 data sets, which confirmed the robustness of our biomarker discovery effort, as evidenced by the resulting AUC of 0.986 (95% CI: 0.944–1.000, P < 0.001). By using a heatmap to visualise the concordant gene expression pattern in the validation data set as in the discovery cohort, we ensured the consistency of the nine-gene panel (Fig. 1c). In summary, our in-depth biomarker discovery effort, which utilised multiple bioinformatic approaches, led us to successfully identify and validate a nine-gene panel that exhibited robust performance in distinguishing GC from noncancerous tissue in independent genome-wide expression profiling data sets of GC patients.
Tissue validation confirms the ability of the nine-gene panel to distinguish GC from normal tissues
Following successful discovery and in silico validation of our gene panel, we next validated these biomarkers in a tissue validation cohort of 82 patients with diffuse-type GC. We used RT-qPCR assays to measure the expression levels of each of the nine genes in all tissue samples. Consistent with their expression profiles in the public data sets, four genes (ALDOB, MSMB, TNFRSF17, and HBB) were significantly downregulated and five genes (KRT7, CEACAM6, UBD, PLA2G2A, and ISG15) were significantly upregulated (Fig. 2a). We used logistic regression to model the ability of the 9-gene panel to distinguish GC in tissues, which resulted in an AUC of 0.914 (95% CI: 0.862–0.953, sensitivity 76.8%, specificity 94.9%, P < 0.001), and was superior to the AUC of individual genes (Fig. 2b). The risk prediction formula was as follows: (0.26431 × ALDOB) + (0.023239 × MSMB) + (0.66401 × TNFRSF17) + (0.42658 × HBB) + (−0.50042 × KRT7) + (−0.30181 × CEACAM6) + (−0.37733 × UBD) + (−0.28293 × PLA2G2A) + (−0.36673 × ISG15) + 3.33614. The performance of the nine-gene panel to discriminate cancer from normal tissue is shown in Fig. 2c. This tissue validation step confirmed the consistency of our nine-gene panel for its diagnostic potential in tissue specimens.
a Patient-matched GC and adjacent normal tissues (NAT) were collected from a tissue validation cohort, and their expression was assessed using RT-qPCR. Consistent with expression in the public gene expression data sets, five genes were significantly upregulated and four were downregulated in cancer tissues. b The discriminatory ability of CEACAM6 was the highest among individual genes. Overall performance of the nine-gene panel reached an AUC of 0.914. c The ability of the nine-gene panel to distinguish GC (red) from NAT (blue) is shown in a scatter plot. (***P < 0.001, **P < 0.01, *P < 0.05).
The five-gene biomarker panel successfully identifies patients with GC in a serum training cohort
To translate our tissue-based gene panel for its noninvasive clinical application, we used RT-qPCR to measure expression of the nine genes in serum samples in a training cohort consisting of 54 patients with GC and 31 HDs. Four genes (ALDOB, MSMB, TNFRSF17, and CEACAM6) were significantly downregulated and not expressed in serum from GC patients and subsequently excluded from the further analysis. The remaining five genes (HBB, KRT7, UBD, PLA2G2A, and ISG15) were upregulated in GC patients, as measured in serum specimens of the training cohort. Among them, three genes (HBB, UBD, and ISG15) were significantly upregulated (Fig. 3a).
a The expression of five genes was measured in serum specimens from a serum training cohort; three genes (HBB, UBD, and ISG) were significantly upregulated in patients with GC relative to healthy donors (HD). b A bootstrapped analysis adjusted the performance of the biomarker and remarkable diagnostic accuracy was maintained after amending optimism. c The five-gene biomarker successfully discriminated patients with GC from HDs in the serum training cohort. d The biomarker further distinguished patients with stage 1 and 2 GC from those with stage 3 and 4 GC and both groups from HDs, based on a logistic regression model. e A waterfall plot exhibits the robust diagnostic performance of the biomarker in the serum validation cohort. (***P < 0.001, **P < 0.01, *P < 0.05, n.s.: not significant).
We built a logistic regression model to distinguish GC patients from controls in serum using the five upregulated genes, which exhibited an AUC of 0.924 (95% CI: 0.845–0.970, sensitivity 75.9%, specificity 100%, P < 0.001). Based on the five genes’ individual coefficients and a constant derived from the model, we developed a risk prediction formulae, defined as (−0.26479 × HBB) + (0.076393 × KRT7) + (0.94302 × UBD) + (−1.75459 × PLA2G2A) + (0.71413 × ISG15) − 2.63663. We thereafter reassessed our five-gene biomarker panel using bootstrap aggregating modules to reduce variance and minimise data overfitting. Based on that algorithm, we measured an optimism-adjusted AUC value of 0.896 (95% CI: 0.894–0.898; Fig. 3b). In addition, we assessed several parameters related to diagnostic performance in the serum training cohort, including accuracy (81.5%), sensitivity (87.1%), specificity (71.7%), positive predictive value (PPV; 84.8%), and negative predictive value (NPV; 76.6%) (Table 1).
We used a waterfall plot to evaluate the performance of the five-gene biomarker panel to identify patients with GC (Fig. 3c). In addition, our gene biomarker successfully stratified patient groups according to tumour stage (stage 1 and 2 vs. stage 3 and 4) and distinguished cancer patients from HDs (Fig. 3d). As the last step, the performance of the biomarker panel was assessed according to patient gender and it was comparable in both groups (an AUC of 0.920 in men and 0.927 in women).
The noninvasive gene biomarker panel robustly identifies GC patients in an independent serum validation cohort
Following development of the 5-gene-based risk prediction formulae, we used a serum validation cohort consisting of 35 patients and 23 HDs to independently assess the diagnostic performance of the biomarkers. We first used RT-qPCR to measure and compare expression of the five genes between patients with GC and HDs. Consistent with our findings in the serum training cohort, the expression of three genes (HBB, UBD, and ISG15) was significantly increased in patients of the validation cohort (Table 2). Our 5-gene biomarker panel discriminated between GC patients and HDs and yielded an AUC of 0.988 (95% CI: 0.916–1.000, sensitivity: 91.4%, specificity: 100%, P < 0.001). We used a waterfall plot to evaluate the diagnostic performance of the five-gene biomarker panel (Fig. 3e). Thereafter, we applied the bootstrap aggregating module to yield an adjusted AUC of 0.947 (95% CI: 0.946–0.949, accuracy 94.0%, sensitivity 94.4%, specificity 93.3%, PPV 95.7%, and NPV 92.0%; Table 1). In line with the serum training cohort, the remarkable performance of 5-gene biomarker irrespective of patient gender was again observed in our study (an AUC of 0.987 in men and 0.990 in women). Through these analyses, we confirmed that our gene biomarker panel was successfully translated into serum specimens and demonstrated high diagnostic performance.
The gene biomarker panel consistently discriminates patients with GC regardless of tumour histology
Our successive efforts revealed the promising potential of the five-gene panel as noninvasive diagnostic biomarkers for GC. Next, we evaluated the performance of our biomarkers according to histologic types (diffuse or intestinal) in the serum training cohort, considering that the five genes were derived from analysis of diffuse-type GC and frequent co-existence of premalignant lesion in intestinal-type tumour might affect diagnostic accuracy of the biomarker in intestinal-type GC. By using the five-gene risk prediction formula, we assessed the performance of the transcriptomic biomarkers, and it demonstrated a remarkably high accuracy in differentiating either tumour type from HD. Bootstrap modules resulted in an AUC of 0.888 (95% CI: 0.886–0.890, accuracy 81.3%, sensitivity 79.7%, specificity 82.6%, PPV 78.7%, and NPV 84.3%) in diffuse-type GC and 0.880 (95% CI: 0.877–0.884, accuracy 88.9%, sensitivity 93.7%, specificity 78.8%, PPV 90.3%, and NPV 87.5%) in intestinal-type GC relative to HDs (Fig. 4a and Table 1).
The diagnostic performance of the biomarker was assessed according to tumour histology (diffuse vs. intestinal type) and stage (stage 1 and 2 vs. 3 and 4). a In the clinical training cohort, regardless of histology, the biomarker consistently identified patients with GC. In addition, the diagnostic performance in patients with stage 1 and 2 tumours was remarkable. b In the clinical validation cohort, the robust performance of the biomarker was again demonstrated relative to tumour histology and pathologic stage. c Compared to preoperative gene expression, three genes (HBB, ISG15, and KRT7) were significantly downregulated after curative surgery in patients with diffuse-type GC. (**P < 0.01, *P < 0.05, n.s.: not significant).
We repeated the analysis in the serum validation cohort, which exhibited an AUC of 0.969 (95% CI: 0.967–0.971, accuracy 95.1%, sensitivity 96.4%, specificity 93.7%, PPV 95.1%, and NPV 95.8%) in diffuse-type GC and 0.960 (95% CI: 0.958–0.962, accuracy 94.6%, sensitivity 95.4%, specificity 93.4%, PPV 95.7%, and NPV 93.8%) in intestinal-type GC relative to HDs (Fig. 4b and Table 1). This validation step yet again validated the consistency of our biomarker panel for the diagnosis of GC across all histologic types.
The 5-gene biomarker panel successfully identifies stage 1 and 2 GC patients
Because tumour detection at earlier stages is closely associated with improved prognosis in patients with GC, we next evaluated the diagnostic potential of the biomarkers in stage 1 and 2 tumours. We dichotomised patients in the serum training and validation cohorts into stage 1 and 2 vs. stage 3 and 4 and assessed their diagnostic performance. In the training cohort, our biomarker panel successfully discriminated patients with stage 1 and 2 GC from HDs, yielding an AUC of 0.928 (95% CI: 0.926–0.931, accuracy 90.9%, sensitivity 87.8%, specificity 92.8%, PPV 89.0%, and NPV 92.7%), which was higher than an AUC of 0.862 (95% CI: 0.859–0.864, accuracy 80.8%, sensitivity 80.4%, specificity 81.2%, PPV 83.7%, and NPV 78.8%) in patients with stage 3 and 4 GC relative to HDs (Fig. 4a and Table 1). We again demonstrated the remarkable performance of the biomarker panel in stage 1 and 2 tumours in the validation cohort, which yielded an AUC of 0.980 (95% CI: 0.979–0.981, accuracy 95.8%, sensitivity 93.5%, specificity 97.8%, PPV 97.4%, and NPV 95.0%) in stage 1 and 2 GC patients, and was higher than an AUC of 0.924 (95% CI: 0.921-0.926, accuracy 91.9%, sensitivity 89.3%, specificity 93.8%, PPV 9.14%, and NPV 93.0%) in stage 3 and 4 GC patients relative to HDs (Fig. 4b and Table 1). These results underscore the potential of the biomarker for early detection of GC.
A significant correlation in pre- vs. post-surgery serum specimens highlights the specificity of the gene biomarker panel
To determine the performance of our transcriptomic panel in post-treatment setting, we analysed postoperative serum samples collected 3 months after curative surgery vs. preoperative serum specimens in the validation cohort. Among the five genes, HBB and KRT7 were significantly downregulated after surgery (Table S5). Postoperative ISG15 expression level was also lower than the preoperative value. When we segregated the samples by histological type, these three genes were significantly downregulated in diffuse-type GC (Fig. 4c). These results highlight the robustness of our biomarkers since they changed in pre- vs. post-treatment serum. Taken together, our comprehensive biomarker discovery and validation effort from tissue to serum specimens allowed us to identify and validate a novel five-gene biomarker panel that might be effective in detecting early-stage tumours, which is a key factor for improving the prognosis of patients with GC.
Discussion
Early detection of tumours is the most desirable approach in the management of GC. Conventional tumour markers are frequently used for detection of other gastrointestinal malignancies; however, they are not recommended for patients with GC due to limited diagnostic accuracy. Although endoscopy is used clinically with high accuracy, it is associated with patient discomfort and poor compliance for disease screening. In addition, repeated endoscopy and biopsies can impose additional risks of bleeding during or after biopsy. In this regard, cell-free nucleic acids in the serum have the potential to be a safer diagnostic biomarker in GC. Cell-free miRNAs have been found to be stable to ribonuclease and freeze–thaw cycles. Several previous studies highlighted cell-free miRNAs for detection of GC and demonstrated promising results [26,27,28,29,30,31]. However, considerable limitations alleviate the clinical implementation of miRNAs as diagnostic biomarkers. Most of the miRNAs identified have not been successfully reproduced in subsequent studies, likely due to the absence of consensus on internal controls to normalise RT-qPCR data in serum. In addition, the accuracy of cell-free miRNAs in the diagnosis of early-stage tumours, an important target for a screening biomarker, either was not evaluated or demonstrated poor overall diagnostic performance. Likewise, although cell-free DNA biomarkers are highly specific, they have limited diagnostic sensitivity due to the low mutational burden in early-stage GC. For mRNA biomarkers, one study showed that human telomerase reverse transcriptase (hTERT) could distinguish cancer patients from healthy people, but its sensitivity was relatively low and no clinical cohort was used for independent validation [32]. In the present study, we developed a five-gene biomarker panel that robustly identified patients with GC, including those with early-stage tumours, by performing comprehensive biomarker discovery and validation steps using both tissue and serum samples from several clinical cohorts.
Our study did not observe identical expression profiles between serum and matched tumour tissue for some of the genes. Discordance in gene expression has been observed in previous studies and it could be explained due to discordant expression profiles between the extracellular and intracellular environments, suggesting that circulating RNA profiles do not coincide with those of tumour tissues [33,34,35,36].
From a functional viewpoint, various genes in our biomarker panel have been shown to be authentic candidates involved in cancer pathogenesis. For instance, haemoglobin beta subunit (HBB) is one of the transcripts most consistently and specifically overexpressed in circulating tumour cells while expression is comparably low in primary tumours. The role of radical oxidative stress has been highlighted for inhibiting cancer dissemination via bloodstream. HBB expression suppresses radical oxidative stress-mediated cytotoxicity, leading to cancer cell survival and facilitating distant metastasis [37]. Moreover, it is associated with cancer cell aggressiveness and poor prognosis in breast cancer [38]. Previous studies have demonstrated that expression of keratin 7 (KRT7) is substantially increased in GC tissues, while it is barely expressed in normal tissues and that its upregulation plays a role in GC cell proliferation and migration suggesting that KRT7 is involved in the development and/or progression of GC [39, 40]. Elevated expression of phospholipase A2 group IIA (PLA2G2A) was reported to be associated with better prognosis in patients with GC, and its activation may suppress progression or metastasis of GC cells [41, 42]. A cell-line-based study showed that PLA2G2A expression was elevated in early-stage tumours but decreased in metastatic or late-stage tumours due to epigenetic silencing, and this could explain better performance of our biomarker panel in early-stage GC in part [43]. The activation of interferon signalling pathways is a key component of innate immunity against cancer; the interferon-stimulated gene 15 ubiquitin-like modifier (ISG15) might allow diffuse-type gastric tumour cells to evade the innate antitumour immune response by suppressing antigen presentation by dendritic cells [44, 45].
We acknowledge some limitations of the present study. First, we used a retrospective design and included only a modest number of patient samples from Asia. A multinational prospective study including a larger number of samples is required in the future. Second, we did not assess the performance of the biomarkers in differentiating malignancy from precancerous lesions in intestinal-type cancer. Regardless, this study is valuable because it demonstrates that a circulating gene expression signature may serve as a robust biomarker with notable diagnostic accuracy across GC, based on clinical validation.
In conclusion, using a comprehensive genome-wide discovery and validation approach in multiple public data sets, we identified a nine-gene panel that distinguishes GC from noncancerous tissue and validated it in tissue samples from a clinical cohort. Subsequently, we established a five-gene biomarker panel, optimised it using a machine learning algorithm, and validated its ability to successfully identify patients with GC, even among patients with early-stage tumours, in serum samples from two independent clinical cohorts. This biomarker panel might serve as a potential noninvasive diagnostic tool for early detection of GC.
Data availability
The data that support the findings of this study are available from the corresponding authors upon reasonable request.
References
Ferlay J, Colombet M, Soerjomataram I, Mathers C, Parkin DM, Pineros M, et al. Estimating the global cancer incidence and mortality in 2018: GLOBOCAN sources and methods. Int J Cancer. 2019;144:1941–53.
Jim MA, Pinheiro PS, Carreira H, Espey DK, Wiggins CL, Weir HK. Stomach cancer survival in the United States by race and stage (2001-2009): findings from the CONCORD-2 study. Cancer. 2017;123:4994–5013.
Shimada H, Noie T, Ohashi M, Oba K, Takahashi Y. Clinical significance of serum tumor markers for gastric cancer: a systematic review of literature by the Task Force of the Japanese Gastric Cancer Association. Gastric Cancer. 2014;17:26–33.
Gupta N, Bansal A, Wani SB, Gaddam S, Rastogi A, Sharma P. Endoscopy for upper GI cancer screening in the general population: a cost-utility analysis. Gastrointest Endosc. 2011;74:610.e2–24.e2.
Levy I, Gralnek IM. Complications of diagnostic colonoscopy, upper endoscopy, and enteroscopy. Best Pract Res Clin Gastroenterol. 2016;30:705–18.
Information Committee of Korean Gastric Cancer Association. Korean Gastric Cancer Association Nationwide Survey on Gastric Cancer in 2014. J Gastric Cancer. 2016;16:131–40.
Katai H, Ishikawa T, Akazawa K, Isobe Y, Miyashiro I, Oda I, et al. Five-year survival analysis of surgically resected gastric cancer cases in Japan: a retrospective analysis of more than 100,000 patients from the nationwide registry of the Japanese Gastric Cancer Association (2001-2007). Gastric Cancer. 2018;21:144–54.
Fournie GJ, Courtin JP, Laval F, Chale JJ, Pourrat JP, Pujazon MC, et al. Plasma DNA as a marker of cancerous cell death. Investigations in patients suffering from lung cancer and in nude mice bearing human tumours. Cancer Lett. 1995;91:221–7.
Stroun M, Anker P, Lyautey J, Lederrey C, Maurice PA. Isolation and characterization of DNA from the plasma of cancer patients. Eur J Cancer Clin Oncol. 1987;23:707–12.
Stroun M, Anker P, Maurice P, Lyautey J, Lederrey C, Beljanski M. Neoplastic characteristics of the DNA found in the plasma of cancer patients. Oncology 1989;46:318–22.
Anker P, Stroun M, Maurice PA. Spontaneous release of DNA by human blood lymphocytes as shown in an in vitro system. Cancer Res. 1975;35:2375–82.
Skog J, Wurdinger T, van Rijn S, Meijer DH, Gainche L, Sena-Esteves M, et al. Glioblastoma microvesicles transport RNA and proteins that promote tumour growth and provide diagnostic biomarkers. Nat Cell Biol. 2008;10:1470–6.
Diehl F, Schmidt K, Choti MA, Romans K, Goodman S, Li M, et al. Circulating mutant DNA to assess tumor dynamics. Nat Med. 2008;14:985–90.
Hamana K, Uzawa K, Ogawara K, Shiiba M, Bukawa H, Yokoe H, et al. Monitoring of circulating tumour-associated DNA as a prognostic tool for oral squamous cell carcinoma. Br J Cancer. 2005;92:2181–4.
Sozzi G, Conte D, Mariani L, Lo Vullo S, Roz L, Lombardo C, et al. Analysis of circulating tumor DNA in plasma at diagnosis and during follow-up of lung cancer patients. Cancer Res. 2001;61:4675–8.
Gevensleben H, Garcia-Murillas I, Graeser MK, Schiavon G, Osin P, Parton M, et al. Noninvasive detection of HER2 amplification with plasma DNA digital PCR. Clin Cancer Res. 2013;19:3276–84.
Spindler KL, Pallisgaard N, Vogelius I, Jakobsen A. Quantitative cell-free DNA, KRAS, and BRAF mutations in plasma from patients with metastatic colorectal cancer during treatment with cetuximab and irinotecan. Clin Cancer Res. 2012;18:1177–85.
Liu R, Zhang C, Hu Z, Li G, Wang C, Yang C, et al. A five-microRNA signature identified from genome-wide serum microRNA expression profiling serves as a fingerprint for gastric cancer diagnosis. Eur J Cancer. 2011;47:784–91.
Shiotani A, Murao T, Kimura Y, Matsumoto H, Kamada T, Kusunoki H, et al. Identification of serum miRNAs as novel non-invasive biomarkers for detection of high risk for early gastric cancer. Br J Cancer. 2013;109:2323–30.
Zhou X, Yin C, Dang Y, Ye F, Zhang G. Identification of the long non-coding RNA H19 in plasma as a novel biomarker for diagnosis of gastric cancer. Sci Rep. 2015;5:11516.
Lauren P. The two histological main types of gastric carcinoma: diffuse and so-called intestinal-type carcinoma. An attempt at a histo-clinical classification. Acta Pathol Microbiol Scand. 1965;64:31–49.
Cristescu R, Lee J, Nebozhyn M, Kim KM, Ting JC, Wong SS, et al. Molecular analysis of gastric cancer identifies subtypes associated with distinct clinical outcomes. Nat Med. 2015;21:449–56.
Sohn BH, Hwang JE, Jang HJ, Lee HS, Oh SC, Shim JJ, et al. Clinical significance of four molecular subtypes of gastric cancer identified by The Cancer Genome Atlas Project. Clin Cancer Res. 2017. https://doi.org/10.1158/1078-0432.CCR-16-2211.
Cohen JF, Korevaar DA, Altman DG, Bruns DE, Gatsonis CA, Hooft L, et al. STARD 2015 guidelines for reporting diagnostic accuracy studies: explanation and elaboration. BMJ Open. 2016;6:e012799.
Edge SB, Compton CC. The American Joint Committee on Cancer: the 7th edition of the AJCC cancer staging manual and the future of TNM. Ann Surg Oncol. 2010;17:1471–4.
Konishi H, Ichikawa D, Komatsu S, Shiozaki A, Tsujiura M, Takeshita H, et al. Detection of gastric cancer-associated microRNAs on microRNA microarray comparing pre- and post-operative plasma. Br J Cancer. 2012;106:740–7.
Tsujiura M, Ichikawa D, Komatsu S, Shiozaki A, Takeshita H, Kosuga T, et al. Circulating microRNAs in plasma of patients with gastric cancers. Br J Cancer. 2010;102:1174–9.
Zhu C, Ren C, Han J, Ding Y, Du J, Dai N, et al. A five-microRNA panel in plasma was identified as potential biomarker for early detection of gastric cancer. Br J Cancer. 2014;110:2291–9.
Toiyama Y, Okugawa Y, Fleshman J, Richard Boland C, Goel A. MicroRNAs as potential liquid biopsy biomarkers in colorectal cancer: a systematic review. Biochim Biophys Acta Rev Cancer. 2018;1870:274–82.
Toiyama Y, Okugawa Y, Goel A. DNA methylation and microRNA biomarkers for noninvasive detection of gastric and colorectal cancer. Biochem Biophys Res Commun. 2014;455:43–57.
Link A, Goel A. MicroRNA in gastrointestinal cancer: a step closer to reality. Adv Clin Chem. 2013;62:221–68.
Kang Y, Zhang J, Sun P, Shang J. Circulating cell-free human telomerase reverse transcriptase mRNA in plasma and its potential diagnostic and prognostic value for gastric cancer. Int J Clin Oncol. 2013;18:478–86.
Chan M, Liaw CS, Ji SM, Tan HH, Wong CY, Thike AA, et al. Identification of circulating microRNA signatures for breast cancer detection. Clin Cancer Res. 2013;19:4477–87.
Cookson VJ, Bentley MA, Hogan BV, Horgan K, Hayward BE, Hazelwood LD, et al. Circulating microRNA profiles reflect the presence of breast tumours but not the profiles of microRNAs within the tumours. Cell Oncol. 2012;35:301–8.
Pigati L, Yaddanapudi SC, Iyengar R, Kim DJ, Hearn SA, Danforth D, et al. Selective release of microRNA species from normal and malignant mammary epithelial cells. PLoS ONE. 2010;5:e13515.
Zhu J, Zheng Z, Wang J, Sun J, Wang P, Cheng X, et al. Different miRNA expression profiles between human breast cancer tumors and serum. Front Genet. 2014;5:149.
Zheng Y, Miyamoto DT, Wittner BS, Sullivan JP, Aceto N, Jordan NV, et al. Expression of beta-globin by cancer cells promotes cell survival during blood-borne dissemination. Nat Commun. 2017;8:14344.
Ponzetti M, Capulli M, Angelucci A, Ventura L, Monache SD, Mercurio C, et al. Non-conventional role of haemoglobin beta in breast malignancy. Br J Cancer. 2017;117:994–1006.
Huang B, Song JH, Cheng Y, Abraham JM, Ibrahim S, Sun Z, et al. Long non-coding antisense RNA KRT7-AS is activated in gastric cancers and supports cancer cell progression by increasing KRT7 expression. Oncogene 2016;35:4927–36.
Yang J. Identification of novel biomarkers, MUC5AC, MUC1, KRT7, GAPDH, CD44 for gastric cancer. Med Oncol. 2020;37:34.
Leung SY, Chen X, Chu KM, Yuen ST, Mathy J, Ji J, et al. Phospholipase A2 group IIA expression in gastric adenocarcinoma is associated with prolonged survival and less frequent metastasis. Proc Natl Acad Sci USA. 2002;99:16203–8.
Wang X, Huang CJ, Yu GZ, Wang JJ, Wang R, Li YM, et al. Expression of group IIA phospholipase A2 is an independent predictor of favorable outcome for patients with gastric cancer. Hum Pathol. 2013;44:2020–7.
Ganesan K, Ivanova T, Wu Y, Rajasegaran V, Wu J, Lee MH, et al. Inhibition of gastric cancer invasion and metastasis by PLA2G2A, a novel beta-catenin/TCF target gene. Cancer Res. 2008;68:4277–86.
Jinawath N, Furukawa Y, Hasegawa S, Li M, Tsunoda T, Satoh S, et al. Comparison of gene-expression profiles between diffuse- and intestinal-type gastric cancers using a genome-wide cDNA microarray. Oncogene 2004;23:6830–44.
Parker BS, Rautela J, Hertzog PJ. Antitumour actions of interferons: implications for cancer therapy. Nat Rev Cancer. 2016;16:131–44.
Acknowledgements
We would like to thank Satoshi Nishiwada, Tatsuhiko Kakisaka, Yuma Wada, and Yasuyuki Okada for their important insights into experiments and data analysis. The biospecimen and data used in this study were provided by Asan Bio-Resource Centre, Korea Biobank Network 2019-05(184), and the Ajou Human Bio-Resource Bank (AHBB), a member of the National Biobank of Korea, which is supported by the Ministry of Health and Welfare.
Author information
Authors and Affiliations
Contributions
I.-S.L.: study concept and design, specimen provider, acquisition of data, analysis and interpretation of data, statistical analysis, drafting of the manuscript. J.A.: analysis and interpretation of data, statistical analysis, drafting of the manuscript. K.K.: analysis and interpretation of data, statistical analysis, drafting of the manuscript. Y.O.: specimen provider, acquisition of data, drafting of the manuscript. Y.T.: specimen provider, acquisition of data, drafting of the manuscript. H.H.: specimen provider, acquisition of data, drafting of the manuscript. A.G.: study concept and design, analysis and interpretation of data, statistical analysis, drafting of the manuscript.
Corresponding author
Ethics declarations
Funding information
The present work was supported by the grants CA72851, CA181572, and CA187956 from the National Cancer Institute, National Institutes of Health, and a pilot grant from the Stupid Strong Foundation to A.G.
Competing interests
The authors declare no competing interests.
Ethics approval and consent to participate
All procedures were conducted in accordance with the Helsinki Declaration. A written informed consent was obtained from all participants. This study was approved by the Institutional Review Boards of participating institutions (Study No. S2019-0237 and AJIRB-BMR-KSP-19-059).
Consent for publication
Not applicable.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Lee, IS., Ahn, J., Kim, K. et al. A blood-based transcriptomic signature for noninvasive diagnosis of gastric cancer. Br J Cancer 125, 846–853 (2021). https://doi.org/10.1038/s41416-021-01461-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41416-021-01461-3
This article is cited by
-
A performance evaluation of drug response prediction models for individual drugs
Scientific Reports (2023)