Abstract
Childhood maltreatment is a major risk factor for chronic and severe mental and physical health problems across the lifespan. Increasing evidence supports the hypothesis that maltreatment is associated with epigenetic changes that may subsequently serve as mechanisms of disease. The current review uses a systematic approach to identify and summarize the literature related to childhood maltreatment and alterations in DNA methylation in humans. A total of 100 empirical articles were identified in our systematic review of research published prior to or during March 2020, including studies that focused on candidate genes and studies that leveraged epigenome-wide data in both children and adults. Themes arising from the literature, including consistent and inconsistent patterns of results, are presented. Several directions for future research, including important methodological considerations for future study design, are discussed. Taken together, the literature on childhood maltreatment and DNA methylation underscores the complexity of transactions between the environment and biology across development.
Similar content being viewed by others
Introduction
Childhood maltreatment is a highly prevalent public health problem that often has devastating effects on physical and mental health. There are now numerous studies linking childhood maltreatment and other early adversities to nearly all forms of mental illness, as well as chronic medical conditions that cut across multiple organ systems1,2,3,4,5,6,7. Unfortunately, the negative sequelae of childhood maltreatment often begin early in life, and persist across adulthood1,6, posing risk for premature mortality. Indeed, adults with numerous adverse experiences in childhood die nearly 20 years earlier than those with no early adversity1. Understanding the mechanisms of biological influence is therefore critical to developing innovative targets for intervention to enhance health outcomes among highly vulnerable children with this major adverse exposure.
Over the past decade a rapidly growing body of literature has underscored the significant role of epigenetics in the sequelae of childhood maltreatment. Epigenetic processes allow the body to respond to environmental influences by altering gene expression through chemical modifications that regulate chromatin structure and/or DNA accessibility without inducing changes to the DNA sequence8. DNA methylation is among the most commonly studied epigenetic processes and involves the addition of a methyl group at sites in the DNA where a cytosine nucleotide occurs next to a guanine nucleotide (CpG dinucleotides). Initial work focused on understanding associations of early life stress and DNA methylation examined these processes in rodents and found that low levels of maternal care (licking and grooming and arched-back nursing) was associated with greater methylation of the glucocorticoid receptor (GR) gene in offspring9,10. Furthermore, this work suggested that methylation was a mechanism of the effect of maternal care on the offspring HPA stress response. Emerging from this groundbreaking work with animal models, some of the earliest studies in humans documenting altered DNA methylation in association with childhood maltreatment focused on the glucocorticoid receptor (GR) gene, NR3C1, which modifies responsiveness of the HPA axis to stress exposure. Our group was the first to demonstrate altered leukocyte DNA methylation of NR3C1 in adults with childhood maltreatment11, associations of NR3C1 methylation with behavior problems and symptoms in children with early adversity12, and maltreatment as a predictor of change in NR3C1 methylation over time13. In 2016, we completed a literature review focused on altered methylation of glucocorticoid signaling genes in association with childhood maltreatment14. Several other recent reviews have also focused on methylation of glucocorticoid signaling genes15,16,17, as well as methylation of genes associated with the serotonin system18,19 in relation to adverse exposures. We expand upon these prior reviews to consider the recent explosion of research in this area that has examined additional genes as well as epigenome-wide effects. Cecil and colleagues recently conducted a systematic review of research focused on childhood maltreatment, specifically experiences of abuse and neglect, and DNA methylation20. The current review provides an important addition to the literature by utilizing a broader conceptualization of childhood maltreatment that includes any experience that involved potential for harm to the child. Our systematic approach to identify and summarize the literature related to childhood maltreatment and other adversities in association with DNA methylation captures research through March, 2020 and includes research focused on candidate genes and studies that leveraged epigenome-wide data in both children and adults.
Methods
In accordance with the Preferred Reporting Items for Systematic Reviews and Meta-Analyses (PRISMA) guidelines21, we conducted a systematic review of human studies investigating the relationship between childhood maltreatment and other adversities and DNA methylation. For the purposes of this review, childhood was defined as events occurring between birth and age 18. Maltreatment was defined as any experience that involved potential for harm to the child, including emotional, sexual, and physical abuse, and emotional and physical neglect. Other adversities included exposure to intimate partner violence or other violence, early parental death or separation, institutional deprivation, and indentured labor. Empirical articles investigating DNA methylation in association with other forms of childhood stress such as low socioeconomic status or famine, without consideration of other adversities, were not included. Studies focusing on experimentally induced stressors were also not included. In addition, because our focus was on childhood maltreatment after birth; we also did not include studies focusing on prenatal stress or substance use and DNA methylation, which have been addressed in other reviews22,23.
Studies were identified by searching PubMed/Medline for empirical articles published in March 2020 or earlier using the following search terms: (((methylation OR epigenetic) AND (child AND (maltreat OR adversity OR trauma OR early life stress OR abuse OR neglect OR ACE))), filtered by Humans. Additional searches in PubMed/Medline for studies published during this timeframe were also conducted using the following search terms: (1) emotion regulation AND methylation; (2) abuse AND methylation NOT substance NOT alcohol; (3) trauma AND methylation; (4) early life stress AND methylation; (5) maltreatment AND methylation; (6) adversity AND methylation. Additional articles published during this timeframe identified through other sources (Google Scholar alerts) were also included.
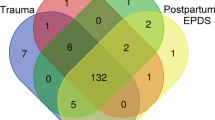
Our qualitative synthesis included empirical articles that: (1) examined the relationship between childhood maltreatment/adversity and DNA methylation; (2) provided statistical indicators to examine the impact of maltreatment during childhood on DNA methylation; and (3) described maltreatment/adversity that occurred prior to age 18. Empirical articles were excluded if they: (1) did not provide information on childhood maltreatment/adversity and DNA methylation; (2) childhood maltreatment/adversity was not clearly distinguished from adult adversities in study measurement/hypothesis testing; (3) only quality (or quantity) of parental care or support was included as a predictor; (4) were not written in English; (5) were only conducted with animals; and (6) did not use a quantitative approach to summarize research findings (i.e., were qualitative, reviews, comments, or other editorials). We did not omit articles based on small sample size; this information is included in the tables so that it can be considered when evaluating results. Figure 1 documents the methods of our systematic review to generate the final empirical articles included in our qualitative synthesis. Of note, consistent with PRISMA guidelines, records screened includes screening of both titles and abstracts.
Results
A total of 100 empirical articles were identified in our systematic review focused on the relationship between childhood maltreatment and DNA methylation. This includes 69 empirical articles focused on candidate genes and 31 empirical articles that leveraged epigenome-wide data. Twenty-eight empirical articles measured DNA methylation in childhood and 72 measured DNA methylation in adulthood. Findings are summarized in Tables 1–4, and examples are described below. Major results of each empirical article are included in the tables, though results of specific CpG sites are not detailed given significant design and analytic variability across studies. Given the focus of the current review, only the major study findings related to the association of childhood maltreatment and other adversities with DNA methylation are included in the tables. Empirical articles that focused on another condition (e.g., parental substance use, psychiatric conditions, as described in Sample description in the tables) but included a result pertaining to maltreatment are presented. Importantly, several empirical articles drew upon overlapping samples. We included information regarding the name of the study in the tables if the study name was provided in the empirical article. However, we were unable to denote all articles that may have utilized overlapping samples as the relevant information was not consistently provided in the literature.
Childhood maltreatment and methylation of candidate genes
Children
As displayed in Table 1, 16 empirical articles focused on childhood maltreatment and methylation of candidate genes in children. Most of these studies involved saliva DNA, but one used buccal cell DNA and several examined DNA from blood. The most commonly studied candidate genes were those that regulate glucocorticoid signaling, including NR3C1 which encodes the glucocorticoid receptor (GR) (six studies represented in eight empirical articles) and FKBP5 which modulates sensitivity of the GR24 (two studies represented in three empirical articles).
Most studies of children support the hypothesis that childhood adversity is associated with higher levels of methylation of NR3C1. In our own study of preschoolers, we found that childhood maltreatment status and the number of stressful life events was associated with greater NR3C1 methylation25, and NR3C1 methylation mediated effects of maltreatment and stressful life events on child internalizing behavior problems12. Likewise, in 534 school-aged children with low socioeconomic status, children who experienced maltreatment beginning in infancy or early childhood had greater NR3C1 methylation than children who had no maltreatment history26. Interestingly, there were no differences between children who experienced maltreatment that started in the preschool period or later, and those children with no maltreatment history. However, more chronic maltreatment and more types of maltreatment, were associated with greater NR3C1 methylation. Using data from the Bucharest Early Intervention Project, time in institutional care was associated with lower levels of methylation of FKBP5, which modulates sensitivity of the GR, when children were 12 years old27. Lower levels of FKBP5 methylation in association with childhood maltreatment were also observed in our sample of preschoolers28. Another study of Tanzanian children found evidence that maltreatment was associated with hypermethylation of CRH, the gene that encodes the hypothalamic corticotropin releasing hormone (CRH) and its downstream target, POMC, which encodes the pituitary precursor of ACTH29. Taken together, these findings indicate that childhood maltreatment may be associated with altered methylation of genes that regulate the child stress response, suggesting a possible mechanism for the effect of maltreatment and other adversities on poor health outcomes.
Epigenetic alterations with childhood maltreatment have been documented in additional candidate genes. Methylation of serotonin signaling genes, including SLC6A4, which encodes the serotonin receptor, and HTR2A, the gene that encodes the serotonin receptor subtype, 5-HT2A, have been investigated in association with childhood maltreatment and other interpersonal adversities27,30,31,32. Childhood adversity was associated with greater SLC6A4 methylation in two studies31,32, but lower levels of methylation in association with institutional care27. Using a subsample (n = 785) of the Avon Longitudinal Study of Parents and Children (ALSPAC), Barker et al.33 generated an inflammation-related epigenetic polygenic risk score (i-ePGS). Adversity between birth and 7 years of age was related to higher i-ePGS, which, in turn, was an indirect pathway by which adversity was associated with internalizing behavior problems in later childhood. The receptor gene for the pituitary hormone oxytocin (OXTR), best known for its role in social behavior and attachment, has also been studied in relation to early adversity in children. Fujisawa et al.34 demonstrated greater methylation of OXTR in Japanese children and adolescents with a maltreatment history (n = 44) compared with controls (n = 41), and found that OXTR methylation was negatively associated with gray matter volume in the left orbitofrontal cortex.
Adults
As displayed in Table 2, 53 empirical articles focused on childhood maltreatment and methylation of candidate genes in adults. Most studies involved analysis of DNA from blood, with a handful of studies examining DNA from saliva or buccal cells, and a few studies of DNA from brain tissue. The most commonly studied candidate genes included NR3C1 (20 empirical articles), FKBP5 (eight empirical articles), SLC6A4 (six empirical articles), and OXTR (six empirical articles).
Taken together, the majority of studies focused on NR3C1 methylation observed significant associations of childhood maltreatment and methylation of this gene, but there is considerable variability in the findings. Several studies found increased NR3C1 methylation with childhood maltreatment, but others found no effect, or reduced methylation with maltreatment. Melas and colleagues35 found that childhood adversity was associated with saliva DNA NR3C1 hypermethylation, with childhood parental loss linked to higher methylation near a transcription factor binding site. Two reports analyzed DNA from the Quebec Suicide Brain Bank in relation to childhood maltreatment retrospectively assessed via psychological autopsy, including a group with suicide and childhood maltreatment, a group with suicide but no maltreatment, and a control group. The suicide with maltreatment group had greater hippocampal NR3C1 methylation and reduced NR3C1 expression across exon 1F36, reduced NR3C1 gene expression across exons 1B, 1C, and 1H, and reduced NR3C1 methylation at exon 1H37, in comparison with the other two groups. In contrast, both suicide groups differed from controls in regions of exon 1B, with hypermethylation at two CpG sites and hypomethylation at one CpG site37. In a diverse sample of adults, childhood maltreatment was associated with greater methylation of leukocyte NR3C1, and maltreatment was associated with lower NR3C1 gene expression38. Likewise, in two samples with psychiatric disorders, childhood maltreatment was associated with greater NR3C1 methylation39,40. In contrast, in a recent study, we found that early adversity, and the number of adverse exposures, was associated with lower NR3C1 methylation in healthy adults41. Other studies found no association of maltreatment and NR3C1 methylation42,43,44,45,46, but one of these found that NR3C1 methylation moderated effects of maltreatment on cortisol response to the Trier Social Stress Test.
The literature focused on FKPB5 methylation in adults also demonstrates inconsistent effects. Klengel et al.47 found that childhood maltreatment was associated with lower methylation of FKBP5 in adults with the rs1360780 T risk allele, and this effect was observed in both a subsample of participants from the Grady Trauma Project and a replication sample. This effect was more recently replicated by Tozzi and colleagues48 who also reported that childhood maltreatment was associated with lower FKBP5 methylation among those with the T risk allele of the rs1360780 SNP. In contrast, Harms et al.49 found significant positive associations between stress in childhood and methylation of FKPB5 in young adulthood using a prospective longitudinal design, and a number of studies did not find significant associations among childhood maltreatment and FKBP5 methylation50,51,52,53, or moderation by FKBP5 risk allele50,51.
Turning to the serotonin system, several studies have examined whether childhood maltreatment is associated with methylation of SLC6A4, the gene for the serotonin transporter. Most studies have found a positive association of childhood maltreatment with methylation of regions of this gene54,55,56,57,58 although some studies show equivocal or trend-level effects59,60 or no effect61 of maltreatment on SLC6A4 methylation.
Based on the hypothesis that oxytocin may play a protective role in the biological response to stress and trauma62, five studies investigated methylation of the oxytocin-receptor-gene (OXTR) in adulthood; all five reported no overall association of childhood maltreatment with OXTR methylation. A number of these studies did report indirect or moderation effects. In a sample of 309 African American men, childhood adversity had a significant indirect effect on OXTR methylation through socioeconomic instability38. No significant direct effect of childhood adversity on OXTR methylation was observed. Likewise, while Smearman and colleagues63 did not observe simple associations of child abuse history and OXTR methylation when accounting for multiple comparisons in their sample of 393 African American adults, OXTR methylation moderated the association of childhood abuse and psychiatric symptoms.
Several other candidate genes have been examined in individual studies. For example, childhood trauma was associated with lower methylation of the proinflammatory IL-6 promoter, and lower methylation in turn was associated with greater salivary IL-6 in response to the Trier Social Stress Test64. Although this study drew upon a relatively small sample, the findings are consistent with other work demonstrating effects of childhood trauma and maltreatment on inflammatory processes.
Childhood maltreatment and epigenome-wide association studies
Arising in part out of replication inconsistencies as well as interest in identifying novel variation, the past decade has included an increase in the number of Genome Wide Association Studies (GWAS), which are agnostic by design and interrogate genetic variation across millions of common genetic variants the entire genome. However, a limitation of GWAS is the extremely large samples needed to detect effects after adjusting for multiple testing. Similar to GWAS approaches, epigenome-wide association studies interrogate markers across the entire genome. Consideration of the biology of DNA methylation as well as optimization of power has led to the development of new frameworks for analyzing multiple markers, including examination of CpG sites that are in close proximity to one another (differentially methylated regions, DMRs), examination of DNA methylation in biological pathways known to be involved in the condition, and application of analytic tools that use a ranking approach rather than relying on p values. Researchers have also developed quality control standards and approaches for addressing both concerns with Type I error as well as population stratification that may arise in diverse samples65.
Given that biological processes function as part of a larger interrelated system, research has increasingly focused on how either systems or patterns of alterations in methylation may be important in efforts to understand epigenetic modifications. Studies have begun to explore DNA methylation as a measure of molecular aging, with evidence that this epigenetic “clock” is associated with age-related disorders and mortality66,67,68,69,70 as well as with age-related development such as menopause71,72. Several analytical methods have been developed to measure epigenetic age as indicated by epigenome-wide methylation profiles73,74,75,76,77. This emerging area of research has the potential to provide exciting evidence for understanding the impact of early adversity on molecular age.
Children
As displayed in Table 3, 12 empirical articles focused on epigenome-wide effects of maltreatment and other adversities in children. These studies suggest that maltreatment is associated with variation in methylation across the genome, and several differentially methylated regions and genes have been identified. In a racially and ethnically diverse sample of 548 children, Cicchetti et al.78 found differential whole-genome methylation in children with a maltreatment history relative to children with no maltreatment. Maltreated children had higher levels of methylation at sites where methylation was generally low, and lower methylation at sites where methylation was generally high, compared to non-maltreated children. Using data from a subsample (n = 774) of the ALSPAC, Dunn et al.79 examined associations among childhood adversity and epigenome-wide methylation in children at 7 years of age. Thirty-eight differentially methylated CpG sites were identified in association with early adversity, and the developmental timing of adversity was the most salient predictor of methylation. Neither the simple presen-ce/absence of adversity, recency of adversity, nor accumulation of adverse experiences was associated with altered methylation.
Studies of early adversity and epigenetic age are just emerging in samples of children. Very recently in a sample of 247 children who were 8–16 years of age, threat-related early adversity (including abuse and violence exposure), but not deprivation (including neglect), was associated with accelerated DNA methylation age80. Furthermore, threat-related early adversity exerted a significant indirect effect on depressive symptoms through accelerated DNA methylation age, suggesting that this measure of early adversity is clinically relevant to psychiatric outcomes. This is consistent with Jovanovic et al.81 that demonstrated that violence exposure was associated with greater DNA methylation age acceleration in African American children who were 6–13 years of age.
Adults
As displayed in Table 4, 19 empirical articles focused on epigenome-wide effects of childhood maltreatment in adults. Collectively, these studies suggest that childhood maltreatment exerts epigenetic effects across the genome, yet these studies differed in their overall focus and methodology. In the ALSPAC and the MRC National Survey of Health and Development, Houtepen et al.82 identified nine differentially methylated regions (DMRs) in the genome that replicated across the two cohorts and were associated with childhood adversity. No individual CpG sites in their epigenome-wide analysis replicated across the cohorts. O’Donnell et al.83 used a different methodological approach to describe variation in DNA methylation, and found that childhood maltreatment was associated with variation in methylation at 27 years of age utilizing principal components scores to describe variation in methylation. Lutz et al.84 examined genome-wide DNA methylation and gene expression in postmortem brain samples of adults with a history of depression who died by suicide. This study found that child abuse history was associated with differential methylation specifically in oligodendrocytes in the cingulate cortex, as well as expression of myelin-related genes.
Investigations using epigenome-wide data to explore effects of early adversity on accelerated epigenetic aging have shown mixed outcomes. Lawn et al.85 used the ALSPAC and the MRC National Survey of Health and Development to examine associations of childhood psychosocial adversity, including abuse and neglect, and DNA methylation age acceleration. Childhood sexual abuse was associated with methylation age acceleration in ALSPAC, and this effect remained significant when controlling for socioeconomic position. Data regarding sexual abuse was not available in the MRC survey, however Tamman et al.86 also found that childhood sexual abuse was associated with increased DNA methylation age. Neither individual adversity types nor a cumulative measure of adversity were associated with methylation age acceleration in ALSPAC and the MRC National Survey of Health and Development85. Han et al.87 also reported a positive association between a history of childhood trauma and epigenetic aging in adults with MDD. DNA methylation age was also examined in the sample of 27 year olds described above, with analyses revealing no differences in epigenetic age as a function of childhood adversity83. Although there have been more studies focused on childhood adversity and accelerated epigenetic age in adults, few studies overall have been completed.
Discussion
This systematic review examined associations of childhood maltreatment and DNA methylation in children and adults. One hundred empirical articles focused on humans were identified. These studies included both candidate gene and epigenome-wide approaches. Strengths of the literature included: (1) rigorous approaches to measure childhood maltreatment in studies focused on DNA methylation in children, including record review methods; (2) several racially and ethnically diverse samples in the child studies; (3) diverse and innovative approaches to measuring epigenome-wide effects of maltreatment, including exploration of how early adversity may lead to epigenetic age acceleration; and (4) several replication studies focused on childhood maltreatment and methylation of glucocorticoid signaling genes in children and adults. Collectively, these studies provide evidence that childhood maltreatment and other adversities are associated with DNA methylation. Genes and pathways observed to have altered methylation in relation to childhood maltreatment, and the expected epigenetic pathways from maltreatment to health and mental health outcomes, are displayed in Fig. 2.
Studies of childhood adversity and DNA methylation often focused on methylation of candidate genes that regulate glucocorticoid signaling including NR3C1 and FKBP5. Most studies in children demonstrated increased NR3C1 methylation with maltreatment. In adults, several studies documented greater NR3C1 methylation in those exposed to childhood maltreatment, whereas others demonstrated no associations of maltreatment and NR3C1 methylation. Furthermore, in our own recent work we found lower NR3C1 methylation in association with childhood adversity41, and other studies have identified CpG sites in NR3C1 that are hypomethylated in association with maltreatment or suicide37 and post-traumatic stress disorder (PTSD)88,89. For FKBP5, Klengel and colleagues first published findings of an interaction of childhood maltreatment and FKBP5 genotype such that maltreatment was associated with lower FKBP5 methylation in adults with the T risk allele47. Lower FKBP5 methylation with maltreatment—although not interacting with genotype—has been seen in the two studies with published data in children. The maltreatment by genotype interaction predicting lower FKBP5 methylation in adults was recently replicated48, but other studies have not observed an effect of maltreatment, or an interaction of maltreatment and genotype, and one study of young adults found that stressful life events in childhood were associated with greater FKBP5 methylation, which in turn mediated effects of early stress on prefrontal brain activity49.
Many studies in adults utilized samples with chronic disease and mental health diagnoses, and did not exclude participants with consistent medication use, which may have influenced the findings. Overall levels of methylation of the NR3C1 gene promoter are generally very low, and this may also limit the ability to reliably detect group differences. However, there is evidence that NR3C1 methylation at exons 1F, 1B, and 1C, is inversely associated with GR gene expression36,37. Since GR mediates negative feedback to the HPA axis, higher methylation—and lower GR gene expression—is expected in conditions with elevated cortisol and there is some evidence for this90,91. But variable findings for methylation of NR3C1 and FKBP5 are perhaps not surprising given that maltreatment, trauma, and stress-related psychiatric disorders have been linked to both exaggerated and diminished basal and provoked cortisol concentrations, which may in part depend on measured or unmeasured individual and trauma characteristics92,93. Moreover, many genes and gene products regulate HPA axis function, and complex interactions among participant characteristics (including age and genetic ancestry), genotype (with allele frequencies that may differ based on genetic ancestry), and stress exposure may best characterize these effects.
Interaction effects may also characterize methylation of oxytocin system genes, given findings of indirect or moderation effects in the absence of simple effects63,94. Although the potential risk for statistical artifacts provide important cautions to investigations exploring interaction effects, interactions and conditional effects are known to occur in biological systems and may be appropriate to examine in large samples and/or using stratification or model invariance strategies. Methylation of serotonin signaling genes was also examined in multiple studies. Most studies of adults and some work in children found elevated methylation of SLC6A4 with maltreatment, but there are also reports of lower SLC6A4 methylation at some CpG sites with adversity, or no difference between adversity groups. Differences in epigenetic findings in SLC6A4 as well as other candidate genes may also be related to inconsistencies in the ways that the number of differentially methylated regions are assessed. For example, SLC6A4 has 81 CpG sites with studies adopting a range of approaches to assessing this large region, including selection of target sites based on the literature, multiple testing of sites, and strategies for binning methylation at multiple sites95. Given evidence for ancestry-related differences in methylation, differences across studies related to ancestry may also potentially contribute to apparent inconsistencies96.
Studies using agnostic approaches to exploring multiple markers across the epigenome also suggested that maltreatment is associated with variation in methylation, though these epigenome-wide studies have generally failed to identify commonly studied candidate genes, leading to questions about candidate approaches. Several reasons for these inconsistencies have been advanced, including large effect sizes needed to detect an effect after multiple testing corrections, biological considerations such as the potential combinatory impact of genotype and DNA methylation (see ref. 83 for an example of integration of genotype and epigenetic influences), interactions among systems of genes, and differences in aspects of the phenotyping and heterogeneity of different studies. Due to traditional approaches to corrections for multiple testing, studies that analyze methylation of multiple markers across the genome tend to require large samples which are often heterogeneous and may have less intensive phenotyping and measurement of maltreatment and other relevant exposures. Promising markers identified with epigenome-wide approaches show some consistency in terms of their roles in neural cell development (BDNF, KITLG, and POU3F1), signaling and apoptosis (LINGO3 and 2NPFF2), neural influences on movement (ALS2), neuroinflammation (ITGB1), and some tentative evidence for immune markers (CXCL1).
A recent area of epigenome-wide research involves DNA methylation as an indicator of molecular age. A number of analytic strategies have been used to examine how early life trauma (and/or symptoms associated with trauma such as depression or post-traumatic stress disorder) may accelerate aging. A number of adult and child studies find age acceleration among individuals exposed to maltreatment (meaning that DNA methylation age is older than the chronological age). Childhood maltreatment and other adverse exposures alter epigenome-wide profiles, thereby likely contributing to chronic disease and physical aging97,98,99,100,101. Interestingly, the particular markers observed to be relevant in methylation age acceleration research may point toward biological systems implicated in the link between maltreatment and disease. For example, the method described by Horvath includes CpG sites enriched for glucocorticoid response elements99. New methods for capturing DNA methylation age continue to be developed, and very recently Belsky and colleagues102 reported a new algorithm that was derived from longitudinal data of 18 biomarkers of organ-system integrity to capture the rate of aging up to the time of measurement. This new algorithm, but not other previously established measures of DNA methylation age, was associated with childhood maltreatment in sample of 1658 young adults who were longitudinally followed since childhood. As new methods to capture age acceleration continue to develop, systematic reviews and meta-analytic approaches will continue to be important to synthesize associations of child maltreatment and the range of measures of methylation age.
Studies of maltreatment and methylation in children were often characterized by careful measurement of adverse exposures using structured record review techniques and in-depth interview methods, and they often utilized prospective and longitudinal designs. In children, DNA methylation was most frequently measured in saliva or buccal epithelial cells, with fewer studies measuring methylation in blood. Many of the samples were racially and ethnically diverse, and these studies covered the full developmental spectrum, from early childhood through adolescence. In contrast, nearly all the adult studies drew upon retrospective reports of childhood maltreatment. Utilization of prospective longitudinal designs and record review techniques would address a significant gap in the literature. Furthermore, in contrast to studies of maltreatment and methylation of candidate genes in children, most adult studies measured methylation in leukocyte DNA.
Moving beyond simple effects of maltreatment on DNA methylation, several studies highlighted the importance of the developmental timing of exposure to adversity26,27,78,79. More research is needed to understand how early adversity at each developmental epoch may be associated with differences in epigenetic marks. Moreover, longitudinal research is needed to explore whether epigenetic changes secondary to early adversity may “reset” during later developmental periods or perhaps may be impacted by later experiences, which either attenuate or compound early adversity. Retrospective research with adults who report childhood adversity has numerous benefits in terms of cost- and time-effectiveness; however, reliance on retrospective reports of childhood adversity present important methodological challenges related to recalling the timing of childhood events, ability to recall very early life events, and systematic recall biases, introducing measurement limitations that are difficult to overcome103. Many studies use the Childhood Trauma Questionnaire104 or other self-report measures. Some self-report measures, such as the Traumatic Life Events Questionnaire105 and the Maltreatment and Abuse Chronology of Exposure106, include trauma characteristics such as the age or frequency of occurrence, which may be important determinants of epigenetic and phenotypic outcomes. Future research may also benefit from interview approaches that can help with retrospective recall such as participant-tailored anchoring.
Although many adult studies reviewed here relied on recall, much can be learned from the few studies that capitalized on longitudinal cohorts. For example, Harms et al.49 assessed stress when children were 9–13 years old and then methylation approximately 10 years later. O’Donnell et al.83 assessed childhood maltreatment in the first 15 years of life and methylation at age 27. Future research should capitalize on existing studies of children that utilized rigorous measures of the environment as they develop over time to expand the literature in adults. Likewise, very few studies drew upon repeated assessments of DNA methylation over time. In our own research, we demonstrated that childhood maltreatment and other adversities are associated with change in methylation of glucocorticoid signaling genes over time in early childhood13,107. Future research should further examine maltreatment as a predictor of change in methylation across development.
Importantly, the majority of studies focused on exposure to childhood maltreatment without consideration of the prenatal environment. Prenatal exposures exert epigenetic effects on several biological systems, particularly the development of the child stress response. For example, intimate partner violence in pregnancy is associated with increased NR3C1 in late childhood and adolescence108. Smoking and depression in pregnancy have also been associated with altered methylation of placental stress-related genes, such as NR3C1 and HSD11B2, which encodes the enzyme that inactivates cortisol109,110,111,112. Associations of childhood maltreatment and DNA methylation may be partially accounted for by prenatal environmental factors. Conversely, maltreatment may exert a unique and independent effect on DNA methylation above and beyond prenatal exposures. Barker et al.33 found that child adversity between birth and 7 years of age was associated with an inflammation-related epigenetic polygenic risk score (i-ePGS) at age 7, but there was no association of prenatal adversity and the i-ePGS. Future research should disentangle prenatal exposures and adversities experienced in childhood to better understand effects of maltreatment on DNA methylation in both childhood and adulthood.
Very few studies examined sex differences in the effects of childhood maltreatment on DNA methylation. Studies focused on in utero stress exposure often find sex differences in epigenetic pathways and observed outcomes in both preclinical and human models113,114,115. For example, Braithwaite et al.116 found that maternal depressive symptoms in pregnancy were associated with greater methylation of NR3C1 in male, but not female, infants. Stroud et al.109 found that the moderating effects of placental HSD11B2 methylation on links between prenatal major depressive disorder and infant cortisol response emerged most strongly for newborn daughters, whereas direct and moderating effects of SLC6A4 gene expression were evident only for sons109. In one of the few studies identified in this review that examined sex differences, Cicchetti et al.78 found that boys and girls showed different directions of the effect of maltreatment on methylation of ALDH2, a gene that encodes a key enzyme in the metabolism of alcohol. Sex differences were also observed in the effect of the developmental timing of adversity. As we have shown in a meta-analysis with ADCYAP1R1, the adenylate cyclase activating receptor gene associated with PTSD, depending on the functional outcomes of the gene(s), sex and developmental differences are reasonable to expect117. Future work on maltreatment and DNA methylation should consider the role of child sex to ensure that important moderation effects are not being overlooked.
An additional future direction is to move from association to causal inference. Maltreatment is often confounded with additional measures of adversity, including poverty and other sociodemographic factors, as well as personality factors, and potentially genetic factors. Many studies reviewed utilized statistical control for potential confounding factors; however, true random assignment designs are not ethical in humans. Thus, synthesizing across preclinical studies (where random assignment is possible) and human association studies will be critical. Additionally, innovative designs in human studies, including random assignment to interventions to reduce maltreatment, and control groups that are demographically and psychosocially matched to maltreatment groups, may allow the field to move closer to causal inference. Research considering intervention effects on change in methylation over time may also address this gap in knowledge. Indeed, we found that service utilization was associated with increases in FKBP5 methylation over time in preschoolers with early adversity107. More recently, Vinkers et al.118 observed changes in methylation among soldiers successfully treated for PTSD such that changes in methylation were observed among soldiers who had reductions in PTSD symptoms. Taken together, this work provides initial evidence that psychosocial interventions exert influence on the epigenome.
DNA methylation has been measured in several tissue types, including blood, saliva, and buccal cells. Yet, the majority of studies focused on DNA methylation in children utilized saliva and buccal cells, and fewer examined methylation in blood. Although some researchers have questioned the value of peripheral markers in research aimed at understanding psychobehavioral outcomes believed to be related to central brain processes, with some researchers suggesting that psychiatric epigenetic research should be limited to brain tissue, a number of studies point to the value of peripheral indicators. Research has shown reasonable concordance between gene-expression signals in blood and brain119,120,121 and primate research has identified correspondence in DNA methylation profiles in the brain and blood122. Similar gene expression was found in the cerebellum and peripheral blood mononuclear cells (PBMCs) across 4000 unique genes123. Correspondence between DNA methylation in PBMCs and postmortem brain tissue was identified with respect to a marker of reward and stress-induced responses124. Recent research showed good correspondence of DNA methylation in blood and saliva125. Studies of methylation age have also shown consistency across tissue types75. Interestingly, there is some evidence126 that saliva samples may be more strongly associated with brain methylation than blood samples, although correlations of methylation in the brain with saliva, blood, and buccal cell DNA were all observed to be high and the strength of these associations appear to depend on the genomic region of interest127. These studies have provided helpful signals regarding the utility of blood, brain, saliva, and buccal cells, and Epigenomic Roadmap datasets now provide some insights into the ways that peripheral methylation profiles may map on to those in brain tissue128,129. Nonetheless, cell type heterogeneity remains a significant challenge for epigenetic research that has been addressed using a number of strategies such as cell counts, cell separation and examination of single cells, and analytic strategies that account for cell type130,131,132,133.
Another concern involves the technology and analytic strategies used to interrogate genome methylation. Power to detect true differences and false positives are both major concerns, so large samples sizes and replication samples are required, but very large studies may not be able to provide rich data on maltreatment. Researchers have also described the importance of considering reliability of BeadChip technology, the most frequently applied technology for interrogating the genome, as a function of: (a) sample type, with lower reliability and replicability in dried blood spots, (b) tissue type, with some probes demonstrating greater cross-tissue concordance, (c) platform (such as Ilumina Infinium HumanMethylation450 BeadChip with nearly 500,000 sites vs Ilumina MethylationEPIC BeadChip with nearly 850,000 sites), and (d) observed variability at each site, with lower reliability at sites with less variability134,135,136,137. Research is needed to carefully document reliability considerations for proper assessment of reproducibility; interested readers are directed to a thorough discussion of reliability, replication, and reproducibility in DNA methylation measurement (see ref. 134). In addition to the importance of clear site level documentation of findings in publications, suggestions for addressing reliability concerns in ongoing research include analytic approaches such as dropping CpG sites with low observed variability, replicating findings with procedures such as pyrosequencing, and integrating preanalysis reliability metrics134,136,137. Focusing the analyses on regions with more variability across participants reduces the likelihood of false positives, and also reduces the number of comparisons, thereby increasing power. It is also important to note that methylation arrays assess only CpG sites; this yields fewer sites than whole-genome bisulfite sequencing approaches, and some research suggests some important variability occurs outside of these CpG islands137,138. Standards in this field continue to be refined, and researchers should follow these developments closely. At a minimum, researchers should implement strategies to address quality control (including thorough evaluations of normalization procedures and strategies for addressing batch effects), cell heterogeneity, and sample ancestry65,139,140,141.
Our systematic review examined associations of maltreatment and other interpersonal adversities during childhood with DNA methylation in children and adults. We identified 100 studies, including studies utilizing both candidate gene approaches and epigenome-wide analyses. Several strengths of the literature were identified, as well as directions for future research. Key challenges facing the field and associated recommendations are described in Fig. 3. We included both studies focused on childhood maltreatment and other adversities, as well as studies that were not focused specifically on early adversity but included measurement of adversity in their examination of another topic (e.g., psychiatric or physical health diagnosis). Inclusion of these studies represents both a strength and limitation due to selection biases inherent to these designs, as well as confounds such as the presence of medical conditions and medications that may influence methylation. We acknowledge several other limitations to the current review. Specifically, (1) the present review focused on DNA methylation and did not include studies addressing other epigenetic modifications (e.g., microRNA, histone modifications); (2) the review did not address the functional impact (e.g., gene expression, proteins) of DNA methylation alterations in various studies; and (3) although systematic, the review was qualitative, and thus, does not provide information regarding effect sizes or heterogeneity of included studies. Meta-analytic reports of the most commonly studied candidate genes and EWAS results would build upon our qualitative analysis; however, the extensive number of individual CpG sites both within each candidate gene and across the EWAS studies poses a significant challenge. Furthermore, a critical limitation of the literature is that the location of individual CpG sites is often not consistently identified in empirical manuscripts, further precluding meta-analytic approaches. Future research should more carefully detail the precise location of CpG sites to facilitate future meta-analysis efforts.
In less than two decades, there has been an explosion of research in childhood maltreatment and epigenetics. Mirroring trends observed in genetic research, studies have increasingly focused on sampling methylation across the genome. As with genetic research, advantages of this approach include opportunities to identify key mechanisms that would otherwise have been overlooked. However, this approach generally requires larger samples to account for multiple testing which in turn has been criticized for less nuanced phenotyping of samples permitted in smaller cohorts. Some of these concerns may be somewhat addressed through collaborations such as efforts undertaken by the Psychiatric Genetics Consortium working groups, particularly when care is taken to balance genomic inflation/deflation and to address concerns that can arise with diverse samples interrogated on differing platforms65. Researchers have also begun to explore ways that methodological approaches may improve understanding of epigenetic factors and adversity outcomes such as repeated measurement, sampling across tissues, twin/family approaches, and psychological/pharmacological treatments. Epigenetic approaches are also perhaps best understood in combination with thorough assessments of genotype, epigenome, and gene expression. Many factors related to the type and timing of adversity, availability and quality of buffers to adversity, and new events and time since early adversity are likely critically important influences on negative outcomes of adversity. Ongoing recognition of the complexity of biological and environmental factors as well as phenotypic nuances is critically needed—apparent inconsistences do not necessarily negate findings from either of the divergent studies but rather may point to important considerations for ongoing research and theoretical conceptualizations.
References
Brown, D. W. et al. Adverse childhood experiences and the risk of premature mortality. Am. J. Prev. Med. 37, 389–396 (2009).
Burns, B. J. et al. Mental health need and access to mental health services by youths involved with child welfare: a national survey. J. Am. Acad. Child Adolesc. Psychiatry 43, 960–970 (2004).
Cohen, S., Janicki-Deverts, D. & Miller, G. E. Psychological stress and disease. JAMA 298, 1685–1687 (2007).
Gilbert, R. et al. Burden and consequences of child maltreatment in high-income countries. Lancet 373, 68–81 (2009).
Grippo, A. J. & Johnson, A. K. Stress, depression and cardiovascular dysregulation: a review of neurobiological mechanisms and the integration of research from preclinical disease models. Stress 12, 1–21 (2009).
Oh, D. L. et al. Systematic review of pediatric health outcomes associated with childhood adversity. BMC Pediatr. 18, 83 (2018).
Schneiderman, N., Ironson, G. & Siegel, S. D. Stress and health: psychological, behavioral, and biological determinants. Annu. Rev. Clin. Psychol. 1, 607–628 (2005).
Kim, J. K., Samaranayake, M. & Pradhan, S. Epigenetic mechanisms in mammals. Cell Mol. Life Sci. 66, 596–612 (2009).
Weaver, I. C. et al. Epigenetic programming by maternal behavior. Nat. Neurosci. 7, 847–854 (2004).
Weaver, I. C. et al. Reversal of maternal programming of stress responses in adult offspring through methyl supplementation: altering epigenetic marking later in life. J. Neurosci. 25, 11045–11054 (2005).
Tyrka, A. R., Price, L. H., Marsit, C., Walters, O. C. & Carpenter, L. L. Childhood adversity and epigenetic modulation of the leukocyte glucocorticoid receptor: preliminary findings in healthy adults. PLoS ONE 7, e30148 (2012).
Parade, S. H. et al. Methylation of the glucocorticoid receptor gene promoter in preschoolers: links with internalizing behavior problems. Child Dev. 87, 86–97 (2016).
Parent, J. et al. Dynamic stress-related epigenetic regulation of the glucocorticoid receptor gene promoter during early development: The role of child maltreatment. Dev. Psychopathol. 29, 1635–1648 (2017).
Tyrka, A. R., Ridout, K. K. & Parade, S. H. Childhood adversity and epigenetic regulation of glucocorticoid signaling genes: Associations in children and adults. Dev. Psychopathol. 28, 1319–1331 (2016).
Liu, P. Z. & Nusslock, R. How stress gets under the skin: early life adversity and glucocorticoid receptor epigenetic regulation. Curr. Genomics 19, 653–664 (2018).
Turecki, G. & Meaney, M. J. Effects of the social environment and stress on glucocorticoid receptor gene methylation: a systematic review. Biol. Psychiatry 79, 87–96 (2016).
Watkeys, O. J., Kremerskothen, K., Quide, Y., Fullerton, J. M. & Green, M. J. Glucocorticoid receptor gene (NR3C1) DNA methylation in association with trauma, psychopathology, transcript expression, or genotypic variation: a systematic review. Neurosci. Biobehav. Rev. 95, 85–122 (2018).
Palma-Gudiel, H. & Fananas, L. An integrative review of methylation at the serotonin transporter gene and its dialogue with environmental risk factors, psychopathology and 5-HTTLPR. Neurosci. Biobehav. Rev. 72, 190–209 (2017).
Provenzi, L., Giorda, R., Beri, S. & Montirosso, R. SLC6A4 methylation as an epigenetic marker of life adversity exposures in humans: A systematic review of literature. Neurosci. Biobehav. Rev. 71, 7–20 (2016).
Cecil, C. A., Zhang, Y. & Nolte, T. Childhood maltreatment and DNA methylation: a systematic review. Neurosc. Biobehav. Rev. 112, 392–409 (2020).
Moher, D., Liberati, A., Tetzlaff, J., Altman, D. G. & Group, P. Preferred reporting items for systematic reviews and meta-analyses: the PRISMA Statement. Open Med. 3, e123–e130 (2009).
Cao-Lei, L. et al. Prenatal stress and epigenetics. Neurosci. Biobehav. Rev. https://doi.org/10.1016/j.neubiorev.2017.05.016 (2017).
Sosnowski, D. W., Booth, C., York, T. P., Amstadter, A. B. & Kliewer, W. Maternal prenatal stress and infant DNA methylation: a systematic review. Dev. Psychobiol. 60, 127–139 (2018).
Binder, E. B. The role of FKBP5, a co-chaperone of the glucocorticoid receptor in the pathogenesis and therapy of affective and anxiety disorders. Psychoneuroendocrinology 34(Suppl 1), S186–S195 (2009).
Tyrka, A. R. et al. Methylation of exons 1D, 1F, and 1H of the glucocorticoid receptor gene promoter and exposure to adversity in preschool-aged children. Dev. Psychopathol. 27, 577–585 (2015).
Cicchetti, D. & Handley, E. D. Methylation of the glucocorticoid receptor gene, nuclear receptor subfamily 3, group C, member 1 (NR3C1), in maltreated and nonmaltreated children: associations with behavioral undercontrol, emotional lability/negativity, and externalizing and internalizing symptoms. Dev. Psychopathol. 29, 1795–1806 (2017).
Non, A. L. et al. DNA methylation at stress-related genes is associated with exposure to early life institutionalization. Am. J. Phys. Anthropol. 161, 84–93 (2016).
Tyrka, A. R. et al. Childhood maltreatment and methylation of FK506 binding protein 5 gene (FKBP5). Dev. Psychopathol. 27, 1637–1645 (2015).
Hecker, T., Radtke, K. M., Hermenau, K., Papassotiropoulos, A. & Elbert, T. Associations among child abuse, mental health, and epigenetic modifications in the proopiomelanocortin gene (POMC): a study with children in Tanzania. Dev. Psychopathol. 28, 1401–1412 (2016).
Parade, S. H. et al. Stress exposure and psychopathology alter methylation of the serotonin receptor 2A (HTR2A) gene in preschoolers. Dev. Psychopathol. 29, 1619–1626 (2017).
Timothy, A. et al. Influence of early adversity on cortisol reactivity, SLC6A4 methylation and externalizing behavior in children of alcoholics. Prog. Neuropsychopharmacol. Biol. Psychiatry 94, 109649 (2019).
van der Knaap, L. J. et al. Adverse life events and allele-specific methylation of the serotonin transporter gene (SLC6A4) in adolescents: the TRAILS study. Psychosom. Med. 77, 246–255 (2015).
Barker, E. D. et al. Inflammation-related epigenetic risk and child and adolescent mental health: A prospective study from pregnancy to middle adolescence. Dev. Psychopathol. 30, 1145–1156 (2018).
Fujisawa, T. X. et al. Oxytocin receptor DNA methylation and alterations of brain volumes in maltreated children. Neuropsychopharmacology https://doi.org/10.1038/s41386-019-0414-8 (2019).
Melas, P. A. et al. Genetic and epigenetic associations of MAOA and NR3C1 with depression and childhood adversities. Int. J. Neuropsychopharmacol. 16, 1513–1528 (2013).
McGowan, P. O. et al. Epigenetic regulation of the glucocorticoid receptor in human brain associates with childhood abuse. Nat. Neurosci. 12, 342–348 (2009).
Labonte, B. et al. Differential glucocorticoid receptor exon 1(B), 1(C), and 1(H) expression and methylation in suicide completers with a history of childhood abuse. Biol. Psychiatry 72, 41–48 (2012).
Bustamante, A. C. et al. Glucocorticoid receptor DNA methylation, childhood maltreatment and major depression. J. Affect Disord. 206, 181–188 (2016).
Martin-Blanco, A. et al. Association between methylation of the glucocorticoid receptor gene, childhood maltreatment, and clinical severity in borderline personality disorder. J. Psychiatr. Res. 57, 34–40 (2014).
Perroud, N. et al. Increased methylation of glucocorticoid receptor gene (NR3C1) in adults with a history of childhood maltreatment: a link with the severity and type of trauma. Transl. Psychiatry 1, e59 (2011).
Tyrka, A. R. et al. Methylation of the leukocyte glucocorticoid receptor gene promoter in adults: associations with early adversity and depressive, anxiety and substance-use disorders. Transl. Psychiatry 6, e848 (2016).
Alexander, N. et al. Glucocorticoid receptor gene methylation moderates the association of childhood trauma and cortisol stress reactivity. Psychoneuroendocrinology 90, 68–75 (2018).
Schur, R. R. et al. Glucocorticoid receptor exon 1F methylation and the cortisol stress response in health and disease. Psychoneuroendocrinology 97, 182–189 (2018).
Steiger, H., Labonte, B., Groleau, P., Turecki, G. & Israel, M. Methylation of the glucocorticoid receptor gene promoter in bulimic women: associations with borderline personality disorder, suicidality, and exposure to childhood abuse. Int. J. Eat. Disord. 46, 246–255 (2013).
Vangeel, E. et al. Chronic fatigue syndrome and DNA hypomethylation of the glucocorticoid receptor gene promoter 1F region: associations with HPA axis hypofunction and childhood trauma. Psychosom. Med. 77, 853–862 (2015).
Wang, W. et al. Increased methylation of glucocorticoid receptor gene promoter 1F in peripheral blood of patients with generalized anxiety disorder. J. Psychiatr. Res. 91, 18–25 (2017).
Klengel, T. et al. Allele-specific FKBP5 DNA demethylation mediates gene–childhood trauma interactions. Nature neuroscience, 16, 33–41 (2013).
Tozzi, L. et al. Epigenetic changes of FKBP5 as a link connecting genetic and environmental risk factors with structural and functional brain changes in major depression. Neuropsychopharmacology 43, 1138–1145 (2018).
Harms, M. B. et al. Early life stress, FK506 binding protein 5 gene (FKBP5) methylation, and inhibition-related prefrontal function: A prospective longitudinal study. Dev. Psychopathol. 29, 1895–1903 (2017).
Bustamante, A. C. et al. FKBP5 DNA methylation does not mediate the association between childhood maltreatment and depression symptom severity in the Detroit Neighborhood Health Study. J. Psychiatr. Res. 96, 39–48 (2018).
Klinger-Konig, J. et al. Methylation of the FKBP5 gene in association with FKBP5 genotypes, childhood maltreatment and depression. Neuropsychopharmacology 44, 930–938 (2019).
Farrell, C. et al. DNA methylation differences at the glucocorticoid receptor gene in depression are related to functional alterations in hypothalamic-pituitary-adrenal axis activity and to early life emotional abuse. Psychiatry Res. 265, 341–348 (2018).
Yeo, S. et al. The influence of FKBP5 genotype on expression of FKBP5 and other glucocorticoid-regulated genes, dependent on trauma exposure. Genes Brain Behav. 16, 223–232 (2017).
Beach, S. R. H., Brody, G. H., Todorov, A. A., Gunter, T. D. & Philibert, R. A. Methylation at SLC6A4 is linked to family history of child abuse: an examination of the Iowa Adoptee sample. Am. J. Med. Genet. B Neuropsychiatr. Genet. 153B, 710–713 (2010).
Beach, S. R., Brody, G. H., Todorov, A. A., Gunter, T. D. & Philibert, R. A. Methylation at 5HTT mediates the impact of child sex abuse on women’s antisocial behavior: an examination of the Iowa adoptee sample. Psychosom. Med. 73, 83–87 (2011).
Beach, S. R. et al. Impact of child sex abuse on adult psychopathology: a genetically and epigenetically informed investigation. J. Fam. Psychol. 27, 3 (2013).
Booij, L. et al. DNA methylation of the serotonin transporter gene in peripheral cells and stress-related changes in hippocampal volume: a study in depressed patients and healthy controls. PLoS ONE 10, e0119061 (2015).
Kang, H. J. et al. Association of SLC6A4 methylation with early adversity, characteristics and outcomes in depression. Prog. Neuropsychopharmacol. Biol. Psychiatry 44, 23–28 (2013).
Peng, H. et al. Childhood trauma, DNA methylation of stress-related genes, and depression: findings from two monozygotic twin studies. Psychosom. Med. 80, 599–608 (2018).
Okada, S. et al. The potential of SLC6A4 gene methylation analysis for the diagnosis and treatment of major depression. J. Psychiatr. Res. 53, 47–53 (2014).
Wankerl, M. et al. Effects of genetic and early environmental risk factors for depression on serotonin transporter expression and methylation profiles. Transl. Psychiatry 4, e402 (2014).
Olff, M., Langeland, W., Witteveen, A. & Denys, D. A psychobiological rationale for oxytocin in the treatment of posttraumatic stress disorder. CNS Spectr. 15, 522–530 (2010).
Smearman, E. L. et al. Oxytocin receptor genetic and epigenetic variations: association with child abuse and adult psychiatric symptoms. Child Dev. 87, 122–134 (2016).
Janusek, L. W., Tell, D., Gaylord-Harden, N. & Mathews, H. L. Relationship of childhood adversity and neighborhood violence to a proinflammatory phenotype in emerging adult African American men: an epigenetic link. Brain Behav. Immun. 60, 126–135 (2017).
Ratanatharathorn, A. et al. Epigenome‐wide association of PTSD from heterogeneous cohorts with a common multi‐site analysis pipeline. Am. J. Med. Genet. Part B Neuropsychiatr. Genet. 174, 619–630 (2017).
Horvath, S. et al. Decreased epigenetic age of PBMCs from Italian semi-supercentenarians and their offspring. Aging 7, 1159–1170 (2015).
Christiansen, L. et al. DNA methylation age is associated with mortality in a longitudinal Danish twin study. Aging cell 15, 149–154 (2016).
Levine, M. E. et al. DNA methylation age of blood predicts future onset of lung cancer in the women’s health initiative. Aging (Albany NY) 7, 690 (2015).
Marioni, R. E. et al. DNA methylation age of blood predicts all-cause mortality in later life. Genome Biol. 16, 1–12 (2015).
Marioni, R. E. et al. The epigenetic clock and telomere length are independently associated with chronological age and mortality. Int. J. Epidemiol. 45, 424–432 (2016).
Levine, M. E. et al. Menopause accelerates biological aging. Proc. Natl Acad. Sci. USA 113, 9327–9332 (2016).
Kanherkar, R. R., Bhatia-Dey, N. & Csoka, A. B. Epigenetics across the human lifespan. Front. Cell Dev. Biol. 2, 49 (2014).
Lu, A. T. et al. DNA methylation GrimAge strongly predicts lifespan and healthspan. Aging (Albany NY) 11, 303–327 (2019).
Zhang, Q. et al. Improved precision of epigenetic clock estimates across tissues and its implication for biological ageing. Genome Med. 11, 54 (2019).
Horvath, S. DNA methylation age of human tissues and cell types. Genome Biol. 14, 1–20 (2013).
Levine, M. E. et al. An epigenetic biomarker of aging for lifespan and healthspan. Aging (Albany NY) 10, 573 (2018).
Hannum, G. et al. Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol. Cell 49, 359–367 (2013).
Cicchetti, D., Hetzel, S., Rogosch, F. A., Handley, E. D. & Toth, S. L. An investigation of child maltreatment and epigenetic mechanisms of mental and physical health risk. Dev. Psychopathol. 28, 1305–1317 (2016).
Dunn, E. C. et al. Sensitive periods for the effect of childhood adversity on DNA methylation: results from a prospective, longitudinal study. Biol. Psychiatry 85, 838–849 (2019).
Sumner, J. A., Colich, N. L., Uddin, M., Armstrong, D. & McLaughlin, K. A. Early experiences of threat, but not deprivation, are associated with accelerated biological aging in children and adolescents. Biol. Psychiatry 85, 268–278 (2019).
Jovanovic, T. et al. Exposure to violence accelerates epigenetic aging in children. Sci. Rep. 7, 8962 (2017).
Houtepen, L. C. et al. Childhood adversity and DNA methylation in two population-based cohorts. Transl. Psychiatry 8, 266 (2018).
O’Donnell, K. J. et al. DNA methylome variation in a perinatal nurse-visitation program that reduces child maltreatment: a 27-year follow-up. Transl. Psychiatry 8, 15 (2018).
Lutz, P. E. et al. Association of a history of child abuse with impaired myelination in the anterior cingulate cortex: convergent epigenetic, transcriptional, and morphological evidence. Am. J. Psychiatry 174, 1185–1194 (2017).
Lawn, R. B. et al. Psychosocial adversity and socioeconomic position during childhood and epigenetic age: analysis of two prospective cohort studies. Hum. Mol. Genet. 27, 1301–1308 (2018).
Tamman, A. J. F. et al. Accelerated DNA methylation aging in U.S. Military Veterans: results from the National Health and Resilience in Veterans Study. Am. J. Geriatr. Psychiatry 27, 528–532 (2019).
Han, L. K. M. et al. Epigenetic aging in major depressive disorder. Am. J. Psychiatry 175, 774–782 (2018).
Labonte, B., Azoulay, N., Yerko, V., Turecki, G. & Brunet, A. Epigenetic modulation of glucocorticoid receptors in posttraumatic stress disorder. Transl. Psychiatry 4, e368 (2014).
Yehuda, R. et al. Lower methylation of glucocorticoid receptor gene promoter 1F in peripheral blood of veterans with posttraumatic stress disorder. Biol. Psychiatry 77, 356–364 (2015).
Oberlander, T. F. et al. Prenatal exposure to maternal depression, neonatal methylation of human glucocorticoid receptor gene (NR3C1) and infant cortisol stress responses. Epigenetics 3, 97–106 (2008).
Dadds, M. R., Moul, C., Hawes, D. J., Mendoza Diaz, A. & Brennan, J. Individual differences in childhood behavior disorders associated with epigenetic modulation of the cortisol receptor gene. Child Dev. 86, 1311–1320 (2015).
Bernard, K., Frost, A., Bennett, C. B. & Lindhiem, O. Maltreatment and diurnal cortisol regulation: a meta-analysis. Psychoneuroendocrinology 78, 57–67 (2017).
Bosch, N. M. et al. Timing matters: long term effects of adversities from prenatal period up to adolescence on adolescents’ cortisol stress response. The TRAILS study. Psychoneuroendocrinology 37, 1439–1447 (2012).
Kogan, S. M., Bae, D., Cho, J., Smith, A. K. & Nishitani, S. Childhood adversity, socioeconomic instability, oxytocin-receptor-gene methylation, and romantic-relationship support among young African American men. Psychol. Sci. 30, 1234–1244 (2019).
Philibert, R. et al. Serotonin transporter mRNA levels are associated with the methylation of an upstream CpG island. Am. J. Med. Genet. B Neuropsychiatr. Genet. 144B, 101–105 (2007).
Galanter, J. M. et al. Differential methylation between ethnic sub-groups reflects the effect of genetic ancestry and environmental exposures. elife https://doi.org/10.7554/eLife.20532 (2017).
Wolf, E. J. & Schnurr, P. P. Posttraumatic stress disorder-related cardiovascular disease and accelerated cellular aging. Psychiatr. Ann. 46, 527–532 (2016).
Wolf, E. J. & Morrison, F. G. Traumatic stress and accelerated cellular aging: from epigenetics to cardiometabolic disease. Curr. Psychiatry Rep. 19, 75 (2017).
Zannas, A. S. et al. Lifetime stress accelerates epigenetic aging in an urban, African American cohort: relevance of glucocorticoid signaling. Genome Biol. 16, 1–12 (2015).
Wolf, E. J. et al. Traumatic stress and accelerated DNA methylation age: a meta-analysis. Psychoneuroendocrinology 92, 123–134 (2018).
Boks, M. P. et al. Longitudinal changes of telomere length and epigenetic age related to traumatic stress and post-traumatic stress disorder. Psychoneuroendocrinology 51, 506–512 (2015).
Belsky, D. W. et al. Quantification of the pace of biological aging in humans through a blood test, the DunedinPoAm DNA methylation algorithm. elife https://doi.org/10.7554/eLife.54870 (2020).
Baldwin, J. R., Reuben, A., Newbury, J. B. & Danese, A. Agreement between prospective and retrospective measures of childhood maltreatment: a systematic review and meta-analysis. JAMA Psychiatry 76, 584–593 (2019).
Bernstein, D. Childhood Trauma Questionnaire: a Retrospective Self-Report Manual (The Psychological Corporation, 1998).
Kubany, E. S. et al. Development and preliminary validation of a brief broad-spectrum measure of trauma exposure: the Traumatic Life Events Questionnaire. Psychol. Assess. 12, 210 (2000).
Teicher, M. H. & Parigger, A. The ‘Maltreatment and Abuse Chronology of Exposure’(MACE) scale for the retrospective assessment of abuse and neglect during development. PLoS ONE 10, e0117423 (2015).
Parade, S. H. et al. Change in FK506 binding protein 5 (FKBP5) methylation over time among preschoolers with adversity. Dev. Psychopathol. 29, 1627–1634 (2017).
Radtke, K. M. et al. Transgenerational impact of intimate partner violence on methylation in the promoter of the glucocorticoid receptor. Transl. Psychiatry 1, e21 (2011).
Stroud, L. R. et al. Prenatal major depressive disorder, placenta glucocorticoid and serotonergic signaling, and infant cortisol response. Psychosom. Med. 78, 979–990 (2016).
Conradt, E., Lester, B. M., Appleton, A. A., Armstrong, D. A. & Marsit, C. J. The roles of DNA methylation of NR3C1 and 11beta-HSD2 and exposure to maternal mood disorder in utero on newborn neurobehavior. Epigenetics 8, 1321–1329 (2013).
Stroud, L. R. et al. Maternal smoking during pregnancy and infant stress response: test of a prenatal programming hypothesis. Psychoneuroendocrinology 48, 29–40 (2014).
Stroud, L. R. et al. Epigenetic regulation of placental NR3C1: mechanism underlying prenatal programming of infant neurobehavior by maternal smoking? Child Dev. 87, 49–60 (2016).
Bale, T. L. Sex differences in prenatal epigenetic programming of stress pathways. Stress 14, 348–356 (2011).
Sutherland, S. & Brunwasser, S. M. Sex differences in vulnerability to prenatal stress: a Review of the Recent Literature. Curr. Psychiatry Rep. 20, 102 (2018).
Sandman, C. A., Glynn, L. M. & Davis, E. P. Is there a viability-vulnerability tradeoff? Sex differences in fetal programming. J. Psychosom. Res. 75, 327–335 (2013).
Braithwaite, E. C., Kundakovic, M., Ramchandani, P. G., Murphy, S. E. & Champagne, F. A. Maternal prenatal depressive symptoms predict infant NR3C1 1F and BDNF IV DNA methylation. Epigenetics 10, 408–417 (2015).
Lind, M. J. et al. Association of posttraumatic stress disorder with rs2267735 in the ADCYAP1R1 gene: a meta-analysis. J. Trauma Stress 30, 389–398 (2017).
Vinkers, C. H. et al. Successful treatment of post-traumatic stress disorder reverses DNA methylation marks. Mol. Psychiatry https://doi.org/10.1038/s41380-019-0549-3 (2019).
Le-Niculescu, H. Convergent functional genomics of genome-wide association data for bipolar disorder: comprehensive identification of candidate genes, pathways and mechanisms. Am. J. Med. Genet. Part B Neuropsychiatr. Genet. 150B, 155 (2009).
Le-Niculescu, H. Identifying blood biomarkers for mood disorders using convergent functional genomics. Mol. Psychiatry 14, 156 (2009).
Kato, T. Comprehensive gene expression analysis in bipolar disorder. Can. J. Psychiatry 52, 763 (2007).
Roth, T. L., Zoladz, P. R., Sweatt, J. D. & Diamond, D. M. Epigenetic modification of hippocampal Bdnf DNA in adult rats in an animal model of post-traumatic stress disorder. J. Psychiatr. Res. 45, 919–926 (2011).
Andrews, J. A. Cells biomarkers, and post-traumatic stress disorder: evidence for peripheral involvement in a central disease. J. Neurochem. 120, 26 (2012).
Colin, L. & Van Lint, C. Molecular control of HIV-1 postintegration latency: implications for the development of new therapeutic strategies. Retrovirology 6, 111 (2009).
Murata, Y. et al. Evaluation of the usefulness of saliva for DNA methylation analysis in cohort studies. Neuropsychopharmacol. Rep. 39, 301–305 (2019).
Smith, A. K. et al. DNA extracted from saliva for methylation studies of psychiatric traits: evidence tissue specificity and relatedness to brain. Am. J. Med. Genet. Part B Neuropsychiatr. Genet. 168, 36–44 (2015).
Braun, P. R. et al. Genome-wide DNA methylation comparison between live human brain and peripheral tissues within individuals. Transl. Psychiatry 9, 47 (2019).
Bernstein, B. E. et al. The NIH roadmap epigenomics mapping consortium. Nat. Biotechnol. 28, 1045–1048 (2010).
Kundaje, A. et al. Integrative analysis of 111 reference human epigenomes. Nature 518, 317–330 (2015).
Jaffe, A. E. & Irizarry, R. A. Accounting for cellular heterogeneity is critical in epigenome-wide association studies. Genome Biol. 15, R31 (2014).
Reinius, L. E. et al. Differential DNA methylation in purified human blood cells: implications for cell lineage and studies on disease susceptibility. PLoS ONE 7, e41361 (2012).
Houseman, E. et al. DNA methylation arrays as surrogate measures of cell mixture distribution. BMC Bioinformatics 13, 1–16 (2012).
Morrison, F. G., Miller, M. W., Logue, M. W., Assef, M. & Wolf, E. J. DNA methylation correlates of PTSD: Recent findings and technical challenges. Prog. Neuro Psychopharmacol. Biol. Psychiatry 90, 223–234 (2019).
Sugden, K. et al. Patterns of reliability: assessing the reproducibility and integrity of DNA methylation measurement. Patterns https://www.sciencedirect.com/science/article/pii/S2666389920300143 (2020).
Dugué, P.-A. et al. Reliability of DNA methylation measures from dried blood spots and mononuclear cells using the HumanMethylation450k BeadArray. Sci. Rep. 6, 30317 (2016).
Logue, M. W. et al. The correlation of methylation levels measured using Illumina 450K and EPIC BeadChips in blood samples. Epigenomics 9, 1363–1371 (2017).
Meng, H. et al. A statistical method for excluding non-variable CpG sites in high-throughput DNA methylation profiling. BMC Bioinformatics 11, 227 (2010).
Bock, C., Walter, J., Paulsen, M. & Lengauer, T. Inter-individual variation of DNA methylation and its implications for large-scale epigenome mapping. Nucleic Acids Res. 36, e55–e55 (2008).
Teschendorff, A. E. et al. A beta-mixture quantile normalization method for correcting probe design bias in Illumina Infinium 450 k DNA methylation data. Bioinformatics 29, 189–196 (2013).
Johnson, W. E., Li, C. & Rabinovic, A. Adjusting batch effects in microarray expression data using empirical Bayes methods. Biostatistics 8, 118–127 (2007).
Barfield, R. T. et al. Accounting for population stratification in DNA methylation studies. Genet. Epidemiol. 38, 231–241 (2014).
Radtke, K. M. et al. Epigenetic modifications of the glucocorticoid receptor gene are associated with the vulnerability to psychopathology in childhood maltreatment. Transl. Psychiatry 5, e571 (2015).
Romens, S. E., McDonald, J., Svaren, J. & Pollak, S. D. Associations between early life stress and gene methylation in children. Child Dev. 86, 303–309 (2015).
Van Der Knaap, L. et al. Glucocorticoid receptor gene (NR3C1) methylation following stressful events between birth and adolescence. The TRAILS study. Transl. Psychiatry 4, e381 (2014).
Barker, V., Walker, R., Evans, K. & Lawrie, S. Methylation of glucocorticoid receptor (NR3C1), BDNF and oxytocin receptor genes in association with childhood maltreatment in schizophrenia and schizoaffective disorder. Schizophr. Res. 216, 529–531 (2019).
Fiacco, S., Gardini, E. S., Mernone, L., Schick, L. & Ehlert, U. DNA methylation in healthy older adults with a history of childhood adversity—findings from the Women 40+ Healthy Aging Study. Front. Psychiatry 10, 777 (2019).
Shields, A. E. et al. Childhood abuse, promoter methylation of leukocyte NR3C1 and the potential modifying effect of emotional support. Epigenomics 8, 1507–1517 (2016).
Suderman, M. et al. Conserved epigenetic sensitivity to early life experience in the rat and human hippocampus. Proc. Natl Acad. Sci. USA 109(Suppl 2), 17266–17272 (2012).
Pape, J. C. et al. DNA methylation levels are associated with CRF1 receptor antagonist treatment outcome in women with post-traumatic stress disorder. Clin. Epigenetics 10, 136 (2018).
Wang, P. et al. HTR1A/1B DNA methylation may predict escitalopram treatment response in depressed Chinese Han patients. J. Affect Disord. 228, 222–228 (2018).
Perroud, N. et al. Methylation of serotonin receptor 3a in Adhd, borderline personality, and bipolar disorders: link with severity of the disorders and childhood maltreatment. Depress Anxiety 33, 45–55 (2016).
Checknita, D. et al. Monoamine oxidase A genotype and methylation moderate the association of maltreatment and aggressive behaviour. Behav. Brain Res. 382, 112476 https://www.sciencedirect.com/science/article/abs/pii/S0166432819312793 (2020).
Gouin, J. P. et al. Associations among oxytocin receptor gene (OXTR) DNA methylation in adulthood, exposure to early life adversity, and childhood trajectories of anxiousness. Sci. Rep. 7, 7446 (2017).
Kogan, S. M., Cho, J., Beach, S. R. H., Smith, A. K. & Nishitani, S. Oxytocin receptor gene methylation and substance use problems among young African American men. Drug Alcohol Depend. 192, 309–315 (2018).
Womersley, J. S. et al. Childhood emotional neglect and oxytocin receptor variants: association with limbic brain volumes. World J. Biol. Psychiatry https://doi.org/10.1080/15622975.2019.1584331 (2019).
Perroud, N. et al. Response to psychotherapy in borderline personality disorder and methylation status of the BDNF gene. Transl. Psychiatry 3, e207 (2013).
Thaler, L. et al. Methylation of BDNF in women with bulimic eating syndromes: associations with childhood abuse and borderline personality disorder. Prog. Neuropsychopharmacol. Biol. Psychiatry 54, 43–49 (2014).
Wang, P. et al. Association of DNA methylation in BDNF with escitalopram treatment response in depressed Chinese Han patients. Eur. J. Clin. Pharmacol. 74, 1011–1020 (2018).
Boks, M. P. et al. SKA2 methylation is involved in cortisol stress reactivity and predicts the development of post-traumatic stress disorder (PTSD) after Military deployment. Neuropsychopharmacology 41, 1350–1356 (2016).
He, Y., Vinkers, C. H., Houtepen, L. C., De Witte, L. D., Boks, M. P. Childhood adversity is associated with increased KITLG methylation in healthy individuals but not in bipolar disorder patients. Frontiers in psychiatry 9, 743 (2019).
Wrigglesworth, J., Ancelin, M. L., Ritchie, K. & Ryan, J. Association between DNA methylation of the KITLG gene and cortisol levels under. Stress 22, 162–168 (2019).
Berent, D. et al. Childhood adversities are not a predictors of SSTR4met in alcoholics. Transl. Neurosci. 8, 127–138 (2017).
Thomas, M. et al. Investigation of MORC1 DNA methylation as biomarker of early life stress and depressive symptoms. J. Psychiatr. Res. 120, 154–162 (2020).
Lapp, H. E., Ahmed, S., Moore, C. L. & Hunter, R. G. Toxic stress history and hypothalamic-pituitary-adrenal axis function in a social stress task: Genetic and epigenetic factors. Neurotoxicol. Teratol. 71, 41–49 (2019).
Misiak, B. et al. Lower LINE-1 methylation in first-episode schizophrenia patients with the history of childhood trauma. Epigenomics 7, 1275–1285 (2015).
Cecil, C. A. et al. Epigenetic signatures of childhood abuse and neglect: implications for psychiatric vulnerability. J. Psychiatr. Res. 83, 184–194 (2016).
Esposito, E. A. et al. Differential DNA methylation in peripheral blood mononuclear cells in adolescents exposed to significant early but not later childhood adversity. Dev. Psychopathol. 28, 1385–1399 (2016).
Kumsta, R. et al. Severe psychosocial deprivation in early childhood is associated with increased DNA methylation across a region spanning the transcription start site of CYP2E1. Transl. Psychiatry 6, e830 (2016).
Marini, S. et al. Adversity exposure during sensitive periods predicts accelerated epigenetic aging in children. Psychoneuroendocrinology 113, 104484 (2020).
Naumova, O. Y. et al. Differential patterns of whole-genome DNA methylation in institutionalized children and children raised by their biological parents. Dev. Psychopathol. 24, 143–155 (2012).
Papale, L. A., Seltzer, L. J., Madrid, A., Pollak, S. D. & Alisch, R. S. Differentially methylated genes in saliva are linked to childhood stress. Sci. Rep. 8, 10785 (2018).
Weder, N. et al. Child abuse, depression, and methylation in genes involved with stress, neural plasticity, and brain circuitry. J. Am. Acad. Child Adolesc. Psychiatry 53, 417–424 e415 (2014).
Yang, B. Z. et al. Child abuse and epigenetic mechanisms of disease risk. Am. J. Prev. Med. 44, 101–107 (2013).
Houtepen, L. C. et al. Genome-wide DNA methylation levels and altered cortisol stress reactivity following childhood trauma in humans. Nat. Commun. 7, 10967 (2016).
Khulan, B. et al. Epigenomic profiling of men exposed to early-life stress reveals DNA methylation differences in association with current mental state. Transl. Psychiatry 4, e448 (2014).
Labonte, B. et al. Genome-wide epigenetic regulation by early-life trauma. Arch. Gen. Psychiatry 69, 722–731 (2012).
Marinova, Z. et al. DNA methylation profiles of elderly individuals subjected to indentured childhood labor and trauma. BMC Med. Genet. 18, 21 (2017).
Marzi, S. J. et al. Analysis of DNA methylation in young people: limited evidence for an association between victimization stress and epigenetic variation in blood. Am. J. Psychiatry 175, 517–529 (2018).
Prados, J. et al. Borderline personality disorder and childhood maltreatment: a genome-wide methylation analysis. Genes Brain Behav. 14, 177–188 (2015).
Robakis, T. K. et al. Epigenetic signatures of attachment insecurity and childhood adversity provide evidence for role transition in the pathogenesis of perinatal depression. Transl. Psychiatry 10, 1–14 (2020).
Roberts, A. L. et al. Exposure to childhood abuse is associated with human sperm DNA methylation. Transl. Psychiatry 8, 194 (2018).
Smith, A. K. et al. Differential immune system DNA methylation and cytokine regulation in post-traumatic stress disorder. Am. J. Med. Genet. B Neuropsychiatr. Genet. 156B, 700–708 (2011).
Suderman, M. et al. Childhood abuse is associated with methylation of multiple loci in adult DNA. BMC Med. Genomics 7, 13 (2014).
Zannas, A. S. et al. Lifetime stress accelerates epigenetic aging in an urban, African American cohort: relevance of glucocorticoid signaling. Genome Biol. 16, 266 (2015).
Zhang, H., Wang, F., Kranzler, H. R., Zhao, H. & Gelernter, J. Profiling of childhood adversity-associated DNA methylation changes in alcoholic patients and healthy controls. PLoS ONE 8, e65648 (2013).
Mehta, T. et al. Childhood maltreatment is associated with distinct genomic and epigenetic profiles in posttraumatic stress disorder. Proceedings of the National Academy of Sciences 110, 8302–8307 (2013).
Acknowledgements
This work was supported by NIH grants R01 HD095837 (SHP), R01 MH101107 (ART), R01 HD086487 (ART), R01 DA044504 (LRS), R01 DA031188 (LRS), R24 ES028507 (LRS), R01 MH10537 (NRN), and R01 MH108641 (NRN). L.H. received support from T32 MH019927 and F32 HD100020. T.D. received support from R25 MH101076. The content is solely the responsibility of the authors and does not necessarily represent the official views of the National Institutes of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Parade, S.H., Huffhines, L., Daniels, T.E. et al. A systematic review of childhood maltreatment and DNA methylation: candidate gene and epigenome-wide approaches. Transl Psychiatry 11, 134 (2021). https://doi.org/10.1038/s41398-021-01207-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-021-01207-y
This article is cited by
-
Mother adversity and co-residence time impact mother–child similarity in genome-wide and gene-specific methylation profiles
Clinical Epigenetics (2024)
-
Effect of parental adverse childhood experiences on intergenerational DNA methylation signatures from peripheral blood mononuclear cells and buccal mucosa
Translational Psychiatry (2024)
-
Associations of altered leukocyte DDR1 promoter methylation and childhood trauma with bipolar disorder and suicidal behavior in euthymic patients
Molecular Psychiatry (2024)
-
Epigenetic analyses in forensic medicine: future and challenges
International Journal of Legal Medicine (2024)
-
Maternal adverse childhood experiences (ACEs) and DNA methylation of newborns in cord blood
Clinical Epigenetics (2023)