Abstract
Machine learning has been suggested as a means of identifying individuals at greatest risk for hospital readmission, including psychiatric readmission. We sought to compare the performance of predictive models that use interpretable representations derived via topic modeling to the performance of human experts and nonexperts. We examined all 5076 admissions to a general psychiatry inpatient unit between 2009 and 2016 using electronic health records. We developed multiple models to predict 180-day readmission for these admissions based on features derived from narrative discharge summaries, augmented by baseline sociodemographic and clinical features. We developed models using a training set comprising 70% of the cohort and evaluated on the remaining 30%. Baseline models using demographic features for prediction achieved an area under the curve (AUC) of 0.675 [95% CI 0.674–0.676] on an independent testing set, while language-based models also incorporating bag-of-words features, discharge summaries topics identified by Latent Dirichlet allocation (LDA), and prior psychiatric admissions achieved AUC of 0.726 [95% CI 0.725–0.727]. To characterize the difficulty of the task, we also compared the performance of these classifiers to both expert and nonexpert human raters, with and without feedback, on a subset of 75 test cases. These models outperformed humans on average, including predictions by experienced psychiatrists. Typical note tokens or topics associated with readmission risk were related to pregnancy/postpartum state, family relationships, and psychosis.
Similar content being viewed by others
Introduction
The ability of prediction tools based on machine learning (ML) to identify high-risk individuals among clinical populations has been embraced across health care1. In psychiatry, a number of reports suggest the ability of these approaches to improve on chance in predicting hospital readmission, suicide attempts, or mortality, for example2,3. In previous work, we demonstrated that incorporation of features based on topic modeling of narrative notes improved prediction among individuals with major depressive disorder. This approach has the important feature of yielding topics that may be more interpretable than individual words or codes4,5.
However, several important questions remain unanswered by prior work. First, most prior investigations focused on a single, homogeneous patient group. In real-world implementation, clinical populations are rarely so selected, which may pose challenges for natural language processing methods. In particular, how well could such models account for populations that are both medically and psychiatrically complex? Second, while prior work indicates improvement over chance, a more relevant question is performance compared to human experts: how well do clinicians perform on a given task, and can ML outperform human learning?
To address these concerns, we drew on 8 years of electronic health records from a large inpatient psychiatric unit. We applied standard ML approaches and then compared these approaches to expert and nonexpert clinical annotation, as a means of understanding whether ML models could equal or surpass human prediction.
Materials and methods
Study design and cohort generation
We utilized a retrospective cohort design, drawing from the electronic health records of a Boston academic medical center’s inpatient psychiatric unit. All consecutive admissions between January 1, 2009 and December 31, 2016 were included. Data were extracted and stored using i2b2 server software6; coded features extracted included age, sex, race/ethnicity, insurance type, as well as admission diagnoses, with ICD9 codes mapped to the HCUP/CCS Level 2 ontology7. The data mart also included all narrative discharge summaries. In total, we analyzed 5076 health records, 75 of which were randomly sampled from the test split for our human prediction study. The primary outcome of interest was all-cause hospital readmission within 180 days, recognizing that psychiatric illness contributes to adverse outcomes that may not be fully captured by psychiatric hospitalization.
The Partners HealthCare Human Research Committee approved the study protocol. As no participant contact was required in this study based on secondary use of data arising from routine clinical care, the committee waived the requirement for informed consent as detailed by 45 CFR 46.116.
Natural language processing of discharge summaries
We preprocessed the raw text of the records by lowercasing the words, removing the punctuation, and replacing all numbers with a placeholder “NUM” token. The text was split into tokens with the regex-based tokenizer of the nltk library v3.2.58, with whitespace characters serving as token delimiters. The resulting vocabulary consisted of 20878 tokens. Length distribution of the discharge summaries is summarized in [Supplement A Fig. 1]. After text preprocessing, we represented the input discharge narratives using several sets of features which are described below. No attempt was made to distinguish discharge summary sections, as note format varied substantially in structure.
TF-IDF features
Term frequency-inverse document frequency scoring9 allows for weighting every word of the given document by its importance relative to the entire corpus. We used the scikit-learn library v0.20.210 to convert every document to a distribution of scores over a pruned vocabulary. The pruning procedure included filtering out common English stopwords, setting the maximum document frequency to 90% of all documents and the minimum document frequency to 10, based on prior work with psychiatric discharge summaries and visual inspection of distributions. This filtering procedure aims to remove noise such as typographic errors in the narrative notes. As a result, the size of the initial vocabulary was reduced to 11062 unique tokens.
LDA features
Latent Dirichlet Allocation11 is an unsupervised generative statistical model that uses word co-occurrence to derive a representation for each input document. The derived representation takes the form of a distribution over a predefined number of topics, with each topic corresponding to a distribution over a set of words. In our experiments, we used the gensim library v3.7.012 that uses raw word counts as input to compute the topic distributions. We varied the number of topics from 25 to 100, selected a priori based on prior work with narrative clinical notes to balance overfitting risk without yielding overly sparse topics, and, based on the held-out validation dataset, found the optimal number to be 75 topics (see [Supplement B Tables 1 and 2]). Stability of Supplement B results was confirmed via multiple runs. The resulting 75-dimensional vectors (henceforth, LDA-75) were used as input to the classifier models. Sample derived topics are shown in Supplement C Table 3.
Prediction models
All models incorporated baseline sociodemographic and clinical features, defined as age at admission, gender, self-reported race (coded as White, African-American, Asian, unknown, and other), age-adjusted Charlson comorbidity index (ACCI)13,14, and insurance type (coded as public payer, public Medicaid, public Medicare, and private payer).
Out of the entire cohort, four patients had missing values for age and ACCI index. These missing values were imputed with the corresponding averages over the dataset and the resulting numeric features were further normalized to fall in the range between zero and one. Categorical features were encoded using the one-hot encoding scheme (i.e., presence/absence of a feature). This procedure resulted in a 13-dimensional demographic baseline feature vector per admission. Additionally, the number of prior psychiatric admissions (PrPA) within the past 12 months was used as a predictive feature.
Using the representation described above, we trained support vector machines with linear kernel (SVM) for different feature combinations, including demographics, TF-IDF, LDA-75, demographics+TF-IDF, demographics+LDA, demographics+TF-IDF + LDA, and demographics+TF-IDF + LDA + PrPA. We experimented with other simple ML models (including logistic regression (LR), gradient boosting (XGB), multilayer perceptrons (MLP)), but all models had very similar levels of performance.
In addition to these standard ML algorithms, in preliminary experiments, we also examined two recurrent neural networks: a standard long short-term memory (LSTM) recurrent neural network15 and a hierarchical LSTM with attention to model a sequence of sentences where each sentence is a sequence of words16. However, due to notably large input size of the tokenized documents, the selected LSTM-based architectures struggled to encode all the relevant information in single fixed-length vectors, leading to consistently poor performance even with optimization of hyperparameters.
Comparison with human prediction
We sought to understand the difficulty of the prediction task for human raters, including both clinical experts and individuals without domain-specific knowledge. To achieve this objective, we used two sets of human raters (n = 7 total). The first consisted of three psychiatrists with at least 5 years of clinical experience, including work on inpatient units and consult-liaison psychiatry. The second consisted of four graduate students without formal postgraduate training in psychiatry or psychology. Human raters were given access to the full discharge summary, as well as the structured demographics information, but were blinded to model features including prior psychiatric admissions (PrPA). Each rater began by scoring the same 25 records, randomly selected to include 9 readmissions and 16 non-readmissions within 180 days. Each record was scored in terms of likely readmission17 and the associated judgment confidence. In order to establish a baseline for prediction accuracy, no feedback was provided for these predictions. In the second phase (henceforth, learning phrase), participants scored a set of additional 50 randomly sampled records (with the same ratio of readmission to non-readmission), receiving feedback about the correct answer after each prediction. These additional 50 records were the same for all human raters, and were shown in the same order.
Results
Characteristics of the full cohort (n = 5076 psychiatric admissions), as well as the training (70%) and testing (30%) cohorts stratified by sex, age, ACCI score, and number of prior admissions, are presented in Table 1. The cohort included a broad range of admission diagnoses, with 44 unique CCS categories in total; the most common diagnosis was mood disorder (5.8.657). The full distribution of the top 10 most frequent admission CCS codes can be found in [Supplement C Fig. 2]. Out of the 5076 patients, 1168 were readmitted within 180 days (23%).
Machine learning
The training set was randomly split into 10 folds to perform cross-validation, which was carried out separately with each of the models (SVM, LR, XGB, and MLP) for the feature representations listed in Methods section above. Using random search over hyperparameter space, we picked the best model hyperparameters based on the average score over the folds. To ensure the reproducibility of the results, we repeated the cross-validation procedure 100 times resulting in 100 best hyperparameter configurations. We tested the resulting 100 fine-tuned models, and we report the average test AUC scores with the corresponding confidence intervals in Table 2.
To identify the most relevant features in ML models and facilitate interpretation, we extracted the top 20 weights (sorted by their magnitude) assigned by the best logistic regression model (with the AUC of .726). The histogram suggests that prior psychiatric admissions, single tokens (pregnancy-related terms; family relationships) as well as age-adjusted Charlson comorbidity index represent major predictors of readmission (Fig. 1). Topics provide additional predictive support, most notably capturing family relationships (Topic 13) and psychosis and antipsychotics (Topic 44).
Human learning
Since human raters did not have access to the information about prior psychiatric admissions, we focused on the SVM results, which yielded the best performance for the demographics+TF-IDF + LDA-75 feature set. Table 3 compares prediction results for 180-day readmission for the SVM models and human raters. Since AUC requires ranked scores (and therefore cannot be reported for human judgments), we report F1, accuracy, and balanced accuracy. Also, for the purposes of comparison, the figures reported for ML models come from a single run, rather than averaged over multiple runs. We also report the baseline performance for two arbitrarily-selected predictors to capture the recurrent nature of psychiatric illness: (a) always predicting that the patient will be readmitted, and (b) always predicting that the patient will be readmitted if there are 5 or more prior psychiatric admissions in the past 12 months.
Overall, all of the ML models, including very basic ones, outperformed the average human performance (balanced accuracy of 0.484 for expert and 0.635 for nonexpert human judges, respectively). Note that the performance of the ML models on the human-annotated learning-phase data (n = 50) is higher than on the full test set (n = 1523), with F1 scores of 0.52–0.63 and 0.41–0.45, respectively. Also, for this smaller subset, models based on the minimal baseline demographic features outperformed the models using NLP-derived features. This suggests that the human-annotated subset was not entirely representative of the full test set, which is in part due to the small sample size.
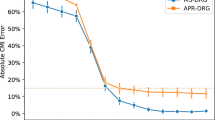
We also looked at human performance over time, both with and without feedback on whether the generated predictions were correct. Figure 2 illustrates the average performance for expert (MD) and nonexpert (non-MD) annotators; each timestep of the x-axis indicates a set of 15 predictions (i.e., 15 hospital discharges) grouped together using a sliding window with a 5-note stride. The first three timesteps (covering notes 0–25) indicate human performance before receiving any feedback. Starting from the 25–40 window, we can see performance over time.
As all raters were shown the same notes in the same order, the figure also enables a comparison of how well each group was able to improve their performance over time. In particular, the nonclinicians were able to improve their performance substantially upon receiving feedback, with a ~10% increase in absolute accuracy between their initial baseline performance and their final set of predictions after training. Performance of expert annotators (i.e., psychiatrists), on the other hand, did not improve notably over time.
Discussion
In this investigation of more than 5000 admissions to a psychiatric inpatient unit, we found that prediction models incorporating topics derived from Latent Dirichlet Allocation in addition to simpler bag-of-words features performed comparably to, or better than, the models relying solely on bag-of-words or on coded data. These findings are consistent with our own and other prior work5; the explanatory power of a single engineered feature, namely prior admissions, is also unsurprising as a marker of illness chronicity and severity. The topics themselves were notable for coherence in capturing predominantly comorbidities (e.g., orthopedic injury), psychosocial features (e.g., family relationships, homelessness), or symptoms (e.g., psychosis, substance abuse).
Conversely, the performance of the models we present is markedly poorer than that of many models predicting medical readmission1. We would underscore that this likely reflects the challenging nature of the task; indeed, clinical features strongly predictive of readmission remain unclear, and there are no validated biomarkers or markers of disease progression.
Although the models we developed improve substantially upon chance, we also sought to understand their ability to improve upon prediction by clinicians and nonclinicians. This task allows us to ground these models using a more understandable frame of reference. We observed that ML models outperformed our human raters, especially domain experts, suggesting both the difficulty of the task and that the models are not overlooking obvious features that would improve prediction. The success of the models suggests that there is some additional signal in the notes not captured fully by human raters, potentially identified by aggregating many small indications. Still, the modest performance overall suggests that the upper bound of performance is probably far below “perfect,” reflecting an element of stochasticity of readmission. Of course, neither human nor machine models have access to information not contained in notes that could be highly relevant to readmission risk, such as undocumented aspects of psychosocial circumstances or comorbidities not addressed during an acute hospitalization.
Perhaps more surprisingly, nonclinician raters outperformed clinicians, which may reflect that this task is distinct from standard clinical practice. Further, as the nonclinician raters were better able to improve their performance with feedback, it may be the case that nonexperts are more easily able to conform to new tasks because they have fewer incorrect priors, whereas experts are harder to shift from their existing frameworks.
This latter set of observations is consistent with decades of prior evidence that clinician predictions, when compared to real-world outcomes, often do not substantially exceed chance. In psychiatry, a seminal paper by Pokorny found that even incorporating a range of rating scales did not yield meaningful prediction18.
Our results also hint at an opportunity to improve prediction in clinical settings by presenting observed outcomes to clinicians in an iterative process. An important question to consider will be whether ML models can be applied to accelerate or enhance this learning process, by highlighting key topics or terms to consider, for example.
We note several important limitations in considering our results. First, they reflect a single inpatient unit, albeit a large and heterogeneous one. The extent to which results generalize to other health systems remains to be established; it is possible that institution-specific models will be required given the heterogeneity of documentation across institutions. Institutional restrictions on accessing large clinical corpora continue to inhibit work in this area and underscore the need for more cross-institution collaboration.
Second, our prediction task likely underestimates the skill of clinicians in practice, who are able to draw on clinical impressions of patients that may not be fully captured in narrative notes. This distinction likely explains the inability to distinguish trained clinicians from graduate students. Moreover, as we use notes generated at time of hospital discharge, we expect that clinicians do not fully document the extent of concern for readmission, insofar as individuals thought to be at high risk would not be candidates for discharge in the first place.
Still, taken together, our results suggest that ML approaches incorporating topic models consistently exceed clinicians’ ability to predict hospital readmission, while retaining inspectability. While such models are far less performant than some recent efforts in more homogenous datasets, this likely reflects the challenging nature of the task itself. As better approaches to staging or characterizing progression emerge, including biomarkers or clinical metrics, we anticipate that the models we describe will provide a valuable baseline to be improved upon in subsequent work.
References
Rajkomar, A. et al. Scalable and accurate deep learning with electronic health records. npj Digital Med. 1, 18 (2018).
McCoy, T. H., Castro, V. M., Roberson, A. M., Snapper, L. A. & Perlis, R. H. Improving prediction of suicide and accidental death after discharge from general hospitals with natural language processing. JAMA Psychiatry 73, 1064 (2016).
McCoy, T. H. et al. Sentiment measured in hospital discharge notes is associated with readmission and mortality risk: an electronic health record study. PLoS ONE 10, e0136341 (2015).
Moss, J. et al. Predictors of readmission to a psychiatry inpatient unit. Compr. Psychiatry 55, 426–430 (2014).
Rumshisky, A. et al. Predicting early psychiatric readmission with natural language processing of narrative discharge summaries. Transl. Psychiatry 6, e921 (2016).
Murphy, S. N. et al. Serving the enterprise and beyond with informatics for integrating biology and the bedside (i2b2). J. Am. Med. Inf. Assoc. 17, 124–130 (2010).
Healthcare Cost and Utilization Project (HCUP). Clinical Classifications Software (CCS) for ICD-9-CM [Internet], https://www.hcup-us.ahrq.gov/toolssoftware/ccs/ccs.jsp (2017).
Bird S., Klein E. & Loper E. Natural Language Processing with Python. (O’Reilly Media., 2009).
Salton, G., Wong, A. & Yang, C. S. A vector space model for automatic indexing. Commun. ACM 18, 613–620 (1975).
Pedregosa, F. et al. Scikit-learn: machine learning in python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Blei, D. M., Ng, A. Y. & Jordan, M. I. Latent dirichlet allocation. J. Mach. Learn. Res. 3, 993–1022 (2003).
Rehurek, R. & Sojka, P. Software framework for topic modelling with large corpora. in Conference Proceedings of the LREC 2010 Workshop on New Challenges for NLP Frameworks. (University of Malta, Valletta,2010).
Charlson, M., Szatrowski, T. P., Peterson, J. & Gold, J. Validation of a combined comorbidity index. J. Clin. Epidemiol. 47, 1245–1251 (1994).
Charlson, M. E., Pompei, P., Ales, K. L. & MacKenzie, C. R. A new method of classifying prognostic comorbidity in longitudinal studies: development and validation. J. Chronic Dis. 40, 373–383 (1987).
Gers, F. A. & Schmidhuber, J. Recurrent nets that time and count. in Proc. IEEE-INNS-ENNS International Joint Conference on Neural Networks. IJCNN 2000. (Neural Computing: New Challenges and Perspectives for the New Millennium. Como, 2000).
Yang, Z. et al. Hierarchical Attention Networks for Document Classification. in Proc. 2016 Conference of the North American Chapter of the Association for Computational Linguistics: Human Language Technologies (2016).
Downar, J., Goldman, R., Pinto, R., Englesakis, M. & Adhikari, N. K. J. The ‘surprise question’ for predicting death in seriously ill patients: a systematic review and meta-analysis. CMAJ 189, E484–E493 (2017).
Pokorny, A. D. Prediction of suicide in psychiatric patients. Report of a prospective study. Arch. Gen. Psychiatry 40, 249–257 (1983).
Acknowledgements
This material is based upon work supported by NIMH 1R01MH106577 and the National Science Foundation Graduate Research Fellowship Program under Grant No. 1122374. Any opinions, findings, and conclusions or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the National Science Foundation.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
R.H.P. has received honoraria or consulting fees from Burrage Capital, Genomind, RID Ventures, and Takeda; he holds equity in Psy Therapeutics and Outermost Therapeutics.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Boag, W., Kovaleva, O., McCoy, T.H. et al. Hard for humans, hard for machines: predicting readmission after psychiatric hospitalization using narrative notes. Transl Psychiatry 11, 32 (2021). https://doi.org/10.1038/s41398-020-01104-w
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-020-01104-w
This article is cited by
-
Predicting which patients with cancer will see a psychiatrist or counsellor from their initial oncology consultation document using natural language processing
Communications Medicine (2024)
-
Natural language processing applied to mental illness detection: a narrative review
npj Digital Medicine (2022)