Abstract
No diagnostic biomarkers are available for obsessive-compulsive disorder (OCD). Here, we aimed to identify magnetic resonance imaging (MRI) biomarkers for OCD, using 46 data sets with 2304 OCD patients and 2068 healthy controls from the ENIGMA consortium. We performed machine learning analysis of regional measures of cortical thickness, surface area and subcortical volume and tested classification performance using cross-validation. Classification performance for OCD vs. controls using the complete sample with different classifiers and cross-validation strategies was poor. When models were validated on data from other sites, model performance did not exceed chance-level. In contrast, fair classification performance was achieved when patients were grouped according to their medication status. These results indicate that medication use is associated with substantial differences in brain anatomy that are widely distributed, and indicate that clinical heterogeneity contributes to the poor performance of structural MRI as a disease marker.
Similar content being viewed by others
Introduction
Obsessive-compulsive disorder (OCD) is a severe and debilitating condition that occurs in 2–3% of the population1. It is characterized by recurrent, intrusive, irrational and distressing thoughts (obsessions) and repetitive behaviors or mental acts (compulsions)2. So far, no biomarkers that aid differential diagnosis are available, and diagnosis relies entirely on recognition of characteristic symptoms assessed by clinical interview3. Many neuroimaging studies have provided evidence for abnormalities in cortico-striato-thalamo-cortical (CSTC) circuits, as well as distributed changes in limbic, parietal and cerebellar regions4,5. These findings have recently been confirmed by different meta-analyses and mega-analyses of neuroimaging studies, based on results that were reported in the literature or by using original data within different consortia6,7,8,9,10. However, inference has been at the group-level, and the small effect sizes reported preclude clinical application.
Analytic tools such as multivariate pattern analysis (MVPA) enable inference at the individual-level, which may result in better discrimination3,11. MVPA techniques can be used to develop predictive models that extract common patterns from neuroimaging data to classify individuals based on their diagnosis. A major advantage of MVPA compared to traditional methods of analysis is its ability to use inter-regional correlations to detect subtle and spatially distributed effects4. Therefore, MVPA seems particularly well suited for neuroimaging analyses in OCD, as abnormalities are typically distributed across the brain12,13. Previous MVPA studies have been able to distinguish OCD patients from controls with accuracies ranging from 66–100%14. Although these results are promising, sample sizes have typically been small, limiting model performance optimization and leading to high variance in estimated accuracy, which may result in reporting optimistic or pessimistic classification rates15. In addition, most studies have been performed using data from one research center to minimize technical (e.g., scanner hardware, protocols, and diagnostic assessment) and clinical (e.g., age, medication status, disease chronicity, and severity) heterogeneity. It is therefore not clear whether the MVPA results obtained from these monocenter studies generalize well to other centers, which would be required for clinical application16,17,18. Interestingly, whereas classification accuracies of monocenter studies only tend to increase with larger samples19,20, accuracies for multicenter studies in other psychiatric disorders such as schizophrenia and autism tend to be lower with increasing sample size12,15,16,17. This paradoxical effect of lower classification accuracy with larger samples has been attributed to larger sample heterogeneity21, which inevitably increases when combining data from different centers. Here, we used data from the Enhancing Neuro-Imaging and Genetics through Meta-Analysis (ENIGMA) OCD consortium, including 4372 participants recruited at 36 research institutes around the world, with a full range of technical and clinical heterogeneity. We assessed the ability of MVPA to distinguish OCD patients from healthy controls using structural neuroimaging data at the individual subject level. We investigated machine learning classification performance in both single-site and multi-site samples using different validation strategies to assess generalizability. Furthermore, the large sample size enables investigation of the influence of clinical heterogeneity by stratification and subsampling, in order to assess the influence of clinical variability on classification accuracy.
Materials and methods
Study population
The ENIGMA-OCD working group includes 46 data sets from 36 international research institutes, with neuroimaging and clinical data from adult (≥18 years) and pediatric (<18 years) samples. In total, we analyzed data from 4372 participants, including 2304 OCD patients (n = 1801 adult, n = 503 child) and 2068 healthy controls (HC; i.e., free of psychopathology; n = 1629 adult, n = 439 child), with 38 of 46 datasets identical to those described in previous mega-analyses by this working group6,7,22. All participating sites obtained permission from their local institutional review boards or ethics committees to provide anonymized data for analysis, and all study participants provided written informed consent. Demographic and clinical characteristics of each site are detailed in supplementary Table S1. A complete overview of instruments used to obtain diagnosis and clinical information can be found elsewhere (Data Supplement 1, Supplementary Section S1)7. Diagnosis was determined in accordance with DSM2; MINI and SCID were used for adult samples and K-SADS, MINI-KIDS and ADIS were used for pediatric samples23,24,25,26,27.
MRI processing
Structural T1-weighted brain MRI scans were acquired and processed locally at each site. Image acquisition parameters are listed elsewhere7. Parcellations were performed using FreeSurfer software version 5.3 (http://surfer.nmr.mgh.harvard.edu), following standardized ENIGMA protocols to harmonize analyses and quality control procedures across multiple sites (see http://enigma.usc.edu/protocols/imaging-protocols/). Mean values of parcellations of 34 cortical (Desikan-Killiany atlas-based28) and 7 subcortical gray matter structures per hemisphere, lateral ventricle volumes, two whole-hemisphere measures and total intracranial volume were extracted, visually inspected and statistically evaluated for outliers (quality assurance is reported elsewhere7). Brain regions (features) used for classification included cortical thickness (CT), surface area (SA) and subcortical volumes of ROIs, two lateral ventricular and intra-cranial volumes (ICV), and two whole-hemisphere measures for SA and CT.
Multivariate classification and validation
Participants with >10% missing entries were excluded (n = 276), and median imputation was used for missing MRI data on the training set. Continuous features were centered around median zero and scaled according to their interquartile range. FreeSurfer variables were combined with covariates age, sex, and site by concatenating individual feature vectors. Categorical covariates were one-hot encoded prior to classification. All analyses were performed separately for pediatric and adult patients, and both groups combined. Common MVPA classifiers were applied: support vector machine (SVM) with linear and non-linear (radial-basis-function (RBF)) kernels, logistic regression (LR) with L1 and L2 regularization, Gaussian processes classification (GPC) with a linear kernel, and two decision-tree based ensemble methods, namely the random forest classifier (RFC) and the XGBoost (XGB) algorithm29,30,31,32. A neural network was also implemented (fully connected; 3 hidden layers with 60, 40, and 20 nodes respectively). SVM and LR classifiers were combined with and without automatic dimensionality reduction via principal component analysis (PCA), using the minimal number of components explaining 90% of the variance. Hyper-parameters for SVM (linear and non-linear), LR and XGB were optimized using nested cross-validation; RFC and GPC were tuned following recommendations. Details on handling missing data, model implementation and hyper-parametrization can be found in Supplementary Methods (and supplementary Fig. S1 for assessment of different imputation strategies). The primary performance metric was the area under the receiver operator curve (AUC) and reported metrics are averaged across CV folds33. Balanced accuracy, sensitivity and specificity are reported in the supplement.
Multi-site classification of OCD patients versus HC was assessed using different cross-validation (CV) approaches. First, we assessed multi-site classification using 10-fold site-stratified CV to obtain maximally homogeneous train-test splits, with approximately the same number of subjects in each fold and the same proportion of samples coming from each site (also referred to as ‘internal validation’). Next, we addressed leave-one-site-out (LOSO) CV, in which all but one site were used to train the models while the left out site was used to assess model performance (external validation). This procedure is then repeated so that each site is used once as a test set. LOSO-CV may result in large between-sample heterogeneity of training and test sets, resulting in lower classification performance34. Because LOSO-CV has different fold sizes, we additionally performed site-stratified CV with varying fold sizes, in which the number of CV folds and respective test-fold sizes are set to match those of LOSO-CV. This was done to evaluate whether differences between site-stratified and LOSO-CV performance were due to differences in heterogeneity or test-fold size variance. Finally, we also performed single-site predictions using repeated 5-fold CV (with 10 repeats) to assess classification performance for individual sites with reduced heterogeneity. For a schematic representation of LOSO and site-stratified CV see supplementary Fig. S2. Statistical significance of model performance and 95% confidence intervals were assessed through the obtained AUC scores using the Mann–Whitney-U statistic for non-parametric testing (see Supplement for details)33,35,36. The Bonferroni-corrected level of significance was set at alpha = 0.05 for the number of classifiers and comparisons (3 CV types × 10 classifiers for multi-site classifications; alpha = 0.05/30). As previous work from ENIGMA-OCD has identified distinct alterations in pediatric and adult patients, we performed all multi-site classification analyses for pediatric (<=18 years of age) and adult (>18 years of age) data separately, as well as both age groups combined6,7.
Clinical variables and sensitivity analysis
To explore the effects of clinical heterogeneity on classification performance, we selected subgroups with particular demographic and clinical characteristics: medication use, OCD severity, age of onset (AO) and duration of illness. Classifications performed were HC vs. low (YBOCS <= 24; mild-moderate37) and high severity (YBOCS > 24; moderate-severe) OCD; HC vs. early (<18 yrs) and late AO (>= 18 yrs) OCD; HC vs. short (<=7 yrs) and long duration (>7 yrs) OCD; and HC vs. unmedicated and medicated OCD. For disease duration and severity, median splits were used to define groups; the 18 year threshold for AO was chosen in line with prior ENIGMA-OCD mega-analyses6,7. Finally, as particular clinical variables can co-occur, we performed a post-hoc sensitivity analysis to investigate the effects of potential clinical covariance for results with AUC ≥ 0.8. First, correlations between all clinical features were computed using point-biserial correlations between dichotomous and continuous variables, phi correlation for dichotomous variables and Pearson correlation for continuous variables. Only those clinical features that were significantly correlated (Bonferroni-corrected) were investigated further by rerunning previously described classifications, but now using samples further split according to their correlated features (e.g., HC vs. unmedicated, short duration OCD patients; etc.). The use of more homogenous subsamples is expected to improve classification performance, while reducing sample size itself is expected to decrease performance. If classifications are relatively unaffected by further splitting, the correlated clinical variable did not have a large influence on the classification results.
Feature importance
To assess which brain regions and clinical variables contributed most to classification we used feature importance extracted from RFC combined with a permutation testing framework (see Supplementary Methods)38.
Results
Multi-site classification
Three different CV approaches were used to assess the influence of sample heterogeneity. Results using various classification algorithms are summarized in Fig. 1. Classification performance (AUC) using site-stratified CV (with training on combined samples and equal fold sizes) ranged between 0.57 (95% confidence intervals (CI) = 0.51–0.63; pcorrected = 0.19) and 0.62 (95% CI = 0.56–0.67; pcorrected < 0.001) across different classifiers. All models had statistically significant performance after multiple comparison corrections except for PCA + LR, PCA + SVM and NN classifiers. LOSO-CV led to lower classification performance; 0.51 (95% CI = 0.4–0.62; pcorrected = 1) to 0.54 (95% CI = 0.42–0.65; pcorrected = 1) AUC with relatively high variance across folds (SD = 0.07–0.11) and no classifiers surviving multiple comparison corrections. AUC values obtained through site-stratified CV with varying fold sizes were similar to site-stratified CV results with equal fold sizes, ranging between 0.56 (95% CI = 0.45–0.67; pcorrected > 0.99) and 0.62 (95% CI = (0.51–0.73); pcorrected = 0.55). However, variance across CV-folds was higher and comparable to that from LOSO-CV (SD; site-stratified fixed: 0.02–0.04; site-stratified variable: 0.05–0.08; LOSO: 0.07–0.11). A complete overview of classification results is provided in supplementary Table S2. Multi-site classification with site-stratified CV (with equal fold sizes), performed separately on pediatric and adult samples yielded similar results, ranging from 0.55 (95% CI = 0.43–0.67; pcorrected = 1) to 0.62 (95% CI = 0.51–0.74; pcorrected = 0.71) and 0.56 (95% CI = 0.5–0.62; pcorrected = 0.69) to 0.61 (95% CI = 0.55–0.67; pcorrected = 0.008) AUC, respectively (see supplementary Tables S3-4). As site-stratified CV with equal fold-sizes resulted in the best performances, we used this strategy for further evaluation of intra-site performance and the influence of clinical variables. RFC classification performance is reported here by default, as differences between classifiers were minimal and this model was also used to extract feature importance.
Boxplots summarize AUC scores obtained across CV-folds; dashed line represents chance-level performance and asterisks indicate scores significantly different from chance (Mann–Whitney-U statistic; p < 0.05 Bonferroni corrected (10 classifiers × 3 CV types), see Supplement for details). SVM Support Vector Machine, PCA Principal Component Analysis, RBF Radial Basis Function, LR Logistic Regression, GPC Gaussian Processes Classification, RFC Random Forest Classifier, XGB XGBoost, NN Neural Network.
Single-site classification
Single-site classification performance with 10-fold CV varied greatly, with AUCs ranging between 0.30–0.89 across different sites and classifiers (see supplementary Table S5). Figure 2 summarizes RFC performances for each individual site. We assessed the correlation between the number of participants in each site and its obtained classification performance (AUC averaged over CV folds), which was significant (rS = 0.37, p = 0.014). In addition, we investigated the relationship between single-site classification performance and the following clinical variables of interest: mean and standard deviation of AO, duration, severity and the proportion of medicated patients and its standard deviation. None of these clinical variables showed a significant correlation with classification performance.
Clinical variables and sensitivity analysis
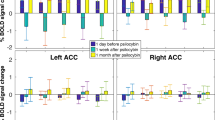
To assess the influence of different clinical variables on classification performance, we repeated the analysis for specific subgroups split according to medication use, AO, disease duration, and severity. A complete overview is provided in supplementary Tables S6(a–d), and results using RFC on combined data with age, sex and site as covariates are reported below. Medicated OCD vs. HC classification resulted in 0.69 AUC (95% CI = 0.63–0.75; pcorrected < 0.001), unmedicated OCD vs. HC in 0.60 (95% CI = 0.54–0.67; pcorrected = 0.03), and medicated vs. unmedicated OCD in 0.78 (95% CI = 0.72–0.85; pcorrected < 0.001) (see Fig. 3). XGB performance was notably higher for medicated vs. unmedicated OCD classification with an AUC of 0.86 (95% CI = 0.78–0.9; pcorrected < 0.001). Early AO OCD vs. HC classification resulted in 0.68 AUC (95% CI = 0.62–0.75; pcorrected < 0.001), late AO OCD vs. HC in 0.73 (95% CI = 0.67–0.79; pcorrected < 0.001), and early vs. late AO in 0.81 (95% CI = 0.74–0.88; pcorrected < 0.001). As no late AO patients were present in pediatric samples, classifications were re-run on adult samples only, resulting in 0.65 AUC (95% CI = 0.57–0.72;pcorrected = 0.01) for early AO vs. HC, 0.70 (95% CI = 0.63–0.76; pcorrected < 0.001) for late AO vs. HC, and 0.73 (95% CI = 0.64–0.82;pcorrected < 0.001) for early vs. late AO. Classification of short disease duration OCD vs. HC resulted in 0.68 AUC (95% CI = 0.61–0.75;pcorrected < 0.001), long disease duration vs. HC in 0.71 (95% CI = 0.65–0.78; pcorrected < 0.001), and short vs. long duration in 0.78 (95% CI = 0.7–0.85; pcorrected < 0.001). Finally, low severity OCD vs. HC classification resulted in 0.60 AUC (95% CI = 0.53–0.67; pcorrected = 0.15), high severity OCD vs. HC in 0.61 (95% CI = 0.54–0.68; pcorrected = 0.04), and low vs. high severity OCD in 0.58 (95% CI = 0.49–0.66; pcorrected = 1).
Only RFC classifier performance for combined (pediatric and adult) data is shown here; Boxplots summarize AUC scores obtained across CV-folds; dashed line represents chance-level performance and asterisks indicate scores significantly different from chance (Mann–Whitney-U statistic; p < 0.05 Bonferroni corrected (10 classifiers × 3 CV types), see Supplement for details). unmed unmedicated, med medicated.
Correlation analysis between medication status and other clinical variables only showed a significant association with disease duration (r = −0.094; pcorrected < 10−05; Bonferroni-corrected). We therefore performed additional medication status classifications after splitting patients for disease duration (e.g., HC vs. medicated + short duration OCD; HC vs. unmedicated + short duration OCD, etc.). Classifications with or without splitting patients for disease duration were comparable (see supplementary Tables S7(a–c) for full overview).
Influence of covariates
As seen in supplementary Table S1, several sites included only medicated patients whereas other sites only included non-medicated patients. To assess whether the high performance reported above for classifying medication status could be explained by site-differences in the covariates (e.g., site ID) rather than neuroimaging data, we performed the following control experiments. We reran both medication and main diagnosis classifications without using covariates (using only brain data), using covariates only, and by using neuroimaging data after removing the effect of the covariates using multiple linear regression. To maintain independence between the training and test sets, regression coefficients were estimated on the training data and applied to the test data.
Results for the main classifications between OCD patients and HC using neuroimaging data only resulted in AUC of 0.61 (95% CI = 0.55–0.66; pcorrected = 0.001), while use of covariates only resulted in AUC of 0.58 (95% CI = 0.52–0.64; pcorrected = 0.04), and using neuroimaging data following correction for covariates resulted in AUC of 0.58 (95% CI = 0.52–0.64; pcorrected = 0.02). These results show that the covariates contain sufficient information to distinguish patients from HC, but the results were otherwise comparable to the AUCs reported above for classifications using the covariates as features.
Classification for medicated patients vs. HC using neuroimaging data only resulted in AUC of 0.66 (95% CI = 0.60–0.73; pcorrected < 0.001) and AUC of 0.70 (95% CI = 0.64–0.76; pcorrected < 0.001) for SVM-RBF. Using covariates only resulted in AUC of 0.71 (95% CI = 0.65–0.77; pcorrected < 0.001), and neuroimaging data following correction for covariates resulted in AUC of 0.59 (95% CI = 0.52–0.66; pcorrected = 0.053), and higher AUC of 0.62 (95% CI = 0.55–0.69; pcorrected = 0.007) for XGB. Next, classification of unmedicated patients vs. HC using neuroimaging data only resulted in an AUC of 0.58 (95% CI = 0.51–0.68; pcorrected = 0.12) with significant classifier performance for PCA + SVM-RBF with AUC of 0.61 (95% CI = 0.55–0.68; pcorrected = 0.007). Using covariates only resulted in AUC of 0.64 (95% CI = 0.58–0.73; pcorrected < 0.001), and using neuroimaging data following correction for covariates resulted in AUC of 0.52 (95% CI = 0.45–0.59; pcorrected = 1). Finally, classification of medicated vs. unmedicated patients using neuroimaging data only resulted in AUC of 0.74 (95% CI = 0.67–0.81; pcorrected < 0.001), using covariates only in AUC of 0.84 (95% CI = 0.78–0.92; pcorrected < 0.001), and using neuroimaging data after correction for covariates resulted in AUC of 0.59 (95% CI = 0.50–0.67; pcorrected = 0.2). A full overview of these control experiment results can be found in supplementary Tables S8–9. These results show that the covariates contained sufficient information to classify these subgroups, and especially to distinguish between medicated vs. unmedicated patients, as subgroups were already partially defined by site. Nevertheless, the classifications using neuroimaging data only were comparable to the classifications reported above when covariates were added as features. However, correcting the neuroimaging data for covariates lowered the performance substantially compared to adding the covariates as features, suggesting that neuroimaging data were partially related to the covariates (e.g., different scanners at different sites).
Feature importance
We investigated which brain regions (features) contributed most to OCD vs. HC classifications for site-stratified CV only, using the feature importance values from RFC and permutation testing, and as we were interested in brain regions rather than the influence of the covariates, we focused on classifications using brain data only following multiple linear regression of confounding factors. No features were selected consistently (survived false discovery rate (FDR) correction in >50% CV-folds) for the main analyses (OCD patients vs. HC classification) in either pediatric, adult or combined samples. However, for medicated vs. unmedicated OCD classification in combined samples, 24 significant and consistently selected features were found. In addition, 12 features were found for early vs. late AO patients classifications in combined samples. A complete overview of these findings (including features importance for medication classification in adult samples) can be found in supplementary Tables S10–11.
Discussion
We found that MVPA of parcellated structural neuroimaging data is unable to provide accurate distinction between OCD cases and HC. Classification of the complete sample using site-stratified CV ranged between an AUC of 0.57 and 0.61, which is not sufficient for clinical application. Differences in performance between classifiers were minimal. Similar results were obtained for classifications performed separately on pediatric and adult samples. When validated on completely new data from other sites using LOSO-CV, model performance hardly exceeded chance-level (0.5 AUC).
Our findings highlight the impact of validation schemes on classification performance and suggest poor discrimination between OCD patients and HC when combining data from multiple sites. In contrast, discrimination between subgroups of patients based on medication status enabled fair individual subject classification. However, our control experiments indicated that non-brain covariates such as age, sex and site can heavily affect classification performance, dependent on the relation between the structural neuroimaging data and those covariates. Yet, even after removal of the covariate effects, the results still indicated that medication use is associated with substantial differences in brain anatomy that are widely distributed, whereas gross gray matter anatomy of patients with OCD was comparable to that of healthy controls. At the same time, this also suggests that clinical heterogeneity contributes to the poor performance of structural MRI as a disease biomarker.
Few diagnostic classifiers have been applied to OCD across multiple scanners and sites. Prior studies using structural MRI data to classify OCD using single-site samples yielded accuracies ranging from 0.72 to 0.9314. The wide range of performances observed in our individual site classification is in agreement with the published literature. Such a wide range may in part be explained by sample size, as larger samples tended to have higher AUC values16,19,34. However, this relationship does not necessarily hold true for large-scale multicenter studies, due to heterogeneity that arises from pooling samples with different scanning parameters, processing pipelines, inclusion criteria, demographic and clinical characteristics14,21. All these factors can impact the data and obscure a pattern of abnormalities shared by all patients. Monocenter studies that minimize heterogeneity may therefore yield higher classification performances, but limit the generalizability to new, unseen data and its use in clinical practice16,17. Thus, whereas small monocenter studies focus on answering a specific question about their patient population, large multicenter studies assume that a fundamental pattern of the disorder of interest can be detected despite the presence of heterogeneity, and both are geared toward answering complementary questions about a particular disorder18,39. Our LOSO-CV results demonstrated that structural MRI features do not provide a biomarker that enables generalization to new sites.
Multi-site classification within subgroups, split according to medication status, resulted in fair performance even after accounting for correlated clinical variables (i.e., disease duration) through additional splits. Evidence from rodent studies suggests that serotonin reuptake inhibitors (SRIs) mediate neuroplasticity in various cortical and subcortical structures through glio-genesis and neuro-genesis40,41,42. However, little is known about how these findings might translate to humans and what the effects of long-term medication use are43. The few longitudinal studies with small samples suggest that SRI treatment normalizes brain volumes. One study reported significantly larger thalamic volumes in twenty-one treatment-naïve pediatric patients compared to HC and that these differences decreased following paroxetine treatment44. Another study reported that smaller putamen volume in treatment-naïve patients was no longer detectable in the thirteen patients that were treated with fluoxetine45. Nonetheless, it remains unclear whether these structural changes are related to medication use or to symptom improvement. In contrast to these normalizing effects of SRIs, the previous univariate meta-analyses and mega-analyses of the ENIGMA-OCD study found significantly thinner cortices in medicated adult OCD patients and smaller cortical surface area in medicated pediatric OCD patients, but could not detect significant differences in cortical and subcortical gray matter between unmedicated OCD patients and HC, with the exception of larger thalamic volumes in unmedicated pediatric OCD patients6,7. Together, these cross-sectional studies suggest that medication use alters brain structure rather than necessarily normalizing it to the level of healthy individuals, an hypothesis which needs to be assessed in appropriately powered longitudinal studies.
The identification of which brain regions contributed most to the classification resulted from a multivariate analysis, and the localization of these regions should therefore be interpreted with caution. MVPA techniques typically result in better discriminative ability between groups compared to standard univariate analyses by taking the distributed nature of effects into account, but they do not provide inherent localization information (i.e., attributing effect sizes to individual ROIs) as all features used for prediction are considered as a whole. We derived individual feature importance from the RFC classifier using permutation-based inference to find brain features that contributed both significantly and consistently (across CV folds) to classification performance. Feature importance was derived from classifications using neuroimaging data after regression of covariates to avoid any undesirable effects on the interpretation of weights caused by non-imaging features. No feature importance obtained for the main classifications (OCD patients vs. HC) in either pediatric, adult or combined samples was statistically significant. This is likely due to the low classification performances obtained, suggesting that the features used are either too noisy or non-informative for main diagnosis predictions and unable to achieve statistical significance and consistency across folds. Similarly, no significant features were found for medicated OCD vs. HC and unmedicated OCD vs. HC classifications. On the other hand, significant features were found that enabled multivariate classifications for medicated vs. unmedicated OCD in adult and combined samples. These included widespread cortical thickness in frontal and temporal regions, including the left inferior temporal gyrus, medial orbital frontal and bilateral transverse temporal cortex, left insula, and bilateral anterior cingulate cortex, as well as surface area of the right entorhinal, left paracentral and bilateral temporal cortex, and left thalamus, pallidum, and ventricle volumes. A full overview of these significant features, as well as those found for early vs. late onset OCD classification, can be found in supplementary Tables S10-11.
The brain regions that were detected in the multivariate analysis are partially consistent with the results from previous univariate ENIGMA-OCD meta-analyses and mega-analyses6,7. Medicated adult OCD patients showed thinner frontal, temporal and parietal cortices, and smaller hippocampal and larger pallidum volumes compared to HC, whereas no differences were found for unmedicated adult patients. Although we were unable to detect significant features for medicated patients and unmedicated patients vs. HC classifications, these earlier results fit with the finding that classification performance for medicated patients vs. HC was better than for unmedicated patients vs. HC, as differences in brain anatomy of unmedicated patients appear to be minimal. Interestingly, the classification between medicated vs. unmedicated patients was even better, which suggests that this results from the minimization of heterogeneity in stratified patient groups as opposed to the larger heterogeneity seen in case-control comparisons. Finally, as the medication used for treating OCD is also used for treatment of many other psychiatric disorders, we anticipate that these results are not specific to OCD.
Another point that deserves emphasis concerns the different ways of dealing with confounding non-imaging variables (e.g., age and sex) when using neuroimaging data for MVPA classification. Although recent studies were unable to detect differences in predictive performance when comparing different approaches for dealing with confounds in MVPA studies (nor differences in the weights assigned by the models46,47), results from our confound control experiments suggest otherwise. We chose to add the covariates age, sex and data collection site ID directly as features to our model as our initial approach. The underlying principle of this approach is that all relevant variables should be included in the model and that their relative contribution to the final predictive model will be recovered during model training, without the need for manual confound adjustment procedures47. This approach resulted in high classification performance for medication classifications (>0.8 AUC). However, several sites included only medicated patients while in others no patients had received medication, which could suggest that this high performance was achieved through classifiers detecting site-effects directly from covariates (e.g., site ID) rather than brain data. The latter is supported by the finding that classifications using covariates only (age, sex, and site) also resulted in high AUC (>0.8 AUC), whereas regressing these covariates out from brain data resulted in lower performance, with only classifications for medicated vs. control classification remaining significant (supplementary Tables S9). Our control experiments also show that the FreeSurfer data itself is likely to be confounded by site-effects as well, as classifications using brain data only (without regressing out covariates) resulted in relatively high classification performance for medicated OCD vs. HC and medicated vs. unmedicated OCD classifications (with 0.70 and 0.75 AUC, respectively). This could be explained by classifiers being able to identify sites through specific sample characteristics (demography and inclusions criteria used) resulting in different brain anatomy, and methodical differences such as types of scanner and imaging protocols used. Interestingly, control experiments for main diagnosis classifications showed that these were relatively unaffected by different ways of dealing with confounds (supplementary Table S8). This is likely due to the fact that the classes for diagnosis classifications (i.e., number of OCD patients and HC) are more balanced across samples.
A number of limitations deserve emphasis. First, we used a sample pooled from existing data across the world, without harmonized protocols for scanning, inclusion criteria or demographic and clinical characteristics. These sources of heterogeneity may limit classification performance, but this also provides an opportunity for model development using independent data sets and the discovery of biomarkers that are reproducible across study sites. Second, standardized FreeSurfer protocols were used for MR data processing to ensure reproducibility across sites. It has been shown that FreeSurfer tends to overestimate subcortical volumes in children48, and that MR field strength can affect regional cortical estimations49. However, these nonsystematic effects are expected to affect patients and HC equally and are therefore not expected to influence our results. Third, limited information on medication use was available. We were therefore only able to distinguish patients on antidepressants with or without adjuvant antipsychotics vs. those who had not received any medication. Medication history, medication dosage, and duration of use were unknown. Nonetheless, these coarsely defined medication groups enabled better case-control discrimination and good classification of medicated vs. unmedicated cases. Fourth, there is a lack of information on OCD subtypes in our dataset. Particular OCD subtypes may have different neural correlates, and this might limit the ability of MVPA models to find generalizable patterns in brain structure14,50. Fifth, it should be noted that the age cut-off used to split the data in pediatric (age below 18) and adult (age 18 and older) samples may not be optimal with respect to the development of the brain, but this was done in accordance with the initial collection of pediatric and adult samples and previous ENIGMA work51,52. Finally, it is possible that the brain features used for classification led to sub-optimal performance. OCD is thought to derive from abnormalities distributed at the network-level rather than focused on a single brain area, and FreeSurfer features might not be sufficiently sensitive to detect subtle alterations associated with OCD.
Taken together, this study provides a realistic estimate of the classification performance that can be achieved in a large, ecologically valid, multi-site sample of OCD participants using data on regional brain structure. Our findings show that parcellated structural MRI data do not enable a good overall distinction between patients with OCD and HC. However, classifying subgroups of patients based on medication status enables fair identification at the individual subject level, which implies that medication use is associated with substantial distributed differences in brain anatomy. This underlines the need for longitudinal studies on the short-term and long-term effects of psychiatric medication on brain structure.
Code availability
The computer code for the above-described analyses is publicly available (https://github.com/WillemB2104/ENIGMA-OCD-2020).
References
Ruscio, A. M. et al. The epidemiology of obsessive-compulsive disorder in the National Comorbidity Survey Replication. Psychiatry Res. 15, 513–517 (2010).
American Psychiatric Association. Diagnostic and statistical manual of mental disorders, 5th edn. (DSM-5). Diagn. Stat. Man. Ment. Disord. 280, https://doi.org/10.1176/appi.books.9780890425596.744053. (2013).
Abramowitz, J. S., Taylor, S. & McKay, D. Obsessive-compulsive disorder. Lancet 374, 491–499 (2009).
Menzies, L. et al. Integrating evidence from neuroimaging and neuropsychological studies of obsessive-compulsive disorder: the orbitofronto-striatal model revisited. Neurosci. Biobehav. Rev. 32, 525–549 (2008).
van den Heuvel, O. A. et al. Brain circuitry of compulsivity. Eur. Neuropsychopharmacol. 26, 810–827 (2016).
Boedhoe, P. S. W. et al. Distinct subcortical volume alterations in pediatric and adult OCD: a worldwide meta- and mega-analysis. Am. J. Psychiatry 174, 60–69 (2017).
Boedhoe, P. S. W. et al. Cortical abnormalities associated with pediatric and adult obsessive-compulsive disorder: findings from the ENIGMA obsessive-compulsive disorder working group. Am. J. Psychiatry 175, 453–462 (2018).
Kong, X. Z. et al. Mapping cortical and subcortical asymmetry in obsessive-compulsive disorder: findings from the ENIGMA Consortium. Biol. Psychiatry 1–13 https://doi.org/10.1016/j.biopsych.2019.04.022. (2019).
de Wit, S. J. et al. Multicenter voxel-based morphometry multi-center mega-analysis of structural brain scans in obsessive-compulsive disorder. Am. J. Psychiatry 171, 340–349 (2013).
Rotge, J. Y. et al. Provocation of obsessive-compulsive symptoms: a quantitative voxel-based meta-analsysis of functional neuroimaging studies. J. Psychiatry Neurosci. 33, 405 (2008).
Orrù, G., Pettersson-Yeo, W., Marquand, A. F., Sartori, G. & Mechelli, A. Using Support Vector Machine to identify imaging biomarkers of neurological and psychiatric disease: a critical review. Neurosci. Biobehav. Rev. 36, 1140–1152 (2012).
Arbabshirani, M. R., Plis, S., Sui, J. & Calhoun, V. D. Single subject prediction of brain disorders in neuroimaging: promises and pitfalls. Neuroimage https://doi.org/10.1016/j.neuroimage.2016.02.079. (2016).
Klöppel, S. et al. Automatic classification of MR scans in Alzheimer’s disease. Brain 131, 681–689 (2008).
Bruin, W., Denys, D. & van Wingen, G. Diagnostic neuroimaging markers of obsessive-compulsive disorder: Initial evidence from structural and functional MRI studies. Prog. Neuro-Psychopharmacology Biol. Psychiatry 0–1 https://doi.org/10.1016/j.pnpbp.2018.08.005. (2018).
Varoquaux, G. Cross-validation failure: small sample sizes lead to large error bars. Neuroimage 1–10 https://doi.org/10.1016/j.neuroimage.2017.06.061. (2017).
Woo, C.-W., Chang, L. J., Lindquist, M. A. & Wager, T. D. Building better biomarkers: brain models in translational neuroimaging. Nat. Neurosci. Rev. 20, 365–377 (2017).
Wolfers, T., Buitelaar, J. K., Beckmann, C. F., Franke, B. & Marquand, A. F. From estimating activation locality to predicting disorder: A review of pattern recognition for neuroimaging-based psychiatric diagnostics. Neurosci. Biobehav. Rev. 57, 328–349 (2015).
Nunes, A. Using structural MRI to identify bipolar disorders–13 site machine learning study in 3020 individuals from the ENIGMA Bipolar Disorders Working Group. Mol. Psychiatry https://doi.org/10.1038/s41380-018-0228-9. (2018).
Nieuwenhuis, M. et al. Classification of schizophrenia patients and healthy controls from structural MRI scans in two large independent samples. Neuroimage 61, 606–612 (2012).
Franke, K., Ziegler, G., Klöppel, S. & Gaser, C. Estimating the age of healthy subjects from T1-weighted MRI scans using kernel methods: exploring the influence of various parameters. Neuroimage 50, 883–892 (2010).
Schnack, H. G. & Kahn, R. S. Detecting neuroimaging biomarkers for psychiatric disorders: Sample size matters. Front. Psychiatry 7, 50 (2016).
Boedhoe, P. S. W. et al. An empirical comparison of meta- and mega-analysis with data from the ENIGMA Obsessive-Compulsive Disorder Working Group. Front. Neuroinform. 12, 1–8 (2019).
Silverman, W. K., Saavedra, L. M. & Pina, A. A. Test-retest reliability of anxiety symptoms and diagnoses with the anxiety disorders interview schedule for DSM-IV: child and parent versions. J. Am. Acad. Child Adolesc. Psychiatry https://doi.org/10.1097/00004583-200108000-00016. (2001).
Kaufman, J. et al. Schedule for affective disorders and schizophrenia for school-age children-present and lifetime version (K-SADS-PL): initial reliability and validity data. J. Am. Acad. Child Adolesc. Psychiatry 36, 980–988 (1997).
Sheehan, D. V. et al. Reliability and validity of the mini international neuropsychiatric interview for children and adolescents (MINI-KID). J. Clin. Psychiatry 71, 313–326 (2010).
First, M. B. Structured Clinical Interview for the DSM (SCID). In The Encyclopedia of Clinical Psychology 1–6 (John Wiley & Sons, Inc., 2015).
Sheehan, D. V. et al. The Mini-International Neuropsychiatric Interview (M.I.N.I.): the development and validation of a structured diagnostic psychiatric interview for DSM-IV and ICD-10. in. J. Clin. Psychiatry 59, 22–33 (1998).
Desikan, R. S. et al. An automated labeling system for subdividing the human cerebral cortex on MRI scans into gyral based regions of interest. Neuroimage 31, 968–980 (2006).
Hastie, T., Tibshirani, R. & Friedman, J. The Elements of Statistical Learning (2nd edn.) (Elements, 2009).
Rasmussen, C. E. & Williams, C. K. I. Gaussian processes for machine learning. Int. J. Neural Syst. 14, 69–106 (2004).
Chang, C. & Lin, C. LIBSVM: a library for support vector machines. ACM Trans. Intell. Syst. Technol. 2, 1–39 (2013).
Chen, T. & Guestrin, C. XGBoost: a scalable tree boosting system. In Proceedings of the ACM SIGKDD International Conference on Knowledge Discovery and Data Mining vols 13-17-785–794 (ACM Press, 2016).
Bradley, A. P. The use of the area under the ROC curve in the evaluation of machine learning algorithms. Pattern Recognit. 30, 1145–1159 (1997).
Abraham, A. et al. Deriving reproducible biomarkers from multi-site resting-state data: an Autism-based example. Neuroimage 147, 736–745 (2017).
Hanley, J. A. & McNeil, B. J. The meaning and use of the area under a receiver operating characteristic (ROC) curve. Radiology 143, 29–36 (1982).
Mason, S. J. & Graham, N. E. Areas beneath the relative operating characteristics (ROC) and relative operating levels (ROL) curves. Q. J. R. Meteorol. Soc. 128, 2145–2166 (2002).
Storch, E. A. et al. Defining clinical severity in adults with obsessive-compulsive disorder. Compr. Psychiatry 63, 30–35 (2015).
Strobl, C., Boulesteix, A. L., Zeileis, A. & Hothorn, T. Bias in random forest variable importance measures: illustrations, sources and a solution. BMC Bioinformatics 8, 25 (2007).
Thomas, R. M., Bruin, W., Zhutovsky, P. & van Wingen, G. Dealing with missing data, small sample sizes, and heterogeneity in machine learning studies of brain disorders. In Machine Learning 249–266 (Elsevier, 2020).
Czéh, B. et al. Chronic social stress inhibits cell proliferation in the adult medial prefrontal cortex: hemispheric asymmetry and reversal by fluoxetine treatment. Neuropsychopharmacology https://doi.org/10.1038/sj.npp.1301275. (2007).
Kodama, M., Fujioka, T. & Duman, R. S. Chronic olanzapine or fluoxetine administration increases cell proliferation in hippocampus and prefrontal cortex of adult rat. Biol. Psychiatry https://doi.org/10.1016/j.biopsych.2004.07.008 (2004).
Soumier, A. et al. Mechanisms contributing to the phase-dependent regulation of neurogenesis by the novel antidepressant, agomelatine, in the adult rat hippocampus. Neuropsychopharmacology https://doi.org/10.1038/npp.2009.72. (2009).
Homberg, J. R., Schubert, D. & Gaspar, P. New perspectives on the neurodevelopmental effects of SSRIs. Trends Pharmacol. Sci. 31, 60–65 (2010).
Gilbert, A. R. et al. Decrease in thalamic volumes of pediatric patients with obsessive-compulsive disorder who are taking paroxetine. Arch. Gen. Psychiatry 57, 449 (2003).
Hoexter, M. Q. et al. Gray matter volumes in obsessive-compulsive disorder before and after fluoxetine or cognitive-behavior therapy: a randomized clinical trial. Neuropsychopharmacology 37, 734–745 (2012).
Linn, K. A., Gaonkar, B., Doshi, J., Davatzikos, C. & Shinohara, R. T. Multivariate pattern analysis and confounding in neuroimaging. Int. J. Biostat. 12, 31 (2016).
Rao, A., Monteiro, J. M. & Mourao-Miranda, J. Predictive modelling using neuroimaging data in the presence of confounds. Neuroimage 150, 23–49 (2017).
Schoemaker, D. et al. Hippocampus and amygdala volumes from magnetic resonance images in children: assessing accuracy of FreeSurfer and FSL against manual segmentation. Neuroimage https://doi.org/10.1016/j.neuroimage.2016.01.038 (2016).
Govindarajan, K. A., Freeman, L., Cai, C., Rahbar, M. H. & Narayana, P. A. Effect of intrinsic and extrinsic factors on global and regional cortical thickness. PLoS ONE 9, 1–15 (2014).
Mataix-Cols, D. & van den Heuvel, O. A. Common and distinct neural correlates of obsessive-compulsive and related disorders. Psychiatr. Clin. North Am. 29, 391–410 (2006).
Leckman, J. F. et al. Obsessive-compulsive disorder: a review of the diagnostic criteria and possible subtypes and dimensional specifiers for DSM-V. Depression Anxiety https://doi.org/10.1002/da.20669 (2010).
Anholt, G. E. et al. Age of onset in obsessive-compulsive disorder: admixture analysis with a large sample. Psychol. Med. https://doi.org/10.1017/S0033291713000470. (2014).
Acknowledgements
The ENIGMA-Obsessive Compulsive Disorder Working-Group gratefully acknowledges support from the NIH BD2K (Big Data to Knowledge) award U54 EB020403 (PI: Dr. Thompson) and Neuroscience Amsterdam, and an IPB grant to Dr. Schmaal and Dr. van den Heuvel. Supported by the Japan Society for the Promotion of Science (JSPS; KAKENHI Grants No. 18K15523 to Dr. Abe, No. 16K19778, No. 18K07608 to Dr. Nakamae, No. 16K04344, 19K03309 to Dr. Hirano, and No. 26461762 to Dr. Nakagawa); the Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP, São Paulo Research Foundation; Grant No. 2011/21357-9; 2018/04654-9 and 2018/21934-5); the EU FP7 project TACTICS (Grant No. 278948 to Dr. Buitelaar); the National Natural Science Foundation of China (No. 81560233 to Dr. Cheng); the International Obsessive-Compulsive Disorder Foundation (IOCDF) Research Award to Dr. Gruner; the Dutch Organization for Scientific Research (NWO) (grants 912-02-050, 907-00-012, 940-37-018, and 916.86.038); the Netherlands Society for Scientific Research (NWO-ZonMw VENI grant 916.86.036 to Dr. van den Heuvel; NWO-ZonMw AGIKO stipend 920-03-542 to Dr. de Vries), a NARSAD Young Investigator Award to Dr. van den Heuvel, and the Netherlands Brain Foundation (2010(1)-50 to Dr. van den Heuvel); the Federal Ministry of Education and Research of Germany (No. BMBF-01GW0724 to Dr. Kathmann); the Alberta Innovates Translational Health Chair in Child and Youth Mental Health and funding from the Ontario Brain Institute to Dr. Arnold; the Deutsche Forschungsgemeinschaft (DFG; Grant No. KO 3744/7-1 to Dr. Koch); the Helse Vest Health Authority (No. 911754, 911880 to Dr. Kvale) and the Norwegian Research Council (No. HELSEFORSK 243675 to Dr. Kvale); the Wellcome Trust and a pump priming grant from the South London and Maudsley Trust, London, UK (Project Grant No. 064846 to Dr. Mataix-Cols); the Generalitat de Catalunya (AGAUR 2017 SGR 1247 to Dr. Menchón); the PhD-iHES program (FCT fellowship Grant No. PDE/BDE/113601/2015 to Dr. Moreira); the Japanese Ministry of Education, Culture, Sports, Science and Technology (Grant-in-Aid for Scientific Research (C) 22591262, 25461732, 16K10253 to Dr. Nakao); the Italian Ministry of Health (No. RC13-14-15-16-17-18A to Dr. Spalletta, Dr. Piras Fabrizio, Dr. Piras Federica); the National Center for Advancing Translational Sciences (Grants No. UL1TR000067/KL2TR00069 to Dr. Stern); the SA MRC funding and the National Research Foundation of South Africa to Dr. Stein and Dr. Lochner; the Canadian Institutes of Health Research, Michael Smith Foundation for Health Research, and British Columbia Provincial Health Services Authority funding Dr. Stewart; the Netherlands Organization for Scientific Research (NWO/ZonMW Vidi 917.15.318 to Dr. van Wingen); the Wellcome-DBT India Alliance (Grant No. 500236/Z/11/Z to Dr. Venkatasubramanian); the National Natural Science Foundation of China (No. 81371340) and the Shanghai Key Laboratory of Psychotic Disorders (No. 13dz2260500) to Dr. Wang; the Government of India grants from the Department of Science and Technology (Grants No. SR/S0/HS/0016/2011 to Prof. Y.C. Janardhan Reddy, and DST INSPIRE faculty grant-IFA12-LSBM-26 to Dr. Janardhanan C. Narayanaswamy) and from the Department of Biotechnology (Grants No. BT/PR13334/Med/30/259/2009 to Prof. Y.C. Janardhan Reddy, and No. BT/06/IYBA/2012 to Dr. Janardhanan C. Narayanaswamy); the Dana Foundation and NARSAD to Dr. Fitzgerald; the NIMH (Grants No. R01 MH107419 and No. K23MH082176 to Dr. Fitzgerald, Grant No. R21MH101441 to Dr. Marsh, Grant No. R21MH093889 to Drs. Marsh and Simpson, Grant No. K23-MH104515 to Dr. Baker; Grant No. R33MH107589 to Dr. Stern, Grant No. R01MH111794 to Dr. Stern; Grant No. R01 MH081864 to Drs. O’Neill and Piacentini; Grant No. R01 MH085900 to Drs. O’Neill and Feusner, Grant No. K23 MH115206 to Dr. Gruner); the NIMH and the David Judah Fund at the Massachusetts General Hospital (Grant No. K23-MH092397 to Dr. Brennan); the Marató TV3 Foundation (Grants No. 01/2010 and 091710 to Dr. Lazaro); the Carlos III Health Institute (PI040829 to Dr Lazaro, Grant CPII16/00048 and Project Grants PI13/01958 and PI16/00889 to Dr. Soriano-Mas), co-funded by FEDER funds/European Regional Development Fund (ERDF), a way to build Europe; the AGAUR (2017 SGR 881 to Dr. Lazaro); the Michael Smith Foundation for Health Research funding Dr. Jaspers-Fayer and Dr. Stewart; the Carlos III Health Institute (PI14/00419 to Dr. Alonso, Grant No. FI17/00294 to Ignacio Martínez-Zalacaín, Project Grant No. PI16/00950 to Dr. Menchón); the Japan Agency for Medical Research and Development (AMED Brain/MINDS Beyond program Grant No. JP19dm0307002 to Dr. Shimizu); Oxford Health Services Research Committee grant to Dr A James and National Institute on Drug Abuse to Dr. Ivanov (NIH/NIDA Grant No. R21/DA046029 and No. R21/DA045218); the Wellcome and Royal Society (211155/Z/18/Z) to Tobias U Hauser; the Jacobs Foundation (2017-1261-04) to Tobias U Hauser, the Medical Research Foundation, and the Brain and Behavior Research Foundation (2018 NARSAD Young Investigator grant; 27023) to Tobias U. Hauser; the Swiss National Science Foundation (SNSF 320030_130237) to Susanne Walitza; the Harmann Müller foundation (1460) to Silvia Brem; NHMRC Career Development Fellowship (1140764 to Dr. Schmaal).
Author information
Authors and Affiliations
Consortia
Corresponding authors
Ethics declarations
Conflict of interest
Dr. Baker has received research support from the National Institute of Mental Health (NIMH) and Valera Health. Dr. Brennan has received consulting fees from Rugen Therapeutics and Nobilis Therapeutics and research grant support from Eli Lilly, Transcept Pharmaceuticals, and Biohaven Pharmaceuticals. Dr. Walitza has received lecture honoraria Opopharma in the last 3 years. Her work was supported in the last 3 years by the Swiss National Science Foundation (SNF), diverse EU FP7s, HSM Hochspezialisierte Medizin of the Kanton Zurich, Switzerland, Bfarm Germany, Zinep, Hartmann Müller Stiftung, Olga Mayenfisch. Dr. Dan J. Stein has received research grants and/or consultancy honoraria from Lundbeck and Sun in the past 3 years. Dr. Paul M. Thompson has received research grant support from Biogen, Inc., for research unrelated to the topic of this manuscript. Dr. Ivanov has received honoraria from Lundbeck as a member of the Data Safety Monitoring Committee and research grants from the National Institute on Drug Abuse in the last 3 years. Dr. Pittenger has received research support and/or honoraria for consultation from Biohaven Pharmaceuticals, Blackthorn Therapeutics, Abide Therapeutics, and Brainsway, and royalties or honoraria from Oxford University Press and Elsevier in the past 3 years. Dr. Feusner has received an honorarium from Pfizer and consultation fees from NOCD, Inc. Dr. Piacentini has received research support from Pfizer Pharmaceuticals for research unrelated to the topic of this manuscript. Dr. Soreni has received support for Investigator Initiated Clinical Trial from Lundbeck LLC unrelated to the topic of this study. Dr. Buitelaar has been in the past 3 years a consultant to/member of advisory board of/and/or speaker for Shire, Roche, Medice, and Servier. He is not an employee of any of these companies, and not a stock shareholder of any of these companies. He has no other financial or material support, including expert testimony, patents, royalties. Dr. Mataix-Cols receives royalties for contributing articles to UpToDate (Wolters Kluwer Health), and for editorial work from Elsevier, all unrelated to the current work. In the last three years, Dr. Simpson has received research support for an industry-sponsored clinical trial from Biohaven Pharmaceuticals, royalties from UpToDate, Inc, and a stipend from JAMA Psychiatry for her role as Associate Editor. Dr. Tolin has received research support from Biohaven Pharmaceuticals. All other individually-named authors in- and outside of the ENIGMA-OCD working group reported no biomedical financial interests or potential conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bruin, W.B., Taylor, L., Thomas, R.M. et al. Structural neuroimaging biomarkers for obsessive-compulsive disorder in the ENIGMA-OCD consortium: medication matters. Transl Psychiatry 10, 342 (2020). https://doi.org/10.1038/s41398-020-01013-y
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-020-01013-y
This article is cited by
-
Brain-based classification of youth with anxiety disorders: transdiagnostic examinations within the ENIGMA-Anxiety database using machine learning
Nature Mental Health (2024)
-
White matter diffusion estimates in obsessive-compulsive disorder across 1653 individuals: machine learning findings from the ENIGMA OCD Working Group
Molecular Psychiatry (2024)
-
In search of environmental risk factors for obsessive-compulsive disorder: study protocol for the OCDTWIN project
BMC Psychiatry (2023)
-
Sampling inequalities affect generalization of neuroimaging-based diagnostic classifiers in psychiatry
BMC Medicine (2023)
-
Association of granulocyte macrophage colony-stimulating factor and interleukin-17 levels with obsessive–compulsive disorder: a case–control study findings
Scientific Reports (2023)