Abstract
The schizophrenia-associated gene, BRD1, encodes an epigenetic regulator in which chromatin interactome is enriched with genes implicated in mental health. Alterations in histone modifications and epigenetic regulation contribute to brain transcriptomic changes in affective disorders and preclinical data supports a role for BRD1 in psychopathology. However, the implication of BRD1 on affective pathology remains poorly understood. In this study, we assess affective behaviors and associated neurobiology in Brd1+/− mice along with their responses to Fluoxetine and Imipramine. This involves behavioral, neurostructural, and neurochemical characterizations along with regional cerebral gene expression profiling combined with integrative functional genomic analyses. We report behavioral changes in female Brd1+/− mice with translational value to depressive symptomatology that can be alleviated by the administration of antidepressant medications. Behavioral changes are accompanied by altered brain morphometry and imbalances in monoaminergic systems. In accordance, gene expression changes across brain tissues reveal altered neurotransmitter signaling and cluster in functional pathways associated with depression including ‘Adrenergic-, GPCR-, cAMP-, and CREB/CREM-signaling’. Integrative gene expression analysis specifically links changes in amygdaloid intracellular signaling activity to the behavioral treatment response in Brd1+/− mice. Collectively, our study highlights the importance of BRD1 as a modulator of affective pathology and adds to our understanding of the molecular mechanisms underlying affective disorders and their treatment response.
Similar content being viewed by others
Introduction
Psychiatric disorders comprise a heterogeneous group of disabling conditions collectively characterized by self-reported and clinically observed changes in state of well-being and abnormal behaviors1. Suggestive of interconnected etiologies, clinical, and therapeutic profiles are overlapping, and risk factors are shared between disorders2,3. In line with the neurodevelopmental hypothesis of psychiatric disorders4, this includes a complex interplay between genetic risks5 and early life adverse exposures6. However, the molecular and biological mechanisms that trigger early life programming and development of psychopathology are poorly understood. Epigenetic processes, such as acetylation of histone lysine residues, are linked with brain development as well as lifelong neural plasticity7 and have been implicated with the pathophysiology of both psychotic and affective disorders8. Correspondingly, a number of clinically effective antidepressants are known to affect the status of cerebral acetylation of histone lysine residues (Kac)9,10,11 and histone deacetylases (HDACs) have been suggested as direct therapeutic targets for depressive disorders12,13. The interpretation of Kac marks is facilitated by the reader domains of the bromodomain (BRD) family of proteins14. Whereas Kac is generally associated with activation of transcription through opening of the chromatin structure, they may also signal for the compaction of chromatin, protein stability, and the regulation of protein-protein interactions15.
Bromodomain containing-1 (BRD1) has been identified in complexes involved with both histone acetylation and chromatin remodeling16,17 and interacts at genomic sites enriched with genes implicated in neurodevelopmental processes17,18. BRD1 is widely expressed in human brain19, differentially regulated in limbic and neocortical tissues upon exposure to external stressors in rats20,21, and involved in the epigenetic regulation of embryonic development, survival, and differentiation of embryonic stem cells16,22,23. Supporting a role for BRD1 in mental health, BRD1 has repeatedly been associated with schizophrenia and bipolar disorder in genetic studies24,25,26,27,28, including gene-wise significant association in the currently largest schizophrenia GWAS mega-analysis29 and genome-wide significance in the Psychiatric Genomics Consortium (PGC1) schizophrenia sample30. The locus does not show significant association in the most recent GWASs of bipolar disorder and major depressive disorder31,32. Despite being highly intolerant to loss of function mutations33, a disruptive nonsense mutation in BRD1 has been reported in a schizophrenia case34. In concordance, we have recently shown that male mice with reduced expression of Brd1 (Brd1+/− mice) recapitulate cardinal features relating to schizophrenia35,36,37,38 (Table S1). In the present study, we assess the impact of reduced Brd1 expression on affective symptomatology, neurochemistry, and neurobiology in mice along with their responsiveness to pharmacological intervention using clinically effective antidepressants. Through global gene expression profiling of selected brain regions, we assess the neuromolecular mechanisms that underly BRD1’s role in affective pathology.
Materials and methods
Animals
A mouse line heterozygous for a targeted deletion within the Brd1 gene, C57BL/6NTac-Brd1tm1.2Arte/AborgMmucd (Brd1+/−) was generated by TaconicArtemis GmbH (Cologne, Germany) using a targeting vector (pBrd1 Final cl 1 (UP0257)) with loxP sites flanking exon 3–5 of the Brd1 gene. For further details, see Supplementary Methods and Materials. The mouse strain has been deposited and is available at Mutant Mouse Resource and Research Center (MMRRC) at University of California at Davis (RRID:MMRRC_065563-UCD). All studies were carried out in accordance with Danish legislation, and permission for the experiments was granted by the animal welfare committee, appointed by the Danish Ministry of Food, Agriculture and Fisheries—Danish Veterinary and Food Administration.
Experimental design
All experiments involved 7–15 mice, 8–11 weeks old, in each group (figure legends present exact numbers). The sample size was chosen on the basis of the resource equation method39. Litter-matched WT and Brd1+/− mice were randomly allocated to individual tests. Observer was blind to mice genotypes. General assessment of neurology, testing in motor coordination tests, prepulse inhibition (PPI), fear conditioning (FC), 8-arm radial maze (8ARM), open field (OF), tail suspension test (TST), forced swim test (FST), bright open field (BOF), light and dark box (LDB), elevated plus maze (EPM) and quantification of neurotransmitters by high-pressure liquid chromatography (HPLC) were performed in parallel on age-matched male and female littermates. However, in this study, PPI, FC, and 8ARM were only reported in female mice, since corresponding male data have already been published elsewhere35,36. One batch of mice completed OF, TST and FST, while another completed BOF, LDB, and EPM. An independent batch of mice was injected subcutaneously with either vehicle (normal saline solution), imipramine (IMN, 1 or 10 mg/kg) or fluoxetine (FLX, 5 mg/kg), dissolved in saline and subjected to OF, TST, and FST. Each mouse underwent 2 days of experiments. Day 1: one hour after the injection, the mice completed TST and were subsequently returned to their cages. Day 2: mice had a second injection. One hour after the injection, the mice completed OF and FST. There was no delay between OF and FST. One hour after the completion of FST, they were sacrificed, and brains were collected.
Amphetamine-induced hyperactivity (AIH), cocaine-induced hyperactivity (CIH), sucrose preference test (SPT), and 24-h locomotor activity (24HLM) and OF, TST and FST following administration of antidepressants were only tested and reported in female mice in this study. Mice were not reused for other experiments.
General assessment of neurology, motor coordination, and behavioral tests
Details on functional observation battery, acute pain response, rotarod, balance beam walking, foot-printing, FST, TST, SPT, OF, BOF, LDB, EPM, FC, 8ARM, 24HLM, PPI, AIH, and CIH can be found in Supplementary Methods and Materials.
Quantification of neurotransmitters
Mice were sacrificed by cervical dislocation and frontal cortical, hippocampal, and striatal tissues were collected by free-hand dissection and processed for quantitative HPLC analyses of dopamine and serotonin. For details on HPLC procedures, see Supplementary Methods and Materials.
Brain morphometry, Golgi-cox staining, and 3-D image analysis
Left cerebral hemispheres (n = 8/group) were stained with FD Rapid Golgi-Stain kit (FD Neurotechnologies, Ellicott City, USA), and cut into 150 µm thick-slices on a vibratome-3000 (Vibratome, St Louis, MO, USA). Anterior cingulate cortex (aCC) pyramidal neurons were identified (×60; oil-immersion; numerical-aperture = 1.4) by their prominent apical dendrites, and 6 neurons/mouse were chosen by systematic uniform random sampling. Image stacks (90-105 consecutive images at 1 µm interval) were captured by optical wide-field microscopy (Olympus BX50, Tokyo, Japan) and newCAST software (Visiopharm, Hoersholm, Denmark). 3-D image reconstruction and analyses were completed using Imaris software version 7.6.3 (Bitplane AG, Zurich, Switzerland). For a description on brain morphometric analyses, see Supplementary Methods and Materials.
Statistical analysis
STATA 15.1 (StataCorp LLC, College Station, TX) and GraphPad Prism 8.1.2 (GraphPad Software, San Diego, CA) software was used for analyzing our data with appropriate tests of statistical significance including t-test and two-way RMANOVA. We checked whether all continuous variables followed Gaussian distribution using Shapiro–Wilk tests. When the study variables did not follow Gaussian distribution, appropriate non-parametric tests such as Mann–Whitney U test were employed. F tests were used for assessing equality of variances and Welch corrections were applied, when needed.
RNA-sequencing and data analyses
Mouse brains from IMN (10 mg/kg) and FLX (5 mg/kg) (n = 9–10/group) were sectioned coronally (1 mm thick) using a slicer matrix (Zivic Instruments, Pittsburgh, USA). For the IMN (10 mg/kg) group, right amygdala (AMG), striatum, caudate putamen (CPu), and aCC were identified, and punched by a punch-needle (1 mm diameter) at −20 °C, whereas for the FLX (5 mg/kg) group only AMG was sampled. RNA was extracted using Maxwell-16 instrument system and LEV simplyRNA Tissue Kit (Promega, Madison, USA). Agilent 2100 Bioanalyzer (Agilent technologies, SantaClara, USA) confirmed the quality of RNA with a mean RNA Integrity Number (RIN) of 7.87 (SD 0.26). For AMG samples, libraries were prepared using TruSeq library preparation kit and RNA-sequencing performed on the Illumina HiSeq2000 platform (Illumina, San Diego, USA). For CPU and aCC samples, libraries were prepared using Beijing Genomics Institute (BGI) library preparation kits and protocols and sequencing performed on the BGISEQ-500 platform. A minimum of 10 million clean 50 bp single-end reads were generated for each sample. Reads that passed quality control (more than 90% bases having less than 1% sequencing error; No ambiguous bases) were aligned to mouse genome (Mus_musculus.GRCm.38.90) by HISAT2 (version 2.1.0)40 and counted by StringTie (version 1.3.4)41. Differentially Expressed Genes (DEGs) were identified by edgeR3.24.142 and reported after Benjamini-Hochberg false discovery rate (FDR) (5%) correction (14, 16) or as nominally significant DEGs (p < 0.01). Functional analyses of all nominally significant DEGs were performed using Ingenuity Pathway Analysis (Ingenuity, Redwood City, USA).
Results
General assessment of neurology and motor coordination
Male and female Brd1+/− mice were overall healthy, as described elsewhere35. However, female mice showed marginally impaired growth and slightly reduced size35. Systematic testing of general neurological functions of female Brd1+/− mice, revealed mildly reduced performance in the grip strength and wire-maneuvering tasks (Table 1). Female Brd1+/− mice did not differ significantly on their pain response (Fig. S1A), but were mildly impaired in their motor coordination, as evident from their rotarod performance (Fig. S1B, two-way ANOVA; p = 0.047)) and gaiting pattern (Fig. S1F, gaiting uniformity, t-test; p = 0.039). However, as they performed at par with their WT littermates in the beam walking task (Fig. S1G–H), we considered female Brd1+/− mice fit for testing in settings assessing complex behaviors. Assessment of motor coordination in male Brd1+/− mice has been reported previously (Table S1)35.
Locomotor activity and sensorimotor response in Brd1 +/− mice
General locomotor activity was assessed in the OF where male (Fig. 1a) and female (Fig. 1b) Brd1+/− mice performed at par with their WT littermates. As reported in male Brd1+/− mice (Table S1)35, female Brd1+/− mice displayed significantly increased acoustic startle responsivity (ASR) (Fig. 1c, two-way ANOVA; p = 0.003), both when initially introduced to the test setting (Fig. 1c, Tukey’s post hoc test; p = 0.004) and before baseline PPI testing (Fig. 1c, Tukey’s post hoc test; p = 0.006). Response latency to the startle was furthermore significantly shorter in female Brd1+/− mice than in WT mice (Fig. 1d, t-test; p = 0.044). The magnitude of baseline startle is known to influence PPI43. Accordingly, female Brd1+/− mice displayed reduced PPI across the span of tested prepulse intensities (Fig. 1e, two-way ANOVA; p = 0.049).
a Male mice (Mann–Whitney U = 94.0; p = 0.46). b Female mice (t = 0.20; p = 0.85): Total distance moved in the open field (OF) (n = 15/group). c Female Brd1+/− mice (n = 18) displayed significantly increased acoustic startle responsivity (ASR) compared to WT mice (n = 17) (genotype effect, F = 10.10, p = 0.003) both when initially introduced to the test setting (Tukey’s post hoc test, t = 3.07; p = 0.004) and before baseline PPI testing (Tukey’s post hoc test, t = 2.94; p = 0.006). d Response latency to the startle was furthermore significantly shorter than in WT mice (t = 2.09, p = 0.044). e Female Brd1+/− mice (n = 18) displayed reduced prepulse inhibition (PPI) compared to WT mice (n = 17) across the span of tested prepulse intensities (genotype effect, F = 4,163, p = 0.049); Cognitive functioning of female Brd1+/− mice. f–h Fear conditioning system (FCS) (n = 15/group). f Time spent as freezing during the repeated presentation of conditioning stimuli (CS; 30 s, 80db, white noise) on day one. Unconditioned stimulus (UCS) was 0.7 mA foot shock for 2 s. Inter-pairing intervals varied with an average of 60 s (30–120 s). Female Brd1 + /− mice displayed significantly less freezing during the conditioning phase of FCS (F = 12.26; p = 0.002). g Duration of freezing behavior within the first 3 min, after entering the conditioned context on the second day. Female Brd1+/− mice displayed significantly reduced freezing behavior (t = 2.23; p = 0.03). h Time spent as freezing during the first presentation of CS (30 s) in the novel context on the third day. Female Brd1+/− mice did not display cue-dependent learning deficits. i, j 8-Arm Radial Maze (8ARM) (n = 12/group). i Working memory errors, measured by the total number of re-entries to baited arms. j Reference memory errors, measured by the total number of entries to non-baited arms. Our 8ARM setup, which provided clear intramaze cues, predominantly assessed non-spatial memory and associative learning. Female Brd1+/− mice did not show working memory deficits, but displayed significant non-spatial reference memory deficits (F = 5.57; p = 0.03). k–m Female mice. k Total time spent in the central zone of bright open field (BOF) (n = 15/group). l Total time spent in the light box of light and dark box test (LDB) (n = 15/group). m Total time spent in the open arms of elevated plus maze (EPM) (n = 15/group). Female Brd1+/− mice did not display any anxiety equivalent behaviors. n–p Male mice. n Total time spent in the central zone of BOF (n = 15/group). o Total time spent in the light box of LDB (n = 15/group). This finding was not corroborated by the BOF and EPM results of male Brd1+/− mice. p Total time spent in the open arms of EPM (n = 15/group). Male Brd1+/− mice spent significantly less time in the light box of LDB (t = 2.97; p = 0.006), but they did not exhibit similar anxiety-like behaviors in BOF or EPM. q Female Brd1+/− mice were significantly more immobile in TST (n = 15/group; t = 3.01; p = 0.007) (r) and in FST (n = 15/group; F = 12.26; p = 0.002). s Male Brd1+/− mice did not display such behavioral despair in TST (n = 15/group) (t) and FST (n = 15/group). u Female Brd1+/− mice did not differ from WT mice in distance moved over 24 h (n = 10/group). Dark indicates the time when the lights were switched off in the stable, while light indicates the time when they were switched on. v Female Brd1+/− mice showed significantly less sucrose preference than the WT mice (n = 11/group; F = 14.03; p = 0.001), sucrose preference (weight of 2% sucrose solution consumed/ weight of total fluid consumed) in percentage. Data shown are mean and SEM for each group *p < 0.05; **p < 0.01.
Cognition
Female Brd1+/− mice froze significantly less than WT mice during the conditioning phase of FCS (Fig. 1f, two-way ANOVA; p = 0.002) and when returning to the same context on the following day (Fig. 1g, t-test; p = 0.03), collectively suggestive of a central acquisition deficit in female Brd1+/− mice. However, female Brd1+/− mice did not differ significantly from the WT mice on their cue dependent learning (Fig. 1h and Table 1) or on working memory errors in 8ARM (Fig. 1i). They did, however, make significantly more entries into the never-baited arms, indicative of impaired reference memory (Fig. 1j, two-way ANOVA; p = 0.03) (Table 1). Cognitive performance of male Brd1+/− mice tested in parallel has previously been reported35,36 and results are summarized in Table S1.
Affective behaviors in Brd1 +/− mice
Affective behaviors were assessed in both male and female Brd1+/− mice (Table 1 and Table S1). Female Brd1+/− mice did not differ significantly in the time spent in the central zone of BOF (Fig. 1k), in the light box of LDB (Fig. 1l) or in the open arms of EPM (Fig. 1m). Although male Brd1+/− mice spent significantly less time in the light box of LDB (Fig. 1n, t-test; p = 0.006), they did not exhibit similar anxiety-like behaviors in BOF (Fig. 1o) or EPM (Fig. 1p). Suggestive of behavioral despair, female Brd1+/− mice were significantly more immobile in TST (Fig. 1q, t-test; p = 0.007) and in FST (Fig. 1r, two-way ANOVA; p = 0.002) compared to WT mice, whereas this was not evident in male Brd1+/− mice (Fig. 1s, t). FST male WT immobility was higher than in female WT mice (Wilcoxon rank-sum; p = 0.001) and the WT male and female mice did not differ significantly on their TST immobility (t-test; p = 0.09)(Fig. 1q–t). Both TST (two-way ANOVA; p = 0.013) and FST (two-level mixed effects GLM; p = 0.014) data confirmed statistically significant interactions between gender and genotypes. Hence, we decided to assess additional affective behaviors in female mice only. Circadian rhythm, measured as 24HLM performance, appeared unaltered in female Brd1+/− mice (Fig. 1u) whereas sucrose preference was significantly reduced in female Brd1+/− mice compared to WT mice (Fig. 1v, two-way ANOVA; p = 0.001).
Neurochemistry and psychotropic drug-induced activity
As reported in male Brd1+/− mice (Table S1)35, female mice displayed unaltered hippocampal serotonin levels (Fig. 2a) and significantly reduced hippocampal dopamine levels (Fig. 2b, t-test; p = 0.045) but unaltered fronto-cortical dopamine levels (Fig. 2c). However, female mice had significantly less fronto-cortical serotonin (Fig. 2d, t-test; p = 0.01) and, noticeably, significantly reduced striatal dopamine (Fig. 2e, t-test; p = 0.02) compared to WT mice. Further analyses confirmed statistically significant interactions between female gender and Brd1+/− genotype on fronto-cortical serotonin (two-way ANOVA; p = 0.002) and striatal dopamine (two-way ANOVA; p = 0.002) levels. Additionally, their sensitivity towards the psychomotor stimulatory effects of amphetamine 5 mg/kg (Fig. 2f) and cocaine 15 or 30 mg/kg (Fig. 2g) did not differ from the sensitivity of WT mice.
a–e Neurotransmitter levels were determined by HPLC in several brain tissues in female Brd1+/− mice in parallel with male Brd1+/− mice as previously reported on ref. 35. a Female mice displayed unaltered hippocampal serotonin level (n = 9/group). b Significantly reduced hippocampal dopamine level (n = 10/group, t = 2.147, p = 0.045). c Unaltered fronto-cortical dopamine (n = 9/group). d Less fronto-cortical serotonin (n = 15/group; t = 2.70; p = 0.01) and; e reduced striatal dopamine (n = 10/group; t = 2.52; p = 0.02) compared to WT mice. 5-HT: 5-hydroxy tryptamine (serotonin); DA: dopamine. f Distance moved before and after amphetamine 5 mg/kg (Amph) injection was similar in female Brd1+/− and WT mice (n = 10/group). g Distance moved before and after cocaine 30 mg/kg injection was similar in female Brd1+/− and WT mice (n = 12/group). h Total brain volume was slightly reduced in female Brd1+/− mice compared to WT mice (n = 7/group; t = 2.31; p = 0.041). i Total dendritic length including apical and basal dendrites. aCC pyramidal neurons had significantly shorter dendrites in female Brd1+/− mice compared to WT mice (t = 3.29; p = 0.008). j Mean branch depth: branching depth was defined by the number of bifurcations from the beginning point to the end of a dendrite. Female Brd1+/− mice had less dendritic branching (t = 3.08; p = 0.01) compared to WT mice. k Mean dendritic spine density (number of spines/ length of dendrites). Female Brd1+/− mice had less dendritic spine density (t = 9.19; p < 0.001) compared to WT mice. l 3-D Sholl analysis: Number of dendritic intersections on concentric spheres (radius interval 20 µm) with their centres at soma. Neurons in female Brd1+/− mice had significantly less dendritic branching (F = 20.60; p < 0.001) than neurons in WT mice. m 3-D reconstruction of left: WT neuron and right: Brd1+/− neuron. Data shown are mean and SEM for each group *p < 0.05; **p < 0.01; ***p < 0.001.
Brain volume and neuronal morphology
Total brain volume, as estimated by stereology, was slightly reduced (~8%) in female Brd1+/− mice (Fig. 2h and Fig. S1A, t-test; p = 0.041), but with no difference in brain symmetry (Fig. S2B) or ventricle volume (Fig. S2C, D). In line with reduced overall brain tissue volume, aCC pyramidal neurons had significantly shorter dendrites in female Brd1+/− mice compared to WT mice (Fig. 2i, t-test; p = 0.008) combined with less dendritic branching (Fig. 2j, t-test; p = 0.01) and less dendritic spine density (Fig. 2k, t-test; p < 0.001). 3-D Sholl analysis counting the dendritic intersections on the concentric spheres with their centres at soma confirmed that these neurons had significantly less dendritic branching (Fig. 2l, m, p < 0.001).
Behavioral response to antidepressants
Provided that the phenomenological and pathophysiological phenotype of female Brd1+/− mice indicate translational relevance to depressive disorders, they may be reversible upon the administration of clinically used antidepressants. In accordance, treatment with Fluoxetine (FLX) and Imipramine (IMN) reversed the despair-like behaviors of female Brd1+/− mice, while having negligible effect on their movements in the OF (Fig. S3). Particularly, differences in immobility between Brd1+/− and WT mice during TST, could be reversed by IMN 1 (Fig. 3a) or 10 mg/kg (Fig. 3a), and by FLX 5 mg/kg (Fig. 3a) with Brd1+/− mice receiving IMN (Fig. 3b; 1 mg/kg, two-way ANOVA; p < 0.001 or 10 mg/kg, two-way ANOVA; p < 0.001) being significantly less immobile during TST than the vehicle-treated Brd1+/− mice. Similarly, the differences in immobility during FST, could be reversed by IMN 10 mg/kg (Fig. 3c) and by FLX 5 mg/kg (Fig. 3c), with IMN (Fig. 3d, two-way ANOVA; p < 0.001) or FLX (Fig. 3d, two-way ANOVA; p < 0.001) causing reduced immobility compared to vehicle-treatment. Vehicle-treated Brd1+/− mice were significantly more immobile in TST (Fig. 3a, t-test; p < 0.001) and FST (Fig. 3c, t-test; p = 0.005) than the vehicle-treated WT mice thus replicating the behavioral despair in untreated female Brd1+/− mice.
a, b Tail suspension test. a Total duration of immobility in the 8 groups (n = 15/group). Comparison between vehicle-treated WT and vehicle-treated Brd1+/− mice confirmed the depression-like behaviors of female Brd1+/− mice (t = 5.42; p < 0.001). Comparisons between other three pairs indicated that the depressive phenotype of female Brd1+/− mice could be reversed by antidepressant medications. b Duration of immobility in the four groups of female Brd1+/− mice, which received four different interventions. Female Brd1+/− mice, that received imipramine 1 (F = 17.55; p < 0.001) and 10 (F = 43.08; p < 0.001) mg/kg, displayed significantly less depression-like behaviors than the vehicle-treated Brd1+/− mice. c, d Forced swim test. c Total duration of immobility in the 8 groups (n = 15/group). Comparison between vehicle-treated WT mice and vehicle-treated Brd1+/− mice confirmed the depression-like behaviors of female Brd1+/− mice (t = 3.14; p = 0.005). Except for 1 mg/kg IMN treated groups, comparisons between other two pairs indicated that the depressive phenotype of female Brd1+/− mice could be reversed by antidepressant medications. d Duration of immobility in the four groups of female Brd1+/− mice, which received four different interventions. Female Brd1+/− mice that received imipramine 1 (F = 9.69; p = 0.004) and 10 (F = 35.71; p < 0.001) mg/kg, and FLX 5 mg/kg (F = 18.05; p < 0.001) displayed significantly less depression-like behaviors than the vehicle-treated Brd1+/− mice. IMN, imipramine; FLX, fluoxetine. Data shown are mean and SEM for each group. *p < 0.05; **p < 0.01; ***p < 0.001.
Global gene expression profiling of antidepressant treatment in Brd1 +/− mice
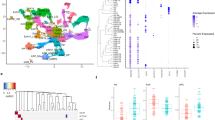
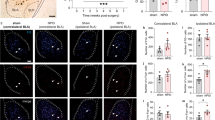
To delineate the molecular signatures accompanying the behavioral response to antidepressants by Brd1+/− mice, we conducted global gene expression profiling of selected brain tissues from mice administered vehicle or antidepressant (IMN or FLX). Amygdaloid (AMG) tissue from vehicle administered mice was characterized by pronounced changes in gene expression involving 144 differentially expressed genes (DEGs) that were significant after Benjamini–Hochberg false discovery rate (FDR) correction at 5% (Fig. 4a (mid panel) and Table S2). As these comprised a high number of predicted/uncharacterized genes (Table S2) we subjected nominally significant DEGs (p < 0.01, 511 genes) to functional analyses using ingenuity pathway analysis (IPA) software. This revealed an overrepresentation of genes acting in biological pathways previously implicated with affective behaviors, including unfolded protein response44 and α-adrenergic45-, chemokine46-, G-protein coupled receptor (GPCR), and Glial cell-derived neurotrophic factor (GDNF) mediated signaling47 (Fig. 4b and Table S3). Notably, fold change of implicated DEGs suggested an activation of G-protein coupled receptor (GPCR) mediated signaling, specifically through the Gaq subunit associated with, among others, the 5HT2 serotonergic-, and Alpha-1 adrenergic receptors (Fig. 4b and Table S3). Supportive of altered amygdaloid intracellular signaling in Brd1+/− mice, predicted upstream DEG regulators comprised a range of stimulus-induced transcription factors, including FOS, CREB1 and, ADORA2A (Table S4).
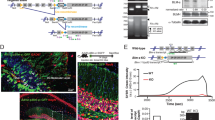
a Gene expression profile in AMG tissue from vehicle, IMN and FLX administered female WT (n = 8/group) and Brd1+/− (n = 8/group) mice. Gray, red, and blue color mark genes that are nominally significantly differentially expressed, significantly upregulated after FDR correction, and significantly downregulated after FDR correction between Brd1+/− and WT mice, respectively. b Functional genomic analysis using ingenuity pathway analysis IPA software identifies pathways that are enriched with nominally significantly AMG DEGs and predicts an activation score (z-score) based on the direction of regulation (up or down) of DEGs. Diameter of circles represent the overlap between DEGs and genes annotated to the respective pathways. c Principal component analysis of all genes expressed (CPM > 0.1) in each sample, cluster vehicle administered Brd1+/− mice separately from vehicle WT mice, and shows that treatment with IMN or FLX normalizes amygdaloid gene expression in Brd1+/− mice. d Heatmap showing the 144 genes surviving correction for multiple testing (FDR). DEGs are generally more abundantly expressed in vehicle administered Brd1+/− mice than in vehicle administered WT mice, whereas IMN or FLX both completely normalizes expression of this subset of genes. e Gene expression profile in aCC tissue from vehicle administered and IMN administered female WT (n = 8–10/group) and Brd1+/− (n = 8–10/group) mice. Gray color marks genes that are nominally significantly differentially expressed between Brd1+/− and WT mice. f Gene expression profile in CPu tissue from vehicle administered and IMN administered female WT (n = 8–10/group) and Brd1+/− (n = 8–10/group) mice. Gray, red, and blue color mark genes that are nominally significantly differentially expressed, significantly upregulated after FDR correction, and significantly downregulated after FDR correction between Brd1+/− and WT mice, respectively. g Functional genomic analysis using ingenuity pathway analysis IPA software identifies pathways that are enriched with nominally significantly CPu DEGs and predicts an activation score (z-score) based on the direction of regulation (up or down) of DEGs. Diameter of circles represent the overlap between DEGs and genes annotated to the respective pathways. h Principal component analysis of all genes expressed (CPM > 0.1) in each sample, cluster vehicle administered Brd1+/− mice separately from vehicle administered WT mice but shows that CPu treatment effect of IMN, in terms of gene expression changes, is largely on WT mice and not Brd1+/− mice.
Overall, administration of FLX and IMN normalized the amygdaloid transcriptome in Brd1+/− mice (Fig. 4c), and, convincingly, FLX and IMN administration nearly completely normalized AMG expression of genes differentially expressed in vehicle administered Brd1+/− mice (Fig. 4d and Table S6 and S6) without affecting Brd1 expression (Fig. 4a). Whereas the tested dose (10 mg/kg) of IMN had only minor effect on AMG gene expression in WT mice, administration of FLX (5 mg/kg) resulted in massive changes in WT mice (Fig. 4a). Accordingly, we proceeded with testing only IMN (10 mg/kg) in tissue samples from aCC and striatum, caudate putamen (CPu). In aCC, only Brd1 was significantly differentially regulated after FDR correction between Brd1+/− and WT vehicle groups (Fig. 4e, lower panel) and only 141 genes were differentially expressed at a nominal significant cut-off set at 1% (Table S7). This set of genes did not cluster in distinct biological pathways (Table S8). Like in AMG, however, administration of antidepressants primarily affected gene expression in Brd1+/−, and to a lesser extent, WT mice (Table S9). This included a reduction in mRNA encoding dopamine receptor 2 (Drd2, Fig. 4e and Table S9), which was significantly overexpressed in vehicle administered Brd1+/− mice compared to WT mice (Fig. 4e and Table S7). In CPu, only Brd1 was significantly downregulated after FDR correction in the vehicle treated group, (Fig. 4f, lower panel and Table S10) whereas a large number of genes (2260) were differentially expressed at a nominally significant cut-off set at 1% (Fig. 4f and Table S10). Although CPu and AMG DEGs generally clustered in the same functional pathways and involved same predicted upstream transcriptional regulators (Tables S2, S11, 12), contrary to AMG DEGs, CPu DEGs consistently reported of inhibited intracellular signaling initiated by GPCR subunits associated with a broad range of neurotransmitter receptors (Fig. 4g and Table S11). Although treatment with IMN overall normalized CPu gene expression between Brd1+/− and WT mice (Fig. 4f (Top panel)), effects were more pronounced in WT (Table S13, 1370 nominally significant DEGs) than in Brd1+/− mice (Table S14, 513 nominally significant DEGs) (Fig. 4h).
Discussion
Despite the clinical heterogeneity of affective disorders, their, etiopathologies are partially overlapping and include an intricate interplay between environmental, genetic, and epigenetic factors1,8,48,49,50. Post translational modifications of histones, such as Kac, have been linked with brain development7 and particularly the pathophysiology of affective disorders8,51. Acting as an epigenetic regulator during neurodevelopment, BRD1 has the potential to integrate intrinsic and environmental signals into the shaping of the maturing brain. Here, we demonstrate that reduced Brd1 expression in female mice results in brain morphometric alterations accompanied by changes in behaviors and underlying neurobiology with broad translational relevance to affective disorders. Supporting the predictive validity of female Brd1+/− mice as a model for depressive pathology, behavioral and molecular changes in Brd1+/− mice are reversible upon the administration of clinically effective antidepressants. Finally, integrative genomic profiling of regional brain transcriptomic effects reveals molecular mechanisms and tissue specificity associated with affective pathology and antidepressant treatment effect.
Neuro-implication of reduced Brd1 expression is sex-biased
Cognitive impairments are common in psychiatric disorders, and although more thoroughly investigated in male Brd1+/− mice36, both sexes display cognitive impairments with broad translational relevance, including central acquisition deficits and impaired reference memory. Sensorimotor deficits in the form of increased baseline startle amplitude, which have been reported in a range of psychiatric disorders52, are similarly seen in both male and female Brd1+/− mice. However, neither male nor female Brd1+/− mice display consistent changes in their risk-taking behaviors. Female Brd1+/− mice, additionally, did not exhibit marked changes in their circadian cycle as measured by 24HLM. However, supporting their translational value as model of depressive symptomatology seen in affective disorders53 and the prodromal stage of schizophrenia54, female Brd1+/− mice displayed increased immobility during FST and TST indicating behavioral despair55, and decreased sucrose preference representing anhedonia56.
Similar to what has been reported in both schizophrenia, bipolar disorder57, and depressed suicide victims58, female Brd1+/− mice display abnormal brain and neuronal morphology with reduced dendritic branching and spine pathology38. Although these parameters have not been assessed in male Brd1+/− mice, detailed structural brain imaging followed by stereological estimation of regional volumes and cell number, have revealed reduced subcortical volume and striatal cell loss in male Brd1+/− mice37. Sex differences in animal models of psychiatric disorders are, however, common and may mirror the documented sex differences in psychiatric disorders where symptom profiles and severity differ between sexes59,60,61,62,63,64 and where depressive disorders are more prevalent among women than men59,63. In line with the reported divergences in behaviors, the neurochemical profile of female Brd1+/− mice varied significantly from what we have previously reported in male Brd1+/− mice35,36. Unlike male Brd1+/− mice35, female Brd1+/− mice are not super-sensitive to the psycho-motor stimulatory effect of cocaine and PCP. Although both male and female mice display increased hippocampal dopamine, only female Brd1+/− mice displayed significantly reduced levels of cortical serotonin and striatal dopamine, consistent with the monoamine hypothesis of depression65. Histone modifications act to epigenetically sexually differentiate the developing brain and consequently behavior66 by regulating the genomic actions of sex steroid hormones66,67. Intriguing in this context, BRD1 is a co-regulator of nuclear hormone receptor-mediated signaling17,18,23 and its chromatin interactome enriched with estrogen and androgen target genes17.
Neuro-molecular effect of reduced Brd1 expression reflect brain regional changes in intracellular signaling activity
Using transcriptomic profiling to broadly capture Brd1-mediated neuro-molecular changes across multiple brain regions, we find that the dopamine receptor 2 (Drd2) and dopamine transporter (Slc6a3) are among the dysregulated genes in aCC and AMG, respectively (Fig. 4a, e). Monoamines act through activation of either G protein or ion channel linked surface receptors, which trigger second messenger systems (e.g. 3′–5′-cyclic adenosine monophosphate (cAMP), Rho family GTPases, inositol 1,4,5-trisphosphate (IP3) or calcium (Ca2+)). They, in turn, activate downstream kinases (e.g. protein kinase A (PKA)), which phosphorylate the transcription factor cAMP response element-binding protein (CREB) that regulates expression of many immediate early and late response genes. Convincingly, DEGs across examined tissues particularly cluster in GPCR-controlled intracellular signaling pathways and comprise immediate early and late response genes like neurotrophic factors (e.g. GDNF68 and NGF). Dysregulation of these signaling cascades are commonly reported in affective disorders69,70,71,72,73, reduced levels of phosphorylated CREB74 and neurotrophin receptors75 have been reported in the post-mortem frontal cortices of people with MDD and cortical IP3 has been suggested as a biomarker for depressive symptoms across diagnostic boundaries76. Interestingly, DEGs in the examined brain tissues reflect regional differences in intracellular signaling activity. Particularly, transcriptomic data from AMG tissue suggest increased Gaq mediated signaling, and thus increased PI3 activity, whereas this is the opposite in CPu tissue. CPu gene expression was further associated with reduced signaling mediated by Gas and Gai and their associated second messenger (cAMP).
The antagonistic effect of antidepressants on BRD1 mediated dysregulation is tissue specific
Supporting the predictive validity of Brd1+/− mice as a model for depressive psychopathology, administration of either of the two tested antidepressants, IMN and FLX, effectively alleviated the behavioral changes displayed by Brd1+/− mice in the TST and FST without affecting basal motor behaviors. This effect was further apparent at the molecular level, where IMN administration essentially normalized AMG gene expression without affecting the expression of Brd1. This was also the case for FLX, but at the selected dose, significant gene-regulatory changes were additionally seen in WT mice. In aCC, IMN treatment completely normalized Drd2 expression and Slc6a3, possibly reflecting a normalization of dopamine signaling in this tissue. Furthermore, expression of Slc25a35, which is a marker of mitochondrial dysfunction in brain regions under experimental mixed anxiety/depression-like disorder77, was additionally normalized following IMN administration (Fig. 4e). However, as basal gene expression, measured in the vehicle administered group, did not differ much between Brd1+/− and WT mice, the neuromolecular effect of antidepressant treatment was less apparent in this tissue. Surprisingly, in CPu, where large changes in basal gene expression were observed between WT and Brd1+/− mice, the effect IMN administration was much more pronounced in WT than in Brd1+/− mice. Similar findings have been reported from other rodent models with depressive-like phenotypes78,79.
Histone modification and chromatin modeling play a role in multiple physiological and pathological processes in the brain, including cognition80, circadian rhythms81, and the development of affective pathology8,51. Here we show that the schizophrenia and bipolar disorder associated epigenetic reader, BRD1, governs affective behaviors and associated neuromolecular and biological pathways in mice. In line with BRD1’s reported function as a co-regulator of nuclear hormone receptors17,18,23 and their targeted transcriptional response to gonadosteroid and corticosteroid signaling17, the effect of reduced Brd1 expression is sex-biased, with only female mice displaying changes in affective behaviors. BRD1 may thus provide an important link between averse environmental risk factors (.e.g. sex and stress) and depression. While equating Brd1 deficiency with depression susceptibility is over-simplistic, female Brd1+/− mice set the stage for further studies evaluating the epigenetic changes and neurodevelopmental abnormalities, pertinent to depression.
References
Lee, S. H. et al. Genetic relationship between five psychiatric disorders estimated from genome-wide SNPs. Nat. Genet 45, 984–994 (2013).
Plana-Ripoll, O. et al. Exploring comorbidity within mental disorders among a Danish national population. JAMA Psychiatry 76, 259 (2019).
Thapar A., Harold G., Rice F., Langley K. & O’Donovan M. The contribution of gene–environment interaction to psychopathology. Dev Psychopathol. 19, 989–1004 (2007).
Owen, M. J. & O’Donovan, M. C. Schizophrenia and the neurodevelopmental continuum:evidence from genomics. World Psychiatry 16, 227–235 (2017).
Anttila, V. et al. Analysis of shared heritability in common disorders of the brain. Science 360, eaap8757 (2018).
Uher, R. & Zwicker, A. Etiology in psychiatry: embracing the reality of poly-gene-environmental causation of mental illness. World Psychiatry 16, 121–129 (2017).
Bakulski, K. M., Halladay, A., Hu, V. W., Mill, J. & Fallin, M. D. Epigenetic research in neuropsychiatric disorders: the “Tissue Issue”. Curr. Behav. Neurosci. Rep. 3, 264–274 (2016).
Nestler, E. J., Pena, C. J., Kundakovic, M., Mitchell, A. & Akbarian, S. Epigenetic basis of mental illness. Neurosci 22, 447–463 (2016).
Réus, G. Z. et al. Ketamine and imipramine in the nucleus accumbens regulate histone deacetylation induced by maternal deprivation and are critical for associated behaviors. Behav. Brain Res. 256, 451–456 (2013).
Tsankova, N. M. et al. Sustained hippocampal chromatin regulation in a mouse model of depression and antidepressant action. Nat. Neurosci. 9, 519–525 (2006).
Onishchenko, N., Karpova, N., Sabri, F., Castrn, E. & Ceccatelli, S. Long-lasting depression-like behavior and epigenetic changes of BDNF gene expression induced by perinatal exposure to methylmercury. J. Neurochem 106, 1378–1387 (2008).
Fuchikami, M. et al. The potential use of histone deacetylase inhibitors in the treatment of depression. Prog. Neuropsychopharmacol. Biol. Psychiatry 64, 320–324 (2016).
Misztak, P., Pańczyszyn-Trzewik, P. & Sowa-Kućma, M. Histone deacetylases (HDACs) as therapeutic target for depressive disorders. Pharm. Rep. 70, 398–408 (2018).
Wu, Q. et al. A chemical toolbox for the study of bromodomains and epigenetic signaling. Nat. Commun. 10, 1–14. (2019).
Eskeland, R., Freyer, E., Leeb, M., Wutz, A. & Bickmore, W. A. Histone acetylation and the maintenance of chromatin compaction by polycomb repressive complexes. Cold Spring Harb. Symp. Quant. Biol. 75, 71–78 (2010).
Mishima, Y. et al. The Hbo1-Brd1/Brpf2 complex is responsible for global acetylation of H3K14 and required for fetal liver erythropoiesis. Blood 118, 2443–2453 (2011).
Fryland, T. et al. Identification of the BRD1 interaction network and its impact on mental disorder risk. Genome Med 8, 53 (2016).
McCullagh, P. et al. The cloning, mapping and expression of a novel gene, BRL, related to the AF10 leukaemia gene. Oncogene 18, 7442–7452 (1999).
Bjarkam, C. R. et al. Further immunohistochemical characterization of BRD1 a new susceptibility gene for schizophrenia and bipolar affective disorder. Brain Struct. Funct. 214, 37–47 (2009).
Christensen, J. H. et al. The schizophrenia and bipolar disorder associated BRD1 gene is regulated upon chronic restraint stress. Eur. Neuropsychopharmacol. 22, 651–656 (2012).
Fryland, T. et al. Electroconvulsive seizures regulates the Brd1 gene in the frontal cortex and hippocampus of the adult rat. Neurosci. Lett. 516, 110–113 (2012).
Kueh, A. J., Dixon, M. P., Voss, A. K. & Thomas, T. HBO1 is required for H3K14 acetylation and normal transcriptional activity during embryonic development. Mol. Cell Biol. 31, 845–860 (2011).
Cho, H. I., Kim, M. S. & Jang, Y. K. The BRPF2/BRD1-MOZ complex is involved in retinoic acid-induced differentiation of embryonic stem cells. Exp. Cell Res. 346, 30–39 (2016).
Severinsen, J. E. et al. Evidence implicating BRD1 with brain development and susceptibility to both schizophrenia and bipolar affective disorder. Mol. Psychiatry 11, 1126–1138 (2006).
Jorgensen, T. H. et al. Search for common haplotypes on chromosome 22q in patients with schizophrenia or bipolar disorder from the faroe islands. Am. J. Med Genet 114, 245–252 (2002).
Nyegaard, M. et al. Support of association between BRD1 and both schizophrenia and bipolar affective disorder. “Am. J. Med Genet Neuropsychiatr. Genet. 153B, 582–591 (2010).
Purcell, S. M. et al. Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 460, 748–752 (2009).
Aberg, K. et al. A comprehensive family-based replication study of schizophrenia genes. JAMA Psychiatry https://doi.org/10.1001/jamapsychiatry.2013.288 (2013).
Pardiñas, A. F. et al. Common schizophrenia alleles are enriched in mutation-intolerant genes and maintained by background selection. Nat. Genet. 50, 381–389 (2018).
Andreassen, O. A., Thompson, W. K. & Dale, A. M. Boosting the power of schizophrenia genetics by leveraging new statistical tools. Schizophr. Bull. 40, 13–17 (2014).
Stahl, E. A. et al. Genome-wide association study identifies 30 loci associated with bipolar disorder. Nat. Genet 51, 793–803 (2019).
Howard, D. M. et al. Genome-wide meta-analysis of depression identifies 102 independent variants and highlights the importance of the prefrontal brain regions. Nat. Neurosci. 22, 343–352 (2019).
Lek, M. et al. Analysis of protein-coding genetic variation in 60,706 humans. Nature 536, 285–291 (2016).
Purcell, S. M. et al. A polygenic burden of rare disruptive mutations in schizophrenia. Nature 506, 185–190 (2014).
Qvist, P. et al. The schizophrenia-associated BRD1 gene regulates behavior, neurotransmission, and expression of schizophrenia risk enriched gene sets in mice. Biol. Psychiatry 82, 62–76 (2017).
Qvist, P. et al. Mice heterozygous for an inactivated allele of the schizophrenia associated Brd1 gene display selective cognitive deficits with translational relevance to schizophrenia. Neurobiol. Learn Mem. 141, 44–52 (2017).
Qvist, P. et al. Brain volumetric alterations accompanied with loss of striatal medium-sized spiny neurons and cortical parvalbumin expressing interneurons in Brd1 +/− mice. Sci. Rep. 8, 1–12. (2018).
Paternoster, V. et al. Brain proteome changes in female Brd1 mice unmask dendritic spine pathology and show enrichment for schizophrenia risk. Neurobiol. Dis. 124, 479–488 (2019).
Festing, M. F. W. & Altman, D. G. Guidelines for the design and statistical analysis of experiments using laboratory animals. ILAR J. 43, 244–258 (2002).
Kim, D., Langmead, B. & Salzberg, S. L. HISAT: a fast spliced aligner with low memory requirements. Nat. Methods 12, 357–360 (2015).
Pertea, M. et al. StringTie enables improved reconstruction of a transcriptome from RNA-seq reads. Nat. Biotechnol. 33, 290–295 (2015).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Csomor, P. A. et al. On the influence of baseline startle reactivity on the indexation of prepulse inhibition. Behav. Neurosci. 122, 885–900 (2008).
Timberlake M.II, & Dwivedi Y. Linking unfolded protein response to inflammation and depression: potential pathologic and therapeutic implications. Mol. Psychiatry 24, 987–994 (2019).
Cottingham, C. & Wang, Q. α2 adrenergic receptor dysregulation in depressive disorders: Implications for the neurobiology of depression and antidepressant therapy. Neurosci. Biobehav Rev. 36, 2214–2225 (2012).
Leighton, S. P. et al. Chemokines in depression in health and in inflammatory illness: a systematic review and meta-analysis. Mol. Psychiatry 23, 48–58 (2018).
Tsybko, A. S., Ilchibaeva, T. V. & Popova, N. K. Role of glial cell line-derived neurotrophic factor in the pathogenesis and treatment of mood disorders. Rev. Neurosci. 28, 219–233 (2017).
Coleman, J. et al. The genetics of the mood disorder spectrum: genome-wide association analyses of over 185,000 cases and 439,000 controls. Biol Psychiatry 88, 169–184 (2020).
McGuffin, P. et al. The heritability of bipolar affective disorder and the genetic relationship to unipolar depression. Arch. Gen. Psychiatry 60, 497 (2003).
Kubota, T., Miyake, K. & Hirasawa, T. Epigenetic understanding of gene-environment interactions in psychiatric disorders: a new concept of clinical genetics. Clin. Epigenetics 4, 1 (2012).
Machado-Vieira, R., Ibrahim, L. & Zarate, C. A. Jr. Histone deacetylases and mood disorders: epigenetic programming in gene-environment interactions. CNS Neurosci. Ther. 17, 699–704 (2011).
Kohl, S., Heekeren, K., Klosterkötter, J. & Kuhn, J. Prepulse inhibition in psychiatric disorders—apart from schizophrenia. J. Psychiatr. Res. 47, 445–452 (2013).
Cryan, J. F. & Mombereau, C. In search of a depressed mouse: utility of models for studying depression-related behavior in genetically modified mice. Mol. Psychiatry 9, 326–357 (2004).
Larson, M. K., Walker, E. F. & Compton, M. T. Early signs, diagnosis and therapeutics of the prodromal phase of schizophrenia and related psychotic disorders. Expert Rev. Neurother. 10, 1347–1359 (2010).
Cryan, J. F., Mombereau, C. & Vassout, A. The tail suspension test as a model for assessing antidepressant activity: review of pharmacological and genetic studies in mice. Neurosci. Biobehav Rev. 29, 571–625 (2005).
Overstreet, D. H. Modeling depression in animal models. Methods Mol. Biol. 829, 125–144 (2012).
Konopaske, G. T., Lange, N., Coyle, J. T. & Benes, F. M. Prefrontal cortical dendritic spine pathology in schizophrenia and bipolar disorder. JAMA Psychiatry 71, 1323 (2014).
Hercher, C., Canetti, L., Turecki, G. & Mechawar, N. Anterior cingulate pyramidal neurons display altered dendritic branching in depressed suicides. J. Psychiatr. Res. 44, 286–293 (2010).
Diflorio, A. & Jones, I. Is sex important? Gender differences in bipolar disorder. Int Rev. Psychiatry 22, 437–452 (2010).
Zagni, E., Simoni, L. & Colombo, D. Sex and gender differences in central nervous system-related disorders. Neurosci. J. 2016, 1–13 (2016).
Canuso, C. M. & Pandina, G. Gender and schizophrenia. Psychopharmacol. Bull. 40, 178–190 (2007).
McLean, C. P., Asnaani, A., Litz, B. T. & Hofmann, S. G. Gender differences in anxiety disorders: prevalence, course of illness, comorbidity and burden of illness. J. Psychiatr. Res. 45, 1027–1035 (2011).
Kuehner, C. Why is depression more common among women than among men? Lancet Psychiatry 4, 146–158 (2017).
Riecher-Rössler, A. Oestrogens, prolactin, hypothalamic-pituitary-gonadal axis, and schizophrenic psychoses. Lancet Psychiatry 4, 63–72 (2017).
Krishnan, V. & Nestler, E. J. The molecular neurobiology of depression. Nature 455, 894–902 (2008).
Matsuda, K. I. et al. Histone deacetylation during brain development is essential for permanent masculinization of sexual behavior. Endocrinology 152, 2760–2767 (2011).
Murray, E. K., Hien, A., de Vries, G. J. & Forger, N. G. Epigenetic control of sexual differentiation of the bed nucleus of the stria terminalis. Endocrinology 150, 4241–4247 (2009).
Hisaoka, K., Maeda, N., Tsuchioka, M. & Takebayashi, M. Antidepressants induce acute CREB phosphorylation and CRE-mediated gene expression in glial cells: a possible contribution to GDNF production. Brain Res. 1196, 53–58 (2008).
Muly, C. Signal transduction abnormalities in schizophrenia: the cAMP system. Psychopharmacol. Bull. 36, 92–105 (2002).
Network and Pathway Analysis Subgroup of Psychiatric Genomics Consortium. Psychiatric genome-wide association study analyses implicate neuronal, immune and histone pathways. Nat. Neurosci. 18, 1861–1861 (2015).
Niciu, M. J., Ionescu, D. F., Mathews, D. C., Richards, E. M. & Zarate, C. A. Second messenger/signal transduction pathways in major mood disorders: moving from membrane to mechanism of action, part I: major depressive disorder. CNS Spectr. 18, 231–241 (2013).
Dwivedi, Y. & Pandey, G. N. Adenylyl cyclase-cyclicAMP signaling in mood disorders: role of the crucial phosphorylating enzyme protein kinase A. Neuropsychiatr. Dis. Treat. 4, 161–176 (2008).
Nurnberger, J. I. et al. Identification of pathways for bipolar disorder. JAMA Psychiatry 71, 657 (2014).
Yamada, S., Yamamoto, M., Ozawa, H., Riederer, P. & Saito, T. Reduced phosphorylation of cyclic AMP-responsive element binding protein in the postmortem orbitofrontal cortex of patients with major depressive disorder. J. Neural Transm. 110, 671–680 (2003).
Tripp, A. et al. Brain-derived neurotrophic factor signaling and subgenual anterior cingulate cortex dysfunction in major depressive disorder. Am. J. Psychiatry 169, 1194–1202 (2012).
Chiappelli, J. et al. Evaluation of myo-inositol as a potential biomarker for depression in schizophrenia. Neuropsychopharmacology 40, 2157–2164 (2015).
Babenko, V. N., Smagin, D. A., Galyamina, A. G., Kovalenko, I. L. & Kudryavtseva, N. N. Altered Slc25 family gene expression as markers of mitochondrial dysfunction in brain regions under experimental mixed anxiety/depression-like disorder. BMC Neurosci. 19, 79 (2018).
O’Leary, A., Kõiv, K., Raudkivi, K. & Harro, J. Antidepressants differentially affect striatal amphetamine-stimulated dopamine and serotonin release in rats with high and low novelty-oriented behaviour. Pharm. Res. 113, 739–746 (2016).
Ichikawa, J. & Meltzer, H. Y. Effect of antidepressants on striatal and accumbens extracellular dopamine levels. Eur. J. Pharm. 281, 255–261 (1995).
Guan, Z. et al. Integration of long-term-memory-related synaptic plasticity involves bidirectional regulation of gene expression and chromatin structure. Cell 111, 483–493 (2002).
Etchegaray, J.-P., Lee, C., Wade, P. A. & Reppert, S. M. Rhythmic histone acetylation underlies transcription in the mammalian circadian clock. Nature 421, 177–182 (2003).
Edgar, R., Domrachev, M. & Lash, A. E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30, 207–210 (2002).
Acknowledgements
We thank Tanja Stenshøj Østergaard for the genotyping of mice and Helene M. Andersen and Maj-Britt Lundorf for assisting in histology experiments. The study was supported by grants from The Danish Council for Independent Research-Medical Sciences (A.D.B. and J.H.C.), The Central Denmark Region (Region Midt) (J.H.C.), The Augustinus Foundation (J.H.C.), The Riisfort Foundation (J.H.C.), The Lundbeck Foundation (A.D.B.), The Faculty of Health Sciences, Aarhus University (A.P.R. and A.D.B.), and The Novo Nordisk Foundation (A.D.B. and J.H.C.). Centre for Stochastic Geometry and Advanced Bioimaging was supported by Villum Foundation (J.R.N.). Part of the data presented in this paper is displayed in a preprint available at BioRxiv (https://doi.org/10.1101/257170).
Author information
Authors and Affiliations
Contributions
A.P.R. and P.Q.: Project administration, methodology, validation, formal analysis, investigation, data curation, writing—original draft, writing—review and editing, visualization, supervision, project administration. M.D., G.W., J.H.C., and A.D.B.: methodology, resources, supervision, project administration, writing—review and editing. J.H.C. and A.D.B.: funding acquisition. A.D.B. and O.M.: conceptualization. J.G.D.: formal analysis, investigation, data curation, writing—original draft, writing—review & editing. M.N., Jon.P. and V.P.: resources, writing—review and editing. K.F., A.M., J.R.N., B.P., N.N., G.W., N.L., T.F., S.H.l.C., and Joh. P.: methodology, writing—review & editing.
Corresponding authors
Ethics declarations
Conflict of interest
Besides being employed by H. Lundbeck A/S, K.F., A.M., and M.D. declare no biomedical financial interests or potential conflicts of interest. A.P.R., P.Q., J.G.D., R.L., Jon.P., N.N., Gu.W., N.L., S.H.l.C., V.P., T.F., Joh.P., M.N., B.P., J.R.N., Gr.W., O.M., J.H.C., and A.D.B. report no biomedical financial interests or potential conflicts of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rajkumar, A.P., Qvist, P., Donskov, J.G. et al. Reduced Brd1 expression leads to reversible depression-like behaviors and gene-expression changes in female mice. Transl Psychiatry 10, 239 (2020). https://doi.org/10.1038/s41398-020-00914-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-020-00914-2
This article is cited by
-
The psychiatric risk gene BRD1 modulates mitochondrial bioenergetics by transcriptional regulation
Translational Psychiatry (2022)
-
Lithium therapy subdues neuroinflammation to maintain pyramidal cells arborization and rescues neurobehavioural impairments in ovariectomized rats
Molecular Neurobiology (2022)
-
Inactivation of the Schizophrenia-associated BRD1 gene in Brain Causes Failure-to-thrive, Seizure Susceptibility and Abnormal Histone H3 Acetylation and N-tail Clipping
Molecular Neurobiology (2021)