Abstract
Autism spectrum disorder (ASD) is a pervasive neurodevelopmental syndrome with a high human and economic burden. The pathophysiology of ASD is largely unclear, thus hampering development of pharmacological treatments for the core symptoms of the disorder. Abnormalities in glutamate and GABA signaling have been hypothesized to underlie ASD symptoms, and may form a therapeutic target, but it is not known whether these abnormalities are recapitulated in humans with ASD, as well as in rodent models of the disorder. We used translational proton magnetic resonance spectroscopy ([1H]MRS) to compare glutamate and GABA levels in adult humans with ASD and in a panel of six diverse rodent ASD models, encompassing genetic and environmental etiologies. [1H]MRS was performed in the striatum and the medial prefrontal cortex, of the humans, mice, and rats in order to allow for direct cross-species comparisons in specific cortical and subcortical brain regions implicated in ASD. In humans with ASD, glutamate concentration was reduced in the striatum and this was correlated with the severity of social symptoms. GABA levels were not altered in either brain region. The reduction in striatal glutamate was recapitulated in mice prenatally exposed to valproate, and in mice and rats carrying Nlgn3 mutations, but not in rodent ASD models with other etiologies. Our findings suggest that glutamate/GABA abnormalities in the corticostriatal circuitry may be a key pathological mechanism in ASD; and may be linked to alterations in the neuroligin–neurexin signaling complex.
Similar content being viewed by others
Introduction
Autism spectrum disorder (ASD) is a pervasive neurodevelopmental syndrome characterized by deficits in social reciprocity and communication, and by restricted interests and repetitive behaviors (DSM-5)1. ASD affects 1–2% of the population and is up to five times more common in males than in females2. The high-societal burden of ASD is exacerbated by the fact that there are no effective pharmacological treatments for the core symptoms of the disorder. This is principally because the etiology and pathophysiology of ASD are largely unclear, thus leading to a lack of therapeutic targets. Most ASD cases are idiopathic, i.e., they cannot be ascribed to any known etiological cause3, and while genetic and environmental risk factors accounting for some cases of ASD have been identified, the neural mechanisms involved remain uncertain.
However, emerging evidence suggests that abnormalities in the balance between excitatory (glutamate-mediated), and inhibitory (GABA-mediated) neurotransmission may be a common pathophysiological mechanism and hence a treatment target in ASD4,5,6. For instance, abnormalities in the expression of glutamate and GABA receptors have been observed in the postmortem brains of people with ASD7. In vivo, proton magnetic resonance spectroscopy ([1H]MRS) has revealed alterations in the levels of glutamate and glutamine in the cortex and basal ganglia of children, and in the basal ganglia in adults with ASD8,9,10,11,12,13,14,15,16. Moreover, reductions in GABA have been reported in several brain regions in children with ASD13,17,18,19,20. While these studies are consistent with and support the hypothesis of an excitation/inhibition (E/I) imbalance in ASD, their respective findings do not always concur in terms of the directionality of the imbalance, and the underlying biological mechanism(s) remain poorly understood7,8,9,10,11,12,13,14,15,16,17,18,19,20. Progress can be made, however, with the help of rodent models of ASD. These models encompass diverse etiologies including genetic, chromosomal, i.e., copy number variations (CNVs), and environmental insults21. This offers a powerful toolbox to explore the underpinnings of an E/I imbalance in ASD. Electrophysiological and biochemical investigations in animal models of ASD have revealed brain region- and model-specific imbalances in E/I function6,22,23,24,25,26, yet it has been difficult to relate these findings to human ASD due to the invasive nature of the techniques, e.g., brain slice recordings, that are typically used. In contrast, [1H]MRS is noninvasive and can be safely used in humans and in animals alike—thus allowing direct inter-species comparisons. Yet, comparable [1H]MRS investigations of glutamate and GABA in both humans and rodent models of ASD do not currently exist.
Therefore, in order to determine whether ASD is associated with abnormal levels of glutamate and GABA in vivo, and to explore its putative biological underpinnings, we conducted a translational [1H]MRS study in both humans with idiopathic ASD and a panel of rodent models of ASD. We selected six etiologically diverse models to cover a wide range of previously identified risk factors for non-syndromic ASD in humans, namely (1) prenatal exposure to valproate (valproic acid, VPA), an environmental risk factor for ASD27, (2) paternal duplication of the 15q11-13 chromosomal region, the most frequent ASD-associated CNV28,29,30,31, and (3) three monogenic models carrying mutations in synapse-related genes that have been linked with ASD in humans: Shank3 knockout (KO) mice26,32, and Nlgn3R451C knock-in (KI) mice and Nlgn3 KO rats, both of which with mutations in the same gene, thus permitting comparisons across two rodent species25,33. In addition, an inbred mouse strain, the BTBR T+tf/J mice34, with strong face validity but unknown genetics was used as a model of ASD of presumed polygenic origin. We performed [1H]MRS in the same brain regions across humans, mice, and rats in order to allow for direct cross-species comparisons. We chose to focus on two brain areas previously implicated in both ASD pathology and social behavior in individuals without ASD, namely the striatum and the medial prefrontal cortex (mPFC)6,11,35,36,37.
Materials and methods
Human participants
A total of 25 individuals with idiopathic ASD were recruited from the Behavioral Genetics Clinic at the Maudsley Hospital, a national referral service for the diagnosis of neurodevelopmental disorders. Control participants were recruited via local advertisements. Inclusion criteria for all human participants included: (1) full-scale IQ of ≥80; (2) not currently taking any psychoactive medication and no psychoactive medication within at least the past six weeks; (3) no history of epilepsy, head injury, brain disease or infection, or serious medical illness (by self-report); (4) no history of bipolar disorder, schizophrenia, or drug or alcohol dependency; (5) no contraindications for MRI scanning i.e., no metallic implants, pacemakers, or history of major surgery to the head or chest. Participants in the ASD group were required to meet International Classification of Disease-Revision 10 (ICD-10) research criteria for autism. All diagnoses were confirmed using the Autism Diagnostic Observation Schedule (ADOS-G) and where possible also with the Autism Diagnostic Interview-Revised. A summary of the recruited participants’ demographics is given in Table 1. The two groups did not differ in age, gender, or in either full-scale IQ, verbal IQ or performance IQ as assessed using the Wechsler Abbreviated Scale of Intelligence. Ethical approval for this study was provided by the Essex 2 National Research Ethics Committee (reference 04-Q0102/26). Full written informed consent according to the Declaration of Helsinki was obtained from all participants.
Human [1H]MRS: data acquisition and processing
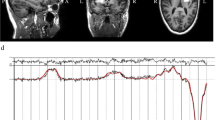
The striatum (including the head of the caudate, the anterior putamen, and the internal capsule) and the mPFC were selected as regions-of-interest (ROI) on the grounds that neuropathological and MRI studies have linked these brain areas to ASD6,11,35,36,37 and that we showed a reduction in Glx (combined pools of glutamate, glutamine, and glutathione) in striatum previously11. As practical considerations limited us to a single unilateral striatal ROI, we chose the left striatum in order to maintain maximum compatibility with our former study11. Moreover, earlier studies reported independency of metabolite levels with the ROI laterality38,39,40. For the cortical ROI, the bilateral mPFC including the cingulate gyrus was selected. Supplementary Figure 1a depicts examples of ROI placements.
[1H]MRS data were acquired on a 3T GE HDxt MRI scanner (GE Medical Systems, Milwaukee, WI, USA). A structural MRI scan for subsequent ROI positioning and tissue segmentation was acquired using a 3D fast inversion-recovery prepared gradient echo acquisition (inversion time (TI) = 450 ms, repetition time (TR) = 7 ms, echo time (TE) = 2.8 ms, matrix = 256 × 256 × 124 with a 0.9375 × 0.9375 × 1.1 mm3 voxel size). [1H]MRS spectra were acquired using Meshcher–Garwood Point-Resolved Spectroscopy (MEGAPRESS). MEGAPRESS is a widely used spectral editing technique that allows the quantification of GABA in humans at magnetic field strengths in which GABA is not readily detectable using plain point-resolved spectroscopy (PRESS) alone41. Excitation and refocusing pulses were shifted 2 ppm upfield from water, such that the voxel’s position was correct for metabolites lying in the center of the spectrum, and to minimize chemical shift displacements in the volume selection for the metabolite range of interest. Glutamate and glutamine were estimated from the “unedited” MEGAPRESS spectra (which is equivalent to a PRESS spectrum), as in the “edited” spectra coediting with GABA occurs42. GABA was quantified from the “difference” (edited–unedited) spectra. One ROI (35 × 30 × 25 mm) was placed in the left striatum. A second ROI (25 × 40 × 30 mm) was placed on the mPFC. Acquisition parameters were: TR = 2000 ms, TE = 68 ms and a total of 368 averages per ROI: 176 edit-on, 176 edit-off, and 16 with unsuppressed water for water scaling. [1H]MRS spectra were processed using LCModel software (version 6.3-0I; http://s-provencher.com/lcmodel.shtml). Water scaling was performed by LCModel, i.e., each metabolite peak was normalized by expressing its magnitude as a ratio against the magnitude of the unsuppressed water peak. Water-scaled estimates were comparable across ROIs and across participants and are expressed in institutional units as described previously11. Spectra were visually inspected, and LCModel quality control parameters were reviewed to verify that spectra were not qualitatively abnormal: all signal-to-noise ratios were ≥14 (striatum: 20.70 ± 3.35; mPFC: 21.10 ± 3.76) and all full widths at half maximums were ≤0.11 ppm (striatum: 0.07 ± 0.015 ppm; mPFC: 0.05 ± 0.022 ppm). Poorly fitted spectra with Cramer–Rao lower bounds equal to or higher than 20% of the metabolic estimate for either glutamate, glutamine or GABA, were excluded from further analysis; this led to the exclusion of one Glx and one GABA estimate from the mPFC ROI of a participant in the control group. The quality of acquired [1H]MRS spectra is shown in Supplementary Figure 1b–d.
To control for interindividual differences in ROI tissue composition, a scaling procedure was used, as previously described11. In brief, the structural MRI was first segmented into gray matter (GM), white matter (WM), and cerebrospinal fluid (CSF) using Statistical Parametric Mapping software (SPM2; http://spm.ion.ucl.ac.uk). For each spectroscopy ROI, its position was registered to the segmented structural images and the GM, WM, and CSF content of the ROI was calculated using in-house software. Absolute metabolite values were then calculated from the raw water-scaled LCModel (version 6.3-0I; http://s-provencher.com/lcmodel.shtml) outputs as follows:
where GM, WM, and CSF together sum up to 1. The (1.55 × CSF) term is included because CSF contains more water than GM or WM, and thus CSF tends to lower water-scaled estimates of the metabolites, in addition to the usual partial volume effects11. In the striatum, the groups did not differ on mean tissue contribution of GM, WM, or CSF as indicated by independent-samples unpaired two-tailed t-tests (all p-values >0.44) (controls: GM 52.5%, WM 43.3%, CSF 8.4%; ASD: GM 52.4%, WM 43.8%, and CSF 8.0%). In the mPFC, there was a significant group difference in GM contribution, which was higher in the ASD group (controls: 57.0%; ASD: 58.7%; p = 0.01; two-tailed t-test). The GM increase possibly reflects altered cortical neuroanatomy in the ASD group43. As brain structure is not the focus of the present study, however, we did not attempt to further characterize this difference but instead entered GM contribution percentage as a covariate in the group comparisons in the mPFC. No differences were seen in WM (controls: 25.8%; ASD: 24.3%; p = 0.19; two-tailed t-test) or CSF contribution (controls: 20.6%; ASD: 20.3%; p = 0.70; two-tailed t-test).
Rodent models
[1H]MRS investigations were carried out in five mouse models and one rat model of ASD. Mouse models included: mice prenatally exposed to VPA and age-matched saline-exposed controls (Harlan Laboratories, Horst, Netherlands; CD1 background), BTBR T+tf/J mice (The Jackson Laboratory, Bar Harbor, ME, USA) and age-matched wild-type C57BL/6J controls (The Jackson Laboratory, Bar Harbor, ME, USA), 15q11-13 patDP mice (kindly provided by T. Takumi, RIKEN, Wako, Japan; C57BL/6J genetic background), Shank3 KO mice (The Jackson Laboratory, Bar Harbor, ME, USA; C57BL/6J genetic background), Nlgn3R451C KI mice (The Rockefeller University, NY, USA; C57BL/6J genetic background), and respective wild-type littermates. In addition, Nlgn3 KO rats and their wild-type littermates (Horizon Discovery, Boyertown, PA, USA; Sprague-Dawley genetic background) were used. An overview of rodent models and their behavioral characteristics is given in Table 2. A total of 75 ASD-like animals and corresponding 73 controls were investigated in groups of 7–15 animals per condition (Table 2). The targeted sample size of ideally ≥12 animals per condition was based on previous in-house variability assessments44 and had to be reduced for logistical reasons in some cases. The animals were selected randomly from different litters. All animals were male adults aged 12–16 weeks (rodent equivalent of the age range of human study-participants)45 and weighing, depending on species and strain, 20–50 g (mice) and 350–500 g (rats) at the time of the study. Animals were group-housed (two to three per cage) in a temperature-, humidity-, and light-controlled environment (22–24 °C, 40–60%, 12 h light/dark cycle) and had access to food and water ad libitum. Animal experiments were approved by the Federal Food Safety and Veterinary Office of Switzerland (references BS-2359 and BS-2523) and conducted in strict adherence to the Swiss federal ordinance on animal protection and welfare, as well as according to the rules of the Association for Assessment and Accreditation of Laboratory Animal Care International.
Rodent [1H]MRS: data acquisition and processing
[1H]MRS in rodents was performed on a Biospec 9.4T/20 cm horizontal bore small animal MR scanner (Bruker, Ettlingen, Germany), equipped with an actively decoupled two-coil system consisting of a resonator for signal excitation, and a head surface coil for reception. For [1H]MRS, mice were initially anesthetized with 2.5–3% isoflurane (Abbott, Baar, Switzerland) in carrier gas composed of oxygen and air (1:5 v/v) supplied with a face mask to the freely breathing animal. Subsequently, etomidate (Etomidate-®Lipuro; B. Braun Melsungen AG, Melsungen, Germany) was administered as a primed continuous intravenous infusion according to the procedures and conditions reported previously45. Rats were anesthetized with isoflurane only: isoflurane concentration was adjusted between 2.0 and 2.3% to maintain stable respiratory rates of 60 breaths/min. Animals were placed in a custom-made cradle and their heads were immobilized in a stereotaxic frame. Respiratory rate, rectal body temperature, and CO2 level in the exhaled air were continuously monitored throughout the experiment as previously described46,47. Body temperature was maintained at 37 °C using a feedback-regulated electric heating blanket.
Anatomical reference images were obtained by rapid acquisition with refocused echoes (RARE)48 (TR = 1800 ms, TEeff = 36 ms, RARE-factor 8, matrix 256 × 256 over 20 × 20 mm and 40 × 40 mm for mice and rats, respectively). Two ROI homologous to those in humans were selected in the right striatum and the mPFC. The right striatum was chosen based on the purported lack of metabolite lateralization and to ensure compatibility with our previous studies44 and an extensive in-house database of rodent [1H]MRS data. The ROIs encompassed ~6 µl in mice (striatum: 2 × 1.8 × 1.8 mm; mPFC: 1.5 × 2.6 × 1.5 mm) and ~16 µl in rats (striatum and mPFC: 2.5 × 2.5 × 2.5 mm). Examples of ROI locations are shown in Supplementary Figure 2a, b. Local magnetic field homogeneity was adjusted for both ROIs separately by applying Fastmap49. [1H]MRS was carried out by PRESS, as the high magnetic field of 9.4T allows direct detection of GABA without using spectral editing techniques such as MEGAPRESS50. Acquisition parameters were: TR = 2000 ms and TE = 10 ms, 4 kHz spectral width sampled with 2048 complex data points and 512 averages collected over 17 min. The PRESS sequence was modified such that the frequencies of the excitation (1.35 ms hermite shape, 4 kHz bandwidth) and refocusing (1.2 ms hermite shape, 2.85 kHz bandwidth) pulses were shifted 2 ppm upfield from water such as to minimize chemical shift displacements in the volume selection for the metabolite range of interest. Water suppression was accomplished using a series of variable power radio frequency pulses on the water resonance51, interleaved with outer volume suppression with adiabatic pulses (1 mm gaps to ROI, 5 mm saturation slabs). An unsuppressed water spectrum was obtained immediately preceding each acquisition of a metabolite spectrum using identical acquisition parameters but only eight averages52. Quantitative analysis of the [1H]MRS data were carried out using LCModel software (version 6.3-0D) in a fully automated pipeline. The basis set comprised model spectra for 17 metabolites and eight macromolecular/lipid components. Absolute metabolite concentrations were determined with reference to unsuppressed water, uncorrected for relaxation. Spectral fits were reviewed visually to verify that spectra were qualitatively normal. The quality of acquired [1H]MRS spectra is shown in Supplementary Figure 2a,b. Predefined exclusion criteria comprised poor signal-to-noise ratio (five or lower) and head movement detected during data acquisition. Furthermore, glutamine estimates were excluded when they exceeded 3.5 mM (this value representing our empirical in-house threshold separating subjects with abnormally high-glutamine levels53 from subjects with normal glutamine levels). Only the latter criterion led to exclusion of data points, namely ten glutamine estimates from mPFC and striatum of five animals.
While there was no blinding with respect to the animals’ genotype/strain during data acquisition, processing, or statistical analysis, the entire pipeline was fully automated, and apart from visual inspection of the spectra there was no step at risk of introducing user bias.
Statistical analyses
Mean metabolite concentrations were compared pairwise between individuals with ASD and controls using two-tailed independent-samples t-tests with p < 0.05 as the significance criterion. The same approach was used to compare the mean metabolite concentrations between the animal model groups and their respective control groups. Tests were performed independently for each group, metabolite and ROI, yielding 44 comparisons (humans: eight; animals: 36) for neurometabolites of primary interest (glutamate, glutamine, Glx, and GABA) and 56 comparisons (humans: eight; animals: 48) for metabolites of secondary interest (creatine, choline, N-acetylaspartate + N-cetylaspartylglutamate (NAA + NAAG), and myo-inositol). To account for these multiple comparisons, the false discovery rate (FDR) was estimated within each of these two priority levels, based on the p-value distribution and its deviation from a uniform distribution, as previously described54,55. Comparisons between ASD cases or animal model groups and their respective control groups yielded 14 significant differences of neurotransmitters and 11 significant differences of further metabolites of secondary interest. Expected FDR levels were estimated to be 5% and 27%, respectively (i.e., expected numbers of false positives are around 1 and 3). Especially for the metabolites of primary interest, this number is reasonably low, while it seems acceptable for the other metabolites, given that the latter are only meant to provide auxiliary aspects of the disease endophenotype. Residuals per group, metabolite, and ROIs were tested for deviations from a normal distribution and for unequal variances between the conditions compared (i.e., ASD (model) vs. control). None of these tests passed FDR control across conditions, indicating no evidence of violation of t-test assumptions.
For the human study, correlations among metabolite concentrations and clinical measures were calculated using Pearson’s product moment coefficient. JMP 10 (SAS Institute, Inc., Cary, NC, USA) and GraphPad Prism 6 (GraphPad Software Inc., La Jolla, CA, USA) software were used for data analysis and preparation of graphs.
Results
[1H]MRS in individuals with ASD
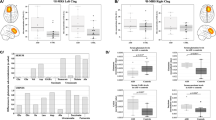
In this cohort of adult males with idiopathic ASD (Table 1), we observed a significant reduction of glutamate in the striatum as compared with healthy controls (Fig. 1a; Supplementary Table 1). Glx (combined pools of glutamate, glutamine and glutathione) was also decreased (controls: 5.80 ± 0.25; ASD: 4.92 ± 0.35; p = 0.04; two-tailed t-test; Supplementary Table 1). Yet, no differences were seen in glutamine (Fig. 1c; Supplementary Table 1) or in GABA (Fig. 1e; Supplementary Table 1) in this brain region. In the mPFC, there was no difference in glutamate (Fig. 1b; Supplementary Table 1), glutamine (Fig. 1d; Supplementary Table 1), Glx (controls: 6.77 ± 0.14; ASD: 6.93 ± 0.16; p = 0.47; two-tailed t-test; Supplementary Table 1), or GABA (Fig. 1f; Supplementary Table 1). For the concomitantly detected cerebral metabolites creatine, NAA + NAAG, and myo-inositol, no differences were observed in either of the two brain regions (Supplementary Table 2). Choline was increased in both striatum and mPFC of people with ASD (Supplementary Table 2).
Left panels: striatal levels (in institutional units) of a glutamate, c glutamine, and e GABA. Right panels: mPFC levels (in institutional units) of b glutamate, d glutamine, and f GABA. Data are depicted as mean ± SEM of n = 25–36 per group; *p < 0.05; two-tailed t-test. This difference remains significant with the removal of the putative outlier in the control group, p = 0.037. ASD autism spectrum disorder, i.u. institutional units, mPFC medial prefrontal cortex. For additional data see Supplementary Table 1
As striatal glutamate and Glx differed between groups, we examined whether these metabolite levels were correlated with current ASD symptom severity scores as measured using the ADOS, the current “gold standard” clinician-rated measure of core ASD behaviors. Striatal glutamate and Glx were both significantly, negatively correlated with ADOS social impairments: ASD individuals with lower (more abnormal) glutamate and Glx displayed worse social function (glutamate: r = −0.56, p = 0.005; Glx: r = −0.63, p = 0.002; both significant after Bonferroni correction for two ROIs × two ADOS scales). There was no correlation between striatal glutamate or Glx and the severity of repetitive behaviors (glutamate: r = −0.07, p = 0.77; Glx: r = −0.07, p = 0.77).
[1H]MRS in rodent models of ASD
In parallel to the human [1H]MRS studies, we evaluated the neurochemical differences in six rodent models of ASD with different underlying etiologies that emulate the breadth of potential risk factors in idiopathic ASD (Table 2). Across the models several neurochemical differences were observed.
A decrease in striatal glutamate was the most consistent finding, being observed in Nlgn3R451C KI mice, Nlgn3 KO rats, and in mice prenatally exposed to VPA (Fig. 2a; Supplementary Table 3). In Nlgn3R451C KI mice and Nlgn3 KO rats, a decrease in glutamate levels was also seen in the mPFC (Fig. 2b; Supplementary Table 3).
Left panels: concentrations (in mM) of a glutamate, c glutamine, and e GABA in striatum. Right panels: concentrations (in mM) of b glutamate, d glutamine, and f GABA in mPFC. All the mouse models (colored bars) were on a C57BL/6J background, except for VPA (CD1) and BTBR T+tf/J mice. The rat model (colored bars) was on a Sprague-Dawley background. Wild-type (gray bars) littermates were used as controls, except for VPA and BTBR T+tf/J mice for which age-matched saline-treated CD1 and wild-type C57BL/6J mice served as respective controls. Note the strain-dependent differences in glutamate levels. Data are depicted as mean ± SEM of n = 7–15 per group; *p < 0.05, **p < 0.01, ***p < 0.001; two-tailed t-test. KI knock-in, KO knockout, mPFC medial prefrontal cortex, patDp paternal duplication, VPA valproic acid. For additional data see Supplementary Table 2
Among the other models, different patterns of changes were seen. For instance, glutamate and glutamine were elevated in the striatum of BTBR T+tf/J mice (Fig. 2a, c; Supplementary Table 3). Glutamine was elevated in the striatum and mPFC of 15q11-13 paternal duplication (patDp/+; henceforth referred to as 15q11-13 patDp) mice, whereas reduced glutamine levels were observed in the mPFC of Nlgn3R451C KI and Shank3 KO mice (Fig. 2d; Supplementary Table 3). Our observation of no alterations in GABA in humans with ASD was recapitulated in most of the rodent models, including Nlgn3R451 KI mice, Nlgn3 KO rats, and VPA mice (Fig. 2e, f; Supplementary Table 3). Elevated GABA levels were observed only in the striatum of BTBR T+tf/J mice (Fig. 2e; Supplementary Table 3).
Several brain region- and model-specific changes were also noted in other commonly reported neurometabolites (Supplementary Table 4). Myo-inositol levels were altered in the striatum and mPFC of Nlgn3 KO rats and BTBR T+tf/J mice (Supplementary Table 4). In BTBR T+tf/J mice, creatine was increased in the striatum but strongly decreased in the mPFC (Supplementary Table 4). In contrast, in Nlgn3 KO rats creatine was elevated in the mPFC (Supplementary Table 4). Levels of NAA + NAAG and choline-containing metabolites were only altered in BTBR T+tf/J mice (Supplementary Table 4).
Discussion
In this translational [1H]MRS study, we examined glutamate and GABA levels in people with ASD and in six rodent models of the disorder. Consistent with the concept of a glutamate/GABA imbalance in ASD, we found that adults with idiopathic ASD have decreased glutamate concentration in the striatum compared to controls. The association between striatal glutamate and the severity of social impairment implies that this abnormality is clinically significant. In contrast to our findings with glutamate, we observed no alterations in GABA in adults with ASD. Further, we showed that reduced striatal glutamate is recapitulated in a subset of rodent models, namely in mice exposed prenatally to valproate, and in both mice and rats carrying mutations in the ASD-associated gene Nlgn3 although in these models mPFC glutamate was also reduced.
Our current human [1H]MRS results replicate and extend our previous findings of reduced Glx in the striatum, but not in the cortex, of people with ASD11. We are now also able to confirm that it is the excitatory neurotransmitter glutamate that is reduced in the striatum in ASD. This is because of the higher magnetic field strength of the 3T MRI scanner used in the present study vs. 1.5T in our previous work11, which allows for the separate quantification of glutamate and glutamine, and minor contributions of glutathione. The glutamate reduction is regionally specific, however, which may explain the differing findings of cortical glutamate across various studies in which [1H]MRS was performed in different cortical areas8,9,10,11,12,13,14,15,16. Brain region- and even neural circuit-specific changes in the E/I balance have also been observed in animal models of ASD6,22,23,24,25,26.
In the present study, we did not observe differences in GABA concentration in adults with ASD in either the striatum or in the mPFC. However, reductions in GABA have been observed in [1H]MRS studies of children with ASD13,17,18,19,20. Therefore, GABA abnormalities in ASD could be age-dependent. Future research should examine GABA longitudinally over the life course—both in humans and in rodent ASD models—in order to determine whether the maturational trajectory of the GABA system is altered in ASD.
We assessed six rodent models of ASD as part of this translational study, and showed that the abnormality seen in individuals with ASD—namely a striatal glutamate deficit—was recapitulated in VPA-exposed mice, Nlgn3R451C KI mice, and Nlgn3 KO rats. Our novel finding is that these rodent models of ASD are similar to each other with regard to this key neurochemical change, and that they recapitulate the change seen in human idiopathic ASD, thus pointing to a common pathophysiology. Literature on valproate and on Nlgn3 offers intriguing clues that support this notion of a common pathophysiology. Valproate is an anticonvulsant and antimanic agent with multiple direct pharmacological effects, such as enhancing GABA neurotransmission56. VPA is also known to modulate gene expression via inhibition of histone deacetylase56, and in particular, VPA has been shown to reduce expression of NLGN357, thus mimicking the effect of the genetic manipulation in Nlgn3R451C KI mice and Nlgn3 KO rats. Both point mutations58 and deletions59,60 of this gene have been identified in people with ASD, and are modeled by Nlgn3R451C KI mice and Nlgn3 KO rats, respectively.
The mechanism by which mutations in the postsynaptic cell-adhesion protein NLGN3 affects the striatal glutamate system is unclear. Nevertheless, its involvement in synapse function and the clustering of synaptic vesicles in the presynaptic buttons25,26,61,62 suggests that NLGN3 deficiency may result in a decreased number of glutamate-containing presynaptic vesicles. Unlike in humans with ASD, Nlgn3 mutant mice and rats also showed reduced glutamate in the mPFC, suggesting that the abnormalities were less regionally specific perhaps due to the pervasiveness of the genetic alterations in the brains of these models.
We also observed a tendency toward reduction in striatal glutamate and glutamine levels in the Shank3 KO mice. SHANK3, like NLGN3, is a synaptic protein implicated in ASD pathology26,63, and it also signals through the neuroligin–neurexin complex64. This implies both a pathophysiological convergence in SHANK3- and NLGN3-deficient rodents and a potential mechanism underpinning the regionally specific neurochemical abnormalities that we observed in humans.
Our finding of altered striatal glutamate might appear surprising, given that the striatum is primarily composed of GABAergic medium spiny neurons in which glutamate levels are very low65,66. However, glutamate is contained within glutamatergic inputs from the cerebral cortex, i.e., corticostriatal projections, in this region. Hence, the [1H]MRS-observable reduction in striatal glutamate in both people with ASD, and in particular animal models of the disorder, might reflect the presence of fewer (or less effective) corticostriatal projections. This interpretation is in line with reports that individuals with ASD have impaired long range connectivity67 and reduced neural activity in corticostriatal circuits in response to a variety of social rewards68,69. Thus our finding of a correlation between striatal glutamate levels and the severity of social deficits is consistent with the role of striatum in social behaviors36,37.
We observed reduced striatal glutamate in humans with ASD and more prevalently in several ASD rodent models. Based on observations that the MRS-amenable glutamate and GABA pools relate to neural function44, this result is opposing the original hypothesis of neural excitability in ASD put forward by Rubenstein and Merzenich5. This hypothesis of increased excitation was largely based on the observation that epilepsy frequently co-occurs with ASD5. However, the majority of ASD individuals do not have seizures70, and epilepsy is not simply a consequence of an increase in neuronal excitation71. Moreover, work on other animal models of ASD supports the idea of a shift in the E/I balance away from excitation24.
Our study has some limitations. While [1H]MRS offers the unique possibility of noninvasively measuring metabolite concentration across species, a restriction of the technique is that it measures the total concentration of metabolites, e.g., glutamate, within a selected ROI and cannot distinguish between synaptic, neuronal and glial pools. However, for glutamate, there is indication that the neurotransmitter and metabolic pools are correlated44,72, and glial and extracellular glutamate pools are several-folds smaller than the neuronal pool65,73,74. Therefore, glutamate concentration measured by [1H]MRS can be viewed as proportional to the neuronal neurotransmitter pool.
Regarding the limitations of the human [1H]MRS study, our sample size was modest. However, it was comparable with, or larger than, similar studies8,9,10,11,12,13,14,15,16,17,18,19,20 and our key finding of reduced striatal Glx in ASD replicates our previous study in a new cohort of participants11. We excluded subjects with a below-normal IQ or a history of epilepsy or seizures. This was done in order to ensure that our sample was broadly representative of the ASD population, since the majority of ASD cases do not have epilepsy70 and recent studies suggest that most have normal range IQ75. Nonetheless, many individuals with ASD do exhibit these features and therefore more research would be required to confirm whether our results hold in such cases. A further limitation is that we were unable to estimate true absolute concentrations in the human [1H]MRS. The estimates we calculated are comparable between individuals, but cannot be directly interpreted as concentrations (in mM) because of a number of assumptions made in the analysis of [1H]MRS data in LCModel. For instance metabolite estimates vary depending on how macromolecules are estimated and the method we used (estimating eight macromolecule peaks) has been scrutinized51. This, however, does not affect the validity of our between-group comparisons because the same analysis was used for all spectra.
A further, conceptual limitation is that our animal models (except BTBR T+tf/J mice) are based on specific etiological factors, and do not model idiopathic ASD as such. However, there are no animal models of idiopathic ASD in this sense, as even BTBR T+tf/J mice were reported to share some genetic features with clinical ASD76,77,78. Our hypothesis was that individuals with ASD and rodent models of ASD may share key pathophysiological mechanisms regardless of etiology79. In order to identify possible points of convergence that would guide future mechanistic investigations, we selected a range of animal models, while restricting our selection to “non-syndromic” ASD models as syndromic forms of ASD, associated with gross neuroanatomical, and morphological abnormalities, could represent unusual subtypes.
In the rodent study, our use of anesthesia to facilitate in vivo [1H]MRS constitutes a difference from our human studies, but the alternative of scanning awake animals, would also have introduced bias due to stress experienced by the rodents80. We have previously demonstrated reliability of [1H]MRS in detecting changes in glutamate and GABA in anesthetized animals44. We cannot completely exclude interactions of anesthesia with neurotransmitter levels, e.g., elicited by the anesthetics’ effect on GABAA receptors81. However, as we used the same anesthetic regime for all the animal models and their controls, it is unlikely that this could explain the differences in glutamate and GABA between rodent groups. Cross-species matching by age (or developmental and maturational stage) for a direct comparison of human and rodent data is complex. Notwithstanding, mice aged 3–4 months, as used in this study, have been reported to reflect a human equivalent of early adulthood45, thus broadly reflecting the human cohorts. Also, we did not perform [1H]MRS and behavioral assessments in the same animals to avoid potential confounds related to the series and sequence of experimental procedures. Previous work has shown that behavioral testing could induce changes in neurometabolite levels82. On the other hand, the imaging procedure with 1H of anesthesia could impact upon subsequent behavioral testing.
As practical considerations limited us to a single unilateral striatal ROI, we chose the left striatum in humans and the right striatum in our animal models in order to maintain maximum compatibility with our former studies11,44 and extensive in-house databases. Earlier studies38,39,40, as well as our separate [1H]MRS study performed in the left striatum of saline-treated C57BL/6J and BTBR T+tf/J mice reported levels for glutamate (C57BL/6J: 4.88 ± 0.08; BTBR T+tf/J: 5.61 ± 0.11), glutamine (C57BL/6J: 2.52 ± 0.08; BTBR T+tf/J: 2.96 ± 0.08), and GABA (C57BL/6J: 2.08 ± 0.05; BTBR T+tf/J: 2.42 ± 0.06) that closely mirrored those measured here in the right striatum of those two mouse strains, thus suggesting the absence of metabolite lateralization.
Finally, another technical difference between our human and animal studies was the difference in [1H]MRS magnetic field strength and pulse sequence used (3T MEGAPRESS vs. 9.4T PRESS) along with ROI size differences. However, both of these are methods optimized and qualified for the two species, and there is no a priori reason to expect them to differ in their ratiometric measurements.
In conclusion, in this first translational [1H]MRS study in ASD, we replicated and extended our previous finding that high-functioning adults with idiopathic ASD have reduced glutamate in the striatum11, while we detected no change in GABA. We further showed that the reduction in striatal glutamate is related to core ASD symptom severity. Moreover, reduction in striatal glutamate was recapitulated in rodents carrying mutations in Nlgn3 or being prenatally exposed to VPA. These translational findings support the notion of glutamatergic dysfunction in the corticostriatal pathway as an underlying core pathophysiological mechanism of ASD. However, contrary to the original hypothesis put forward by Rubenstein and Merzenich5 that ASD might be rooted in increased cortical excitability, our data on humans, as well as ours and recent work24 on several rodent models of ASD, are suggestive of a region-specific imbalance slanted toward reduced excitation.
References
American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders 5th edn (American Psychiatric Press, Arlington, TX, 2013).
Lai, M. C., Lombardo, M. V. & Baron-Cohen, S. Autism. Lancet 383, 896–910 (2014).
De Rubeis, S. & Buxbaum, J. D. Genetics and genomics of autism spectrum disorder: embracing complexity. Hum. Mol. Genet. 24, 24–31 (2015).
Coghlan, S. et al. GABA system dysfunction in autism and related disorders: from synapse to symptoms. Neurosci. Biobehav. Rev. 36, 2044–2055 (2012).
Rubenstein, J. L. & Merzenich, M. M. Model of autism: increased ratio of excitation/inhibition in key neural systems. Genes Brain Behav. 2, 255–267 (2003).
Yizhar, O. et al. Neocortical excitation/inhibition balance in information processing and social dysfunction. Nature 477, 171–178 (2011).
Blatt, G. J. & Fatemi, S. H. Alterations in GABAergic biomarkers in the autism brain: research findings and clinical implications. Anat. Rec. 294, 1646–1652 (2011).
Bejjani, A. et al. Elevated glutamatergic compounds in pregenual anterior cingulate in pediatric autism spectrum disorder demonstrated by 1H MRS and 1H MRSI. PLoS ONE 7, e38786 (2012).
Bernardi, S. et al. In vivo 1H-magnetic resonance spectroscopy study of the attentional networks in autism. Brain Res. 1380, 198–205 (2011).
Brown, M. S., Singel, D., Hepburn, S. & Rojas, D. C. Increased glutamate concentration in the auditory cortex of persons with autism and first-degree relatives: a (1)H-MRS study. Autism Res. 6, 1–10 (2013).
Horder, J. et al. Reduced subcortical glutamate/glutamine in adults with autism spectrum disorders: a [(1)H]MRS study. Transl. Psychiatry 3, e279 (2013).
Joshi, G. et al. Magnetic resonance spectroscopy study of the glutamatergic system in adolescent males with high-functioning autistic disorder: a pilot study at 4T. Eur. Arch. Psychiatry Clin. Neurosci. 263, 379–384 (2013).
Kubas, B. et al. Metabolite alterations in autistic children: a 1H MR spectroscopy study. Adv. Med. Sci. 57, 152–156 (2012).
Naaijen, J. et al. Fronto-striatal glutamate in children with Tourette’s disorder and attention-deficit/hyperactivity disorder. Neuroimage Clin. 13, 16–23 (2017).
Page, L. A. et al. In vivo 1H-magnetic resonance spectroscopy study of amygdala-hippocampal and parietal regions in autism. Am. J. Psychiatry 163, 2189–2192 (2006).
Tebartz van Elst, L. et al. Disturbed cingulate glutamate metabolism in adults with high-functioning autism spectrum disorder: evidence in support of the excitatory/inhibitory imbalance hypothesis. Mol. Psychiatry 19, 1314–1325 (2014).
Gaetz, W. et al. GABA estimation in the brains of children on the autism spectrum: measurement precision and regional cortical variation. Neuroimage 86, 1–9 (2014).
Harada, M. et al. Non-invasive evaluation of the GABAergic/glutamatergic system in autistic patients observed by MEGA-editing proton MR spectroscopy using a clinical 3 tesla instrument. J. Autism Dev. Disord. 41, 447–454 (2011).
Puts, N. A. J. et al. Reduced GABA and altered somatosensory function in children with autism spectrum disorder. Autism Res. 10, 608–619 (2017).
Rojas, D. C., Singel, D., Steinmetz, S., Hepburn, S. & Brown, M. S. Decreased left perisylvian GABA concentration in children with autism and unaffected siblings. Neuroimage 86, 28–34 (2014).
Chen, J. A., Penagarikano, O., Belgard, T. G., Swarup, V. & Geschwind, D. H. The emerging picture of autism spectrum disorder: genetics and pathology. Annu. Rev. Pathol. 10, 111–144 (2015).
Baudouin, S. J. et al. Shared synaptic pathophysiology in syndromic and nonsyndromic rodent models of autism. Science 338, 128–132 (2012).
Gandal, M. J. et al. GABAB-mediated rescue of altered excitatory-inhibitory balance, gamma synchrony and behavioral deficits following constitutive NMDAR-hypofunction. Transl. Psychiatry 2, e142 (2012).
Nelson, S. B. & Valakh, V. Excitatory/inhibitory balance and circuit homeostasis in autism spectrum disorders. Neuron 87, 684–698 (2015).
Tabuchi, K. et al. A neuroligin-3 mutation implicated in autism increases inhibitory synaptic transmission in mice. Science 318, 71–76 (2007).
Yoo, J., Bakes, J., Bradley, C., Collingridge, G. L. & Kaang, B. K. Shank mutant mice as an animal model of autism. Philos. Trans. R. Soc. Lond. B Biol. Sci. 369, 20130143 (2014).
Roullet, F. I., Lai, J. K. & Foster, J. A. In utero exposure to valproic acid and autism - a current review of clinical and animal studies. Neurotoxicol. Teratol. 36, 47–56 (2013).
Bolton, P. F. et al. Chromosome 15q11-13 abnormalities and other medical conditions in individuals with autism spectrum disorders. Psychiatr. Genet. 14, 131–137 (2004).
Nakatani, J. et al. Abnormal behavior in a chromosome-engineered mouse model for human 15q11-13 duplication seen in autism. Cell 137, 1235–1246 (2009).
Urraca, N. et al. The interstitial duplication 15q11.2-q13 syndrome includes autism, mild facial anomalies and a characteristic EEG signature. Autism Res. 6, 268–279 (2013).
Veltman, M. W. et al. A paternally inherited duplication in the Prader-Willi/Angelman syndrome critical region: a case and family study. J. Autism Dev. Disord. 35, 117–127 (2005).
Peca, J. et al. Shank3 mutant mice display autistic-like behaviours and striatal dysfunction. Nature 472, 437–442 (2011).
Hamilton, S. M. et al. Fmr1 and Nlgn3 knockout rats: novel tools for investigating autism spectrum disorders. Behav. Neurosci. 128, 103–109 (2014).
Meyza, K. Z. et al. The BTBR T(+)tf/J mouse model for autism spectrum disorders-in search of biomarkers. Behav. Brain Res. 251, 25–34 (2012).
Amaral, D. G., Schumann, C. M. & Nordahl, C. W. Neuroanatomy of autism. Trends Neurosci. 31, 137–145 (2008).
Baez-Mendoza, R. & Schultz, W. The role of the striatum in social behavior. Front. Neurosci. 7, 233 (2013).
Shepherd, G. M. Corticostriatal connectivity and its role in disease. Nat. Rev. Neurosci. 14, 278–291 (2013).
Sekine, Y. et al. Metabolite alterations in basal ganglia associated with methamphetamine-related psychiatric symptoms. A proton MRS study. Neuropsychopharmacology 27, 453–461 (2002).
Quetscher, C. et al. Striatal GABA-MRS predicts response inhibition performance and its cortical electrophysiological correlates. Brain Struct. Funct. 220, 3555–3564 (2015).
Zahr, N. M. et al. In vivo glutamate measured with magnetic resonance spectroscopy: behavioral correlates in aging. Neurobiol. Aging 34, 1265–1276 (2013).
Mullins, P. G. et al. Current practice in the use of MEGA-PRESS spectroscopy for the detection of GABA. Neuroimage 86, 43–52 (2014).
Abdallah, C. G. et al. Glutamate metabolism in major depressive disorder. Am. J. Psychiatry 171, 1320–1327 (2014).
Raznahan, A. et al. Cortical anatomy in autism spectrum disorder: an in vivo MRI study on the effect of age. Cereb. Cortex 20, 1332–1340 (2010).
Waschkies, C. F. et al. Neuropharmacological and neurobiological relevance of in vivo (1)H-MRS of GABA and glutamate for preclinical drug discovery in mental disorders. Neuropsychopharmacology 39, 2331–2339 (2014).
Flurkey K., Currer J. M., Harrison D. E. Mouse models in aging research. In Fox J. G., Davisson M. T., Quimby F. W., Barthold S. W., Newcomer C. E., Smith A. L. (eds) The Mouse in Biomedical Research 2nd edn, p. 637–672 (Elsevier, Burlington, VT, 2007).
Petrinovic, M. M. et al. A novel anesthesia regime enables neurofunctional studies and imaging genetics across mouse strains. Sci. Rep. 6, 24523 (2016).
Bruns, A., Kunnecke, B., Risterucci, C., Moreau, J. L. & von Kienlin, M. Validation of cerebral blood perfusion imaging as a modality for quantitative pharmacological MRI in rats. Magn. Reson. Med. 61, 1451–1458 (2009).
Hennig, J., Nauerth, A. & Friedburg, H. RARE imaging: a fast imaging method for clinical MR. Magn. Reson. Med. 3, 823–833 (1986).
Gruetter, R. Automatic, localized in vivo adjustment of all first- and second-order shim coils. Magn. Reson. Med. 29, 804–811 (1993).
Bottomley, P. A. Spatial localization in NMR spectroscopy in vivo. Ann. N. Y. Acad. Sci. 508, 333–348 (1987).
Tkac, I., Starcuk, Z., Choi, I. Y. & Gruetter, R. In vivo 1H NMR spectroscopy of rat brain at 1 ms echo time. Magn. Reson. Med. 41, 649–656 (1999).
Thulborn, K. R. & Ackerman, J. J. H. Absolute molar concentrations by NMR in inhomogeneous B1. A scheme for analysis of in vivo metabolites. J. Magn. Reson. 55, 357–371 (1983).
Cudalbu, C. et al. The C57BL/6J mouse exhibits sporadic congenital portosystemic shunts. PLoS ONE 8, e69782 (2013).
Storey, J. D. A direct approach to false discovery rates. J. R. Stat. Soc. B. 64, 479–498 (2002).
Storey, J. D. & Tibshirani, R. Statistical significance for genomewide studies. Proc. Natl Acad. Sci. USA 100, 9440–9445 (2003).
Rosenberg, G. The mechanisms of action of valproate in neuropsychiatric disorders: can we see the forest for the trees? Cell. Mol. Life Sci. 64, 2090–2103 (2007).
Kolozsi, E., Mackenzie, R. N., Roullet, F. I., deCatanzaro, D. & Foster, J. A. Prenatal exposure to valproic acid leads to reduced expression of synaptic adhesion molecule neuroligin 3 in mice. Neuroscience 163, 1201–1210 (2009).
Jamain, S. et al. Mutations of the X-linked genes encoding neuroligins NLGN3 and NLGN4 are associated with autism. Nat. Genet. 34, 27–29 (2003).
Gilman, S. R. et al. Rare de novo variants associated with autism implicate a large functional network of genes involved in formation and function of synapses. Neuron 70, 898–907 (2011).
Sanders, S. J. et al. Multiple recurrent de novo CNVs, including duplications of the 7q11.23 Williams syndrome region, are strongly associated with autism. Neuron 70, 863–885 (2011).
Sudhof, T. C. Neuroligins and neurexins link synaptic function to cognitive disease. Nature 455, 903–911 (2008).
Patel, S., Roncaglia, P. & Lovering, R. C. Using Gene Ontology to describe the role of the neurexin-neuroligin-SHANK complex in human, mouse and rat and its relevance to autism. BMC Bioinformatics 16, 186 (2015).
Penagarikano, O. et al. Absence of CNTNAP2 leads to epilepsy, neuronal migration abnormalities, and core autism-related deficits. Cell 147, 235–246 (2011).
Arons, M. H. et al. Autism-associated mutations in ProSAP2/Shank3 impair synaptic transmission and neurexin-neuroligin-mediated transsynaptic signaling. J. Neurosci. 32, 14966–14978 (2012).
Ottersen, O. P., Zhang, N. & Walberg, F. Metabolic compartmentation of glutamate and glutamine: morphological evidence obtained by quantitative immunocytochemistry in rat cerebellum. Neuroscience 46, 519–534 (1992).
Storm-Mathisen, J. et al. First visualization of glutamate and GABA in neurones by immunocytochemistry. Nature 301, 517–520 (1983).
Rane, P. et al. Connectivity in autism: a review of MRI connectivity studies. Harv. Rev. Psychiatry 23, 223–244 (2015).
Delmonte, S. et al. Social and monetary reward processing in autism spectrum disorders. Mol. Autism 3, 7 (2012).
Kohls, G. et al. Reward system dysfunction in autism spectrum disorders. Soc. Cogn. Affect. Neurosci. 8, 565–572 (2013).
Bolton, P. F. et al. Epilepsy in autism: features and correlates. Br. J. Psychiatry 198, 289–294 (2011).
Engel, J. Jr. Excitation and inhibition in epilepsy. Can. J. Neurol. Sci. 23, 16–174 (1996).
Stagg, C. J. et al. Relationship between physiological measures of excitability and levels of glutamate and GABA in the human motor cortex. J. Physiol. 589, 5845–5855 (2011).
Danbolt, N. C. Glutamate uptake. Prog. Neurobiol. 65, 1–105 (2011).
Yudkoff, M. et al. Brain glutamate metabolism: neuronal-astroglial relationships. Dev. Neurosci. 15, 343–350 (1993).
Elsabbagh, M. et al. Global prevalence of autism and other pervasive developmental disorders. Autism Res. 5, 160–179 (2012).
Erickson, C. A. et al. Initial analysis of peripheral lymphocytic extracellular signal related kinase activation in autism. J. Psychiatr. Res. 84, 153–160 (2017).
Faridar, A. et al. Mapk/Erk activation in an animal model of social deficits shows a possible link to autism. Mol. Autism 5, 57 (2014).
Weng, N., Weiler, I. J., Sumis, A., Berry-Kravis, E. & Greenough, W. T. Early-phase ERK activation as a biomarker for metabolic status in fragile X syndrome. Am. J. Med. Genet. B Neuropsychiatr. Genet. 147B, 1253–1257 (2008).
Ghosh, A., Michalon, A., Lindemann, L., Fontoura, P. & Santarelli, L. Drug discovery for autism spectrum disorder: challenges and opportunities. Nat. Rev. Drug Discov. 12, 777–790 (2013).
Haensel, J. X., Spain, A. & Martin, C. A systematic review of physiological methods in rodent pharmacological MRI studies. Psychopharmacology 232, 489–499 (2014).
Garcia, P. S., Kolesky, S. E. & Jenkins, A. General anesthetic actions on GABA(A) receptors. Curr. Neuropharmacol. 8, 2–9 (2010).
Zhou, I. Y., Ding, A. Y., Li, Q., McAlonan, G. M. & Wu, E. X. Magnetic resonance spectroscopy reveals N-acetylaspartate reduction in hippocampus and cingulate cortex after fear conditioning. Psychiatry Res. 204, 178–183 (2012).
Acknowledgements
The research leading to these results has received support from the Innovative Medicines Initiative Joint Undertaking under grant agreement no. 115300, resources of which are composed of financial contribution from the European Union’s Seventh Framework Programme (FP7/2007-2013) and EFPIA companies’ in kind contribution. M.M.P. was supported by the Roche Postdoctoral Fellowship program. We thank S. Coghlan, D. De La Harpe Golden, L. Brennan, I. Dud, D. Spain, L. Ajram, R. Edden, and R. Turra (formerly O’Gorman) for invaluable assistance with the human [1H]MRS study. We thank S. Schöppenthau, S. Debilly, and C. Wälti for their excellent technical support and data acquisition, and T. Bielser for data management and processing. We are grateful to L. Ozmen and her team for maintaining animal colonies.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
M.M.P., A.B., W.S., and B.K. received salaries for employment with F. Hoffmann-La. The remaining authors declare that they have no conflict of interest.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Horder, J., Petrinovic, M.M., Mendez, M.A. et al. Glutamate and GABA in autism spectrum disorder—a translational magnetic resonance spectroscopy study in man and rodent models. Transl Psychiatry 8, 106 (2018). https://doi.org/10.1038/s41398-018-0155-1
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41398-018-0155-1
This article is cited by
-
Amygdalar neurotransmission alterations in the BTBR mice model of idiopathic autism
Translational Psychiatry (2024)
-
The Lack of Alterations in Metabolites in the Medial Prefrontal Cortex and Amygdala, but Their Associations with Autistic Traits, Empathy, and Personality Traits in Adults with Autism Spectrum Disorder: A Preliminary Study
Journal of Autism and Developmental Disorders (2024)
-
Altered behavior, brain structure, and neurometabolites in a rat model of autism-specific maternal autoantibody exposure
Molecular Psychiatry (2023)
-
Neurometabolite levels in the brains of patients with autism spectrum disorders: A meta-analysis of proton magnetic resonance spectroscopy studies (N = 1501)
Molecular Psychiatry (2023)
-
Gamma-Aminobutyric Acid Type A Receptor Variants are Associated with Autism Spectrum Disorders
Journal of Molecular Neuroscience (2023)