Abstract
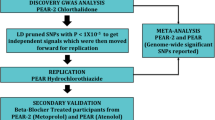
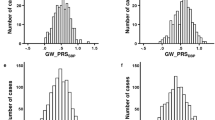
Resistant hypertension (RHTN), defined as uncontrolled blood pressure (BP) ≥ 140/90 using three or more drugs or controlled BP (<140/90) using four or more drugs, is associated with adverse outcomes, including decline in kidney function. We conducted a genome-wide association analysis in 1194 White and Hispanic participants with hypertension and coronary artery disease from the INternational VErapamil-SR Trandolapril STudy—GENEtic Substudy (INVEST-GENES). Top variants associated with RHTN at p < 10−4 were tested for replication in 585 White and Hispanic participants with hypertension and subcortical strokes from the Secondary Prevention of Subcortical Strokes GENEtic Substudy (SPS3-GENES). A genetic risk score for RHTN was created by summing the risk alleles of replicated RHTN signals. rs11749255 in MSX2 was associated with RHTN in INVEST (odds ratio (OR) (95% CI) = 1.50 (1.2–1.8), p = 7.3 × 10−5) and replicated in SPS3 (OR = 2.0 (1.4–2.8), p = 4.3 × 10−5), with genome-wide significance in meta-analysis (OR = 1.60 (1.3–1.9), p = 3.8 × 10−8). Other replicated signals were in IFLTD1 and PTPRD. IFLTD1 rs6487504 was associated with RHTN in INVEST (OR = 1.90 (1.4–2.5), p = 1.1 × 10−5) and SPS3 (OR = 1.70 (1.2–2.5), p = 4 × 10−3). PTPRD rs324498, a previously reported RHTN signal, was among the top signals in INVEST (OR = 1.60 (1.3–2.0), p = 3.4 × 10−5) and replicated in SPS3 (OR = 1.60 (1.1–2.4), one-sided p = 0.005). Participants with the highest number of risk alleles were at increased risk of RHTN compared to participants with a lower number (p-trend = 1.8 × 10−15). Overall, we identified and replicated associations with RHTN in the MSX2, IFLTD1, and PTPRD regions, and combined these associations to create a genetic risk score.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 print issues and online access
$259.00 per year
only $43.17 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Egan BM, Zhao Y, Axon RN. US trends in prevalence, awareness, treatment, and control of hypertension, 1988-2008. JAMA. 2010;303:2043–50.
Calhoun DA, Jones D, Textor S, Goff DC, Murphy TP, Toto RD, et al. Resistant hypertension: diagnosis, evaluation, and treatment: a scientific statement from the American Heart Association Professional Education Committee of the Council for High Blood Pressure Research. Circulation. 2008;117:e510–26.
Persell SD. Prevalence of resistant hypertension in the United States, 2003-2008. Hypertension. 2011;57:1076–80.
Sarafidis PA, Georgianos P, Bakris GL. Resistant hypertension—its identification and epidemiology. Nat Rev Nephrol. 2013;9:51–8.
Whelton PK, Carey RM, Aronow WS, Casey DE, Collins KJ, Dennison Himmelfarb C, et al. 2017 ACC/AHA/AAPA/ABC/ACPM/AGS/APhA/ASH/ASPC/NMA/PCNA guideline for the prevention, detection, evaluation, and management of high blood pressure in adults: a report of the American College of Cardiology/American Heart Association Task Force on Clinical Practice Guidelines. Hypertension. 2018;71:1269–324.
Daugherty SL, Powers JD, Magid DJ, Tavel HM, Masoudi FA, Margolis KL, et al. Incidence and prognosis of resistant hypertension in hypertensive patients. Circulation. 2012;125:1635–42.
Smith SM, Gong Y, Handberg E, Messerli FH, Bakris GL, Ahmed A, et al. Predictors and outcomes of resistant hypertension among patients with coronary artery disease and hypertension. J Hypertens. 2014;32:635–43.
El Rouby N, Cooper-DeHoff RM. Genetics of resistant hypertension: a novel pharmacogenomics phenotype. Curr Hypertens Rep. 2015;17:583.
Pepine CJ, Handberg EM, Cooper-DeHoff RM, Marks RG, Kowey P, Messerli FH, et al. A calcium antagonist vs a non-calcium antagonist hypertension treatment strategy for patients with coronary artery disease. The International Verapamil-Trandolapril Study (INVEST): a randomized controlled trial. JAMA. 2003;290:2805–16.
Benavente OR, White CL, Pearce L, Pergola P, Roldan A, Benavente MF, et al. The secondary prevention of small subcortical strokes (SPS3) study. Int J Stroke. 2011;6:164–75.
Dumitrescu L, Ritchie MD, Denny JC, El Rouby NM, McDonough CW, Bradford Y, et al. Genome-wide study of resistant hypertension identified from electronic health records. PLoS ONE. 2017;12:e0171745.
Fontana V, McDonough CW, Gong Y, El Rouby NM, Sa AC, Taylor KD, et al. Large-scale gene-centric analysis identifies polymorphisms for resistant hypertension. J Am Heart Assoc. 2014;3:e001398.
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MA, Bender D, et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet. 2007;81:559–75.
Willer CJ, Li Y, Abecasis GR. METAL: fast and efficient meta-analysis of genomewide association scans. Bioinformatics. 2010;26:2190–1.
DIAbetes Genetics Replication And Meta-analysis (DIAGRAM) Consortium, Asian Genetic Epidemiology Network Type 2 Diabetes (AGEN-T2D) Consortium; South Asian Type 2 Diabetes (SAT2D) Consortium, Mexican American Type 2 Diabetes (MAT2D) Consortium, Type 2 Diabetes Genetic Exploration by Nex-generation sequencing in muylti-Ethnic Samples (T2D-GENES) Consortium, et al. Genome-wide trans-ancestry meta-analysis provides insight into the genetic architecture of type 2 diabetes susceptibility. Nat Genet. 2014;46:234–44.
Gorlov IP, Moore JH, Peng B, Jin JL, Gorlova OY, Amos CI. SNP characteristics predict replication success in association studies. Hum Genet. 2014;133:1477–86.
Hou L, Zhao H. A review of post-GWAS prioritization approaches. Front Genet. 2013;4:280.
Barrett JC, Hansoul S, Nicolae DL, Cho JH, Duerr RH, Rioux JD, et al. Genome-wide association defines more than 30 distinct susceptibility loci for Crohn’s disease. Nat Genet. 2008;40:955–62.
Lee TH, Ko TM, Chen CH, Chang YJ, Lu LS, Chang CH, et al. A genome-wide association study links small-vessel ischemic stroke to autophagy. Sci Rep. 2017;7:15229.
McDonough CW, Gong Y, Padmanabhan S, Burkley B, Langaee TY, Melander O, et al. Pharmacogenomic association of nonsynonymous SNPs in SIGLEC12, A1BG, and the selectin region and cardiovascular outcomes. Hypertension. 201362:48–54.
Ward LD, Kellis M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 2012;40(Database issue):D930–4.
Boyle AP, Hong EL, Hariharan M, Cheng Y, Schaub MA, Kasowski M, et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. 2012;22:1790–7.
Cohen MM Jr. Craniofacial disorders caused by mutations in homeobox genes MSX1 and MSX2. J Craniofac Genet Dev Biol. 2000;20:19–25.
Towler DA, Bidder M, Latifi T, Coleman T, Semenkovich CF. Diet-induced diabetes activates an osteogenic gene regulatory program in the aortas of low density lipoprotein receptor-deficient mice. J Biol Chem. 1998;273:30427–34.
Shao JS, Cheng SL, Pingsterhaus JM, Charlton-Kachigian N, Loewy AP, Towler DA. Msx2 promotes cardiovascular calcification by activating paracrine Wnt signals. J Clin Invest. 2005;115:1210–20.
Fox CS, Heard-Costa N, Cupples LA, Dupuis J, Vasan RS, Atwood LD. Genome-wide association to body mass index and waist circumference: the Framingham Heart Study 100K project. BMC Med Genet. 2007;8:S18.
Levy D, Larson MG, Benjamin EJ, Newton-Cheh C, Wang TJ, Hwang SJ, et al. Framingham Heart Study 100K Project: genome-wide associations for blood pressure and arterial stiffness. BMC Med Genet. 2007;8:S3.
Gong Y, McDonough CW, Beitelshees AL, El Rouby N, Hiltunen TP, O’Connell JR, et al. PTPRD gene associated with blood pressure response to atenolol and resistant hypertension. J Hypertens. 2015;33:2278–85.
Visser M, Palstra RJ, Kayser M. Human skin color is influenced by an intergenic DNA polymorphism regulating transcription of the nearby BNC2 pigmentation gene. Hum Mol Genet. 2014;23:5750–62.
Paterson AD, Waggott D, Boright AP, Hosseini SM, Shen E, Sylvestre MP, et al. A genome-wide association study identifies a novel major locus for glycemic control in type 1 diabetes, as measured by both A1C and glucose. Diabetes. 2010;59:539–49.
Li C, He J, Hixson JE, Gu D, Rao DC, Shimmin LC, et al. Abstract P253: genome-wide gene-potassium interaction analyses on blood pressure: The GenSalt Study. Circulation. 2016;133:AP253.
Li C, He J, Chen J, Zhao J, Gu D, Hixson JE, et al. Genome-wide gene-potassium interaction analyses on blood pressure: the gensalt study (genetic epidemiology network of salt sensitivity). Circ Cardiovasc Genet. 2017;10:6.
Rosenbloom KR, Sloan CA, Malladi VS, Dreszer TR, Learned K, Kirkup VM, et al. ENCODE data in the UCSC Genome Browser: year 5 update. Nucleic Acids Res. 2013;41:D56–63.
Williams B, MacDonald TM, Morant S, Webb DJ, Sever P, McInnes G, et al. Spironolactone versus placebo, bisoprolol, and doxazosin to determine the optimal treatment for drug-resistant hypertension (PATHWAY-2): a randomised, double-blind, crossover trial. Lancet. 2015;386:2059–68.
Wright JT, Williamson JD, Whelton PK, Snyder JK, Sink KM, Rocco MV, et al. A randomized trial of intensive versus standard blood-pressure control. N Engl J Med. 2015;373:2103–16.
Group SPSS, Benavente, Coffey OR, et al. Blood-pressure targets in patients with recent lacunar stroke: the SPS3 randomised trial. Lancet. 2013;382:507–15.
Cooper-DeHoff R, Handberg E, Heissenberg C, Johnson K. Electronic prescribing via the internet for a coronary artery disease and hypertension megatrial. Clin Cardiol. 2001;24:V14–6.
Newton KM, Peissig PL, Kho AN, Bielinski SJ, Berg RL, Choudhary V, et al. Validation of electronic medical record-based phenotyping algorithms: results and lessons learned from the eMERGE network. J Am Med Inform Assoc. 2013;20:e147–54.
Funding
INVEST was supported by grants from the University of Florida Opportunity Fund and Abbott Pharmaceuticals. INVEST-GENES was supported by NIH grants U01-GM074492, NIH R01 HL074730. The SPS3 trial was funded by the National Institute of Health and Neurological Disorders and Stroke Grant No. U01NS38529-04A1. The SPS3-GENES was funded by R01 NS073346 and U01-GM074492-05S109. Dr. El Rouby is supported by NIH grant T32HL083810 and Dr. McDonough is supported by NIH Grant 1 KL2 TR001429. The eMERGE Network is funded by NHGRI, with additional funding from NIGMS through the following grants: U01HG04599 and U01HG006379 to Mayo Clinic; U01HG004610 and U01HG006375 to Group Health Cooperative and University of Washington, Seattle; U01HG004608 to Marshfield Clinic; U01HG006389 to Essentia Institute of Rural Health; U01HG004609 and U01HG006388 to Northwestern University; U01HG04603 and U01HG006378 to Vanderbilt University; U01HG006385 to the Coordinating Center; U01HG006382 to Geisinger Clinic; U01HG006380 to Mount Sinai School of Medicine. A portion of the dataset(s) used for the analyses described were obtained from Vanderbilt University Medical Center’s BioVU, supported by institutional funding and by the Vanderbilt CTSA grant UL1 TR000445 from NCATS/NIH, and the Mayo Clinic Biobank supported by the Mayo Clinic Center for Individualized Medicine.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Conflict of interest
Dr. Shuldiner is employed by Regeneron Pharmaceuticals, Inc. The remaining authors declare that they have no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
El Rouby, N., McDonough, C.W., Gong, Y. et al. Genome-wide association analysis of common genetic variants of resistant hypertension. Pharmacogenomics J 19, 295–304 (2019). https://doi.org/10.1038/s41397-018-0049-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41397-018-0049-x
This article is cited by
-
A genome-wide association study identifies a novel association between SDC3 and apparent treatment-resistant hypertension
BMC Medicine (2022)
-
A genome-wide association study identifies a novel candidate locus at the DLGAP1 gene with susceptibility to resistant hypertension in the Japanese population
Scientific Reports (2021)