Abstract
Microbial activity in the deep sea is cumulatively important for global elemental cycling yet is difficult to quantify and characterize due to low cell density and slow growth. Here, we investigated microbial activity off the California coast, 50–4000 m water depth, using sensitive single-cell measurements of stable-isotope uptake and nucleic acid sequencing. We observed the highest yet reported proportion of active cells in the bathypelagic (up to 78%) and calculated that deep-sea cells (200–4000 m) are responsible for up to 34% of total microbial biomass synthesis in the water column. More cells assimilated nitrogen derived from amino acids than ammonium, and at higher rates. Nitrogen was assimilated preferentially to carbon from amino acids in surface waters, while the reverse was true at depth. We introduce and apply the Gini coefficient, an established equality metric in economics, to quantify intracommunity heterogeneity in microbial anabolic activity. We found that heterogeneity increased with water depth, suggesting a minority of cells contribute disproportionately to total activity in the deep sea. This observation was supported by higher RNA/DNA ratios for low abundance taxa at depth. Intracommunity activity heterogeneity is a fundamental and rarely measured ecosystem parameter and may have implications for community function and resilience.
Similar content being viewed by others
Introduction
The majority of the marine habitat—~70% of its volume [1]—is permanently dark. Microbial cell density in the water column decreases with water depth, but the dark ocean nonetheless contains about 55% of total microbial cells in the pelagic ocean [2, 3]. Respiration rates of these cells determine the extent to which sinking organic matter exported from the photic zone is degraded and remineralized [4,5,6,7], with implications for carbon burial and the sequestration of carbon dioxide. Microbial activity in the deep sea also has implications for surface water nutrient availability, as deep water-masses will eventually up-well in coastal sites [8]. However, while there is growing appreciation for the phylogenetic and functional diversity of cells in the deep sea [9, 10], efforts to investigate microbial activity in the deep sea are rare, especially at the single-cell level, fueling a debate about the percent of cells that are actually active and leaving the variability in activity levels between cells unknown. Investigations with microautoradiography (using radiolabelled amino acids) and cytometric analysis with CTC+ dye (5-cyano-2,3-ditolyl tetrazolium chloride) reported that 3–20% of prokaryotic cells were active in meso- and bathypelagic waters [11,12,13]. However, these numbers are likely underestimates of total active cells. For instance, not all microorganisms incorporate intact amino acids, including the majority of active Marine Group I Thaumarchaeota (recently reclassified as members of the Nitrososphaeria, a class within the Crenarchaeota), which are abundant in the meso- and bathypelagic [14]. Furthermore, these techniques either cannot or are difficult to obtain quantitative cell-specific activity, and therefore have not been employed to resolve the variation in activity levels between cells.
Nanoscale secondary ion mass spectrometry (nanoSIMS) has made it possible to accurately quantify anabolic activity within uncultured individual microbial cells using a wide range of stable-isotope-labeled substrates [15,16,17,18]. Furthermore, nanoSIMS has allowed the single-cell analysis of large microbial communities [14, 19, 20], making it possible to determine the intracommunity variability in activity. The distribution of activity within a community is a fundamental parameter of an ecosystem and may have implications for the resilience of ecosystem function, yet this is still an emerging concept for complex natural communities. Heterogeneity in activity has been estimated in pure cultures using the standard deviation [21, 22] and within specific, sorted environmental populations by calculating the coefficient of variation of the mean of all cells [23]. These approaches are suitable for normal distributions, however, are not optimal for environmental microbial communities with non-normal, multimodal distributions. To overcome this limitation, Calabrese et al. [24] proposed a new calculation based on rank-activity distribution analysis, defined as differentiation tendency index (DTI), and have used it to demonstrate differentially active subpopulations of non-normally distributed pure cultures [25]. However, since the DTI focuses on differences between the active cells, it is not optimized for characterizing or comparing natural communities with a high abundance of non-active cells, for instance under those under low or variable nutrient conditions. Therefore, there is a need to further develop additional microbial heterogeneity metrics that would allow the heterogeneity comparison under a variety of natural settings.
In the current work we use nanoSIMS and 16S rRNA amplicon analysis to study microbial activity in the deep sea. We aim to achieve a more nuanced perspective of deep-sea microbiology by characterizing the distribution and single-cell anabolic activity of microorganisms from surface to seafloor. We use the uptake of 15N-13C-labeled amino acids and 15N-labeled ammonium as a proxy for total anabolic activity in thousands of individual microbial cells living from 50 to 4000 m water depth at two sites offshore San Francisco in the northeast Pacific Ocean, 67 and 281 km from the coast. We assess the proportion of active cells, single-cell uptake rates, preference for nitrogen uptake from ammonium versus amino acids, and the relative-use efficiency of carbon and nitrogen from amino acids. Additionally, we introduce a comparison of different metrics to quantify the intracommunity heterogeneity of anabolic activity and investigate trends in the distribution of activity. First, we propose using the Gini coefficient to quantify microbial anabolic heterogeneity (inequality), which provides robust and comparable values while incorporating all members of the community (including positive and zero values). The Gini coefficient was developed over a 100 years ago to quantify income inequality in human populations [26], and it is still used routinely in economics today. It has also recently been applied as an indicator of ecological population distribution [27, 28], including bacterial diversity and species distribution [29, 30]. Additionally, we calculate the percent of cells responsible for 90% of total activity for comparison. We compare and contrast these methods together with previous approaches for characterizing the intracommunity distribution of microbial activity in environmental samples.
Material and methods
Sample collection
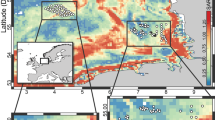
Seawater collection in the northeast Pacific Ocean was carried onboard the R/V Oceanus in March 2017. Seawater was sampled with Niskin bottles at 6 sites along a 300 km transect (Fig. 1). Samples were collected in the epipelagic at 50 and 150 m, in the mesopelagic at 500 and 1000 m, and in the bathypelagic at 2000, 3000, and 4000 m, as the total water depth allowed. Different physicochemical water properties (including temperature, conductivity, pressure, and fluorescence) were determined with a CTD (SeaBird, USA).
Maps of the northeastern Pacific Ocean showing (A) sampling locations and (B) sampling water depths. Asterisks indicate the two sites where stable isotope labeling experiments were conducted. Maps were generated with GeoMapApp version 3.6.12. C Neighbor-joining tree generated from the Bray–Curtis dissimilarity matrix for DNA samples for the different study sites and depths. Water depth at which the sample was collected is indicated by different colors.
16S rRNA gene DNA/cDNA samples collection and processing
Samples for 16S rRNA gene and 16S rRNA sequencing were collected for all depths at all sites. Seawater (5–25 l, depending on the depth) was filtered through 0.2 µm Sterivex filter units (Millipore, Germany) immediately after collection. Filters were flash frozen in liquid N2 and stored at −80 °C. DNA and RNA extraction were performed on shore by an All Prep DNA/RNA mini kit (Qiagen, CA, USA). Samples were prepared for sequencing using the 515F-Y/926R primers [31] and sequencing was performed using the MiSeq (Illumina) platform at the UC Davis DNA Technologies Core Facility. See Supplementary Information for further details. All sequences have been submitted to the European Nucleotide Archive under the accession number PRJEB54878.
Seawater incubations with stable isotopes
Bottle incubations amended with stable isotope-labeled substrates were conducted at two sites: a site 281 km from shore (OC6, hereon called Open Ocean site) and a site located immediately beyond the continental slope (OC3, 67 km from shore; hereon called Slope site). Two sets of incubations were carried out, one using 50 nM of 15N-labeled ammonium chloride (99% 15N, Cambridge Isotope Laboratories, USA), and the other using 50 nM of 15N13C-labeled algal amino acid mixture (AA, 97–99% 13C and 97–99% 15N, Cambridge Isotope Laboratories, USA; amino acid mixture composition included in SI Table 1). Incubations were conducted using 2 or 4 l polycarbonate bottles, in the dark. Water from 50 m and 150 m depth were incubated at 10.5 °C and from 500 to 4000 m at 4 °C, as close as possible to the average in situ temperatures (see Fig. S1). At 0 and 72 h, 100 ml of seawater of each incubation was fixed using 3% formaldehyde and filtered onto polycarbonate filters (25 mm diameter, 0.2 µm pore size; GTTP type, Millipore). Incubation time was selected with the goal of being long enough to achieve low detection limits but short enough to avoid major community shifts, based on previous results [14]. Filtered, fixed cells were washed with PBS, 1:1 PBS:EtOH and EtOH before storage at −80 °C for nanoSIMS analysis. At the same timepoints, 1.875 l (50 and 150 m depth) and 3.875 l (500 to 4000 m depth) unfixed seawater were filtered through anodisc filters (47 mm diameter, 0.2 µm pore size; Whatman, Maidstone, UK) and stored at −80 °C for analysis by isotope ratio mass spectrometry (IRMS).
Bulk isotope uptake by IRMS analysis
Anodisc filters were packed in small tin cups and dried overnight at 55 °C. Bulk isotope analysis was performed at the UC Davis stable isotope facility using an Elementar Vario EL Cube and Micro Cube elemental analyzer (Elementar Analysensysteme GmbH, Hanau, Germany).
Single-cell isotope uptake by nanoSIMS
Single-cell uptake rates for 13C and 15N were analyzed by nanoSIMS. Details of sample preparation and nanoSIMS analysis are described as Supplementary Information. The resulting isotope images were analyzed using LANS software [32] allowing for the accumulation of the scans with lateral drift correction and quantitative analysis of isotopic ratios (13C−12C−/12C2− and 12C15N−/12C14N−). To determine cell-specific isotope ratios, regions of interest were drawn manually inside the cells, avoiding the cell-surrounding areas, using the 12C14N− channel. Cells were considered active if their isotope ratio was greater than 2 standard deviations above the mean isotope ratio of the 0 h cells from each site [14] (Supplementary Information). Cell-specific growth rates and the amount of N and C assimilated by each cell in fg cell−1 h−1 was calculated according to Stryhanyuk et al. [33], and as also described in Arandia-Gorostidi et al. [34]. Integrated rates for each oceanic region were calculated by multiplying cell abundance (see Supplementary Information) at each depth with the single-cell assimilation rate and by extrapolating it to the total volume of each region considering an area of 1 m2. The epipelagic was considered to be 0–200 m, the mesopelagic 201–1000 m, and the bathypelagic 1001–3000 or −4000 m (depending on site). Single-cell C/N relative use efficiency (RUE) from amino acids was calculated according to Mayali et al. [35], only considering cells that were enriched in both 13C and 15N.
Intracommunity activity heterogeneity was calculated using the Gini coefficient and visualized with Lorenz curves [26] using the DescTools package in R (version 3.6.2). The cell-specific assimilation of nitrogen (in fg) over the course of the incubation was used as the input value for each cell, and therefore as a proxy for cellular activity. Additionally, heterogeneity in activity levels was calculated as the percent of cells responsible for 90% of total anabolic activity by determining the number of cells (starting from the most active ones) whose 15N uptake summed to 90% of the total 15N uptake from each substrate, amino acids and ammonium, at each depth. Calculations for the DTI value were carried out following Calabrese et al. [24].
Additional methodological details can be found in the Supplementary Information.
Results
Environmental analysis
Analysis of temperature, salinity, and fluorescence at each site showed standard trends (Fig. S1). As expected, an oxygen minimum zone was detected, with a minimum of 4.3% oxygen saturation at 1000 m water depth (surface values were near 100%). Ammonium concentrations were lowest at the Open Ocean site where it was below the detection limit (<10 nM) throughout the water column and were below 100 nM at most sites (Fig. S2). Nitrate concentrations increased throughout the epi- and mesopelagic and then decreased slightly in the bathypelagic.
Cell abundance and microbial community composition
We observed the maximum cell abundance at 150 m in the Slope site and at 50 m in the Open Ocean site (13.1 × 105 and 4.7 × 105 cell ml−1, respectively). Cell-abundances decreased until 1000 and 2000 m for the Slope and Open Ocean site respectively and remained constant below these depths (Table 1).
Microbial community composition analyzed by 16S rRNA gene (DNA) amplicon sequencing showed that the microbial communities separated taxonomically by water depth rather than distance from shore (Fig. 1C). Alphaproteobacteria and Cyanobacteria dominated the communities at 50 m depth, while Crenarchaeota, Deltaproteobacteria, and Marinimicrobia were relatively more abundant at 150 m depth and below. The Crenarchaeota detected were members of the Nitrososphaeria, previously known as Marine Group I Thaumarchaeota. Other groups such as Gammaproteobacteria showed consistent relative abundance throughout the water column. The greatest dissimilarities separated the 50 m samples from the others, but consistent vertical structuring was observed even within the meso- and bathypelagic regions (Fig. 1C).
Bulk uptake of ammonium and amino acids with depth
Bulk community-level activity measured with IRMS showed maximum incorporation of 15N-ammonium and 15N-amino acids at 50 m, except for ammonium uptake at the Slope site, where it peaked at 150 m depth (Fig. S4). Uptake of 15N from both substrates was low at 500 m depth and was undetected at 1000 m and in most samples within the bathypelagic. 13C-amino acid uptake also peaked at either 50 m (Open Ocean site) or 150 m (Slope Site). Consistent with the pattern of 15N-uptake, uptake of 13C-amino acid was undetected in the bathypelagic except at 2000 m at the slope site.
Single-cell uptake of ammonium and amino acids with depth
Unlike with the bulk IRMS approach, using nanoSIMS we detected cell-specific activity throughout the water column at both sites. Proportions of active cells were maximum in the epipelagic (50 or 150 m) at both sites, with around 90% of the cells incorporating 15N from each substrate (max of 98.7% from amino acids at the Slope Site), and then generally decreased with increasing water depth (Fig. 2). The extent of the decline varied between sites. The proportion of active cells decreased to <25% just below 1000 m at the Open Ocean site, but around 75% of cells assimilated 15N-derived from amino acids throughout the meso- and bathypelagic at the Slope site. At both sites, the proportion of 15N-enriched cells remained relatively constant below 1000 m water depth, although amino acid uptake hit a minimum at 2000 m at the Open Ocean site.
Cell-specific N-based growth and assimilation rates were higher at the Slope site than at the Open Ocean site and decreased with water depth at both sites (Fig. 3 and Table 1). Similar to the proportion of active cells, this decline with depth was more extensive at the Open Ocean site. In contrast, while the proportion of 15N-enriched cells was similar for amino acids and ammonium, the N-based growth and assimilation rates were different for the two substrates. Mean single-cell N-based growth and assimilation rates for amino acids were considerably higher than that for ammonium; greater than one order of magnitude in the epipelagic and in the bathypelagic region (Fig. 3 and Table 1).
A Histograms of N-based cell-specific growth for 15N-amino acids (blue) and 15N-ammonium (green) in log10 scale for each depth of the Open ocean (left) and Slope sites (right). Histogram bar-width represent a range of Ka = 0.15 for 50 m and Ka = 0.3 for the rest of the depths. Only active cells are displayed. Boxplots indicate the median and the upper and lower quartiles. Inset text indicate the total number of analyzed cells and the percentage of active cells in parentheses. B NanoSIMS 12C15N−/12C14N− isotope ratio images showing enrichment of 15N from 15N-amino acids (AA) at the Slope site. Only active (15N-enriched) cells are visible.
Cell-specific assimilation rates and proportion of active cells were also determined for 13C derived from amino acids (Table 1 and Fig. S5). Similar to the trend for 15N-based growth, the highest assimilation rates for 13C-amino acids were both found at 50 m for both sites (0.0103 and 0.0162 fg cell−1 h−1 and 74.8% and 48.3% for Slope and Open Ocean sites, respectively). The percent of cells enriched in 13C from amino acids was lower than that from 15N throughout the water column, decreasing to 4% (or less) around 1000 m at both sites, and then creeping up again at both sites to 25% and 5% at the bottom of the water column at the Slope and Open Ocean site, respectively. The low single-cell 13C enrichment could be due to inclusion of carbon from the polycarbonate filters in the analysis, which would dilute the 13C enrichment in the cells and underestimate uptake [36]. However, it could also be due to selective usage of nitrogen from amino acids, and/or inflated rates of 15N assimilation from amino acids due to some cells assimilating the 15N-products of amino acid degradation (e.g., 15N-ammonium) via cross-feeding rather than 15N directly from amino acids.
C:N RUE were calculated for those cells incorporating both C and N from amino acids. A value of one indicates stoichiometric assimilation of the two elements and therefore likely assimilation of intact amino acids [35]. The C:N RUE varied significantly with depth (ANOVA, p value <0.001 for both sites). The maximum RUE values (i.e., high preference for carbon over nitrogen) were found at the deepest depth at both sites (with a ratio of 0.77 and 2.55 at 3000 m for the Slope site and at 4000 m at Open Ocean site, respectively, Fig. 4), indicating an increasing preferential assimilation of carbon with depth. The RUE values were also significantly higher in the Ocean Site (1.25 on average) than in the Slope site (0.36), indicating a higher demand for carbon relative to nitrogen in the open ocean (t-test, p value <0.001).
Intracommunity heterogeneity of microbial activity
The distribution of activity (cell-specific uptake rates) between cells generally did not follow a normal distribution (as tested using the Shapiro–Wilk’s method) and was often multimodal with an overall positive skew (Fig. 3, note the log scale on the x-axis). At the Slope site, the Gini coefficient varied from 0.48 to 0.83, and generally increased with increasing water depth, indicating increasing heterogeneity in activity with depth (Fig. 5). The Gini coefficient for ammonium assimilation at 150 m was a notable exception to this trend. In the Open Ocean site, the Gini coefficient varied from 0.29 to 0.96, with trends differing by substrate but generally showing a peak in heterogeneity within the bathypelagic (2000 m) as well. For reference, a coefficient value of 0 indicates that anabolic activity is equally distributed between all the cells of the community (indicating that all cells have the same isotope enrichment per unit of biovolume), while a value of 1 indicates a single-cell is responsible for all activity (complete inequality).
A Schematic representation of Lorenz curve and the Gini coefficient calculations. B Lorenz curves and Gini coefficients by depth for the Open ocean (left) and Slope sites (right). Dotted line shows perfectly even distribution of cell-specific activity. C Changes of Gini coefficient with depth for amino acid (blue) and ammonium (green) incubations, in the Slope Site (right) and Open Ocean (left) sites.
As an alternative way to quantify the distribution of activity, the proportion of cells that were responsible for 90% of total 15N-uptake from amino acids and ammonium were calculated (hereon called P90), with low proportions indicating that few cells were responsible for most activity (and therefore higher activity heterogeneity). The observed trends with this metric were very similar to the ones determined using the Gini coefficient, and the values ranged from 16 to 47% (Table 1). The two metrics showed a significant correlation between them (t-test, p value = 0.03).
RNA/DNA ratios of individual amplicon sequence variants (ASVs)
In addition to assessing microbial community composition with 16S rRNA genes (DNA), potential translational activity was calculated by analyzing 16S rRNA amplicons (RNA) (Fig. 3B). Gammaproteobacteria and Marinimicrobia groups represented most of the RNA reads in the meso- and bathypelagic zones (21.5% and 16.4% of the total reads in the meso- and bathypelagic zones combined, respectively). Nitrososphaeria RNA reads were relatively low at all depths (3.7%, 5.3% and 5.5% of all reads in the epi-, meso- and bathypelagic regions respectively), despite being the most abundant taxa in the DNA reads at water depths of 150 m and below (Fig. S3B).
The potential activity for each amplicon sequence variant (ASV) was estimated using their relative 16S rRNA RNA/DNA ratio, as proposed by Kemp et al. [37] for marine microbes. The least-abundant microbial ASVs (<25% of DNA reads) had higher RNA/DNA ratios on average than the rest of the community in the epi-, meso-, and bathypelagic regions (Fig. 6). These cells belonged almost exclusively to the Proteobacteria phylum, and mostly to the Deltaproteobacteria Class (30% of all cells with an RNA/DNA ratio >25%). Furthermore, the average RNA/DNA ratio of this group—the least abundant ASVs—increased with water depth (ANOVA, p value <0.01).
Discussion
Quantification of microbial anabolic activity in the deep sea
Quantitative measurements of microbial activity in the deep-sea are relatively rare, especially at the single-cell level. Bulk analysis of uptake of stable isotope tracers via IRMS is problematic due to the low density of cells (and therefore large volumes of water required for incubations), the low rates of activity (relative to the detection limits of the assay), and the need to use non-carbon-based filters due to the total combustion during analysis. Here, we used aluminum oxide anodisc filters with a pore size of 0.2 μm to overcome the potential loss of small deep-sea cells through traditional glass-fiber filters (GF/F, nominal pore size 0.7 μm) for our IRMS analysis. However, we were still only able to detect isotope uptake via IRMS in the epipelagic region, with uptake levels mostly below the limit of detection below 500 m water depth. As a result of such difficulties, bulk anabolic activity in the deep sea has been mostly performed with radioisotopes to date [3, 38, 39]. Although a sensitive bulk assay, relying on radioisotopes constrains the range of tracer substrates significantly, and limits the potential for cell-specific metabolic rate measurements.
Previous studies examining microbial activity at the community level using 3H-leucine have detected significant rates of uptake throughout the deep-water column [12, 40], in some cases rivaling that in the epipelagic when integrated with depth [3]. However, studies using microautoradiography to visually detect 3H- or 14C-uptake in individual cells have consistently detected activity in only 3–20% of cells in the bathypelagic, leaving open the possibility that most deep-sea cells are dead or anabolically dormant [11,12,13, 41]. Here, using 15N-ammonium and 15N-amino acids and quantitative single-cell analysis by nanoSIMS, we not only detect activity throughout the water column but exceed previous estimates for the proportion of active cells in the deep sea by several fold in some samples (up to 78% in the bathypelagic). Our values for the epipelagic, where 90–98% of the cells were detected as active, were also greater than previous estimates in the epipelagic with microautoradiography, and consistent with our recent nanoSIMS observations made closer to shore (<20 km) [14]. A difference between ours and previous studies is the higher concentration of substrates added to the incubations (50 nM versus 10 nM), lowering the detection limit but also increasing the chance that activity was inadvertently stimulated. Although we cannot eliminate that possibility, the lower proportions of active cells found at the Ocean Site, where the percent of added substrate over the in situ concentration was highest, in comparison to the Slope Site, indicates that any stimulation was likely minor and could not explain the high values at the Slope Site alone. At a minimum, the high proportions of active cells in our incubations demonstrate the high proportion of viable cells in the deep sea. Additionally, it suggests that the highly reproducible vertical structuring in microbial community composition throughout the water column (Fig. S3 and [42]) is due to environmental selection of specific physiologies and metabolic capabilities as conditions change with depth rather than stochastic processes distributing dormant cells.
Both the proportion of active cells and the 15N-enrichment level of the active cells decreased with water depth (Figs. 2 and 3). Previous work noted that bulk levels of activity decreased faster than the decrease in cell numbers indicated, suggesting that the rate of activity in individual cells as well as the number of cells was decreasing with depth [38,39,40]. Our single-cell rate results provide direct evidence that this is indeed the case. However, we found that when integrated by water column volume, the microbes in the meso- and bathypelagic zones were still responsible for 14 and 34% of the total anabolic activity of the water column at the Open Ocean and Slope sites, respectively, measured by uptake of 15N-amino acids. The integrated values are lower than the 48.2% estimated by Aristegui et al. [3] using bulk 3H-leucine uptake in the North Atlantic Ocean. Such difference, however, can be due to the low cell-specific production rates estimated in Aristegui et al. [3] for the epipelagic region, which is typical for oligotrophic surface waters such as the North Atlantic. Contrary to Aristegui et al. [3], our study was performed in an upwelling zone, which explains the higher surface microbial activity and likely the greater difference in activity between the epipelagic and deeper waters. Additionally, a recent study has reported that marine prokaryotic production can be overestimated by several fold due to inaccuracy of leucine-carbon conversion factors, with overestimation more likely and severe in deeper waters [43]. Studies using these conversion factors therefore might calculate artificially high relative proportions of activity in the deep sea. Still, the difference between the two sites within our study also indicates that the contribution of the dark ocean to the total pelagic activity can be variable even within a single region, potentially dependent on a combination of biological, chemical, and physical processes.
While bulk activity levels and the percent of active cells were similar between the Open Ocean and Slope sites at 50 m depth, in the meso- and bathypelagic the Slope site had 42% more active cells, ~15 times higher cell-specific uptake rates, and nearly twice the integrated contribution to water column activity as the Open Ocean site. The observed difference in activity may be related to (1) differences in microbial community composition, and/or (2) differences in environmental parameters between sites. The highly similar community composition of the two sites at both the class (Fig. S3A) and ASV level (see Fig. S6) indicates that the impact of taxonomic composition was likely minor. The difference in chemical properties was, most likely, the major cause of the difference in activity observed between both sites. The higher ammonium in situ concentration found in the slope site, probably supporting both heterotrophic and (chemo-) autotrophic metabolisms, and the potentially higher availability of organic carbon in this site due to its proximity to coast [44] likely explains the larger activity rates found in the meso- and bathypelagic regions of the Slope Site. This suggests that the community present at the Open Ocean site could be capable of higher activity rates if additional organic matter became available and is not inherently limited by enzymatic availability or physical limitation (e.g., temperature or pressure). This may have implications for the rate at which natural or anthropogenic bursts of organic matter to the deep sea in the open ocean would be respired.
We observed higher proportions of active cells with 15N-amino acids than 15N-ammonium, apparently contrary to our initial hypothesis that amino acid uptake underestimates active cells due to lack of uptake of intact amino acids by some abundant deep-sea taxa. However, enrichment in 15N does not require assimilation of intact 15N-amino acids, as some organisms might consume 15N via the 15N-labeled products of amino acid degradation performed by other cells, i.e., cross-feeding, or even preferentially assimilate nitrogen over carbon from amino acids [35]. The latter could be achieved if after cleaving the amine group the cells either excreted the C-skeleton or respired it, releasing the 13C as 13C-CO2. The lower levels of 13C-assimilation from 13C-amino acids in this study supports these alternative explanations (lack of assimilation of intact amino acids), as has been observed previously for members of the Nitrososphaeria [14]. The combined observations suggest 15N-amino acids are a taxonomically inclusive tracer of viability, more so than 13C-amino acids, with the caveat that 15N-enrichment may reflect assimilation of a variety of 15N-molecules. The higher proportions and rates compared to ammonium uptake could reflect a stimulation due to alleviation of carbon limitation for some cells, particularly heterotrophs. In this respect, ammonium uptake may be more representative of in situ activity than amino acid uptake, however, ammonium may stimulate activity in other cells, particularly ammonia-oxidizing microorganisms (e.g., members of the Nitrososphaeria). Another potential explanation for greater uptake of amino acids over ammonium is simply a preference for organic versus inorganic nitrogen; amino acid-specific transporters have been shown to be more abundant relative to those for ammonium (Amt transporters) in the meso and bathypelagic regions of the Atlantic Ocean [9].
We observed an increasing C/N RUE from amino acids with depth, and overall higher values at the Open Ocean Site. This is likely due to increasing C limitation with depth and with distance from coast. Microbes in coastal upper waters likely have more access to alternative organic C sources (non 13C labeled, thus decreasing the C/N use efficiency), due to photosynthetic production, with C limitation increasing with depth [45] and distance from shore [44]. This, combined with increasingly abundant alternative N sources in the deep sea (e.g., nitrate), may result in a higher C- than N-affinity from amino acids in the meso- and bathypelagic regions of the Open Ocean site.
Recovering the samples with standard niskin bottles and incubating in non-pressurized vessels resulted in a decompression of the cells, which normally experience 5 to 400x sea-level pressure in situ (assuming a hydrostatic pressure of 1 atm/10 m water depth). A previous survey revealed that 76% of total deep-sea samples investigated (n = 252 comparisons; water and sediment) and 90% of the unmixed water column samples (n = 120 comparisons) demonstrated decreased microbial activity when measured at sea-level relative to in situ pressure [46]. This suggests that most assays of deep-sea activity conducted at sea level underestimate activity, and that the values for proportion of active cells and the average cell-specific growth rates measured here are likely lower than if pressure had been maintained. Furthermore, since the difference between in situ and sea-level pressure increases with sampling depth, it is possible that activity in deeper sites is systematically underestimated. Therefore, the attenuation of activity with depth may be less steep in reality than observed here. It will be illuminating to perform the type of single-cell measurements conducted here while maintaining in situ pressures, to confirm and/or refine the values and trends observed here.
Distribution of anabolic heterogeneity with depth
To assess heterogeneity in cell-specific growth rates within marine communities, we used a previously proposed approach for non-normal multimodal cell-specific distribution (DTI [24]) and two alternative approaches proposed in the current work: (1) Gini coefficient and (2) proportion of cells responsible for 90% of total anabolic activity (P90). We compared all three approaches at the Slope site, and observe similar trends (Table 1 and Fig. S7), yet only the Gini coefficient and the P90 metric showed a significant correlation with each other. The DTI values also showed lower reproducibility than the other two approaches as it is sensitive to somewhat subjective manual identification of activity subpopulations. As DTI heterogeneity calculations focus on differences between active cells, we did not test it in the Open Ocean site, where we observed up to 89% inactive cells. Although the DTI still represents a very useful approach for highly active communities, or for microbial cultures, we consider that the Gini coefficient better addresses the activity distribution in communities with very variable activity rates, including many at or below detection levels. While the P90 is perhaps the most intuitive of all, we recommend it only as a first pass to understand activity heterogeneity in a sample because its simplicity fails to capture more subtle differences in activity distribution.
The lowest Gini coefficient values (lowest heterogeneity, highest evenness), were usually found at 50 m (particularly in the Slope Site), coinciding with the maximum proportion of active cells and highest mean N-based growth rates. Below the epipelagic zone, we observed an increasing Gini coefficient in the Slope site (indicating higher activity heterogeneity with depth), and a decrease in the P90, both indicating that an increasingly small subset of cells is responsible for the majority of total anabolic activity in water samples from this oceanic realm (Table 1). Such increasing heterogeneity with depth might be related to slower growth due to nutrient limitation. Phenotypic heterogeneity has been shown to increase in a pure culture due to nutrient limitation, including specifically in Staphylococcus aureus when growth was slowed by carbon limitation [21, 22]. A similar effect might therefore happen in the deep sea, where waters are typically more oligotrophic than at the surface [47].
In contrast to the Slope site, however, the Open Ocean site showed decreasing heterogeneity values below 2000 m depth. Since carbon availability is generally thought to decrease unidirectionally with depth, the decreasing variability below 2000 m depth indicates that if nutrient availability is a driving factor in the intracommunity distribution of activity, carbon limitation is not the only, or even primary driver in the bathypelagic at the Open Ocean site. In a complex community, there are likely many factors affecting intracommunity heterogeneity in activity, including both genotypic and phenotypic diversity, and the non-linearity or even lack of unidirectionality at the Open Ocean site likely reflects their complicated interplay. Additionally, some deep-sea cells might be less active after decompression (or not active at all), as described above, while others, for instance those transported to depth via particles recently, might be more active at decreased pressure [46], adding another layer of complexity to these trends. However, effects of decompression likely did not dominate our observations because while pressure changes consistently and linearly with depth, the trends we observe in heterogeneity in activity are more variable.
Difference in heterogeneity was also observed between the 15N-ammonium and amino acid uptake. While the Gini coefficient was overall higher (reflecting greater heterogeneity in uptake) for ammonium than amino acids in the epipelagic zone (average Gini coefficient values of 0.73 and 0.53, respectively) in the meso- and bathypelagic regions the reverse was true (average values of 0.68 and 0.76, respectively). Although previous studies have shown distinct amino acid uptake rates for different microbial taxa in the surface waters [34, 48], our results suggest that heterogeneity in the incorporation of ammonium may exceed the one of amino acid in the epipelagic zone, which agrees with previous single-cell studies showing a high variability in the uptake of ammonium [23, 49]. The larger amino acid uptake heterogeneity in the meso- and bathypelagic regions on the other hand, might be due to the lack of amino acid incorporation in some deep-sea taxa, such as members of the Nitrososphaeria [14] which would increase the overall heterogeneity of the community as these community members become more abundant.
ASV-specific activity in the dark ocean
RNA/DNA ratios have previously been used to estimate relative microbial activity, as the relationship between 16S rRNA copies (RNA) and 16S rRNA genes (DNA) may reflect the growth rate for environmental microbial taxa [36, 50, 51]. However, there are significant limitations to this approach [52], as factors other than growth rate can affect this relationship, including physiological changes and different life strategies [53, 54]. Direct measurements of activity, including observations of stable isotope uptake, are therefore more definitive than RNA-based assessments of activity. However, direct measurements can be subject to their own biases, including bottle effects and limited sample sizes, making a combination of independent techniques to assess potential activity beneficial [55]. Here, we explored RNA/DNA ratios not to quantify the relative activity of specific taxa, but rather to compare overall trends with depth with our direct activity measurements performed by nanoSIMS analysis.
The observation that the RNA/DNA ratio of low abundance ASVs (<25% of DNA reads) increases more rapidly with depth than other ASVs (Fig. 6) is consistent with our isotope data in suggesting that a minority of the community is responsible for an increasingly disproportionate share of total activity at depth. Higher potential activity of the low-abundant microbes has been previously reported in the surface of coastal Atlantic Ocean [50] as well as in the meso- and bathypelagic regions of the Pacific Ocean [56]. Although increasing RNA/DNA ratio can be a result of higher stochasticity during the PCR and/or amplicon sequencing of the low abundant taxa, the increasing average RNA/DNA ratios of the low-abundance taxa relative to that of the other taxa with increasing water depth cannot be explained by known artifacts. The few, highly active deep-sea cells in the isotope analysis may or may not be these low abundance ASVs; heterogeneity in activity within ASVs can also explain the highly anabolically active rare cells, as explained above, or a combination of these mechanisms. Still, both the molecular and isotope-uptake datasets suggest that a small fraction of the community is responsible for an increasingly large part of the total activity with increasing depth in the ocean.
Conclusions
Our work provides a first look at cell-specific microbial activity and its intracommunity heterogeneity throughout the marine water column. Despite levels of anabolic activity below the detection limit of a traditional bulk stable isotope incorporation assay, we detect activity throughout the water column on the single-cell level and report the highest yet observed proportions of active cells in the bathypelagic. The high activity observed at the Slope Site relative to the Open Ocean Site suggests the importance of greater nutrient availability within near-shore deep waters, and furthermore that nutrient limitation might therefore outweigh physical stressors in limiting activity at depth. This highlights the fact that intermittent events such as the degradation of sinking particles by attached microbial communities and natural or artificial events that temporarily increase carbon export to the deep sea may result in a substantial activity boost to deep-sea communities.
We propose the Gini coefficient as an intracommunity activity distribution metric. It can be easily applied across ecosystems in future studies and provide an additional parameter with which we can compare the basic functional structure of a community, analogous to diversity indices. Here, the Gini coefficient indicates that an increasingly small fraction of the community is responsible for a disproportionate share of activity with depth. The increasing heterogeneity with depth could reflect the higher resistance of some rare taxa to nutrient limitation and/or other environmental stress related to increasing depth, or a cumulative effect of each individual taxa demonstrating greater phenotypic heterogeneity. The effects of depressurization during the isotope incubations may also influence these trends, however the ASV RNA/DNA ratios support our overall conclusion. The effect of increasing activity heterogeneity in natural microbial communities is not yet known. While phenotypic heterogeneity can increase the chance of survival in a subset of a genetically identical population (i.e., “bet hedging” [22, 57]), the overall community may be less resilient due to the vulnerability inherent when a few individuals dominate total community function. Investigating the identity of the few, highly anabolically active cells as well as the causes and implications of this trend with water depth and other environmental gradients will be fruitful directions for future work.
Data availability
The authors declare that all data supporting the findings of this study are available within the article and its Supplementary Information file, or from the corresponding author upon request. All sequences have been submitted to the European Nucleotide Archive (ENA) under the project accession number PRJEB54878.
References
Orcutt BN, Sylvan JB, Knab NJ, Edwards KJ. Microbial ecology of the dark ocean above, at, and below the seafloor. Microbiol Mol Biol Rev. 2011;75:361–422.
Whitman WB, Coleman DC, Wiebe WJ. Prokaryotes: the unseen majority. Proc Natl Acad Sci USA. 1998;95:6578–83.
Arístegui J, Gasol JM, Duarte CM, Herndl GJ. Microbial oceanography of the dark ocean's pelagic realm. Limnol Oceanogr. 2009;54:1501–29.
Ebrahimi A, Schwartzman J, Cordero OX. Cooperation and spatial self-organization determine rate and efficiency of particulate organic matter degradation in marine bacteria. Proc Natl Acad Sci USA. 2019;116:23309–16.
Fontanez KM, Eppley JM, Samo TJ, Karl DM, DeLong EF. Microbial community structure and function on sinking particles in the North Pacific Subtropical Gyre. Front Microbiol. 2015;6:469.
Simon M, Grossart HP, Schweitzer B, Ploug H. Microbial ecology of organic aggregates in aquatic ecosystems. Aquat Microb Ecol. 2002;28:175–211.
Alldredge AL, Silver MW. Characteristics, dynamics and significance of marine snow. Progr Oceanogr. 1988;20:41–82.
Zehr JP, Kudela RM. Nitrogen cycle of the open ocean: from genes to ecosystems. Ann Rev Mar Sci. 2011;3:197–225.
Bergauer K, Fernandez-Guerra A, Garcia JAL, Sprenger RR, Stepanauskas R, Pachiadaki MG, et al. Organic matter processing by microbial communities throughout the Atlantic water column as revealed by metaproteomics. Proc Natl Acad Sci USA. 2018;115:E400–8.
Acinas SG, Sánchez P, Salazar G, Cornejo-Castillo FM, Sebastián M, Logares R, et al. Deep ocean metagenomes provide insight into the metabolic architecture of bathypelagic microbial communities. Commun Biol. 2021;4:604.
Teira E, Lebaron P, van Aken H, Herndl GJ. Distribution and activity of Bacteria and Archaea in the deep water masses of the North Atlantic. Limnol Oceanogr. 2006;51:2131–44.
Herndl GJ, Reinthaler T, Teira E, van Aken H, Veth C, Pernthaler A, et al. Contribution of Archaea to total prokaryotic production in the deep Atlantic Ocean. Appl Environ Microbiol. 2005;71:2303–9.
Gasol JM, Alonso-Sáez L, Vaqué D, Baltar F, Calleja ML, Duarte CM, et al. Mesopelagic prokaryotic bulk and single-cell heterotrophic activity and community composition in the NW Africa-Canary Islands coastal-transition zone. Progr Oceanogr. 2009;83:189–96.
Dekas AE, Parada AE, Mayali X, Fuhrman JA, Wollard J, Weber PK, et al. Characterizing chemoautotrophy and heterotrophy in marine archaea and bacteria with single-cell multi-isotope NanoSIP. Front Microbiol. 2019;10:2682.
Musat N, Foster R, Vagner T, Adam B, Kuypers MMM. Detecting metabolic activities in single cells, with emphasis on nanoSIMS. FEMS Microbiol Rev. 2012;36:486–511.
Orphan VJ, House CH. Geobiological investigations using secondary ion mass spectrometry: microanalysis of extant and paleo-microbial processes. Geobiology. 2009;7:360–72.
Pett-Ridge J, Weber PK. NanoSIP: NanoSIMS applications for microbial biology. Methods Mol Biol. 2012;881:375–408.
Nuñez J, Renslow R, Cliff JB, Anderton CR. NanoSIMS for biological applications: current practices and analyses. Biointerphases. 2018;13:03B301.
Dawson KS, Scheller S, Dillon JG, Orphan VJ. Stable isotope phenotyping via cluster analysis of NanoSIMS data as a method for characterizing distinct microbial ecophysiologies and sulfur-cycling in the environment. Front Microbiol. 2016;7:774.
Arandia-Gorostidi N, Weber PK, Alonso-Sáez L, Morán XAG, Mayali X. Elevated temperature increases carbon and nitrogen fluxes between phytoplankton and heterotrophic bacteria through physical attachment. ISME J. 2017;11:641–50.
Kopf SH, McGlynn SE, Green-Saxena A, Guan Y, Newman DK, Orphan VJ. Heavy water and (15) N labelling with NanoSIMS analysis reveals growth rate-dependent metabolic heterogeneity in chemostats. Environ Microbiol. 2015;17:2542–56.
Schreiber F, Littmann S, Lavik G, Escrig S, Meibom A, Kuypers MMM, et al. Phenotypic heterogeneity driven by nutrient limitation promotes growth in fluctuating environments. Nat Microbiol. 2016;1:16055.
Berthelot H, Duhamel S, L'Helguen S, Maguer JF, Wang S, Cetinic I, et al. NanoSIMS single cell analyses reveal the contrasting nitrogen sources for small phytoplankton. ISME J. 2019;13:651–62.
Calabrese F, Voloshynovska I, Musat F, Thullner M, Schlömann M, Richnow HH, et al. Quantitation and comparison of phenotypic heterogeneity among single cells of monoclonal microbial populations. Front Microbiol. 2019;10:1–23.
Calabrese F, Stryhanyuk H, Moraru C, Schlömann M, Wick LY, Richnow HH, et al. Metabolic history and metabolic fitness as drivers of anabolic heterogeneity in isogenic microbial populations. Environ Microbiol. 2021;23:6764–76.
Gini C. Variabilità e Mutuabilità. Contributo allo Studio delle Distribuzioni e delle Relazioni Statistiche. C. Cuppini, Bologna; 1912.
Fernández-Tschieder E, Binkley D. Linking competition with growth dominance and production ecology. Ecol Manag. 2018;414:99–107.
Cordonnier T, Kunstler G. The Gini index brings asymmetric competition to light. Perspect Plant Ecol Evol Syst. 2015;17:107–15.
Harch BD, Correll RL, Meech W, Kirkby CA, Pankhurst CE. Using the Gini coefficient with BIOLOG substrate utilisation data to provide an alternative quantitative measure for comparing bacterial soil communities. J Microbiol Methods. 1997;30:91–101.
Li J, Ma YB, Hu HW, Wang JT, Liu YR, He JZ. Field-based evidence for consistent responses of bacterial communities to copper contamination in two contrasting agricultural soils. Front Microbiol. 2015;6:31.
Parada AE, Needham DM, Fuhrman JA. Every base matters: assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples. Environ Microbiol. 2016;18:1403–14.
Polerecky L, Adam B, Milucka J, Musat N, Vagner T, Kuypers MM. Look@NanoSIMS-a tool for the analysis of nanoSIMS data in environmental microbiology. Environ Microbiol. 2012;14:1009–23.
Stryhanyuk H, Calabrese F, Kümmel S, Musat F, Richnow HH, Musat N. Calculation of single cell assimilation rates from SIP-nanoSIMS-derived isotope ratios: a comprehensive approach. Front Microbiol. 2018;9:2342.
Arandia‐Gorostidi N, Alonso‐Sáez L, Stryhanyuk H, Richnow HH, Morán XAG, Musat N. Warming the phycosphere: Differential effect of temperature on the use of diatom‐derived carbon by two copiotrophic bacterial taxa. Environ Microbiol. 2020;22:1381–96.
Mayali X, Weber PK, Pett-Ridge J. Taxon-specific C/N relative use efficiency for amino acids in an estuarine community. FEMS Microbiol Ecol. 2013;83:402–12.
Meyer NR, Fortney JL, Dekas AE. NanoSIMS sample preparation decreases isotope enrichment: magnitude, variability and implications for single-cell rates of microbial activity. Environ Microbiol. 2021;23:81–98.
Kemp PF, Lee S, Laroche J. Estimating the growth rate of slowly growing marine bacteria from RNA content. Appl Environ Microbiol. 1993;59:2594–601.
Baltar F, Arístegui J, Gasol J, Sintes E, van Aken H, Herndl G. High dissolved extracellular enzymatic activity in the deep central Atlantic Ocean. Aquat Micro Ecol. 2010;58:287–302.
Lønborg C, Nieto-Cid M, Hernando-Morales V, Hernández-Ruiz M, Teira E, Álvarez-Salgado XA. Photochemical alteration of dissolved organic matter and the subsequent effects on bacterial carbon cycling and diversity. FEMS Microbiol Ecol. 2016;92:fiw048.
Nagata T, Fukuda H, Fukuda R, Koike I. Bacter-ioplankton distribution and production in deep Pacific waters: large-scale geographic variations and possible coupling with sinking particle fluxes. Limnol Oceanogr. 2000;45:426–35.
Teira E, Reinthaler T, Pernthaler A, Pernthaler J, Herndl GJ. Combining catalyzed reporter deposition-fluorescence in situ hybridization and microautoradiography to detect substrate utilization by bacteria and Archaea in the deep ocean. Appl Environ Microbiol. 2004;70:4411–4.
Mestre M, Hofer J. The microbial conveyor belt: connecting the globe through dispersion and dormancy. Trends Microbiol. 2020;29:482–92.
Giering SLC, Evans C. Overestimation of prokaryotic production by leucine incorporation—and how to avoid it. Limnol Oceanogr. 2022;67:726–38.
Amos CM, Castelao RM, Medeiros PM. Offshore transport of particulate organic carbon in the California Current System by mesoscale eddies. Nat Commun. 2019;10:4940.
Bauer JE, Druffel ERM. Ocean margins as a significant source of organic matter to the deep open ocean. Nature. 1998;392:482–5.
Tamburini C, Boutrif M, Garel M, Colwell RR, Deming JW. Prokaryotic responses to hydrostatic pressure in the ocean—a review. Environ Microbiol. 2013;15:1262–74.
Arrieta JM, Mayol E, Hansman RL, Herndl GJ, Dittmar T, Duarte CM. Dilution limits dissolved organic carbon utilization in the deep ocean. Science. 2015;348:331–3.
Alonso C, Musat N, Adam B, Kuypers M, Amann R. HISH-SIMS analysis of bacterial uptake of algal-derived carbon in the Río de la Plata estuary. Syst Appl Microbiol. 2012;35:541–8.
Klawonn I, Bonaglia S, Whitehouse MJ, Littmann S, Tienken D, Kuypers MMM, et al. Untangling hidden nutrient dynamics: rapid ammonium cycling and single-cell ammonium assimilation in marine plankton communities. ISME J. 2019;13:1960–74.
Campbell BJ, Yu L, Heidelberg JF, Kirchman DL. Activity of abundant and rare bacteria in a coastal ocean. Proc Natl Acad Sci USA. 2011;108:12776–81.
Kirchman DL. Growth rates of microbes in the oceans. Annu Rev Mar Sci. 2016;8:150720190448005.
Blazewicz SJ, Barnard RL, Daly RA, Firestone MK. Evaluating rRNA as an indicator of microbial activity in environmental communities: limitations and uses. ISME J. 2013;7:2061–8.
Licht TR, Tolker-Nielsen T, Holmstrøm K, Krogfelt KA, Molin S. Inhibition of Escherichia coli precursor-16S rRNA processing by mouse intestinal contents. Environ Microbiol. 1999;1:23–32.
Sukenik A, Kaplan-Levy RN, Welch JM, Post AF. Massive multiplication of genome and ribosomes in dormant cells (akinetes) of Aphanizomenon ovalisporum (Cyanobacteria). ISME J. 2012;6:670–9.
Dekas AE, Connon SA, Chadwick GL, Trembath-Reichert E, Orphan VJ. Activity and interactions of methane seep microorganisms assessed by parallel transcription and FISH-NanoSIMS analyses. ISME J. 2016;10:678–92.
Wu J, Gao W, Johnson R, Zhang W, Meldrum D. Integrated metagenomic and metatranscriptomic analyses of microbial communities in the meso- and bathypelagic realm of North Pacific Ocean. Mar Drugs. 2013;11:3777–801.
Ackermann M. A functional perspective on phenotypic heterogeneity in microorganisms. Nat Rev Microbiol. 2015;13:497–508.
Acknowledgements
This work was supported the Simons Foundation (Early Career Investigator Award 507798 to AED). The cruise on the R/V Oceanus was supported by the National Science Foundation (OCE-1634297 to AED). We thank the captain and crew of OC1703A for their support during the fieldwork, and Julian Fortney, Bennett Kapili and Nicolette Meyer for their help during sample collection. The nanoSIMS analyses were performed at the Stanford Nano Shared Facilities (SNSF), supported by the National Science Foundation under award ECCS-2026822. We thank Christie Jilly-Rehak and Chuck Hitzman at the SNSF for their support during the nanoSIMS analysis.
Author information
Authors and Affiliations
Contributions
AEP and AED collected samples and designed and conducted the stable isotope experiment. AEP performed nutrient analyses. NAG performed DNA/RNA preparation for sequencing and nanoSIMS measurements. NAG analyzed the data and generated the figures. NAG and AED interpreted the data and wrote the paper with input from AEP.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Arandia-Gorostidi, N., Parada, A.E. & Dekas, A.E. Single-cell view of deep-sea microbial activity and intracommunity heterogeneity. ISME J 17, 59–69 (2023). https://doi.org/10.1038/s41396-022-01324-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-022-01324-6