Abstract
Honey bees have suffered dramatic losses in recent years, largely due to multiple stressors underpinned by poor nutrition [1]. Nutritional stress especially harms larvae, who mature into workers unable to meet the needs of their colony [2]. In this study, we characterize the metabolic capabilities of a honey bee larvae-associated bacterium, Bombella apis (formerly Parasaccharibacter apium), and its effects on the nutritional resilience of larvae. We found that B. apis is the only bacterium associated with larvae that can withstand the antimicrobial larval diet. Further, we found that B. apis can synthesize all essential amino acids and significantly alters the amino acid content of synthetic larval diet, largely by supplying the essential amino acid lysine. Analyses of gene gain/loss across the phylogeny suggest that four amino acid transporters were gained in recent B. apis ancestors. In addition, the transporter LysE is conserved across all sequenced strains of B. apis. Finally, we tested the impact of B. apis on developing honey bee larvae subjected to nutritional stress and found that larvae supplemented with B. apis are bolstered against mass reduction despite limited nutrition. Together, these data suggest a novel role of B. apis as a nutritional mutualist of honey bee larvae.
Similar content being viewed by others
Introduction
All metazoan evolution has occurred in the context of microorganisms, which exist in and around all living things. Consequently, mutualism between microbes and animal hosts is an ancient and widespread phenomenon (see [3]). Bacterial symbionts can have dramatic effects on animal hosts, including nutrient supplementation of incomplete host diets [4,5,6,7], protection from parasites and pathogens [8,9,10], and providing developmental cues [8, 11,12,13,14,15]. Many eukaryotic hosts rely on bacterial partners for fundamental aspects of their metabolism, such as providing key nutrients absent or insufficient in the host diet [4, 6, 16, 17]. In fact, the ecological variety of insects on earth is due in part to their ability to form new niches in previously inhospitable nutritional conditions, an accomplishment often achieved via association with bacterial partners [18,19,20]. The European honey bee, Apis mellifera, is an excellent exemplar of this phenomenon, whereby the gut microbiome allows the colony to subsist upon recalcitrant plant pollen and nectar [21,22,23,24].
Honey bees are essential for pollinating food crops, resulting in a multi-billion-dollar global industry [25]. However, managed honey bee colonies have suffered substantial losses in the last two decades [26]. Beekeepers in the United States reported losing 40.5% of their managed colonies between 2015 and 2016 alone [27]. These declines are often credited to a combination of stressors, the crux of which is poor nutrition [1, 28, 29]. Poor nutrition for managed honey bee colonies is partly due to dwindling natural floral resources and an increase in reliance on large monoculture crops [30, 31]. These large monocultures pose especially difficult nutritional landscapes for honey bees, as pollen from most individual crops only barely provides a colony’s base nutritional requirements [32, 33]. This discrepancy between available and required protein is most problematic for larvae, the immature stage of honey bee workers [34]. Ample multifloral pollen protein is required for honey bee larval development, and insufficient protein during this stage can have cascading effects through the colony [2, 34]. Larvae deprived of adequate protein mature into adults who are stunted in size and deficient in their ability to forage for floral resources, further exacerbating nutritional stress to the next generation of brood and compromising colony dynamics [2, 35]. Additionally, honey bees raised on pollen containing inadequate protein are significantly more likely to fall victim to secondary stressors such as viral pathogens, Varroa destructor mites, and Nosema apis infection [1, 29, 36, 37].
However, the microbiome can significantly modulate larval nutrition in holometabolous insects [6, 38, 39]. The adaptive decoupling of larval and adult phases in holometabolous insects creates an opportunity for growth-promoting nutritional symbionts to associate specifically with growing larvae [40, 41]. The most extensively studied examples of nutritional symbionts of larvae come from studies in Drosophila melanogaster, where just a single member of the microbiome can rescue larval growth despite severe protein limitation [11, 42]. Yet less is known about how the microbial communities associated with honey bee larvae contribute to their nutrition and development. Honey bee larvae are nurtured by their adult nestmates, who feed them a larval diet of nectar, pollen, and royal jelly [43]. This larval diet is relatively low in bacterial diversity and is occupied predominantly by the bacterium Bombella apis (formerly Parasaccharibacter apium), Lactobacillus, and Fructobacillus species [44, 45]. B. apis is consistently associated with honey bee larvae, larval diet, and the adult glands which secrete royal jelly, but is not found in large numbers in the adult worker gut [44,45,46,47]. Therefore, B. apis seems particularly well positioned to serve a nutritional role in honey bee larval development.
Here, we present data showing that indeed, B. apis is supplementing honey bee larvae through its secretion of essential amino acids. We first asked which bacterial members of the honey bee larval microbiome community can survive in the in vitro larval diet by subjecting a panel of strains to media containing a gradient of royal jelly. We subsequently performed a comparative genomic analysis across all sequenced strains of Bombella and related Saccharibacter to identify significant gene conservation and gene gain/loss. This study confirmed that all B. apis strains can synthesize all amino acids and have acquired multiple amino acid transporters. We then selected one strain of Bombella apis, A29, and performed a microbiological and metabolomic analysis of its metabolic potential as a nutritional mutualist. Finally, we modified an established in vitro larval rearing protocol to measure the effect of B. apis dietary supplementation on the growth of larvae experiencing nutritional stress. Our results strongly support the hypothesis that B. apis is a nutritional symbiont of honey bee larvae.
Materials and methods
Culturing bacteria
Bombella apis A29 was grown at 34 °C, ambient oxygen, shaking at 250 rpm, in Bacto-Schmehl (BS) liquid media. BS growth medium is derived from the larval diet outlined in Schmehl et al., 2011 conceived for growth of the honey bee larvae. BS bacterial growth medium is composed of 5% w/v D-glucose, 5% w/v D-fructose, 1% yeast extract, 4% v/v 5X Sigma (St. Louis, MO, U.S.A.) M9 salt solution (catalog #M9956), 0.2% v/v cation solution (1% [v/v]; 100 mM MgSO4 and 10 mM CaCl2 in diH2O), and 84.8% milli-Q H2O. The cation solution must be autoclaved separately from the M9 salt solution. The final pH of BS is 6.5. The designation BS is used with the written permission of Dr. Daniel Schmehl. Minimal Bacto-Schmehl Media (mBS) replaces the yeast extract portion of the base BS recipe with 1× Sigma MEM vitamin solution (catalog #M6895), plus either 1× Sigma MEM amino acids solution (catalog #M5550) or 1× Sigma non-essential amino acid solution (catalog #M7145).
Bacterial growth in honey bee larval diet
Overnight cultures of each bacterial strain were grown in BS broth (Bombella apis A29, B. apis B8, B. apis C6, B. apis SME1, B. apis MRM1T, Lactobacillus kunkeei AJP1, and Fructobacillus fructosus AJP3). Cultures were washed (cells spun down in microcentrifuge tubes and resuspended) twice in sterile phosphate-buffered saline (PBS), then normalized to 107 CFU/ml. 50 μl of each bacterial suspension was added to 500 μl of BS media with an increasing proportion of Glory Bee (Eugene, OR, U.S.A.) royal jelly up to 50%, in triplicate. After 24 h incubating at 34 °C, samples were serially diluted and plated on BS agar, in triplicate. CFUs were counted to determine numbers of viable cells.
Comparative genomic content analyses, identification of orthologs, and gain/loss analysis
To define orthologs, protein sequences were extracted from NCBI annotated sequence files for the Acetobacteraceae clade rooted on Gluconobacter. Reciprocal best BLAST hits were calculated, and genes clustered into ortholog groups using complete linkage. Conserved core orthologs were used to generate the species tree for these genera and this was used, in conjunction with GLOOME [48, 49] to infer branch-specific gene gain/loss events (Supplementary Table 2). To define presence/absence of amino acid biosynthesis genes (Supplementary Table 1), ortholog representatives were run against GapMind [50] to find amino acid biosynthetic genes in the proteomes. In addition, annotation based on NCBI’s PGAP was used, in conjunction with DOE’s IMG/M, to confirm the putative function of orthologous groups of genes.
Minimal media assay
An overnight culture of B. apis A29 was grown in BS broth, then washed in sterile PBS before inoculating mBS media containing either a complete amino acid solution or a non-essential amino acid solution. Cultures were diluted 1:100 in mBS and incubated at 34 °C for 48 h. Optical density (OD600) was measured using a spectrophotometer at the start of the experiment and at 48 h.
Larval collection and rearing
All larvae were collected from hives at the Indiana University Research and Teaching Preserve Bayles Road field site in October 2019. To minimize any effects of manipulation of the bee larvae and of genetic differences between colonies, all larvae were evenly distributed between treatments with respect to colony of origin and order of collection.
First-instar larvae were grafted from comb using a plastic grafting tool and were deposited into plastic queen cups in 48-well plates. Grafting and rearing protocols were conducted according to the Schmehl et al. 2011 protocol with several deviations noted here. Each larva was grafted into 10 μl UV-sterilized larval Diet A at the field site over the course of 2 h [50]. The larvae were transported back to the laboratory and incubated in darkness at 34 °C and 90% relative humidity. After overnight incubation, the larvae that did not survive grafting were removed and the remaining larvae were divided into experimental groups.
All larvae were fed according to the diet recipes of Schmehl et al., 2011. Larval diet was made no more than 48 h in advance of each feeding. All diet was UV-sterilized for 20 min to remove any potential bacterial contamination. Diet was refrigerated at 4 °C between feedings and warmed to 34 °C prior to each feeding. Larval diet was dispensed to individual larvae under a laminar flow hood using sterile pipettes. Larvae were fed mid-afternoon across all experiments.
The larval feeding and bacterial supplementation timeline is as follows: On day zero, larvae were grafted from the field. Larvae were fed 10 μl of Diet A on day 1, 20 μl Diet B on day 2, 30 μl Diet C on day 3, 40 μl of Diet C on day 4, and 50 μl of Diet C on day 5. Larvae were given 5 μl of the bacterial cell suspension on days 2 through 5. Diets A, B, and C are composed of varying proportions of royal jelly, glucose, fructose, yeast extract, and water. Details of the larval diet recipes can be found in Schmehl et al., 2011.
In all supplementation experiments, high-diet larvae received undiluted larval diet according to Schmehl et al., 2011 recipes. Low-diet larvae received larval diet from the same batches, divided and diluted by 25% using sterile deionized water. Each batch of diet was divided and diluted prior to UV-sterilization.
On the 6th day following larval grafting, larvae had consumed all remaining diet and defecated in their cells. Each larva was then removed from its cell and individually weighed. Residual diet and excrement were removed from the surface of each larva using a modified plastic grafting tool prior to weighing. Any larvae that were accidentally punctured during the weighing process were not weighed.
Bacterial supplementation of larval diet
An overnight culture of B. apis A29 was washed twice to remove excess media, then resuspended in sterile PBS. Absorbance was measured using a spectrophotometer to confirm adherence to known OD/CFU. Prior to larval feeding, bacterial suspensions were normalized to 104 CFU/ml using PBS. For experiments involving heat-killed controls, this normalized solution was then divided, and half was subjected to boiling for 10 min. Five μl of bacterial suspensions or PBS was pipetted into each queen cup containing a single larva. Bacterial suspensions were given immediately after daily feeding. All feeding and supplementation was performed using sterile technique under a laminar flow hood. Larval masses were compared in R using pairwise Mann–Whitney U-tests, then Bonferroni corrected for multiple comparisons.
Metabolomic analysis
On the 5th day following larval grafting, samples were taken from the larval diet of in vitro-reared larvae for metabolomic analysis. Eight hours after diet administration and bacterial supplementation, 3 μl of diet was removed from each larval cell. Samples were combined based on treatment, yielding 12 μl samples representing diet from four individual larvae. These 12 μl samples were immediately flash-frozen in liquid nitrogen and stored at −80 °C before gas chromatography-mass spectrometry (GC-MS). Samples were randomized prior to GC-MS to control for variation between individual GC-MS runs.
GC-MS analysis of larval diet samples were conducted using a modified version of a previously described method [51]. Briefly, 12 ml of larval diet was dissolved in 800 ml of prechilled (−20 °C) 90% methanol containing 2 μg/ml succinic-d4 acid. The sample was incubated at −20 °C for 1 h and centrifuged at 20,000 × g for 5 min at 4 °C. 600 ml of the supernatant was transferred into a new 1.5 ml microcentrifuge tube and dried overnight in a vacuum centrifuge. Dried samples were resuspended in 40 μl of 40 mg/ml methoxylamine hydrochloride (MOX) dissolved in anhydrous pyridine and incubated at 37 °C for 1 h in a thermal mixer shaking at 600 rpm. Samples were then centrifuged for 5 min at 20,000 × g and 25 μl of supernatant was transferred into an autosampler vial with a 250 μl deactivated glass microvolume insert (Agilent 5181-8872). 40 μl of N-methyl-N-trimethylsilyltrifluoracetamide (MSTFA) containing 1% TMCS was then added to the sample, at which point the autosampler vial was capped and placed at 37 °C for 1 h with shaking (250 rpm).
One μl of sample was injected into an Agilent GC7890-5977 mass spectrometer equipped with a Gerstel MPS autosampler. Samples were injected with a 10:1 split ratio and an inlet temperature of 300 °C. Chromatographic separation was achieved using a 0.25 mm × 30 m Agilent HP-5ms Ultra Insert GC column with a helium carrier gas flow rate of 1.98 ml/min. The GC temperature gradient was as follows: Hold at 95 °C for 1 min. Increase temperature to 110 °C with a 40 °C/min ramp. Hold 2 min. Increase temperature to 250 °C with a 25 °C/min ramp. Increase temperature to 330 °C with a 25 °C/min ramp. Hold for 4 min. Extraction and GC-MS was performed by the Indiana University Mass Spectrometry Facility. Metabolites were initially identified by analyzing a set of standards and subsequently identified using the NIST2017 library. Metabolite concentrations were compared in R using pairwise Mann–Whitney U-tests, then Bonferroni corrected for multiple comparisons, or were normalized using Box Cox transformation prior to one-way ANOVA and Tukey HSD correction.
Results
Only Bombella apis persists in honey bee larval diet
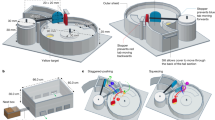
The honey bee larval diet comprises nectar, pollen, and royal jelly [34, 52]. Royal jelly has long been known to possess potent antimicrobial properties, due to its acidity, viscosity, and the presence of antimicrobial peptides [53,54,55]. Therefore, any bacterial mutualist exposed to the larval diet must be capable of tolerating this strongly antimicrobial environment. The honey bee larval microbiome has been characterized using 16 S rRNA gene amplicons and comprises a limited number of bacterial taxa, predominately Bombella apis, Lactobacillus kunkeei, and Fructobacillus fructosus (plus uncharacterized Lactobacillus spp. and Fructobacillus spp., and occasionally Bifidobacterium spp.) [46, 56]. To assess the ability of larvae-associated microbes to survive in royal jelly, we subjected bacterial strains to media containing a gradient of royal jelly (up to 50%) for 24 h and determined strain persistence by counting resulting CFUs (Fig. 1A). The inoculum concentrations were normalized to 107 CFU/ml for each strain. The 50% royal jelly treatment recapitulates the highest proportion of royal jelly used in standard in vitro larval rearing diets [50, 57]. The bacterial media used here and in following assays was developed by our lab for the growth of larvae-associated strains and downstream metabolic analysis and is based on the components of the honey bee larval diet (BS, see Methods). All five strains of Bombella apis assayed were able to survive at all levels of royal jelly (Fig. 1B). Some B. apis strains show a dip in the number of CFUs recovered between the media control and lowest concentration of royal jelly, indicating a degree of susceptibility to royal jelly inhibition (Fig. 1C). CFUs varied between strains, with B8 and C6 displaying the least reduction when royal jelly is first introduced. A29 and MRM1T show the greatest sensitivity to royal jelly addition, each exhibiting a 10-fold reduction in CFUs compared to media alone. In contrast, Lactobacillus kunkeei AJP1 and Fructobacillus fructosus AJP3 were highly sensitive to royal jelly addition. L. kunkeei AJP1 displayed a dose-dependent decline in the number of CFUs recovered at each increase in royal jelly in the media, with near total inhibition at 50%. F. fructosus AJP3 was unable to survive in any concentration of royal jelly beyond 10%. Overall, B. apis survival when challenged with royal jelly appears robust compared to other bacteria often identified as larvae-associated.
A The survival of a panel of larvae-associated bacteria in the presence of royal jelly was assessed by subjecting each to media containing a gradient of royal jelly from 0 to 50%. Strains were incubated overnight and plated on agar media to count CFUs. B Representative images of the spot-dilution plates used to count CFUs after incubation in 50% royal jelly. C Boxplots containing the total counts of CFUs resulting from each strain across all concentrations of royal jelly. Each concentration of all strains was calculated across three biological replicates (N = 3). Means are shown, with the upper and lower quartiles denoted by the upper and lower box boundaries, respectively. The upper and lower extremes of the data are indicated by the upper and lower whiskers.
B. apis can produce all essential amino acids
As B. apis can persist in larval diet, we reasoned that it may be able to metabolically transform it. To first consider how B. apis may modify the host diet, we used tools provided by the DOE’s (IMG/M) website to explore the metabolic capabilities of a sequenced strain, A29. The B. apis genomes encode the complete biosynthetic pathways required to produce all canonically essential amino acids (Fig. 2A). We then extended this result to all sequenced B. apis strains using an analysis of conserved orthologs; all sequenced B. apis strains retain the ability to synthesize all amino acids (Supplementary Table 1).
A Schematic diagram depicting the amino acid metabolic potential of B. apis A29 and highlighting a putative lysine/arginine exporter. Arrows represent enzymatic steps in biosynthetic pathways. Each amino acid that can be synthesized by B. apis A29 is positioned at the end of a pathway, with essential amino acids labeled in pink. B Boxplots representing the optical density achieved by B. apis A29 after incubating for 48 h in media containing either all 20 amino acids required for protein synthesis, or only non-essential amino acids. Each group contained at least five biological replicates (N = 5). Means are shown, with the upper and lower quartiles denoted by the upper and lower box boundaries, respectively. The upper and lower extremes of the data are indicated by the upper and lower whiskers.
To validate the genomic prediction that B. apis A29 can synthesize all essential amino acids, we created two minimal media (see Methods) containing either all amino acids, or nonessential amino acids only. When provided only nonessential amino acids, B. apis A29 showed no growth defects after 48 hours, arriving at a final OD600 similar to that observed when provided all essential amino acids in minimal media, confirming that B. apis A29 can synthesize all essential amino acids from nonessential precursors (Fig. 2B).
B. apis secretes lysine in the larval diet
After validating our metabolic predictions in culture, we next assessed how the presence of B. apis may modify the nutritional composition of the larval diet itself. We reasoned that if B. apis supplements the host with amino acids, it should encode amino acid transporters. Indeed, the B. apis A29 genome encodes a LysE/ArgO cationic amino acid transporter (Fig. 2A). We therefore used a gene gain/loss analysis across the phylogeny of all sequenced B. apis strains and related bacteria to identify branches at which amino acid transporters may have been acquired (Fig. 3). In the process of performing this analysis, using a larger number of strains, we recapitulated prior results, identifying the acquisition of gluconolactonase, of CRISPR-Cas cassettes, and of several restriction modification systems by Bombella (Supplementary Table 2) [58]. In addition to LysE/ArgO (WP_043561507), which appears to have been present in the ancestor of all taxa we assessed (Fig. 3), two cationic amino acid permeases (WP_154981533 and WP_154981532) were acquired in the ancestor of Saccharibacter and Bombella (Fig. 3, node N6), and two (WP_052349139 and WP_086431440) were more recently acquired in the ancestor of Bombella (Fig. 3, node N11). The above Genbank accessions reference the corresponding genes in Bombella apis strain SME1 (NZ_WHNS00000000). See Supplementary Table 3 for accessions of other species.
Phylogenetic tree generated from conserved core orthologs across the included strains (accessions found in Supplementary Table 3). Predicted acquisitions of cationic amino acid transporters indicated at arrowheads and nodes are numbered to facilitate in-text reference.
In order to experimentally confirm the secretion of lysine by B. apis, we next performed metabolomics on larval diet. Due to the logistical challenge of acquiring large quantities of natural larval diet, we relied on the synthetic larval diet used in the process of in vitro rearing of honey bee larvae, developed by Schmehl et al, 2011. This synthetic larval diet is compositionally like natural larval diet and is sufficient to rear honey bee larvae to adulthood [50]. To determine how B. apis A29 modifies the larval diet, we, performed GC-MS on samples of diet incubated with either live or heat-killed B. apis A29. We selected this strain because its genome has been previously sequenced, and it grows reliably under laboratory conditions. The heat-killed control allowed us to subtract the nutritional contribution of raw bacterial biomass provided by lysed cells and focus on the output of B. apis A29’s active metabolism in the larval diet. We were able to identify and quantify the following 13 amino acids in our GC-MS output: alanine, valine, leucine, isoleucine, proline, glycine, serine, threonine, methionine, glutamate, aspartate, asparagine, and lysine. Of these, we considered the 7 amino acids glycine, alanine, asparagine, aspartate, glutamate, proline, and serine to be “non-essential,” and to correspond to our minimal, defined medium (mBS), which included MEM non-essential amino acids. Larval diet supplemented with live B. apis A29 contains significantly higher levels of total essential amino acids (Fig. 4A, one-way ANOVA, Tukey HSD, p < 0.001). Conversely, nonessential amino acids are significantly lower when live B. apis A29 is present (Fig. 4A, one-way ANOVA, Tukey HSD, p = 0.03), while total TCA cycle intermediates are not significantly affected (Supplementary Fig 1, one-way ANOVA, Tukey HSD, p = 0.631). These patterns suggest an upcycling by B. apis of nonessential dietary amino acids into essential amino acids, which are then taken up by the host. The significant increase in total essential amino acids is driven largely by a more than twofold increase in lysine (Fig. 4B, one-way ANOVA, Tukey HSD, p < 0.001). Essential amino acids significantly decreased by live B. apis include methionine and threonine (one-way ANOVA, Tukey HSD, p < 0.001 and p < 0.001, respectively), both of which rely on the same metabolic precursors as lysine. Nonessential amino acids significantly decreased by B. apis were glutamate and serine (Supplementary Fig 2, one-way ANOVA, Tukey HSD, p = 0.002 and p = 0.011, respectively). Though not statistically significant, proline was the only nonessential amino acid increased by the presence of live B. apis (Supplementary Fig. 1, one-way ANOVA, Tukey HSD, p = 0.087). Together these data reveal that B. apis can dramatically impact the amino acid content of the honey bee larval diet, largely by increasing dietary lysine.
A Boxplots showing the total peak areas of essential and non-essential amino acids in synthetic larval diet after incubating with either live (blue) or heat-killed (red) B. apis A29. Larval diet incubated with live B. apis A29 contained significantly higher total essential amino acids (p < 0.001) and significantly lower total non-essential amino acids (p = 0.03). B Boxplots showing the peak areas of individual essential amino acids in synthetic larval diet after incubating with either live (blue) or heat-killed (red) B. apis A29. Live B. apis A29 results in significantly higher dietary lysine (p < 0.001) and significantly lower methionine (p < 0.001) and threonine (p < 0.001). Each group contains six biological replicates (N = 6). The following 13 amino acids were identified in our GC-MS output: alanine, valine, leucine, isoleucine, proline, glycine, serine, threonine, methionine, glutamate, aspartate, asparagine, and lysine. Of these, we considered the 7 amino acids glycine, alanine, asparagine, aspartate, glutamate, proline, and serine to be “non-essential.” Means are shown, with the upper and lower quartiles denoted by the upper and lower box boundaries, respectively. The upper and lower extremes of the data are indicated by the upper and lower whiskers. Significant differences in peak area were determined using. one-way ANOVA and corrected using Tukey HSD. Prior to ANOVA, data was normalized using Box Cox transformation.
B. apis bolsters honey bee larval growth under nutrient scarcity
To assess whether the observed metabolic modifications of the larval diet by B. apis translate to ecologically meaningful outcomes for honey bee larvae, we conducted an in vitro rearing experiment testing the impact of dietary B. apis under different diet conditions. Larvae were grafted at first instar from naturally mated colonies into sterile multiwell plates containing axenic in vitro rearing diet. This approach allowed us to modify the microbial content of the diet as well as the nutrition the larvae received; however we are unable to sterilize field-collected larvae to create axenic individuals. We raised larvae under sterile conditions on either synthetic diet (nutrient-rich) or diet that had been diluted with water by 25% (nutrient-poor) and supplemented them daily with 5 μl of sterile PBS or with a 105 CFU/ml solution of live B. apis A29 in PBS. As B. apis does not proliferate in the larval diet (Fig. 1), the bacterial biomass in this inoculum likely does not a constitute a significant nutritional source for larvae. Larvae were individually weighed at the end of their larval period, just before pupation and after evacuation of the larval gut. In this experiment, larvae reached masses between 50.9 mg and 158.5 mg (mean 133.4 mg, standard deviation 22.4 mg), well within range of published masses for honey bee 5th instar larvae [59, 60]. As expected, dropping the nutritional content of the larval diet by 25% resulted in an average weight drop of 17% in PBS-supplemented control larvae. Therefore, our nutrient limitation treatment translated to phenotypic differences in the 5th instar larvae. Larvae absent their symbiont but subjected to nutrient-poor conditions were significantly smaller than those in nutrient-rich conditions (Fig. 5, Mann–Whitney U test, Bonferroni correction, p = 0.013). In contrast, larvae in the nutrient-poor condition supplemented with B. apis A29 were able to reach the same masses as those in nutrient-rich conditions (Fig. 5, Mann–Whitney U test, Bonferroni correction, p = 0.179). We also noticed that the variance in the masses reached by larvae in the nutrient-poor condition absent their symbiont was significantly greater than the variance of those in the nutrient poor condition but given B. apis (Fig. 5, Levene’s Test of Equal Variance, Bonferroni correction, p = 0.017)—from 53.71 mg in the B. apis-supplemented group to 1000.25 mg in the un-supplemented group. PBS control larvae in nutrient-limited conditions reached masses as low as 50.9 mg, while those supplemented with B. apis under the same nutrient-limited conditions reached more than twice the mass (114 mg for the smallest individual). While there was no statistically significant difference between PBS and B. apis-supplemented larvae in the nutrient poor condition (Fig. 5, Mann–Whitney U test, p = 0.527), on average, nutrient-poor PBS control larvae were 7% smaller than those supplemented with B. apis. Overall, these results indicate that B. apis can rescue growth under nutrient limitation. Though nutrient-limited larvae in the PBS group were smaller at prepupation, developmental time was the same between all groups, as indicated by the purging of gut contents on the fifth day of feeding. Coupled with our metabolomic findings, these data showing a growth buffering phenotype of larval bees experiencing poor nutrition indicate a role for B. apis as a nutritional mutualist of honey bee larvae.
Boxplot showing the masses of individual larvae after receiving either synthetic larval diet or diet diluted 25% with water, plus either live B. apis A29 or sterile phosphate-buffered saline (PBS). Larvae given diet supplemented daily with B. apis A29 show no significant difference in mass between full or diluted diet (p = 0.193). Among larvae given PBS only, those receiving diluted diet are significantly smaller than those receiving undiluted diet (p = 0.013). For larvae receiving undiluted diet, B. apis-supplemented larvae N = 12, and PBS-supplemented larvae N = 12. For larvae receiving diluted diet, B. apis-supplemented larvae N = 10, and PBS-supplemented larvae N = 16. Means are shown, with the upper and lower quartiles denoted by the upper and lower box boundaries, respectively. The upper and lower extremes of the data are indicated by the upper and lower whiskers. Significant differences in mass were determined using the Mann–Whitney U test with Bonferroni correction.
Discussion
The ability of Bombella apis to not only survive, but meaningfully modify the honey bee larval diet and buffer larval growth strongly implicates this bacterium as a nutritional mutualist of honey bee larvae. Insects who feed on nutritionally challenging diets must often rely on bacterial partners to supplement missing nutrients [4,5,6,7]. Many of the best documented examples of bacterial supplementation of host diet involve insects gaining essential amino acids from intracellular symbionts [4, 5, 38, 61, 62]. Such endosymbionts are typically ancient and obligate, resulting in loss of bacterial genes essential for life outside the host [19, 63,64,65,66]. These losses often result in metabolic dependencies between host and symbiont, or multiple symbionts [5, 6, 61, 67,68,69,70]. However, in the case of B. apis, its position in the larval diet niche appears to maintain selective pressure on free living traits such as complete biosynthetic pathways for the generation of all amino acids (Fig. 2A). The retention of these complete pathways suggests that at least some of the environments B. apis inhabits are amino acid poor; indeed, nectar and honey would be particularly nutrient poor [71, 72]. Since B. apis is associated with multiple in-hive environments such as the nurse crop, queen gut, and nectar, it is likely that B. apis is faced with a variety of nutritional environments that necessitate metabolic autonomy.
In previous studies of the honey bee microbiome, bacterial taxa categorized as larvae-associated may have been identified based on DNA that was extracted from living cells or from DNA that was ephemerally maintained from cells lysed in the diet [46, 56]. Our findings suggest that Bombella apis and, to a lesser degree, Lactobacillus kunkeei are living in this niche, and other frequently sequenced bacteria such as Fructobacillus fructosus are likely not. This observation is in line with previous genomic evidence establishing B. apis as a honey bee-associated bacterium [58]. Hosts who associate with horizontally acquired symbionts require a means of winnowing beneficial partners from environmental microbes, a phenomenon that has been predominantly studied in the context of selectivity of host tissues [73,74,75,76]. However, honey bees mature in a built environment which can itself exert selective pressures on bacterial assemblages. In the case of the larval niche, it appears that the presence of royal jelly selects strongly for Bombella apis strains to associate with growing larvae, where they may improve colony health by supplementing the larval diet.
B. apis A29 appears to be shunting its metabolic energies into production of the essential amino acid lysine, which may be particularly valuable to developing larvae. Lysine appears to play an important role in other holometabolous symbioses; bacterial lysine synthesis and export is crucial for whitefly reproduction, and lysine synthesis is maintained in two Campotonus ant symbionts despite genome-wide erosion of central metabolic genes [70, 77, 78]. Indeed, our finding that cationic amino acid permeases have been gained and maintained suggests that in the evolution of Bombella in honey bee association, the transport of amino acids, such as lysine, was an important trait. This trait may have been maintained by selective pressures unrelated to symbioses, as the secretion of lysine by bacteria can help to alleviate feedback inhibition for the biosynthesis of multiple amino acids (such as lysine, threonine, isoleucine, and methionine). The phosphorylation of aspartate, the first step in the synthesis of all four of these amino acids, is inhibited by concentrations of any one of them. In addition, secretion of lysine can alleviate inhibition via lysine sensitive riboswitches caused by accumulation of intracellular lysine [79]. It is also important to note that without a characterized model, we do not know which direction the LysE/ArgO transporter we have identified in B. apis transports lysine. In some model systems, LysE/ArgO transporters have been observed to secrete lysine and be characterized as lysine efflux permeases [79, 80]. Indeed, the amino acid sequence of our LysE/ArgO homolog is 43% identical to that of the E. coli homolog, which mediates export of L-lysine [81]. However, without further characterization, we can only correlate the acquisition of two cationic amino acid permeases and the presence of the LysE/ArgO transporter in B. apis with the metabolomic phenotype we observe. An essential role of lysine on adult honey bee mass was revealed by de Groot in a 1952 study where the author measured adult mass after withholding individual amnio acids from the adult diet. Adult bees deprived of lysine suffered greatly reduced mass relative to those on complete diets [33]. Further, many commercial crops which rely on honey bee pollination services only barely meet an adult bee’s minimum lysine requirements [32]. Both of these studies focused on adult bees, yet we know that the nutritional demands of larvae are more dire [2, 34, 35]. It is easy, therefore, to imagine a scenario where a honey bee colony must rely on mutualistic bacteria such as B. apis to fill in the nutritional gaps in the larval diet.
Our finding that B. apis can bolster larval growth against poor nutrition is especially striking considering the limitations of the honey bee in vitro larval rearing system. In our experiment, though raised in sterile conditions, larvae were grafted from field honey bee colonies and could not be made axenic. It is therefore possible that the larvae in our larval rearing experiment (Fig. 5) harbored residual B. apis from the colony. In addition, the synthetic diet fed to in vitro-reared larvae was developed to ensure maximal survival of larvae in a laboratory setting. In nature, the larval diet fed to developing larvae depends on both the nutritional quality of foraged nectar and pollen, and on the nutritional status of the nurse bees who produce royal jelly. It is therefore possible that in a pollen-stressed colony, the nutritional content of the royal jelly would be highly variable as nurses themselves experience starvation. In such a context, the contribution of B. apis may be more crucial to larval growth than our in vitro experiment can capture. Conversely, in a colony experiencing ample multifloral protein, the contribution of B. apis to larval growth may be small. During such times, the established role of B. apis as an antifungal symbiont may favor its maintenance [82]. These and other context-dependencies of the symbiotic nature of B. apis in the larval niche should be explored in future work.
Conclusions and future work
Bombella apis is the only honey bee larvae-associated microbe that can survive in royal jelly. It synthesizes all essential amino acids and secretes lysine in larval diet. The presence of B. apis bolsters honey bee larval mass during nutrient scarcity, which can have dramatic downstream consequences for honey bee colony health. All these data point to the importance of B. apis in a colony. However, we have not linked the lysine secretion directly to honey bee nutritional supplementation. Future work will focus on genetic modification of B. apis to squarely implicate the cationic amino acid transporters and/or amino acid biosynthetic pathways to honey bee nutrition. Also, although we have performed a comparative genomic analysis on multiple strains and determined that all strains are capable of synthesizing and secreting lysine, it remains to be determined whether there is variation in their nutritional bolstering. It is conceivable, that some strains are better mutualists than others, and the link between genetic and phenotypic diversity in B. apis is an active area of research in the lab. Indeed, based on our data (Fig. 1), we might suspect that different strains are better able to survive and supplement larvae in the royal jelly diet. In addition, genetic differences in the honey bee larvae in our experiments may account for some of the variance we observed in our experiments, although we were careful to randomize our sampling of larvae across treatments. The interaction between honey bee genetics and the microbiome is only starting to be explored and it would be a benefit in future experiments to at least control for genetic variation in a colony by using single drone inseminated queens. Finally, our experiments were performed on B. apis alone and it is likely that Lactobacillus kunkeei also plays some role in larval development given that (1) it is routinely isolated from larval niches and honey bee hive environments and (2) it can survive in the presence of some royal jelly. Understanding the interaction between B. apis and L. kunkeei will be important to understanding the role that these microbes play in honey bee larval development and nutritional supplementation.
Data availability
The datasets generated during and/or analyzed during the current study are available from the Dryad repository: https://doi.org/10.5061/dryad.n5tb2rbz1.
References
Dolezal AG, Toth AL. Feedbacks between nutrition and disease in honey bee health. Curr Opin Insect Sci. 2018;26:114–9.
Scofield HN, Mattila HR. Honey bee workers that are pollen stressed as larvae become poor foragers and waggle dancers as adults. PLoS ONE. 2015;10:e0121731.
McFall-Ngai M, Hadfield MG, Bosch TCG, Carey HV, Domazet-Lošo T, Douglas AE, et al. Animals in a bacterial world, a new imperative for the life sciences. Proc Natl Acad Sci USA. 2013;110:3229–36.
Akman Gündüz E, Douglas AE. Symbiotic bacteria enable insect to use a nutritionally inadequate diet. Proc R Soc B Biol Sci. 2009;276:987–91.
Wu D, Daugherty SC, Van Aken SE, Pai GH, Watkins KL, Khouri H, et al. Metabolic Complementarity and Genomics of the Dual Bacterial Symbiosis of Sharpshooters. PLoS Biol 2006;4:e188.
Bing X, Attardo GM, Vigneron A, Aksoy E, Scolari F, Malacrida A, et al. Unravelling the relationship between the tsetse fly and its obligate symbiont Wigglesworthia: transcriptomic and metabolomic landscapes reveal highly integrated physiological networks. Proc R Soc B Biol Sci. 2017; 284:20170360.
Itoh H, Jang S, Takeshita K, Ohbayashi T, Ohnishi N, Meng X-Y, et al. Host–symbiont specificity determined by microbe–microbe competition in an insect gut. Proc Natl Acad Sci USA. 2019;116:22673–82.
Flórez LV, Scherlach K, Miller IJ, Rodrigues A, Kwan JC, Hertweck C, et al. An antifungal polyketide associated with horizontally acquired genes supports symbiont-mediated defense in Lagria villosa beetles. Nat Commun. 2018;9:2478.
Kaltenpoth M, Göttler W, Herzner G, Strohm E. Symbiotic bacteria protect wasp larvae from fungal infestation. Curr Biol. 2005;15:475–9.
Oliver KM, Degnan PH, Hunter MS, Moran NA. Bacteriophages encode factors required for protection in a symbiotic mutualism. Science 2009;325:992–4.
Shin SC, Kim SH, You H, Kim B, Kim AC, Lee KA, et al. Drosophila microbiome modulates host developmental and metabolic homeostasis via insulin signaling. Science 2011;334:670–4.
Dedeine F, Vavre F, Fleury F, Loppin B, Hochberg ME, Boulétreau M. Removing symbiotic Wolbachia bacteria specifically inhibits oogenesis in a parasitic wasp. Proc Natl Acad Sci USA. 2001;98:6247–52.
Koropatnick TA, Engle JT, Apicella MA, Stabb EV, Goldman WE, McFall-Ngai MJ. Microbial factor-mediated development in a host-bacterial mutualism. Science 2004;306:1186–8.
Chun CK, Troll JV, Koroleva I, Brown B, Manzella L, Snir E, et al. Effects of colonization, luminescence, and autoinducer on host transcription during development of the squid-vibrio association. Proc Natl Acad Sci USA. 2008;105:11323–8.
Shikuma NJ, Pilhofer M, Weiss GL, Hadfield MG, Jensen GJ, Newman DK. Marine tubeworm metamorphosis induced by arrays of bacterial phage tail-like structures. Science 2014;343:529–33.
Freiberg C, Fellay R, Bairoch A, Broughton WJ, Rosenthal A, Perret X. Molecular basis of symbiosis between Rhizobium and legumes. Nature 1997;387:394–401.
Médigue C, Masson-Boivin C, Gilbert LB, Cruveiller S, Gris C, Batut J, et al. Experimental evolution of a plant pathogen into a legume symbiont. PLoS Biol. 2010;8:e1000280.
Brucker RM, Bordenstein SR. Speciation by symbiosis. Trends Ecol Evol. 2012;27:443–51.
Moran NA, McCutcheon JP, Nakabachi A. Genomics and evolution of heritable bacterial symbionts. Annu Rev Genet. 2008;42:165–90.
Moran NA, Tran P, Gerardo NM. Symbiosis and insect diversification: an ancient symbiont of sap-feeding insects from the bacterial phylum Bacteroidetes. Appl Environ Microbiol. 2005;71:8802–10.
Lee FJ, Miller KI, McKinlay JB, Newton ILG. Differential carbohydrate utilization and organic acid production by honey bee symbionts. FEMS Microbiol Ecol. 2018;94:fiy113.
Lee FJ, Rusch DB, Stewart FJ, Mattila HR, Newton ILG. Saccharide breakdown and fermentation by the honey bee gut microbiome. Environ Microbiol. 2015;17:796–815.
Zheng H, Nishida A, Kwong WK, Koch H, Engel P, Steele MI, et al. Metabolism of toxic sugars by strains of the bee gut symbiont Gilliamella apicola. MBio 2016;7:e01326–16.
Kešnerová L, Mars RAT, Ellegaard KM, Troilo M, Sauer U, Engel P. Disentangling metabolic functions of bacteria in the honey bee gut. PLoS Biol. 2017;15:e2003467.
Gallai N, Salles JM, Settele J, Vaissière BE. Economic valuation of the vulnerability of world agriculture confronted with pollinator decline. Ecol Econ. 2009;68:810–21.
Brodschneider R, Gray A, Adjlane N, Ballis A, Brusbardis V, Charrière JD, et al. Multi-country loss rates of honey bee colonies during winter 2016/2017 from the COLOSS survey. J Apic Res. 2018;57:452–7.
Kulhanek K, Steinhauer N, Rennich K, Caron DM, Sagili RR, Pettis JS, et al. A national survey of managed honey bee 2015-6 annual colony losses in the USA. J Apic Res. 2017;56:328–40.
Goulson D, Nicholls E, Botías C, Rotheray EL. Bee declines driven by combined stress from parasites, pesticides, and lack of flowers. Science 2015;347:1255957.
Dolezal AG, Carrillo-Tripp J, Judd TM, Allen Miller W, Bonning BC, Toth AL. Interacting stressors matter: Diet quality and virus infection in honeybee health. R Soc Open Sci. 2019;6:81803.
St Clair AL, Zhang G, Dolezal AG, O’Neal ME, Toth AL, et al. Diversified farming in a monoculture landscape: effects on honey bee health and wild bee communities. Environ Entomol. 2020;49:753–64.
Naug D. Nutritional stress due to habitat loss may explain recent honeybee colony collapses. Biol Conserv. 2009;142:2369–72.
Taha EKA, Al-Kahtani S, Taha R. Protein content and amino acids composition of bee-pollens from major floral sources in Al-Ahsa, eastern Saudi Arabia. Saudi J Biol Sci. 2019;26:232–7.
de Groot AP. Amino acid requirements for growth of the honeybee (Apis mellifica L.). Experientia 1952;8:192–4.
Brodschneider R, Crailsheim K. Nutrition and health in honey bees. Apidologie 2010;41:278–94.
Keller I, Fluri P, Imdorf A. Pollen nutrition and colony development in honey bees - Part II. Bee World. 2005;86:27–34.
Huang Z. Pollen nutrition affects honey bee stress resistance. Terr Arthropod Rev. 2012;5:175–89.
van Dooremalen C, Stam E, Gerritsen L, Cornelissen B, van der Steen J, van Langevelde F, et al. Interactive effect of reduced pollen availability and Varroa destructor infestation limits growth and protein content of young honey bees. J Insect Physiol. 2013;59:487–93.
Feldhaar H, Straka J, Krischke M, Berthold K, Stoll S, Mueller MJ, et al. Nutritional upgrading for omnivorous carpenter ants by the endosymbiont Blochmannia. BMC Biol. 2007;5:48.
Sannino DR, Dobson AJ, Edwards K, Angert ER, Buchon N. The Drosophila melanogaster gut microbiota provisions thiamine to its host. MBio 2018;9:e00155–18.
Hammer TJ, Moran NA. Links between metamorphosis and symbiosis in holometabolous insects. Philos Trans R Soc B Biol Sci. 2019;374:20190068.
Kowallik V, Mikheyev AS. Honey bee larval and adult microbime life stages are effectively decoupled with vertical transmisson overcoming early life perturbations. mBio 2021;12:e02966–21.
Storelli G, Defaye A, Erkosar B, Hols P, Royet J, Leulier F. Lactobacillus plantarum promotes Drosophila systemic growth by modulating hormonal signals through TOR-dependent nutrient sensing. Cell Metab. 2011;14:403–14.
Wright GA, Nicolson SW, Shafir S. Nutritional physiology and ecology of honey bees. Annu Rev Entomol. 2017;63:327–44.
Tarpy DR, Mattila HR, Newton ILG. Development of the honey bee gut microbiome throughout the queen-rearing process. Appl Environ Microbiol. 2015;81:3182–91.
Corby-Harris V, Snyder LA, Schwan MR, Maes P, McFrederick QS, Anderson KE. Origin and effect of Alpha 2.2 Acetobacteraceae in honey bee larvae and description of Parasaccharibacter apium gen. nov., sp. nov. Appl Environ Microbiol. 2014;80:7460–72.
Vojvodic S, Rehan SM, Anderson KE. Microbial gut diversity of Africanized and European honey bee larval instars. PLoS ONE. 2013;8:72106.
Kwong WK, Medina LA, Koch H, Sing KW, Soh EJY, Ascher JS, et al. Dynamic microbiome evolution in social bees. Sci Adv. 2017;3:e1600513.
Cohen O, Ashkenazy H, Belinky F, Huchon D, Pupko T. GLOOME: Gain loss mapping engine. Bioinformatics 2010;26:2914–5.
Price MN, Deutschbauer AM, Arkin AP. GapMind: Automated annotation of amino acid biosynthesis. mSystems 2020;5:e00291–20.
Schmehl DR, Tomé HVV, Mortensen AN, Martins GF, Ellis JD. Protocol for the in vitro rearing of honey bee (Apis mellifera L.) workers. J Apic Res. 2016;55:113–29.
Li H, Tennessen JM. Preparation of Drosophila larval samples for gas chromatography-mass spectrometry (GC-MS)-based metabolomics. J Vis Exp. 2018;136:e57847.
Rortais A, Arnold G, Halm MP, Touffet-Briens F. Modes of honeybees exposure to systemic insecticides: Estimated amounts of contaminated pollen and nectar consumed by different categories of bees. Apidologie 2005;36:71–83.
Buttstedt A, Mureşan CI, Lilie H, Hause G, Ihling CH, Schulze SH, et al. How honeybees defy gravity with royal jelly to raise queens. Curr Biol. 2018;28:1095–1100.
Fratini F, Cilia G, Mancini S, Felicioli A. Royal jelly: An ancient remedy with remarkable antibacterial properties. Microbiol Res. 2016;192:130–41.
Fontana R, Mendes MA, De Souza BM, Konno K, César LMM, Malaspina O, et al. Jelleines: A family of antimicrobial peptides from the royal jelly of honeybees (Apis mellifera). Peptides 2004;25:919–28.
Rokop ZP, Horton MA, Newton ILG. Interactions between cooccurring lactic acid bacteria in honey bee hives. Appl Environ Microbiol. 2015;81:7261–70.
Crailsheim K, Brodschneider R, Aupinel P, Behrens D, Genersch E, Vollmann J, et al. Standard methods for artificial rearing of Apis mellifera larvae. J Apic Res. 2013;52:1–16.
Smith EA, Newton ILG. Genomic signatures of honey bee association in an acetic acid symbiont. Genome Biol Evol. 2020;12:1882–94.
Kaftanoglu O, Linksvayer TA, Page RE. Rearing honey bees, Apis mellifera, in vitro 1: Effects of sugar concentrations on survival and development. J Insect Sci. 2011;11:96.
Aupinel P, Fortini D, Dufour H, Tasei J-N, Michaud B, Odoux J-F, et al. Improvement of artificial feeding in a standard in vitro method for rearing Apis mellifera larvae. Bull Insectol. 2005;58:107–11.
Hansen AK, Moran NA. Aphid genome expression reveals host-symbiont cooperation in the production of amino acids. Proc Natl Acad Sci USA. 2011;108:2849–54.
McCutcheon JP, McDonald BR, Moran NA. Convergent evolution of metabolic roles in bacterial co-symbionts of insects. Proc Natl Acad Sci USA. 2009;106:15394–9.
Shigenobu S, Watanabe H, Hattori M, Sakaki Y, Ishikawa H. Genome sequence of the endocellular bacterial symbiont of aphids Buchnera sp. Aps Nat. 2000;407:81–86.
Gil R, Silva FJ, Zientz E, Delmotte F, González-Candelas F, Latorre A, et al. The genome sequence of Blochmannia floridanus: Comparative analysis of reduced genomes. Proc Natl Acad Sci USA. 2003;100:9388–93.
McCutcheon JP, Moran NA. Extreme genome reduction in symbiotic bacteria. Nat Rev Microbiol. 2012;10:13–26.
Wernegreen JJ, Lazarus AB, Degnan PH. Small genome of Candidatus Blochmannia, the bacterial endosymbiont of Camponotus, implies irreversible specialization to an intracellular lifestyle. Microbiology 2002;148:2551–6.
McCutcheon JP, Moran NA. Parallel genomic evolution and metabolic interdependence in an ancient symbiosis. Proc Natl Acad Sci USA. 2007;104:19392–7.
Bennett GM, Mccutcheon JP, Macdonald BR, Romanovicz D, Moran NA. Differential genome evolution between companion symbionts in an insect-bacterial symbiosis. mBio 2014;5:e01697–14.
Husnik F, Nikoh N, Koga R, Ross L, Duncan RP, Fujie M, et al. Horizontal gene transfer from diverse bacteria to an insect genome enables a tripartite nested mealybug symbiosis. Cell 2013;153:1567.
Bao XY, Yan JY, Yao YL, Wang Y, Bin, Visendi P, Seal S, et al. Lysine provisioning by horizontally acquired genes promotes mutual dependence between whitefly and two intracellular symbionts. PLOS Pathog. 2021;17:e1010120.
Cotte JF, Casabianca H, Giroud B, Albert M, Lheritier J, Grenier-Loustalot MF. Characterization of honey amino acid profiles using high-pressure liquid chromatography to control authenticity. Anal Bioanal Chem. 2004;378:1342–50.
Baker HG. Non-sugar chemical constituents of nectar. Apidologie 1977;8:349–56.
Nyholm SV, McFall-Ngai MJ. The winnowing: Establishing the squid - Vibrios symbiosis. Nat Rev Microbiol. 2004;2:632–42.
Kikuchi Y, Hosokawa T, Fukatsu T. Insect-microbe mutualism without vertical transmission: a stinkbug acquires a beneficial gut symbiont from the environment every generation. Appl Environ Microbiol. 2007;73:4308–16.
Itoh H, Jang S, Takeshita K, Ohbayashi T, Ohnishi N, Meng XY, et al. Host–symbiont specificity determined by microbe–microbe competition in an insect gut. Proc Natl Acad Sci USA. 2019;116:22673–82.
Oono R, Anderson CG, Denison RF. Failure to fix nitrogen by non-reproductive symbiotic rhizobia triggers host sanctions that reduce fitness of their reproductive clonemates. Proc R Soc B Biol Sci. 2011;278:2698–703.
Brown BP, Wernegreen JJ. Genomic erosion and extensive horizontal gene transfer in gut-associated Acetobacteraceae. BMC Genom. 2019;20:1–15.
Vitreschak AG, Rodionov DA, Mironov AA, Gelfand MS. Riboswitches: the oldest mechanism for the regulation of gene expression? Trends Genet. 2004;20:44–50.
Meijuan X, Rao Z, Yang J, Dou W, Xu Z. The effect of a LYSE exporter overexpression on L-arginine production in Corynebacterium crenatum. Curr Microbiol. 2013;67:271–8.
Indurthi SM, Chou H-T, Lu C-D. Molecular characterization of lysR-lysXE, gcdR-gcdHG and amaR-amaAB operons for lysine export and catabolism: a comprehensive lysine catabolic network in Pseudomonas aeruginosa PAO1. Microbiology 2016;162:876–88.
Pathania A, Sardesai AA. Distinct paths for basic amino acid export in Escherichia coli: YbjE (LysO) mediates export of L-lysine. J Bacteriol. 2015;197:2036–47.
Miller DL, Smith EA, Newton ILG. A bacterial symbiont protects honey bees from fungal disease. mBio 2021;12:e00503–21.
Acknowledgements
We thank the IU Mass Spectrometry Facility and the graduate and undergraduate students in the Newton lab who helped with honey bee husbandry. This work was financially supported by a Project Apis m. research grant and an NSF Collaborative Research grant (2005306). Components of Figs. 1 and 2 were created with BioRender.com.
Author information
Authors and Affiliations
Contributions
Designed and performed research, AJ Parish and ILG Newton; Contributed new reagents or analytic tools, DW Rice, VM Tanquary, JM Tennessen; Analyzed data, AJP, DWR, ILGN; Wrote the paper AJP and ILGN.
Corresponding author
Ethics declarations
Competing interests
AJP and ILGN are cofounders of VitaliBee.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Parish, A.J., Rice, D.W., Tanquary, V.M. et al. Honey bee symbiont buffers larvae against nutritional stress and supplements lysine. ISME J 16, 2160–2168 (2022). https://doi.org/10.1038/s41396-022-01268-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-022-01268-x
This article is cited by
-
The honeybee microbiota and its impact on health and disease
Nature Reviews Microbiology (2024)
-
A phylogenomic and comparative genomic analysis of Commensalibacter, a versatile insect symbiont
Animal Microbiome (2023)
-
Hive Transplantation Has Minimal Impact on the Core Gut Microbiome of the Australian Stingless Bee, Tetragonula carbonaria
Microbial Ecology (2023)
-
Honey bee symbiont gives larvae a boost
Nature Reviews Microbiology (2022)