Abstract
Predicting the response of ocean primary production to climate warming is a major challenge. One key control of primary production is the microbial loop driven by heterotrophic bacteria, yet how warming alters the microbial loop and its function is poorly understood. Here we develop an eco-evolutionary model to predict the physiological response and adaptation through selection of bacterial populations in the microbial loop and how this will impact ecosystem function such as primary production. We find that the ecophysiological response of primary production to warming is driven by a decrease in regenerated production which depends on nutrient availability. In nutrient-poor environments, the loss of regenerated production to warming is due to decreasing microbial loop activity. However, this ecophysiological response can be opposed or even reversed by bacterial adaptation through selection, especially in cold environments: heterotrophic bacteria with lower bacterial growth efficiency are selected, which strengthens the “link” behavior of the microbial loop, increasing both new and regenerated production. In cold and rich environments such as the Arctic Ocean, the effect of bacterial adaptation on primary production exceeds the ecophysiological response. Accounting for bacterial adaptation through selection is thus critically needed to improve models and projections of the ocean primary production in a warming world.
Similar content being viewed by others
Introduction
Microorganisms dominate ocean biodiversity and play a major role in global ecosystem function. As Falkowski, Fenchel, and DeLong [1] aptly phrased it, Earth’s biogeochemical cycles are driven by microbial engines. The ocean microbes’ response to current climate changes has the potential to alter the global cycles of carbon and nutrients with likely feedbacks to the climate system [2, 3]. To improve our projections of ecosystem function such as primary production and better understand the future of climate, we need to assess both how ocean microorganisms respond to climate change and how their response impacts to the global environment [4].
The rise of sea surface temperature causes dramatic changes in the oceanic environment, such as increased stratification resulting in weaker nutrient fluxes from the deep sea [5], deoxygenation [6], and sea-level rise [7]. How ocean microorganisms are affected, and how these potential effects might propagate through the ocean ecological web is poorly known. In particular, primary production response to warming is hard to predict, even on short temporal horizons [8]. In some regions, not only the magnitude but even the very direction of these responses remains uncertain [9, 10], notably because of complex interactions between temperature, nutrient supply, and light.
Among ocean microorganisms, heterotrophic bacteria are key actors of nutrient cycling. They remineralize dissolved organic matter into nutrients (the “recycling pathway”), and re-direct otherwise lost organic matter to higher trophic links via grazing. Even though this “microbial loop” [11] is estimated to process about half of all primary production [12, 13], an explicit recycling pathway is often missing in models of global carbon and nutrient cycles, and instead heterotrophic bacteria are treated implicitly [14]. Models that represent the microbial loop explicitly [15, 16] point to complex interaction effects between recycling and sea-surface warming: at a given temperature, models that include the microbial loop often predict a reduction in net primary production (NPP) [16]; however, when warming is predicted to decrease NPP by models without a microbial loop, inclusion of the microbial loop can reverse that prediction [17]. It thus seems that the direction of changes in primary production can in part be explained by the microbial loop and the balance between new and regenerated production. Our work aims at providing a mechanistic explanation for the direction of primary production variations by focusing on the microbial loop response to sea-surface warming.
A potentially important component of this response that is completely missing from previous models is bacterial adaptation through selection, i.e., the selection of different, more adapted bacterial strains. Throughout this paper, “adaptation” should thus be understood as “adaptation through selection”, and be contrasted to the physiological responses of individual bacteria to ecological changes in the environment that we call “ecophysiological response”. The combined effects of bacterial adaptation through selection and ecophysiological response will be referred to as the “eco-evolutionary response”. Bacterial adaptation may have important consequences for the future of primary production: as individual cells respond physiologically to warming, population- and ecosystem-level effects may feed back to the microbial community and drive the selection of different bacterial strains. In return, such bacterial adaptation may alter the ecological state of the system, thus entangling ecological and evolutionary dynamics in a closed eco-evolutionary feedback loop [18]. The capacity of bacterial populations to adapt rapidly to temperature change through selection mechanisms has long been established in the laboratory [19] and evidence is mounting for the important role of adaptation in the response of whole microbial communities to environmental change [20, 21]. Because of heterotrophic bacteria’s large population sizes and short generation time [22] relative to the timescale of climate change, we expect bacterial adaptation to play a role in their response to ocean warming [23], potentially altering the balance between new and regenerated primary production. How much and where then become key questions.
Here we extend an ecological model of the ocean’s surface to include the microbial loop and account for bacterial adaptation in the ecosystem’s response to warming. As eco-evolutionary feedbacks drive joint and reciprocal changes in adaptive bacterial traits and the ecosystem, we computationally explore a large parameter space to predict the response of the microbial loop efficiency and the consequences for new and regenerated primary production. We seek to compare and contrast these eco-evolutionary responses to the responses predicted in the absence of selection, as in current global ocean models. By applying the model to environmental scenarios differing in baseline temperature and nutrient influx, we identify general biogeographic conditions under which heterotrophic bacterial adaptation through selection is expected to have a large influence on the response of ocean productivity to climate warming.
Methods
First, we introduce a simple ecosystem model describing fluxes of nutrients between phytoplankton, grazers, and an explicit compartment of heterotrophic bacteria. Temperature influences the physiology of all organisms in the system. Then we model bacterial adaptation using the microbial life-history evolution framework of Malik et al. [24] to focus on Bacterial Growth Efficiency (BGE), the fraction of resources allocated by a cell to growth, as a key integrative bacterial trait. Assuming that BGE can vary among bacterial strains [25, 26] under the constraint of a trade-off with the cell’s resource acquisition capacity [25, 27], we use evolutionary game theory (a mathematical framework to model selection among competing populations “playing” different strategies) to predict the optimal (adapted) BGE as a function of temperature. By feeding the temperature-dependent adapted BGE into the ecosystem model, we can evaluate the system’s “eco-evolutionary” response to warming. This can then be compared to the purely ecophysiological response (in the absence of bacterial adaptation), the difference between the two measuring the impact of bacterial adaptation.
Ecosystem model
The ecosystem model is adapted from biogeochemical modules of global circulation models, mainly Hasumi and Nagata [16] and Aumont et al. [14]. The backbone structure of our model is a standard “NPZ” model with three pools: nitrogen (often the common currency in models with only one nutrient, as stated in Sarmiento and Gruber [28]), phytoplankton, and zooplankton. We included the microbial loop as in Hasumi and Nagata [16], based on the seminal work of Bendsten et al. [29].
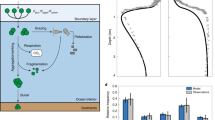
We designed our model to accommodate a 0D setting to study the balance between new and regenerated production at sea surface. This is why we divided the nitrogen pool into three: nitrate tracks new production, ammonium tracks regenerated production, and Dissolved Organic Nitrogen (DON) tracks microbial loop activity. Phytoplankton species are grouped into one compartment, and so are heterotrophic bacteria; both share zooplankton as one compartment of a common predator, allowing for both emergent bottom-up and top-down limitations. The outcome is a 6-compartment model called “NPZB” (Fig. 1), of which a complete mathematical description can be found in Supplementary note S1.
B, bacterial biomass. P phytoplankton biomass, Z zooplankton biomass, DON dissolved organic nitrogen. Fluxes driven by mortality are represented with dashed lines and all end up in the DON pool. Respiration fluxes are represented with dotted lines, ending in the ammonium pool. Red lines represent the incoming and outgoing fluxes of the system. All parameters and fluxes are defined in Supplementary Note S1.
As is done classically, we assume a type II (Monod) response of the uptake rates, U, to nutrient concentrations. To keep the model mathematically tractable, we assume that the response of grazing rates, G, to population densities is type I; this means that phytoplankton consumption by zooplankton is seldom at saturation. Maximum rates of uptake and grazing are temperature-dependent through an Arrhenius relationship, with different activation energies between autotrophic and heterotrophic organisms, ca. 0.3 eV for phytoplankton and 0.6 eV for bacteria and zooplankton [30]. See Supplementary note S1 for more detail.
Bacterial adaptation and the growth efficiency – resource acquisition trade-off
Predicting how microbial processes influence primary production can be addressed with phenotypic models in which bacterial metabolic traits influence microbial life history strategies [24, 31, 32], which in turn shape the interaction between mircrobial populations and their environment through resource consumption and waste production. Here we use Malik et al.’s [24] framework (rooted in Grimes’ [33] classical theory of life-history evolution) to describe and parametrize microbial life history variation in three principal dimensions: growth efficiency, resource acquisition capacity, and stress tolerance. As such, BGE acts as a “master trait”, resources not allocated to growth being distributed by the cell between resource acquisition mechanisms and tolerance to environmental stressors. Our model assumes no variation among strains in stress tolerance in order to focus on bacterial adaptation along the growth efficiency–resource acquisition trade-off and its ecological consequences. BGE is also a key determinant of the bacterial influence on nutrient cycling in the ocean [18], as different BGE strategies will lead to different fluxes of nutrients through the system, thus potentially affecting biogeochemical cycles at large.
Because of the direct influence of BGE on the life history of cells, we expect genetic variation in BGE [26, 27, 34] to be under intense selection [25], and thus BGE adaptation through selection to be a significant factor of variation between populations exposed to different environments. Evidence for this has been provided by the genomic studies of Roller et al. [26] and Saifuddin et al. [25] where support for the growth efficiency - resource acquisition trade-off is also presented. Here we extend our ecosystem model to predict BGE adaptation through selection under the constraint of this trade-off and investigate the ecological consequences.
In our model, BGE is represented by the parameter ω, and the fraction (1 − ω) of resources that is not invested by a cell into growth is respired. Bacterial respiration is used as a proxy for all processes involved in maintaining cell functions [35]. This includes processes that are central to resource acquisition and uptake, such as hydrolysis exoenzymes production [36], ATP production [37] or rRNA copy number [26]. As a consequence, a cell that invests more in bacterial growth will be less efficient in acquiring nutrients. To represent this trade-off between BGE ω and resource acquisition, we use bacteria-DON specific affinity [38], defined as the initial rate of DON uptake per capita per unit of DON concentration. Specific affinity is an increasing function of the respired fraction (1 − ω), so that a bacterium that invests quasi exclusively into growth (i.e., ω close to one) will be almost unable to perform DON uptake (i.e., an uptake rate nearing zero). Conversely, a bacterium that invests quasi exclusively into uptake (i.e., ω close to zero) will not grow efficiently. The optimal value ω∗ must then be between these two extremes. See Supplementary note S2 for the corresponding mathematical formalism.
For any given temperature within relevant limits, we use an evolutionary game theory approach [39, 40] to compute the optimal BGE value ω∗ as an evolutionary stable strategy. This means that a population of a single strain of bacteria with BGE ω∗ cannot be replaced by any bacterial strain with different ω value. Unless stated otherwise, we assume that at any given temperature the bacterial population evolves towards a single (monomorphic) evolutionary stable value ω∗. The optimal ω∗ can be calculated for that temperature and given all other environmental parameters (see Supplementary note S2 for details). A similar approach was developed in Abs and Ferrière [41] to model the eco-evolutionary dynamics of soil microbial populations exposed to changes in the physical or chemical properties of their environment.
Model analysis
We use analytical and numerical simulations to quantitatively predict the ecophysiological and eco-evolutionary responses of the ecosystem to a temperature increase denoted by ∆T. Specifically, we compare ecological and evolutionary steady states under three conditions:
-
1.
Initial adaptation: Initially, the system is locally adapted to a given sea-surface temperature, T0. The corresponding bacterial evolutionary stable strategy is denoted by ω0.
-
2.
Ecophysiological scenario: Here, temperature-dependent parameters respond to sea surface warming, from T0 to T1 = T0 + ∆T, but we control the model to prevent bacterial adaptation: BGE remains at ω = ω0. In this scenario, bacterial populations are ill-adapted to the new environment.
-
3.
Eco-evolutionary scenario: Here, as temperature rises from T0 to T1 = T0 + ∆T, the adaptation capacity of heterotrophic bacteria is included, and BGE evolves from ω0 at T0 to the new evolutionary stable value ω1 at T1. Selection may act on strains with slightly different trait values arising by mutations of small effect; as well as strains or species that were already present at low density and have larger differences in their trait value, as would be the case in an ecological guild. Irrespective of the mechanism of trait variation among strains, the adapted trait value will be the same, because of the uniqueness of the fitness optimum that our model predicts.
By comparing the ecosystem steady states under the ecophysiological and eco-evolutionary scenarios to the initial adaptation, we compute the ecophysiological and eco-evolutionary responses of the system to the warming increment ∆T. The difference between these two responses then measures the impact of bacterial adaptation through selection.
To analyze the responses of the microbial loop to warming, we focus on BGE, ω, and the microbial loop efficiency (MLE), η, which represents the fraction of primary production that cycles through bacteria and thus contributes to bacterial growth or respiration. To analyze the responses of primary production to warming, we focus on both new and regenerated production; we refer to the f − ratio as the ratio of new production over total primary production. The f − ratio is relevant for the study of the balance of new and regenerated production, but also export production: at ecological equilibrium, it is equal to the export ratio, or e − ratio [42], which can be denoted by the ef − ratio.
Here we report the results of 1000 simulations of the ecosystem equilibrium with parameters sampled from a Latin cube with credible parameter ranges (Supplementary Table 1). Parameter ranges were drawn from two main sources: Bendsten et al. [29] for microbial loop parameters and Aumont et al. [14] for other ecosystem processes. For parameters that are specific to our model (as in the functional response for zooplankton grazing), we followed [14] and selected ranges of values so that the ecosystem model outputs are of the same order of magnitude as empirical measurements reported in classical sources [28, 43]. A complementary set of simulations is run specifically to compare ecophysiological and eco-evolutionary responses under four contrasted environmental conditions of biogeographical significance: cold vs. warm and low vs. high nutrient input.
Results
Initial adaptation
Following adaptation to the initial temperature T0 (Fig. 2), all equilibrium state variables correlate negatively with T0, except for inorganic nutrients (Supplementary Fig. 1). Warmer temperatures accelerate fluxes between ecosystem compartments but do not increase nutrient input: this acceleration is thus made at the expense of equilibrium concentrations and biomass, which decrease across the temperature gradient (Fig. 2a). Conversely, higher nutrient input correlates with higher nutrient concentrations and population biomass.
The adaptive effect represents variation in the ecosystem state driven by bacterial adaptation through selection after a temperature increase. Black dots indicate median values. a Equilibrium biomass and concentrations. See Fig. 1 for notations. b Microbial loop and ecosystem production outputs (separated by the black vertical line). BGE bacterial growth efficiency. MLE microbial loop efficiency. All parameters vary in the default parameter ranges (Supplementary Table 1).
Correlations across state variables at equilibrium bear the signatures of top-down controls, with strong, negative correlations between inorganic nutrients concentrations and phytoplankton growth rate, and between phytoplankton biomass and grazing by zooplankton.
The f − ratio positively correlates with both BGE, ω0, and MLE, η. By re-injecting in the trophic chain a fraction of the primary production that would otherwise be lost in the DON pool, the microbial loop acts as a recycling process, thus driving the export ratio up.
Ecophysiological scenario: the direction of primary production variation is controlled by ecophysiological changes in the microbial loop
An increase in temperature changes multiple physiological parameters, causing a shift in the ecosystem equilibrium even in the absence of bacterial adaptation (green distributions in Fig. 2). The biomass of all populations (heterotrophic bacteria, phytoplankton, zooplankton) and all nutrients concentrations tend to decrease. Even though individual process rates tend to increase with temperature, the overall effect on the ecosystem equilibrium is negative because nutrient consumption by phytoplankton and grazing both increase.
The general decrease in phytoplankton biomass observed across the parameter space (Fig. 2a) is to be contrasted with the distribution of primary production. In some cases, primary production increases despite phytoplankton biomass decreasing (Fig. 2b). This decoupling is explained by faster phytoplankton metabolism, which increases per capita primary production. Whether the effect of faster metabolism outweighs the decrease in biomass and yields a gain of primary production depends on the balance between new and regenerated production. While new production always increases with temperature, this is not the case of regenerated production (Fig. 2b). Total primary production increases when the decrease in regenerated production is smaller than the increase in new production.
Both nutrient limitation and the relative temperature sensitivities of phytoplankton and zooplankton are at play in this balance (Supplementary Fig. 2) with the effect of nutrient limitation being stronger. Indeed, total Dissolved Inorganic Nitrogen (DIN) concentration (i.e., the sum of nitrate and ammonium concentrations) at initial temperature T0 is a key factor of the response of primary production to warming (Fig. 3a): positive in nutrient-rich ecosystems, negative in nutrient-poor ecosystems. Nutrient-rich ecosystems can sustain faster metabolism of phytoplankton in warmer conditions, driving primary production up. In nutrient-poor ecosystems, the increased maximum uptake rates result in stronger nutrient limitation, driving primary production down.
a Influence of nutrient availability. b Influence of microbial loop efficiency. Distribution of 1000 simulation outputs with parameters sampled in the ranges given in Supplementary Table 1 (see “Methods” for more detail). In (a), the dashed line is the linear regression through the whole set of simulation outputs. In (b), the red region indicates the kernel density of outputs for the resampled set of simulations that yielded dissolved inorganic nitrogen concentrations (DIN) lower than 1.5 mmol·m−3 (see Supplementary Table 2). The dashed line represents the linear regression for the resampled simulations.
In the case of nutrient-poor ecosystems, the negative effect of warming on primary production can be directly traced to a decrease in microbial loop activity (Fig. 3b). By resampling the set of parameters corresponding to nutrient-poor ecosystems (Supplementary Table 2), we find a significant positive correlation between MLE and the ecophysiological response of primary production to warming, which approaches zero in ecosystems where little to no change occurs in MLE. This pattern is due to the fact that in nutrient-poor ecosystems, the main component of primary production is regenerated production, which strongly depends on microbial loop activity. In nutrient-rich ecosystems, primary production is driven mainly by new production, and ammonium production relies equally on bacterial and zooplankton respiration, thus causing a decoupling between primary production and microbial loop activity. In other words, only in nutrient-poor ecosystems does primary production hinge on the microbial loop.
Eco-evolutionary scenario: bacterial adaptation in the microbial loop drives system export
We now assume that heterotrophic bacterial population can adapt to warming through selection. In response to a temperature increase ∆T, the adapted BGE shifts from the ω0 value adapted to the initial sea-surface temperature T0 to the new optimal value ω1 adapted to temperature T1 = T0 + ∆T (see Supplementary Note S2 and Supplementary Fig. 3). The concurrent change in ecosystem equilibrium state is the eco-evolutionary response of the system to the ∆T warming increment (red distributions in Fig. 2). By measuring the difference between the ecophysiological response (without bacterial adaptation) and the eco-evolutionary response (with bacterial adaptation) on the ecosystem state variables we can quantify the ecosystem impact of bacterial adaptation to warming (blue distributions in Fig. 2).
Bacterial strains with BGE lower than the initially adapted value turn out to be competitively superior in a warming ocean; as they replace the initial strain, BGE measured at the scale of the bacterial population decreases (Fig. 2b). This occurs in response to the change in ecosystem state following from the ecophysiological response to warming. The selection gradient of BGE (equation (20) in Supplementary Note S2) reveals the two main controls of selection on bacteria: top-down control by mortality (including grazing), and bottom-up control by DON-limitation. Because the eco-physiological response of mortality to warming is always positive, the corresponding top-down agent of selection on BGE always intensifies with warming, which favors bacterial strains with higher BGE. Yet the result of adaptation is a decrease in BGE, showing that the system is under a stronger bottom-up limitation. Indeed, there is a strong correlation between the decrease in DON concentration and the adaptive response of bacteria (Fig. 4a). In ecosystems with higher initial temperature, the increased mortality and relatively low decrease in DON concentrations results in weaker adaptive responses. In ecosystems with lower initial temperature, the decrease in DON concentration is stronger, resulting in a decrease in BGE. This is a consequence of the growth efficiency—resource acquisition trade-off, which favors resource acquisition when resource availability (here, DON concentration) drops.
a From ecology to evolution: BGE adaptation as a function of the ecophysiological response of bacterial resource (DON) to warming. Individual simulations are color-coded according to the initial temperature. b From evolution to ecology: ecological impact of BGE adaptation on new and regenerated production. All parameters vary in the default parameter ranges (Supplementary Table 1).
The effect of bacterial adaptation then ripples through the whole ecosystem in a trophic cascade (Figs. 2 and 4c, d, Supplementary Figs. 4–7): the decrease in BGE drives bacterial production and microbial loop activity down (Supplementary Fig. 4) while increasing ammonium concentrations (Supplementary Fig. 5a). This nutrient increase drives phytoplankton biomass up (Supplementary Fig. 5b), which adds pressure to the nitrate pool, thus decreasing the nitrate concentration (Supplementary Fig. 6a). These variations in nutrient concentrations cause an increase in new and regenerated production (Supplementary Fig. 7), but not evenly so, resulting in a decrease in f − ratio (Supplementary Fig. 6b).
The adaptation of BGE to warming causes both new and regenerated production to increase (Fig. 4b). Importantly, the net primary production variation due to bacterial adaptation to warming is of the same magnitude, or even greater than the ecophysiological response of the system (Fig. 2b). This underscores the importance of the microbial loop and its evolutionary adaptative capacity to predict the response of primary production and other ecosystem functions to warming.
While the ecophysiological response to warming is controlled primarily by the availability of nutrients (Fig. 3), the initial temperature of the system is also a major determinant of the eco-evolutionary response. In cold waters, warming causes a strong ecophysiological decrease in DON concentration, which selects for lower BGE. In warmer environments, the ecophysiological decrease in DON concentration is weaker, leading to a smaller adaptive response of BGE (Fig. 4a). Thus, we expect a two-dimensional gradient of temperature and nutrient availability to shape the pattern of microbial adaptation (Fig. 4a) and eco-evolutionary responses (Fig. 4b).
Warming causes different eco-evolutionary responses in different biogeographic regions
To further evaluate the ecosystem impact of bacterial adaptation to warming, we focus on four hypothetical biogeographic regions characterized by two different initial temperatures, T0 = 5o (cold) or 20o (warm), and nutrient input rates, φ = 0.03 day−1 (poor) or 0.1 day−1 (rich).
The temperature difference captures the expected contrast between tropical zones and high-latitude areas. The difference in nutrient input was chosen so that the low φ value yields nitrate concentrations around 0.5 mmol·m−3, which is typical of oligotrophic zones; and the high φ value yields intermediate nutrient concentrations, ca. 2.0 mmol·m−3. Warm and poor environments are typical of subtropical gyres, while high latitude ecosystems are typically cold and nutrient-rich.
For each of these four hypothetical bioregions, we performed 1000 simulations with parameters sampled from the same ranges as before (Supplementary Table 1) while T0 and φ are fixed at values assigned to the region. All output distributions are shown in Supplementary Figs. 8–19. Hereafter we focus on the responses of the microbial loop and primary production (Fig. 5).
a, c, e, g Ecosystem equilibrium at initial temperature, T0. b, d, f, h Ecophysiological (green) and eco-evolutionary (red) responses to warming with the difference between the two quantifying the effects of adaptation through selection (blue). Initial temperature, T0, and nutrient input, φ, are fixed, respectively at 5 °C or 20 °C for cold vs. warm regions, and 0.03 day−1 or 0.1 day−1 for nutrient-poor vs. nutrient-rich regions. All other parameters vary in the default parameter ranges (see Supplementary Table 1).
The initially adapted value of BGE tends to be larger in cold and/or nutrient-rich regions (Fig. 5a). With warming, these regions are also the ones where the adaptive change in BGE is the largest (Fig. 5b). As expected, primary production is in general much larger in nutrient-rich regions (Fig. 5c), with little influence of the initial temperature. In nutrient-poor regions, ecophysiological effects of warming tend to be negative, whereas the effect of bacterial adaptation is positive (Fig. 5d). Thus, in nutrient-poor regions, the decrease in primary production driven by the cophysiological response to warming may be compensated by bacterial adaptation. In regions that are both nutrient-poor and cold, the positive effect of bacterial adaptation may even exceed the negative ecophysiological effect, causing an increase in primary production (Fig. 5d).
Primary production shows contrasted ecophysiological and eco-evolutionary responses to warming. This pattern can be understood from the responses of new and regenerated production (Fig. 5e–h). The response of new production is similar in all regions (Fig. 5f), and the contrast in primary production comes from regenerated production variation (Fig. 5e). As seen before, the ecophysiological response of primary production is driven by regenerated production variation induced by nutrient availability--positive in nutrient-rich regions, negative in nutrient-poor regions. In contrast, the effect of bacterial adaptation is always positive (even if small on new production), but depends on the initial temperature of the environment: the effect is larger in initially cold environments, smaller in initially warm environments. This implies that the ecophysiological response of the system and the effect of bacterial adaptation add up in regions that are both cold and nutrient-rich, such as the Arctic Ocean, resulting in a strong increase in total primary production.
Discussion
Our study aims at improving our mechanistic understanding of nutrient cycling in the surface oceans and its response to climate warming. We asked how the rise of sea surface temperature alters the microbial loop activity and primary production, both ecophysiologically and through bacterial adaptation. Nutrient limitation turns out to be the main control of the ecophysiological response of primary production to warming, controlling microbial loop activity and thus the balance between new and regenerated production. We show that bacterial adaptation is an equally important factor of the response of primary production to warming, especially in cold oceanic regions. Our results highlight the importance of two often underestimated components of biogeochemical ocean models, namely the dependence of ecosystem production on the microbial loop, and the adaptive potential of microbial populations [44, 45].
Ecophysiological vs. eco-evolutionary predictions
When used to resolve the ecophysiological response of the ecosystem to sea surface warming, our model predicts general trends such as a decrease bacterial, phytoplankton, and zooplankton biomass and an increase in top-down controls. This is consistent with other predictions suggesting that oceans may become more oligotrophic over time [46], with stronger top-down control by grazing [30]. Our model predicts that despite this general decrease in population biomass, primary production may either increase or decrease with warming, confirming that biomass measurements alone are poor predictors of changes in primary production [47].
Changes in the balance of top-down and bottom-up controls of phytoplankton abundance are thought to be important to assess the future of primary production in the face of climate warming [10]. Our ecophysiological model provides a simple framework to evaluate this balance and predict the net response of primary production to warming. In agreement with existing data [5, 10], our model predicts nutrient-poor areas to be more prone to a decrease in primary production than richer regions, and provides a mechanistic explanation for this pattern. In nutrient-poor environments exposed to warming, regenerated production decreases faster than new production increases, resulting in an overall decrease in primary production. The decline in primary production is a result of a change in the balance between new and regenerated production due to a decrease in microbial loop activity, further confirming the important feedback of the recycling pathway in oligotrophic environments, as previously reported in Fenchel [12].
At ecosystem equilibrium, the f − ratio and the e − ratio are equal [28], meaning that new production equals export production. This allows us to predict the ecophysiological response of export production to sea-surface warming. Even though total primary production can decrease due to increasing temperatures in nutrient-poor environments, this response is fully driven by the decrease in regenerated production. New production increases across all simulated systems, which means that export production increases in all regions. Due to their fast metabolic rates and short generation time [22] relative to the timescale of climate change, heterotrophic bacteria have a strong potential to evolve and adapt rapidly to their changing environment. Our model was designed to predict the optimal strategy of heterotrophic bacteria for resource allocation into growth in a given environment, or Bacterial Growth Efficiency (BGE), under a trade-off with resource acquisition. As the environment changes, the optimal strategy changes, and the consequences for primary production can be evaluated.
As sea surface temperature rises, the optimal BGE responds to two opposing selective pressures, namely increased bacterial mortality and the depletion of dissolved organic matter. While nutrient abundance is the main driver of ecophysiological responses to warming, environmental temperature shapes BGE adaptation. We find that increasing temperature favors a bottom-up control of BGE, with decreasing concentrations of DON driving the optimal BGE down. The DON decrease is stronger in cold regions, resulting in a stronger adaptive response of bacterial populations.
These results provide new insights into the “link or sink” behavior of the microbial loop [12]. By recapturing otherwise lost organic matter and recycling or transferring it to higher trophic levels, the microbial loop can increase export production (the “link” behavior). On the other hand, by fixing nutrients in bacterial biomass, the microbial loop can effectively decrease export production (the “sink” behavior). Across all simulated systems, adaptation to warming drives BGE down and microbial loop activity decreases, with antagonistic consequences for export production. One effect is to decrease the export ratio (equal to the f − ratio in our model), acting as a sink. The opposing effect is to increase overall primary production, which increases export production, acting as a link. Both effects on export ratio and primary production are significant here, confirming the importance of taking both into account when assessing export production and its response to warming [17]. The increase in primary production is larger than the decrease of f − ratio, which eventually makes the “link” component of the microbial loop stronger.
We predict eco-evolutionary processes mediated by heterotrophic bacterial adaptation to shape contrasted biogeographical responses to warming, as bacterial optimal BGE follows a temperature gradient while ecophysiological responses follow a nutrient gradient. In nutrient-rich regions, bacterial adaptation and ecophysiological responses combine synergistically to increase both new and regenerated production. This is of particular significance in already productive cold and rich regions such as the Arctic Ocean [48], where productivity is expected to increase in the coming years [49]. Our model predicts an even larger increase due to bacterial adaptation. In nutrient-poor regions, the effects of bacterial adaptation oppose the ecophysiological decrease of primary production. In cold and poor regions, adaptation mitigates the decrease in primary production, going as far as to reverse it, whereas ecophysiology drives the overall response in regions that are both nutrient-poor and warm.
Our results underscore the need to take adaptive processes into account in predictive models of ocean productivity [50]. As BGE is often assumed to be independent of temperature for a given strain of bacteria [51, 52], it is usually set as constant in biogeochemical models, but we showed that natural selection acting on bacterial strains with varying BGE could impact our prediction of future primary production in the ocean.
Bridging eco-evolutionary modeling and sequence data
Our modeling approach derived from evolutionary game theory [53] is purely phenotypical in essence: BGE is treated as a quantitative character, heritable variation is assumed, and adaptation to a changing environment is predicted as the outcome of optimization under the constraint of a trade-off with resource acquisition. Under broad conditions [54, 55], the underlying genetic architecture of the traits and genetic mechanisms of their variation do not alter the phenotypic results. However, the study of microbial communities has benefited tremendously from the expansion of our molecular sequencing capability and genomic and other “omics” analytical toolbox. These advances provide support for underlying hypotheses in our models and present new opportunities to evaluate our key predictions, as we briefly summarize hereafter.
“Omics” studies can support and inform phenotypic models by providing data on genes, transcripts, proteins and metabolites to infer phenotypic trait variation within and between populations, and uncover some of the underlying mechanisms such as selection and constraints. BGE is a challenging trait to extract from omics data [24] because it compounds multiple underlying traits related to cellular maintenance, protein synthesis, and metabolic and respiratory pathways [26, 56]. Saifuddin et al. [25] circumvent these difficulties and predict BGE by using genome-scale metabolic modeling subject to flux balance analysis [57]. They resolve BGE variation across 200 bacterial taxa and provide support for the adaptive nature of this variation and for the growth efficiency—resource acquisition trade-off used in our model. This is in support of other studies combining comparative genomics with direct trait measurement by laboratory assays [26, 27]
Evidence from omics studies is also growing for the role of adaptive evolution in the response of whole microbial communities to environmental change [58,59,60]. New metagenomic computational pipelines hold much promise to extend pioneering evolutionary analyses done in small-scale systems such as the human gut microbiome [20, 21, 61] to ocean microbial communities. For example, bioinformatic pipelines for microbial community metagenome assembly such as GraftM [62] and MetaPop [63, 64] can resolve microbial metagenomic sequence data at intra-population level, identify Single-nucleotide polymorphisms (SNPs), calculate nucleotide diversity, and detect positive selection within populations. We expect such computational tools to greatly improve our ability to assess bacterial adaptive evolution within natural communities such as those involved in the ocean microbial loop.
Finally, ecosystem models that assume a simple relation between phenotypic traits and gene-encoded biochemical pathways [65] pave the way for the development of phenotype-based models of evolutionary adaptation that will directly simulate metagenomes and metatranscriptomes, hence opening the possibility to quantitatively test the predictions of models like ours with omics data.
Model extensions and conclusion
To our knowledge, this is the first model aimed to predict the effect of microbial adaptation on ocean productivity. Previous models of biological adaptation in warming oceans addressed the impact of adaptation through selection on phytoplankton community diversity [65,66,67,68,69]. Some of these studies suggest that adaptation may reduce community diversity especially in tropical regions, leading to a potential decrease in primary production in these regions [66]. Future work is warranted to probe the generality of this result, integrate bacterial and phytoplankton adaptation in a common framework, and extend the scope of potential adaptive responses to warming to a broader set of traits and trade-offs, and to other functional groups. We highlight two areas of interest.
Regarding the set of traits and trade-offs, our model focuses on the growth efficiency—resource acquisition trade-off, thus assuming that bacterial populations do not change along the stress-tolerance axis of life-history variation. This assumption could be lifted in future work. By using sequence data and computational tools to extract quantitative information about stress tolerance traits, we should be able to extend existing comparative genomic analyses to obtain a more comprehensive and quantitative understanding of bacterial phenotypic variation in all three axes of life-history evolution: growth efficiency, resource acquisition, and stress tolerance [24]. Such information is needed to account for stress tolerance in predictive models of bacterial adaptation. Once done, it will be possible to address the specific ecophysiological and eco-evolutionary effects of stressors such as extreme climatic events on microbial loop activity, and their consequences for carbon and nutrient cycling.
Regarding the inclusion of other functional groups, viruses are of particular interest because of their dramatic direct ecological impact (e.g., the viral shunt of material fluxes [12, 70, 71]), their indirect role as physiological hijackers and selective agents of their hosts [72], and their own capacity for record fast evolution and adaptation to ecological and environmental change. Existing ecosystem models provide a solid foundation for such developments [73], which will contribute to a general workflow for integrating micro-organisms evolutionary adaptation in global earth system models. Meta-omics tools and analyses that have been developed specifically for marine viral communities [74] should help identify key adaptive traits and trade-offs that would be captured in eco-evolutionary models.
In conclusion, bacteria adaptation to warming can drive changes in ocean primary production of the same magnitude as the ecophysiological response, especially in the most productive areas such as the Arctic Ocean. While ecophysiological mechanisms may accurately predict short-term responses of ecosystem function to seasonal variation in temperature, we expect eco-evolutionary responses to be important on multi-annual timescales, over which bacterial populations may evolve and adapt to long-term trends in temperature. Our model provides a critical step towards the integration of microbial eco-evolutionary processes in ocean ecosystem models, necessary for improving our projections of ocean nutrient cycle in a warming world.
References
Falkowski PG, Fenchel T, Delong EF. The microbial engines that drive Earth’s biogeochemical cycles. Science. 2008;320:1034–9.
Riebesell U, Körtzinger A, Oschlies A. Sensitivities of marine carbon fluxes to ocean change. Proc Natl Acad Sci USA. 2009;106:20602–9.
Hutchins DA, Fu F. Microorganisms and ocean global change. Nat Microbiol. 2017;2:1–11.
Cavicchioli R, Ripple WJ, Timmis KN, Azam F, Bakken LR, Baylis M, et al. Scientists’ warning to humanity: microorganisms and climate change. Nat Rev Microbiol. 2019;17:569–86.
Bopp L, Resplandy L, Orr JC, Doney SC, Dunne JP, Gehlen M, et al. Multiple stressors of ocean ecosystems in the 21st century: projections with CMIP5 models. Biogeosciences. 2013;10:6225–45.
Oschlies A, Brandt P, Stramma L, Schmidtko S. Drivers and mechanisms of ocean deoxygenation. Nat Geosci. 2018;11:467–73.
Cazenave A, Llovel W. Contemporary sea level rise. Ann Rev Mar Sci. 2010;2:145–73.
Frölicher TL, Ramseyer L, Raible CC, Rodgers KB, Dunne J. Potential predictability of marine ecosystem drivers. Biogeosciences. 2020;17:2061–83.
Taucher J, Oschlies A. Can we predict the direction of marine primary production change under global warming? Geophys Res Lett. 2011;38:L02603.
Laufkötter C, Vogt M, Gruber N, Aita-Noguchi M, Aumont O, Bopp L, et al. Drivers and uncertainties of future global marine primary production in marine ecosystem models. Biogeosciences. 2015;12:6955–84.
Azam F, Fenchel T, Field JG, Gray J, Meyer-Reil L, Thingstad F. The ecological role of water-column microbes in the sea. Mar Ecol Prog Ser. 1983:257–63.
Fenchel T. The microbial loop–25 years later. J Exp Mar Biol Ecol. 2008;366:99–103.
Kirchman DL, Morán XAG, Ducklow H. Microbial growth in the polar oceans—role of temperature and potential impact of climate change. Nat Rev Microbiol. 2009;7:451–9.
Aumont O, Éthé C, Tagliabue A, Bopp L, Gehlen M. PISCES-v2: An ocean biogeochemical model for carbon and ecosystem studies. Geosci Model Dev Discuss. 2015;8:2465–513.
Vichi M, Masina S. Skill assessment of the PELAGOS global ocean biogeochemistry model over the period 1980–2000. Biogeosciences. 2009;6:2333–53.
Hasumi H, Nagata T. Modeling the global cycle of marine dissolved organic matter and its influence on marine productivity. Ecol Model. 2014;288:9–24.
Laufkötter C, Vogt M, Gruber N, Aumont O, Bopp L, Doney SC, et al. Projected decreases in future marine export production: the role of the carbon flux through the upper ocean ecosystem. Biogeosciences. 2016;13:4023–47.
Monroe JG, Markman DW, Beck WS, Felton AJ, Vahsen ML, Pressler Y. Ecoevolutionary dynamics of carbon cycling in the anthropocene. Trends Ecol Evol. 2018;33:213–25.
Bennett AF, Dao KM, Lenski RE. Rapid evolution in response to high-temperature selection. Nature. 1990;346:79–81.
Garud NR, Good BH, Hallatschek O, Pollard KS. Evolutionary dynamics of bacteria in the gut microbiome within and across hosts. PLoS Biol. 2019;17:e3000102.
Zhao S, Lieberman TD, Poyet M, Kauffman KM, Gibbons SM, Groussin M, et al. Adaptive evolution within gut microbiomes of healthy people. Cell Host Microbe. 2019;25:656–67.
Pomeroy LR, Williams PJleB, Azam F, Hobbie JE. The microbial loop. J Oceanogr. 2007;20:28–33.
Walworth NG, Zakem EJ, Dunne JP, Collins S, Levine NM. Microbial evolutionary strategies in a dynamic ocean. Proc Natl Acad Sci USA. 2020;117:5943–8.
Malik AA, Martiny JB, Brodie EL, Martiny AC, Treseder KK, Allison SD. Defining trait-based microbial strategies with consequences for soil carbon cycling under climate change. ISME J. 2020;14:1–9.
Saifuddin M, Bhatnagar JM, Segrè D, Finzi AC. Microbial carbon use efficiency predicted from genome-scale metabolic models. Nat Commun. 2019;10:1–10.
Muscarella ME, Howey XM, Lennon JT. Trait‐based approach to bacterial growth efficiency. Environ Microbiol. 2020;22:3494–3504.
Roller BR, Stoddard SF, Schmidt TM. Exploiting rRNA operon copy number to investigate bacterial reproductive strategies. Nat Microbiol. 2016;1:1–7.
Sarmiento JL, Gruber N. Ocean biogeochemical dynamics. Princeton University Press, 2006.
Bendtsen J, Lundsgaard C, Middelboe M, Archer D. Influence of bacterial uptake on deep-ocean dissolved organic carbon. Glob Biogeocehm Cycles. 2002;16:74–1.
Chen B, Landry MR, Huang B, Liu H. Does warming enhance the effect of microzooplankton grazing on marine phytoplankton in the ocean? Limnol Oceanogr. 2012;57:519–26.
Krause S, Le Roux X, Niklaus PA, Van Bodegom PM, Lennon JT, Bertilsson S, et al. Trait-based approaches for understanding microbial biodiversity and ecosystem functioning. Front Microbiol. 2014;5:251.
Kiørboe T, Visser A, Andersen KH. A trait-based approach to ocean ecology. ICES Int J Mar Sci. 2018;75:1849–63.
Grime JP. Evidence for the existence of three primary strategies in plants and its relevance to ecological and evolutionary theory. Am Nat. 1977;111:1169–94.
Polz MF, Cordero OX. Bacterial evolution: genomics of metabolic trade-offs. Nat Microbiol. 2016;1:1–2.
Carlson CA, Del Giorgio PA, Herndl GJ. Microbes and the dissipation of energy and respiration: from cells to ecosystems. J Oceanogr. 2007;20:89–100.
Arnosti C. Patterns of microbially driven carbon cycling in the ocean: links between extracellular enzymes and microbial communities. Adv Oceanogr. 2014;2014:706082.
Pfeiffer T, Schuster S, Bonhoeffer S. Cooperation and competition in the evolution of ATP-producing pathways. Science. 2001;292:504–7.
Button D. Biochemical basis for whole-cell uptake kinetics: specific affinity, oligotrophic capacity, and the meaning of the Michaelis constant. Appl Environ Microbiol. 1991;57:2033–8.
Metz JA, Nisbet RM, Geritz SA. How should we define ‘fitness’ for general ecological scenarios? Trends Ecol Evol. 1992;7:198–202.
Geritz SA, Metz JA, Kisdi E, Meszéna G. Dynamics of adaptation and evolutionary´ branching. Phys Rev Lett. 1997;78:2024.
Abs E, Ferrière R. Modeling microbial dynamics and heterotrophic soil respiration: effect of climate change. Biogeochemical cycles: ecological drivers and environmental impact. 2020:103–29.
Lipson DA. The complex relationship between microbial growth rate and yield and its implications for ecosystem processes. Front Microbiol. 2015;6:615.
Hansell DA, Carlson CA. Biogeochemistry of marine dissolved organic matter. Academic Press, 2014.
Urban MC, De Meester L, Vellend M, Stoks R, Vanoverbeke J. A crucial step toward realism: responses to climate change from an evolving metacommunity perspective. Evol Appl. 2012;5:154–67.
Norberg J, Urban MC, Vellend M, Klausmeier CA, Loeuille N. Eco-evolutionary responses of biodiversity to climate change. Nat Clim Change. 2012;2:747–51.
Sarmento H, Montoya JM, Vázquez-Domínguez E, Vaqué D, Gasol JM. Warming effects on marine microbial food web processes: how far can we go when it comes to predictions? Philos Trans R Soc Long B Biol Sci. 2010;365:2137–49.
Walther S, Voigt M, Thum T, Gonsamo A, Zhang Y, Köhler P, et al. Satellite chlorophyll fluorescence measurements reveal large-scale decoupling of photosynthesis and greenness dynamics in boreal evergreen forests. Glob Change Biol. 2016;22:2979–96.
Williams RG, Follows MJ. Ocean dynamics and the carbon cycle: Principles and mechanisms. Cambridge University Press, 2011.
Lewis K, Van Dijken G, Arrigo KR. Changes in phytoplankton concentration now drive increased Arctic Ocean primary production. Science. 2020;369:198–202.
Ward B, Collins S, Dutkiewicz S, Gibbs S, Bown P, Ridgwell A, et al. Considering the role of adaptive evolution in models of the ocean and climate system. J Adv Model Earth Syst. 2019;11:3343–61.
Vázquez-Domínguez E, Vaque D, Gasol JM. Ocean warming enhances respiration and carbon demand of coastal microbial plankton. Glob Change Biol. 2007;13:1327–34.
López-Urrutia A, Morán XAG. Resource limitation of bacterial production distorts´ the temperature dependence of oceanic carbon cycling. Ecology. 2007;88:817–22.
Parker GA, Smith JM. Optimality theory in evolutionary biology. Nature. 1990;348:27–33.
Hammerstein P. Darwinian adaptation, population genetics and the streetcar theory of evolution. J Math Biol. 1996;34:511–32.
Eshel I, Feldman MW, Bergman A. Long-term evolution, short-term evolution, and population genetic theory. J Theor Biol. 1998;191:391–6.
Hagerty SB, Allison SD, Schimel JP. Evaluating soil microbial carbon use efficiency explicitly as a function of cellular processes: implications for measurements and models. Biogeochemistry. 2018;140:269–83.
Segre D, Vitkup D, Church GM. Analysis of optimality in natural and perturbed metabolic networks. Proc Natl Acad Sci USA. 2002;99:15112–7.
Marx CJ. Can you sequence ecology? Metagenomics of adaptive diversification. PLoS Biol. 2013;11:e1001487.
O’Brien S, Hodgson DJ, Buckling A. The interplay between microevolution and community structure in microbial populations. Curr Opin Biotechnol. 2013;24:821–5.
Scheuerl T, Hopkins M, Nowell RW, Rivett DW, Barraclough TG, Bell T. Bacterial adaptation is constrained in complex communities. Nat Commun. 2020;11:1–8.
Schloissnig S, Arumugam M, Sunagawa S, Mitreva M, Tap J, Zhu A, et al. Genomic variation landscape of the human gut microbiome. Nature. 2013;493:45–50.
Boyd JA, Woodcroft BJ, Tyson GW. GraftM: a tool for scalable, phylogenetically informed classification of genes within metagenomes. Nucleic Acids Res. 2018;46:e59–9.
Gregory AC, Gerhardt K, Zhong ZP, Bolduc B, Temperton B, Konstantinidis KT, et al. MetaPop: a pipeline for macro-and micro-diversity analyses and visualization of microbial and viral metagenome-derived populations. bioRxiv 2020. https://doi.org/10.1101/2020.11.01.363960.
Coles VJ, Stukel MR, Brooks MT, Burd A, Crump BC, Moran MA, et al. Ocean biogeochemistry modeled with emergent trait-based genomics. Science. 2017;358:1149–1154.
Scheinin M, Riebesell U, Rynearson TA, Lohbeck KT, Collins S. Experimental evolution gone wild. J R Soc Interface. 2015;12:20150056.
Thomas MK, Kremer CT, Klausmeier CA, Litchman E. A global pattern of thermal adaptation in marine phytoplankton. Science. 2012;338:1085–8.
Grimaud GM, Le Guennec V, Ayata SD, Mairet F, Sciandra A, Bernard O. Modelling the effect of temperature on phytoplankton growth across the global ocean. IFACPapersOnLine. 2015;48:228–33.
Sauterey B, Ward B, Rault J, Bowler C, Claessen D. The implications of ecoevolutionary processes for the emergence of marine plankton community biogeography. Am Nat. 2017;190:116–30.
Beckmann A, Schaum CE, Hense I. Phytoplankton adaptation in ecosystem models. J Theor Biol. 2019;468:60–71.
Wilhelm SW, Suttle CA. Viruses and nutrient cycles in the sea: viruses play critical roles in the structure and function of aquatic food webs. Bioscience. 1999;49:781–8.
Danovaro R, Corinaldesi C, Dell’Anno A, Fuhrman JA, Middelburg JJ, Noble RT, et al. Marine viruses and global climate change. FEMS Microbiol Rev. 2011;35:993–1034.
Breitbart M, Bonnain C, Malki K, Sawaya NA. Phage puppet masters of the marine microbial realm. Nat Microbiol. 2018;3:754–66.
Weitz JS, Stock CA, Wilhelm SW, Bourouiba L, Coleman ML, Buchan A, et al. A multitrophic model to quantify the effects of marine viruses on microbial food webs and ecosystem processes. ISME J. 2015;9:1352–64.
Gregory AC, Zayed AA, Conceição-Neto N, Temperton B, Bolduc B, Alberti A, et al. Marine DNA viral macro-and microdiversity from pole to pole. Cell. 2019;177:1109–23.
Acknowledgements
We thank Olivier Aumont, Laurent Bopp, and Boris Sauterey for discussion and comments.
Funding
This work is supported by France Investissements d’Avenir program (ANR-10-LABX-54 MemoLife, ANR-10-IDEX-0001-02 PSL) and PSL - University of Arizona Mobility Program. PC is supported by a doctoral fellowship from the French IPEF program. RF acknowledges support from the U.S. National Science Foundation, Dimensions of Biodiversity (DEB-1831493), Biology Integration Institute-Implementation (DBI-2022070), and National Research Traineeship (DGE-2022055) programs; and from the United States National Aeronautics and Space Administration, Interdisciplinary Consortium for Astrobiology Research program.
Author information
Authors and Affiliations
Contributions
RF conceived the study. All authors developed the model. PC performed the analysis. All authors wrote the first version of the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Cherabier, P., Ferrière, R. Eco-evolutionary responses of the microbial loop to surface ocean warming and consequences for primary production. ISME J 16, 1130–1139 (2022). https://doi.org/10.1038/s41396-021-01166-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-021-01166-8