Abstract
The recent application of macroecological tools and concepts has made it possible to identify consistent patterns in the distribution of microbial biodiversity, which greatly improved our understanding of the microbial world at large scales. However, the distribution of microbial functions remains largely uncharted from the macroecological point of view. Here, we used macroecological models to examine how the genes encoding the functional capabilities of microorganisms are distributed within and across soil systems. Models built using functional gene array data from 818 soil microbial communities showed that the occupancy-frequency distributions of genes were bimodal in every studied site, and that their rank-abundance distributions were best described by a lognormal model. In addition, the relationships between gene occupancy and abundance were positive in all sites. This allowed us to identify genes with high abundance and ubiquitous distribution (core) and genes with low abundance and limited spatial distribution (satellites), and to show that they encode different sets of microbial traits. Common genes encode microbial traits related to the main biogeochemical cycles (C, N, P and S) while rare genes encode traits related to adaptation to environmental stresses, such as nutrient limitation, resistance to heavy metals and degradation of xenobiotics. Overall, this study characterized for the first time the distribution of microbial functional genes within soil systems, and highlight the interest of macroecological models for understanding the functional organization of microbial systems across spatial scales.
Similar content being viewed by others
Introduction
The functional potential of microbes relies on the collection of metabolic capabilities encoded by the genes contained in their genomes, and that, once expressed, define the traits of the microorganism carrying these genes. While several functional genes are specific to certain taxa [1], many genes are common to most microorganisms and compose the “core genome” [2, 3]. This results in high levels of functional redundancy among microbial taxa [4,5,6]. In addition, the wide occurrence of mobile accessory genes exchanged through horizontal transfer [7, 8] plays significant contribution in these systems and adds to their complexity. Further, some genes exist with different sequences despite encoding similar products, which correspond to the functional redundancy among variants of the same gene. For all these reasons, the insights provided by taxonomic approaches into the role of microbial communities in ecosystem functioning are limited. As an alternative, the use of functional approaches has been widely advocated, notably through the direct study of the gene content of microbial communities and regarding genes as potential microbial functional traits [6, 9, 10].
In the recent years, researchers have used macroecological models to disentangle the complexity of microbiomes [11, 12]. Such approaches have been notably used to explore commonness and rarity patterns in microbial systems and successfully uncovered regularities in the distribution of microorganisms across various spatial scales (species, communities, or ecosystems) and revealed similarities or idiosyncrasies in the processes underlying these distributions [12,13,14]. Such macroecology-based frameworks do not rely on arbitrarily defined thresholds that oppose rare vs. abundant biological units. Instead, full distributions are used to classify units along gradients ranging from rarity to commonness. To investigate patterns in local abundance, rank-abundance distributions (RAD) place biological units from a given area or community along a gradient from low to high abundance. RAD have been used to describe the distribution of taxa within microbial taxa, revealing the presence of a long tail of rare organisms composing the so-called “rare biosphere” [15,16,17]. In spatial occupancy studies, occupancy-frequency distribution (OFD) describes how biological units are spatially distributed across a set of communities [18], and classifies these units along a distribution gradient, from spatially restricted to ubiquitous. OFD models describing both macro and microorganisms were found to be either unimodal or bimodal, and to exhibit a higher left mode [19], that is, high proportion of taxa represents small fractions of communities. Finally, the relationship between local abundance and spatial occupancy (occupancy-abundance relationships—OAR) is one of the most reported trends in macroecology and has been shown to be positive for a wide range of macro- [20,21,22,23,24,25] and microorganisms [12, 26,27,28,29,30,31,32,33,34]. The positive OAR for biological units in natural systems predicts that some units have a restricted spatial distribution with low abundance (i.e., “satellites”) while others are ubiquitous and found in high abundance (i.e., “core”) [35,36,37,38]. In microbial ecology, observations of these distribution patterns contributed to unveil community dynamics, which later led to the concept of “conditionally rare taxa”, i.e., satellite organisms having the potential to bloom and temporarily influence community dynamics [39,40,41]. Such life strategy is known to be related to the metabolic capabilities of certain microbes [42, 43]. Therefore, if we aim to understand the mechanisms underpinning the macroecological patterns of microbial communities, applying these concepts in the context of functional genes could provide valuable information.
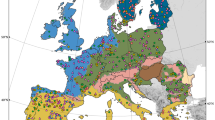
In this study, we aimed to address a simple, and yet unanswered, question: how are the functions carried by microorganisms distributed within soil systems? To tackle this question, we aimed to assess the distribution of microbial functional genes at different scales in soils using macroecological models, to identify abundant and rare functions across these systems. For that, we constructed a database with functional gene array (FGA) [44] data from 818 topsoil microbial communities sampled from ten sites located around the globe and representing various ecosystem types (i.e., tundra, grassland, forest, shrubland and pasture; Fig. 1). The FGA was used to hybridize microbial community DNA to a set of 39,681 probes that correspond to variants of 194 functional genes encoding various microbial functions involved in biogeochemical cycles, pollutant breakdown, virulence and resistance to various types of physical and chemical stress. By considering genes instead of taxa as the unit of our study, we can make the following predictions about their distribution patterns: the presence of a set of core genes shared among most microorganisms should lead to (i) OFD displaying either unimodal with a right mode or bimodal with a stronger right mode; and (ii) RAD characterized by few dominant genes and a long tail of rare genes. Consequently, as observed for most of the biological units from the smallest to the largest, the two previous patterns should result in positive OAR. By using this approach, we classified genes along a continuum from low abundance and limited occurrence to high abundance and ubiquity, with the two ends of this gradient representing satellite and core genes, respectively. Then, by investigating the functions carried by these genes we show that rare and common microbial genes encode different functions in soil ecosystems.
Material and methods
Composition of the database
The database consisted of ten datasets collected in the frame of previous projects and comprised a total of 818 surface topsoil samples from three continents, representing a wide gradient of environmental conditions (Fig. S1 and Table S1). The spatial scale covered by each dataset range from hundreds of meters in some experimental sites to dozens of kilometers across natural landscapes. We did not investigate within site differences among samples as our goal was to look for repeatable patterns across geographically distant sites, considered as separate entities from the physico-chemical, climatic and pedoclimatic standpoint. Here, we opted for a large spatial scale macroecological approach that did not consider local contingencies and focused on comparing the distribution of functional gene variants within and across communities from isolated ecosystems [45].
Five sites were located in the United States, including three grassland [46,47,48] and two Alaskan tundra ecosystems [49, 50]. Climate change experiments were conducted in four of these sites (i.e., variation in temperature, CO2 concentration, etc.). Four sites were located in China, comprising two grassland ecosystems from the Qinghai province [51, 52], in addition to forests and shrubland sites from the Hubei Province [53, 54]. The last site corresponded to pasture areas located in the Brazilian Amazon basin [55, 56].
All the samples were analyzed using a FGA [44] composed of 39,681 DNA probes targeting protein-coding genes. Probe design was done as described elsewhere [44, 57, 58], by searching keywords against the NCBI nr database. Candidate sequences were validated with HMM models and 50-mer oligonucleotide probes were designed using CommOligo 2.0 [59]. These probes (herafter termed genes variants or variants) served as the unit of our study to characterize the macroecological distribution of microbial functions. The potential role of these gene variants in the microbial communities were defined according to a functional classification performed using information available in databases such as NCBI, UniProt, or EXpasy and were also based on extensive literature reviews [10]. The 39,681 variants correspond to 194 genes (e.g., nirB, ureC, exochitinase, arsB), defined as collections of variants encoding a similar product but with slightly different DNA sequence and originating from different organisms [60]. Genes were further classified into 56 gene families, defined as collections of genes that, together, represent a coherent set of microbial functions (e.g., resistance to oxygen or heat stress, C fixation, denitrification). Finally, these families were grouped into nine broad categories of microbial functions (e.g., C, N, P or S cycling, antibiotic resistance and virulence; cf. Table S2 for a full description of the distribution of variants in this different levels of functional resolution). This classification allowed the linkage of genes with the function they carry.
Functional gene array analyses
FGA hybridizations were performed according to standardized laboratory procedure from the Institute for Environmental Genomics (IEG, OK, USA), as described in [44]. Total community DNA was quantified using picogreen and, for each sample, 800 ng were labeled with Cy-5 (GE Healthcare), dried in a Speedvac at 45 °C for 45 min and stored at −20 °C before hybridization. The pellet was re-hydrated in 2.68 µl of tracking control completed with 7.12 µl of hybridization solution (Formamide, SSC, SDS, oligo Cy-3, oligo Cy-5 and universal standard). Labeled DNA was incubated at 95 °C for 5 min before loading onto the array. The hybridization was done at 42 °C in the presence of 40% formamide for 16 h. After washing and drying, arrays were scanned and gridded before signal intensity quantification using ImaGene 6.0 (Biodiscovery Inc., El Segundo, CA, USA). Original raw hybridization signal intensity data were retrieved from the IEG microarray data repository (http://ieg2.ou.edu/NimbleGen/analysis.cgi). To estimate the abundance of functional genes, noise data were removed using a hybridization signal cutoff of 2000 intensity [57, 61].
Distribution patterns of functional gene variants
The abundance of the 39,681 genes variants within a given sample was estimated as the logged hybridization signal intensity on the FGA. The shape of variants RAD within each of the 818 samples was assessed using four widely used rank-abundance models (Logseries, Poisson lognormal, Negative binomial and Zipf). Models were fitted using maximum likelihood estimation of parameters and their goodness of fit compared using the AIC (Akaike Information Criterion). For each sample, the model with the lowest AIC value was considered to be the best fitting model. Models were fitted and compared using the python package macroecotools [62].
Gene variants occupancy was estimated within each site by counting the number of samples in which the variant was detected and dividing it by the total number of samples in the site (ranged between 0 and 1). The shape of OFD of variants across multiple soil communities was analyzed using the Mitchell-Olds & Shaw test as implemented in the MOStest function of the R package vegan [63]. This approach fits a quadratic generalized linear models of the type µ = b0 + b1x + b2x2 to the OFD, where b0 corresponds to the intercept, b1 to rate of change and b2 determines whether the model is convex or concave and was used to estimate the model shape: if b2 < 0 the model is unimodal and if b2 > the model is bimodal.
To test OAR within each site, variants abundance was estimated as the average abundance across samples from that site, occupancy in each site was estimated as described above and the relationships were analyzed using linear models (lm() function in R) relating variants occupancy with their average abundance across samples.
Translation of gene variants distribution into microbial functions distribution
To determine how the distribution of genes variants translates into the distribution of microbial functions in soil systems, we associated the variants to the functions they encode (Fig. 1) using the classification provided by the FGA (genes, gene families and broad categories, cf. Table S2). First, variant distributions were used to rank the 39,681 variants and to group them into bins that represented gradients of abundance within a community (RAD), occupancy across communities (OFD) or commonness within sites (OAR). Bins were defined by splitting variants into six sets of equal number based on their rank (see supplementary information for an explanation of the choice of bin number). Then, for each bin (ranked 1 for the lowest end of the gradient to 6 for the highest end), the importance of a function at a given level of functional classification was estimated as the proportion of the summed hybridization signal of all the variants from that bin. However, different functions (e.g., gene families) differed in numbers of variants on the FGA design (Table S2), and this must be accounted for when estimating function importance. If the importance of a function in a bin is simply estimated by counting the number of variants from this function, or their summed signal intensity, then functions represented by many variants are more likely to be considered important than those represented by few variants. To avoid this bias, and thus take into account unequal sampling effort across function on the FGA, function importance within a bin was estimated by dividing the observed proportion of the total signal intensity in the bin represented by variants from this function by the proportion of the total number of variants represented by this function on the FGA design (Fig. 1). The obtained ratio, here termed weight of the function, describes how much the proportion of the signal represented by a function in the bin departs from a null expectation, which corresponded to the proportion of the signal represented by this function if variants were randomly sampled on the FGA. Functions with weight values > 1 were considered over-represented in a given bin, i.e., more abundant than expected by chance, while functions with a weight < 1 were considered under-represented, i.e., less abundant than expected by chance. This provided matrices describing the composition of each bin (column) in terms of microbial functions (rows), with each function being associated with a weight. As bins (B1–B6) represented gradients of increasing abundance within communities, increasing occupancy across communities and increasing commonness within sites, we were able to identify the functions encoded by variants along these gradients.
Analysis of the distribution of microbial functions in soil systems
To characterize the distribution of microbial functions in soil systems, we analyzed the weighted matrices described above (i.e., functions × bins). The dissimilarity between bins was estimated with the Bray–Curtis index and visualized using Detrended Correspondence Analysis (DCA). We tested for differences in the composition of bins using permutational multivariate analysis of variance (PERMANOVA) [64], implemented as the adonis function in the R package vegan [63]. We tested the differences between bins of different ranks (1–6) and originating from different sites. This was done after associating genes to function weight at the three levels of functional classification (genes, gene families, and broad categories).
For each of the 194 genes, we fitted linear models describing the relationship between the weight of genes in each bin and the rank of occupancy-abundance bins (1-6). By looking at the slopes of these models, we identified the genes, and the corresponding gene families, that were under- or over-represented along the occupancy-abundance gradient. Negative relationships (significant negative slopes) corresponded to genes over-represented in rare variants, whereas positive ones (significant positive slopes) corresponded to genes over-represented in abundant variants. When the slope of the linear model was not significant the case was classified as “no relation”. Finally, we characterized the composition in terms of function weight of in rare (B1) and common (B6) occupancy-abundance bins.
Results
Macro-ecological distribution patterns of functional gene variants in soil ecosystems
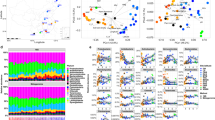
The RAD of gene variants within communities (i.e., samples) was described using four different RAD models (Logseries, Poisson lognormal, Negative binomial and Zipf). Poisson lognormal was found to be the best model to describe variants RAD in 100% of the samples (n = 818, Fig. 2). The OFDs of variants across communities were significantly bimodal in all ten sites, with a maximum at low and high occupancy (Fig. 3, MOS test, p value < 0.001). In all but one site (CiPEHR), we observed increased variant frequencies at high occupancy, in comparison to low occupancy (i.e., the right mode of the OFD was stronger). This was supported by the observation of higher F values when testing the presence of a frequency maximum at high occupancy (F = 103 ± 74) compared with low occupancy (F = 59 ± 57, Table S3). F values at high occupancy were 1.7 ± 0.7 times higher than at low occupancy. F values of the left mode was less pronounced for the two datasets with the lowest number of samples (KAEFS, n = 12 and Fazenda nova vida, n = 24), suggesting that the sampling effort was not high enough to capture variants with spatially restricted distribution. The relationships between average variant abundance and occupancy (OAR) were linear, positive and highly significant in all ten sites (Fig. 4). This linear trend represents a gradient ranging from rarity, i.e., low abundance and restricted spatial distribution (bottom left), to commonness, i.e., ubiquitous distribution across communities and high abundance (top right). At the two ends of this gradients lie satellite (B1) and core gene variants (B1), respectively.
For each community (i.e., sample), we fitted four rank-abundance models (Logseries, Poisson lognormal, Negative binomial and Zipf) using maximum likelihood estimation (MLE). Each subplot corresponds to one site and each gray line represents the RAD of variants within a sample (logged hybridization signal intensity). The thick lines correspond to the average model for each site.
Colored lines correspond to the best model describing the relationship. The color of data points corresponds to the colors used in Fig. 1. The p values of the MOS test of bimodality along with the F values associated with the test of the presence of local maxima at low and high occupancy are depicted.
In each site, the occupancy of the 39,681 gene variants present on the FGA was estimated as the proportion of samples in which it was detected. Their abundance was estimated as the average abundance across all the samples from the site. Black lines represent the best linear models describing the occupancy-abundance relationship.
Distribution of microbial functions in soil ecosystems
We found that abundance bins from different samples but with similar rank had a more similar distribution of function weights than bins from the same sample but with different ranks. This was validated for each site, as bins with similar rank clustered together on the DCA, based on the Bray–Curtis dissimilarity estimated on function weights (Fig. S2). In addition, bins located at the two ends of the abundance gradient within samples were the most dissimilar (B1 and B6). This result was also validated when comparing the weight of functions in abundance bins across sites (n = 818 samples, times 6 abundance bins). The weight of genes in abundance bins was better predicted by the rank of the bin along abundance gradient (B1–B6, PERMANOVA, p < 0.01) than by its site of origin (Table 1). Bin rank explained between 65 and 84% of the variation in gene weight while the site explained between 8 and 16%. This trend was confirmed when higher levels of variants classification were used (e.g., gene families and broad categories), as suggested by higher F values of bin rank compared with the site effect (i.e., 18.6, 8.3 and 8.1 times higher for broad categories, gene families and genes, respectively).
We performed a similar analysis using occupancy bins (B1–B6) and found that variants with similar occupancy within sites exhibited similar function weights. Bin rank explained from 31 to 57% of the variation in gene weight distribution between bins from the ten sites, while the factor site explained only 8 to 15% (Table 1). According to the F values, the effect of occupancy rank was 5.4–13.3 times higher than the site effect. As observed for abundance bins, the greatest differences in distribution of function weight among occupancy bins were observed between the two extremes of the gradient, B1 and B6 (Table S4).
Functions of satellites and core genes in soil ecosystems
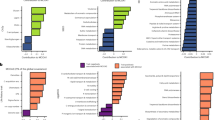
We observed clear trends in the distribution of the 194 gene families along the occupancy-abundance gradient in soil systems, and we identified the gene families, and the corresponding broad ecological categories that were systematically over-represented at one end of this gradient (Figs. 5 and S3-S4-S5, Table S5). Among the 194 linear models fitted between genes weight and bin rank, only 22 (11%) were not significant (p value > 0.05, Fig. 5-A and Fig. S3). This corresponded to genes that were not associated with rare or abundant variants. We observed 91 (47%) negative relationships (p value < 0.05 and slope < 0, Fig. 5-A and Fig. S4), corresponding to genes that were over-represented in rare (i.e., satellites) variants and under-represented in abundant (i.e., core) variants. Among these, 38.5% of the genes were related to stress responses (e.g., osmotic, oxygen or radiation stress, cold or heat shocks, sigma factors, N or P limitations), 18.7% to metal resistance, 14.3% to C cycling, 11% to virulence, and the remaining 6% comprised three categories (antibiotic resistance, N and S cycling). The 20 genes with the strongest negative slope were related to various forms of stress responses, virulence proteins (toxin, adhesin, aerobactin), metal resistance (cadmium, cobalt, aluminum), broad biological functions (blue copper protein, thioredoxin), C cycling (acetogenesis) and energy processes (hydrogenase). Significant positive relationships were found for 81 (42%) genes (p value < 0.05 and slope > 0, Fig. 5-A and Fig. S5). These genes were under-represented in rare and over-represented in abundant variants. From those, 39.5% were related to C cycling, 18.5% to metal resistance, 12.3% to N cycling, 9.9% to stress responses, 4.9% to antibiotic resistance and S cycling, 3.7% to energy processes and P cycling and 2.5% to virulence. The 20 genes with the strongest positive slope were related to the degradation of C-based substrates, the N cycle (denitrification, assimilatory-N-reduction and ammonification), metal resistance (lead, silver and mercury), C fixation (pcc and CODH genes), S oxidation (sox gene) and energy processes (hydrogenase).
For each of the 194 genes, we fitted linear models describing the relationship between the weight of genes in each occupancy-abundance bin and the rank of the bin (1–6). Negative relationships (significant negative slopes) corresponded to genes over-represented in rare gene variants whereas positive ones (significant positive slopes) corresponded to genes over-represented in abundant variants, when the slope of the linear model was not significant the case was classified as “no relation”. A Relative proportions of genes across broad categories for the models with non-significant (n = 22), negative (n = 91) and positive (n = 81) relationships. B Slopes of the models classified by broad categories. C–D In each of the ten studied sites, satellite and core variants were defined as those from the extreme occupancy-abundance bins (B1 and B6, respectively). Boxplots represent the weight of ecological processes in bins from each of the ten sites for satellite (C) and core (D) variants. Gene families were ranked according to their average weight in core variants across the ten sites. Colors represent different broad categories of functions as depicted in the legend in (D).
We also looked at the weight of each gene family across the ten sites in the first (B1, satellite variants) and in the sixth (B6, core variants) occupancy-abundance bins (Fig. 5C, D, Fig. S6). Core variants were enriched in functions related to the C, N, P and S biogeochemical cycles, but were depleted in functions related to stress response, virulence, heavy metal and antibiotics resistance. Satellite variants were more evenly distributed across the categories, despite notable depletion in functions related to the N cycle and enrichment in stress response and virulence related functions. Twelve processes were clearly enriched in core variants: S oxidation, denitrification, C fixation, ammonification, assimilatory-N-reduction, C degradation, P utilization and, surprisingly, resistance to mercury, lead and silver contamination. On the contrary, the processes enriched in satellites variants included stress response (e.g., oxygen limitation, heat shocks, radiation, osmotic and protein stresses, P and N limitation), antibiotics resistance (e.g., membrane transporters), resistance to heavy metal (e.g., Cr, Cu, As, Te and Al) and virulence (e.g., hemolysin, capsule formation, pilin, aerobactin and pilin). Interestingly, two C-related processes (methane metabolism and acetogenesis) were also enriched in satellites variants.
Discussion
Macro-ecological distribution patterns of microbial gene variants in soil systems
In this study, we applied an analytical framework derived from macroecological concepts to describe the distribution of microbial gene variants at two scales, i.e., within and across communities, in ten different soil ecosystems. We showed that RAD of gene variants within soil communities can be adequately described using classic macroecological models that were designed to capture the intrinsically uneven distribution of species within natural assemblages. Here the Poisson lognormal model was the best one to describe variants RAD. While many RAD models have been developed over time to describe these data, the Poisson lognormal model is often considered as the most widely applicable due to its “positive range, right skewness, heavy right tail, and easily computed parameter estimates” [65]. In microbes, it was identified as the best model to characterize bacterial RAD at the global scale [14], in the marine environment [66] and in wastewater treatment plants [67], and it was used to predict the total number of microbial OTUs at a global scale [68, 69]. Overall, our results highlight that the wide applicability of the lognormal model to describe RAD of biological units can be extended to microbial gene variants. Furthermore, we found that RAD of microbial taxa and gene variants are very similar, which demonstrates the usefulness of macroecological tools beyond the dichotomy micro- vs. macro-organisms [12], and toward a wider range of biological units (e.g., genes, interactions, viruses).
When looking at the spatial distribution of gene variants across samples, the observed OFDs differed from what is generally reported for taxa in communities of both macro and microorganisms (i.e., the “hollow” distribution), with a higher number of taxa being found in a few sites and only a small number of ubiquitous taxa [70]. In their review, McGeoch & Gaston (2002), analyzed OFD models describing the distribution of macro-organisms (e.g., plants, insects, birds, fishes) from small (<1 km²) to continental scales. Among the 68 reported models, some were unimodal (57%), other bimodal (31%), but the large majority exhibited a higher left mode (68%), that is a higher proportion of taxa observed in a small proportion of communities than widely distributed. Similar right-skewed OFD have been reported for microbial taxa, from the microscale [33] to hundreds of km in both marine [31, 32, 71, 72] and soil environments [27, 73]. Here, we found that the OFD of microbial functional gene variants contrast with these general trends reported for taxa, as they exhibit a stronger right mode with a much higher proportion of variants that were ubiquitously distributed within a site than unique to a single community. This pattern was expected and due to the functional redundancy among microbes [4,5,6, 74], i.e., the fact that most microbes share a common set of functional genes that can detected in any soil sample collected within a given site. Despite this stronger right mode, the observed OFD were bimodals in the ten studied ecosystems, a pattern known as the Raunkiaer’s law of distribution of frequencies. It has been suggested that this pattern can emerge from random sampling of biological units from a lognormal rank-abundance distribution [75], which seems to be the case in our study.
The combination of within-community abundance distribution (RAD) and across-communities occupancy distribution (OFD) corresponds to OAR, and these OAR have been a reported to be positive for a wide range of macro- and microbial taxa [12, 20,21,22,23,24,25,26, 28,29,30,31,32,33,34]. Several theories have been proposed to explain the existence of positive OAR in taxa, including stochastic processes resulting from neutral dynamics [19] or differences between species in terms of ecological niche [76]. However, there is currently no consensus on the underlying mechanisms of OAR for taxa and no studies for functional genes. Positive OAR can be seen as a gradient of commonness (or rarity) across a set of communities, with biological units that are both spatially restricted and locally scarce at one end (i.e., the satellites), and the biological units that are widespread and very abundant at the other end (i.e., the core). Here, we observed that some gene variants were present in a small number of communities within each site and exhibited low abundance in these communities, and that other gene variants exhibited high abundance in all the communities from all the sites. This resulted in positive OAR of microbial functional gene variants within each studied site, and allowed the identification of rare and common microbial gene variants in soil systems, along with the function they encode. However, despite the fact that OAR of microbial taxa and gene variants are both positive, they differ greatly regarding the distribution of biological units along the rarity to commonness gradient. As mentioned, while the RAD are quite similar the OFD are very different. These differences resulted in taxonomic OAR with many satellite and few core taxa while functional OAR had only few satellite and many core gene variants (Fig. 4). Interestingly, ecological theories suggest that core taxa are more likely to be generalists while satellite ones are more likely to be specialists [77, 78], which leads to the question whether core and satellite gene variants encode general and specialized functions, respectively.
From gene variants distribution to the functional organization of soil microbial systems
Overall, we found strong differences in the functions encoded by satellite and core gene variants. There are several possibilities for gene variants to be identified as core. It could be present in the genome of a single widespread and abundant taxa (generalist), in the genomes of several widespread and low abundance taxa or in the genomes of many spatially restricted and low abundance taxa (specialists). Unfortunately, we could not quantify the contributions of these different scenarii, as the FGA did not allow to link functional genes with the identity of the taxa carrying them. Our results show that core functional gene variants correspond mostly to genes related to the main biogeochemical cycles (C, N, P and S) and support the hypothesis that a wide range of microorganisms have the abilities to carry out fundamental ecological processes such as degradation of C-based substrates, denitrification or assimilatory-N-reduction [79, 80]. This is not surprising from a functional perspective, as microbial systems are well known for their redundancy in the metabolic capabilities between organisms [6] and, at larger scale, across communities [4, 5, 81,82,83,84,85].
By contrast, there are fewer possibilities for a gene variant to be identified as a satellite as it must be present in only spatially restricted and low abundance taxa. Consequently, core variants are expected to represent the functions that are shared by microorganisms with many different macroecological distributions, while satellite variants represent the function that are found only in spatially restricted and low abundance microorganisms. We found that satellite gene variants encoded the capabilities of microorganisms to cope with environmental stresses (e.g., osmotic, oxygen or radiation, cold or heat shocks, sigma factors), withstand nutrient limitations (N and P) and resist to pollutants or potentially toxic compounds (i.e., heavy metals, antibiotics). This result could explain why rare microbes that likely carry these variants appears less affected by disturbances and abiotic changes compared with dominant ones, which tend to respond to a higher number of disturbances and oscillate in abundance when facing them [86]. A step further, these results support previous observations that the ecological strategy of some microorganisms is to maintain a low abundance and a slow growth, while prioritizing the expression of maintenance and survival functions [87, 88]. Our results also support recent findings showing that rare microorganisms are particularly important for the adaptation of microbial communities to environmental variation and their ability to withstand perturbations and maintain ecosystem functions across spatio-temporal scales [41, 89]. In fact, rare and dominant microorganisms are thought to carry redundant metabolic potential regarding major functions (C, N, P cycles), but the rare ones harbor distinct abilities to cope with environmental changes. Hence, they may temporarily thrive and support the functioning at the community level by replacing dominant taxa that were affected by these changes [90,91,92].
It is worth mentioning that we characterized the functional content of microbial communities using FGA, which was constrained by the array design and represented only a fraction of the gene diversity that can be assessed using deep shotgun sequencing [93]. This could have resulted in an underestimation of the functional potential represented by rare genes (and thus the rare biosphere). With the advance in sequencing techniques and expansion of databases, our ability to detect rare genes is rapidly increasing. It is likely that the differences observed here between the functional potential encoded by core and satellite genes would have been even greater using deep shotgun sequencing. However, the FGA approach also offered several advantages over sequencing that are particularly relevant for our study, as it provided a level of reproducibility and standardization that could not be matched by sequencing approaches. Such a standardize microbial data system was recently termed as “highly needed” for pursuing questions related to,microbial macroecology [45]. In addition, microarrays are often more accurate for genes quantification and are more sensitive to rare genes than sequencing approaches [94, 95], making them particularly well suited for analyzing occurrence and abundance patterns of functional genes. To conclude, we foresee that the conceptual approach proposed here could be adapted to the analysis of publicly available metagenomic datasets in order to characterize the distribution of microbial functions across a wide range of environments.
In this study, we showed that the distribution of microbial gene variants can be adequately described using concepts and tools derived from the field of macroecology. This approach allowed us to classify gene variants along a gradient from rarity to commonness, showing that variants with low abundance and limited spatial distribution encode functions that are distinct from those encoded by variants with high abundance and ubiquitous distribution. Common variants encode microbial traits involved in the major biogeochemical cycles (C, N, P and S) while rare ones encode traits allowing microorganisms to withstand environmental stresses and nutrient limitation, along with their resistance to heavy metals and xenobiotics. Our results support the hypothesis that the rare biosphere carries different functional capabilities compared with more prevalent microbes and that these capabilities may determine the essential role of rare microbes in the resilience of microbial communities and their ability to sustain ecological processes across temporal and spatial scales.
References
Gupta A, Sharma VK. Using the taxon-specific genes for the taxonomic classification of bacterial genomes. BMC Genom. 2015;16:1–15.
Gil R, Silva FJ, Pereto J, Moya A. Determination of the Core of a Minimal Bacterial Gene Set. Microbiol Mol Biol Rev. 2004;68:518–37.
Mira A, Martín-Cuadrado AB, D’Auria G, Rodríguez-Valera F. The bacterial pan-genome: a new paradigm in microbiology. Int Microbiol. 2010;13:45–57.
Escalas A, Troussellier M, Yuan T, Bouvier T, Bouvier C, Mouchet MA, et al. Functional diversity and redundancy across fish gut, sediment and water bacterial communities. Environ Microbiol. 2017;19:3268–82.
Jurburg SD, Salles JF. Functional Redundancy and Ecosystem Function — The Soil Microbiota as a Case Study. In: Lo Y-H, Blanco JA, Shovonlal R, editors. Biodiversity in Ecosystems—Linking Structure and Function. BoD–Books on Demand; 2015. p. 29–49.
Louca S, Polz MF, Mazel F, Albright MBN, Huber JA, O’Connor MI, et al. Function and functional redundancy in microbial systems. Nat Ecol Evol. 2018;2:936–43.
Polz MF, Hunt DE, Preheim SP, Weinreich DM. Patterns and mechanisms of genetic and phenotypic differentiation in marine microbes. Philos Trans R Soc Lond B Biol Sci. 2006;361:2009–21.
Young JPW. Bacteria Are Smartphones and Mobile Genes Are Apps. Trends Microbiol. 2016;24:931–2.
Boon E, Meehan CJ, Whidden C, Wong DHJ, Langille MGI, Beiko RG. Interactions in the microbiome: communities of organisms and communities of genes. FEMS Microbiol Rev. 2014;38:90–118.
Escalas A, Hale L, Voordeckers JW, Yang Y, Firestone MK, Alvarez-Cohen L, et al. Microbial Functional Diversity: from Concepts to Applications. Ecol Evol. 2019;5:12000–16.
Barberán A, Casamayor EO, Fierer N. The microbial contribution to macroecology. Front Microbiol. 2014;5:1–8.
Shade A, Dunn RR, Blowes SA, Keil P, Bohannan BJM, Herrmann M, et al. Macroecology to Unite All Life, Large and Small. Trends Ecol Evol. 2018;33:731–44.
Chase AB, Martiny JB. The importance of resolving biogeographic patterns of microbial microdiversity. Microbiol Aust. 2018;1:5–8.
Shoemaker WR, Locey KJ, Lennon JT. A macroecological theory of microbial biodiversity. Nat Ecol Evol. 2017;1:e1450v4.
Bachy C, Worden AZ. Microbial ecology: finding structure in the rare biosphere. Curr Biol. 2014;24:R315–R317.
Lynch MDJ, Neufeld JD. Ecology and exploration of the rare biosphere. Nat Rev Microbiol. 2015;13:217–29.
Pedrós-Alió C. The Rare Bacterial Biosphere. Ann Rev Mar Sci. 2012;4:449–66.
Rabinowitz D. Seven forms of rarity and their frequency in the flora of the British Isles. In: Soulé ME, editors. Conservation biology: the science of scarcity and diversity. Sinauer Associates; Massachusetts; 1986.
McGeoch MA, Gaston KJ. Occupancy frequency distributions: patterns, artefacts and mechanisms. Biol Rev Camb Philos Soc. 2002;77:311–31.
Blackburn TM, Cassey P, Gaston KJ. Variations on a theme: Sources of heterogeneity in the form of the interspecific relationship between abundance and distribution. J Anim Ecol. 2006;75:1426–39.
Buckley HL, Freckleton RP. Understanding the role of species dynamics in abundance-occupancy relationships. J Ecol. 2010;98:645–58.
Gaston KJ, Blackburn TM, Greenwood JJD, Gregory RD, Quinn RM, Lawton JH. Abundance-occupancy relationships. J Appl Ecol. 2000;37:39–59.
Miranda LE, Killgore KJ. Abundance–occupancy patterns in a riverine fish assemblage. Freshw Biol. 2019;64:2221–33.
Suhonen J, Jokimäki J. Temporally stable species occupancy frequency distribution and abundance-occupancy relationship patterns in urban wintering bird assemblages. Front Ecol Evol. 2019;7:129.
Webb TJ, Barry JP, McClain CR. Abundance–occupancy relationships in deep sea wood fall communities. Ecography. 2017;40:1339–47.
Amend AS, Oliver TA, Amaral-Zettler LA, Boetius A, Fuhrman JA, Horner-Devine MC, et al. Macroecological patterns of marine bacteria on a global scale. J Biogeogr. 2013;40:800–11.
Barberán A, Bates ST, Casamayor EO, Fierer N. Using network analysis to explore co-occurrence patterns in soil microbial communities. ISME J. 2012;6:343–51.
Barnes CJ, Burns CA, van der Gast CJ, McNamara NP, Bending GD. Spatio-temporal variation of core and satellite arbuscular mycorrhizal fungus communities in Miscanthus giganteus. Front Microbiol. 2016;7:1–12.
Fillol M, Auguet JC, Casamayor EO, Borrego CM. Insights in the ecology and evolutionary history of the Miscellaneous Crenarchaeotic Group lineage. ISME J. 2016;10:665–77.
Jeanbille M, Gury J, Duran R, Tronczynski J, Agogué H, Saïd OBen, et al. Response of core microbial consortia to chronic hydrocarbon contaminations in coastal sediment habitats. Front Microbiol. 2016;7:1–13.
Lindh MV, Sjöstedt J, Ekstam B, Casini M, Lundin D, Hugerth LW, et al. Metapopulation theory identifies biogeographical patterns among core and satellite marine bacteria scaling from tens to thousands of kilometers. Environ Microbiol. 2017;19:1222–36.
Logares R, Audic SS, Bass D, Bittner L, Boutte C, Christen R, et al. Patterns of Rare and Abundant Marine Microbial Eukaryotes. Curr Biol. 2014;24:813–21.
Michelland R, Thioulouse J, Kyselková M, Grundmann GL. Bacterial Community Structure at the Microscale in Two Different Soils. Micro Ecol. 2016;72:717–24.
Unterseher M, Jumpponen A, Öpik M, Tedersoo L, Moora M, Dormann CF, et al. Species abundance distributions and richness estimations in fungal metagenomics - Lessons learned from community ecology. Mol Ecol. 2011;20:275–85.
Grime JP. Benefits of plant diversity to ecosystems: Immediate, filter and founder effects. J Ecol. 1998;86:902–10.
Grime JP. Dominant and subordinate components of plant communities: implications for succession, sta- bility and diversity. In: Gray AJ, Crawley MJ, editors. Colonization, Succession and Stability. Oxford:Blackwell Scientific Publications; 1984. p. 413–28.
Hanski I. Dynamics of Regional Distribution: the Core and Satellite Species Hypothesis. Oikos. 1982;38:210.
Magurran AE, Henderson PA. Explaining the excess of rare species in natural species abundance distributions. Nature. 2003;422:714–6.
Newton R, Shade A. Lifestyles of rarity: understanding heterotrophic strategies to inform the ecology of the microbial rare biosphere. Aquat Micro Ecol. 2016;78:51–63.
Shade A, Jones SE, Caporaso JG, Handelsman J, Knight R, Fierer N, et al. Conditionally rare taxa disproportionately contribute to temporal changes in microbial diversity. MBio. 2014;5:e01371–14.
Shade A, Gilbert JA. Temporal patterns of rarity provide a more complete view of microbial diversity. Trends Microbiol. 2015;23:335–40.
Koch AL. Oligotrophs versus copiotrophs. BioEssays. 2001;23:657–61.
Cobo-Simón M, Tamames J. Relating genomic characteristics to environmental preferences and ubiquity in different microbial taxa. BMC Genom. 2017;18:1–11.
Tu Q, Yu H, He Z, Deng Y, Wu L, Van Nostrand JD, et al. GeoChip 4: a functional gene-array-based high-throughput environmental technology for microbial community analysis. Mol Ecol Resour. 2014;14:914–28.
Xu X, Wang N, Lipson D, Sinsabaugh R, Schimel J, He L, et al. Microbial macroecology: in search of mechanisms governing microbial biogeographic patterns. Glob Ecol Biogeogr. 2020;29:1870–86.
Reich PB, Knops J, Tilman D, Craine J, Ellsworth D, Tjoelker M, et al. Plant diversity enhances ecosystem responses to elevated CO2 and nitrogen deposition. Nature. 2001;410:809–12.
Field CB, Chapin FS, Chiariello NK, Holland EA, Mooney HA. The Jasper Ridge CO2 Experiment: Design and Motivation. In: Mooney HA, Koch GW, (Editors). Carbon Dioxide and Terrestrial Ecosystems. San Diego, California: Academic Press; 1996. p. 121–45.
Luo C, Rodriguez-R LM, Johnston ER, Wu L, Cheng L, Xue K, et al. Soil microbial community responses to a decade of warming as revealed by comparative metagenomics. Appl Environ Microbiol. 2014;80:1777–86.
Mauritz M, Bracho R, Celis G, Hutchings J, Natali SM, Pegoraro E, et al. Nonlinear CO2 flux response to 7 years of experimentally induced permafrost thaw. Glob Chang. Biol. 2017;23:3646–66.
Natali SM, Schuur EAG, Mauritz M, Schade JD, Celis G, Crummer KG, et al. Permafrost thaw and soil moisture driving CO2 and CH4 release from upland tundra. J Geophys Res Biogeosci. 2015;120:525–37.
Yang Y, Gao Y, Wang S, Xu D, Yu H, Wu L, et al. The microbial gene diversity along an elevation gradient of the Tibetan grassland. ISME J. 2014;8:430–40.
Yang Y, Wu L, Lin Q, Yuan M, Xu D, Yu H, et al. Responses of the functional structure of soil microbial community to livestock grazing in the Tibetan alpine grassland. Glob Chang Biol. 2013;19:637–48.
Zhang Y, Cong J, Lu H, Li G, Xue Y, Deng Y, et al. Soil bacterial diversity patterns and drivers along an elevational gradient on Shennongjia Mountain, China. Micro Biotechnol. 2015;8:739–46.
Zhang Y, Cong J, Lu H, Deng Y, Liu X, Zhou J, et al. Soil bacterial endemism and potential functional redundancy in natural broadleaf forest along a latitudinal gradient. Sci Rep. 2016;6:1–8.
Paula FS, Rodrigues JLM, Zhou J, Wu L, Mueller RC, Mirza BS, et al. Land use change alters functional gene diversity, composition and abundance in Amazon forest soil microbial communities. Mol Ecol. 2014;23:2988–99.
Rodrigues JLM, Pellizari VH, Mueller R, Baek K, Jesus EDC, Paula FS, et al. Conversion of the Amazon rainforest to agriculture results in biotic homogenization of soil bacterial communities. Proc Natl Acad Sci USA. 2013;110:988–93.
He Z, Deng Y, Van Nostrand JD, Tu QC, Xu MY, Hemme CL, et al. GeoChip 3.0 as a high-throughput tool for analyzing microbial community composition, structure and functional activity. Isme J. 2010;4:1167–79.
He Z, Gentry TJ, Schadt CW, Wu L, Liebich J, Chong SC, et al. GeoChip: a comprehensive microarray for investigating biogeochemical, ecological and environmental processes. ISME J. 2007;1:67–77.
Li X, He Z, Zhou J. Selection of optimal oligonucleotide probes for microarrays using multiple criteria, global alignment and parameter estimation. Nucleic Acids Res. 2005;33:6114–23.
Tu Q, He Z, Deng Y, Zhou J. Strain/species-specific probe design for microbial identification microarrays. Appl Environ Microbiol. 2013;79:5085–8.
Wu L, Liu X, Schadt CW, Zhou J. Microarray-based analysis of subnanogram quantities of microbial community DNAs by using whole-community genome amplification. Appl Environ Microbiol. 2006;72:4931–41.
Wu L, Liu X, Schadt CW, Zhou J. Microarray-based analysis of subnanogram quantities of microbial community DNAs by using whole-community genome amplification. Applied and Environmental Microbiology. 2006;72:4931–41.
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’hara RB, et al. Package ‘vegan’. Community ecology package, version. 2013;2:1–295.
Anderson MJ, Bueno AS. A new method for non-parametric multivariate analysis of variance. Austral Ecol. 2001;26:32–46.
Crow EL, Patil GP. Applications in Ecology. In: Cros E, Shimizu K, editors. Lognormal Distributions. New York and Basel:Marcel Dekker; 1988. p. 303–30.
Ser-Giacomi E, Zinger L, Malviya S, De Vargas C, Karsenti E, Bowler C, et al. Ubiquitous abundance distribution of non-dominant plankton across the global ocean. Nat Ecol Evol. 2018;2:1243–9.
Wu L, Ning D, Zhang B, Li Y, Zhang P, Shan X, et al. Global diversity and biogeography of bacterial communities in wastewater treatment plants. Nat Microbiol. 2019;4:1183–95.
Locey KJ, Lennon JT. Scaling laws predict global microbial diversity. Proc Natl Acad Sci. 2016;113:5970–5.
Louca S, Mazel F, Doebeli M, Parfrey LW. A census-based estimate of Earth’s bacterial and archaeal diversity. PLoS Biol. 2019;2:1–30.
Tokeshi M. Dynamics of distribution in animal communities: theory and analysis. Res Popul Ecol (Kyoto). 1992;34:249–73.
Logares R, Deutschmann IM, Junger PC, Giner CR, Krabberød AK, Schmidt TSB, et al. Disentangling the mechanisms shaping the surface ocean microbiota. Microbiome. 2020;8:55.
Azovsky A, Mazei Y. Do microbes have macroecology? Large-scale patterns in the diversity and distribution of marine benthic ciliates. Glob Ecol Biogeogr. 2013;22:163–72.
Noguez AM, Arita HT, Escalante AE, Forney LJ, García-Oliva F, Souza V. Microbial macroecology: highly structured prokaryotic soil assemblages in a tropical deciduous forest. Glob Ecol Biogeogr. 2005;14:241–8.
Thompson LR, Sanders JG, McDonald D, Amir A, Ladau J, Locey KJ, et al. A communal catalogue reveals Earth’s multiscale microbial diversity. Nature. 2017;551:457–63.
Papp L, Izsák J, Papp L, Izsak J. Bimodality in Occurrence Classes: a Direct Consequence of Lognormal or Logarithmic Series Distribution of Abundances- A Numerical Experimentation. Oikos. 1997;79:191.
Verberk WCEP, van der Velde G, Esselink H. Explaining abundance-occupancy relationships in specialists and generalists: A case study on aquatic macroinvertebrates in standing waters. J Anim Ecol. 2010;79:589–601.
Liao J, Cao X, Zhao L, Wang J, Gao Z, Wang MC, et al. The importance of neutral and niche processes for bacterial community assembly differs between habitat generalists and specialists. FEMS Microbiol Ecol. 2016;92:fiw174.
Slatyer RA, Hirst M, Sexton JP. Niche breadth predicts geographical range size: a general ecological pattern. Ecol Lett. 2013;16:1104–14.
Fierer N, Barberán A, Laughlin DC. Seeing the forest for the genes: using metagenomics to infer the aggregated traits of microbial communities. Front Microbiol. 2014;5:1–6.
Rivett DW, Bell T. Abundance determines the functional role of bacterial phylotypes in complex communities. Nat Microbiol. 2018;3:767–72.
Wertz S, Degrange V, Prosser JI, Poly F, Commeaux C, Guillaumaud N, et al. Decline of soil microbial diversity does not influence the resistance and resilience of key soil microbial functional groups following a model disturbance. Environ Microbiol. 2007;9:2211–9.
Wertz S, Degrange V, Prosser JI, Poly F, Commeaux C, Freitag T, et al. Maintenance of soil functioning following erosion of microbial diversity. Environ Microbiol. 2006;8:2162–9.
Mendes LW, Tsai SM, Navarrete AA, de Hollander M, van Veen JA, Kuramae EE. Soil-Borne microbiome: linking diversity to function. Micro Ecol. 2015;70:255–65.
Sunagawa S, Coelho LP, Chaffron S, Kultima JR, Labadie K, Salazar G, et al. Structure and function of the global ocean microbiome - SM. Science. 2015;348:1261359–1261359.
Wohl DL, Arora S, Gladstone JR. Functional redundancy supports biodiversity and ecosystem function in a cloased and constant environment. Ecology. 2008;85:1534–40.
Kurm V, Geisen S, Gera Hol WH. A low proportion of rare bacterial taxa responds to abiotic changes compared with dominant taxa. Environ Microbiol. 2019;21:750–8.
Bergkessel M, Basta DW, Newman DK. The physiology of growth arrest: Uniting molecular and environmental microbiology. Nat Rev Microbiol. 2016;14:549–62.
Hofer U. Life in the slow lane. Nat Rev Microbiol. 2019;26:266–7.
Baho DL, Peter H, Tranvik LJ. Resistance and resilience of microbial communities - Temporal and spatial insurance against perturbations. Environ Microbiol. 2012;9:2283–92.
Jousset A, Bienhold C, Chatzinotas A, Gallien L, Gobet A, Kurm V, et al. Where less may be more: how the rare biosphere pulls ecosystems strings. ISME J. 2017;11:853–62.
Aanderud ZT, Jones SE, Fierer N, Lennon JT. Resuscitation of the rare biosphere contributes to pulses of ecosystem activity. Front Microbiol. 2015;6:1–11.
Lawson CE, Strachan BJ, Hanson NW, Hahn AS, Hall ER, Rabinowitz B, et al. Rare taxa have potential to make metabolic contributions in enhanced biological phosphorus removal ecosystems. Environ Microbiol. 2015;17:4979–93.
Zhou J, He Z, Yang Y, Deng Y, Tringe SG, Alvarez-Cohen L. High-throughput metagenomic technologies for complex microbial community analysis: open and closed formats. MBio. 2015;6:e02288–14.
Zhou J, Wu L, Deng Y, Zhi X, Jiang YH, Tu Q, et al. Reproducibility and quantitation of amplicon sequencing-based detection. ISME J. 2011;5:1303–13.
Shi Z, Yin H, Van Nostrand JD, Voordeckers JW, Tu Q, Deng Y, et al. Functional Gene Array-Based Ultrasensitive and Quantitative Detection of Microbial Populations in Complex Communities. mSystems. 2019;4:99–117.
Acknowledgements
The authors would like to thank all the persons that contributed to sample collection and laboratory analyses. This synthesis was primarily funded by the U.S. Department of Energy (DOE), Office of Science, Office of Biological and Environmental Research’s (OBER) Systems Biology Research to Advance Sustainable Bioenergy Crop Development (DE-SC0014079), Biological Systems Research on the Role of Microbial Communities in Carbon Cycling program (DE-SC0004730, DE-SC001057, DE-SC0004601 and DE-SC0010715), by the U.S. National Science Foundation MacroSystems Biology program under the contract (NSF EF-1065844), and by the Office of the Vice President for Research at the University of Oklahoma, all to JZ. This work was also supported by the National Natural Science Foundation of China (No.31670614) to YY.
Author information
Authors and Affiliations
Contributions
AE developed the original concept of the study. FP, MY, FL, JF, LW, YZ, and YY provided experimental data. AE and FG performed data analyses. AE wrote the first draft of the paper with help from FP, FG, MY, YY, FL, JF, LW, YZ, and JZ.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Escalas, A., Paula, F.S., Guilhaumon, F. et al. Macroecological distributions of gene variants highlight the functional organization of soil microbial systems. ISME J 16, 726–737 (2022). https://doi.org/10.1038/s41396-021-01120-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-021-01120-8
This article is cited by
-
Microbial Journey: Mount Everest to Mars
Indian Journal of Microbiology (2022)