Abstract
Microorganisms drive soil carbon mineralization and changes in their activity with increased temperature could feedback to climate change. Variation in microbial biodiversity and the temperature sensitivities (Q10) of individual taxa may explain differences in the Q10 of soil respiration, a possibility not previously examined due to methodological limitations. Here, we show phylogenetic and taxonomic variation in the Q10 of growth (5–35 °C) among soil bacteria from four sites, one from each of Arctic, boreal, temperate, and tropical biomes. Differences in the temperature sensitivities of taxa and the taxonomic composition of communities determined community-assembled bacterial growth Q10, which was strongly predictive of soil respiration Q10 within and across biomes. Our results suggest community-assembled traits of microbial taxa may enable enhanced prediction of carbon cycling feedbacks to climate change in ecosystems across the globe.
Similar content being viewed by others
Introduction
Soil is the largest terrestrial reservoir of carbon (C), yet this C pool is not well constrained in global climate models due to uncertainty regarding microbial feedbacks to climatic change [1,2,3]. Soil organic carbon (SOC) is mineralized by microorganisms and returns to the atmosphere as CO2. Changes in this soil C flux, therefore, could substantially influence the direction and magnitude of future climate changes [4]. Since the decomposition of SOC is temperature-dependent, most C cycling models and studies predict that increasing temperature will lead to decreasing soil C storage due to warming-induced increases in microbial activity and soil respiration [5, 6]. Yet, experimental studies show conflicting effects of warming on soil C storage [7] and suggest our understanding of microbial feedbacks to temperature is incomplete. Understanding the variation in microbial community traits and their responses to temperature under climate warming could elucidate the contradictory effects of temperature on soil C cycling and improve our ability to make accurate predictions.
Soil microbial respiration is sensitive to temperature [8, 9], reflecting the temperature sensitivities of microbial growth and metabolism, such as enzyme activities and C use efficiency [10, 11]. The considerable variation observed in temperature sensitivity of microbial processes at the community level may reflect dependencies on community membership [12]. Classical culture-based approaches provide clear evidence that microbial species differ in their minimum, optimum, and maximum growth temperatures [13,14,15,16]. Consequently, differences in community composition, and the temperature sensitivities of individual taxa, could explain differences in temperature sensitivity of soil respiration. Indeed, consistent shifts in relative abundance have been reported for ubiquitous soil bacteria and archaea in response to warming [17]. But the knowledge of microbial community composition alone offers weak predictive power for microbial process rates [18, 19]. Direct measurements of growth and temperature sensitivity of individual microbial taxa could illuminate and help to predict their impact on ecosystem functioning.
We applied quantitative stable isotope probing (qSIP) [20, 21], a technique that tracks the incorporation of 18O-water into newly synthesized DNA, to quantify the growth of individual soil microbial taxa and their temperature sensitivities in four ecosystems, representing Arctic, boreal, temperate, and tropical biomes (Supplementary Fig. 1). These biomes include an Arctic tundra in Alaska, a boreal forest in northern Minnesota, a temperate mixed conifer forest in northern Arizona and a tropical forest in Puerto Rico and encompass a large range in mean annual temperature (MAT) from −7 to 24 °C (Supplementary Table 1). Each soil was incubated under four temperatures (5, 15, 25, and 35 °C) for 5 days. The temperature sensitivity (Q10) of whole community respiration or growth of individual taxa was calculated as Q10 = Rtemp + 10/Rtemp, where R is the microbial respiration (μg C g−1 soil) or growth (day−1) at the lower temperature (temp) and higher temperature (temp + 10) [22]. We hypothesized that when the temperature sensitivities of individual taxa were aggregated, the community level microbial growth Q10 would correlate with the soil respiration Q10. We also predicted that the microbial growth Q10 would be phylogenetically clustered because the temperature sensitivities of organisms are likely to be dependent upon central gene systems encoding metabolic enzymes that evolve slowly and are rarely transferred between organisms [23, 24].
Method
Sample collection and incubation
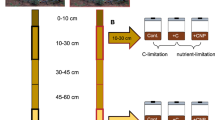
In August 2017, soil samples were collected from four ecosystems, with one site in each of the Arctic, boreal, temperate, and tropical biomes (Supplementary Fig. 1). The Arctic samples were from the Arctic LTER site at Toolik Lake Field Station. The boreal samples were from the SPRUCE experimental site (Spruce and Peatland Responses Under Changing Environments) in northern Minnesota. The temperate samples were collected from a mixed conifer forest site, located at the Hart Prairie Nature Reserve in northern Arizona, and the tropical samples were from the Sabana Field Research Station in the Luquillo Experimental Forest in Puerto Rico. The sites represent common ecosystem types within each climactic zone and have relevant data available from past studies [25, 26] but are merely samples from each biome. We acknowledge that our sites do not encompass the diversity of ecosystems within the Artic, boreal, temperate and tropical climatic zones, and advise caution when generalizing our results. We selected these sites as a first attempt to compare microbial growth and respiration responses to temperature at a broad scale. Detailed information of sampling sites can be found in Supplementary Table 1. Five independent replicate samples were collected from each site at 0–10-cm depth and stored at either room temperature (temperate and tropical soils) or 4 °C (boreal and Arctic) until they arrived in the lab.
Respiration measurements
For the respiration measurements, the homogenized soils, ranging from 1.2 to 12.5-g dry weight due to the different amount of soil we had for different sites, were weighed into specimen cups and placed into mason jars with a septum in the lid for headspace gas sampling (n = 5 for each temperature and biome). Soil moisture was adjusted to 60% water holding capacity (WHC) and the samples were incubated 5 days at 5, 15, 25, and 35 °C, respectively. To measure CO2 concentrations, a t0 gas sample was taken ~15 min after the jar was closed. Headspace gas samples were then taken every ~24 h from the 25 and 35 °C temperatures, and every ~48 h from the 5 and 15 °C temperatures. The headspace gas samples removed from the jars with 15-mL syringes were analyzed with a LI-Cor 6262 (LI-Cor Biosciences Inc. Lincoln, Nebraska, USA). The cumulative CO2–C was calculated as the difference in CO2–C from t0 to t24–48 h. Following gas sampling at 24 or 48 h, jars were opened for ~15–30 min to release accumulated CO2 and equilibrate to atmospheric CO2 concentrations before being closed again and measurements were repeated over the 5-day incubation period.
Microbial biomass C measurements
Microbial biomass C was measured after 5-day incubation following a revised protocol based on the chloroform-fumigation extraction methods described by Witt et al. [27]. Soils were weighed into Nalgene bottles and extracted in replicate pairs (fumigated and non-fumigated) using a 1:5 soil: extractant ratio with 0.5-M K2SO4. The fumigated replicate had EtOH-free chloroform added directly. All samples were shaken at low speed (~150 rpm) for 4 h before being vacuum-filtered through a Whatman GF/A filter. All fumigated samples containing chloroform were sparged with compressed air for 1 h in a Falcon tube, allowing the chloroform to pool at the bottom and be bubbled out. Filtered and sparged samples were frozen until further analysis. Thawed samples were analyzed within 48 h for total organic carbon (TOC) on a Shimadzu TOC-L analyzer with an ASI-L autosampler (Shimadzu Corporation, Kyoto, Japan). The average blank values were subtracted from TOC to correct for any C present in the extractant solution. Microbial biomass C was calculated as the difference between the fumigated and non-fumigated replicates.
Quantitative stable isotope probing (qSIP)
For qSIP, 1–2-g dry mass soil from each biome was weighed into Falcon tubes. Samples were preincubated at 5, 15, 25, and 35 °C for 12 h. After 12 h, samples (n = 5) were adjusted to 60% WHC by adding 18O-enriched water (98 atom%) and were incubated at their respective temperatures for an additional 5 days. This resulted in 80 18O-enriched t5 samples (five replicates soil samples from four biomes at four different temperatures). An additional 25 t0 samples were analyzed for one or two of the temperatures for each replicate from each biome (15 °C for Arctic, 15 °C boreal, 5 and 15 °C for temperate, and 25 °C for tropical). The samples stored at −80 °C immediately following natural abundance water addition (t0) or after the incubation (t5). Total DNA was extracted from 0.5 g of soil using a MoBio PowerSoil Kit (Carlsbad, CA, USA) following the manufacturer’s instructions. To separate DNA by density, 1 μg of DNA was added to 3.6 mL of a saturated cesium chloride solution (1.9 g mL−1), ~1.1 mL of gradient buffer (200-mM trisaminomethane) (Tris), pH 8, 200-mM potassium chloride, 2-mM ethylenediaminetetraacetic acid in an OptiSeal ultracentrifuge tube (Beckman Coulter, Brea, CA). The tubes were spun at 127,000 × g at 18 °C for 72 h using a Beckman TLN-100 rotor in an Optima MAX ultracentrifuge (Beckman Coulter, Brea, CA).
After spinning, ~3.5 mL of the cesium chloride gradient was separated into twenty-two 150-μL fractions. The density of each fraction was measured using a Reichert AR 200 handheld digital refractometer (Reichert Technologies, Buffalo, NY). Following, DNA in each fraction was purified using an isopropanol precipitation method and quantified by PicoGreen fluorescence on a BioTek Synergy HT plate reader (Winooski, VT, USA). The 16S rRNA gene was quantified and sequenced in each fraction within the density range 1.64–1.73 g/mL (~17 fractions). Standard curves were generated using tenfold serial dilutions of genomic Escherichia coli DNA.
For qPCR, triplicate 10-μL reactions contained 0.25 μM of each primer (Eub 338 5′-ACT CCT ACG GGA GGC AGC AG-3′/Eub 518 5′-ATT ACC GCG GCT GCT GG-3′), 1X Forget-Me-Not EvaGreen qPCR Master Mix, and 1.5-mM MgCl2. The assay was performed on a CFX 384 (Bio-Rad, Hercules, USA), using a program of 95 °C for 2 min followed by 40 cycles of 95 °C for 10 s, 62 °C for 10 s, and 72 °C for 10 s. Bacterial gene copy numbers were calculated using a regression equation for each assay relating the cycle threshold (Ct) value to the known number of copies in the standards.
For sequencing, two PCR steps were used [28]. Each sample was first amplified using primers 515F (5′-GTGYCAGCMGCCGCGGTA-3′) and 806R (5′-GGACTACNVGGGTWTCTAAT-3′) [29, 30]. This was done in triplicate 10-µL PCR reactions containing 1 mM of each primer, 0.01-U/µL Phusion HotStart II Polymerase (Thermo Fisher Scientific, Waltham MA, USA), 1X Phusion HF buffer (Thermo Fisher Scientific, Waltham MA, USA), 3.0-mM MgCl2, 6% glycerol, and 200-µM dNTPs. PCR conditions were 95 °C for 2 min; 15 cycles of 95 °C for 30 s, 55 °C for 30 s, and 60 °C for 30 s. Initial PCR reaction products were pooled, checked on a 1% agarose gel, tenfold diluted, and used as template in the subsequent tailing PCR reaction with region-specific primers that included the Illumina flow cell adapter sequences and a 12 nucleotide Golay barcode (15 cycles identical to initial amplification conditions). Products of the tailing reaction were purified with carboxylated SeraMag Speed Beads (Sigma-Aldrich, St. Louis, USA) at a 1:1 v/v ratio as described in Rohland and Reich [31], and quantified by Picogreen fluorescence (Thermo Fisher, USA). Equal concentrations of the reaction products were then pooled; the library was bead-purified once again (1:1 ratio), quantified by qPCR using the Library Quantification Kit for Illumina (Kapa Biosciences, Woburn, MA), and loaded at 11 pM (including a 30% PhiX control) onto an Illumina MiSeq instrument (San Diego, CA, USA) using 2 × 150 paired-end read chemistry.
Sequence data processing and analysis
Sequencing data were analyzed for each biome separately. The paired-end reads were joined [32] and analyzed with the QIIME software package (version 1.8, available at http://www.qiime.org) [33]. The most abundant sequence for each operational taxonomic unit (OTU) was aligned [34], and taxonomy was assigned with UCLUST [35] using the SILVA 16S rRNA gene database (version 128, available at https://www.arb-silva.de/documentation/release-128) [36] at 97% identity. Any OTUs that accounted for <0.05% of the total sequences were discarded [37]. In addition, samples with <3500 sequences were excluded from the analysis. Isotopic composition of individual taxa after exposure to H218O was calculated as described by Hungate et al. [20]. Briefly, the density for each bacterial taxon was computed as a weighted average, summing the densities across all fractions multiplied by the total number of 16S rRNA gene copies in that fraction expressed as a proportion of the total 16S rRNA gene copies. The increase in weighted density relative to the weighted density of the unlabeled treatments was calculated. We determined the GC content of the DNA for each taxon, based on its density, using the relationship of GC content and density based on a pure culture study. The GC content was then used to calculate the molecular weights and the corresponding values of 18O isotope composition for each taxon. Bootstrap resampling (with replacement, 1000 iterations) of replicates within each treatment was used to estimate taxon-specific 90% confidence intervals for the change in density and the corresponding value of 18O excess atom fraction (EAF) isotope composition. Preliminary data analysis revealed an effect of ultracentrifuge tube on estimation of weighted average density of a taxon, probably a consequence of slight differences in CsCl density gradients between tubes. This technical error was corrected as previously described [38]. We then corrected for the differences in pore water enrichment by dividing the EAF of a taxon in a sample by the proportion of 18O–H2O in that sample. The relative growth rate (day−1) was estimated as a function of the increase of 18O–EAF of microbial DNA, which assumed that 33% of oxygen in microbial DNA was derived from water and that the community is at steady state: growth = 18O–EAF/(0.33 × 5 days) [20, 39]. The production of new microbial biomass (μg C g−1 soil) at different temperatures was estimated, which considered the increase in community-weighted 18O–EAF of DNA as the proportion of new biomass produced in total microbial biomass. Code associated with qSIP calculations is publicly available at https://bitbucket.org/QuantitativeSIP/qsip_repo. All sequences in this study are available in Sequence Read Achieve database of NCBI under accession numbers PRJNA649787, PRJNA649546, PRJNA649571, and PRJNA649802.
Null model approach
The null model approach has widely been applied to study the community assembly processes [40,41,42]. Appropriate null models randomize the characteristics of the observed data that are involved in the null hypothesis, while maintaining structures in the dataset unrelated to the null hypothesis [41, 43]. Here, we tested if the relationship between community-traits-based growth Q10 and respiration Q10 could be observed in randomly assembled microbial communities. We randomized species abundances within samples, which maintained the observed species richness within communities. This procedure limited any bias in null predictions due to differences in the number of species observed in the communities, as this feature of microbial communities is a key driver of multiple ecosystem functions [44]. We ran 10,000 null model randomizations for each temperature range using species abundance matrices that included all study sites within a given temperature range (Supplementary Fig. 2). For each of the 10,000 randomizations, we calculated the community-traits-based growth Q10 using mean trait values for each species at each temperature range. This analysis was conducted using “Picante” package in R 3.1.1 [45].
Using the null distribution of Q10 values, we calculated standardized effect sizes as the difference between the randomized community-traits-based growth Q10 of a given community and its observed value, divided by the standard deviation. We then calculated p value as the number of randomizations with modeled growth Q10 ≤ observed Q10 divided by the total number of randomizations [46]. Further, we calculated the 10,000 randomly generated relationships between respiration Q10 and growth Q10 and extracted the slope, the y-intercept and the adjusted R2 of the 10,000 random relationships. We compared coefficients from the randomized dataset with observed parameters for the null models. The p values associated with regression coefficients were calculated as described previously.
Statistical analyses
Temperature sensitivity (Q10) at each of three temperature ranges (5–15 °C, 15–25 °C and 25–35 °C) was estimated as Q10 = Rtemp + 10/Rtemp, where R is the growth or respiration rate at the higher temperature (temp + 10) and lower temperature (temp) [22]. A microbial community-weighted mean (CWM) growth Q10 was calculated following [47]:
where pi is the relative abundance of microbial species to abundance of community, n is the number of species, and Q10_i is the Q10 value of species i. While we acknowledge that Q10 is an imperfect measure of temperature sensitivity, newer approaches such as the macromolecular rate theory require a large number of independent measurements to produce robust model estimates [48]. Given the constraints of our experiment, Q10 was selected because it could be applied to individual replicates of both growth and respiration allowing a robust statistical analysis of the relationship between these variables. Two-way ANOVA was applied to test for the differences in respiration, growth, and Q10 with incubation temperature, biome, and their interaction. The ordinary least square method was applied for all regressions in this study. To determine how the temperature sensitivity of growth varied among microbial species and phylogenetic groups, we assessed phylogenetic clustering using Blomberg’s K and Pagel’s λ phylogenetic signal tests [38]. The phylogenetic analysis was conducted using the “phytools” and “phylobase” packages in R 3.5.5 [45]. The growth Q10 of individual taxa were categorized based on their maximum growth Q10 among 5–15, 15–25, and 25–35 °C as “cold,” “moderate,” or “warm” responders. The phylogenetic tree was visualized using the online tool, Interactive Tree Of Life (version 5.7, https://itol.embl.de) [49].
Results and discussions
Q 10 of soil respiration
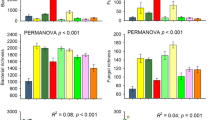
Cumulative soil respiration increased with temperature (Fig. 1A) and was generally higher in soil from the cold biomes (Arctic and boreal) relative to soil from the warm biomes (temperate and tropical) (Supplementary Table 2). In general, the Q10 values of respiration were high at the low temperature range (5–15 °C) and declined as temperature increased (Fig. 1B). This trend held for all biomes, although the differences were not significant in soil from the Arctic (Supplementary Table 3). This negative relationship between Q10 and temperature is consistent with numerous empirical studies [9, 22, 50].
Cumulative soil respiration (A) and microbial growth rate (C) for soils from four biomes over a 5-day incubation at four temperatures (5–35 °C). Soil respiration temperature sensitivity (Q10, B) and microbial growth rate temperature sensitivity (Q10, D) were calculated for each soil between incubation temperatures (mean ± standard error). Statistical analysis of the differences for each variable among temperatures can be found in Supplementary Tables 2 and 3.
Q 10 of microbial growth
In order to relate soil respiration to microbial activity, we used qSIP with 18O–H2O to measure the relative growth rate (day−1) of each individual taxon and aggregated these values to produce relative abundance weighted, community level measures of growth rate [20, 21]. Microbial growth generally increased with increasing incubation temperature for all biomes except for the temperate biome, which showed a maximum growth rate at 15 °C (Fig. 1C). The magnitude of temperature effects was dependent upon the biome from which the soil was collected (Supplementary Table 2). These differences likely result from biotic and abiotic differences between these ecosystems such as variation in microbial community composition (discussed below) and SOC content, which can influence microbial growth rate [51]. We also estimated new microbial biomass production (μg C g−1 soil); as expected, more soil microbial biomass was produced at high temperatures. Biomass production was strongly correlated with soil respiration rate for each biome (Supplementary Fig. 3) and across all biomes (Fig. 2, R2 = 0.68, p < 0.01). These results suggest that microbial growth dynamics are responsible for changes in soil respiration rate with temperature [12]. When comparing across biomes, the slope of relationship between biomass production and respiration covaried with biome MAT. Higher slope values were observed at the sites with high MAT (Supplementary Fig. 4), which suggests that microbes have higher C use efficiency in the soil with low MAT under increasing temperature. Alternatively, the accumulation of SOC in the colder boreal and Arctic ecosystems may facilitate high microbial carbon use efficiency (CUE) in these systems, as past work has shown microbial CUE to be positively correlated with SOC content at a regional scale [52].
Using the same principle as for soil respiration [9, 22], we determined the Q10 of microbial growth for individual taxa and aggregated to determine CWM values [53, 54]. The mean Q10 of microbial growth for the whole community (weighted by microbial relative abundance) declined with increasing incubation temperature and varied across biomes (Fig. 1D and Supplementary Table 3). Consistent with our first hypothesis, community level microbial growth Q10 correlated strongly with the Q10 of soil respiration for each biome (Supplementary Fig. 5) and across all biomes (Fig. 3, R2 = 0.62, p < 0.01). Our findings are consistent with vegetation-based studies, which find CWM traits to be strong predictors of ecosystem properties aboveground [55, 56] and demonstrate a similar approach may be applied with microorganisms belowground. Moreover, our results suggest the temperature dependency of microbial growth and respiration are coupled across a wide range of biomes with different soil properties (e.g., SOC, pH, etc.) and climates. This suggest warming-induced changes in microbial physiological parameters, such as growth rate, may be important factors in regulating the sensitivity of organic C decomposition to temperature in soil [57]. Consequently, understanding the temperature sensitivity of microbial growth and including this information in climate change models may better predict soil C dynamics under global warming.
Significance of microbial community composition
Variation in the temperature sensitivity of microbial growth across biomes could be due to differences in soil community composition. To determine the significance of community membership for microbial growth Q10, we tested our observed data against a null model of random community assembly (Supplementary Fig. 2). Approximately 40% of community-level measures of microbial growth Q10 had a low probability of occurrence (p < 0.05) when the presence and relative abundance of taxa were randomized (Supplementary Fig. 6). Similarly, the null model was unable to reproduce the strong relationship observed between the temperature sensitivities of microbial growth and soil respiration (Supplementary Fig. 6). Taken together these results suggest that presence, relative abundance, and trait (i.e., Q10) values of individual microbial taxa drive the community level temperature sensitivity in microbial growth, which in turn influences the temperature sensitivity of soil respiration. This finding highlights the importance of considering biodiversity and taxon-specific functional traits to understand ecosystem processes [40, 55] including those performed by microorganisms.
Phylogeny of Q 10 of microbial growth
The temperature sensitivity of growth varied among microbial species and phylogenetic groups, exhibiting phylogenetic clustering according to Blomberg’s K and Pagel’s λ phylogenetic signal tests [38]. The Q10 of microbial growth rates were non-randomly distributed in boreal, temperate, and tropical biomes across all temperature ranges (Supplementary Table 4), with weaker (and generally nonsignificant) phylogenetic signals in the Arctic soil. Taken together, these patterns support our second hypothesis and show a nonrandom phylogenetic distribution for temperature sensitivity for microbial taxa: the growth rates of closely related organisms exhibited similar temperature sensitivities, indicating that temperature sensitivity is evolutionarily conserved. Other microbial traits also tend to be distributed in phylogenetic clusters, including oxygenic photosynthesis [23], nitrogen fixation [23], nitrogen assimilation [58], C decomposition [38], and oxygen consumption [59], all traits that govern element transformations in nature. Such phylogenetic organization could help build a framework for how microbial diversity underpins the functioning of ecosystems. While our analysis focused on the diversity and community composition of soil bacteria, future studies should aim to encompass all soil organisms including fungi and protozoa, which contribute soil respiration [60] and likely influence soil C responses to temperature.
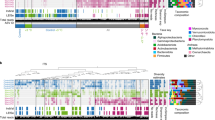
To visualize phylogenetic patterns within and across biomes, we grouped taxa into three categories based on the temperature at which maximum growth Q10 was observed: “cold” (5–15 °C), “moderate” (15–25 °C), and “warm” (25–35 °C) responders. Then, the proportion of taxa in each category was summarized at the class level for each biome (Supplementary Fig. 7) and across four biomes (Fig. 4). The observed maximum growth Q10 of microorganisms within our soils is likely to depend upon both direct effects of temperature on microbial physiology [10, 15] and indirect effects, as microbes respond to temperature-induced changes in their physiochemical environment (e.g., changes in dissolved resource availability [61]). Despite likely temperature–environment interactions [17], the maximum growth Q10 of growth for some taxonomic groups was consistent across the biomes. For instance, 67% of the taxa in Planctomycetacia as well as most Sphingobacteriia (83%) and Betaproteobacteria (68%) were most sensitive at lower temperatures (5–15 °C) across biomes. Many Nitrospira (58%) and the Subgroup 2 (36%) of Acidobacteria were most sensitive at higher temperatures (25–35 °C). A majority of Acidimicrobiia (55%), Spartobacteria (54%), Deltaproteobacteria (51%), and many Thermoleophilia (48%) were more sensitive at a moderate temperature range (15–25 °C). Consistency in the maximum growth Q10 of taxa within these groups in soils from distinct biomes suggests that these temperature responses may be robust to environmental variation. In addition, the majority of taxa were cold responders in the Arctic (58%) and temperate (53%) biomes, while moderate temperature responders were most common in boreal (45%) and tropical (46%) biomes. Therefore, these results indicate that assuming all soil microbes exhibit identical temperature sensitivities at large scale could result in substantial uncertainty of microbial respiration rate [62].
A Each phylotype was categorized based on its maximum Q10 among 5–15, 15–25, and 25–35 °C as “cold,” “moderate,” or “warm” responders. The pie charts represent proportion of growth Q10 values in each response category at class level (with exception for Chloroflexi phyla) using data from all four sites. B Total proportion of growth Q10 response for each biome and across all biomes. Phylogenetic statistical analysis for growth Q10 was provided in Supplementary Table 4.
Conclusions
In summary, growth and its temperature sensitivity varied among microbial taxa within natural soil communities across a broad range of temperatures and biomes, and the variation was phylogenetically constrained. When taxon-specific measures were aggregated to CWMs, the temperature sensitivities of microbial growth were strongly and positively related to total soil respiration. This relationship could not be reproduced by random community assembly, indicating that biodiversity shapes microbial growth aggregated at the community level, in turn regulating soil C mineralization in response to warming. This work demonstrates how the traits of individual microbial taxa can be aggregated to predict community level processes such as changes in soil respiration due to temperature. The strong relationships we observed between microbial and biogeochemical parameters (Figs. 2 and 3) suggest that incorporation of microbial data, perhaps via the development of trait-based models that consider biodiversity, may reduce model uncertainties and allow us to better understand the future of our warming world.
Data availability
Soil respiration, microbial biomass, and figure data are available in the Supplementary materials. Raw sequence data for this study are available in Sequence Read Achieve (SRA) database of NCBI under accession numbers PRJNA649787, PRJNA649546, PRJNA649571, and PRJNA649802.
Code availability
Code associated with qSIP calculations is publicly available at https://bitbucket.org/QuantitativeSIP/qsip_repo.
References
Bradford MA, Wieder WR, Bonan GB, Fierer N, Raymond PA, Crowther TW. Managing uncertainty in soil carbon feedbacks to climate change. Nat Clim Chang. 2016;6:751–8.
Cavicchioli R, Ripple WJ, Timmis KN, Azam F, Bakken LR, Baylis M, et al. Scientists’ warning to humanity: microorganisms and climate change. Nat Rev Microbiol. 2019;17:569–86.
Crowther TW, Hoogen JVD, Wan J, Mayes MA, Keiser AD, Mo L, et al. The global soil community and its influence on biogeochemistry. Science. 2019;365:eaav0550.
Heimann M, Reichstein M. Terrestrial ecosystem carbon dynamics and climate feedbacks. Nature. 2008;451:289–92.
Wieder WR, Bonan GB, Allison SD. Global soil carbon projections are improved by modelling microbial processes. Nat Clim Chang. 2013;3:909–12.
Walker TWN, Kaiser C, Strasser F, Herbold CW, Leblans NIW, Woebken D, et al. Microbial temperature sensitivity and biomass change explain soil carbon loss with warming. Nat Clim Chang. 2018;8:885–9.
Crowther TW, Todd-Brown KEO, Rowe CW, Wieder WR, Carey JC, Machmuller MB, et al. Quantifying global soil carbon losses in response to warming. Nature. 2016;540:104–8.
Li JQ, Pei JM, Pendall E, Fang CM, Nie M. Spatial heterogeneity of temperature sensitivity of soil respiration: a global analysis of field observations. Soil Biol Biochem. 2020;141:107675.
Wang QK, Zhao XC, Chen LC, Yang QP, Chen S, Zhang WD, et al. Global synthesis of temperature sensitivity of soil organic carbon decomposition: latitudinal patterns and mechanisms. Funct Ecol. 2019;33:514–23.
Nottingham AT, Baath E, Reischke S, Salinas N, Meir P. Adaptation of soil microbial growth to temperature: using a tropical elevation gradient to predict future changes. Glob Chang Biol. 2019;25:827–38.
Ye JS, Bradford MA, Dacal M, Maestre FT, García-Palacios P. Increasing microbial carbon use efficiency with warming predicts soil heterotrophic respiration globally. Glob Chang Biol. 2019;25:3354–64.
Smith TP, Thomas TJH, Garcia-Carreras B, Sal S, Yvon-Durocher G, Bell T, et al. Community-level respiration of prokaryotic microbes may rise with global warming. Nat Commun. 2019;10:5124.
Schipper LA, Hobbs JK, Rutledge S, Arcus VL. Thermodynamic theory explains the temperature optima of soil microbial processes and high Q10 values at low temperatures. Glob Chang Biol 2014;20:3578–86.
Pietikainen J, Pettersson M, Baath E. Comparison of temperature effects on soil respiration and bacterial and fungal growth rates. FEMS Microbiol Ecol. 2005;52:49–58.
Bárcenas-Moreno G, Gómez-Brandón M, Rousk J, Bååth E. Adaptation of soil microbial communities to temperature: comparison of fungi and bacteria in a laboratory experiment. Glob Chang Biol. 2009;15:2950–7.
Engqvist MKM. Correlating enzyme annotations with a large set of microbial growth temperatures reveals metabolic adaptations to growth at diverse temperatures. BMC Microbiol. 2018;18:177.
Oliverio AM, Bradford MA, Fierer N. Identifying the microbial taxa that consistently respond to soil warming across time and space. Glob Chang Biol. 2017;23:2117–29.
Bier RL, Bernhardt ES, Boot CM, Graham EB, Hall EK, Lennon JT, et al. Linking microbial community structure and microbial processes: an empirical and conceptual overview. FEMS Microbiol Ecol. 2015;91:fiv113.
Dubey A, Malla MA, Khan F, Chowdhary K, Yadav S, Kumar A, et al. Soil microbiome: a key player for conservation of soil health under changing climate. Biodivers Conserv. 2019;28:2405–29.
Hungate BA, Mau RL, Schwartz E, Caporaso JG, Dijkstra P, van Gestel N, et al. Quantitative microbial ecology through stable isotope probing. Appl Environ Microbiol. 2015;81:7570–81.
Koch BJ, McHugh TA, Hayer M, Schwartz E, Blazewicz SJ, Dijkstra P, et al. Estimating taxon-specific population dynamics in diverse microbial communities. Ecosphere. 2018;9:e02090.
Hamdi S, Moyano F, Sall S, Bernoux M, Chevallier T. Synthesis analysis of the temperature sensitivity of soil respiration from laboratory studies in relation to incubation methods and soil conditions. Soil Biol Biochem. 2013;58:115–26.
Martiny AC, Treseder K, Pusch G. Phylogenetic conservatism of functional traits in microorganisms. ISME J. 2013;7:830–8.
DeAngelis KM, Pold G, Topçuoğlu BD, van Diepen LTA, Varney RM, Blanchard JL, et al. Long-term forest soil warming alters microbial communities in temperate forest soils. Front Microbiol. 2015;6:104.
Euskirchen ES, Bret-Harte MS, Shaver GR, Edgar CW, Romanovsky VE. Long-term release of carbon dioxide from Arctic Tundra ecosystems in Alaska. Ecosystems. 2017;20:960–74.
Reed SC, Reibold R, Cavaleri MA, Alonso-Rodríguez AM, Berberich ME, Wood TE. Chapter six—soil biogeochemical responses of a tropical forest to warming and hurricane disturbance. In: Dumbrell AJ, Turner EC, Fayle TM, editors. Advances in ecological research. (Academic Press, Cambridge MA, 2020) pp 225–52.
Witt C, Gaunt JL, Galicia CC, Ottow JCG, Neue HU. A rapid chloroform-fumigation extraction method for measuring soil microbial biomass carbon and nitrogen in flooded rice soils. Biol Fertil Soils. 2000;30:510–9.
Berry D, Ben Mahfoudh K, Wagner M, Loy A. Barcoded primers used in multiplex amplicon pyrosequencing bias amplification. Appl Environ Microbiol. 2012;78:612.
Apprill A, McNally S, Parsons R, Weber L. Minor revision to V4 region SSU rRNA 806R gene primer greatly increases detection of SAR11 bacterioplankton. Aquat Micro Ecol. 2015;75:129–37.
Parada AE, Needham DM, Fuhrman JA. Every base matters: assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples. Environ Microbiol. 2016;18:1403–14.
Rohland N, Reich D. Cost-effective, high-throughput DNA sequencing libraries for multiplexed target capture. Genome Res. 2012;22:939–46.
Aronesty E. ea-utils: “Command-line tools for processing biological sequencing data”. 2011. https://github.com/ExpressionAnalysis/ea-utils.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods. 2010;7:335–6.
Caporaso JG, Bittinger K, Bushman FD, Desantis TZ, Andersen GL, Knight R. PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics. 2010;26:266–7.
Edgar RC. Search and clustering orders of magnitude faster than BLAST. Bioinformatics. 2010;26:2460–1.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2013;41:D590–6.
Bokulich NA, Subramanian S, Faith JJ, Gevers D, Gordon JI, Knight R, et al. Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat Methods. 2013;10:57–9.
Morrissey EM, Mau RL, Schwartz E, McHugh TA, Dijkstra P, Koch BJ, et al. Bacterial carbon use plasticity, phylogenetic diversity and the priming of soil organic matter. ISME J. 2017;11:1890–9.
Li J, Mau RL, Dijkstra P, Koch BJ, Schwartz E, Liu X-JA, et al. Predictive genomic traits for bacterial growth in culture versus actual growth in soil. ISME J. 2019;13:2162–72.
Gross N, Bagousse-Pinguet YL, Liancourt P, Berdugo M, Gotelli NJ, Maestre FT. Functional trait diversity maximizes ecosystem multifunctionality. Nat Ecol Evol. 2017;1:0132.
Laliberté E, Norton DA, Scott D. Contrasting effects of productivity and disturbance on plant functional diversity at local and metacommunity scales. J Veg Sci. 2013;24:834–42.
Plass-Johnson JG, Taylor MH, Husain AAA, Teichberg MC, Ferse SCA. Non-random variability in functional composition of coral reef fish communities along an environmental gradient. PLOS ONE. 2016;11:e0154014.
Götzenberger L, Botta-Dukát Z, Lepš J, Pärtel M, Zobel M, de Bello F. Which randomizations detect convergence and divergence in trait-based community assembly? A test of commonly used null models. J Veg Sci. 2016;27:1275–87.
Delgado-Baquerizo M, Trivedi P, Trivedi C, Eldridge DJ, Reich PB, Jeffries TC, et al. Microbial richness and composition independently drive soil multifunctionality. Funct Ecol. 2017;31:2330–43.
R Core Team. R: a language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing; 2020. https://www.R-project.org/.
Bruelheide H, Dengler J, Purschke O, Lenoir J, Jiménez-Alfaro B, Hennekens SM, et al. Global trait–environment relationships of plant communities. Nat Ecol Evol. 2018;2:1906–17.
Piton G, Legay N, Arnoldi C, Lavorel S, Clément J-C, Foulquier A. Using proxies of microbial community-weighted means traits to explain the cascading effect of management intensity, soil and plant traits on ecosystem resilience in mountain grasslands. J Ecol. 2020;108:876–93.
Alster CJ, von Fischer JC, Allison SD, Treseder KK. Embracing a new paradigm for temperature sensitivity of soil microbes. Glob Chang Biol. 2020;26:3221–9.
Letunic I, Bork P. Interactive Tree Of Life (iTOL) v4: recent updates and new developments. Nucleic Acids Res. 2019;47:W256–9.
Li J, Nie M, Pendall E, Reich PB, Pei J, Noh NJ, et al. Biogeographic variation in temperature sensitivity of decomposition in forest soils. Glob Chang Biol. 2020;26:1873–85.
Lipson DA. The complex relationship between microbial growth rate and yield and its implications for ecosystem processes. Front Microbiol. 2015;6:615.
Buckeridge KM, Mason KE, McNamara NP, Ostle N, Puissant J, Goodall T, et al. Environmental and microbial controls on microbial necromass recycling, an important precursor for soil carbon stabilization. Commun Earth Environ. 2020;1:36.
Ali A, Yan E-R, Chang SX, Cheng J-Y, Liu X-Y. Community-weighted mean of leaf traits and divergence of wood traits predict aboveground biomass in secondary subtropical forests. Sci Total Environ. 2017;574:654–62.
Buzzard V, Michaletz ST, Deng Y, He Z, Ning D, Shen L, et al. Continental scale structuring of forest and soil diversity via functional traits. Nat Ecol Evol. 2019;3:1298–308.
Luo Y-H, Cadotte MW, Burgess KS, Liu J, Tan S-L, Zou J-Y, et al. Greater than the sum of the parts: how the species composition in different forest strata influence ecosystem function. Ecol Lett. 2019;22:1449–61.
Díaz S, Lavorel S, de Bello F, Quétier F, Grigulis K, Robson TM. Incorporating plant functional diversity effects in ecosystem service assessments. Proc Natl Acad Sci USA. 2007;104:20684–9.
Bradford MA. Thermal adaptation of decomposer communities in warming soils. Front Microbiol. 2013;4:333.
Morrissey EM, Mau RL, Schwartz E, Koch BJ, Hayer M, Hungate BA. Taxonomic patterns in the nitrogen assimilation of soil prokaryotes. Environ Microbiol. 2018;20:1112–9.
Coskun OK, Ozen V, Wankel SD, Orsi WD. Quantifying population-specific growth in benthic bacterial communities under low oxygen using H218O. ISME J. 2019;13:1546–59.
Zhou G, Zhou X, Liu R, Du Z, Zhou L, Li S, et al. Soil fungi and fine root biomass mediate drought-induced reductions in soil respiration. Funct Ecol. 2020;34:2634–43.
Melillo JM, Frey SD, Deangelis KM, Werner WJ, Bernard MJ, Bowles FP, et al. Long-term pattern and magnitude of soil carbon feedback to the climate system in a warming world. Science. 2017;358:101–5.
Johnston ASA, Sibly RM. The influence of soil communities on the temperature sensitivity of soil respiration. Nat Ecol Evol. 2018;2:1597–602.
Acknowledgements
This work was supported by the U.S. Department of Energy, Program in Genomic Sciences (DE-SC0020172 and DE-SC0016207). Work at LLNL was performed under the auspices of the U.S. Department of Energy under Contract DE-AC52-07NA27344 and Awards SCW1590 and SCW1679. CW acknowledges the funding of Key Research Program of Frontier Sciences, CAS (ZDBS-LY-DQC019), Youth Innovation Promotion Association CAS to CW (2018231), and CAS scholarship.
Author information
Authors and Affiliations
Contributions
Experimental work was conceived by BAH, EMM, ES, KSH, JP-R, and SJB. Samples were collected and experiments performed by RLM, MH, MCM, JCM, SLB, SNM, BJK, BWS, AMP, SJB, KSH, EMM, and JP-R. CW and EMM analyzed the data, with null model analysis by JP and feedback from ES, PD, and BAH. The manuscript was written by CW and EMM with the contributions from all co-authors.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Wang, C., Morrissey, E.M., Mau, R.L. et al. The temperature sensitivity of soil: microbial biodiversity, growth, and carbon mineralization. ISME J 15, 2738–2747 (2021). https://doi.org/10.1038/s41396-021-00959-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-021-00959-1
This article is cited by
-
Emergent temperature sensitivity of soil organic carbon driven by mineral associations
Nature Geoscience (2024)
-
Experimental warming increases respiration and affects microbial communities of soil wetlands at different elevations of the Argentinean Puna
Soil Ecology Letters (2024)
-
Soil bacteria mediate organic carbon stability during alpine wetland biogeomorphic succession in the arid region of Central Asia
Plant and Soil (2024)
-
Viral lysing can alleviate microbial nutrient limitations and accumulate recalcitrant dissolved organic matter components in soil
The ISME Journal (2023)
-
The predictive power of phylogeny on growth rates in soil bacterial communities
ISME Communications (2023)