Abstract
Intracellular symbionts in insects often have reduced genomes. Host acquisition of genes from bacteria is an important adaptation that supports symbionts. However, the function of horizontally transferred genes in insect symbiosis remains largely unclear. The primary symbiont Portiera housed in bacteriocytes lacks pantothenate synthesis genes: panB and panC, which is presumably complemented by a fused gene panB-panC (hereafter panBC) horizontally transferred from bacteria in Bemisia tabaci MEAM1. We found panBC in many laboratory cultures, and species of B. tabaci shares a common evolutionary origin. We demonstrated that complementation with whitefly panBC rescued E. coli pantothenate gene knockout mutants. Portiera elimination decreased the pantothenate level and PanBC abundance in bacteriocytes, and reduced whitefly survival and fecundity. Silencing PanBC decreased the Portiera titer, reduced the pantothenate level, and decreased whitefly survival and fecundity. Supplementation with pantothenate restored the symbiont titer, PanBC level, and fitness of RNAi whiteflies. These data suggest that pantothenate synthesis requires cooperation and coordination of whitefly PanBC expression and Portiera. This host–symbiont co-regulation was mediated by the pantothenate level. Our findings demonstrated that pantothenate production, by the cooperation of a horizontally acquired, fused bacteria gene and Portiera, facilitates the coordination of whitefly and symbiont fitness. Thus, this study extends our understanding on the basis of complex host–symbiont interactions.
Similar content being viewed by others
Introduction
Intracellular symbionts, which include primary and secondary symbionts, are widespread in insects [1, 2]. Primary symbionts can provide specific nutrients that are deficient in the diet and which the host insect cannot synthesize [3,4,5,6,7,8]. Secondary symbionts may affect insect fitness under certain conditions [2]. Intracellular symbionts, particularly those restricted to specialized host cells (bacteriocytes), have reduced genomes [1]. Horizontal gene transfer is an important host adaptation that supports and controls insect symbionts [3, 8,9,10]. Horizontally transferred genes (HTGs) may complement the missing genes involved in synthesis of metabolites by the symbiont [1]. Revealing how genomes from divergent lineages form a functional biosynthetic pathway will help unravel the processes underlying the origin of eukaryotic cells [1, 11].
The important role of HTGs has been increasingly appreciated in insect symbiosis. For example, horizontally transferred RlpA4 encodes a protein that is transported to the symbiont Buchnera [12]; silencing horizontally transferred amiD and ldcA1 decreases the abundance of Buchnera in aphids [13]. Horizontally transferred MurF encodes a protein that is transported to the symbiont Moranella in mealybugs for peptidoglycan synthesis [11]. B vitamins perform various important functions in animals [14]. Animals cannot synthesize these compounds, so they must acquire them from their diet or the microbiome [14]. However, the plant sap diet of Hemiptera is deficient in B vitamins. Horizontally transferred whitefly biotin genes with a bacterial origin can synthesize biotin in the whitefly [15]. Because functional validation for most HTGs is lacking, the role of HTGs in insect–symbiont interactions and coevolution remains unclear.
Pantothenate is an indispensable B vitamin in animals [14]. Available studies indicate the importance of pantothenate synthesis in insect symbiosis systems [3, 16,17,18,19]. The genomes of the symbiont Buchnera in the aphid Acyrthosiphon pisum [14, 16, 17] and Moranella in the mealybug Planococcus citri have panB and panC. However, the genomes of the aphid and mealybug lack these genes [3, 20]. In contrast, the genomes of the primary symbiont Portiera in Bemisia tabaci MEAM1 and Carsonella in the psyllids Ctenarytaina eucalypti and Heteropsylla cubana lack panB and panC [18, 19]. These genes are present in the genomes of secondary symbionts of the psyllids C. eucalypti and H. cubana [19]. Whether the genomes of C. eucalypti and H. cubana have panB and panC or not is unknown due to unavailable genome data. But these genes are not present in the genome of the related psyllid species Pachypsylla venusta [9]. Fused gene panB-panC (hereafter called panBC) horizontally transferred from bacteria occurs in B. tabaci MEAM1 [18]. These data show the prevalence of panB and panC in the bacterial symbiosis of sternorrhynchan insects.
The whitefly B. tabaci is a complex of more than 35 cryptic species [21,22,23]. All B. tabaci species harbor the primary symbiont “Candidatus Portiera aleyrodidarum” (hereafter Portiera) in bacteriocytes and may harbor up to four secondary symbiont lineages [24, 25]. Within the B. tabaci species complex, B. tabaci MEAM1 is a globally important and invasive agricultural pest [26]. In China, B. tabaci MEAM1 bears Portiera and “Candidatus Hamiltonella defensa” (hereafter Hamiltonella) in the same bacteriocyte and Rickettsia sp. (hereafter Rickettsia) in the whole body cavity [15, 27, 28]. The genome of Portiera is highly reduced and mainly contains genes involved in synthesis of essential amino acids [29]. Portiera also retains genes, except for panB and panC, involved in pantothenate synthesis. Horizontally transferred panBC encoded in the genome of B. tabaci MEAM1 is presumably involved in pantothenate synthesis [14, 18]. In contrast, Hamiltonella and Rickettisa have totally lost the capability for pantothenate synthesis [18], and they have pantothenate symporters that transport pantothenate from the bacteriocytes where the whitefly and Portiera cooperate to synthesize this metabolite. However, the manner in which panBC contributes to whitefly–symbiont interactions and coevolution is unclear. In this study, the function of the horizontally transferred panBC gene in the B. tabaci MEAM1 symbiosis system was studied. We reveal that pantothenate provisioned by whitefly panBC facilitates the regulation of whitefly symbiosis with Portiera.
Materials and methods
Insect rearing and plants
The whitefly B. tabaci MEAM1 colony (mtCO1 GenBank accession no. GQ332577) was maintained on cotton plants (Gossypium hirsutum, cv. Shiyuan 321) grown in compost supplemented with Miracle-Gro Water Soluble All Purpose Plant Food as described previously [15, 30, 31]. Cotton plants were cultivated to the 6–7 true-leaf stage for use in experiments. For more details, see the Supplementary Text.
Quantitative PCR (qPCR) and RT-PCR
DNA and RNA were extracted following the protocol previously described [15, 32]. Symbionts were quantified by qPCR using the CFX96 Real-Time PCR Detection System (Bio-Rad, Hercules, CA, USA) with 2×SYBR Green Master Mix (Bio-Rad). All primers used in this study are shown in Supplementary Table 1. Relative symbiont density was calculated using the 2−ΔCt method [33]. RNAs were first strand reverse transcribed using a kit (Bimake, Houston, TX, USA) as described previously [15]. For more details, see the Supplementary Text.
Amino acid sequence alignment and phylogenetic tree analysis
To determine the homologous genes in other B. tabaci species, verified sequences of panBC of the whitefly B. tabaci MEAM1 were subjected to TBLASTX against the genome of B. tabaci MED [34] and SSA-ECA (GenBank accession No.: GCA_004919745.1) and transcriptome of B. tabaci MEAM [10], MED [35], and Asia II 3 [36]. Amino acid sequence alignments were conducted using BioEdit v7.1.3.0 among the three B. tabaci species, B. aphidicola, and E. coli. To construct the molecular phylogenetic tree, a Bayesian inference (BI) analysis was conducted as described previously [10, 15]. For more details, see the Supplementary Text.
Fluorescence in situ hybridization (FISH)
Localization of Portiera in bacteriocytes of female adult whiteflies was investigated by FISH following a previously described protocol [15, 30, 37]. For more details, see the Supplementary Text.
Recombinant enzyme generation and antibody preparation
The recombinant enzyme for whitefly panBC was generated, and custom-made polyclonal antibodies against PanBC (predicted size, 68KD) protein were produced by ProbeGene Life Sciences Co. Ltd. following previously described methods [12, 15, 38]. For more details, see the Supplementary Text.
Immunofluorescence microscopy
Bacteriocytes from Portiera-infected and Portiera-cured adult female whiteflies as well as dsRNA-injected adult female whiteflies were dissected, fixed, permeabilized, and incubated with antibodies. Images were collected and analyzed on a FV3000 confocal microscope (Olympus, Japan). For more details, see the Supplementary Text.
Pantothenate measurement
A microbiological assay was used for pantothenate quantification in whiteflies using Lactobacillus plantarum ATCC 8014 (Beijing Landbridge Technology Limited, Beijing, China) referring to the protocol of [15, 39]. Briefly, 50 female adult whiteflies from each treatment were collected for each of three replicates using a mouth aspirator and flash-frozen in liquid N2. Insects from each treatment were pooled, weighed, and homogenized in citrate buffer using homogenizer MP FastPrep-24 (MP Biomedicals LLC, Santa Ana, CA, USA) and incubated at 95 °C for 30 min. Then samples were sterilized using a 0.2-μm pore size filter (PALL Life Science, Port Washington, NY, USA), mixed with vitamin B5-deficient pantothenate assay medium (hereafter called as B5DP; Beijing Landbridge Technology Limited, Beijing, China), and inoculated with log-phase L. plantarum ATCC 8014. Cultures were incubated for 20 h at 37 °C. Standard concentrations of pantothenate (0–80 ng/mL; Dr. Ehrenstorfer Gmbh, Augsburg, Germany) were mixed with the L. plantarum culture to create a standard curve. A negative control consisting of citrate buffer only was subjected to the complete procedure to ensure the retention of the initial pantothenate and a lack of additional pantothenate. An additional negative control included only the assay medium lacking L. plantarum to exclude contamination by other bacteria. The growth of L. plantarum was measured using a microplate reader (Versa Max Molecular Devices, San Jose, CA, USA) taking absorbance readings at 630 nm. Pantothenate in whiteflies was quantified using the standard curve and normalized to the weight of insects used in the homogenate.
Functional complementation of E. coli pantothenate auxotrophs with whitefly panBC
To examine the metabolic function of horizontally transferred panBC, E. coli pantothenate gene (panB and panC) knockout mutants were generated and functional complementation with whitefly panBC was performed. The kanamycin resistance site was amplified from E. coli G11 using knockout primers (Supplementary Table 1). Then pKD46 plasmids expressing Lambda Red recombinase were transformed into the E. coli K-12 BW25113 by electroporation. Subsequently, the E. coli K-12 BW25113 panB and panC knockout mutants (i.e., −ΔpanB and −ΔpanC) were generated following the Lambda Red protocol as previously described [40,41,42]. E. coli mutants were screened using a LB media agar plate with 0.1-mg/mL kanamycin. PCR amplification was carried out using primers specific for deleted genes, the kanamycin resistance site, and the flanking locus to verify the loss of the parental fragment and gain of the kanamycin resistance site (Supplementary Table 1). The E. coli mutants were then made into competent cells by electroporation. The whole CDS region of panBC was amplified from panBC-PUC57, as described above, and was ligated to the pMD19-T vector. For the functional complement experiments, E. coli K-12 mutant cells were transformed with the plasmids pMD19-T-panBC and pMD19-T empty vector (negative control), respectively, following previous methods [15, 16]. The E. coli wild-type K-12, mutant K-12 and mutant K-12 transformants were grown overnight in LB agar media with 0.1 mg/mL ampicillin at 37 °C. The mutant K-12 transformants were verified by PCR amplification using primers specific for inserted genes. Then E. coli wild-type K-12, mutant K-12, and mutant K-12 transformants were grown overnight in pantothenate-deficient B5DP medium (Beijing Landbridge Technology Limited, Beijing, China) at 37 °C. All E. coli cells were washed twice in sterile distilled water and resuspended to measure the cell density at OD600 using a microplate reader (Versa Max Molecular Devices, San Jose, CA, USA).
Effects of Portiera elimination by antibiotic treatment on PanBC level, symbiont localization, pantothenate level, and whitefly performance
Previous studies have demonstrated that treatment with antibiotic rifampicin at 30 μg/ml for 2 days was effective in curing Portiera, Hamiltonella, and Rickettsia in the F1 B. tabaci [43, 44]. Thus, to eliminate Portiera, hundreds of adult whiteflies of B. tabaci (F0, 0–7 days after emergence) were released into each feeding chamber and fed on 25% sucrose solution (w/v) supplemented with the antibiotic rifampicin (BBI Life Sciences, Shanghai, China) dissolved in 5-mM phosphate buffer (pH 7.0), at 30 μg/ml for 2 days as described previously [43, 44]. Control whiteflies were allowed to feed on sucrose solution not supplemented with antibiotics. Following the antibiotic treatment, B. tabaci were transferred to cotton plants. F1 female and male adults were collected. The DNA was extracted from eight female adults of B. tabaci (at 3–7 days after eclosion) and used for symbiont quantification by qPCR. The F1 B. tabaci with reduced Portiera titer (−PBt), which were obtained by antibiotic treatment, and control F1 B. tabaci (+PBt), which were obtained by feeding sucrose solution not supplemented with antibiotics, were identified. To test whether Portiera elimination affects the PanBC level and symbont localization in bacteriocytes, bacteriocytes of female adults of B. tabaci (at 3–7 days after eclosion) were dissected, fixed, permeabilized, and incubated with antibodies and a fluorescent probe for Portiera as described above. Three biological replicates were conducted. Images were collected and analyzed on a FV3000 confocal microscope (Olympus, Japan). In parallel, 50 female adult whiteflies (at 3–7 days after eclosion) in each of three biological replicates were collected for pantothenate analysis as described above.
To determine if Portiera elimination by antibiotic treatment influences whitefly performance, 20 female adult whiteflies within 3 days post emergence per replicate of +PBt and −PBt lines were transferred onto a cotton leaf disk kept on a 1.5% agar plate at 26 ± 2 °C, with 14:10 h (L:D) photoperiod and 60–80% RH. Four biological replicates were conducted. The mortality of injected female whiteflies was recorded at days 3 and 6. In addition, one pair of female and male adult whiteflies at 2 h post emergence from +PBt and −PBt lines were released onto one cotton leaf disk kept on 1.5% agar plate, and allowed to lay eggs for 6 days. Ten biological replicates were conducted. After 6 days, eggs were recorded.
dsRNA preparation
dsRNAs specific to whitefly panBC (dspanBC) and GFP (dsGFP) were synthesized using a T7 RiboMAX™ Express RNAi System kit (Promega, USA), following the manufacturer’s instructions. For more details, see the Supplementary Text.
Effects of silencing panBC on PanBC level, symbiont titer, pantothenate levels, and whitefly performance
To investigate whether silencing of horizontally transferred panBC gene influences PanBC level, symbiont titer, and pantothenate levels, ~3000 female adult whiteflies infected with Portiera within 4–6 days after emergence were injected with 1.5-μg/μL dspanBC in injection buffer using an Eppendorf microinjection system (Hamburg, Germany) referring to a previously described method with some modifications [45]. Control whiteflies were injected with dsGFP. The average injection volume used was 10 nl. After injection, whiteflies were transferred onto cotton leaf disks kept on 1.5% agar plates in the incubator at 26 ± 2 °C, with a 14:10 h (L:D) photoperiod and 60–80% RH. The survival rate of injected whiteflies was 85–90% at 24 h after injection, showing that the microinjection protocol had little physical effect on the whiteflies. To examine whether silencing whitefly panBC affects the PanBC level and symbiont localization in bacteriocytes, whiteflies were collected at days 1, 3, 5, and 7, and the whitefly bacteriocytes were dissected, fixed, permeabilized, and incubated with antibodies and a fluorescent probe for Portiera as described above. Three biological replicates were conducted. Images were analyzed on a FV3000 confocal microscope (Olympus, Japan). To test whether silencing whitefly panBC influences symbiont abundance, DNA was extracted from individual female adult whiteflies for each of ten biological replicates at day 3 after whiteflies were microinjected with dspanBC. Then, qPCR was performed as described above. In parallel, 50 female adult whiteflies in each of three biological replicates were collected for pantothenate analysis as described above.
To determine if panBC silencing influences whitefly performance, ~700 female adult whiteflies infected with Portiera within 4 days after emergence were injected using the microinjection procedures described above. After injection, 30 female adults per replicate of panBC-injected and dsGFP-injected whiteflies were transferred onto a cotton leaf disk kept on a 1.5% agar plate at 26 ± 2 °C, with 14:10 h (L:D) photoperiod and 60–80% RH. Five biological replicates were conducted. The mortality of injected female whiteflies was recorded at days 1, 3, 5, and 7. In addition, after injection, individual whiteflies were transferred onto cotton leaf disks as described above. Egg numbers were recorded for living whiteflies with 13 biological replicates of individuals at day 7 post injection.
Effects of pantothenate supplementation on symbiont localization, PanBC level, and whitefly performance
To investigate whether pantothenate supplementation restores the symbiont localization and PanBC level of dsRNA-injected whiteflies, ~500 female adult whiteflies within 4–6 days after emergence were microinjected with dspanBC and dsGFP. After recovery on a cotton leaf disk for 12 h, these whiteflies were fed 30% (w/v) sucrose solution supplemented with or without pantothenate at a final concentration of 250 ng/mL for 2 days. The controls were dsGFP-injected whiteflies fed with 30% (w/v) sucrose solution. Then, bacteriocytes were dissected for localization of Portiera in bacteriocytes of female adult whiteflies by FISH following the protocol as described above. In addition, bacteriocytes were dissected for examination of PanBC level by immunofluorescence microscopy as described above. Three biological replicates were conducted.
To examine whether pantothenate supplementation restores the survival of dsRNA-injected whiteflies, ~800 female adult whiteflies within 4–6 days after emergence were microinjected with dspanBC and dsGFP. After recovery on a cotton leaf disk for 12 h, 30 female adults per replicate of panBC-injected and dsGFP-injected whiteflies were fed 30% (w/v) sucrose solution supplemented with or without pantothenate at a final concentration of 250 ng/mL for 2 days. Then the mortality of injected female whiteflies was recorded. Six biological replicates were conducted.
To examine whether pantothenate supplementation restores the fecundity of dsRNA-injected whiteflies, dsRNA-injected whiteflies were fed 30% (w/v) sucrose solution supplemented with or without pantothenate at a final concentration of 250 ng/mL for 2 days as described above. Then, individual female adult whiteflies were released on cotton leaf disks. After 3 days, egg numbers were recorded. Ten replicates were conducted for each treatment.
Statistical analyses
For the OD values of the E. coli wild-type K-12, mutant K-12 and mutant K-12 transformants, statistical differences were evaluated using one-way ANOVA at a significance level of 0.05 followed by LSD post-hoc tests. For the symbiont titer, pantothenate amount, and mortality and egg numbers of dsGFP, and dspanBC-injected female whiteflies, statistical differences were evaluated using one-way ANOVA at a significance level of 0.05. Percentage data were transformed by arcsine square root before analysis. All data analyses were conducted using the STATISTICA v6.1 software (StatSoft, Inc., Tulsa, OK, USA).
Results
Prevalence of panBC in the bacterial symbiosis of whiteflies
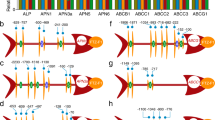
To investigate whether panBC are ubiquitous in B. tabaci, the presence of panBC was determined in several B. tabaci species. We found that panBC is present in seven lab cultures of four B. tabaci species, which are distributed in Asia, America, and Africa (Supplementary Table 2). The B. tabaci panBC has five introns and five CDSs with two enzymatic functions [18] (Fig. 1 and Supplementary Fig. 1a). Thus, the domains corresponding to the bacterial PanB and PanC proteins as shown in Fig. 1 were analyzed separately for sequence alignment and phylogenetic tree construction. To examine the divergence of protein sequences, amino acid sequences were aligned among three whitefly species, B. aphidicola, and E. coli for PanB and PanC. The amino acid sequence identity was high among all B. tabaci species (88.1% for PanB and 93.81% for PanC, respectively) but low among B. tabaci, B. aphidicola, and E. coli (68.91% for PanB and 72.36% for PanC, respectively) (Supplementary Fig. 1b, c). To gain insight into the evolution of B. tabaci PanBC, a phylogenetic tree was constructed. Interestingly, PanBC of all whitefly species clustered within the genus Pseudomonas (Supplementary Fig. 2). These data suggest that panBC has a common evolutionary origin in B. tabaci.
Gene structure of panBC was adapted from ref. [18] and the whitefly genome database (http://www.whiteflygenomics.org/cgi-bin/bta/geneinfo.cgi?&gene=Bta05339). The conserved domain presented is based on BLASTP results.
Functional complementation of E. coli pantothenate auxotrophs with whitefly panBC
To test the hypothesis that whitefly panBC functions in pantothenate synthesis, E. coli K-12 panB and panC knockout mutants (−ΔpanB or −ΔpanC) were generated using the Lambda Red protocol and functionally complemented E. coli K-12 mutant with whitefly panBC, respectively. Compared to wild-type E. coli, E. coli K-12 knockout mutants (−ΔpanB and −ΔpanC) did not grow on B5DP medium lacking pantothenate (Fig. 2). Although whitefly PanBC shared low amino acid sequence identities with E. coli homolog genes (41% and 42.57% for PanB and PanC, respectively) (Supplementary Fig. 1b, c), complementation with whitefly panBC rescued E. coli K-12 knockout mutants on B5DP medium (Fig. 2). In contrast, cells transformed with the pMD19-T empty vector did not grow on B5DP medium without pantothenate supplementation (Fig. 2). Significant differences in OD values among treatments were detected (Fig. 2; F3,8 = 51.34, P < 0.0001 for panB and F3,8 = 50.09, P < 0.0001 for panC).
E. coli K-12 knockout mutant cells were transformed with expression plasmids containing whitefly panBC or the negative control pMD19-T empty vector. The E. coli wild-type K-12, mutant K-12 (−Δ), and mutant K-12 transformants (+Δ) were grown overnight in pantothenate-deficient B5DP medium at 37 °C. All E. coli cells were washed and re-suspended to measure cell density at OD600. Data are means ± SE. Different letters above the bars indicate significant differences between treatments at P < 0.05.
Portiera elimination reduced pantothenate level, PanBC localization, and whitefly performance
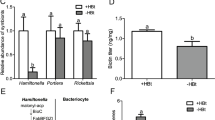
To investigate whether Portiera contributes to the generation of pantothenate in whiteflies, Portiera were cured by antibiotic treatments. The Portiera titer was reduced by 97% and the titer of Hamiltonella and Rickettisa was reduced by 95–97%, by rifampicin treatment (Fig. 3A) (F1,14 = 68.1, P < 0.0001 for Portiera; F1,14 = 43.77, P < 0.0001 for Hamiltonella and F1,14 = 12.06, P = 0.0037 for Rickettisa). The pantothenate level was decreased significantly in Portiera-cured (−PBt) compared to Portiera-infected (+PBt) whiteflies (Fig. 3B; F1,4 = 14.37, P = 0.019).
A Effects of antibiotic treatments on the abundance of symbionts in B. tabaci. B Effects of Portiera elimination on pantothenate levels in B. tabaci. +PBt and −PBt represent Portiera-infected and Portiera-cured whiteflies, respectively. Localization of Portiera (red) and PanBC (green) in the bacteriocytes of +PBt (C) and −PBt (D) female adult whiteflies. DNA was stained with DAPI. E Mortality of +PBt and −PBt female adult whiteflies after feeding on cotton leaf disks for 3 days. F Fecundity of +PBt and −PBt female adult whiteflies after feeding on cotton leaf disks for 6 days. Data are means ± SE. The significant differences between treatments are indicated by asterisks in A, B, and F (*P < 0.05; **P < 0.01; ***P < 0.001).
To test whether Portiera influences expression of PanBC, we examined the subcellular location of the whitefly PanBC protein in bacteriocytes. The polyclonal antibody against PanBC protein was produced using the purified recombinant protein (Supplementary Fig. 3A–C). Immunofluorescence microscopy showed that PanBC has a high expression level in whitefly bacteriocytes. PanBC was located in the cytoplasmic region of bacteriocytes and had some clusters (Fig. 3C). Confocal microscopy observation confirmed that Portiera was completely depleted in bacteriocytes after antibiotic treatments (Fig. 3D). After Portiera was cured, the PanBC protein level decreased significantly and the PanBC protein localization pattern was altered with less protein clusters in whiteflies (Fig. 3D). These data suggest that the abundance of Portiera impacts the pantothenate level and the localization of whitefly PanBC.
To test whether Portiera affects whitefly performance, we examined whitefly survival and fecundity after Portiera was cured. Portiera elimination decreased the survival of female adult whiteflies (Fig. 4E; F1,6 = 4.21, P = 0.086 for day 3 and F1,6 = 3.0, P = 0.13 for day 6) and significantly reduced the whitefly fecundity (Fig. 4F; F1,18 = 43.94, P < 0.0001).
A Localization of PanBC (green) and Portiera (red) in bacteriocytes of dsGFP-injected and dspanBC-injected female adult whiteflies at 1, 3, 5, and 7 days after whiteflies were microinjected with dsRNA. DNA was stained with DAPI. B The symbiont titer in whiteflies at day 3 after whiteflies were microinjected with dspanBC. C Pantothenate levels in whiteflies at 3 and 5 days after whiteflies were microinjected with dspanBC. D Female mortality at 1, 3, 5, and 7 days after whiteflies was microinjected with dspanBC. E Female fecundity at day 7 after whiteflies was microinjected with dspanBC. Data are means ± SE. The significant differences between treatments are indicated by asterisks in B, C, and E (*P < 0.05).
Silencing panBC reduces symbiont titer, pantothenate level, and whitefly performance
To study the metabolic function of horizontally transferred panBC, whitefly panBC was silenced by microinjection with dspanBC. The level of PanBC was significantly reduced in bacteriocytes at day 1, 3, and 5 after RNAi treatment but recovered at day 7 after microinjection with dspanBC (Fig. 4A). In parallel with the changes in PanBC level, the titer of Portiera also decreased in bacteriocytes at day 1, 3, and 5 but recovered at day 7 after RNAi treatment (Fig. 4A), as shown by FISH observation. panBC gene silencing in whitefly bacteriocytes may not persist for more than 7 days after microinjection with dsRNA, which could lead to the recovery of PanCB and Portiera levels at day 7. A qPCR test also showed that the titer of Portiera was significantly decreased at day 3 after RNAi treatment (Fig. 4B; F1,18 = 4.6, P = 0.046). In contrast, the titer of Hamiltonella remained unchanged at day 3 after RNAi treatment (Fig. 4B; F1,18 = 0.04, P = 0.85). As a result of gene silencing, the pantothenate level was decreased significantly in dspanBC-injected compared to dsGFP-injected whiteflies (Fig. 4C; F1,4 = 12.45–13.42, P = 0.022–0.024). In addition, silencing the horizontally transferred panBC gene significantly increased the mortality of female adult whiteflies at days 3–7 after microinjection with dsRNAs (Fig. 4D; F1,8 = 10.62–28.39, P = 0.0007–0.012) and reduced female fecundity at day 7 after microinjection with dsRNAs (Fig. 4E; F1,24 = 4.67, P = 0.041).
Supplementation with pantothenate restores symbiont titer, PanBC level, and fitness of RNAi whiteflies
To examine whether B5 pantothenate supplementation can alter the titer of Portiera, PanBC level, and fitness of dsRNA-injected whiteflies, dsRNA-injected whiteflies were fed an artificial diet supplemented with pantothenate for 2 days. The titer of Portiera increased in bacteriocytes of dspanBC-injected whiteflies supplemented with pantothenate, which was similar to dsGFP-injected whiteflies (Fig. 5A). The level of PanBC also recovered in bacteriocytes of RNAi whiteflies after pantothenate supplementation treatment (Fig. 5B). The mortality of dspanBC-injected whiteflies was increased significantly compared to dsGFP-injected whiteflies; after pantothenate supplementation, the mortality of dspanBC-injected whiteflies was decreased, which is close to that of dsGFP-injected whiteflies, while the mortality of dsGFP-injected whiteflies supplemented with pantothenate decreased but not significantly compared to dsGFP-injected whiteflies without pantothenate supplementation (Fig. 5C; F5,18 = 102.15, P < 0.0001). The fecundity of dspanBC-injected whiteflies was decreased significantly compared to dsGFP-injected whiteflies; after pantothenate supplementation, the fecundity of dspanBC-injected whiteflies was restored to a level comparable to that of dsGFP-injected whiteflies, while the fecundity of dsGFP-injected whiteflies supplemented with pantothenate increased but not significantly compared to dsGFP-injected whiteflies without pantothenate supplementation (Fig. 5D; F3,36 = 7.72, P = 0.00042).
A Localization of Portiera (red) in bacteriocytes of dsGFP-injected and dspanBC-injected female adult whiteflies feeding on the artificial diet supplemented with or without B5 pantothenate for 2 days. DNA was stained with DAPI. B Localization of PanBC (green) in bacteriocytes of dsGFP-injected and dspanBC-injected female adult whiteflies feeding on the artificial diet supplemented with or without B5 pantothenate for 2 days. DNA was stained with DAPI. C Mortality of dsGFP-injected and dspanBC-injected female adult whiteflies after feeding on artificial diet supplemented with B5 pantothenate for 2 days. D Fecundity of dsGFP-injected and dspanBC-injected female adult whiteflies after feeding on artificial diet supplemented with B5 pantothenate for 2 days and then feeding on cotton leaf disk for 3 days. Data are means ± SE. Different letters above the bars indicate significant differences between treatments at P < 0.05.
Discussion
Host adaptations are critical in enabling extreme gene loss of symbionts [1]. Insect genes can compensate for missing genes in symbionts [1, 3, 9, 10, 46, 47]. However, the regulation of insect bacterial symbiosis is not well known. We found co-regulation of whitefly fitness (PanBC protein level and whitefly performance) and symbiont titer. This co-regulation was mediated by the pantothenate level. Therefore, we revealed that a fused gene of bacterial origin can cooperate with Portiera to synthesize pantothenate. This collaboration facilitates the regulation of whitefly and symbiont fitness. Our findings demonstrate the key role of the horizontally transferred panBC gene in whitefly symbiosis.
The host–symbiont relationship is complex and dynamic [48]. System-level in silico analysis suggests that the biosynthesis of essential amino acids by Buchnera and Portiera is exclusively controlled by the aphid and whitefly host [49, 50]. The relative uniformity in the location and numbers of resident microbes also indicates that the insect host exerts tight control over its symbionts [51]. Aphids use the amino acid transporter to regulate the precursor glutamine supply to the bacteriocyte, thereby regulating amino acid biosynthesis in Buchnera [52]. Although the aphid seems to play the key role in regulating the metabolic output of bacteriocytes, Buchnera can regulate its gene expression by small RNAs [53, 54]. The host and symbiont gene expression between aphids and Buchnera appears to be coordinated [48]. We found co-regulation of whitefly PanBC protein abundance, whitefly performance, and symbiont titer. Portiera elimination or panBC silencing reduced the pantothenate level. Pantothenate is a vitamin (i.e. coenzyme) and the precursor of coenzyme A, which is required for the function of key enzymes in central carbon metabolism, especially involved in the TCA cycle and fatty acid metabolism [14]. The reduced pantothenate level likely inhibits the generation of lipids or other metabolites, which may repress protein synthesis, whitefly fecundity and survival, and symbiont proliferation. In contrast, the additional pantothenate allows Porteria to grow to high enough levels so that it may stimulate production of PanBC or directly facilitate PanBC synthesis and whitefly performance, regardless of RNAi treatment. Therefore, whitefly fitness (PanBC protein level and whitefly performance) and Portiera abundance can be co-regulated by the pantothenate level. This is supported by evidence that dietary supplementation with pantothenate restored Portiera abundance and whitefly PanBC protein level and whitefly performance. The coordination of whitefly and symbiont fitness reflects the coadaptation and coevolution between host and its symbiont. The pantothenate-mediated coordination of whitefly and symbiont fitness suggests that the host and symbiont interactions are also modulated by nutrition status. These findings explain why whitefly fitness and symbiont abundance can vary among whitefly lineages. Furthermore, this study proposes that the regulation of metabolite synthesis in insect symbiosis may depend on the enzyme type, metabolite function, metabolite synthesis pathway, and the species involved. It extends our understanding the basis of complex host–symbiont interactions.
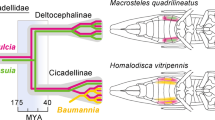
All of the four representative sternorrhynchan insect–bacterial symbiosis systems have distinct pantothenate synthesis pathways. The primary smbionts Buchnera of the aphid and Moranella of the mealybug retain the intact pantothenate synthesis pathway [3, 14, 16]. In contrast, the primary symbiont Carsonella of psyllids lacks panBC, which can be complemented by secondary symbionts in psyllids [9, 19]. The primary symbiont Portiera of B. tabaci only retains incomplete synthesis pathways for a few B vitamins including pantothenate. A fused gene panBC horizontally transferred from bacteria in whiteflies can complement for the missing gene in Portiera [18]. This result indicates the importance of pantothenate synthesis in insect symbiosis systems. All seven lab cultures of the four B. tabaci species harbored horizontally transferred panBC with a common evolutionary origin of Pseudomonas, which is absent in the B. tabaci species complex. Therefore, panB and panC were likely co-transferred to the common ancestor of B. tabaci and then later fused into the gene panBC. We demonstrated that whiteflies are able to synthesize pantothenate via horizontally transferred panBC. Because pantothenate is essential for whitefly fitness and panBC is widespread in B. tabaci, these results show that panBC contributes to the ability of whiteflies to feed on plant phloem deficient in B vitamins. Our findings suggest that pantothenate provisioning in animal symbiosis evolutionarily transitioned from bacterial symbionts to animal hosts through horizontal gene transfer events.
Rifampicin treatment leds to elimination of Portiera, Hamiltonella, and Rickettisa in B. tabaci MEAM1. Because Hamiltonella and Rickettisa have no pantothenate synthesis pathway [18], the reduced titer of Hamiltonella and Rickettisa could not have effects on pantothenate levels in B. tabaci MEAM1. B. tabaci MED harbors Portiera and Hamiltonella. After Portiera and Hamiltonella were cured in B. tabaci MED by rifampicin treatment, the survival rate and fecundity were also significantly reduced as that in Portiera, Hamiltonella, and Rickettisa-depleted B. tabaci MEAM1 [43, 44]. So Rickettisa elimination may play a minor role in the altered mortality rate and fecundity of symbionts-depleted B. tabaci MEAM1. Furthermore, our previous studies demonstrated that Hamiltonella elimination only influenced the sex ratio and didn’t affect mortality rate and fecundity of B. tabaci MEAM1 [15, 28]. Thus, it is an elimination of Portiera but not other symbionts that impacts the mortality rate and fecundity in B. tabaci MEAM1. Overall, Portiera elimination reduced the pantothenate level, thereby repressing the survival rate and fecundity of B. tabaci MEAM1.
PanBC has a high expression level in whitefly bacteriocytes. Both PanBC and Portiera cells were located in the cytoplasmic region of bacteriocytes. The high expression level and bacteriocyte localization of whitefly PanBC facilitates its cooperation with Portiera to synthesize pantothenate in B. tabaci MEAM1. In this case, pantothenate is produced in the cytoplasmic region of bacteriocytes, which may benefit its utilization by Portiera cells. Therefore, Portiera does not need the pantothenate symporter, which helps explain why it is lacking. In contrast, Hamiltonella and Rickettisa have totally lost the capability for pantothenate synthesis. However, they have pantothenate symporters to transport pantothenate into the cells of Hamiltonella or Rickettisa from the bacteriocytes where the whitefly and Portiera cooperate to synthesize this metabolite. There is lower titer of Hamiltonella in whiteflies as compared to Portiera [15, 27, 30], and changes of pantothenate levels may not have a large effect on the titer of Hamiltonella over short time periods. So we did not observe changes of pantothenate levels influencing the titer of Hamiltonella over 3 days after silencing panBC.
Acquisition of HTGs in the insect genome confers novel genetic information. Intron gain of HTGs is a critical step for them to attain functionality [15, 55]. In other species, panB and panC are separately present in the genome. However, the B. tabaci panBC has become a single gene and acquired introns [18]. This reveals that another way for HTGs to become functional is intron gain and further or simultaneous gene fusion or gene fusion and further intron gain. The PanB and PanC domains of whitefly PanBC mediate the proximal and final reactions in pantothenate synthesis [18]. Fused panBC in B. tabaci could make the chemical reaction more efficient during pantothenate production. In addition, the case of PanBC helps us to better understand the mechanisms underlying the formation of multiple domains of proteins.
Data availability
All relevant data supporting the findings of this study are included within the article and its Supplementary Information files.
References
Moran NA, Bennett GM. The tiniest tiny genomes. Annu Rev Microbiol. 2014;68:195–215.
Douglas AE. Multiorganismal insects: diversity and function of resident microorganisms. Annu Rev Entomol. 2015;60:17–34.
Husnik F, Nikoh N, Koga R, Ross L, Duncan RP, Fujie M, et al. Horizontal gene transfer from diverse bacteria to an insect genome enables a tripartite nested mealybug symbiosis. Cell. 2013;153:1567–78.
Nikoh N, Hosokawa T, Moriyama M, Oshima K, Hattori M, Fukatsu T. Evolutionary origin of insect-Wolbachia nutritional mutualism. Proc Natl Acad Sci USA. 2014;111:10257–62.
Salem BauerE, Kirsch R, Berasategui A, Cripps M, Weiss B, et al. Drastic genome reduction in an herbivore’s pectinolytic symbiont. Cell. 2017;171:1520–31.
Moran NA, McCutcheon JP, Nakabachi A. Genomics and evolution of heritable bacterial symbionts. Annu Rev Genet. 2008;42:165–90.
Moya A, Peretó J, Gil R, Latorre A. Learning how to live together: genomic insights into prokaryote-animal symbioses. Nat Rev Genet. 2008;9:218–29.
Wybouw N, Pauchet Y, Heckel DG, Van, Leeuwen T. Horizontal gene transfer contributes to the evolution of arthropod herbivory. Genome Biol Evol. 2016;8:1785–801.
Sloan DB, Nakabachi A, Richards S, Qu J, Murali SC, Gibbs RA, et al. Parallel histories of horizontal gene transfer facilitated extreme reduction of endosymbiont genomes in sap-feeding insects. Mol Biol Evol. 2014;31:857–71.
Luan JB, Chen WB, Hasegawa DK, Simmons AM, Wintermantel WM, Ling KS, et al. Metabolic coevolution in the bacterial symbiosis of whiteflies and related plant sap-feeding insects. Genome Biol Evol. 2015;7:2635–47.
Bublitz DC, Chadwick GL, Magyar JS, Sandoz KM, Brooks DM, Mesnage S, et al. Peptidoglycan production by an insect-bacterial mosaic. Cell. 2019;179:703–12.
Nakabachi A, Ishida K, Hongoh Y, Ohkuma M, Miyagishima SY. Aphid gene of bacterial origin encodes a protein transported to an obligate endosymbiont. Curr Biol. 2014;24:640–1.
Chung SH, Jing X, Luo Y, Douglas AE. Targeting symbiosis-related insect genes by RNAi in the pea aphid-Buchnera symbiosis. Insect Biochem Mol Biol. 2018;95:55–63.
Douglas AE. The B vitamin nutrition of insects: the contributions of diet, microbiome and horizontally acquired genes. Curr Opin Insect Sci. 2017;23:65–9.
Ren FR, Sun X, Wang TY, Yao YL, Huang YZ, Zhang X, et al. Biotin provisioning by horizontally transferred genes from bacteria confers animal fitness benefits. ISME J. 2020;14:2542–53.
Price DR, Wilson AC. A substrate ambiguous enzyme facilitates genome reduction in an intracellular symbiont. BMC Biol. 2014;12:110–9.
Wilson ACC, Duncan RP. Signatures of host/symbiont genome coevolution in insect nutritional endosymbioses. Proc Natl Acad Sci USA. 2015;112:10255–61.
Chen W, Hasegawa DK, Kaur N, Kliot A, Pinheiro PV, Luan JB, et al. The draft genome of whitefly Bemisia tabaci MEAM1, a global crop pest, provides novel insights into virus transmission, host adaptation, and insecticide resistance. BMC Biol. 2016;14:110.
Sloan DB, Moran NA. Genome reduction and co-evolution between the primary and secondary bacterial symbionts of psyllids. Mol Biol Evol. 2012;29:3781–92.
Nikoh N, McCutcheon JP, Kudo T, Miyagishima S, Moran NA, Nakabachi A. Bacterial genes in the aphid genome: absence of functional gene transfer from Buchnera to its host. PloS Genet. 2010;6:e1000827.
De Barro PJ, Liu SS, Boykin LM, Dinsdale AB. Bemisia tabaci: a statement of species status. Annu Rev Entomol. 2011;56:1–19.
Liu SS, Colvin JJ, De, Barro PJ. Species concepts as applied to the whitefly Bemisia tabaci systematics: how many species are there? J Integr Agr. 2012;11:176–86.
Firdaus S, Vosman B, Hidayati N, Supena EDJ, Visser RGF, van Heusden AW. The Bemisia tabaci species complex: additions from different parts of the world. Insect Sci. 2013;20:723–33.
Gottlieb Y, Ghanim M, Gueguen G, Kontsedalov S, Vavre F, Fleury F, et al. Inherited intracellular ecosystem: symbiotic bacteria share bacteriocytes in whiteflies. FASEB J. 2008;22:2591–9.
Škaljac M, Zanic K, Ban SG, Kontsedalov S, Ghanim M. Co-infection and localization of secondary symbionts in two whitefly species. BMC Microbiol. 2010;10:142–57.
Liu SS, De Barro PJ, Xu J, Luan JB, Zang LS, Ruan YM, et al. Asymmetric mating interactions drive widespread invasion and displacement in a whitefly. Science. 2007;318:1769–72.
Luan JB, Shan HW, Isermann P, Huang JH, Lammerding J, Liu SS, et al. Cellular and molecular remodelling of a host cell for vertical transmission of bacterial symbionts. Proc R Soc B. 2016;283:20160580.
Shan HW, Luan JB, Liu YQ, Douglas AE, Liu SS. The inherited bacterial symbiont Hamiltonella influences the sex ratio of an insect host. Proc R Soc B. 2019;286:20191677.
Sloan DB, Moran NA. Endosymbiotic bacteria as a source of carotenoids in whiteflies. Biol Lett. 2012;8:986–9.
Wang YB, Ren FR, Yao YL, Sun X, Walling LL, Li NN, et al. Intracellular symbionts drive sex ratio in the whitefly by facilitating fertilization and provisioning of B vitamins. ISME J. 2020;14:2923–35.
Qin L, Pan LL, Liu SS. Further insight into reproductive incompatibility between putative cryptic species of the Bemisia tabaci whitefly complex. Insect Sci. 2016;23:215–24.
Luan JB, Sun XP, Fei ZJ, Douglas AE. Maternal inheritance of a single somatic animal cell displayed by the bacteriocyte in the whitefly Bemisia tabaci. Curr Biol. 2018;28:459–65.
Schmittgen TD, Livak KJ. Analyzing real-time PCR data by the comparative CT method. Nat Protoc. 2008;3:1101–8.
Xie W, Chen C, Yang Z, Guo L, Yang X, Wang D, et al. Genome sequencing of the sweetpotato whitefly Bemisia tabaci MED/Q. Gigascience. 2017;6:1–7.
Wang XW, Luan JB, Li JM, Bao YY, Zhang CX, Liu SS. De novo characterization of a whitefly transcriptome and analysis of its gene expression during development. BMC Genom. 2010;11:400–11.
Wang XW, Zhao QY, Luan JB, Wang YJ, Yan GH, Liu SS, et al. Analysis of a native whitefly transcriptome and its sequence divergence with two invasive whitefly species. BMC Genom. 2012;13:529–42.
Gottlieb Y, Ghanim M, Chiel E, Gerling D, Portnoy V, Steinberg S, et al. Identification and localization of a Rickettsia sp. in Bemisia tabaci (Homoptera: Aleyrodidae). Appl Environ Microbiol. 2006;72:3646–52.
Russell CW, Bouvaine S, Newell PD, Douglas AE. Shared metabolic pathways in a coevolved insect-bacterial symbiosis. Appl Environ Microbiol. 2013;79:6117–23.
Ren FR, Bai B, Hong JS, Huang YZ, Luan JB. A microbiological assay for biotin determination in insects. Insect Sci. 2020. https://doi.org/10.1111/1744-7917.12827.
Datsenko KA, Wanner BL. One-step inactivation of chromosomal genes in Escherichia coli K-12 using PCR products. Proc Natl Acad Sci USA. 2000;97:6640–5.
Baba T, Ara T, Hasegawa M, Takai Y, Okumura Y, Baba M, et al. Construction of Escherichia coli K-12 in-frame, single-gene knockout mutants: the Keio collection. Mol Syst Biol. 2006;2:2006–8.
Mori H, Baba T, Yokoyama K, TakeuchiR, Nomura W, Makishi K, et al. Identification of essential genes and synthetic lethal gene combinations in Escherichia coli K-12. Methods Mol Biol. 2015;1279:45–65.
Zhang CR, Shan HW, Xiao N, Zhang FD, Wang XW, Liu YQ, et al. Differential temporal changes of primary and secondary bacterial symbionts and whitefly host fitness following antibiotic treatments. Sci Rep. 2015;5:15898.
Shan HW, Zhang CR, Yan TT, Tang HQ, Wang XW, Liu SS, et al. Temporal changes of symbiont density and host fitness after rifampicin treatment in a whitefly of the Bemisia tabaci species complex. Insect Sci. 2016;23:200–14.
Ghanim M, Kontsedalov S, Czosnek H. Tissue-specific gene silencing by RNA interference in the whitefly Bemisia tabaci (Gennadius). Insect Biochem Mol Bio. 2007;37:732–8.
Nakabachi A, Shigenobu S, Sakazume N, Shiraki T, Yoshihide Hayashizak Y, Carninci P, et al. Transcriptome analysis of the aphid bacteriocyte, the symbiotic host cell that harbors an endocellular mutualistic bacterium, Buchnera. Proc Natl Acad Sci USA. 2005;102:5477–82.
Hansen AK, Moran NA. Aphid genome expression reveals host-symbiont cooperation in the production of amino acids. Proc Natl Acad Sci USA. 2011;108:2849–54.
Smith TE, Moran NA. Coordination of host and symbiont gene expression reveals a metabolic tug-of-war between aphids and Buchnera. Proc Natl Acad Sci USA. 2020;117:2113–21.
Thomas GH, Zucker J, Macdonald SD, Sorokin A, Goryanin I. A fragile metabolic network adapted for cooperation in the symbiotic bacterium Buchnera aphidicola. BMC Syst Biol. 2009;3:24.
Ankrah N, Luan JB, Douglas AE. Cooperative metabolism in a three-partner insect-bacterial symbiosis revealed by metabolic modeling. J Bacteriol. 2017;199:e00872–16.
Douglas AE. Lessons from studying insect symbioses. Cell Host Microbe. 2011;10:359–67.
Price DR, Fen H, Baker JD, Bavan S, Luetje CW, Luetje CW, et al. Aphid amino acid transporter regulates glutamine supply to intracellular bacterial symbionts. Proc Natl Acad Sci USA. 2014;111:320–5.
Hansen AK, Degnan PH. Widespread expression of conserved small RNAs in small symbiont genomes. ISME J. 2014;8:2490–502.
Thairu MW, Cheng S, Hansen AK. A sRNA in a reduced mutualistic symbiont genome regulates its own gene expression. Mol Ecol. 2018;27:1766–76.
Husnik F, Mccutcheon JP. Functional horizontal gene transfer from bacteria to eukaryotes. Nat Rev Microbiol. 2018;16:67–79.
Acknowledgements
The authors thank Professor Angela E. Douglas from Cornell University for constructive comments. We thank Professor Liu Shu-Sheng from Zhejiang University for providing the B. tabaci MEAM1 culture, and Dr Zhang De-Xian, Liu Bing-Qi, Li Ce, and Wang Yan-Bin for their assistance with the experiments. We thank Dr Zhang Xue from China Agricultural University for help and advice on E. coli functional complementation experiments. This work was supported by the National Natural Science Foundation of China (No. 31871967), High-Level Talent Support Foundation from Liaoning and Shenyang Agricultural University (Project XLYC1902104 and 880418001).
Author information
Authors and Affiliations
Contributions
J-BL conceived the study. F-RR conducted symbiont elimination, pantothenate assays, and with C-QL ecology experiments. J-YY, F-RR, and T-YW carried out gene silencing. F-RR, XS, and T-YW performed the complementation experiments. XS carried out FISH and immunofluorescence experiments. Y-LY constructed the phylogenetic tree. F-RR, XS, Y-LY, and J-BL analyzed the data. J-BL wrote the manuscript. All authors edited and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Ren, FR., Sun, X., Wang, TY. et al. Pantothenate mediates the coordination of whitefly and symbiont fitness. ISME J 15, 1655–1667 (2021). https://doi.org/10.1038/s41396-020-00877-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-020-00877-8
This article is cited by
-
Application of bacteria and bacteriophage cocktails for biological control of houseflies
Parasites & Vectors (2024)
-
Symbioses shape feeding niches and diversification across insects
Nature Ecology & Evolution (2023)
-
Silencing horizontally transferred genes for the control of the whitefly Bemisia tabaci
Journal of Pest Science (2023)
-
Genetic innovations in animal–microbe symbioses
Nature Reviews Genetics (2022)
-
Mutualism promotes insect fitness by fungal nutrient compensation and facilitates fungus propagation by mediating insect oviposition preference
The ISME Journal (2022)