Abstract
Sporocarps (fruit bodies) are the sexual reproductive stage in the life cycle of many fungi. They are highly nutritious and consequently vulnerable to grazing by birds and small mammals, and invertebrates, and can be infected by microbial and fungal parasites and pathogens. The complexity of communities thriving inside sporocarps is largely unknown. In this study, we revealed the diversity, taxonomic composition and host preference of fungicolous fungi (i.e., fungi that feed on other fungi) in sporocarps. We carried out DNA metabarcoding of the ITS2 region from 176 sporocarps of 11 wood-decay fungal host species, all collected within a forest in northeast Finland. We assessed the influence of sporocarp traits, such as lifespan, morphology and size, on the fungicolous fungal community. The level of colonisation by fungicolous fungi, measured as the proportion of non-host ITS2 reads, varied between 2.8–39.8% across the 11 host species and was largely dominated by Ascomycota. Host species was the major determinant of the community composition and diversity of fungicolous fungi, suggesting that host adaptation is important for many fungicolous fungi. Furthermore, the alpha diversity was consistently higher in short-lived and resupinate sporocarps compared to long-lived and pileate ones, perhaps due to a more hostile environment for fungal growth in the latter too. The fungicolous fungi represented numerous lineages in the fungal tree of life, among which a significant portion was poorly represented with reference sequences in databases.
Similar content being viewed by others
Introduction
The fungal life cycle is dominated by an inconspicuous mycelial stage, where the mycelia grow hidden belowground or within various substrates. Most Basidiomycota (=Dikarya), produce macroscopic fruit bodies (hereafter referred to as sporocarps) at some point(s) in their lives, where karyogamy, meiosis and formation of meiospores occur. For these fungi, the sporocarp is a key structure of the life cycle, as it bears meiotic spores that are fundamental for dispersal and population persistency. Though not always reflecting phylogenetic relationships [1,2,3], fungi have evolved an extensive morphological diversity of sporocarps [4] that are adapted to different strategies for maximising reproductive effort [5].
Sporocarps are complex structures varying widely in size, shape, colour, persistency and odour. Among wood-decay fungi, which are important nutrient recyclers in forest ecosystems, some species have evolved tough and resistant long-lived (perennial) sporocarps that can persist for several years, while others produce short-lived (ephemeral) sporocarps lasting for only a few days or weeks [6, 7]. Some sporocarps can extend relatively far out from the substrate with a stalk or being bracket-shaped (pileate), while others are relatively simple and “crust-like” (resupinate) adhering to the surface of the substrate [2].
The fruiting stage is a vulnerable phase of the fungal life cycle, as sporocarps are easily damaged by drought, heat, frost, or eaten and destroyed by other organisms. Being highly nutritious [8], sporocarps host a large diversity of bacteria [9,10,11], they can be consumed by small mammals [12] and birds [13], and also provide refuge and resources to a wide diversity of invertebrates such as slugs, snails and worms, as well as arthropods, including insects and mites [14]. Many of the fungivorous organisms play an important role as vectors for the dispersal of hyphal fragments or spores, either attached to their bodies or are excreted undigested, as they are enriched in recalcitrant compounds like chitin [15].
Whilst studies have revealed obligate associations between insects and fungi [16,17,18,19], including highly specific associations, far less is known about the diversity and richness of other species of fungi residing inside sporocarps. These fungi, which feed on other fungi, are termed fungicolous [20, 21] but little is known about their diversity, occurrence and interactions with host fungi.
So far, about 1500 fungicolous fungal taxa have been recorded [22]. Considering a global fungal diversity estimated between about two and four million species [23], the number of fungicolous fungi is probably a gross underestimate. Among fungicolous fungi adapted to colonise sporocarps, some can themselves produce macroscopic sporocarps through a rapid life cycle [24], but many do not, which is probably why they have largely been overlooked, despite being widespread in nature. One such overlooked group is yeasts, which can also be found as endophytes in plants and trees [25, 26], lichens [27] as well as in tripartite interactions with parasitic fungi and mushrooms [28]. Apart from several taxonomic studies [29,30,31,32,33] and studies addressing the parasitic activity of fungicolous fungi (see ref. [22]), their broad diversity is largely unexplored and their ecological roles uncharacterised. There are, therefore, many unaddressed questions, such as: how common are such fungal–fungal associations? How diverse is the fungicolous fungal community associated with the sporocarp of particular species, and does the community composition vary among and between sporocarps of the same and different species? Do fungal–fungal co-occurrences arise from random opportunistic use of the sporocarps, or do they represent adaptation to the specific niche(s) each host offers?
One reason why these fundamental questions remain unresolved is that fungicolous fungi have mainly been studied using culture-dependent approaches and morphology-based identification [34], which provide a limited view of the diversity. DNA metabarcoding, based on high-throughput sequencing of amplified marker(s), has become a powerful approach to scrutinise complex fungal communities and does not require prior knowledge of the community composition. Recently, a DNA-based study by Koskinen et al. [9] revealed that soft and short-lived agaricoid sporocarps (of the orders Agaricales, Boletales and Russulales) house a high diversity of fungicolous fungi, but without any distinct host preference. The relatively short lifespan of fleshy agarics (compared to other annual fungi) could partially explain this lack of host specificity.
Sporocarp characteristics, as well as environmental variables such as climate, are potential drivers of fungicolous fungal community composition. The structural composition of sporocarps vary from a simple organisation (monomitic hyphal system) in ephemeral sporocarps, to a more complex structure (di- or trimitic hyphal system) in the persistent ones. In addition to sporocarp morphology and persistency, extensive differentiation in their biochemical composition may also explain differences in fungicolous fungal community composition. Further, short-lived, fleshy and soft sporocarps typically have a higher water content than the resistant long-lived sporocarps, which may favour germination and growth of fungicolous fungi. Together, these sporocarp-related traits may lead to a variable degree of host-specialisation among fungicolous fungi. There are indications that some fungicolous fungi are host-specific [35,36,37,38] and, therefore, constrained to the geographic ranges of their hosts.
The overarching aim of this study was to reveal the diversity of fungicolous fungi residing in sporocarps of wood-inhabiting fungi. This was done by DNA metabarcoding analyses of sporocarp tissue from eleven wood-decay fungi, using the rDNA ITS2 region as a target marker. We expected that a vast majority of the ITS2 sequences will be derived from the host sporocarp tissue, but that a minor fraction of non-host ITS2 sequences, representing fungicolous fungi, will vary systematically according to the hosts, reflecting different levels of colonisation. We hypothesised that the short-lived fleshy sporocarps with higher water content are a more suitable habitat for fungicolous fungi, thus resulting in a higher proportion of non-host ITS2 sequences and higher diversity of fungicolous fungi (H1-1). Alternatively, the long-lived resistant sporocarps may accumulate a higher diversity of fungicolous fungi since they are available for colonisation for considerably longer, up to several years (H1-2). Second, we asked how the diversity of fungicolous fungi varies according to sporocarp morphology. We hypothesised that resupinate sporocarps house a higher diversity of fungicolous fungi due to a larger surface area to volume ratio, thus providing relatively more area for colonisation of fungicolous fungi (H2). Next, we ask how species-specific fungicolous fungi are. Due to differences in physical structure and chemical composition of the polypore sporocarps, we expected numerous specific co-occurrences between fungal hosts and fungicolous fungi (H3). Finally, the abundance of fungal hosts, surveyed at local and regional scales, allowed us to investigate whether there is a correlation between host density and the species richness of fungicolous fungi. Assuming that some degree of host preference exists, we expected that widespread and abundant hosts, acting as small islands in the forest landscape, provide habitat and resources for a larger pool of fungicolous fungi compared to rare hosts (H4). According to island biogeography theory, reduced distance separating hosts (i.e., distance effect) may increase the colonisation rate of fungicolous fungi.
Material and methods
Sample collection
The sampling was conducted in October 2014 in an old-growth forest dominated by Norway spruce (Picea abies) in northeast Finland, Kuhmo, Issakka (62°38′42.4″N 30°47′47.8″E). Compared to southern Finland, forestry has been less intensive in the northeast, including the Kuhmo region, where intensive forestry also lasted over a shorter period, which is today reflected in a relatively large area of old-growth forest with high connectivity between the remaining patches of old-growth boreal forest. Consequently, this area is relatively rich in dead wood and represents a biodiversity hotpot for many wood-dependent organisms, including wood-decay fungi where several species fruit at the same time, allowing intra- and inter-species comparative studies. To understand the natural dynamics of fungicolous fungi, choosing an area of this kind is preferable and allows a comparison across several species while maintaining other environmental variables constant. In addition, to limit the effect of confounding variables attributed to substrate, landscape and seasonality, we collected sporocarps only from logs of Picea abies, in one area (20 ha) within a short time frame (2 days).
Due to their importance in dead wood decomposition, their conspicuous size and well-characterised species traits and ecologies, polypores are a good study system to investigate the natural dynamics of fungicolous fungi in old-growth forests. We targeted common and rare boreal species having dead wood of Picea abies as main resource, that fruit in a shared time frame and with sufficient occurrences thus allowing a comparative population-level study. Eleven wood-decay fungal species were collected: the polypores Amylocystis lapponica (amylap), Antrodia serialis (antser), Gloeophyllum sepiarium (glosep), Fomitopsis (Rhodofomes) rosea (fomros), Fomitopsis pinicola (fompin), Phellopilus nigrolimitatus (phenig), Phellinidium ferrugineofuscus (phefer), Phellinus (Fuscoporia) viticola (phevit), Postia cyanescens in Postia caesia complex (poscae), Trichaptum abietinum (triabi) and the corticioid species Phlebia centrifuga (phecen). For each of the focal species, we included 16 sporocarps (individuals) collected on different spruce logs (Fig. S1), totalling 176 sporocarps.
Sample processing and DNA extraction
We processed both annual and perennial species similarly, by removing the outer surface layer to avoid aerial contaminants and processing the subiculum layer. Between 10 and 15 small pieces of ~5 mm3, each were cut from the subiculum layer, placed in 2-mL tubes containing 800 µl of 2% CTAB and 1% beta-mercaptoethanol, and stored at −20 °C until DNA extraction. The samples were then grinded 4 × 45 s at 25 oscillations s−1 with two tungsten-carbide beads (2 mm) using the Retsch MM200 mixer mill and stored at −80 °C overnight. Subsequently, DNA was extracted using a modified CTAB/chloroform extraction protocol [39, 40]. The DNA extracts were cleaned with an E.Z.N.A Soil DNA kit (Omega Biotek) by adding the HTR reagent and then following the manufacturer´s guidelines, and eluted into 100-µL elution buffer. For each sample, we quantified the DNA concentration using the Qubit dsDNA BR Assay kit (Life Technologies) and standardised with 10-mM Tris in the range of 5–10 ng µL−1.
Preparation of metabarcoding libraries and sequencing
In addition to the 176 DNA extracts, twelve technical replicates (at least one sample for each species), two mock samples (artificial fungal community composed of six other fungal species) and two PCR-negative controls were added in the library preparation, resulting in a total of 192 samples processed in 2 × 96 PCR plates. Amplicon libraries were constructed using a combination of 96 uniquely tagged primers designed to target the variable ITS2 region, with the reverse primer ITS4 (5′-xCTCCGCTTATTGATATG from [41]) and modified forward primer gITS7 (5′-xGTGARTCATCGARTCTTTG from [42]), barcodes x ranging from 6 to 9 base pair. One microlitre of DNA template was used for the 25-µl PCR reaction containing 14.6 µl of Milli-Q water, 2.5 µl of 10x Gold buffer, 0.2-µl dNTPs (25 nM), 1.5 µl each of reverse and forward primers (10 µM), 2.5-µl MgCl2 (50 mM), 1.0-µl BSA (20 mg/ml) and 0.2-µl AmpliTaq Gold polymerase (5 U/µl). DNA was amplified as follows: initial denaturation at 95 °C for 5 min, followed by 25 cycles of denaturation at 95 °C for 30 s, annealing at 55 °C for 30 s, and elongation at 72 °C for 1 min, and a final elongation step at 72 °C for 10 min. Amplifications were assessed by electrophoresis on a 1.5% agarose gel, normalised using the SequalPrep Normalisation Plate Kit (Invitrogen) and eluted into a total volume of 20-μl Elution Buffer. The 96 PCR products within each plate were pooled, concentrated and cleaned using Agencourt AMPure XP magnetic beads (Nerliens Meszansky AS). Quality control was performed using ds DNA 1000 Bioanalyzer (Agilent Technologies) and Qubit (Life Technologies). The two libraries were barcoded with Illumina adapters, indexed with 20% PhiX and sequenced in one Illumina MiSeq (Illumina, San Diego, CA, USA) lane with 2*300-bp paired-end reads at StarSEQ (StarSEQ GmbH, Mainz, DE).
Bioinformatics analyses
The entire raw dataset comprised 25,953,804 sequences. Sequencing errors, including substitutions and indels were corrected using bayeshammer software [43]. The resulting 25,935,562 reads were merged using pear [44], with a minimum of sequence length of 200 bp, and filtered by quality and length using Fastx-toolkit [45] (quality = 30, percentage = 90) and vsearch [46] (fastq_maxee = 0.5, fastqminlen = 100, fastq_truncqual = 21). Paired-end reads were demultiplexed using sdm software [47]. Both forward and reverse primers were removed using cutadapt [48] followed by ITS extractor program [49]. The ITS2 sequences were dereplicated and sorted by abundance using vsearch. Operational taxonomic units (OTUs) were clustered at 97% similarity and chimeras were removed with vsearch (abskew = 2, mindiffs = 3, mindiv = 0.8, minh = 0.28). As we wanted to restrict our analyses to the most abundant fungi, singletons were excluded, and only OTUs supported by a total observation count of >10 sequences in the dataset were retained. Taxonomy of the ITS2 representative sequences was assigned using BLAST in qiime [50] against the NCBI and unite databases [51] (v. 7_99_s_01.12.2017). The raw OTU table comprised 16,196,863 reads assembled into 2188 OTUs. To correct for over-splitting of OTUs due to e.g., PCR and sequencing errors, we processed the raw OTU table using the lulu algorithm [52] and retained 1632 OTUs. We removed the 12 PCR replicates from the dataset after checking the repeatability by DCA (Fig. S2). OTUs without corresponding blast hit and non-fungal OTUs (mainly Rhizaria), were excluded from the dataset (1385 OTUs retained). To minimise the proportion of tag-switching inherent to library construction, Illumina sequencing platforms and bioinformatics steps, we ran the owi_renormalized script (https://github.com/metabarpark/) with a 10% threshold. We subsequently removed the OTUs corresponding to the mock community composed of six fungal species, the two negative PCR controls and manually curated for tag-switching, thus removing the mis-assigned reads corresponding to the host species in other samples. The final filtered OTU table was composed of 1794,144 reads accounting for 1367 OTUs. We then used the rrarefy function in the R package vegan ([53], v 2.4-5) to make abundances comparable between samples (10,000 reads/sample) and excluded two samples (fompin3 and fompin4).
Compilation of metadata
The focal fungal hosts are characterised by measurable traits that vary across species. A series of these trait variables were compiled from the literature [6, 7, 54, 55]. Life-history traits included (i) specialisation to main host tree species, and traits-related to sporocarp such as (ii) sporocarp lifespan (short- or long-lived), (iii) morphology (pileate or resupinate), (iv) sporocarp hyphal system (mono, di- or trimitic), (v) size, which were categorised in three classes, with 1 = small (up to the size of a fingertip for pileate or a few fingerprints for resupinate), 2 = intermediate (up to the size of an apple-half for pileate or a palm for resupinate) and 3 = large (larger than the size of a fist for pileate or two palms for resupinate); (vi) thickness (minimum and maximum) and (vii) average hymenophore surface area (J. Nordén, unpublished data). Abundance data were inferred at the (viii) forest scale and were registered as the number of logs with the focal species (local abundance) [56], (ix) regional abundance of the species [55] and finally, (x) the proportion of fungicolous fungal reads in each sample was estimated from the current dataset and inferred as a response variable.
Statistical analyses
To estimate whether the sampling effort (n = 16 sporocarps per fungal host) reached an acceptable level of efficiency, we constructed species-accumulation curves for the 11 fungal hosts using the function specaccum in vegan with the method rarefaction (Fig. S3). The effects of sporocarp-related traits were evaluated using OTU richness and Shannon H index, both calculated per sporocarp, as incorporated in vegan and the visualisations were made with ggplot2 package v 2.2.1 [57]. To investigate the effect of sporocarp lifespan and morphology on OTU richness (log-transformed) and Shannon diversity, linear mixed effect models were fitted using LME4 package in R, with host species as a random effect. Full models were specified with all the explanatory variables, and to identify the most parsimonious model, the models were subjected to a backward elimination procedure based on Akaike information criterion.
To account for the phylogenetic relationship between hosts, we extracted partial 28S rRNA sequences for the 11 fungal species from NCBI, performed a multiple sequence alignment in muscle [58] and calculated a pairwise distance matrix based on the Maximum Composite Likelihood model in MEGA [59]. Likewise, we computed a distance matrix from the fungicolous fungal community, aggregated both matrices and performed a Procrustes analysis with 9999 permutations in vegan to correlate the two matrices.
To visualise the variation in the community composition of OTUs across species, we used non-metric multidimensional scaling (NMDS) implemented in vegan, with parameter settings as recommended by Liu et al. [60]. To confirm convergence of the NMDS, the two best solutions were compared using Procrustes comparisons with 999 permutations. Since it was not possible to produce NMDS ordinations with acceptable stress values and convergence, we reduced the dimensions of the data by removing 11 samples with <5 OTUs. In addition, we performed a DCA and tested for consistency with NMDS; correspondence of the results and absence of artefacts, such as arch-effect (NMDS), tongue effect (DCA) and absence of extreme outliers, were interpreted as reliable gradients found. The effects of sporocarp morphology (pileate vs. resupinate), lifespan (short vs. long-lived) and size, local abundance of the fungal host species (occurrence) and the proportion of fungicolous fungi were visualised using vectors from the envfit function in vegan. The fit (R2) of each variable to the NMDS was assessed with a Monte-Carlo analysis of 999 permutations (Table 1). Using variance partitioning, canonical correspondence analysis with 999 permutations, we quantified the independent components of variation in the fungal community composition explained by hosts, sporocarp-related traits (lifespan, morphology and size), abundance of hosts (local and regional) and proportion of fungicolous fungi. To reveal detailed information on the multiple responses of the fungal community to each variable, we applied Bayesian generalised linear modelling implemented in R-INLA [61], in which host species was employed as a factor to predict the proportion of fungicolous fungi. To investigate the effects of host species abundance (applied as log of counts data) at local and regional scales on the richness of fungicolous fungi, we estimated an additional alpha index, Chao1 using phyloseq [62] that accounts for rare OTUs in low abundance in the dataset.
To identify significant co-occurrence patterns between host fungi and fungicolous fungi (i.e., species that are significantly correlated in the dataset), we performed an indicator species analysis using the R package Indicspecies [63]. First, we applied the multipatt function to determine the list of OTUs that are correlated to a particular host. Second, based on a permutation test, we filtered the indicator value index to retain only significant associations (p value < 0.05) with at least two fungal host occurrences. We then used Cytoscape [64] to visualise the fungicolous fungi OTUs interaction networks among the eleven host fungi.
Results
Proportion and taxonomic composition of fungicolous fungi
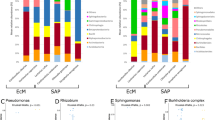
Using a metabarcoding approach, we analysed the diversity of fungicolous fungi inhabiting sporocarps of eleven fungal host species collected within a forest. As expected, a large proportion of the ITS2 sequences corresponded to the fungal host species, while the relative abundances of non-host ITS2 sequences, representing fungicolous fungi, ranged from 2.8% in Fomitopsis rosea to 39.8% in Amylocystis lapponica (Fig. 1A). Most of the fungicolous fungi belonged to Ascomycota and varied between 77% in Phellopilus nigrolimitatus to 96% in Antrodia serialis, with Helotiales and Hypocreales as the most abundant orders (Fig. 1B). However, in the polypore Fomitopsis pinicola, ascomycete yeasts of Saccharomycetales were highly dominant, accounting for 47% of the fungicolous sequences. Basidiomycota, with 6.7% of the fungicolous reads, was the second most abundant phylum, with Atheliales and Cantharellales as predominant orders. Mucoromycota (6.3%) were present in variable proportions in eight of the focal hosts, mainly represented by the orders Mucorales and Umbelopsidales. In addition to these predominant phyla, we detected at lower frequencies (<1%) OTUs belonging to Chytridiomycota, Mortierellomycota, Olpidiomycota and Rozellomycota, mainly present in the hosts Trichaptum abietinum and Phellinus viticola.
A Bar plot representing the proportion of ITS2 sequences corresponding to non-host fungicolous fungi relative to all fungal sequences including the hosts. For each species, an average was calculated from 16 individual sporocarps and error bars represent 95% confidence intervals, indicating systematic differences across species. B Average proportion of 15 main orders (alphabetical listing) of fungicolous fungi, where blue colours represent Basidiomycota, red Ascomycota and green Mucoromycota.
Fungicolous fungal diversity varies with life-history traits of host fungi
The fungal host species were classified into two categories based on the lifespan of their sporocarps: short-lived (four species with 64 individuals) versus long-lived sporocarps (seven species with 110 individuals). A significant difference in OTU richness was observed between these two categories (Fig. 2A), with short-lived sporocarps hosting on average 2.58 (p = 0.03) more OTUs than the long-lived. In contrast, the Shannon diversity index was not significantly different between short- and long-lived sporocarps (p = 0.16, Fig. 2C). Among the short-lived species, T. abietinum (triabi) and P. centrifuga (phecen), producing resupinate sporocarps with large surface area, showed particularly high OTU richness. Regarding sporocarp morphology, we observed significantly higher OTU richness (estimate = 3.99 OTUs, p value = 0.004) as well as Shannon diversity index (estimate = 1.13 OTUs, p value = 0.001) in resupinate compared to pileate sporocarps. Species with pileate sporocarps showed higher variability in both OTU richness and Shannon diversity than resupinate species.
Box plots illustrating variation in OTU richness (A, B) and Shannon diversity (C, D) of fungicolous fungi across 11 wood-decay fungal species. For each fungal host species, 16 sporocarps were analysed (excluding two fompin samples). The black dots represent data falling outside the interquartile range while the median is indicated by the dark line. The smaller frames (down left corner) compare long- and short-lived sporocarps (A, C) and pileate and resupinate sporocarps (B, D), respectively. Statistical differences between the different categories were evaluated using a Wilcoxon test with p value < 0.05.
To test whether the diversity of fungicolous fungi is correlated to host abundance, we related the abundance data of the host species, both at the local and regional scales, against OTU richness, Shannon and Chao1 indices of fungicolous fungi (Fig. S4). We expected hosts with smaller population sizes to be associated with a lower diversity of fungicolous fungi. However, all correlations were weak and non-significant.
Tentative drivers of the community pattern
Multivariate analyses indicated that all the assessed variables were significantly related to the compositional variation (Table 2). The analyses revealed that the fungicolous fungal communities were distinct between host species (Fig. 3, R2 = 0.56, p value = 0.001 and see Fig. S5 for individual sporocarps). Interestingly, species with long-lived sporocarps (e.g., F. rosea and F. pinicola) varied more in composition compared to many short-lived fungi (e.g., A. serialis or P. caesia), indicating that the ephemeral sporocarps host more species-specific fungicolous fungi than the perennial species (Fig. 3). Overall, the structure of the community composition clearly indicated strong associations between fungicolous fungi and fungal host species. Although non-significant (p = 0.19, N = 11), there was a high correlation (0.68) between the genetic distance of the host fungi, assessed using a 28S rRNA phylogeny, and the community distance of the fungicolous fungi.
Fungal–fungal co-occurrences
The correlation-based network shown in Fig. 4 illustrates how the fungicolous fungal OTUs were associated to the different fungal hosts. Out of the 454 fungal OTUs having significant associations (p < 0.01) with the eleven host fungi, 236 fungicolous OTUs were associated with a single host, whereas 218 OTUs co-occurred in at least two host species (up to 11 host species), the latter more likely reflecting generalists. The majority of the host fungi had species-specific associations with fungicolous fungal OTUs, with the exception of Fomitopsis rosea, which had only shared co-occurrences with other fungal hosts. In contrast, in four host species (Phellopilus nigrolimitatus, Phlebia centrifuga, Phellinus viticola and Trichaptum abietinum) we detected a high number (30–90) of species-specific fungicolous fungi that were not present in other hosts. At the same time, these species also shared the most fungicolous fungi with the other hosts, suggesting that these four hosts species harbour many specialists as well as generalists. Three hosts with short-lived sporocarps (Amylocystis lapponica, Antrodia serialis and Gloeophyllum sepiarium) were associated with a moderate number (7–14) of host-specific fungicolous OTUs. In general, species with smaller sporocarps and/or a smaller total hymenophore surface area per log have fewer specific associations, except for P. ferrugineofuscus and F. pinicola.
Fungal hosts are represented by filled-square symbols while circles represent fungicolous fungal OTUs coloured according to phyla. The 236 OTUs of fungicolous fungi that are specific to a single host are located on the outer edge while 218 OTUs shared between two hosts or more are indicated on the inner edge. All edges represent a significant link (999 permutations test, p value < 0.05) between a host species and a fungicolous fungal OTU.
Among the species-specific OTUs, there was a large proportion of fungi without taxonomic assignment below the kingdom level, accounting for 13% of all the significant co-occurrences (52% unidentified OTUs at genus level). We could assign functions with some level of confidence to only a proportion of the identified OTUs. Of these, there were a few wood saprotrophs, including the brown rot Calocera sp. (Dacrymycetes) and the two white rotters Dichostereum granulosum and Athelia decipiens, all three species known to decay Norway spruce. In addition, several species known as soil saprotrophs in the orders Mortierellales and Mucorales were associated with long-lived sporocarps. The indicator species analysis supported numerous associations between P. nigrolimitatus and OTUs affiliated to Mucoromycota (Umbelopsis and Mucor). Besides fungal species known to be associated with plants (endophytes, mycorrhizal fungi and plant pathogens), we also detected fungal pathogens of animals, which were mainly from the class Leotiomycetes (Cryptomyces, Cryptosporiopsis, Leconeurospora and Pseudogymnoascus). Fungi previously reported to be associated with other fungi (facultative and obligate), formed both specific and generalist associations with the hosts. Endomycete yeasts described as parasites e.g., classes Microbotryomycetes and Tremellomycetes, made up a large proportion of the Basidiomycota, particularly in the hosts P. caesia and G. sepiarium. Several Ascomycota (e.g., Coleophoma empetri, Hypomyces spp., Tolypocladium spp., Talaromyces infraolivaceus) reported as parasites of fungi were also present in many of the hosts.
Discussion
This study is one of the very first providing a DNA-based overview of fungal communities residing within sporocarps of various fungal hosts. We unveiled a species-specific community of fungicolous fungi inside sporocarps of wood-decay fungi in the boreal forest ecosystem. A significant proportion of the fungicolous fungi was poorly represented with associated reference sequences in databases, indicating that this diversity has hitherto been poorly explored. Our results support fungicolous fungi being widespread across different lineages within the fungal kingdom [65, 66], and dominated by Ascomycota, most with unknown ecological functions.
Are short-lived sporocarps more suitable habitat for fungicolous fungi?
In concordance with our first hypothesis (H1-1), we obtained evidence that short-lived sporocarps hosted more fungicolous fungi, both in terms of OTUs richness and Shannon diversity. Most likely, the fleshy, spongy and ephemeral sporocarps, with a higher moisture content, represent a more favourable niche, facilitating germination and growth of fungicolous fungi, as compared to tough perennial sporocarps that might be more resistant to colonisation by fungicolous fungi. Our alternative hypothesis (H1-2), that long-lived sporocarps accumulate a higher diversity due to longer exposure time, was not supported. Colonisation by fungicolous fungi may represent a neglected but strong selection pressure on fruit body evolution, giving rise to different fruiting strategies. Fungi with short-lived sporocarps, predominant in the fungal kingdom, have developed a ‘boom and bust’ strategy, involving rapid sporulation and little investment in structural and physical defence against fungicolous fungi, leading to rapid senescence [67]. The opposite strategy is the production of complex and resistant long-lived sporocarps, limiting the colonisation of fungicolous fungi. However, there is considerable variation in sporocarp longevity between those enduring for several months (e.g., P. ferrugineofuscus) to several years (P. nigrolimitatus), which may partly explain the variation in OTU richness and Shannon diversity of fungicolous fungi observed among these species. Regardless of the fruiting strategies, the mycelia inside the wood are generally long-lived [68].
Compared to the other host species producing short-lived sporocarps, Amylocystis lapponica had a lower alpha diversity of fungicolous fungi. Yearly variation in climatic conditions and sampling time (e.g., late autumn) can directly influence the diversity and composition of fungicolous fungi in short-lived species, a plausible reason for the lower diversity detected in A. laponica. Interestingly, we detected a high abundance of Hypomyces sequences in A. lapponica, which could also be seen in the field as orange to reddish spots on the host sporocarps. Hypomyces is a diverse worldwide genus of Pezizomycotina, including mycoparasitic species altering the original colour and shapes of their hosts, e.g., Amanita, Agaricus, Lactarius, Russula and Polyporales [69,70,71,72]. In several patho-systems when species-specific parasites dominate, like Hypomyces in A. lapponica, this could lead to competitive exclusion of other parasites (see ref. [73]), resulting in low alpha diversity as observed in sporocarps of A. lapponica. In addition, the relatively high density of A. lapponica in Issakka forest could directly amplify the abundance of specific-specific parasites.
Is sporocarp morphology of importance?
We found solid support for our second hypothesis (H2), implying that resupinate sporocarps house a higher diversity of fungicolous fungi compared to pileate species. Resupinate sporocarps possess a significantly higher surface area to volume ratio. Consequently, a larger surface area of the sporocarps is in contact with the wooden substrate in resupinate species, as well as exposed to air and wind, precipitation and (in-)vertebrates, which are potential vectors of fungicolous fungi [74]. Though we processed the subiculum layer of the sporocarps, nonetheless we cannot exclude that a proportion of these non-host fungi might only be present as spores or mycelium fragments, without real ecological roles within the host fungi.
How species-specific are the fungicolous fungi?
As hypothesised, we observed a relatively high number of species-specific co-occurrences between the focal host fungi and fungicolous fungi (H3). The multivariate analyses also supported that the assembly process of fungicolous fungi in sporocarps is not a random process, but instead governed by species-specific filtering processes. Indeed, at both forest scale and broader landscape level, biogeographic processes affecting the host fungi may also impact the fungicolous fungi. Among the possible probabilistic filtering processes maintaining species-specific associations over broader scales are local habitat suitability in terms of abiotic factors within the limits of variation for growth of fungicolous fungi (e.g., climatic variables, moisture content inside hosts, pH), sporocarp abundance, nutrient availability in the sporocarps, host susceptibility (defence mechanisms) and the capacity of colonisation by fungicolous fungi (types of interaction).
To establish and grow, fungicolous fungi must be able to withstand the biochemical defences of the host fungi. Polypore fungi produce secondary metabolites, including triterpenoids, organic acids and volatile organic compounds, known for their wide range of biological activities such as antimicrobial, anti-inflammatory, cytotoxic, antioxidant, immunostimulant and anti-thrombin properties [75]. Adaption to the various secondary metabolites may lead to host specificity.
Striking specific co-occurrences were observed between the host P. nigrolimitatus and several OTUs of Umbelopsidales (Umbelopsis ramanniana, U. isabellina and U. angularis). Commonly isolated from rhizosphere soils worldwide [76], there is increasing evidence that species of Umbelopsis occur as root endophytes of boreal trees [77]. Either sporocarps of P. nigrolimitatus, commonly produced beneath large and well-decayed logs, enable the recruitment of soil fungi as a result of ground contact or Umbelopsis species may have biotrophic relationships with the long-lived sporocarps of P. nigrolimitatus.
To what extent does host abundance matter?
Island biogeography theory suggests that remote islands host less biodiversity than connected islands purely due to dispersal limitations. Likewise, if there are species-specific associations between fungicolous fungi and polypore hosts, as we have seen, one might expect that host species with large populations and numerous sporocarps will harbour a higher richness of fungicolous fungi compared to rarer species. The availability of abundance data of the host species, measured as number of colonised substrates (logs) at local and regional scales, allowed us to test this hypothesis (H4). However, we observed no significant correlations between the abundance of hosts and OTU richness (all diversity indices) at either spatial scale. Hence, other factors than host abundance are more important in driving the diversity of fungicolous fungi. In addition to the focal host species being historically restricted to old-growth boreal forests and regionally more abundant in northeast Finland, the number of host species examined here (n = 11) might be too low to draw any meaningful conclusions on this result.
Plausible ecological roles of fungicolous fungi?
We revealed a high diversity of fungicolous fungi, widely spread across different lineages in the fungal kingdom. Among the fungicolous fungi described, most are classified as mycoparasites, probably based on the symptoms (modification of colour and shape and reduced growth) observed on mushrooms in nature or in the farming industry [78, 79]. Though, it is tempting to speculate that they have different ecological roles, as mycoparasites, saprotrophs of dead fungal host tissue or even mutualists, but from our data it is not possible to clearly determine their life strategies. It is well known that many fungi living as plant endophytes have beneficial functions to their hosts, such as Epichloë species [80,81,82,83]. These fungi employ different mechanisms which protect their host plants from grazing [84], either by acting directly (release of growth promoter and secondary metabolites) or indirectly (modification of host physiology and nutrient balance). An interesting parallel would be if fungicolous fungi could also produce metabolites protecting the host sporocarps against grazing e.g., by invertebrates. One might expect such beneficial fungi to coevolve with the host fungus and develop host specificity. Given the relatively short lifespan of sporocarps of some fungi, one may ask why fungicolous fungi adapt to hosts with such fruiting strategy? Obviously, the ease of colonisation and growth, together with low level of host defence is plausible arguments. However, the reasons may be more complex as some fungicolous fungi may not be solely restricted to the colonisation of sporocarps. For instance, some invasive necrotrophs are also capable to penetrate host cell walls and grow within the vegetative mycelial [85], just as endosymbiotic bacteria dwell inside fungal mycelium [86]. In such cases, the whole life cycle of the host is parasitised.
Further work and implications
Our findings raise questions about how widespread fungal–fungal co-occurrences are across different ecosystems. This study was conducted on eleven fungal host species within one forest. A further step would be to assess whether these co-occurrence patterns, between fungicolous fungi and their fungal hosts, are stable at broader geographic scales, and to what degree they are also influenced by site characteristics, e.g., climate and forest connectivity. If the observed co-occurrence patterns are stable across broader geographic scales, this will support a strong coevolution. Since the composition of secondary metabolites in the host sporocarps likely play an important role, another step would be to characterise the active secondary metabolites of the different host species. In this regard one might expect that long-lived sporocarps have a more complex profile of secondary metabolites, leading to higher resistance to fungicolous fungi compared to short-lived ones. Owing to their ecological importance in nutrient recycling, widespread distribution, large diversity of sporocarp characteristics and the production of broad secondary metabolites, polypore fungi are clearly a well-suited study system for investigating host specificity and fungal–fungal coevolution.
Another aspect that may play an important role in structuring the community of fungicolous fungi is their tentative interactions with arthropods. Colonisation by arthropods may shape the fungal community, either directly by vectoring fungicolous fungi or indirectly by altering the physical structure and chemical composition of the sporocarps. Arthropods very likely act as vectors, not only of the host fungi, but also of its associated fungicolous fungi. Considering the community composition of arthropods colonising the host sporocarps together with the fungicolous fungi may provide valuable information about the community assembly processes in the sporocarps.
Data availability
The MiSeq raw sequence data is available on the NCBI short read archive (SUB8582638) under Bioproject PRJNA680258. The scripts for performing the statistical analyses and generating the main and Supplementary fig. together with the OTU table, the metadata, intermediate files are deposited on Dryad https://doi.org/10.5061/dryad.573n5tb66. Supplementary Material is available for download on the ISME website.
References
Hibbett DS, Pine EM, Langer E, Langer G, Donoghue MJ. Evolution of gilled mushrooms and puffballs inferred from ribosomal DNA sequences. Proc Natl Acad Sci. 1997;94:12002–6.
Hibbett DS, Binder M. Evolution of complex fruiting–body morphologies in homobasidiomycetes. Proc R Soc Lond Ser B: Biol Sci. 2002;269:1963–9.
Varga T, Krizsán K, Földi C, Dima B, Sánchez-García M, Sánchez-Ramírez S, et al. Megaphylogeny resolves global patterns of mushroom evolution. Nat Ecol Evol. 2019;3:668.
Kües U, Khonsuntia W, Subba S. Complex fungi. Fungal Biol. Rev. 2018;32:205–18.
Halbwachs H, Simmel J, Bässler C. Tales and mysteries of fungal fruiting: how morphological and physiological traits affect a pileate lifestyle. Fungal Biol Rev. 2016;30:36–61.
Ryvarden L, Gilbertson RL. European polypores, vol. 1. Synopsis Fungorum 6. Fungiflora A/S, Oslo, Norway, 1993.
Ryvarden L, Gilbertson RL. European polypores: part 2: Meripilus-Tyromyces. Fungiflora A/S; Oslo, Norway, 1994. p. 394–43.
Kalač P. A review of chemical composition and nutritional value of wild‐growing and cultivated mushrooms. J Sci Food Agric. 2013;93:209–18.
Koskinen J, Roslin T, Nyman T, Abrego N, Michell C, Vesterinen EJ. Finding flies in the mushroom soup: host specificity of fungus‐associated communities revisited with a novel molecular method. Mol Ecol. 2019;28:190–202.
Pent M, Põldmaa K, Bahram M. Bacterial communities in boreal forest mushrooms are shaped both by soil parameters and host identity. Front Microbiol. 2017;8:836.
Warmink JA, Nazir R, Van Elsas JD. Universal and species‐specific bacterial ‘fungiphiles’ in the mycospheres of different basidiomycetous fungi. Environ Microbiol. 2009;11:300–12.
Fogel R, Trappe JM. Fungus consumption (mycophagy) by small animals. Northwest Sci. 1978;52:1–31.
Elliott TF, Jusino MA, Trappe JM, Lepp H, Ballard GA, Bruhl JJ, et al. A global review of the ecological significance of symbiotic associations between birds and fungi. Fungal Divers. 2019;98:161–94.
Boddy L, Jones TH. Interactions between basidiomycota and invertebrates. In: British Mycological Society Symposia Series, vol. 28. Academic Press, Amsterdam, 2008. p. 155–79.
Smrž J, Soukalová H, Čatská V, Hubert J. Feeding patterns of Tyrophagus putrescentiae (Sarcoptiformes: Acaridae) indicate that mycophagy is not a single and homogeneous category of nutritional biology. J Insect Sci. 2016;16:94.
Beaver RA, Wilding N, Collins N, Hammond P, Webber J. Insect-fungus relationships in the bark and ambrosia beetles. (Eds Wilding N, Collins NM, Hammond PM & Weber JF). Insect-fungus Interactions. Academic Press, New York, 1989. p. 121–43.
Boddy L. Interactions between fungi and other microbes. (Eds Watkinson S, Boddy L, Money N). The fungi. Academic Press, Oxford, 2016. p. 337–60.
Schigel DS. Fungivory and host associations of Coleoptera: a bibliography and review of research approaches. Mycology. 2012;3:258–72.
Six DL. Ecological and evolutionary determinants of bark beetle—fungus symbioses. Insects. 2012;3:339–66.
Barnett HL. The nature of mycoparasitism by fungi. Annu Rev Microbiol. 1963;17:1–14.
Rudakov OL. Physiological groups in mycophilic fungi. Mycologia. 1978;70:150–9.
Sun JZ, Liu XZ, McKenzie EH, Jeewon R, Liu JKJ, Zhang XL, et al. Fungicolous fungi: terminology, diversity, distribution, evolution, and species checklist. Fungal Divers. 2019;95:337–430.
Hawksworth DL, Lücking R. Fungal diversity revisited: 2.2 to 3.8 million species. (Eds Heitman J, Howlett B, Crous P, Stukenbrock E, James T & Gow NAR). The fungal kingdom. ASM Press, Washington, DC, 2017: p. 79–95.
Marano AV, Pires-Zottarelli CLA, Barrera MD, Steciow MM, Gleason FH. Diversity, role in decomposition, and succession of zoosporic fungi and straminipiles on submerged decaying leaves in a woodland stream. Hydrobiologia. 2011;659:93–109.
Doty SL. Endophytic yeasts: biology and applications. In: Symbiotic endophytes. Berlin, Heidelberg: Springer; 2013. p. 335–43.
Pirttilä AM, Pospiech H, Laukkanen H, Myllylä R, Hohtola A. Two endophytic fungi in different tissues of Scots pine buds (Pinus sylvestris L.). Microb Ecol. 2003;45:53–62.
Spribille T, Tuovinen V, Resl P, Vanderpool D, Wolinski H, Aime MC, et al. Basidiomycete yeasts in the cortex of ascomycete macrolichens. Science. 2016;353:488–92.
Yurkov A, Krüger D, Begerow D, Arnold N, Tarkka MT. Basidiomycetous yeasts from Boletales fruiting bodies and their interactions with the mycoparasite Sepedonium chrysospermum and the host fungus Paxillus. Microb Ecol. 2012;63:295–303.
Lazarus KL, Benny GL, Ho HM, Smith ME. Phylogenetic systematics of Syncephalis (Zoopagales, Zoopagomycotina), a genus of ubiquitous mycoparasites. Mycologia. 2017;109:333–49.
Põldmaa K, Samuels GJ. Aphyllophoricolous species of Hypomyces with KOH-negative perithecia. Mycologia. 1999;91:177–99.
Põldmaa K. Three species of Hypomyces growing on basidiomata of Stereaceae. Mycologia. 2003;95:921–33.
Reynolds NK, Benny GL, Ho HM, Hou YH, Crous PW, Smith ME. Phylogenetic and morphological analyses of the mycoparasitic genus Piptocephalis. Mycologia. 2019;111:54–68.
Spirin V, Nordén J, Svantesson S, Larsson KH. New records of intrahymenial heterobasidiomycetes (Basidiomycota) in north Europe. Nord J Bot. 2016;34:475–7.
Pacioni G, Leonardi M, Aimola P, Ragnelli AM, Rubini A, Paolocci F. Isolation and characterization of some mycelia inhabiting Tuber ascomata. Mycological Res. 2007;111:1450–60.
Hargreaves J, Brickle P, Van West P. The fungal ecology of seabird nesting sites in the Falkland Islands indicates a niche for mycoparasites. Fungal Ecol. 2018;36:99–108.
Siozios S, Tosi L, Ferrarini A, Ferrari A, Tononi P, Bellin D, et al. Transcriptional reprogramming of the mycoparasitic fungus Ampelomyces quisqualis during the powdery mildew host-induced germination. Phytopathology. 2015;105:199–209.
Spirin V, Malysheva V, Yurkov A, Miettinen O, Larsson KH. Studies in the Phaeotremella foliacea group (Tremellomycetes, Basidiomycota). Mycological Prog. 2018;17:451–66.
Tamm H, Põldmaa K. Diversity, host associations, and phylogeography of temperate aurofusarin-producing Hypomyces/Cladobotryum including causal agents of cobweb disease of cultivated mushrooms. Fungal Biol. 2013;117:348–67.
Gardes M, Bruns TD. ITS primers with enhanced specificity for basidiomycetes‐application to the identification of mycorrhizae and rusts. Mol Ecol. 1993;2:113–8.
Murray MG, Thompson WF. Rapid isolation of high molecular weight plant DNA. Nucleic Acids Res. 1980;8:4321–6.
White TJ, Bruns T, Lee SJWT, Taylor J. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In: PCR protocols: a guide to methods and applications, vol. 18. 1990. p. 315–22.
Ihrmark K, Bödeker I, Cruz-Martinez K, Friberg H, Kubartova A, Schenck J, et al. New primers to amplify the fungal ITS2 region–evaluation by 454-sequencing of artificial and natural communities. FEMS Microbiol Ecol. 2012;82:666–77.
Nikolenko SI, Korobeynikov AI, Alekseyev MA. BayesHammer: Bayesian clustering for error correction in single-cell sequencing. In: BMC genomics, vol. 14. BioMed Central; 2013. p. S7.
Zhang J, Kobert K, Flouri T, Stamatakis A. PEAR: a fast and accurate Illumina Paired-End reAd mergeR. Bioinformatics. 2014;30:614–20.
Gordon A., Hannon GJ. Fastx-toolkit. FASTQ/A short-reads preprocessing tools (unpublished). 2010. http://hannonlab.cshl.edu/fastx_toolkit, 5.
Rognes T, Flouri T, Nichols B, Quince C, Mahé F. VSEARCH: a versatile open source tool for metagenomics. PeerJ. 2016;4:e2584.
Hildebrand F, Tadeo R, Voigt AY, Bork P, Raes J. LotuS: an efficient and user-friendly OTU processing pipeline. Microbiome. 2014;2:30.
Martin M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J. 2011;17:10–2.
Nilsson RH, Veldre V, Hartmann M, Unterseher M, Amend A, Bergsten J, et al. An open source software package for automated extraction of ITS1 and ITS2 from fungal ITS sequences for use in high-throughput community assays and molecular ecology. Fungal Ecol. 2010;3:284–7.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods. 2010;7:335.
Kõljalg U, Larsson KH, Abarenkov K, Nilsson RH, Alexander IJ, Eberhardt U, et al. UNITE: a database providing web‐based methods for the molecular identification of ectomycorrhizal fungi. N Phytol. 2005;166:1063–8.
Frøslev TG, Kjøller R, Bruun HH, Ejrnæs R, Brunbjerg AK, Pietroni C, et al. Algorithm for post-clustering curation of DNA amplicon data yields reliable biodiversity estimates. Nat Commun. 2017;8:1–11.
Oksanen J, Kindt R, Legendre P, O’Hara B, Stevens MHH, Oksanen MJ, et al. The vegan package. Community Ecology Package. 2007;10:631–7.
Niemelä T. Polypores, lignicolous fungi. Norrlinia. 2005;13:1–320.
Nordén J, Penttilä R, Siitonen J, Tomppo E, Ovaskainen O. Specialist species of wood‐inhabiting fungi struggle while generalists thrive in fragmented boreal forests. J Ecol. 2013;101:701–12.
Ovaskainen O, Hottola J, Siitonen J. Modeling species co‐occurrence by multivariate logistic regression generates new hypotheses on fungal interactions. Ecology. 2010;91:2514–21.
Wickham H. Ggplot: using the grammar of graphics with R. 2009.
Edgar RC. MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinform. 2004;5:113.
Kumar S, Stecher G, Li M, Knyaz C, Tamura K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol. 2018;35:1547–9.
Liu H, Økland T, Halvorsen R, Gao J, Liu Q, Eilertsen O, et al. Gradients analyses of forests ground vegetation and its relationships to environmental variables in five subtropical forest areas, S and SW China. Sommerfeltia. 2008;32:3–196.
Rue H, Martino S, Chopin N. Approximate Bayesian inference for latent Gaussian models by using integrated nested Laplace approximations. J R Stat Soc: Ser B (Stat Methodol). 2009;71:319–92.
McMurdie PJ, Holmes S. phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PloS ONE. 2013;8:4.
De Caceres M, Jansen F, De Caceres MM. Package ‘indicspecies’. Relationship between species and groups of sites. R package version. 2016. 1(6).
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13:2498–504.
Deighton FC. Microfungi: some hyperparasitic hyphomycetes, and a note on Cercosporella Uredinophila Sacc. IV. Commonwealth Mycological Institute; 1969.
Poinar GO Jr, Buckley R. Evidence of mycoparasitism and hypermycoparasitism in Early Cretaceous amber. Mycological Res. 2007;111:503–6.
Boddy L, Hiscox J. Fungal ecology: principles and mechanisms of colonization and competition by saprotrophic fungi. (Eds Heitman J, Howlett B, Crous P, Stukenbrock E, James T & Gow NAR). The fungal kingdom. ASM Press, Washington, DC, 2017, p. 293–308.
Ovaskainen O, Schigel D, Ali-Kovero H, Auvinen P, Paulin L, Nordén B, et al. Combining high-throughput sequencing with fruit body surveys reveals contrasting life-history strategies in fungi. ISME J. 2013;7:1696–709.
Põldmaa K, Larsson E, Kõljalg U. Phylogenetic relationships in Hypomyces and allied genera, with emphasis on species growing on wood-decaying homobasidiomycetes. Can J Bot. 2000;77:1756–68.
Rogerson CT, Samuels GJ. Boleticolous species of Hypomyces. Mycologia. 1989;81:413–32.
Rogerson CT, Samuels GJ. Polyporicolous species of Hypomyces. Mycologia. 1993;85:213–72.
Rogerson CT, Samuels GJ. Agaricicolous species of Hypomyces. Mycologia. 1994;86:839–66.
Parratt SR, Laine AL. Pathogen dynamics under both bottom‐up host resistance and top‐down hyperparasite attack. J Appl Ecol. 2018;55:2976–85.
Jacobsen RM, Kauserud H, Sverdrup-Thygeson A, Bjorbækmo MM, Birkemoe T. Wood-inhabiting insects can function as targeted vectors for decomposer fungi. Fungal Ecol. 2017;29:76–84.
Grienke U, Zöll M, Peintner U, Rollinger JM. European medicinal polypores—a modern view on traditional uses. J Ethnopharmacol. 2014;154:564–83.
Meyer W, Walter GAMS. Delimitation of Umbelopsis (Mucorales, Umbelopsidaceae fam. nov.) based on ITS sequence and RFLP data. Mycol Res. 2003;107:339–50.
Hoff JA, Klopfenstein NB, McDonald GI, Tonn JR, Kim MS, Zambino PJ, et al. Fungal endophytes in woody roots of Douglas‐fir (Pseudotsuga menziesii) and ponderosa pine (Pinus ponderosa). For Pathol. 2004;34:255–71.
Foulongne-Oriol M, Rodier A, Savoie JM. Genetic architecture of crop production traits in the button mushroom Agaricus bisporus II: relationship between yield components and partial resistance to Lecanicillium fungicola. Appl Environ Microbiol. 2012;78:2435–42.
Kim M, Ahn C, Kim C. The fungicolous ascomycetes genus Hypomyces in Korea. Mycobiology. 2017;45:209–12.
Compant S, Saikkonen K, Mitter B, Campisano A, Mercado-Blanco J. Editorial special issue: soil, plants and endophytes. Plant Soil 2016;405:1–11.
Lugtenberg B, Kamilova F. Plant-growth-promoting rhizobacteria. Annu Rev Microbiol. 2009;63:541–56.
Saikkonen K, Young CA, Helander M, Schardl CL. Endophytic Epichloë species and their grass hosts: from evolution to applications. Plant Mol Biol. 2016;90:665–75.
Clay K, Holah J. Fungal endophyte symbiosis and plant diversity in successional fields. Science. 1999;285:1742–4.
Hume DE, Ryan GD, Gibert A, Helander M, Mirlohi A, Sabzalian MR. Epichloë Fungal endophytes for grassland ecosystems. In: Lichtfouse E, editor. Sustainable agriculture reviews, vol 19. Cham: Springer; 2016.
Jeffries P. Biology and ecology of mycoparasitism. Can J Bot. 1995;73:1284–90.
Bonfante P, Desirò A. Who lives in a fungus? The diversity, origins and functions of fungal endobacteria living in Mucoromycota. ISME J. 2017;11:1727–35.
Acknowledgements
The authors acknowledge Teppo Helo, Ari Meriruoko from Metsähallitus and Gergely Várkonyi from the Friendship Park Research Center in Kuhmo for help with organising fieldwork. Funding for this research was provided by a Miljøforsk Grant No. 254746 from the Research Council of Norway (SM, JN and HK), by NansenFondet (Grant to HK) and SM was partly supported by a Marie Curie FP7-PEOPLE-2013-IEF (Grant No. 628326). GA is grateful to École Normale Supérieure de Lyon for partly supporting his 4-month internship in 2018 at UiO. The authors acknowledge Lucie Zinger for constructive comments on preliminary results, and Lynne Boddy and two anonymous reviewers for improving the quality of the manuscript.
Author information
Authors and Affiliations
Contributions
SM, JN and HK designed the research and applied for funding. SM, JN, OM and HK organised sampling. SM and HK performed sampling and conceptualised the study. JN contributed in metadata. SM conducted molecular work and processed data. SM and GA did the bioinformatics analyses. SM performed statistical analyses with contributions from GA and SSB. SM prepared figures and wrote the manuscript with contributions from HK. All authors edited and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Maurice, S., Arnault, G., Nordén, J. et al. Fungal sporocarps house diverse and host-specific communities of fungicolous fungi. ISME J 15, 1445–1457 (2021). https://doi.org/10.1038/s41396-020-00862-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-020-00862-1
This article is cited by
-
Fungal and Bacterial Diversity in the Tuber magnatum Ecosystem and Microbiome
Microbial Ecology (2023)
-
Community composition and trophic mode diversity of fungi associated with fruiting body of medicinal Sanghuangporus vaninii
BMC Microbiology (2022)
-
Truffle species strongly shape their surrounding soil mycobiota in a Pinus armandii forest
Archives of Microbiology (2021)