Abstract
The marine ciliate Mesodinium rubrum is famous for its ability to acquire and exploit chloroplasts and other cell organelles from some cryptophyte algal species. We sequenced genomes and transcriptomes of free-swimming Teleaulax amphioxeia, as well as well-fed and starved M. rubrum in order to understand cellular processes upon sequestration under different prey and light conditions. From its prey, the ciliate acquires the ability to photosynthesize as well as the potential to metabolize several essential compounds including lysine, glycan, and vitamins that elucidate its specific prey dependency. M. rubrum does not express photosynthesis-related genes itself, but elicits considerable transcriptional control of the acquired cryptophyte organelles. This control is limited as light-dependent transcriptional changes found in free-swimming T. amphioxeia got lost after sequestration. We found strong transcriptional rewiring of the cryptophyte nucleus upon sequestration, where 35% of the T. amphioxeia genes were significantly differentially expressed within well-fed M. rubrum. Qualitatively, 68% of all genes expressed within well-fed M. rubrum originated from T. amphioxeia. Quantitatively, these genes contributed up to 48% to the global transcriptome in well-fed M. rubrum and down to 11% in starved M. rubrum. This tertiary endosymbiosis system functions for several weeks, when deprived of prey. After this point in time, the ciliate dies if not supplied with fresh prey cells. M. rubrum represents one evolutionary way of acquiring photosystems from its algal prey, and might represent a step on the evolutionary way towards a permanent tertiary endosymbiosis.
Similar content being viewed by others
Introduction
Endosymbiotic events have enabled eukaryotes to photosynthesize. More than a billion years ago, during a primary endosymbiosis event, a photosynthesizing cyanobacterium was retained by a non-plastidic unicellular eukaryote. Since then, chloroplasts have spread throughout the eukaryotic tree of life by secondary and tertiary endosymbiosis.
Teleaulax amphioxeia is an ecologically important, phototrophic marine unicellular eukaryote (protist) with a worldwide distribution [1]. It is 8–11 µm long and a member of the enigmatic group of cryptophytes, a group that is challenging to place in the evolutionary tree of life [2]. Most cryptophytes have permanent chloroplasts, originating from a secondary endosymbiosis event between a red alga and a phylogenetically distinct, non-photosynthetic host [3, 4]. Due to this origin, cryptophyte chloroplasts have a complex membrane topology with four membranes that enclose a nucleomorph between the outer two and the inner two membranes [5,6,7]. The nucleomorph is a highly reduced remnant of the endosymbiotic red algal nucleus. Cryptophytes hence possess DNA of different origin: red algal nuclear DNA in the nucleomorph, chloroplast DNA, cryptophyte mitochondrial DNA, and cryptophyte nuclear DNA [8].
Being primary producers, phototrophic cryptophytes are at the base of the marine food web, and grazed upon by heterotrophic and mixotrophic protists alike [9]. One of these grazers is Mesodinium rubrum, an abundant and ecologically important ciliate. M. rubrum is widely distributed in coastal ecosystems and known for causing nontoxic red tides [10,11,12]. Acquisition of phototrophy by retaining a chloroplast that originated from a secondary endosymbiosis event is regarded as a tertiary endosymbiosis [13, 14]. M. rubrum preys on cryptophytes belonging to the genera Geminigera, Teleaulax, and Plagioselmis. M. rubrum cells keep around 20 chloroplasts from its cryptophyte preys, and usually a single enlarged prey nucleus located close to the nuclei of the ciliate (ciliates have two macronuclei and one micronucleus) [15,16,17]. In order to sustain its maximum growth rate of ~0.5 per day, M. rubrum has to ingest ~one cryptophyte per day [18, 19]. M. rubrum covers typically >98% of its carbon need via photosynthesis at natural prey concentrations, and can replicate the acquired chloroplasts approximately four times after prey deprivation [16]. Eventually, the chloroplasts are degraded, and M. rubrum dies unless new cryptophyte prey cells are ingested [18,19,20,21]. Thus, this tertiary endosymbiosis between a cryptophyte and M. rubrum is not permanent and stable, but species-specific [22, 23].
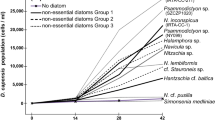
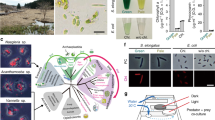
The regulation of cryptophyte genes within M. rubrum has previously been studied using RNA-seq, or Expressed Sequence Tags and microarray approaches [24, 25]. These studies found a remarkable cellular and metabolic chimerism between host and prey, and showed that M. rubrum not only sequesters the organelle machinery of its prey, but also the anabolic potential of the sequestered organelles [25]. Most cryptophyte genes involved in photosynthesis were upregulated after sequestration of the cryptophyte nucleus and chloroplasts into the ciliate [24]. However, previous studies had the challenge to distinguish between transcripts originating from M. rubrum and transcripts originating from the prey cryptophytes. We used genomic DNA (gDNA) data from free-swimming T. amphioxeia and prey-starved M. rubrum to overcome this problem. By screening for k-mers shared between gDNA reads and transcripts, we were able to assign transcripts to the right species by sequence signature. Using this approach, we could follow the transcriptional changes upon sequestration for cryptophyte and ciliate genes separately. We investigated changes in the level of T. amphioxeia genes expressed before and after ingestion by M. rubrum and compared those with starved M. rubrum cells that had lost the prey nucleus (Fig. 1). We explored changes in the regulation of the sequestered cryptophyte nuclei in response to changing light and time conditions (night, morning, and day) corresponding to darkness, 20 min after turning on the light, and full light, and focused for the first time on transcriptional changes of ciliate genes under different light conditions and prey availabilities.
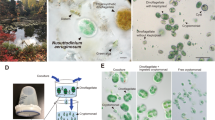
a Free-swimming T. amphioxeia with chloroplast, nucleomorph, mitochondrion, and nucleus. The outer membrane of the nucleus is connected to the outer membrane of the chloroplast. b Well-fed M. rubrum with two macronuclei, one micronucleus, and one enlarged cryptophyte nucleus. M. rubrum contains its own mitochondria, cryptophyte mitochondria, and cryptophyte chloroplasts that are arranged along the periphery of the cell. c Starved M. rubrum with two macronuclei, one micronucleus, and ciliate mitochondria. Starved M. rubrum was defined as cultures where at least 90% of cells had lost the cryptophyte nucleus. Well-fed cells of M. rubrum have one enlarged cryptophyte nucleus, which is always located in the center of the cell, termed CPN (centered prey nucleus) [24]. Well-fed cells might keep some extra prey nuclei in the periphery of the cell. Upon ciliate cell division, one of the two daughter cells receives the CPN, while in the other, one of the extra prey nuclei migrate close to the ciliate nuclei and enlarges [16]. Scale bar equals 5 µm in a, and 10 µm in b and c.
Materials and methods
Cultures
Cultures were established from single-cell isolates of Teleaulax amphioxeia (SCCAP K-1837, collected in Elsinore Harbor, Denmark; available at https://niva-cca.no/shop/cryptophyceae/teleaulax/k-1837), and Mesodinium rubrum (MBL-DK2009 collected in September 2009 in Elsinore Harbor, Denmark, available from PJH upon request). Cultures (T. amphioxeia, M. rubrum fed T. amphioxeia) were kept in triplicates and grown in glass bottles in F/2 medium at 15 °C in a light/dark cycle of 16/8 h with a light intensity of 100-µmol photons m−2s−1. During the exponential phase of growth, the ciliates were transferred to new media when cell concentrations reached 5000 ml−1 or more.
RNA extraction
For RNA extraction, cultures were harvested in full light (7 h into the light cycle), in darkness (6 h into dark cycle), and in the transition between dark and light (20 min into the light cycle). Cells of M. rubrum were harvested in a well-fed and a starved stage.
For the well-fed condition, we checked before extraction that no free cryptophyte cells remained in the medium (while counting M. rubrum cell numbers in a 1 mL Sedgewick Rafter Counting Chamber under a light microcope) and that at least 90% of all M. rubrum cells contained a cryptophyte nucleus. This was done by staining the nuclei with Hoechst reagent (#33342, Thermo Fisher Scientific, Waltham, USA), and checking 20 stained cells under a fluorescent microscope. Harvesting of starved cells was done ~4 weeks after the last cryptophytes had been seen in the culture. We confirmed the loss of cryptophyte nuclei by staining with Hoechst reagent and checking for prey nuclei under a fluorescence microscope. Cells were harvested after at least 90% of all M. rubrum cells had lost their cryptophyte nucleus. Cells were harvested by centrifugation in 10-ml glass tubes at 3220 rcf for 10 min (see Supplementary Table 1 for cell numbers in each harvest). Pellets were transferred to 1.5-mL LoBind Eppendorf tubes, and liquid nitrogen was directly added onto the pellets. The Eppendorf tubes were stored on ice without allowing the pellets to thaw until the lysis buffer was added. RNA was extracted using the column-based Exiqon Cell and Plant RNA Isolation Kit (#300110, Exiqon, Vedbæk, Denmark) following the “plant” protocol. In addition, a separate round of harvest has been transferred to hot Trizol and stored at −80 °C as backup. Two samples (10 and 11) from this backup have been used for RNA extraction using the Trizol method. Extracted RNA was stored at −80 °C until library preparation for sequencing.
DNA extraction
For DNA extraction T. amphioxeia cells as well as starved M. rubrum (fed T. amphioxeia) cells were harvested as described above and DNA extracted using a KingFisher Duo Prime System (#5400110, Thermo Fisher Scientific, Waltham, USA) using the Plant DNA Kit and following the manufacturers recommendations.
Library construction and sequencing
The RNA-seq libraries were mainly prepared using the MGIEasy RNA Library Prep Set (V1.0, MGI Tech) with 1-μg total RNA as input and sequenced on the BGISEQ-500RS platform using the PE100 chemistry according to the standard protocols provided by MGI Tech Co., Ltd (Shenzhen, China). The only exception was one of the three biological replicates of Tamp-day, of which the amount of total RNA was <1 µg and failed to meet the requirement of the MGI kit. The RNA-seq library of this sample was prepared using the TruSeq Stranded mRNA LT Sample Prep kit (RS-122–2101, Illumina) with 500-ng total RNA as input, and sequenced on the Illumina HiSeq 4000 platform using the PE100 chemistry, according to the standard Illumina protocols (San Diego, CA, USA).
The DNA sequencing libraries of free-swimming T. amphioxeia and starved M. rubrum were prepared using the MGIEasy DNA Library Prep Kit (V1.1, MGI Tech) with 1-μg gDNA as input, and sequenced on the BGISEQ-500RS platform using the PE100 chemistry according to the standard protocols provided by MGI Tech Co., Ltd (Shenzhen, China).
Quality control of raw sequencing data
Prior to subsequent analyses, all the DNA- and RNA-seq raw reads were passed to SOAPnuke (v1.5.3) [26] for quality control by removal of adapter-contaminated reads and low-quality reads. Specifically, all the DNA-seq data were filtered by SOAPnuke with parameters -n 0.02 -l 20 -q 0.3 -Q 2 -G -d. For the RNA-seq data, we generated two versions of clean data. The first one was used for de novo transcriptome assembly and generated by parameters -n 0.02 -l 20 -q 0.3 -p 1 -t 15,0,15,0 -Q 2 -G -d (i.e., removing adapter-contaminated, low-quality, and duplicated reads). The second version was used for gene-expression measurement and generated by parameters -n 0.02 -l 20 -q 0.3 -p 1 -t 15,0,15,0 -Q 2 -G (i.e., removing adapter-contaminated and low-quality reads but keeping duplicated reads). We kept the duplicated reads for gene-expression analysis, as it is impossible to discriminate real PCR-duplicates (i.e., reads created from the same cDNA molecule by PCR) from meaningful duplicated reads with identical sequences but from different cDNA molecules of the same transcript in our RNA-seq data [27].
Construction of reference gene sets for T. amphioxeia and M. rubrum
A hierarchical strategy was employed to construct the reference gene sets for T. amphioxeia and M. rubrum with the RNA-seq and DNA-seq clean data.
De novo transcriptome assembly for each RNA sample
The clean RNA reads (without duplicates) of each sample were first assembled using Trinity (v2.4.0) [28, 29] with parameters --min_contig_length 150 --min_kmer_cov 2 --min_glue 3. Then the highly similar sequences were clustered, and the redundant transcripts were removed from each transcriptome assembly using cd-hit-est (v4.6.8) [30, 31] with a sequence identity threshold of 0.95. Finally, the clean RNA reads (with duplicates) were aligned to each assembly to quantify the abundance of each gene defined by Trinity (i.e., a transcript cluster) using Salmon (v0.13.1) [32] with parameters --validateMappings -l IU –allowDovetail. The lowly expressed genes with TPM < 1 were removed from each transcriptome assembly, then only the longest transcript of each gene was kept. Detailed statistics for the transcriptome assembly for each RNA sample are available in Fig. 2 and Supplementary Table 2.
a Sampling strategy. b Analysis workflow. c Summary of the nonredundant reference gene sets constructed from the de novo transcriptome assembly. nt nucleotides, aa amino acids, ORF open reading frame, KEGG Kyoto Encyclopedia of Genes and Genomes, GO gene ontology. d Comparison of gene length and GC content for M. rubrum and T. amphioxeia genes, respectively.
Assignment of transcripts to T. amphioxeia and M. rubrum
Species assignment was conducted based on the average nucleotide identity (ANI) of each transcript in relation to the DNA sequences of T. amphioxeia and M. rubrum estimated by Mash (v2.1) [33, 34]. To eliminate potential prokaryotic contamination in the transcriptomes, we also built a data set containing 10,243,458 prokaryotic nucleotide sequences by extracting all Archaea and Bacteria sequences from the NCBI nt database (release 20190315). Specifically, we sketched at most 5000 nonredundant 21-mers from each transcript, and compared them with the nonredundant 21-mers generated from all of the T. amphioxeia DNA reads, M. rubrum DNA reads, and the prokaryotic nucleotide sequences, respectively, to estimate ANI of T. amphioxeia (ANItamp), M. rubrum (ANImrub), and prokaryotic (ANIprok) for each transcript. Of note, even so, more than 90% of the cells in the starved M. rubrum cultures had lost the cryptophyte nucleus, genomic sequences of T. amphioxeia were still detectable in the starved M. rubrum DNA reads (although occurring at low abundances). Considering this, a transcript was assigned to: (1) T. amphioxeia if ANItamp ≥ 0.95 and ANIprok < 0.95; (2) M. rubrum if ANItamp < 0.95, ANImrub ≥ 0.95 and ANIprok < 0.95; (3) unknown sequences for other conditions (Supplementary Table 2). The unknown sequences were discarded from subsequent analyses.
Hierarchical removal of redundant transcripts
The reference gene sets of T. amphioxeia and M. rubrum were generated by combining T. amphioxeia and M. rubrum transcripts from all the samples, respectively. The highly similar sequences were clustered, and the redundant transcripts for the same species were removed using cd-hit-est (v4.6.8) [30, 31] with a sequence identity threshold of 0.95. To further remove redundant transcripts that failed to be clustered by cd-hit-est (caused by alternative splicing such as exon skipping, intron retention, etc.), two rounds of Minimap2 spliced alignment and MCL clustering processes were performed. Specifically, pairwise spliced alignment for all transcripts was generated using the all-vs-all mode of Minimap2 (v2.10-r764) [35] with parameters -aX -x splice. Then, a graph was built with the best alignment of each query that satisfied identity >0.95 and coverage >70% against the shorter sequence. The vertices of the graph were the transcripts and the edges were weighted by identity x coverage. Next, the graph was inputted to Markov Clustering algorithm (mcl v14–137) [36] to cluster similar sequences with the default power and inflation setting, and the longest transcript of each cluster was kept as the representative. The Minimap-MCL process ran iteratively based on the cluster representatives generated by the last iteration until no more pairwise alignments satisfying the threshold were found. Then a second round of the Minimap-MCL process was performed with the threshold identity >0.98 and coverage of the shorter sequence >50% to generate the final nonredundant T. amphioxeia and M. rubrum reference gene sets. The longest ORF for each gene was detected by TransDecoder (v5.5.0) [29] with parameters -m 50 --genetic_code universal for the T. amphioxeia genes and -m 50 --genetic_code Mesodinium for the M. rubrum genes.
Estimation of the representativeness of the reference gene sets
To evaluate the representativeness of the reference gene sets, we aligned the RNA clean reads (with duplicates) from each of the T. amphioxeia samples to the T. amphioxeia genes, and aligned the reads from each of the M. rubrum samples to the collection of T. amphioxeia and M. rubrum genes using BWA-MEM (v0.7.16) [37] with default parameters. Then, the numbers and ratios of reads being aligned, being uniquely aligned (as defined by mapping quality ≥ 30), and being aligned in proper pairs, in relative to the total numbers of inputted reads, were counted by samtools flagstat (SAMtools v1.7) [38, 39]. The completeness of the gene sets was assessed by the Benchmarking Universal Single-Copy Orthologs (BUSCO v3) [40] with the eukaryota odb9 database (Supplementary Table 3).
Identification of T. amphioxeia genes present in prey-starved M. rubrum DNA data
To determine whether the whole prey nuclei were kept inside M. rubrum (Fig. 3a), DNA read alignment and alignment-free k-mer screening methods were used to identify T. amphioxeia genes in the two starved M. rubrum DNA samples. For the alignment-based method, we aligned the DNA reads of the two starved M. rubrum DNA samples to the T. amphioxeia gene set using BWA-MEM (v0.7.16) [37] and calculated the coverage of each gene using genomeCoverageBed (v2.26.0) [41] with default parameters. T. amphioxeia genes with coverage larger than 70% were considered present in the starved M. rubrum cells. The alignment-free k-mer screening method was described in the above section “Assignment of transcripts to T. amphioxeia and M. rubrum.” The T. amphioxeia genes present in the starved M. rubrum DNA samples were defined as ANItamp ≥ 0.95 and ANIprok < 0.95.
Expression-level quantification
We aligned the RNA clean reads of each sample to the database containing all T. amphioxeia and M. rubrum genes and quantified the abundance of each gene using Salmon (v0.13.1) [32] with parameters --validateMappings -l IU --allowDovetail. Of note, more than 99.98% of the aligned reads from the free-swimming T. amphioxeia samples were mapped to the T. amphioxeia genes by Salmon, highlighting the reliability of our species assignment process described above. There was some disparity in the amount of data originating from T. amphioxeia across different sample groups. To eliminate this disparity for subsequent analyses, we downsampled the clean data of each sample to adjust the number of T. amphioxeia aligned reads in each sample to at most ~40 M. The raw counts of the T. amphioxeia genes in all the 27 samples were collected (Supplementary Table 4). Then, we balanced the data amount originating from M. rubrum across all M. rubrum samples in the same way and generated the raw counts of the M. rubrum genes in all the 18 M. rubrum samples (Supplementary Table 4).
Transcriptome features of M. rubrum
We investigated actively transcribed T. amphioxeia and M. rubrum genes across different sample groups to reveal the transcriptome features of M. rubrum. Genes with TPM ≥ 1 were deemed to be actively transcribed. The TPM of a given gene i was calculated as follows: TPMi = 106 * (Countsi/EffectiveLengthi)/Σj (Countsj/EffectiveLengthj). We performed two separate analyses: (1) the proportion of actively transcribed T. amphioxeia genes before and after sequestration (Fig. 3b), and (2) the relative contribution of actively transcribed T. amphioxeia and M. rubrum genes to the global M. rubrum transcriptomes (Fig. 3c, d). The two analyses used different scaling factors Σj (Countsj/EffectiveLengthj) for TPM calculation. Analysis 1 used the total abundance of the T. amphioxeia genes. Analysis 2 included the total abundance of all T. amphioxeia and M. rubrum genes. For analysis 1, we collected EffectiveLength and Count of each T. amphioxeia gene in each sample from the Salmon output and recalculated the TPMs of T. amphioxeia genes with the above formula. Then we measured the proportion of actively transcribed T. amphioxeia genes in all samples. For analysis 2, the TPM matrices output by Salmon was used directly to identify actively transcribed genes (NumGenesTamp vs. NumGenesMrub) and accumulated abundance of those genes (ΣTPMTamp vs. ΣTPMMrub) in the M. rubrum samples. The proportion of actively transcribed M. rubrum genes at different prey availabilities (Fig. S1) was counted in the same manner as Fig. 3b.
Identification of differentially expressed genes (DEGs)
The DEG candidates between sample groups were detected by DESeq2 (v1.22.0) [42]. First, the library sizes across samples were normalized using the default median ratio method, taking the raw counts as input. Next, to obtain dispersion estimates, the type of fitting of dispersions to the mean intensity was set to be parametric. Then, we used Wald significance tests (nbinomWaldTest) for model fitting and test statistics. The Benjamini–Hochberg false discovery rate (FDR) was employed to adjust p value for multiple test correction. Finally, the significant DEGs were defined with the criteria of basemean ≥20, adjusted p value < 0.01, and |log2 (fold change)| > 1.5 (|log2 (fold change)| > 1.5 and |log2 (fold change)| > 1 were both used to detect significant M. rubrum DEGs).
Principal component analysis (PCA) clustering
We used the normalized counts as described in the above DEG section to perform PCA clustering. We filtered the gene counts with narrow variance (standard deviation of normalized count < 10 across all samples) and then variance stabilizing transformation (vst provided by DESeq2) transformed the normalized counts with parameter blind = FALSE. Finally, we generated the PCA plots with the vst-transformed matrices using plotPCA function provided by DESeq2.
Identification of co-expression modules
Co-expression network analyses were performed using the weighted gene correlation network analyses (WGCNA) R package (WGCNA v1.68) [43]. After filtering lowly expressed genes (mean normalized count <10 across all samples), the normalized read count matrices were vst-transformed with parameter blind = FALSE. Then the vst-transformed matrices were passed to the blockwiseModules function implemented in the WGCNA package for identification of the signed co-expression modules with parameters maxBlockSize = 50,000, networkType = signed, minModuleSize = 100, minKMEtoStay = 0.6, minCoreKME = 0.5, mergeCutHeight = 0.15, numericLabels = TRUE, pamRespectsDendro = FALSE, and power = 16 for the T. amphioxeia genes and power = 18 for the M. rubrum genes.
Functional annotation and enrichment analysis
We aligned the gene sequences to the UniProtKB/Swiss-Prot+TrEMBL database (release 20190408) with BLASTX (blast-2.2.26) using parameters -F F -e 1e-5. The best hit of each query was retained based on the BLASTX bit score. The GO annotation of the best-aligned UniProt protein was then assigned to the query gene. To determine what pathways the genes might be involved in, gene sequences were searched against the KEGG database (v89.1) [44] with BLASTX (blast-2.2.26) using parameters -F F -e 1e-5. The best hit of each query was retained based on the BLASTX bit score.
A set of better quality and reliable functional annotations was generated (the BLASTX hits with e value lower than 1e−10) for the functional enrichment analyses. Hypergeometric tests were employed to examine whether a list of DEGs was enriched in a specific GO term in relation to background genes as previously described [45], by comparing the number of target DEGs annotated to this GO term, the number of target DEGs not annotated to this GO term, the number of background genes (i.e., all the T. amphioxeia or M. rubrum genes excluding the target DEGs) annotated to this GO term, and the number of background genes not annotated to this GO term. P values were adjusted for multiple testing by applying FDR [46], and enriched GO terms were considered significant for adjusted p value < 0.05. The GO enrichment results were visualized with EnrichmentMap (v3.2.1) [47] in Cytoscape (v3.7.2) [48].
KEGG enrichment analysis was done using the same principle as the GO enrichment. The regulation of gene expression involved in the enriched KEGG pathways was visualized with Pathview (v1.23.3) [49].
Identification of pathways absent in M. rubrum but expressed by the sequestered prey nucleus
The BLASTX hits against the KEGG database with e value lower than 1e−10 were used to compare the biological process compositions of the host M. rubrum and its prey T. amphioxeia in this endosymbiotic system. Specifically, we first manually excluded the KEGG pathways that are clearly inapplicable in unicellular eukaryotes (i.e., pathways involved in disease or cancer genesis, nervous and immune systems). Next, we calculated the KO assignment rate of each pathway in M. rubrum and T. amphioxeia, respectively, and measured the lowest KO assignment rate that covered 95% of all assigned KOs in each species. Then, the background expectation of the overall KO assignment rate in a given species was estimated to be the proportion of nonredundant assigned KOs out of the total KOs in the pathways above the lowest KO assignment rates. After that, we employed binomial tests to examine whether the observed number of assigned KOs in a pathway was significantly lower than the background expectation based on the binomial distribution B(k, n, p), where k was the observed number of assigned KOs in a given pathway, n was the total number of KOs in this pathway, and p was the background expectation of overall KO assignment rate. P values were then adjusted by the Benjamini–Hochberg FDR procedure. We finally required that a pathway with adjusted p value < 0.05 in M. rubrum but > 0.1 in T. amphioxeia to be present in the prey but absent in the host. We also extracted and counted the KO assignments of pathways present in the model ciliates Paramecium tetraurelia and Tetrahymena thermophile directly from the KEGG database for reference.
Results and discussion
Transcriptomic profiles and reference gene set constructions for Teleaulax amphioxeia and Mesodinium rubrum
We performed RNA-seq on cultures of free-swimming T. amphioxeia, M. rubrum well-fed, and M. rubrum prey-starved for more than 4 weeks (i.e., more than 90% of the cells in the M. rubrum culture had lost the central cryptophyte nucleus) (Fig. 1). Each culture was sampled at three time points during the light–dark cycle: night (6 h after the light was switched-off), morning (20 min after the light was switched-on), and day (7 h after the light was switched-on) (Fig. 2a). Biological triplicates were collected for each condition, and an average of 184 million reads were generated for each biological replicate (Supplementary Table 5). To accurately discriminate the species origin of each assembled transcript, we also performed genome sequencing for DNA extracted from free-swimming T. amphioxeia and starved M. rubrum, respectively (Fig. 2a and Supplementary Table 5).
As no reference nuclear genomes were available for T. amphioxeia and M. rubrum, we de novo assembled the transcriptome of each sample separately in a first step, followed by stepwise combining the transcripts assembled from each sample. The species identity of each transcript was determined by screening the k-mers shared between gDNA reads and transcript sequences (Fig. 2b and Supplementary Table 2). This allowed us to identify 72,061 and 22,250 nonredundant transcripts (i.e., genes) as T. amphioxeia and M. rubrum origin, respectively (Fig. 2c). To assess the representativeness of the reference gene sets, we aligned the RNA-seq reads from free-swimming T. amphioxeia samples to the 72,061 T. amphioxeia genes, and aligned the reads from M. rubrum samples to the collection of 72,061 T. amphioxeia and 22,250 M. rubrum genes (note: M. rubrum samples transcribed genes from both the host and prey genomes). On average, 97.7% of the reads (ranging 96.2–98.6%) could be mapped back to the reference gene sets, 95.7% (ranging 93.1–97.2%) were aligned in proper pairs, and 88.0% (ranging 84.9–89.6%) had mapping quality ≥30 (Supplementary Table 6), demonstrating that most sequences in the transcriptomes are present uniquely in the two reference gene sets. We also aligned the RNA-seq reads from free-swimming T. amphioxeia samples to the collection of T. amphioxeia and M. rubrum genes, and observed <0.02% of the aligned reads being mistakenly mapped to M. rubrum genes, highlighting the reliability of our DNA-based species assignment process.
We annotated 62.9% of T. amphioxeia and 50.6% of M. rubrum genes by searching against different functional databases (Fig. 2c and Supplementary Table 7). Interestingly, the GC content of T. amphioxeia genes was around 59%, thus considerably higher than the GC content of M. rubrum with 35% (Fig. 2c, d). This GC deviation further supports that the genes were assigned to the right species.
M. rubrum keeps all the genetic material and transcribes most genes from the acquired cryptophyte nuclei
By searching for the T. amphioxeia genes in the M. rubrum gDNA sequence reads, we retrieved almost all (97.3–99.9%) of the 72,061 T. amphioxeia genes in the two starved M. rubrum DNA samples (Fig. 3a; see “Materials and methods”), suggesting that M. rubrum keeps all the genetic material from the acquired cryptophyte nuclei. Next, we examined the transcriptional activity of the T. amphioxeia nuclei upon sequestration by M. rubrum. Gene-expression measurement indicated that on average 63% of the T. amphioxeia genes were actively transcribed (TPM ≥ 1) inside M. rubrum at some time-point during the sampling cycle, which comprised 82% of the T. amphioxeia genes, when considering all sampling points together (Fig. 3b). Even though these ratios were lower than those observed in the free-swimming T. amphioxeia samples (77% in average and 96% in combination), they did indicate that the majority of T. amphioxeia genes (82%) were actively transcribed inside the M. rubrum cells (Fig. 3b). At the same time, an average of 90% of the M. rubrum genes was actively transcribed regardless of light and prey availability (Fig. S1).
a Percentage of T. amphioxeia genes identified in the starved M. rubrum DNA data by read alignment and k-mer screening methods. b Proportion of actively transcribed T. amphioxeia genes before and after sequestration. c Global transcriptome of M. rubrum with proportion of contributing T. amphioxeia and M. rubrum genes. d Global transcriptome of M. rubrum with proportion of transcript abundance originating from T. amphioxeia or M. rubrum.
Up to 68.4 ± 1.2% of the genes transcribed within the well-fed M. rubrum cells originated from T. amphioxeia. This proportion was maintained at 50.4 ± 1.3% for the starved M. rubrum samples (Fig. 3c). The contribution of cryptophyte genes to the global transcriptome of well-fed M. rubrum in the present study (68.4 ± 1.2%) is higher than previous estimates (13.5% in [24], 58–62% in [25]). However, when taking the transcriptional abundance of each gene into account, the contribution of T. amphioxeia transcripts to the global M. rubrum transcriptomes was much lower, ranging from 47.5 ± 1.3% (well-fed morning) to 10.2 ± 0.9% (starved night) (Fig. 3d). Thus, despite the fact that most T. amphioxeia genes were transcribed inside M. rubrum, the gene products from M. rubrum dominated the mRNA pools of the host cells even in well-fed cells. In a well-integrated endosymbiotic system, one would expect to find a lower qualitative expression of endosymbiont genes: only genes that are beneficial to the host will be expressed, while genes not needed by the host will suffer depletion. Given that 82% of all T. amphioxeia genes were expressed at some time-point within M. rubrum, it is likely that many of the T. amphioxeia transcripts are not photosynthesis-related.
Cryptophyte nuclei presents dramatic transcriptional rewiring upon sequestration
PCA with the T. amphioxeia gene-expression matrix separated all the 27 samples into three distinct clusters of free-swimming T. amphioxeia, well-fed M. rubrum, and prey-starved M. rubrum (Fig. 4a), with the distance separating free-swimming T. amphioxeia samples from all M. rubrum samples being larger than the distance separating well-fed and starved M. rubrum groups. This suggests that the condition of sequestration alone induced an overwhelming amount of transcriptional changes when compared with other experimental conditions for the T. amphioxeia genes. Consistently, this was supported also by WGCNA using the same matrix after filtration for lowly expressed genes (44,241 T. amphioxeia genes with mean normalized count ≥10), which identified six modules. The first two modules comprised 81% of the input genes and enriched T. amphioxeia genes that were prevailingly down- and upregulated after sequestration by M. rubrum, respectively (Fig. S2). More specifically, differential gene-expression analyses between the three sample groups revealed by PCA showed that 34.8% and 31.9% of T. amphioxeia genes were significantly differentially expressed (|log2FC| > 1.5 and FDR < 0.01) upon sequestration in well-fed and prey-starved M. rubrum, respectively (Fig. 4b). All these results consistently emphasize that a profound transcriptional rewiring occurs in the T. amphioxeia nuclei after sequestration by M. rubrum, as reported previously [24, 25].
a Principal component analysis of T. amphioxeia genes show a clear segregation between free-swimming, well-fed, and starved samples. b Amount of significantly differentially expressed genes (|log2FC| > 1.5 and FDR < 0.01) upon sequestration. c GO enrichment results for T. amphioxeia genes up-/downregulated after sequestration in well-fed samples visualized as an enrichment map. Nodes represent enriched gene-sets and edges represent mutual overlap between gene sets, thus clustering highly redundant gene-sets. d Changes in T. amphioxeia gene expression in the eukaryote replication complex pathway. Left part of each box shows log2 fold change in gene expression for free-swimming vs. well-fed samples. Right part of each box shows log2 fold change in gene expression for free-swimming vs. starved samples.
Functional enrichment analyses for the sequestration-induced DEGs revealed that T. amphioxeia genes related to ion transmembrane transport, signal transduction, cell motility, and regulation of metabolic processes were downregulated. On the other hand, upregulation after sequestration was observed for genes involved in photosynthesis, RNA processing, DNA replication and repair, lipid and protein metabolism, and metabolism of diverse compounds (e.g., nucleic acid, carbohydrate, amino acid, carboxylic acid, and pigment) (Fig. 4c and Supplementary Tables 8, 9). These results were generally consistent with previous observations by Kim et al. [24] and Lasek-Nesselquist et al. [25]. Interestingly, we also found that the upregulated DEGs were enriched in DNA replication, repair and recombination, and cell cycle. These comprised T. amphioxeia genes encoding cyclins (e.g., CycA, CycH), cyclin-dependent kinases (e.g., CDK2, CDK7), cell division control proteins (e.g., Cdc6, Cdc7, Cdc45), and almost all the genes involved in the eukaryotic replication complex (Figs. 4d and S3). This suggests that the sequestered nuclei are able to replicate their DNA. In contrast to Lasek-Nesselquist et al. [25], we did not observe downregulation of genes involved in protein processing pathways (Fig. S4). On the contrary, many genes involved in endoplasmic reticulum membrane and mRNA surveillance pathway were upregulated after sequestration (Figs. S4 and S5). This implies that the sequestered prey nuclei play an active regulatory role in transcription, translation, and also in transportation of T. amphioxeia gene products.
Of note, only few T. amphioxeia genes (927) were identified as DEGs between well-fed and prey-starved M. rubrum samples (Fig. 4b), and the majority of DEGs were shared between free-swimming-vs.-inside well-fed M. rubrum and free-swimming-vs.-inside starved M. rubrum (Fig. S6). This demonstrates that the global transcriptional patterns of the T. amphioxeia nuclei inside well-fed M. rubrum cells were highly similar with those inside starved M. rubrum cells.
These findings are unexpected as most of the starved cells had lost their prey nuclei; <10% of the starved M. rubrum cells had preserved the enlarged centered prey nucleus (CPN). A typical well-fed M. rubrum cell has about 20 chloroplasts and a single enlarged CPN, that is located at more or less the same position anterior to the two macronuclei within M. rubrum [16]. With each chloroplast, M. rubrum takes up one cryptophyte nucleus. Well-fed M. rubrum cells can contain multiple prey nuclei, i.e., the CPN and some extra prey nuclei that are kept in the periphery of the cell [16]. The finding that well-fed and starved cells have similar gene-expression patterns suggests that only the CPN is actively transcribed inside M. rubrum. Otherwise, the multiple periphery nuclei have to perform a somehow concerted gene transcription with the CPN inside well-fed M. rubrum cells.
Experimental evidence suggests that chloroplasts can divide within M. rubrum without the presence of cryptophyte nuclei [16, 18]. It is also known that photosynthesis in M. rubrum is related to the percentage of cells with a CPN, not to the number of chloroplasts [16]. Starved M. rubrum cells that have lost the CPN will usually have some chloroplasts remaining in the cell. A reason why those chloroplasts can survive within M. rubrum might be that they are particularly robust, with a comparatively large gene set [50, 51]. It is possible that the nucleomorph plays a crucial role in enabling the T. amphioxeia chloroplasts to divide within M. rubrum and renders it a favored prey in comparison to chloroplasts with smaller gene sets [52].
Cryptophyte responses to light and time changes get lost upon sequestration by M. rubrum
Free-swimming T. amphioxeia is expected to adjust its gene-expression pattern according to light and time changes during night, morning, and day as any other photosynthetic organism with permanent chloroplasts. This was confirmed by the PCA result (Fig. 4a). To get a closer look at the light- and time-dependent transcriptional responses of the cryptophyte genes before and after sequestration, we conducted pairwise correlation analyses of the free-swimming T. amphioxeia samples and the M. rubrum samples, respectively. While relatively low correlations were observed among T. amphioxeia samples from different light and time conditions (Fig. 5a), we found all the M. rubrum samples showing consistently high pairwise correlations (Fig. 5b), implying that the cryptophyte nuclei had lost the ability to adjust gene expression according to light and time changes upon sequestration. This conclusion is further confirmed by the overwhelming amount of dark/light-responding DEGs (10,828; |log2FC| > 1.5 and FDR < 0.01) identified in free-swimming T. amphioxeia samples in comparison to those identified in M. rubrum samples (157; Fig. 5c).
a Pearson correlation analysis of T. amphioxeia genes among different samples show differences according to time and light condition. b Pearson correlation analysis of T. amphioxeia genes after sequestration by M. rubrum reveals an expression pattern that is independent of light and prey availability. c Amount of T. amphioxeia genes that were differentially expressed according to time and light condition in free-swimming cells and after sequestration by M. rubrum. d Small panels show expression fold change of light-dependent T. amphioxeia DEGs at night, night-versus-morning, and night-versus-day in free-swimming, inside well-fed M. rubrum and inside starved M. rubrum condition. The heat map shows T. amphioxeia genes that got differentially expressed according to time and light condition before sequestration by M. rubrum but maintained at high-expression levels at night after sequestration.
T. amphioxeia genes, significant differentially expressed according to light and time changes, were functionally enriched in photosynthesis, oxidative phosphorylation, glycolysis and circadian entrainment-related pathways (Supplementary Table 10), consistent with the expectation for a free-living photosynthetic organism. Of note, T. amphioxeia genes involved in circadian entrainment were generally downregulated upon sequestration (Supplementary Table 9). This might partly account for the loss of time/light response of the cryptophyte nuclei upon sequestration. The expression of many genes in free-swimming T. amphioxeia responded to light (i.e., DEGs upregulated in morning and day versus night). After sequestration, these light-responding genes maintained high-expression levels at night. Including DEGs encoding for light-harvesting complex and light reaction of photosynthesis, which are responsible for harvesting and transferring light energy and obviously not needed at night (Fig. 5d and Supplementary Table 11). The loss of the dark/light response together with the over-expression of potentially undesired genes strongly suggests that M. rubrum can elicit only one expression pattern out of its acquired cryptophyte nucleus regardless of light condition and prey availability (i.e., the number of acquired prey nuclei).
Interestingly, in a different system, the Antarctic Ross Sea dinoflagellate acquires transient chloroplasts from haptophyte prey, and the expression of kleptoplast-targeted genes is also unaffected by environmental parameters such as light [53]. The cryptophyte Chroomonas sp., which gets sequestered by the dinoflagellate Nusuttodinium aeruginosum, also loses its transcriptional responses to light after sequestration [54].
During evolution, foreign chloroplasts have ended up in other protists in many different ways [13]. In some protists, intact endosymbionts are well integrated into host cells [55]. Other protists reduce ingested algal cells, and keep prey nuclei as well as other cell organelles beside the chloroplasts (like M. rubrum). Yet, other protists retain exclusively the chloroplasts for shorter, (i.e., the ciliate Strombidium) or longer time (i.e., the dinoflagellate Dinophysis) [56, 57]. This can be interpreted as evolutionary steps toward permanent endosymbiosis. In a first step, a prey cell is taken up by a host and not digested. In a second step, the host gets some control over the gene expression of the acquired cell via the ingested prey nuclei—that is where M. rubrum is right now. In a third step, only the chloroplasts are retained, but need to be replaced with time (i.e. Dinophysis [58]). In the final step, the genes from the host and the acquired cell (or organelles such as chloroplasts and nuclei) need to align in order to fine-tune the gene expression according to environmental conditions. Whether or not M. rubrum is on its way toward a permanent tertiary endosymbiosis is speculative. Such a step will depend on the ability of M. rubrum to divide and keep the sequestered prey nuclei permanently, or rely on gene transfer from the algal prey to the ciliate nuclei.
M. rubrum fine-tunes gene expression in response to prey availability and upregulates genes involved in transport when well-fed
The problem that M. rubrum faces is that it has to deal with different genetic codes. Ciliates show deviations in the genetic code and it has been suggested that these deviations have occurred multiple times independently [59]. M. rubrum uses a genetic code that is different from cryptophytes and other eukaryotes, for instance it translates UAA and UAG into tyrosine and not into STOP codons [60]. By retaining organelles from its cryptophyte prey, M. rubrum can use the prey nucleus to serve the chloroplast gene products using the standard code. Given that this is possible for several cryptophytes such as Teleaulax amphioxeia, T. acuta, and Geminigera cryophila (TPG clade), the question remains as to why only these taxa and no other cryptophytes apparently can be exploited [15]. M. rubrum is known to feed on cryptophyte species belonging to different clades, but cannot utilize them for growth and photosynthesis, with the exception of the TPG clade [15, 61].
The construction of reference gene sets separately for M. rubrum and T. amphioxeia allowed us to compare the gene compositions of the host and its prey in this endosymbiotic system. By mapping the M. rubrum and T. amphioxeia genes to KEGG pathways, we found that the majority of pathways were present in both species, whereas M. rubrum lacked genes involved in photosynthesis, antenna proteins for photosynthesis, and carotenoid biosynthesis (Fig. 6a and Supplementary Table 12), although we could not completely rule out the possibility that these genes were presented in the M. rubrum genome but not expressed. Interestingly, we also found that the pathways related to the biosynthesis or metabolism of some essential compounds such as lysine, glycan, and several kinds of vitamins (pantothenate, riboflavin, and biotin) were absent or non-expressed in M. rubrum (Fig. 6a), indicating that M. rubrum has to obtain the metabolic potential for these compounds from the prey. This explains its dependency on grazing. Of note, most of these M. rubrum-absent pathways were actually upregulated in T. amphioxeia after sequestration (Fig. 6a), implicating that M. rubrum is able to obtain these nutrients from its cryptophyte prey without digesting the acquired organelles, which is a critical step toward a permanent endosymbiosis. It can be speculated that M. rubrum has lost the ability to biosynthesize some essential compounds due to the symbiotic relationship with T. amphioxeia. These micronutrients might be a reason why M. rubrum sequesters exclusively species of the TPG clade.
a Comparison of the presence of genes from selected pathways in the ciliates M. rubrum, Paramecium tetraurelia, and Tetrahymena thermophile and the cryptophyte T. amphioxeia with differential expression of T. amphioxeia genes upon sequestration (free-swimming versus well-fed), showing services provided by T. amphioxeia to M. rubrum. b Principal component analysis of M. rubrum genes. c Barplot showing the amount of significantly differentially expressed M. rubrum genes (|log2FC| > 1.5 or |log2FC| > 1) according to prey and light conditions. d GO enrichment analysis of M. rubrum genes upregulated in the well-fed samples visualized as an enrichment map. e Heat map showing the differential expression (up/down fold change > 1.5) of active and passive transmembrane transporters in well-fed and starved M. rubrum cells.
Next, we investigated the transcriptional changes of M. rubrum genes in response to different light conditions and prey availability (well-fed or prey-starved). PCA with the M. rubrum gene expression matrix revealed that the M. rubrum samples were clustered according to light condition as well as according to prey availability (Fig. 6b). The response to the supply of prey was stronger than the response to light (Fig. 6b). This finding was confirmed by a WGCNA analysis, which uncovered ten co-expression modules. The two largest modules comprised up to 35% of the input genes and enriched M. rubrum genes that were prevailingly down- and upregulated after starvation (Fig. S7). However, DEG analyses using the same cutoff as the T. amphioxeia genes (|log2FC| > 1.5 and FDR < 0.01), or even lower cutoff (|log2FC| > 1 and FDR < 0.01), only identified a small number of genes as DEGs between well-fed and prey-starved M. rubrum samples (Fig. 6c). Actually, the negligible number of DEGs identified between samples from different light conditions suggests that M. rubrum is not sensitive to light changes, and that the few responses to light take comparatively long time (Fig. 6b, c).
To uncover the functional preference of M. rubrum genes in response to prey availability, we conducted functional enrichment analysis for genes clustered in the two largest co-expression modules of the WGCNA analysis. Module 1 comprised 3285 genes that got downregulated upon prey starvation (Fig. S7). Module 2 contained 2387 M. rubrum genes that got upregulated upon starvation (Fig. S7). Genes downregulated upon prey starvation (upregulated in well-fed condition) were enriched in small molecule metabolism, ribonucleoside metabolism, and transmembrane transport (Fig. 6d). Interestingly, genes related to active transmembrane transporter activity (GO:0022804; adjusted p = 0.001) were enriched in module 1 (Supplementary Table 13). This indicates that M. rubrum-derived active transmembrane transporters play an important role in well-fed M. rubrum (Fig. 6e). In contrast, passive transmembrane transporter activity is more prominent in prey-starved M. rubrum (Fig. 6e), indicating that well-fed M. rubrum is transporting molecules among different cell compartments and actively coordinating biological processes of itself and the cryptophyte prey within the cell.
Conclusions
We found very strong transcriptional changes of T. amphioxeia genes after sequestration by M. rubrum. Upregulated prey genes were related to photosynthesis and metabolism, as well as biosynthesis of lysine and glycan, several kinds of vitamins and gene replication. These processes provide a gain for the host, M. rubrum, and demonstrate its prey dependency. Light-dependent transcriptional regulation of T. amphioxeia genes found in free-swimming condition got lost upon sequestration. The transcriptional pattern of T. amphioxeia genes in well-fed and prey-starved M. rubrum was highly similar, indicating that M. rubrum can only induce the expression of one particular pattern out of the acquired prey nucleus. M. rubrum shows only very few adjustments in its gene expression in response to different light conditions. Noticeable is the upregulation of active transmembrane transporters in well-fed M. rubrum and the role of passive transmembrane transporters in starved M. rubrum.
Data availability
The raw sequencing reads produced in this study are deposited in the CNGB Nucleotide Sequence Archive (CNSA) [62, 63] with accession number CNP0000925. The nucleotide sequences and functional annotations of the reference gene sets for Teleaulax amphioxeia and Mesodinium rubrum are deposited in the figshare repository under the link 10.6084/m9.figshare.12360836.
Change history
24 September 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41396-021-01115-5
References
Altenburger A, Blossom HE, Garcia-Cuetos L, Jakobsen HH, Carstensen J, Lundholm N, et al. Dimorphism in cryptophytes—the case of Teleaulax amphioxeia/Plagioselmis prolonga and its ecological implications. Sci Adv. 2020;6:eabb1611.
Burki F, Kaplan M, Tikhonenkov DV, Zlatogursky V, Minh BQ, Radaykina LV, et al. Untangling the early diversification of eukaryotes: a phylogenomic study of the evolutionary origins of Centrohelida, Haptophyta and Cryptista. Proc R Soc B. 2016;283:20152802.
Cavalier-Smith T. Membrane heredity and early chloroplast evolution. Trends Plant Sci. 2000;5:174–82.
Douglas SE, Murphy CA, Spencer DF, Gray MW. Cryptomonad algae are evolutionary chimaeras of two phylogenetically distinct unicellular eukaryotes. Nature. 1991;350:148–51.
Cavalier-Smith T. Principles of protein and lipid targeting in secondary symbiogenesis: euglenoid, dinoflagellate, and sporozoan plastid origins and the eukaryote family tree. J Eukaryot Microbiol. 1999;46:347–66.
Curtis BA, Tanifuji G, Burki F, Gruber A, Irimia M, Maruyama S, et al. Algal genomes reveal evolutionary mosaicism and the fate of nucleomorphs. Nature. 2012;492:59–65.
Douglas S, Zauner S, Fraunholz M, Beaton M, Penny S, Deng LT, et al. The highly reduced genome of an enslaved algal nucleus. Nature. 2001;410:1091–6.
Hoef-Emden K, Archibald JM. Cryptophyta (Cryptomonads). In: Archibald JM, Simpson AGB, Slamovits CH, editors. Handbook of the protists. Cham: Springer International Publishing; 2017. p. 851–91.
Ward BA, Follows MJ. Marine mixotrophy increases trophic transfer efficiency, mean organism size, and vertical carbon flux. Proc Natl Acad Sci U S A. 2016;113:2958–63.
Herfort L, Peterson TD, Campbell V, Futrell S, Zuber P. Myrionecta rubra (Mesodinium rubrum) bloom initiation in the Columbia River estuary. Estuar Coast Shelf Sci. 2011;95:440–6.
Johnson MD, Beaudoin DJ, Laza-Martinez A, Dyhrman ST, Fensin E, Lin S, et al. The genetic diversity of Mesodinium and associated cryptophytes. Front Microbiol. 2016;7:2017.
Lindholm T. Mesodinium rubrum—a unique photosynthetic ciliate. Adv Aquat Microbiol. 1985;3:1–48.
Nowack EC, Melkonian M. Endosymbiotic associations within protists. Philos Trans R Soc B. 2010;365:699–712.
Johnson MD, Oldach D, Delwiche CF, Stoecker DK. Retention of transcriptionally active cryptophyte nuclei by the ciliate Myrionecta rubra. Nature. 2007;445:426–8.
Hansen PJ, Moldrup M, Tarangkoon W, Garcia-Cuetos L, Moestrup O. Direct evidence for symbiont sequestration in the marine red tide ciliate Mesodinium rubrum. Aquat Micro Ecol. 2012;66:63–75.
Kim M, Drumm K, Daugbjerg N, Hansen PJ. Dynamics of sequestered cryptophyte nuclei in Mesodinium rubrum during starvation and refeeding. Front Microbiol. 2017;8:1–14.
Nam SW, Park JW, Yih W, Park MG, Shin W. The fate of cryptophyte cell organelles in the ciliate Mesodinium cf. rubrum subjected to starvation. Harmful Algae. 2016;59:19–30.
Juel Hansen P, Fenchel T. The bloom-forming ciliate Mesodinium rubrum harbours a single permanent endosymbiont. Mar Biol Res. 2006;2:169–77.
Smith M, Hansen PJ. Interaction between Mesodinium rubrum and its prey: importance of prey concentration, irradiance and pH. Mar Ecol Prog Ser. 2007;338:61–70.
Matthew DJ, Diane KS. Role of feeding in growth and photophysiology of Myrionecta rubra. Aquat Micro Ecol. 2005;39:303–12.
Fenchel T, Hansen PJ. Motile behaviour of the bloom-forming ciliate Mesodinium rubrum. Mar Biol Res. 2006;2:33–40.
Gustafson DE, Stoecker DK, Johnson MD, Van Heukelem WF, Sneider K. Cryptophyte algae are robbed of their organelles by the marine ciliate Mesodinium rubrum. Nature. 2000;405:1049–52.
Peltomaa E, Johnson M. Mesodinium rubrum exhibits genus-level but not species-level cryptophyte prey selection. Aquat Micro Ecol. 2017;78:147–59.
Kim GH, Han JH, Kim B, Han JW, Nam SW, Shin W, et al. Cryptophyte gene regulation in the kleptoplastidic, karyokleptic ciliate Mesodinium rubrum. Harmful Algae. 2016;52:23–33.
Lasek-Nesselquist E, Wisecaver JH, Hackett JD, Johnson MD. Insights into transcriptional changes that accompany organelle sequestration from the stolen nucleus of Mesodinium rubrum. BMC Genom. 2015;16:805.
Chen Y, Chen Y, Shi C, Huang Z, Zhang Y, Li S, et al. SOAPnuke: a MapReduce acceleration-supported software for integrated quality control and preprocessing of high-throughput sequencing data. Gigascience. 2018;7:1–6.
Parekh S, Ziegenhain C, Vieth B, Enard W, Hellmann I. The impact of amplification on differential expression analyses by RNA-seq. Sci Rep. 2016;6:25533.
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol. 2011;29:644–52.
Haas BJ, Papanicolaou A, Yassour M, Grabherr M, Blood PD, Bowden J, et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat Protoc. 2013;8:1494–512.
Fu L, Niu B, Zhu Z, Wu S, Li W. CD-HIT: accelerated for clustering the next-generation sequencing data. Bioinformatics. 2012;28:3150–2.
Li W, Godzik A. Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics. 2006;22:1658–9.
Patro R, Duggal G, Love MI, Irizarry RA, Kingsford C. Salmon provides fast and bias-aware quantification of transcript expression. Nat Methods. 2017;14:417–9.
Ondov BD, Treangen TJ, Melsted P, Mallonee AB, Bergman NH, Koren S, et al. Mash: fast genome and metagenome distance estimation using MinHash. Genome Biol. 2016;17:132.
Ondov BD, Starrett GJ, Sappington A, Kostic A, Koren S, Buck CB, et al. Mash screen: high-throughput sequence containment estimation for genome discovery. Genome Biol. 2019;20:232.
Li H. Minimap2: pairwise alignment for nucleotide sequences. Bioinformatics. 2018;34:3094–100.
Enright AJ, Van Dongen S, Ouzounis CA. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. 2002;30:1575–84.
Li H, Durbin R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics. 2009;25:1754–60.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, et al. The sequence Alignment/Map format and SAMtools. Bioinformatics. 2009;25:2078–9.
Li H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics. 2011;27:2987–93.
Simao FA, Waterhouse RM, Ioannidis P, Kriventseva EV, Zdobnov EM. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics. 2015;31:3210–2.
Quinlan AR, Hall IM. BEDTools: a flexible suite of utilities for comparing genomic features. Bioinformatics. 2010;26:841–2.
Love MI, Huber W, Anders S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014;15:550.
Langfelder P, Horvath S. WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 2008;9:559.
Kanehisa M, Goto S. KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 2000;28:27–30.
Boyle EI, Weng S, Gollub J, Jin H, Botstein D, Cherry JM, et al. GO::TermFinder-open source software for accessing Gene Ontology information and finding significantly enriched Gene Ontology terms associated with a list of genes. Bioinformatics. 2004;20:3710–5.
Benjamini Y, Hochberg Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B (Stat Method). 1995;57:289–300.
Merico D, Isserlin R, Stueker O, Emili A, Bader GD. Enrichment map: a network-based method for gene-set enrichment visualization and interpretation. PLoS ONE. 2010;5:e13984.
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 2003;13:2498–504.
Luo W, Brouwer C. Pathview: an R/Bioconductor package for pathway-based data integration and visualization. Bioinformatics. 2013;29:1830–1.
Allen JF, de Paula WBM, Puthiyaveetil S, Nield J. A structural phylogenetic map for chloroplast photosynthesis. Trends Plant Sci. 2011;16:645–55.
Qiu H, Lee Jun M, Yoon Hwan S, Bhattacharya D. Hypothesis: gene-rich plastid genomes in red algae may be an outcome of nuclear genome reduction. J Phycol. 2017;53:715–9.
Grzebyk D, Schofield O, Vetriani C, Falkowski PG. The mesozoic radiation of eukaryotic algae: the portable plastid hypothesis. J Phycol. 2003;39:259–67.
Hehenberger E, Gast RJ, Keeling PJ. A kleptoplastidic dinoflagellate and the tipping point between transient and fully integrated plastid endosymbiosis. Proc Natl Acad Sci USA. 2019;116:17934–42.
Onuma R, Hirooka S, Kanesaki Y, Fujiwara T, Yoshikawa H, Miyagishima S-Y. Changes in the transcriptome, ploidy, and optimal light intensity of a cryptomonad upon integration into a kleptoplastic dinoflagellate. ISME J. 2020;14:2407–23.
McFadden GI. Origin and evolution of plastids and photosynthesis in eukaryotes. Cold Spring Harb Perspect Biol. 2014;6:a016105.
Park MG, Kim M, Kim S. The acquisition of plastids/phototrophy in heterotrophic Dinoflagellates. Acta Protozool. 2014;53:39–50.
Johnson MD, Beaudoin DJ. The genetic diversity of plastids associated with mixotrophic oligotrich ciliates. Limnol Oceanogr. 2019;64:2187–201.
Kim M, Kim S, Yih W, Park MG. The marine dinoflagellate genus Dinophysis can retain plastids of multiple algal origins at the same time. Harmful Algae. 2012;13:105–11.
Tourancheau AB, Tsao N, Klobutcher LA, Pearlman RE, Adoutte A. Genetic code deviations in the ciliates: evidence for multiple and independent events. EMBO J. 1995;14:3262–7.
Heaphy SM, Mariotti M, Gladyshev VN, Atkins JF, Baranov PV. Novel ciliate genetic code variants including the reassignment of all three stop codons to sense codons in Condylostoma magnum. Mol Biol Evol. 2016;33:2885–9.
Johnson MD, Beaudoin DJ, Frada MJ, Brownlee EF, Stoecker DK. High grazing rates on cryptophyte algae in Chesapeake Bay. Front Mar Sci. 2018;5:1–13.
Guo X, Chen F, Gao F, Li L, Liu K, You L, et al. CNSA: a data repository for archiving omics data. Database. 2020;2020:baaa055.
Chen Fengzhen YL, Fan Yang, Lina Wang, Xueqin Guo, Fei Gao, Cong Hua, et al. CNGBdb: China National GeneBank DataBase. Hereditas. 2020;42:799–809.
Acknowledgements
This project was supported by the Danish Council for Independent Research (Grant number 4181–00484 to PJH), the Major scientific and technological projects of Hainan Province (ZDKJ2019011), the National Key Research and Development Program of China (2018YFC0308401), and the Carlsberg Foundation (2012_01_0515 to LG-C). We acknowledge the China National GeneBank (CNGB) for support with computing resources.
Author information
Authors and Affiliations
Contributions
PJH, NL, UJ, and AA conceived the work. AA, KD, MK, and LG-C did the culturing; AA and LG-C extracted RNA; QL, YZ, and XZ did the sequencing; HC, AA, and QL analyzed the data under the supervision of SL and NL; AA, HC, and QL drafted the manuscript. All authors contributed to writing of the final version of the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised due to an error in a sub-section of the methods.
Rights and permissions
About this article
Cite this article
Altenburger, A., Cai, H., Li, Q. et al. Limits to the cellular control of sequestered cryptophyte prey in the marine ciliate Mesodinium rubrum. ISME J 15, 1056–1072 (2021). https://doi.org/10.1038/s41396-020-00830-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-020-00830-9
This article is cited by
-
Prey preference in a kleptoplastic dinoflagellate is linked to photosynthetic performance
The ISME Journal (2023)
-
De novo transcriptome assembly and gene annotation for the toxic dinoflagellate Dinophysis
Scientific Data (2023)
-
Taming the perils of photosynthesis by eukaryotes: constraints on endosymbiotic evolution in aquatic ecosystems
Communications Biology (2023)
-
Multiyear analysis uncovers coordinated seasonality in stocks and composition of the planktonic food web in the Baltic Sea proper
Scientific Reports (2023)
-
Microzooplankton grazing and its key group composition in subtropical eutrophic coast of Southern China: in relation to environmental changes
Ecotoxicology (2021)