Abstract
Microbially induced corrosion of metallic iron (Fe0)-containing structures is an environmental and economic hazard. Methanogens are abundant in low-sulfide environments and yet their specific role in Fe0 corrosion is poorly understood. In this study, Sporomusa and Methanosarcina dominated enrichments from Baltic Sea methanogenic sediments that were established with Fe0 as the sole electron donor and CO2 as the electron acceptor. The Baltic-Sporomusa was phylogenetically affiliated to the electroactive acetogen S. silvacetica. Baltic-Sporomusa adjusted rapidly to growth on H2. On Fe0, spent filtrate enhanced growth of this acetogen suggesting that it was using endogenous enzymes to retrieve electrons and produce acetate. Previous studies have proposed that acetate produced by acetogens can feed commensal acetoclastic methanogens such as Methanosarcina. However, Baltic-methanogens could not generate methane from acetate, plus the decrease or absence of acetogens stimulated their growth. The decrease in numbers of Sporomusa was concurrent with an upsurge in Methanosarcina and increased methane production, suggesting that methanogens compete with acetogens for electrons from Fe0. Furthermore, Baltic-methanogens were unable to use H2 (1.5 atm) for methanogenesis and were inhibited by spent filtrate additions, indicating that enzymatically produced H2 is not a favorable electron donor. We hypothesize that Baltic-methanogens retrieve electrons from Fe0 via a yet enigmatic direct electron uptake mechanism.
Similar content being viewed by others
Introduction
Microbially induced corrosion (MIC) accounts for 20% of the total corrosion costs for the oil and gas industries [1, 2]. Additionally, chemical leaks from corroded waste containers cause health and environmental problems [3, 4]. Previous studies have primarily focused on MIC in sulfide-rich marine environments, where sulfate-reducing bacteria are predominantly causing corrosion [5]. However, in low-sulfide environments such as the Bothnian Bay, Baltic Sea [6], cooperative interactions between microorganisms (i.e. methanogens and acetogens) may be responsible for corrosion of infrastructure [7,8,9,10]. Microbial cooperation was proposed to enhance corrosion rates [11], but it is poorly understood. Of the methanogens, particularly Methanosarcina have been suggested to play an essential role in corrosion. Methanosarcina are frequently associated with corroded Fe0-structures from oil, gas, sewage, water storage, and transportation facilities [7,8,9,10], and in aquifers where radionuclide-waste is stored underground in steel containers [12].
Nevertheless, of the highly corrosive methanogens described to date (Methanococcus maripaludis strains KA1, Mic1c10, MM1264, and Methanobacterium strain IM1 [13,14,15,16]), none belongs to the genus Methanosarcina. Studies of corrosive Methanococcus and Methanobacterium have shown that H2 generated abiotically by Fe0 cannot provide sufficient electrons to account for all the methane being produced by these species during growth in the presence of Fe0 [13,14,15,16,17]. Consequently, the electron uptake mechanisms proposed for these different methanogenic strains included (1) a direct uptake route [15, 17] or (2) an extracellular enzyme-mediated electron uptake route [18, 19].
(1) Direct electron uptake from Fe0 by Methanobacterium strain IM1 was suggested as an alternative to abiotic-H2 uptake because this strain generated more methane (CH4) from Fe0 oxidation than a H2-utilizing M. maripaludis strain [15] with low H2-uptake thresholds [20]. IM1 also produced methane when a cathode poised at a potential unfavorable for abiotic H2 evolution was the sole source of electrons [17]. However, the mechanism utilized by IM1 for electron uptake directly from Fe0 or electrodes is unknown. Also, it is unknown whether other methanogens have similar capabilities.
Methanosarcina species were previously shown to carry out Fe0-dependent methanogensis presumably by using abiotic-H2 evolved at the Fe0-surface [21]. However, some Methanosarcina cannot use H2 [22,23,24,25,26], and even those Methanosarcina species that are capable of hydrogenotrophic methanogenesis have high thresholds for H2-uptake [27, 28]. Thus, Methanosarcina species would not be competitive at abiotic-H2 removal from the Fe0 surface. Nonetheless, Methanosarcina-species dominate on corroded Fe0-structures [7,8,9,10]. Therefore, we propose that Methanosarcina retrieves electrons directly from Fe0, extrapolating from recent findings that Methanosarcina can retrieve extracellular electrons directly from poised electrodes [29, 30], electrogenic syntrophic partners [31, 32], or electrically conductive particles [31,32,33,34]. Plausible scenarios for direct electron uptake in Methanosarcina have only recently been substantiated, using comparative transcriptomics [35].
(2) In addition to direct extracellular electron uptake, a second strategy making use of extracellular enzymes to capture electrons from Fe0 was described in methanogens [16, 18, 19]. Enzymes like hydrogenases, formate dehydrogenases, or the heterodisulfide reductase supercomplex produced by methanogens can generate H2 or formate from Fe0-derived electrons [16, 18, 19]. It is not clear whether an extracellular enzyme-dependent strategy would be competitive in corrosive environments. Moribund cells may release enzymes like hydrogenases into their extracellular milieu [19] that can capture electrons freed during Fe0-oxidation to reduce protons from solution to H2 [36, 37]. H2 could then be used non-specifically by a variety of H2-utilizers. Sensitive anaerobic enzymes tend to only be stable for a few days outside of the cell [38]. However, Fe2+ released during the corrosion process may further stabilize these enzymes [39]. Hydrogenase-mediated H2 production at the Fe0 surface appears to generate sufficient H2 for the growth of hydrogenotrophic methanogens [16, 19].
Corroded infrastructure often harbors both Methanosarcina-methanogens and acetogens, where Methanosarcina is thought to play a role in Fe0-corrosion [7,8,9,10]. However, Methanosarcina’s role was assumed to be indirect and dependent on cooperation with other corrosive organisms. For example, Methanosarcina was proposed to utilize acetate produced by acetogens actively corroding Fe0. In this study, we investigated the theory that acetoclastic methanogens like Methanosarcina require cooperative interactions with acetogens to corrode Fe0. We used Fe0 to enrich for Methanosarcina species from sediments collected off the coast of Bothnia. Molecular and physiological tests were used to investigate the role of methanogens and their possible synergy with co-occurring microbes during Fe0 corrosion. We present evidence that Baltic-methanogens perform Fe0-dependent methanogenesis and compete with acetogens for access to Fe0. Specific inhibition experiments indicate that two different mechanisms for Fe0-dependent electron uptake by Baltic-acetogens and methanogens are feasible.
Materials and methods
Baltic-Sea-enrichment cultures
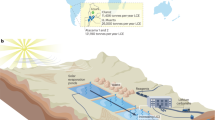
We collected sediment cores from the Bothnian Bay, Baltic Sea at a water depth of 15 m (65°43.6′N and 22°26.8′E; station RA2) during August 2014 [40]. The sediment had a temperature of 15 °C and an in situ salinity of 0.5. The mineral content was low in insoluble manganese oxides, high in insoluble FeS, and high in crystalline iron oxides, such as semiconductive goethite or conductive magnetite, as previously described [40].
Enrichment cultures were prepared using sediment from the methanogenic zone (30–36 cm) under aseptic and anoxic conditions as previously described [40], but with the addition of 100 g/L iron granules, and exclusion of sulfide as a reducing agent, which was instead replaced with an additional 2 mM cysteine (cf = 3 mM). We prepared all subsequent transfers in 50 mL blue chlorobutyl-rubber-stoppered glass vials with an anoxic headspace of CO2/N2 (20:80, v/v). For all enrichment incubations, we used a DSM120-modified medium (modifications: 0.6 g/L NaCl, without casitone, sodium acetate, methanol, or Na2S×9H2O). For Fe0 incubations, we added to the media as sole electron donor iron granules (99.98%, ThermoFisher, Germany) or iron coupons (3 cm × 1 cm × 1 mm). Other electron donors tested included H2 (ca. 1.5 atm) and acetate (10 mM).
All culture experiments were carried out in at least triplicate and sometimes up to 10 replicates. As soon as methane production reached stationary phase, we transferred cells into fresh media with Fe0. Cultures were shaken vigorously to remove cells from Fe0-granules. We then used 10–20% of the dispersed cells to inoculated fresh Fe0-media for subsequent transfers. The temperature used for all incubations was 20 °C. To reach stationary, initial enrichments required circa 6 months, whereas later transfers (T3) took circa 3 months and most recent incubations (T10) took only 1–2 months.
Abiotic control experiments lacked cells. However, we used the same Fe0-media and incubation conditions as done for experiments with cells.
Inhibition experiments were carried out by the addition of inhibitors specific for methanogens or bacteria. A methanogenesis inhibitor (2 mM 2-bromoethane sulfonate [41]) was added to the culture media to generate a bacteria-only culture. In order to generate a methanogen-only culture, we added a mixture of antibiotics (200 µg/mL kanamycin and 100 µg/mL ampicillin) to the culture media. Experiments with inhibitors were run parallel to controls lacking inhibitors.
Spent filtrate addition experiments were carried out by the addition of 1 mL spent media from a stationary culture to a new culture as described previously [40]. The spent media of a Sporomusa acetogen and a Methanococcus methanogen were previously shown to contain electroactive enzymes, which retrieve electrons from Fe0 or electrodes for proton reduction to H2 [42]
Downstream analyses, DNA extractions, substrate evaluations, and microscopy were performed after the fifth consecutive transfer on Fe0.
Chemical analyses
To determine methane and H2 concentrations, we used a Trace 1300 gas chromatography system (Thermo Scientific, Italy) equipped with a thermal conductivity detector (TCD), an injector operated at 150 °C and a detector at 200 °C with 1.0 mL/min reference gas flow. The oven temperature was constant at 70 °C. A TG-BOND Msieve 5A column (Thermo Scientific; 30-m length, 0.53-mm i.d., and 20-μm film thickness) was used with argon as the carrier gas with a set flow at 25 mL/min. The GC was controlled and automated with the Chromeleon software (Dionex, Version 7). Using this setup, the minimum detection limit for methane and H2 was 5 µM.
For determination of acetate concentrations, we used a Dionex ICS-1500 Ion Chromatography System (ICS-1500) equipped with the AS50 autosampler, and an IonPac AS22 column coupled to a conductivity detector (31 mA). For separation of volatile fatty acids, we used 4.5 mM Na2CO3 with 1.4 mM NaHCO3 as eluent. The run was isothermic at 30 °C with a flow rate of 1.2 mL/min.
For determination of ferrous iron (Fe2+) produced by Fe0-oxidation in our cultures, we dissolved Fe2+ in 0.67 M HCl (containing 0.67 M hexamethylenetetramine to avoid dissolution of metallic iron) and quantified Fe2+ concentrations colorimetrically with the ferrozine assay [43].
For elemental analyses of the gray-black crust that formed on Fe0-coupons after 2 months when cells were present, the crust was scraped off the Fe0-coupons and dried in an anoxic glove box. Mass spectrometry informed on the content of carbonate and organic carbon. For total reduced inorganic sulfur determination (including iron monosulfides, pyrite, and S0) we performed hot chromium distillation [44]. The organic carbon quantification took place after acidification with HCl. We calculated the value for the carbon in carbonates by subtracting the organic-carbon values remaining after acidification from the total unacidified carbon (C-total).
DNA purification from microbial enrichments
DNA purification was performed using a combination of two commercially available kits; the MasterPure™ Complete DNA and RNA Purification Kit (Epicenter, Madison, Wi, USA), and the Fast Prep spin MPTM kit for soil (Mobio/Qiagen, Hildesheim, Germany). For DNA extraction, we pelleted 10 mL cells, either by harvesting an entire culture grown on Fe0 or by removing 10 mL from a larger volume after vigorous shaking the Fe0-cultures in order to detach cells from the Fe0-surface. We used an Epicenter kit to initiate the DNA extraction with the following modifications to the manufacturer’s protocol: a three-fold higher concentration of proteinase K was added to ensure cell lysis, and a prolonged incubation time at 65 °C was performed until the color of the samples changed from black to brown (the brown pellet gave higher DNA extraction efficiencies). After DNA extraction, we used the Fast Prep spin MPTM kit for soil to carry out RNase treatment and protein precipitation. An advantage of this kit is that it allowed removal of the high iron content, while simultaneously purifying DNA on a binding matrix. Quality and quantity of genomic DNA were determined by electrophoresis on a 1% agarose gel and by UV spectrophotometry on a mySPEC spectrophotometer (VWR®, Germany).
Metagenome analyses
After a single whole-genome amplification cycle, random shotgun metagenome sequencing was performed commercially (Macrogen/Europe) using the Illumina HiSeq2500 platform. We merged the unassembled DNA sequences, checked for quality, and annotated using the Metagenomics Rapid Annotation (MG-RAST) server (vs. 4.03) with default parameters [45]. Shotgun metagenome sequencing resulted in 10,739 high-quality assembled reads of a total of 10,749 with an average length of 167 bp. We obtained metagenome taxonomy information using the databases available in MG-RAST, including Silva [46], RDP [47], Greengenes [48], and RefSeq [49]. For the metagenome taxonomy, the horizontal asymptote of the rarefaction curve indicated complete coverage of the prokaryotic diversity in these samples. For metagenome taxonomy analyses, we used the default MG-RAST cutoff parameters: e-value of 1E–5, a minimum identity of 60%, and a maximum alignment length of 15 bp. The metagenome data is available at MG-RAST with this ID: MGM4796413.3.
16S rRNA gene sequence analyses
General Archaeal and Bacterial primers (Table 1SM) were used to perform PCR amplification of the 16S rRNA gene from the isolated DNA. PCR reactions contained in a final volume of 50 μL, 1.5 mM MgCl2, 0.2 mM dNTPs, 0.2 μM of each primer, and 1U Promega Taq polymerase, and 10x PCR reaction buffer. PCR reactions included an initial denaturation step at 94 °C for 10 min; then 35 cycles of denaturation at 94 °C for 30 s, annealing at the specific annealing temperature for the primer pair (Table 1SM) for 30 s, and extension at 72 °C for 90 s; and a final extension cycle at 72 °C for 10 min. Next, we cloned PCR products with the TOPO® TA Cloning® Kit for Sequencing (Invitrogen, Carlsbad, CA, USA). PCR products from individual clones were amplified with M13-vector primers and sent to the Institute of Clinical Molecular Biology in Kiel for Sanger sequencing. Sequences were analyzed using the Geneious® software package, version 11.0.4 [50], and compared against the NCBI GenBank DNA database using BLAST. Consensus sequences for Archaea and Bacteria (97% identity) were assembled using ClustalW implemented within Geneious. Consensus 16S rRNA gene sequences were used to construct maximum likelihood phylogenetic trees in Geneious using RaxML [51]. We deposited sequences in GenBank, under the accession number: MK433201.
Quantitative PCR
Extracted DNA was used for 16S rRNA gene quantification via qPCR with specific Sporomusa, Methanosarcina, and general Bacteria primers (Table 1SM). For quantification of the members within the corrosive Baltic Sea community, we carried out qPCR assays on duplicate cultures harvested at different times during their growth; 18 days (T10) and 60 days (T9). For each biological replicate, we run quadruplicate qPCR reactions alongside quadruplicate standards (101–108 16S rRNA gene copies per ml). All standards were prepared as previously described [40, 52].
We prepared the qPCR reaction mix as described before in a final volume of 25 µl of which 10 µl were a 5Prime Hot Master Mix, 0.25 µl BSA (stock 10 mg/ml), 1 µl or forward and reverse primer (10 µM stock each) and 1 µl of a template [40]. The qPCR amplification ran as follows: 2 min hot start at 94 °C, 1 min denaturation at 94 °C, 1 min at the annealing temperature appropriate for the primer pair used (Table 1SM) and 2 min extension at 72 °C. Steps two to four (denaturation, annealing, and extension) were repeated 40 times. The final step was a 10 min elongation step at 72 °C and storage at 4 °C.
Fluorescence in situ hybridization
To fix cells, we added 2% microscopy grade paraformaldehyde (PFA, 16%) directly to anaerobic cultures and incubated them for 2 h at room temperature. Then all cells were collected with the Fe0-granules via centrifugation at 10,000 rpm for 10 min. We gently sonicated (20% intensity; 5 × 5 s) to detach cells from the Fe0-granules. Then we collected 50 µL of the resuspended cells by filtration on 0.22 µm filters. We hybridized cells with specific probes (final concentration 5 ng/µL) for Methanosarcina (MX821) using the formamide concentration specified in Table 1SM, followed by 2 h hybridization at 46 °C and 15 min. washing at 48 °C [53]. For counterstaining, we incubated the air-dried filters for 3–5 min with a DNA fluorescent stain: 4′,6-diamidino-2-phenylindole (DAPI; 1 µg/ml).
Epifluorescence microscopy
To confirm the presence or absence of methanogens, we also used their natural autofluorescence due to coenzyme F420 and visualized the cells on an epifluorescence microscope equipped with a 420 nm excitation filter as previously described [54]. To visualize cells, we used an upright epifluorescence microscope from Zeiss (Axioscope A1) equipped with a Cy3 (excitation 549 nm, emission 562 nm), a DAPI (excitation 359 nm, emission 461 nm), and an F420-filter set (excitation 420 nm, emission 480 nm). For image acquisition, we used a digital CCD camera (Axiocam) controlled by an Axiovision vs. 4.7 software.
Scanning electron microscopy and sample preparation
We carried out SEM visualization on cells from the fifth transfer that have been growing on Fe0-coupons for circa 3 months. We removed the excess culture media and directly fixed cells attached to the Fe0-coupon in the anaerobic culture vials using a mix of 2.5% (v/v) glutaraldehyde in 0.1 M phosphate buffer (pH 7.3). Cells were incubated at 4 °C for 12 h, washed in phosphate buffer, dehydrated with anoxic ethanol at increasing concentrations (35%, 50%, 70%, 80%, 90%, 95%, 100%, and three times in 100% v/v; each step for 10 min). The Fe0-coupons were then chemically dried with hexamethyldisilazane for 30 min [55] and traces of hexamethyldisilazane were evaporated under N2. We stored dried-out Fe0-coupons in the culture bottle under N2-gas before electron microscopy. Scanning electron microscopy (SEM) was performed with a FESEM Magellan 400 at 5.0 kV at the microscopy facility of the University of Massachusetts, Amherst, USA.
Results and discussion
Previous studies of corrosive non-sulfidic environments attributed corrosion to syntrophic interactions between acetogens (e.g. Clostridium) and acetate-utilizing methanogens (Methanosarcinales) [7,8,9,10]. Here we challenge this assumption and demonstrate that acetogens and methanogens compete for electrons from Fe0, rather than operating cooperatively.
A corrosive community became enriched from methanogenic sediment collected off the Swedish coast using Fe0 as the sole electron donor and CO2 as the sole electron acceptor. After the establishment of original slurries on Fe0 (25% sediment), cultures were transferred sequentially into fresh Fe0-containing media using a 10–20% inoculum for the next 3 years.
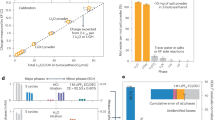
Original slurries from Baltic Sea sediments provided with Fe0 generated circa five times more methane (Fig. 1a) and four times more acetate (Fig. 1a) than parallel incubations without Fe0 (Fig. 1b). Surprisingly, these Fe0-containing slurries accumulated acetate (Fig. 1), which was not consumed by the Methanosarcina known to harbor these sediments [40].
Initial slurries established from methanogenic sediments collected off the coast of the Baltic Sea and third transfer incubations with Fe0 or acetate as electron donors vs. parallel control incubations without electron donors. a Slurries incubated with Fe0 or b without Fe0 (n = 3). Electron conversions into products (methane and acetate) are presented as mM electron equivalents (mM eeq) taking into account that a mol methane/acetate requires 8 mols electrons according to the reactions: CO2 + 8e- + 8H+ → CH4 + 2H2O (methanogenesis) and 2 CO2 + 8e- + 8H+ → CH3COOH + 2H2O (acetogenesis). c Fe0-dependent methane production in the third successive passage (sediment-free). However, methane was undetected in electron-donor free controls and when 10 mM acetate replaced Fe0 as sole electron donor. The electron donor-free control was used to probe for sediment carryover substrates (c-inset). A gray-black corrosion product was observed only in the presence of cells and not in their absence (c-inset)
After three transfers, incubations became sediment-free as determined by (i) visual inspection, (ii) microscopy, and (iii) the fact that the inoculum did not lead to product formation from carryover of electron donors in incubations without external electron donors (Figs. 1c and 1SM). To verify for acetate utilization by acetoclastic methanogens, Fe0 was replaced with 10 mM acetate as the sole substrate for growth. Acetate did not lead to methane production after 2 months of incubation, whereas Fe0 did (Fig. 1c), indicating that enriched Baltic-methanogens became adjusted to Fe0-dependent methanogenesis, and were not capable of acetoclastic methanogenesis during the given time frame of 2 months.
These sediment-free cultures formed a black crust on the surface of the metal, which was absent in abiotic incubations (Fig. 1c—inset). Under similar conditions (non-sulfidic, carbonate-buffered, pH~7), Fe0 is oxidized primarily into Fe2+-carbonates such as siderite [15, 56, 57]. Microorganisms like methanogens or acetogens accelerate Fe0-oxidation to Fe2+ via processes that are energetically more favorable than the abiotic reaction (Table 1). During the eighth transfer, we removed and analyzed the gray-black precipitate formed on Fe0 by the Baltic Sea methanogenic community. We determined that the precipitate had a high carbonate content (ca. 50% by weight in FeCO3 equivalents), but low reduced inorganic sulfur content (ca. 0.1%; including sulfides), consistent with the formation of iron carbonates, like siderite. The remaining organic carbon content was ~1%.
Enhanced Fe0-oxidation to Fe2+ by Baltic Sea acetogens and methanogens
Fe0 corrosion was assessed using ferrous iron (Fe2+) accumulation as a proxy for corrosion (Fig. 2a, b), as done before, particularly in studies documenting corrosion by methanogens [13,14,15, 58, 59].
Products of Fe0 exposure to microorganisms from the Baltic Sea. a Cells enhanced Fe0 oxidation to Fe2+ above the background observed in cell-free (abiotic) controls. b Absolute rates of Fe2+-yield over 25 days in the presence or absence of cells. c In abiotic controls, electrons were recovered as H2. d The entire community grown on Fe0 for eight transfers recovered electrons as methane and acetate. e Inhibition of all bacteria (including acetogens) with a mixture of antibiotics led to enhanced methanogenesis (filled line) compared to the mixed-community (dotted line). f Specific inhibition of methanogens with BES, enhanced acetogenesis (filled line) compared to the mixed-community (dotted line) (n > 5)
In our incubations, the corrosive microbial community started Fe0-oxidation to Fe2+ immediately and persisted for circa 25 days (Fig. 2a). The presence of cells led to a tripling of the Fe2+ yield per day (44.7 ± 4.6 µM/day) compared to the background Fe2+ yield observed in abiotic controls (16.9 ± 10 µM/day) (Fig. 2b). The daily increase of Fe2+ in the presence of a Baltic-community indicates that the community was more corrosive than abiotic controls.
Fe0-corrosion under non-sulfidic, carbonate-rich conditions in the absence of electron acceptors other than CO2 can be attributed to two possible metabolisms: Fe0-dependent methanogenesis or Fe0-dependent acetogenesis. Theoretically, Fe0-dependent methanogenesis is energetically more favorable than Fe0-dependent acetogenesis, but both are energetically more favorable than abiotic Fe0-dissolution (Table 1). Therefore, to better understand the interplay between acetogens and methanogens within the corrosive microbial community we (1) monitored product evolution and (2) inhibited various metabolic groups in order to determine the corrosive potential of each surviving group.
Electron recoveries from Fe0
After eight transfers on Fe0-containing medium, the Baltic Sea corrosive enrichment cultures exhibited a quick Fe0-dependent acetogenic phase (days 5–10) followed by a slow methanogenic phase (days 10–25) (Fig. 2d). During the acetogenic phase, acetogens were able to convert 2.2 ± 0.1 mM/day electron equivalents from Fe0 to form acetate (Fig. 2d). As soon as the acetogenic phase stopped, the methanogenic-phase began, and methanogens recuperated 0.2 ± 0.05 mM electron equivalents from Fe0 into methane daily. After 25 days, the community routed more electrons into acetate than into methane (Fig. 2d). Thus, it appears that acetogens outcompeted methanogens for access to electrons from Fe0.
Competition between acetogens and methanogens
We evaluated whether methanogens were in competition with acetogens for electrons from Fe0 by testing whether methanogens functioned better without acetogens. In order to test this, we inhibited acetogens and other bacteria using antibiotic additions (kanamycin and ampicillin). With acetogens inhibited, methanogens oxidized Fe0 to Fe2+ at rates above abiotic controls (Fig. 2e) and similar to those observed for the entire community (Fig. 2d). After 15 days, methanogens alone produced six-times more methane (3.8 ± 0.7 mM electron equivalents/eeq CH4) (Fig. 2e) than they did when they were co-existing with bacteria in a mixed community (0.6 ± 0.2 mM eeq CH4) (Fig. 2d). During the first 5 days of the methanogenic-phase (days 10–15), electron recoveries into methane were higher (0.81 ± 0.06 mM eeq CH4) than expected from the rates achievable if cells were dependent on the production of abiotic H2 production (0.65 ± 0.09 mM eeq H2). Nevertheless, electron recoveries decreased by half (0.42 ± 0.1 mM eeq CH4) (Fig. 2e), possibly due to competition for Fe0 with acetogens that developed antibiotic resistance and generated 2.5 mM eeq acetate. However, even brief inhibition periods of the acetogens led to significantly higher methanogenic activity (six-fold), indicating that acetogens inhibited methanogenesis on Fe0. Sub-optimal methane production suggests that methanogens may experience decreased access to electrons from Fe0 due to the competitive exclusion by acetogens.
To test whether methanogens impacted the growth of acetogens, we inhibited methanogens with BES, a methyl–CoA analog [41]. In the presence of the methanogenic inhibitor, methanogens were rendered inactive throughout the incubation (Fig. 2f). Nevertheless, acetogens alone were able to oxidize Fe0 to Fe2+ (Fig. 2a, b), while producing more acetate (14%; p = 0.0001) than they did within the mixed community (Fig. 2f). These data suggest that methanogens constrain the growth of acetogens.
These results indicate that acetogens and methanogens negatively affect one another when competing for Fe0 as the sole electron donor (Fig. 3).
Contrasting mechanisms of electron uptake from Fe0 in Baltic-acetogens and methanogens
To investigate the possible mechanisms of electron uptake by Baltic-acetogens and methanogens, we compared electron recoveries in abiotic controls vs. those in the presence of cells. In the absence of cells, Fe0 released 0.65 ± 0.09 mM/day electron equivalents as H2 (Fig. 2c) continuously for 25 days. When acetogens were present, electron equivalents were recovered 3.5 times faster than expected from abiotic H2 (Fig. 2f), excluding electron recovery into biomass unaccounted for during the experiments. Thus, acetogens likely used an alternative mechanism to access electrons from Fe0 easily and accelerate acetogenesis.
Acetogens have been shown to use two different mechanisms for electron uptake from Fe0 facilitated by enzymes evolving H2 [16, 19] or by direct-electron uptake [60,61,62]. The latter is plausible because several acetogens can grow on electrodes poised at potentials that do not generate abiotic H2 [60,61,62]. On the other hand, previous studies illustrated efficient enzymatic-mediated electron uptake from Fe0 using a purified Clostridium [FeFe]-hydrogenase, which retrieves electrons directly from Fe0 for proton oxidation to H2 [36, 37]. Unlike [NiFe]-hydrogenases from methanogens, the [FeFe]-hydrogenases of Clostridium are effective at oxidizing H+ [63] and quickly evolving H2 that could serve as an electron donor for Baltic-acetogens. Therefore, we had to determine whether Baltic-acetogens (i) utilize H2, (ii) are stimulated by endogenous enzymes, or (iii) use an alternative direct electron uptake mechanism.
To verify whether Baltic-acetogens could rapidly switch to H2 after being adapted to Fe0 as the sole electron donor for eight transfers, we incubated the acetogens on H2 (after BES-inhibition of methanogens). H2-dependent acetogenesis took 5 days to commence, similar to Fe0-incubations (Fig. 4). Unlike a 5-day long Fe0-dependent acetogenesis (Fig. 2f), H2-dependent acetogenesis continued steadily for 20 days (Fig. 4). Although Baltic-acetogens were effective H2-utilizers, the rates of abiotic H2 formation from Fe0 could not explain the tripling in electron recovery rates by acetogens on Fe0 (Fig. 2c, f). Therefore, we assumed that extracellular hydrogenases might stimulate electron uptake from Fe0 by inducing enzymatic H2-formation and subsequently enhancing the rates of acetogenesis from Fe0. To determine if such enzymes had a stimulatory effect, we filtered the spent medium of a pre-grown Fe0-culture into a fresh culture provided with Fe0. If the active enzymes present in the spent-filtrate stimulated the growth of acetogens then the rate of electron recovery from Fe0 into acetate would increase. Indeed, spent filtrate stimulated acetogenesis, which started 5 days earlier (Fig. 5a) than it did in Fe0-grown or H2-grown cultures of Baltic-acetogens. Moreover, acetate recoveries were the highest after the addition of spent filtrate (Fig. 5a) compared to the unamended community (22% increase, n = 10, p < 0.00001), or unamended acetogens (7% increase; n = 10, p < 0.02). These results suggest that Baltic-acetogens use an enzyme-mediated mechanism to enhance electron uptake from Fe0, similar to other acetogens [19, 55].
Incubations (9th transfer) replacing Fe0 with H2 as the electron donor for a Baltic-methanogens vs. b Baltic-acetogens. To generate culture conditions favorable only for Baltic-methanogens, we added kanamycin and ampicillin to inhibit the acetogens. To generate culture conditions favorable only for Baltic-acetogens, we added BES a specific inhibitor for the methanogens
We determined the possible impact of extracellular enzymes/shuttles from spent-filtrate by incubating 10 new cultures in Fe0/CO2-media spiked with 1 mL spent filtrate (n = 10, T8). These incubations were run in parallel with cultures without spent-filtrate (see Fig. 2). For clarity, we separated the acetogens and methanogens into two panels. In the spent-media spiked community we monitored a the early on-set for acetogenesis after the addition of spent filtrate to Fe0-cultures and b inhibition of methanogenesis after the addition of spent-filtrate. Gray lines depict trends for product formation by the acetogens-alone and methanogens-alone after specific inhibition of their competitors. Black lines represent product formation by acetogens and methanogens within the mixed-community
Like acetogens, methanogens are believed to retrieve electrons from Fe0 via an enzyme-mediated electron uptake mechanism [16, 18, 19, 64] or a poorly understood direct electron uptake system [15, 17]. Thus, to distinguish between these two mechanisms, we tested the impact of H2 and spent-media filtrate on the growth of Baltic-methanogens.
Unlike Baltic-acetogens, Baltic-methanogens could not utilize H2 for methanogenesis (Fig. 4), also their methane productivity declined after the addition of spent media filtrate (−23%; n = 10; p < 0.03; Fig. 5b). These results suggest that Baltic-methanogens did not use an enzyme-mediated electron uptake mechanism. These data corroborate with previously published results initiated from the same sediment and in which we observed that Methanosarcina was capable of mineral-mediated syntrophy independent of enzymes from spent filtrate additions [40]. Additionally, in the present study we show that Baltic-Methanosarcina preferred Fe0 as electron donor and were unable to consume acetate or abiotic-H2, demonstrating that they were likely retrieving electrons directly from Fe0 (Figs. 1b and 4).
Sporomusa and Methanosarcina dominate the corrosive microbial community
16S rRNA gene and metagenome sequence (MGS) analyses of the corrosive community identified Proteobacteria, Firmicutes, and Euryarchaeota (Fig. 6a, b) as representative phylotypes.
MGS and 16S rRNA gene library studies identified the same representative phylotypes in Baltic Sea corrosive enrichments during sequential transfers. a Phylogenetic assignment of Bacteria and Archaea according to metagenome sequencing (MGS) of duplicate-cultures harvested at the end of transfer #5. b Phylogenetic assignment of Bacteria and Archaea according to 16S rRNA gene libraries assembled from duplicate-cultures harvested at the end of transfer #8; Phylogenetic trees were built using sequences from the 16S rRNA gene libraries. c Maximum-likelihood tree showing complete 16S rRNA gene sequences of Baltic-Archaea. d Maximum-likelihood tree showing 16S rRNA gene sequences of Baltic-Bacteria. The numbers in parenthesis represent the number of clones with >97% similarity to each other. The scale bar reflects the mean number of nucleotide substitutions per site as inferred by the RAxML algorithm. The numbers close to the nodes reflect bootstrap values (>70% good support) as inferred from 100 tree iterations using RAxML
Proteobacteria clustered primarily with the genus Desulfovibrio. All Desulfovibrio sequences were most similar to D. idahonensis (97.5% identity; Fig. 6d) which was isolated from a metal(oid) contaminated sediment [65]. Baltic-Desulfovibrio were also related (98.5% identity) to a Desulfovibrio from a drinking water system contaminated by iron oxides [66]. Desulfovibrio species are capable of corrosion [5] under high-sulfate conditions common in marine environments (ca. 28 mM sulfate [67]). However, the methanogenic zone of the Baltic Sea contains no sulfate [6] and thus is a low-sulfide environment (0–2 mM [68]). Therefore, under the low-sulfate conditions in our media (ca. 0.4 mM) Desulfovibrio could only (i) use trace sulfate for its metabolism or (ii) ferment dead-biomass organics (e.g. pyruvate, fumarate) alone [69] or syntrophically [70].
Acetogens identified by MGS and 16S rRNA gene libraries belonged to the Firmicutes genera Sporomusa, Clostridium, and Acetobacterium (Fig. 6a, b, d). Spore-bearing curved rods resembling Sporomusa were visually observed in our cultures (Fig. 7a). Sporomusa dominated (>88%) the bacterial community according to 16S rRNA-gene qPCR analyses, independent of the incubation period (Fig. 7j). The closest isolated relative for our Baltic-Sporomusa was S. silvacetica (97.4% identity), previously shown to be capable of electroacetogenesis on a cathode at −400 mV vs. SHE [61]. Its closest uncultured relative (99.3% identity) was a Sporomusa kenriched on a cathode (Genbank KJ600503 Fig. 6d).
Morphotypes and abundant phylotypes from a Baltic corrosive community. a Scanning electron microscopy (SEM) image of a spore-forming curved rod resembling a Sporomusa sporulating cell. b SEM micrograph of tetrads of cocci resembling Methanosarcina and their usual cocci aggregates. SEM was performed at the end of transfer #5 (ca. 3 months). c Epifluorescence micrographs of DAPI-stained cells detached from Fe0-granules by shaking and sonication during day 18 of transfer #10. We observed two morphotypes: a banana-shaped rod and diplococci. d The diplococci were Methanosarcina as identified by a specific probe for in situ hybridization (Cy3/red-MX821). e Baltic-Methanosarcina sometimes also formed tetrads and could never be visualized as single cells in these Fe0-dependent cultures. f Epifluorescence micrograph of DAPI-stained cells detached from Fe0-granules by vigorous shaking during day 60 of transfer #10. Only two morphotypes were observed—banana-shaped rods and cocci joined in large aggregates. g Methanosarcina formed cocci aggregates which could be detected by their natural F420-autofluorescence. No other morphotypes of methanogens could be detected. h and i Methanosarcina- aggregates with various morphologies and compactness. j Group-specific qPCR to determine the abundance of Methanosarcina and Sporomusa after 18 and 60 days of incubation on Fe0
Methanogens identified by MGS and 16S rRNA gene libraries belonged to Methanomicrobia represented by the genera Methanosarcina, Methanocalculus, and Methanocella. Because the Baltic-methanogens could not utilize H2 (1.5 atm; Fig. 4) for methanogenesis, we expected Methanosarcina to be the dominant member of the methanogenic community. As such, Methanosarcina became 26 times more abundant at the end of the incubation period (day 60) than they were during the first stage of incubation (day 18; Fig. 7j). During the early stages of Fe0-dependent growth (18 days), Methanosarcina cells formed diplococci (Fig. 7d) or tetrads (Fig. 7e), but after 2 months of incubation, multicellular aggregates could be visualized by F420-autofluorescence specific for methanogens (Fig. 7g–i). No other methanogenic morphotypes were observed with F420-autofluorescence [54], indicating that Methanosarcina was the dominant methanogen.
Methanosarcina is the only known genus that includes species incapable of methanogenesis from H2 [22, 25, 26], acetate [71], or both [23, 24]. Additionally, Methanosarcina includes species capable of direct electron uptake from electrodes [29, 30] and other cells either directly [29, 31, 32], or via conductive minerals [29, 31,32,33,34, 72]. The closest relative of Baltic-Methanosarcina was the non-acetoclastic and non-hydrogenotrophic M. subterranea (Fig. 6c). The inability of their closest relative to use H2 and acetate aligns with physiological evidence that Baltic-Methanosarcina was also incapable of methanogenesis using these substrates (Figs. 1c, 4). Conclusively, Baltic-Methanosarcina was unlikely to consume acetate produced by Baltic-acetogens. Hence, our results contest previous suppositions that Methanosarcina and acetogens mainly interact syntrophically, via acetate-transfer, within a corrosive microbial community [73]. Instead, we provide evidence that Baltic-Methanosarcina were more metabolically active on Fe0 in the absence of bacterial partners (Fig. 2e). Indeed, the methanogens appeared to compete with acetogens to access Fe0, since Fe0-dependent methanogenesis decreased approximately threefold when acetogens were active (Fig. 2d, e).
Moreover, our results indicate that Baltic-Methanosarcina might be using a direct mechanism of electron uptake from Fe0, since they could not use H2, independent of its origin (Figs. 4 and 5b). Extracellular enzyme-facilitated Fe0 corrosion has only been demonstrated in Methanococcus species [16, 19], while Methanosarcina species were thought to retrieve electrons from Fe0 via abiotic H2 uptake [21]. However, some Methanosarcina cannot use H2 at all [22,23,24,25,26], while others have high H2-uptake thresholds (296–376 nM) [27, 28]. Therefore, when using abiotic or enzymatic H2 Methanosarcina should be outcompeted by strict hydrogenotrophic methanogens with low H2-uptake thresholds (e.g. 6 nM for Methanobacterium formicicum) [27, 28], yet this was not the case in our enrichments indicating they may use an alternative electron uptake mechanism. Similar to other studies Methanosarcina, rather than strict hydrogenotrophic methanogens dominated the Fe0 corroding community, suggesting that Methanosarcina can in fact outcompete strict hydrogenotrophic methanogens from a corrosive community. The mechanism of direct electron uptake from Fe0 or any other insoluble electron donors employed by Methanosarcina is unknown. Extracellular electron uptake in Methanosarcina has been examined recently using a comparative transcriptomics approach contrasting Methanosarcina provided either with electrons from a current-producing syntrophic partner (electrogenic Geobacter [31, 32]) or with H2 from a H2-producing syntrophic partner (Pelobacter) [35]. During extracellular electron uptake from an electrogenic bacterium, Methanosarcina up-regulated cell-surface proteins with redox properties, such as cupredoxins, cytochromes, and other Fe-S-proteins [35]. However, the exact role of these redox-active proteins in Methanosarcina’s extracellular electron uptake from insoluble extracellular electron donors (Fe0, other cells or electrically conductive particles) remains enigmatic and requires future exploration.
Conclusion
Methanosarcina and acetogens often cohabit on the surface of corroded Fe0-structures from low-sulfate environments. However, the role of Methanosarcina was assumed to be commensal, feeding on the acetate produced by acetogens. Our results demonstrate that Baltic-Methanosarcina does not establish a syntrophic partnership with acetogens based on acetate transfer as often reported. Instead, Baltic-Methanosarcina and Baltic-Sporomusa competed with one another to reclaim electrons from Fe0, and each group became favored when specific inhibitors for their competitors were added to the medium. While Baltic-acetogens seem to be stimulated by enzymes/shuttles from spent filtrate, Methanosarcina were not. Moreover, Baltic-Methanosarcina were unable to utilize acetate and H2 as electron donors, suggesting that they may be retrieving electrons directly via a largely unexplored mechanism.
References
AlAbbas FM, Bhola R, Spear JR, Olson DL, Mishra B. Electrochemical characterization of microbiologically influenced corrosion on linepipe steel exposed to facultative anaerobic Desulfovibrio sp. Int J Electrochem Sci. 2013;8:859–71.
Hays GF. World Corrosion Organization. CORRODIA—NACE European Board. http://events.nace.org/euro/corrodia/Fall_2010/wco.asp.
Medvedeva N, Polyak Y, Kankaanpää H, Zaytseva T. Microbial responses to mustard gas dumped in the Baltic Sea. Mar Environ Res. 2009;68:71–81.
Tornero V, Hanke G. Chemical contaminants entering the marine environment from sea-based sources: a review with a focus on European seas. Mar Pollut Bull. 2016;112:17–38.
Enning D, Garrelfs J. Corrosion of iron by sulfate-reducing bacteria: new views of an old problem. Appl Environ Microbiol. 2014;80:1226–36.
Egger M, Rasigraf O, Sapart CJ, Jilbert T, Jetten MSM, Röckmann T, et al. Iron-mediated anaerobic oxidation of methane in brackish coastal sediments. Environ Sci Technol 2014;49:277–83.
Kato S, Yumoto I, Kamagata Y. Isolation of acetogenic bacteria that induce biocorrosion by utilizing metallic iron as the sole electron donor. Appl Environ Microbiol. 2015;81:67–73.
Mand J, Park HS, Jack TR, Voordouw G. The role of acetogens in microbially influenced corrosion of steel. Front Microbiol. 2014;5:1–14.
Zhang T, Fang HHP, Ko BCB. Methanogen population in a marine biofilm corrosive to mild steel. Appl Microbiol Biotechnol. 2003;63:101–6.
Zhu XY, Lubeck J, Kilbane JJ. Characterization of microbial communities in gas industry pipelines. Appl Environ Microbiol. 2003;69:5354–63.
Lee JS, Mcbeth JM, Ray RI, Little BJ, Emerson D. Iron cycling at corroding carbon steel surfaces. Biofouling. 2013;29:1243–52.
Rajala P, Bomberg M, Vepsäläinen M, Carpén L. Microbial fouling and corrosion of carbon steel in deep anoxic alkaline groundwater. Biofouling. 2017;33:195–209.
Uchiyama T, Ito K, Mori K, Tsurumaru H, Harayama S. Iron-corroding methanogen isolated from a crude-oil storage tank. Appl Environ Microbiol. 2010;76:1783–8.
Mori K, Tsurumaru H, Harayama S. Iron corrosion activity of anaerobic hydrogen-consuming microorganisms isolated from oil facilities. J Biosci Bioeng. 2010;110:426–30.
Dinh HT, Kuever J, Mußmann M, Hassel AW, Mussman M, Hassel AW, et al. Iron corrosion by novel anaerobic microorganisms. Nature. 2004;427:829–32.
Lienemann M, Deutzmann JS, Milton RD, Sahin M, Spormann AM. Mediator-free enzymatic electrosynthesis of formate by the Methanococcus maripaludis heterodisulfide reductase supercomplex. Bioresour Technol. 2018;254:278–83.
Beese-Vasbender PF, Grote J-P, Garrelfs J, Stratmann M, Mayrhofer KJ. Selective microbial electrosynthesis of methane by a pure culture of a marine lithoautotrophic archaeon. Bioelectrochemistry. 2015;102:50–5.
Tsurumaru H, Ito N, Mori K, Wakai S, Uchiyama T, Iino T, et al. An extracellular [NiFe] hydrogenase mediating iron corrosion is encoded in a genetically unstable genomic island in Methanococcus maripaludis. Sci Rep. 2018;8:15149.
Deutzmann JS, Merve S, Spormann AM, Jörg D, Spormann AM. Extracellular enzymes facilitate electron uptake in biocorrosion and bioelectrosynthesis. MBio. 2015;6:1–8.
Brileya KA, Connolly JM, Downey C, Gerlach R, Fields MW. Taxis toward hydrogen gas by Methanococcus maripaludis. Sci Rep. 2013;3:3140.
Daniels L, Belay N, Rajagopal B, Weimer P. Bacterial methanogenesis and growth from CO2 with elemental iron as the sole source of electrons. Science. 2019;237:509–11.
Sowers KR, Baron SF, Ferry JG. Methanosarcina acetivorans sp. nov., an acetotrophic methane-producing bacterium isolated from marine sediments. Appl Environ Microbiol. 1984;47:971–8.
Ni S, Boone DR. Isolation and characterization of a dimethyl sulfide-degrading methanogen, Methanolobus siciliae HI350, from an oil well, characterization of M. siciliae T4/MT, and emendation of M. siciliae. Int J Syst Bacteriol. 1999;41:410–6.
Lyimo TJ, Pol A, Op den Camp H, Harhangi HR, Vogels GD. Methanosarcina semesiae sp. nov., a dimethylsulfide-utilizing methanogen from mangrove sediment. Int J Syst Evol Microbiol. 2000;50:171–8.
Von KleinD, Arab H, Völker H. Methanosarcina baltica, sp. nov., a novel methanogen isolated from the Gotland Deep of the Baltic Sea. Extremophiles. 2002;6:103–10.
Shimizu S, Upadhye R, Ishijima Y, Naganuma T. Methanosarcina horonobensis sp. nov., a methanogenic archaeon isolated from a deep subsurface miocene formation. Int J Syst Evol Microbiol. 2011;61:2503–7.
Kral TA, Brink KM, Miller SL, McKay CP. Hydrogen consumption by methanogens on the early Earth. Orig Life Evol Biosph. 1998;28:311–9.
Lovley DR. Minimum threshold for hydrogen metabolism in methanogenic bacteria. Appl Environ Microbiol. 1985;49:1530–1.
Yee MO, Snoeyenbos-West OL, Thamdrup B, Ottosen LDM, Rotaru A-E. Extracellular electron uptake by two Methanosarcina species. Front Energy Res. 2019;7:458091.
Rowe AR, Xu S, Gardel E, Bose A, Girguis P, Amend JP, et al. Methane-linked mechanisms of electron uptake from cathodes by Methanosarcina barkeri. mBio. 2019;10:e02448–18.
Rotaru A-E, Shrestha PM, Liu F, Markovaite B, Chen S, Nevin KP, et al. Direct interspecies electron transfer between Geobacter metallireducens and Methanosarcina barkeri. Appl Environ Microbiol. 2014;80:4599–605.
Rotaru A-E, Woodard TL, Nevin KP, Lovley DR. Link between capacity for current production and syntrophic growth in Geobacter species. Front Microbiol. 2015;6:744.
Liu F, Rotaru A-EE, Shrestha PM, Malvankar NS, Nevin KP, Lovley DR. Promoting direct interspecies electron transfer with activated carbon. Energy Environ Sci. 2012;5:8982.
Chen S, Rotaru A-E, Shrestha PM, Malvankar NS, Liu F, Fan W, et al. Promoting interspecies electron transfer with biochar. Sci Rep. 2014;4:5019.
Holmes DE, Rotaru A, Ueki T, Shrestha PM, James G, Lovley DR. Electron and proton flux for carbon dioxide reduction in Methanosarcina barkeri during direct interspecies electron transfer. Front Microbiol. 2018;9:1–11.
Mehanna M, Basseguy R, Delia ML, Girbal L, Demuez M, Bergel A. New hypotheses for hydrogenase implication in the corrosion of mild steel. Electro Acta. 2008;54:140–7.
Mehanna M, Rouvre I, Delia M-L, Feron D, Bergel A, Basseguy R. Discerning different and opposite effects of hydrogenase on the corrosion of mild steel in the presence of phosphate species. Bioelectrochemistry. 2016;111:31–40.
Karyakin AA, Morozov SV, Karyakina EE, Varfolomeyev SD, Zorin NA, Cosnier S. Hydrogen fuel electrode based on bioelectrocatalysis by the enzyme hydrogenase. Electrochem commun. 2002;4:417–20.
Partridge CDP, Yates MG. Effect of chelating agents on hydrogenase in Azotobacter chroococcum. Evidence that nickel is required for hydrogenase synthesis. Biochem J. 1982;204:339–44.
Rotaru A-E, Calbrese F, Stryhanyuk H, Musat F, Shrestha PM, Weber HS, et al. Conductive particles enable syntrophic acetate oxidation between Geobacter and Methanosarcina from coastal sediments. mBio. 2018;49:1–14.
Zhou Z, Meng Q, Yu Z. Effects of methanogenic inhibitors on methane production and abundances of methanogens and cellulolytic bacteria in in vitro ruminal cultures. Appl Environ Microbiol. 2011;77:2634–9.
Lohner ST, Deutzmann JS, Logan BE, Leigh J, Spormann AM. Hydrogenase-independent uptake and metabolism of electrons by the archaeon Methanococcus maripaludis. ISME J. 2014;8:1673–81.
Lovley DR, Phillips EJP. Rapid assay for microbially reducible ferric iron in aquatic sediments. Appl Environ Microbiol. 1987;53:1536–40.
Canfield DE, Raiswell R, Westrich JT, Reaves CM, Berner RA. The use of chromium reduction in the analysis of reduced inorganic sulfur in sediments and shales. Chem Geol. 1986;54:149–55.
Meyer F, Paarmann D, D’Souza M, Olson R, Glass E, Kubal M, et al. The metagenomics RAST server—a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinforma. 2008;9:386.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, et al. The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res. 2012;41:D590–6.
Cole JR, Wang Q, Fish JA, Chai B, McGarrell DM, Sun Y, et al. Ribosomal Database Project: data and tools for high throughput rRNA analysis. Nucleic Acids Res. 2014;42:D633–42.
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol. 2006;72:5069–72.
Tatusova T, Ciufo S, Federhen S, Fedorov B, McVeigh R, O’Neill K, et al. Update on RefSeq microbial genomes resources. Nucleic Acids Res. 2015;43:D599–605.
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, et al. Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics. 2012;28:1647–9.
Stamatakis A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics. 2014;30:1312–3.
Loescher CR, Großkopf T, Desai FD, Gill D, Schunck H, Croot PL, et al. Facets of diazotrophy in the oxygen minimum zone waters off Peru. ISME J. 2014;8:2180–92.
Pernthaler J, Glöckner FO, Schönhuber W, Amann R. Fluorescence in situ hybridization with rRNA-targeted oligonucleotide probes. Methods Microbiol. 2001;30:1–31.
Doddema HJ, Vogels GD. Improved identification of methanogenic bacteria by fluorescence microscopy. Appl Environ Microbiol. 1978;36:752–4.
Philips J, Monballyu E, Georg S, De Paepe K, Prévoteau A, Rabaey K, et al. An Acetobacterium strain isolated with metallic iron as electron donor enhances iron corrosion by a similar mechanism as Sporomusa sphaeroides. FEMS Microbiol Ecol. 2019;95:1–13.
Usher KM, Kaksonen aH, Bouquet D, Cheng KY, Geste Y, Chapman PG, et al. The role of bacterial communities and carbon dioxide on the corrosion of steel. Corros Sci. 2015;98:354–65.
Ingham B, Ko M, Laycock N, Kirby NM, Williams DE. First stages of siderite crystallisation during CO2 corrosion of steel evaluated using in situ synchrotron small- and wide-angle X-ray scattering. Faraday Discuss. 2015;180:171–90.
Noubactep C, Schöner A. Fe0-based alloys for environmental remediation: Thinking outside the box. J Hazard Mater. 2009;165:1210–4.
Venzlaff H, Enning D, Srinivasan J, Mayrhofer KJJ, Hassel AW, Widdel F, et al. Accelerated cathodic reaction in microbial corrosion of iron due to direct electron uptake by sulfate-reducing bacteria. Corros Sci. 2013;66:88–96.
Nevin KP, Woodard TL, Franks AE. Microbial electrosynthesis: feeding microbes electricity to convert carbon dioxide and water to multicarbon extracellular organic compounds. mBio. 2010;1:1–4.
Nevin KP, Hensley SA, Franks AE, Summers ZM, Ou J, Woodard TL, et al. Electrosynthesis of organic compounds from carbon dioxide is catalyzed by a diversity of acetogenic microorganisms. Appl Environ Microbiol. 2011;77:2882–6.
Zaybak Z, Logan BE, Pisciotta JM. Electrotrophic activity and electrosynthetic acetate production by Desulfobacterium autotrophicum HRM2. Bioelectrochemistry. 2018;123:150–5.
Peters JW, Schut GJ, Boyd ES, Mulder DW, Shepard EM, Broderick JB, et al. [FeFe]- and [NiFe]-hydrogenase diversity, mechanism, and maturation. Biochim Biophys Acta. 2015;1853:1350–69.
Milton RD, Ruth JC, Deutzmann JS, Spormann AM. Methanococcus maripaludis employs three functional heterodisulfide reductase complexes for flavin-based electron bifurcation using hydrogen and formate. Biochemistry. 2018;57:4848–57.
Sass H, Ramamoorthy S, Yarwood C, Langner H, Schumann P, Kroppenstedt RM, et al. Desulfovibrio idahonensis sp. nov., sulfate-reducing bacteria isolated from a metal(loid)-contaminated freshwater sediment. Int J Syst Evol Microbiol. 2009;59:2208–14.
Li D, Li Z, Yu J, Cao N, Liu R, Yang M. Characterization of bacterial community structure in a drinking water distribution system during an occurrence of red water. Appl Environ Microbiol. 2010;76:7171–80.
Brunner B, Arnold GL, Røy H, Müller IA, Jørgensen BB. Off limits: sulfate below the sulfate–methane transition. Front Earth Sci. 2016;4:75.
Pester M, Knorr KH, Friedrich MW, Wagner M, Loy A. Sulfate-reducing microorganisms in wetlands - fameless actors in carbon cycling and climate change. Front Microbiol. 2012;3:1–19.
Olivier B, Cord-Ruwisch R, Hatchikian E, Garcia J. Characterization of Desulfovibrio fructosovorans sp. nov. Arch Microbiol. 1988;149:447–50.
Bryant MP, Campbell LL, Reddy CA, Crabill MR. Growth of Desulfovibrio in lactate or ethanol media low in sulfate in association with H2-utilizing methanogenic bacteria. Appl Environ Microbiol. 1977;33:1162–9.
Simankova MV, Parshina SN, Tourova TP, Kolganova TV, Zehnder AJB, Nozhevnikova AN. Methanosarcina lacustris sp. nov., a new psychrotolerant methanogenic archaeon from anoxic lake sediments. Syst Appl Microbiol. 2001;24:362–7.
Wang O, Zheng S, Wang B, Wang W. Necessity of electrically conductive pili for methanogenesis with magnetite stimulation. PeerJ. 2018;2:1–14.
Skovhus LT, Enning D, Lee JS. Microbiologically influenced corrosion in the upstream oil and gas industry. Boca Raton, FL: CRC Press; 2017.
Acknowledgements
This work is a contribution to a Sapere Aude Danish Research Council grant number 4181-00203 awarded to A-ER. We want to thank E. Laursen, L. Ørum Smidt, H. Grøn Jensen, and D. Holmgaard Skov for lab assistance. We thank J. Ward for assistance with SEM preparation and the microscopy facility unit at the University of Massachusetts Amherst who provided us with access to scanning electron microscopy. We thank the Institute of Clinical Molecular Biology in Kiel for Sanger sequencing partly supported by the DFG Clusters of Excellence “Inflammation at Interfaces” and “Future Ocean” and the ICMB-Kiel technicians S. Greve, T. Henke, and C. Noack for lab assistance. We thank Prof. Dawn Holmes and Assist. Prof. Nicole Posth for editing the final version of this manuscript.
Author information
Authors and Affiliations
Contributions
PAP and A-ER designed the experiments. A-ER carried out sampling, and processing of the Baltic Sea sediment, as well as the initial incubations and in situ hybridization experiments. BT carried out elemental identification and analyses of corrosion products. PAP carried all downstream growth experiments and analyses. PAP did all molecular experiments and analyses with support from A-ER, CRL, and OS-W. PAP and A-ER wrote the manuscript, and all authors contributed to the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Palacios, P.A., Snoeyenbos-West, O., Löscher, C.R. et al. Baltic Sea methanogens compete with acetogens for electrons from metallic iron. ISME J 13, 3011–3023 (2019). https://doi.org/10.1038/s41396-019-0490-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-019-0490-0
This article is cited by
-
MmcA is an electron conduit that facilitates both intracellular and extracellular electron transport in Methanosarcina acetivorans
Nature Communications (2024)
-
Anaerobic degradation of organic carbon supports uncultured microbial populations in estuarine sediments
Microbiome (2023)
-
Microbially induced corrosion impacts on the oil industry
Archives of Microbiology (2022)
-
Stainless steel corrosion via direct iron-to-microbe electron transfer by Geobacter species
The ISME Journal (2021)