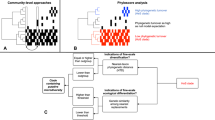
Abstract
Microdiversity can lead to different ecotypes within the same species. These are assumed to provide stability in time and space to those species. However, the role of microdiversity in the stability of whole microbial communities remains underexplored. Understanding the drivers of microbial community stability is necessary to predict community response to future disturbances. Here, we analyzed 16S rRNA gene amplicons from eight different temperate bog lakes at the 97% OTU and amplicon sequence variant (ASV) levels and found ecotypes within the same OTU with different distribution patterns in space and time. We observed that these ecotypes are adapted to different values of environmental factors such as water temperature and oxygen concentration. Our results showed that the existence of several ASVs within a OTU favored its persistence across changing environmental conditions. We propose that microdiversity aids the stability of microbial communities in the face of fluctuations in environmental factors.
Similar content being viewed by others
Introduction
Microbial communities have a profound effect on the planet’s surface and atmosphere. These communities are composed of hundreds to thousands of different species of microbes, but the nature of the relationships between microbial diversity and community and ecosystem functioning remains one of the big unanswered questions in general ecology [1]. Subtle relationships between the environment and microbial species can affect microbial community structure and therefore ecosystem-level processes [2]. Moreover, high diversity has been tied to resilience in some microbial communities [3, 4].
Traditionally, species have been considered to be the units of diversity. In the case of microorganisms, the formal definition of a species requires isolation in pure culture, phenotypic characterization, and genome sequencing [5]. Nevertheless, the definition of a microbial species is an extensively debated issue in microbiology [6, 7]. Furthermore, the usefulness of species as the most significant unit in microbial ecology is also controversial [8,9,10] because two individuals classified within the same species may carry out different functions in the environment [11, 12]. Each species is formed by several strains that share a core genome and differ in the accessory genome. The term “pan-genome” has been coined to refer to all the genes that can be found in the known strains of a species, including the core and all the accessory genes [13]. Variations in the accessory genome are assumed to be responsible for niche differentiation and allow the discrimination of different ecotypes, pragmatically defined as populations of cells adapted to a given ecological niche [14,15,16]. When dealing with complex microbial communities, 16S ribosomal RNA (rRNA) gene profiling allows to perform a preliminary characterization of diversity and population dynamics without having to resort to whole-metagenome sequencing. This technique gives no information on the genome content of the taxa present in the community, but has a high taxonomic resolution and can nonetheless be useful to detect groups of taxa associated with different environmental variables. The analysis of 16S rRNA data has traditionally involved the definition of operational taxonomic units (OTUs). Commonly, OTUs have been defined at the 97% similarity level (assumed to correspond approximately to species), although an increasing number of authors have proposed higher cutoffs [17].
This perceived necessity for higher taxonomic resolutions was countered by the concern that, at very high similarity cutoffs, sequencing errors would lead to artifactual clusters [18]. Recent methodological developments, however, have overcome this limitation [19,20,21,22,23,24], allowing the discrimination of amplicon sequence variants (ASVs) with only one nucleotide difference in their 16S rRNA gene sequence. Analyzing these ASVs may reveal the distribution of ecotypes across environments, unveiling previously overlooked ecological patterns [19, 25,26,27].
The existence of ecotypes with overlapping ecological functions (but adapted to different niches) has been proposed to confer stability to microbial ecosystems [28], guaranteeing the long-lasting persistence of bacterial populations [29]. Several species have been shown to consist of many different subpopulations that sustain their distribution across broad environmental gradients [12, 25, 26, 30]. However, it is still not clear whether this microdiversity also contributes to the preservation of the higher-order inter-species interactions that are fundamental for community functioning and stability. Stability has been studied extensively [31,32,33], especially in relation with diversity [32, 34]. Diverse communities will contain functionally equivalent organisms able to respond to different perturbations, resulting in increased stability [32]. The stability of microbial communities is also influenced by the type and strength of ecological interactions among their members. Species capable of forming versatile interactions with multiple partners will be less affected by variations in community composition; a community dominated by such weak interactions will thus be more resilient to change [32]. Understanding the drivers of microbial community stability is necessary in order to predict community response to future disturbances (for a thorough review see ref. [35]), such as those resulting from global change.
Here, we examined the identity and distribution of 97% similarity OTUs and ASVs using a large dataset of samples collected in eight different temperate bog lakes in Wisconsin, U.S.A. [36, 37]. These lakes are located in the Northern Highlands, a rocky, high elevation (for Wisconsin) area with a high density of lakes. These bog lakes are in the southernmost edge of the North American boreal biome. Lakes have been studied intensively [38,39,40,41,42] because they play an important role in human activities, and can be used as sentinels and integrators of environmental change [43]. Besides, the fact that lakes are relatively similar to each other, yet differ in environmental parameters such as temperature, oxygen, or mixing regimes, make these systems fitting to address core questions in microbial ecology such as how community assembly takes place, how interactions among taxa influence community composition, or how microbial communities respond to environmental perturbations [3, 37, 44]. To shed light on these questions, we took advantage of the natural environmental variability typical of lacustrine systems to distinguish ecotypes within microbial communities. We compared OTUs with ASVs and examined whether ASVs corresponded to different ecotypes associated with environmental variables such as mixing regime, thermal layer, water temperature, or dissolved oxygen. We also studied the dynamics of these ecotypes across space and time and their influence on the stability of the whole community.
Materials and methods
Samples
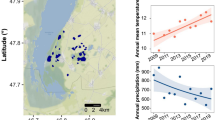
We analyzed a set of 1505 samples from eight different temperate bog lakes mainly during the ice-free periods of 2005, 2007, 2008, and 2009 [37]. Lakes are located in Wisconsin (U.S.A.) (Table 1 and Fig. S1) (see ref. [37] for more information). These lakes vary by mixing regime; three are polymictic, three dimictic, and two meromictic. Integrated samples were taken from the epilimnion and hypolimnion.
Sequence data used in this study were publicly available in the European Bioinformatics Institute with accession number ERP016854 [37]. The metadata linked to these samples were available in the MG-RAST database (accession code: mgp127) and in the McMahon Lab GitHub repository “North_Temperate_Lakes-Microbial_Observatory” (https://github.com/McMahonLab/North_Temperate_Lakes-Microbial_Observatory). The 150 base pairs sequences corresponded to amplicons from the V4 region of the 16S rRNA gene, which was amplified and sequenced using the Illumina HiSeq platform. A complete description of the sample collection and processing protocols can be found in refs. [36, 37].
High-resolution microbial diversity
We quality-filtered raw sequences using moira [45], truncating them to 140 bp and discarding every sequence with a probability > 0.005 of having more than one error according to its quality scores (maxerrors = 1, alpha = 0.005). Next, we processed the sequences using DADA2 [22] to distinguish amplicon sequence variants (ASVs). The SILVA nr v.128 database [46] was used for taxonomic assignment and removing non-bacterial sequences. Sequences were also classified with the FreshTrain database [41] using mothur v.1.39.1 [47]. Samples with <10,000 sequences or without associated metadata were removed, and the remaining samples were rarefied to 10,000 sequences. After this step, we retained 26,128 ASVs from 1108 samples. More detailed information about the successive steps of the procedure can be found in Table S1. Additionally, we clustered the ASVs into 97% similarity OTUs using the Opticlust algorithm [48]. This resulted in 13,940 OTUs (Data S1). For the sake of brevity, we will simply use the terms “OTU” (for 97% similarity OTUs) and “ASV” throughout the manuscript, even although ASVs also fall within the definition of operational taxonomic units.
Statistics
We normalized the data with a Hellinger transformation. This transformation is particularly suited to abundance data because it gives low weights to variables (OTUs or ASVs) with low counts and many zeros [49, 50]. Then, we performed a detrended correspondence analysis (DCA) [51] using the relative abundance of OTUs or ASVs across epilimnion and hypolimnion samples. Procrustes analysis was used to compare both DCAs [52].
Finally, we tested whether the samples clustered significantly by lake or mixing regime using Bray–Curtis dissimilarities calculated from the non-normalized OTU and ASV abundance tables and applying a PERMANOVA test (adonis). All statistical analyses were done with R version 3.4.4 and the package vegan [53].
Environmental distribution of ASVs
To determine the environmental distribution (lakes and layers) for each ASV, we calculated their mean abundances in each lake-layer combination and standardized them to z-scores. This normalization facilitated visualization of the habitat distribution for low-abundance ASVs.
Influence of microdiversity on OTU persistence and variability
The effective microdiversity was calculated as follows. First, we selected the OTUs with a global abundance higher-than 5000 counts, as the Shannon diversity index has been shown to lose accuracy for abundance values below that threshold [54]. For each OTU, we calculated the global abundances of its constituent ASVs and rarefied the resulting vector to 5000 counts. This was done in order to avoid OTUs with higher global abundances being assigned a higher microdiversity. Finally, for each OTU, the effective microdiversity was calculated from the rarefied abundances of its constituent ASVs as the exponential of the Shannon index [55]. This value represents the effective number of ASVs in a given OTU, and its calculation is conceptually analogous to the calculation of the effective number of species in a community, but using ASVs as units of diversity and restricting the sampling space to each individual OTU.
For each OTU with a global abundance above 5000, we also rarefied its abundance to 5000 counts and calculated its persistence (fraction on samples in which the taxon had at least one count) and variability (coefficient of variation of the abundance of the taxon across the samples), as described in ref. [37]. Rarefaction was used to decouple persistence and variability from global abundance. We then tested for differences in persistence and variability between OTUs with low (less than two effective ASVs) and high (two or more effective ASVs) microdiversity via Welch’s t-tests.
OTU and ASV distribution modeling
We selected the most stable OTUs (global abundance > 5000, persistence > 0.75 and variability < 2), and their constituent ASVs (global abundance > 1000). OTU and ASV distributions in samples were modeled as a function of lake, year, month, depth, temperature, and dissolved oxygen concentration using boosted regression trees. This technique allows to model the effect of multiple predictive variables, works well with a variety of response types and is robust to missing values and outliers [56]. The gbm.step function from the dismo R package [56] was used to select an optimal number of boosting trees with the following parameters: tree.complexity = 5, n.trees = 1000, max.trees = 20,000, step.size = 10, bag.fraction = 0.75, learning.rate = 0.001. The influence of each environmental variable on the distribution of each OTU/ASV was measured as its relative influence on reducing the loss function during model computation. OTUs with only one high-abundance ASV and models with a cross-validation correlation smaller than 0.6 were excluded from further analyses.
We also tested whether temperature and oxygen influenced the abundance ratios between ASVs belonging to the same clades. For each lake, we selected three ecologically relevant ubiquitous freshwater clades (Polynucleobacter PnecC, Actinobacteria acI-B2 and Bacteroidetes bacI; Data S2) and the three most abundant ASVs from each one. These clades were described as key members of a model freshwater community identified by Garcia et al. [57]. For each possible pair of ASVs from the same clade (1–2, 1–3, and 2–3), we then fitted a linear model of their abundance ratio (in log scale) against temperature separately for each lake. The resulting p-values were corrected for multiple testing with the Bonferroni method. For each lake and clade, the two ASVs with a higher correlation between abundance ratio and either temperature or oxygen were selected for plotting.
Results
Diversity and dynamics of the bacterioplankton
First, we analyzed the bacterial diversity in the lakes at the OTU level (Fig. S2). The most abundant OTUs across all lakes belonged to the phyla Proteobacteria, Actinobacteria, Acidobacteria, Bacteroidetes, and Verrucomicrobia. The most prevalent OTU was the Betaproteobacteria Polynucleobacter necessarius (PnecC), which had a high relative abundance in all the lakes, followed by Actinobacteria acI-B2. Lakes with the same mixing regime shared the same abundant taxa through the seasons (Fig. S2). For instance, Polynucleobacter PnecC and Actinobacteria acI-B2 were the OTUs with the highest relative abundance in polymictic and dimictic lakes, but they were accompanied by Actinobacteria acI-B3 and Verrucomicrobia in polymictic, and Gammaproteobacteria and Acidobacteria in dimictic lakes. In contrast, these OTUs were less abundant in meromictic lakes.
To evaluate the similarity in community composition among samples, we applied a detrended correspondence analysis (DCA) to both the OTUs and ASVs abundance data, distinguishing between epilimnion and hypolimnion (Fig. 1). At both the OTU and ASV levels, the samples clustered by lake and mixing regime (PERMANOVA significance test p < 0.001, Table S2). Samples from meromictic lakes were very different from the others, accounting for most of the variance. In contrast, samples from the other two mixing regimes were compositionally more similar. Samples for WSB clustered independently from the other polymictic lakes. This lake is larger in surface area and deeper than the other two polymictic lakes, which may explain the differences in community composition.
Detrended correspondence analysis (DCA) of the microbial community composition in OTUs and ASVs for the epilimnia and hypolimnia of the different lakes. Each point represents a sample. Lakes are indicated by point color. Procrustes analysis in the middle column shows the differences between the OTU and ASV-level ordinations
The clustering of samples with OTUs and with ASVs exhibited good agreement, as shown by Procrustes analysis (Fig. 1), since samples clustered by lake of origin in both cases. Information from ASVs, however, suggested finer distinctions in community composition. For example, hypolimnion samples from lakes HK and MA clustered together in the OTU analysis, but they were separated along DCA component 2 when using ASVs (Fig. 1).
To search for possible seasonal patterns, we also plotted time-decay curves for the four lakes that were sampled for two or more consecutive years (Fig. 2; Lakes MA, NSB, TB, and SSB). Regardless of whether we considered OTUs or ASVs, Bray–Curtis dissimilarities were maximal for pairs of samples from opposite seasons (i.e. separated 6, 18, 30 months) and minimal for pairs of samples collected at the same time of the year (i.e., separated 12, 24, 36 months). However, even in the latter case, Bray–Curtis dissimilarities were relatively high (0.521 for ASVs and 0.462 for OTUs), showing that, although seasonal trends could be discerned, there was high inter-year variability in community composition.
ASVs from the same OTU show different habitat distributions
To explore the ecological significance of microdiversity, we analyzed the distribution and habitat preferences of ASVs within the OTUs with the highest relative abundance in the samples. The upper row of radar charts in Fig. 3 and Fig. S3 shows the average abundance of ASVs from each of the most abundant OTUs in different lakes (color-coded inner ring) and layers (grayscale-coded outer ring). OTU-1 (Polynucleobacter PnecC), which had the highest relative abundance in the dataset, comprises three main ASVs (ASV-1, ASV-2, and ASV-48). ASV-1 was more abundant than ASV-2 in the hypolimnion of NSB, but less abundant in the epilimnion of FB. Both ASVs, in turn, were always more abundant than ASV-48. To determine the habitat preferences of ASVs irrespective of their abundance, z-scores were plotted in the lower row of radar charts (Fig. 3). Positive z-scores represent lake/layer combinations in which the ASV had higher-than-average abundance, and vice-versa. The three most abundant ASVs from Polynucleobacter PnecC showed different distributions: ASV-1 showed a higher relative abundance in the hypolimnion of dimictic lakes, ASV-48 in the epilimnion of polymictic lakes and in the epilimnion of NSB, and ASV-2, although present in most lakes, was most abundant in FB. Differences in the distribution of ASV-1 and ASV-2 in different lakes can be seen in Fig. S4. While ASV-2 was prevalent in polymictic lakes, ASV-1 and ASV-2 coexisted in the remaining lakes.
ASVs from the same OTU show different habitat distributions. Distribution and habitat preference of four relevant OTUs, which are further discussed in the text. OTUs 1, 2, and 7 correspond to the ubiquitous freshwater clades PnecC, acI-B2, and bacI-A, while OTU-3 corresponds to a bloomer from the gamI clade. The radar charts show a the abundances (over 10,000 counts, logarithmic scale) and b z-scores of the three most abundant ASVs for each of the four OTUs. The colors of the inner-ring represent the mixing regime of the lake: green for polymictic, blue for dimictic, and red for meromictic. The shades of gray in the outer ring indicate the thermal layer (epilimnion and hypolimnion). Zero values have been plotted as log10 (0.01). z-scores indicate the environmental preference of each ASV. Since the z-score normalization was made independently for each ASV, z-scores cannot be compared quantitatively among ASVs. The red dotted line indicates the mean abundance of each ASV, separating positive (white background) and negative (gray background) environmental preferences as schematically represented at the bottom right of the figure
Similar situations were found for the other OTUs. In the case of OTU-2 (Actinobacteria acI-B2), ASV-4 had a higher relative abundance in CB, while ASV-18 tended to appear in the hypolimnion of dimictic and polymictic lakes. ASV-5, on the other hand, was observed less frequently in meromictic lakes, especially in HK, but showed some preference for several epilimnetic samples and the hypolimnion of CB. OTU-3 (Gammaproteobacteria gamI) illustrates a case in which two strains from the same OTU had similar spatial preferences: ASV-3 and ASV-31 both preferred TB. However, ASV-31 was absent from several lakes where ASV-3 was present. Finally, in OTU-7 (Bacteroidetes bacI-A), ASV-29 had a higher relative abundance in CB and NSB, while ASV-44 was more abundant in FB (Fig. 3).
We found similar situations in other OTUs. For instance, Acidobacteria Geothrix (the fifth most abundant OTU) was present in lakes TB and SSB (Fig. S2), but looking at the ASV level, ASV-6 had a higher relative abundance in TB, while ASV-15 did so in SSB (Figs. S3 and S4). OTU-4 (Actinobacteria acI-B3) was present in the three polymictic lakes (Fig. S4). ASV-8 was dominant in FB, but both ASV-8 and ASV-10 co-dominated in the other two lakes (Figs. S3 and S4). In general, abundant OTUs were present in most lakes and layers, but their constituent ASVs showed preferences for only some of them.
ASVs from the same OTU show different temporal dynamics
ASVs from the same OTU not only showed different spatial distributions, but also different temporal dynamics, as exemplified in NSB (Fig. 4). For instance, Polynucleobacter PnecC (OTU-1) had a fairly constant relative abundance during the 2007–2008 period (Fig. 4a). However, its two most abundant ASVs (ASV-1 and ASV-2) changed their relative abundances over time: in general, ASV-1 was more abundant in the hypolimnion and ASV-2 in the epilimnion, but ASV-1 came to dominate also in the epilimnion during Spring 2008, when the surface water was still cold (Fig. 4a). Similar trends were observed in the dimictic lakes TB and SSB (Fig. S4). This suggests that each ASV may be adapted to different water temperatures.
ASVs from the same OTU show different temporal dynamics in NSB. Both plots show the relative abundance of the 20 most abundant ASVs in each mixing layer and year and the relative abundance of the OTUs to which they belong to. Taxa were colored in gray except for OTU-1 and OTU-3, whose dynamics are discussed in detail in the main text. a Temporal abundance pattern of the most abundant persister OTU (OTU-1, PnecC clade), and its corresponding ecotypes with time. b Temporal abundance pattern of the most abundant bloomer OTU (OTU-3, gamI clade) and its corresponding ecotypes with time. In both cases, the middle row shows the temperature profiles along time and depth. Homogeneous vertical temperature profiles correspond to mixing events. In 2008, there was an artificial mixing event from 2nd July to 10th July, when homogeneous temperature was observed. Black vertical lines in the different panels indicate sampling points
Another example was Gammaproteobacteria gamI (OTU-3), which could be assigned to the methanotrophic genus Methylobacter. The OTU could be decomposed into two main ASVs: ASV-31, which bloomed during the fall mixing of 2007, and ASV-3, which remained at an intermediate abundance until the next spring mixing (Fig. 4b). The bloom of ASV-31 was concurrent with a transient oxygen depletion throughout the whole-water column (Fig. S5).
Microdiversity imparts stability at higher taxonomic levels
In general, the OTUs with the highest relative abundance were represented by at least two ASVs with different distributions in space and seasonal dynamics. This points to the existence of different ecotypes with slightly different environmental preferences. We explored whether rich microdiversity was linked to increased stability at the OTU level. To that end, we calculated the effective microdiversity of the OTUs from the abundances of their constituent ASVs. OTUs were rarefied to 5000 counts to account for the effect of abundance in diversity, persistence and variability (see materials and methods). We found out that OTUs with two or more effective ASVs were more persistent (i.e., appeared in a larger proportion of samples) and less variable (i.e., showed a more constant abundance across samples) than OTUs with less than two effective ASVs (Welch’s t-test, p < 0.001 in both cases, Fig. 5). In general, persistence increased and variability decreased with effective microdiversity (linear model p = 0.038 and p = 0.030, respectively), although OTUs restricted to particular lake-layer combinations (and thus having intermediate global persistences and variabilities) could still show a high number of effective ASVs (Fig. S6).
Effect of microdiversity on OTU persistence (top) and variability (bottom). Boxplots show the effect of microdiversity (left: low- vs. high-effective microdiversity; right: full range) in a the persistence and b the variability of OTUs. Significant (Welch’s t-test p < 0.001) differences between OTUs with low- and high-effective microdiversity are denoted with an asterisk. OTUs with a high-effective number of ASVs are indicated in the right part; their habitat distribution can be found in Fig. S6
Environmental factors drive the generation of microdiversity
We were interested in finding whether microdiversification occurred in response to a particular environmental gradient. To test this, we modeled the distribution of the most stable OTUs (persistence > 0.75, variability < 2) and their constituent ASVs across samples as a function of lake, year, month, depth, temperature, and dissolved oxygen concentration, using boosted regression trees [56]. The median cross-validation correlation of our models was 0.79, indicating that the relative abundances of OTUs and ASVs in these lakes could be reasonably modeled from the available parameters. We then compared the contributions of different environmental parameters to the OTU and ASV models (Fig. S7). Temperature and oxygen concentration had a significantly higher influence on the distribution of individual ASVs than on the distribution of their parent OTUs (Welch’s t-test p = 0.041 and p = 0.014, respectively), suggesting that microdiversification in response to temperature and oxygen gradients is a general phenomenon in these freshwater lakes.
Dynamics of a freshwater model community
To identify potential associations between the taxa present in the samples, we further focused on representatives from a freshwater model community identified by Garcia et al. [57]. This community was originally enriched from the dimictic Lake Grosse Fuchskuhle and consisted of four organisms commonly found in freshwater (Polynucleobacter PnecC, two different Actinobacteria and a Bacteroidetes), which were able to grow in a mixed culture presumably by establishing mutually beneficial metabolic interactions [57] (Fig. 6a). Even though each strain had different auxotrophies that precluded axenic isolation, the consortium was stable over time [57], thus revealing a powerful model of mutualism in natural microbial communities. However, as noted by the authors, mixed cultures provide limited information on how the system may evolve under changing environmental conditions [57]. We therefore sought to identify members of that model community in the dataset and characterize their dynamics over time.
Microdiversity provides community stability across environmental gradients in freshwater ecosystems. a Freshwater model community proposed by García et al. [57]. b Amplicon sequence variants (ASVs) from the different member clades of the model community, classified according to their environmental preferences. c Relative abundance of low-temperature/oxygen (LTO) vs. high-temperature/oxygen (HTO) ecotypes of the model community in dimictic lake samples, plotted as a function of water temperature. Symbol color indicates dissolved oxygen concentration in shades of blue. Symbol shape indicates sampling year. Samples above the dashed red line are dominated by LTO ecotypes, and vice-versa. Significant linear regressions (FDR < 0.05) and 95% confidence intervals are shown with a red line and a shaded gray area, respectively
Of the four members, Polynucleobacter PnecC, Actinobacteria acI-B2 and a Bacteroidetes from the Cytophagaceae/bacI clade were found in high relative abundances in the data, particularly in summer. Conversely, Actinobacteria acIII was either absent or present in very low abundance in almost all samples. This might be due to acIII being more successful when growing in culture than in the environment or differences between Lake Grosse Fuchskuhle and the lakes analyzed in this study. Free-living freshwater bacteria have recently been proposed to establish promiscuous rather than specific interactions, fulfilling their metabolic needs by associating with whatever suitable partner they happen to encounter [58]. The ecological role of acIII might thus be fulfilled by a different microorganism in different lakes.
The PnecC, acI-B2, and bacI clades were represented by more than one ASV in the dataset, and the relative abundance of each ASV was largely controlled by temperature and oxygen. The most abundant ASVs for each clade and each of the dimictic lakes are shown in Fig. 6b, together with their temperature and oxygen preferences. Both parameters significantly explained the abundance ratios of the different ASVs from the same clade in the three dimictic lakes analyzed in this study (Fig. 6c, FDR < 0.05), as well as in most of the other lakes (Figs. S8 and S9). For the most part, ecotypes appeared to vary similarly with temperature and oxygen concentrations, which were significantly correlated in the data. The only exceptions were the two main ecotypes of clade bacI in NSB (dependent on oxygen, but not on temperature) and of clade acI-B2 in SSB (dependent on temperature, but not on oxygen). The existence of niche partitioning among ecotypes of the same species has been reported previously [59,60,61,62,63,64], and our results suggest that this phenomenon might be generalized in freshwater ecosystems.
Discussion
We studied the microbial composition of eight temperate lakes with different characteristics using 140 bp 16S rRNA amplicons and analyzing them at the 97%-OTU and the ASV (single-nucleotide difference, 99.3% similarity) levels. We note that ASVs simply represent the maximum taxonomic resolution attainable in a given study, and that in some cases this might not be enough to distinguish between closely related taxa [65]. For example, Hoetzinger et al. [66] reported on two different Polynucleobacter species sharing the full length of the 16S rRNA gene. Nonetheless, our ASVs showed coherent distributions across space, time, and environmental gradients, which suggests that, even if a finer diversity could be revealed using longer sequences or additional taxonomic markers, our taxonomic resolution was sufficient to yield ecologically meaningful results.
We first focused on evaluating whether OTUs and ASVs offered different perspectives on the compositional dynamics of freshwater lakes. Ordination methods showed a good agreement between the OTU and ASV resolutions in explaining the general compositional trends of the samples (Fig. 1), suggesting that community structure was at least partially determined above the ASV level. This agrees well with previous reports of ASVs and OTUs revealing similar broad scale ecological results [67]. Samples from the same lakes and different years clustered together (Fig. 1), showing that each lake had a characteristic compositional signature that remained relatively stable over time.
In spite of this, there was a high-temporal OTU turnover in the four lakes that were sampled in consecutive years (Fig. 2); although seasonal patterns could be observed, the communities did not return to their exact original composition after a year had passed. Freshwater microbial communities are composed of metabolically streamlined microorganisms, which thrive by promiscuously complementing with a wide array of possible partners [58, 64]. This versatility should result in a compositionally fluid microbial community, which may remain robust without necessarily showing strong seasonal trends. Lake NSB, for example, hosted blooms associated with the fall mixing in both 2007 and 2008, which involved two different methanotrophic OTUs (Fig. 4b). It is likely that their respective blooms responded to the same underlying environmental trigger (the combined availability of hypolimnetic methane and epilimnetic oxygen during the mixing event) and had a similar effect on the ecosystem (a transient depletion of oxygen associated with methane consumption, Fig. S5). In this sense, even if community composition changed from year to year, the associated functional turnover may have been much smaller.
Despite the interannual variability in community composition, some taxa were found in all communities and layers, as previously described in ref. [37]. When we focused on the eight OTUs with the highest relative abundance in the dataset, we found that they were composed of different ASVs with appeared to have specific preferences for particular lakes and layers (Fig. 3 and Fig. S3). This hinted at an important role of microdiversity in the dynamics of the microbial communities of lakes. While it is true that OTUs and ASVs provided a similar overview of community structure (Fig. 1), some ecosystem processes might be determined by the presence and interplay of particular ecotypes, which can be only distinguished by delineating ASVs [18,19,20,21].
Several studies have described the existence of ecotypes within microbial species in diverse environments, including host-associated microbiomes [19,20,21, 68], granular biofilms [69], marshes [21], the ocean [12, 27, 30, 70,71,72], oil spills [25, 26], and freshwater lakes [65, 73,74,75]. Moreover, it has been claimed that a rich biodiversity guarantees the long-lasting maintenance of bacterial populations [29, 35, 76, 77]. However, the role of microdiversity as a stabilizing force in microbial communities remains insufficiently explored. In this study, microdiversity appeared to be crucial at preserving the higher-order community composition in the face of environmental change. OTUs with more ASVs were more stable than OTUs with one or few ASVs, showing higher persistence and reduced variability (Fig. 5). In a recent work, Needham et al. [71] reported a similar trend in terms of persistence for marine taxa and hypothesized that microdiversity may provide defense against viruses and flexibility in terms of substrate specialization. In our study, temperature and oxygen were better predictors of the distribution of ASVs than of the distribution of their parent OTUs (Fig. S7), suggesting that both environmental factors could be driving the generation of microdiversity. Indeed, when studying the dynamics of taxa from a model community in the samples, we found that three of the most ubiquitous freshwater clades (Polynucleobacter PnecC, Actinobacteria acI-B2 and Bacteroidetes bacI) were each represented by two main ecotypes that predominated under different temperature and oxygen conditions (Fig. 6c and Figs. S8 and S9). In addition to this, other variables not analyzed in this study (such as pH or nutrient availability) might be also influencing microdiversification in freshwater lakes, resulting in an array of closely related strains, each optimized for a particular ecological niche.
Our results point to microdiversity as an important factor determining community stability: fluctuations over time (seasonal patterns) and space (thermal layer) may affect the ratio between different ecotypes, but the OTUs in the community remain detectable throughout the year. This is exemplified in Fig. 4a, in which the relative abundance of Polynucleobacter PnecC was fairly constant during years 2007 and 2008 in both thermal layers, except for blooms of other taxa during the autumn mixing events. This stability was possible due to the existence of two well-differentiated ecotypes of PnecC (ASV-1 and ASV-2), which replaced each other as environmental conditions changed. In particular, ASV-1, which for the most part was observed in the hypolimnion (Figs. 3 and 4a), came to dominate in the epilimnion during early spring, where temperatures were lower and a thermal inversion was still in place. In general, it was difficult to separate the effects of temperature and oxygen, as most of the samples were taken during summer, where water is hotter and more oxygenated in the surface than in deeper locations. However, in early spring, temperature and oxygen concentrations followed opposite trends (Fig. S5). Thus, the dynamics of both ecotypes of Polynucleobacter PnecC during this period suggests that their abundance might be controlled by temperature, rather than by oxygen concentrations. This agrees with an increasing body of literature highlighting the critical role of temperature in shaping microbial ecosystems [78,79,80], and in particular, controlling the distribution of Polynucleobacter ecotypes [73, 81]. Other authors, however, have pointed to oxygen as a main factor driving Polynucleobacter diversification [61]. These options are not mutually exclusive and would both result in an increased persistence of Polynucleobacter under variable conditions.
ASV resolution also provided new insights into the behavior of OTUs experiencing episodic blooms. Gammaproteobacteria gamI, assigned to the methanotrophic genus Methylobacter, bloomed during the fall mixing of 2007, remained abundant until the spring mixing, and dropped to a very low abundance during summer stratification (Fig. 4b). This methanotroph was actually represented by two different ASVs in NSB: ASV-3 and ASV-31. ASV-31 was only abundant at the beginning of the autumn mixing event, where the methane accumulated in the hypolimnion during summer suddenly gets mixed with oxygen throughout the water column. At that point, we observed a sharp decrease in oxygen concentrations (Fig. S5), which we attribute to the rapid oxidation of the methane burst by ASV-31. ASV-3, on the other hand, dominated afterwards, when oxygen concentrations returned to higher levels (Fig. 4b and Fig. S5, ref. [3]).
We hypothesize that the two ASVs represent two ecotypes of Methylobacter, with ASV-31 being adapted to compete for oxygen when there is an excess of methane, and ASV-3 being suited to low-methane/high-oxygen scenarios. This would be consistent with the known existence of Methylobacter groups with different oxygen preferences, some of which are able to live at undetectable concentrations of oxygen [82]. Interestingly, in the fall mixing of 2008, a different methanotrophic OTU (Methyloparacoccus, ASV-50) bloomed at about the same time (Fig. 4b). These data point to a double layer of functional redundancy: there were two OTUs carrying out methanotrophy and blooming after fall mixing in different years, and at least two ASVs with different temporal dynamics within Gammaproteobacteria gamI.
For simplicity, we have assumed that ecotypes from the same OTUs are functionally equivalent and that their relative abundances in the samples were only controlled by environmental factors. The observed trends in community composition agreed with such a basic model (Fig. 6c), and while this study presents a taxonomic, rather than functional, characterization of the samples, we find it reasonable to assume that phylogenetically close organisms will have similar functional characteristics [80, 83]. If this was indeed true for the taxa detected in this study, our observations would fit well with a scenario of functional redundancy [35, 84], with microdiversity guaranteeing the persistence of higher taxonomic ranks—and therefore functional stability—across environmental gradients.
Freshwater lakes are complex ecosystems that host highly diverse microbial communities and have a major role in the global biogeochemical cycles [85,86,87]. They are also good candidates for studying bacterial community dynamics [3, 37, 88] because they experience seasonal variations in environmental conditions, are subjected to periodic perturbations (mixing), and contain strong environmental gradients (stratification). Lakes have been traditionally believed to show highly seasonal patterns with regards to community composition [73, 89,90,91], although some multi-year studies have partially challenged this view, in particular during the summer season [37]. Lacustrine microbial communities also show variability at short timescales, being sensitive to high-frequency/high-locality perturbations, such as hourly changes in wind speed [92]. Under such unpredictable conditions, adaptability becomes of capital importance for lake-dwelling microorganisms. As recently reported, the ability to establish promiscuous metabolic complementations might be a way to maximize survival within an ever-changing biotic landscape [58]. Here, we show that microdiversification is another factor leading to the persistence of ubiquitous freshwater taxa, in this case across abiotic gradients. Microdiversity generation might thus be a key process contributing to microbial community stability. While we have mainly focused on dichotomic relationships between ecotypes of the same OTU, much more complex dynamics might be unraveled in the future. Ongoing long-term time series experiments characterizing not only the taxonomic but also the functional composition of the North Temperate Lakes Microbial Observatory will surely help to elucidate the links between abiotic and biotic variability, inter and intra-specific diversity, and ecosystem functioning, shedding further light on how microbial communities adapt to environmental change.
References
Sutherland WJ, Freckleton RP, Godfray HCJ, Beissinger SR, Benton T, Cameron DD, et al. Identification of 100 fundamental ecological questions. J Ecol. 2013;101:58–67.
Graham EB, Knelman JE, Schindlbacher A, Siciliano S, Breulmann M, Yannarell A, et al. Microbes as engines of ecosystem function: when does community structure enhance predictions of ecosystem processes? Front Microbiol. 2016;7:214.
Shade A, Read JS, Youngblut ND, Fierer N, Knight R, Kratz TK, et al. Lake microbial communities are resilient after a whole-ecosystem disturbance. ISME J. 2012;6:2153–67.
Erkus O, De Jager VCL, Spus M, Van Alen-Boerrigter IJ, Van Rijswijck IMH, Hazelwood L, et al. Multifactorial diversity sustains microbial community stability. ISME J. 2013;7:2126–36.
On SLW, Miller WG, Houf K, Fox JG, Vandamme P. Minimal standards for describing new species belonging to the families Campylobacteraceae and Helicobacteraceae: Campylobacter, Arcobacter, Helicobacter and Wolinella spp. Int J Syst Evol Microbiol. 2017;67:5296–311.
Doolittle WF, Zhaxybayeva O. On the origin of prokaryotic species. Genome Res. 2009;19:744–56.
Rosselló-Mora R, Amann R. Past and future species definitions for bacteria and Archaea. Syst Appl Microbiol. 2015;38:209–16.
Koeppel A, Perry EB, Sikorski J, Krizanc D, Warner A, Ward DM, et al. Identifying the fundamental units of bacterial diversity: a paradigm shift to incorporate ecology into bacterial systematics. Proc Natl Acad Sci USA 2008;105:2504–9.
Shapiro BJ, Polz MF. Ordering microbial diversity into ecologically and genetically cohesive units. Trends Microbiol. 2014;22:235–47.
Tikhonov M. Theoretical microbial ecology without species. Phys Rev E. 2017;96:032410.
Rasko DA, Rosovitz MJ, Myers GSA, Mongodin EF, Fricke WF, Gajer P, et al. The pangenome structure of Escherichia coli: comparative genomic analysis of E. coli commensal and pathogenic isolates. J Bacteriol. 2008;190:6881–93.
Kashtan N, Roggensack SE, Rodrigue S, Thompson JW, Biller SJ, Coe A, et al. Single-cell genomics reveals hundreds of coexisting subpopulations in wild Prochlorococcus. Science. 2014;344:416–20.
Tettelin H, Masignani V, Cieslewicz MJ, Donati C, Medini D, Ward NL, et al. Genome analysis of multiple pathogenic isolates of Streptococcus agalactiae: implications for the microbial “pan-genome”. Proc Natl Acad Sci USA. 2005;102:13950–55.
Cohan FM. Bacterial species and speciation. Syst Biol. 2001;50:513–24.
Coleman ML, Chisholm SW. Code and context: Prochlorococcus as a model for cross-scale biology. Trends Microbiol. 2007;15:398–407.
Koeppel AF, Wertheim JO, Barone L, Gentile N, Krizanc D, Cohan FM. Speedy speciation in a bacterial microcosm: new species can arise as frequently as adaptations within a species. ISME J. 2013;7:1080–91.
Yarza P, Yilmaz P, Pruesse E, Glöckner FO, Ludwig W, Schleifer KH, et al. Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nat Rev Microbiol. 2014;12:635–45.
Koeppel AF, Wu M. Surprisingly extensive mixed phylogenetic and ecological signals among bacterial operational taxonomic units. Nucleic Acids Res. 2013;41:5175–88.
Eren AM, Maignien L, Sul WJ, Murphy LG, Grim SL, Morrison HG, et al. Oligotyping: differentiating between closely related microbial taxa using 16S rRNA gene data. Methods Ecol Evol. 2013;4:1111–9.
Tikhonov M, Leach RW, Wingreen NS. Interpreting 16S metagenomic data without clustering to achieve sub-OTU resolution. ISME J. 2014;9:68–80.
Eren AM, Morrison HG, Lescault PJ, Reveillaud J, Vineis JH, Sogin ML. Minimum entropy decomposition: unsupervised oligotyping for sensitive partitioning of high- throughput marker gene sequences. ISME J. 2015;9:968–79.
Callahan BJ, Mcmurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP. DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods. 2016;13:581–3.
Edgar RC. UNOISE2: improved error-correction for Illumina 16S and ITS amplicon sequencing. bioRxiv 2016;81257.
Amir A, McDonald D, Navas-Molina JA, Kopylova E, Morton JT, Zech Xu Z, et al. Deblur rapidly resolves single-nucleotide community sequence patterns. mSystems. 2017;2:e00191–16.
Kleindienst S, Grim S, Sogin M, Bracco A, Crespo-Medina M, Joye SB. Diverse, rare microbial taxa responded to the Deepwater Horizon deep-sea hydrocarbon plume. ISME J. 2016;10:400–15.
Terrisse F, Cravo-Laureau C, Noël C, Cagnon C, Dumbrell AJ, McGenity TJ, et al. Variation of oxygenation conditions on a hydrocarbonoclastic microbial community reveals Alcanivorax and Cycloclasticus Ecotypes. Front Microbiol. 2017;8:1549.
Chafee M, Fernàndez-Guerra A, Buttigieg PL, Gerdts G, Eren AM, Teeling H, et al. Recurrent patterns of microdiversity in a temperate coastal marine environment. ISME J. 2018;12:237–52.
Konopka A, Lindemann S, Fredrickson J. Dynamics in microbial communities: unraveling mechanisms to identify principles. ISME J. 2015;9:1488–95.
Schloter M, Lebuhn M, Heulin T, Hartmann A. Ecology and evolution of bacterial microdiversity. FEMS Microbiol Rev. 2000;24:647–60.
Kashtan N, Roggensack SE, Berta-Thompson JW, Grinberg M, Stepanauskas R, Chisholm SW. Fundamental differences in diversity and genomic population structure between Atlantic and Pacific Prochlorococcus. ISME J. 2017;11:1997–2011.
Pimm SL. The complexity and stability of ecosystems. Nature. 1984;307:321–26.
McCann KS. The diversity-stability. Nature. 2000;405:228–33.
Wohl DL, Arora S, Gladstone JR. Functional redundancy supports biodiversity and ecosystem function in a closed and constant environment. Ecology. 2004;85:1534–40.
Loreau M, Naeem S, Inchausti P, Bengtsson J, Grime JP, Hector A, et al. Biodiversity and ecosystem functioning: current knowledge and future challenges. Science. 2001;294:804–8.
Shade A, Peter H, Allison SD, Baho DL, Berga M, Bürgmann H, et al. Fundamentals of microbial community resistance and resilience. Front Life Sci. 2012;3:417.
Shade A, Jones SE, McMahon KD. The influence of habitat heterogeneity on freshwater bacterial community composition and dynamics. Environ Microbiol. 2008;10:1057–67.
Linz AM, Crary BC, Shade A, Owens S, Gilbert JA, Knight R, et al. Bacterial community composition and dynamics spanning five years in freshwater bog lakes. mSphere. 2017;2:e00169–17.
Pace ML, McManus GB, Findlay SEG. Planktonic community structure determines the fate of bacterial production in a temperate lake. Limnol Oceano. 1990;35:795–808.
Zwart G, Crump BC, Kamst-van Agterveld MP, Hagen F, Han SK. Typical freshwater bacteria: an analysis of available 16S rRNA gene sequences from plankton of lakes and rivers. Aquat Micro Ecol. 2002;28:141–155.
Jaspers E, Overmann J. Ecological significance of microdiversity: identical 16S rRNA gene sequences can be found in bacteria with highly divergent genomes and ecophysiologies. Appl Environ Microbiol. 2004;70:4831–39.
Newton RJ, Jones SE, Eiler A, McMahon KD, Bertilsson S. A guide to the natural history of freshwater lake bacteria. Microbiol Mol Biol Rev. 2011;75:14–49.
Pernthaler J. Competition and niche separation of pelagic bacteria in freshwater habitats. Environ Microbiol. 2017;19:2133–50.
Williamson CE, Dodds W, Kratz TK, Palmer MA. Lakes and streams as sentinels of environmental change in terrestrial and atmospheric processes. Front Ecol Environ. 2008;6:247–54.
Shade A, Read JS, Welkie DG, Kratz TK, Wu CH, McMahon KD. Resistance, resilience and recovery: aquatic bacterial dynamics after water column disturbance. Environ Microbiol. 2011;13:2752–67.
Puente-Sánchez F, Aguirre J, Parro V. A novel conceptual approach to read-filtering in high-throughput amplicon sequencing studies. Nucl Acids Res. 2016;44:e40.
Yilmaz P, Parfrey LW, Yarza P, Gerken J, Pruesse E, Quast C, et al. The SILVA and ‘All-species Living Tree Project (LTP)’ taxonomic frameworks. Nucl Acids Res. 2014;42:D643–8.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, et al. Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol. 2009;75:7537–41.
Westcott SL, Schloss PD. OptiClust, an improved method for assigning amplicon-based sequence data to operational taxonomic units. mSphere. 2017;2:e00073–17.
Legendre P, Gallagher ED. Ecologically meaningful transformations for ordination of species data. Oecologia. 2001;129:271–80.
Buttigieg PL, Ramette A. A guide to statistical analysis in microbial ecology: a community-focused, living review of multivariate data analyses. FEMS Microbiol Ecol. 2014;90:543–50.
Hill MO, Gauch HG. Detrended correspondence analysis: an improved ordination technique. Vegetatio. 1980;42:47–58.
Peres-Neto PR, Jackson DA. How well do multivariate data sets match? The advantages of a Procrustean superimposition approach over the Mantel test. Oecologia. 2001;129:169–178.
Oksanen J. Multivariate analysis of ecological communities in R: vegan tutorial. University of Oulu, Oulu, 2007.
Lundin D, Severin I, Logue JB, Östman Ö, Andersson AF, Lindström ES. Which sequencing depth is sufficient to describe patterns in bacterial α- and β-diversity? Environ Microbiol Rep. 2012;4:367–72.
Jost L. Entropy and diversity. Oikos. 2006;113:363–75.
Elith J, Leathwick JR, Hastie T. A working guide to boosted regression trees. J Anim Ecol. 2008;77:802–13.
Garcia SL, Buck M, McMahon KD, Grossart HP, Eiler A, Warnecke F. Auxotrophy and intrapopulation complementary in the ‘interactome’ of a cultivated freshwater model community. Mol Ecol. 2015;24:4449–59.
Garcia SL, Buck M, Hamilton JJ, Wurzbacher C, Grossart HP, McMahon KD, et al. Model communities hint at promiscuous metabolic linkages between ubiquitous free-living freshwater bacteria. mSphere. 2018;3:e00202–18.
Johnson ZI, Zinser ER, Coe A, Mcnulty NP, Malcolm E, Woodward S, et al. Niche partitioning among Prochlorococcus ecotypes along ocean-scale environmental gradients. Science. 2006;311:1737–40.
Newton RJ, Jones SE, Helmus MR, McMahon KD. Phylogenetic ecology of the freshwater Actinobacteria acI lineage. Appl Environ Microbiol. 2007;73:7169–76.
Jezbera J, Jezberová J, Brandt U, Hahn MW. Ubiquity of Polynucleobacter necessarius subspecies asymbioticus results from ecological diversification. Environ Microbiol. 2011;13:922–31.
Biller SJ, Berube PM, Lindell D, Chisholm SW. Prochlorococcus: the structure and function of collective diversity. Nat Rev Microbiol. 2015;13:13–27.
Hall MW, Rohwer RR, Perrie J, McMahon KD, Beiko RG. Ananke: temporal clustering reveals ecological dynamics of microbial communities. Peer J. 2017;5:e3812.
Neuenschwander SM, Ghai R, Pernthaler J, Salcher MM. Microdiversification in genome-streamlined ubiquitous freshwater Actinobacteria. ISME J. 2018;12:185–98.
Hahn MW, Jezberová J, Koll U, Saueressig-Beck T, Schmidt J. Complete ecological isolation and cryptic diversity in Polynucleobacter bacteria not resolved by 16S rRNA gene sequences. ISME J. 2016;10:1642–55.
Hoetzinger M, Schmidt J, Pitt A, Koll U, Lang E, Hahn MW. Polynucleobacter paneuropaeus sp. nov., characterized by six strains isolated from freshwater lakes located along a 3000 km north-south cross-section across Europe. Int J Syst Evol Microbiol. 2019;69:203–13.
Glassman SI, Martiny JBH. Broadscale ecological patterns are robust to use of exact sequence variants versus operational taxonomic units. mSphere. 2018;3:e00148–18.
Ellegaard KM, Engel P. Beyond 16S rRNA community profiling: intra-species diversity in the gut microbiota. Front Microbiol. 2016;7:1475.
Leventhal GE, Boix C, Kuechler U, Enke TN, Sliwerska E, Holliger C, et al. Strain-level diversity drives alternative community types in millimetre-scale granular biofilms. Nat Microbiol. 2018;3:1295–303.
Farrant GK, Doré H, Cornejo-Castillo FM, Partensky F, Ratin M, Ostrowski M, et al. Delineating ecologically significant taxonomic units from global patterns of marine picocyanobacteria. Proc Natl Acad Sci USA 2016;113:E3365–74.
Needham DM, Sachdeva R, Fuhrman JA. Ecological dynamics and co-occurrence among marine phytoplankton, bacteria and myoviruses shows microdiversity matters. ISME J. 2017;11:1614–29.
Kent AG, Baer SE, Mouginot C, Huang JS, Larkin AA, Lomas MW, et al. Parallel phylogeography of Prochlorococcus and Synechococcus. ISME J. 2019;13:430–41.
Wu QL, Hahn MW. High predictability of the seasonal dynamics of a species-like Polynucleobacter population in a freshwater lake. Environ Microbiol. 2006;8:1660–66.
Sangwan N, Zarraonaindia I, Hampton-Marcell JT, Ssegane H, Eshoo TW, Rijal G, et al. Differential functional constraints cause strain-level endemism in Polynucleobacter populations. mSystems. 2016;1:e00003–16.
Hoetzinger M, Schmidt J, Jezberová J, Koll U, Hahn MW. Microdiversification of a pelagic Polynucleobacter species is mainly driven by acquisition of genomic islands from a partially interspecific gene pool. Appl Environ Microbiol. 2017;83:e02266–16.
Doak DF, Bigger D, Harding EK, Marvier MA, O’Malley RE, Thomson D. The statistical inevitability of stability-diversity relationships in community ecology. Am Nat. 1998;151:264–76.
Yachi S, Loreau M. Biodiversity and ecosystem productivity in a fluctuating environment: the insurance hypothesis. Proc Natl Acad Sci USA 1999;96:1463–8.
Tamames J, Abellán JJ, Pignatelli M, Camacho A, Moya A. Environmental distribution of prokaryotic taxa. BMC Microbiol. 2010;10:85.
Sunagawa S, Coelho LP, Chaffron S, Kultima JR, Labadie K, Salazar G, et al. Structure and function of the global ocean microbiome. Science. 2015;348:1261359.
Tamames J, Sánchez PD, Nikel PI, Pedrós-Alió C. Quantifying the relative importance of phylogeny and environmental preferences as drivers of gene content in prokaryotic microorganisms. Front Microbiol. 2016;7:433.
Lindström ES, Agterveld MPK, Zwart G. Distribution of typical freshwater bacterial groups is associated with pH, temperature, and lake water retention time. Appl Environ Microbiol. 2005;71:8201–06.
Biderre-Petit C, Jézéquel D, Dugat-Bony E, Lopes F, Kuever J, Borrel G, et al. Identification of microbial communities involved in the methane cycle of a freshwater meromictic lake. FEMS Microbiol Ecol. 2011;77:533–45.
Cobo-Simón M, Tamames J. Relating genomic characteristics to environmental preferences and ubiquity in different microbial taxa. BMC Genom. 2017;18:499.
Allison SD, Martiny JBH. Resistance, resilience, and redundancy in microbial communities. Proc Natl Acad Sci USA 2008;105:11512–19.
Bastviken D, Cole J, Pace M, Tranvik L. Methane emissions from lakes: dependence of lake characteristics, two regional assessments, and a global estimate. Glob Biogeochem Cycles. 2004;18:GB4009.
Harrison JA, Maranger RJ, Alexander RB, Giblin AE, Pierre-Andre J, Mayorga E, et al. The regional and global significance of nitrogen removal in lakes and reservoirs. Biogeochemistry. 2009;93:143–57.
Frings PJ, Clymans W, Jeppesen E, Lauridsen TL, Struyf E, Conley DJ. Lack of steady-state in the global biogeochemical Si cycle: emerging evidence from lake Si sequestration. Biogeochemistry. 2014;117:255–77.
Shade A, Chiu CY, McMahon KD. Seasonal and episodic lake mixing stimulate differential planktonic bacterial dynamics. Micro Ecol. 2010;59:546–54.
Graham JM, Kent AD, Lauster GH, Yannarell AC, Graham LE, Triplett EW. Seasonal dynamics of phytoplankton and planktonic protozoan communities in a northern temperate humic lake: diversity in a dinoflagellate dominated system. Micro Ecol. 2004;48:528–40.
Allgaier M, Grossart HP. Diversity and seasonal dynamics of Actinobacteria populations in four lakes in northeastern Germany. Appl Environ Microbiol. 2006;72:3489–97.
Eiler A, Heinrich F, Bertilsson S. Coherent dynamics and association networks among lake bacterioplankton taxa. ISME J. 2012;6:330–42.
Rusak JA, Tanentzap AJ, Klug JL, Rose KC, Hendricks SP, Jennings E, et al. Wind and trophic status explain within and among-lake variability of algal biomass. Limnol Oceano Lett. 2018;3:409–18.
Acknowledgements
We thank the North Temperate Lakes Microbial Observatory and Katherine McMahon’s research group for the collection of the original data. We also thank Guillermo Martín Serrano for his contribution during early stages of data analysis. This work was funded by grant CTM2016-80095-C2-1-R from the Spanish Ministerio de Economía, Industria y Competitividad. NG-G was funded by a student scholarship from the Severo Ochoa Program at CNB (SEV-2013-0347-17-2).
Author information
Authors and Affiliations
Contributions
FP-S, JT, and CP-A designed research. NG-G and FP-S performed research and analyzed the data. NG-G, FP-S, and CP-A wrote the manuscript. All authors discussed the results and revised the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
García-García, N., Tamames, J., Linz, A.M. et al. Microdiversity ensures the maintenance of functional microbial communities under changing environmental conditions. ISME J 13, 2969–2983 (2019). https://doi.org/10.1038/s41396-019-0487-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-019-0487-8
This article is cited by
-
Decoding populations in the ocean microbiome
Microbiome (2024)
-
Biodiversity patterns of cyanobacterial oligotypes in lakes and rivers: results of a large-scale metabarcoding survey in the Alpine region
Hydrobiologia (2024)
-
Lower viral evolutionary pressure under stable versus fluctuating conditions in subzero Arctic brines
Microbiome (2023)
-
Assembly processes and functional diversity of marine protists and their rare biosphere
Environmental Microbiome (2023)
-
Role in aromatic metabolites biodegradation and adverse implication of denitrifying microbiota in kitchen waste composting
Environmental Microbiome (2023)