Abstract
The susceptibility of soil organic carbon (SOC) in tundra to microbial decomposition under warmer climate scenarios potentially threatens a massive positive feedback to climate change, but the underlying mechanisms of stable SOC decomposition remain elusive. Herein, Alaskan tundra soils from three depths (a fibric O horizon with litter and course roots, an O horizon with decomposing litter and roots, and a mineral-organic mix, laying just above the permafrost) were incubated. Resulting respiration data were assimilated into a 3-pool model to derive decomposition kinetic parameters for fast, slow, and passive SOC pools. Bacterial, archaeal, and fungal taxa and microbial functional genes were profiled throughout the 3-year incubation. Correlation analyses and a Random Forest approach revealed associations between model parameters and microbial community profiles, taxa, and traits. There were more associations between the microbial community data and the SOC decomposition parameters of slow and passive SOC pools than those of the fast SOC pool. Also, microbial community profiles were better predictors of model parameters in deeper soils, which had higher mineral contents and relatively greater quantities of old SOC than in surface soils. Overall, our analyses revealed the functional potential of microbial communities to decompose tundra SOC through a suite of specialized genes and taxa. These results portray divergent strategies by which microbial communities access SOC pools across varying depths, lending mechanistic insights into the vulnerability of what is considered stable SOC in tundra regions.
Similar content being viewed by others
Introduction
In response to climate change, soil carbon (C) at high latitudes is considered to be the single largest component of the terrestrial C pool susceptible to substantial loss over the century time-scale [1,2,3]. The potential release of previously frozen soil C in Arctic regions through microbial decomposition sparks widespread concern over positive feedbacks to climate change [4]. These concerns have been corroborated by recent findings. Rising summer temperatures correlated with high respiration rates during early winter periods, which tip tundra ecosystems into net atmospheric carbon dioxide (CO2) sources [5, 6]. This can be associated with soil microbial communities rapidly responding to warmer soil temperatures and increasing thaw depths [7]. Thaw can result in prolonged microbial exposure to unfrozen SOC and in some areas ice melt increases waterlogged and anaerobic conditions contributing to substantial methane release in addition to CO2 [8]. However, laboratory incubations indicated substantially greater (averaging 3.4 times more) SOC loss under aerobic than anaerobic conditions, suggesting that aerobic SOC decomposition plays a crucial role in permafrost thaw feedbacks [9, 10]. In tundra soils, positive feedbacks to climate warming through enhanced mid- to long-term temperature sensitivity of respiration were observed with the strongest enhancing responses in soils with high C to nitrogen (N) ratios [11, 12]. Altogether, previous results indicated that tundra SOC from varying depths, which represents a massive terrestrial SOC pool, is vulnerable to enhanced decomposition under warmed conditions.
Many studies that assessed the influences of climate variables on tundra SOC loss focused on soil temperature and moisture and utilized only respiration and total soil microbial biomass data, which do not reflect underlying microbial community compositions and functions associated with SOC decomposition [13, 14]. The quality and quantity of SOC have been shown to be important drivers shaping microbial community composition, abundances of bacteria, archaea, and fungi [15, 16], and microbial C use efficiency (CUE) [17]. Additionally, in response to thaw, bacterial/archaeal and fungal community abundances and compositions exhibited significant shifts over depth profiles [18,19,20] and across landscapes [21]. Mineral-organic associations, which increase with depth, can protect SOC from decomposition [22] and the proportion of the passive SOC pool has also been shown to increase with depth and to be higher in mineral soils [12, 23]. Furthermore, colder temperatures and waterlogging potentially slow microbial SOC decomposition with depth and could explain higher percentages of old carbon in deeper tundra soils [24]. Hence, to better understand the temperature sensitivity of tundra SOC it is important to assess microbial SOC decomposition from soils of varying organic matter content and quality and depths and simultaneously assay changes in population dynamics and functional potentials of the soil microbiome.
We previously reported tundra SOC decomposition kinetics under aerobic conditions using 280-day laboratory incubations of soils obtained from the Alaskan tundra treated with experimental field warming or ambient conditions [23]. Bracho et al. [23] demonstrated the sensitivity of the slow SOC pool to microbial decomposition, which accounted for most of the respiration throughout the experiment. We continued the incubation for a total of 3 years without fresh C inputs and employed a three-pool C model to estimate parameters related to decomposition and respiration of fast, slow, and passive SOC pools. For each pool size, the following model parameters were estimated; cumulative CO2 respiration (CR), percentages of CR attributed to each pool, CO2 respiration rates (R), percentages of R from each pool, and decomposition rate constants. The temperatures and time points in the incubation were selected for SOC modeling purposes and do not reflect field temperatures at this site (mean annual temperature is −1 °C) or time points that signify important microbial community shifts in the field (i.e. seasonal variations or response feedbacks to environmental changes). Hence, most analyses focus on the linkages between the model parameters and community profiles. We identified the functional gene and taxonomic variations between microbial communities throughout the incubation and assessed community changes that correlated to the estimated SOC decomposition parameters. We hypothesized that shifts in community taxa and traits would correlate with SOC decomposition parameters and that these associations would be divergent across the depth gradient, owing to the varying C content and SOC pool sizes over depth. Our results indicated that microbial community profiles could predict model parameters for fast, slow and passive SOC pools and revealed a suite of specialized taxa and traits important for stable SOC decomposition.
Materials and methods
Site description and sample collection
Samples for this study were collected from the Carbon in Permafrost Experimental Heating Research (CiPEHR) project, after exposure to two consecutive winter seasons wherein warmed treatments were derived using snow fences. Details pertaining to field site characteristics and warming experiment design are detailed in SI methods and Fig. S1. Soil cores were collected to a depth of 60 cm from 6 control and 6 warmed soil plots. Soil cores were sectioned based on organic matter compositions (0–15; 15–25 cm; and 35–58 cm). Measurement methods for determining soil moisture content, bulk density, mass-based N and C contents, and pH were previously described in Bracho et al. [23]. Soil collected from the plots had higher pH values with increasing depth (4.6–5.15; Table S1). The surface depth (0–15 cm) was a fibric O horizon with litter and course roots and the mid-depth (15–25 cm) was an O horizon with decomposing litter and roots. The lowest depth range (35–58 cm) was composed of soils with a mineral-organic mix, laying just above the permafrost. Averaging all field plots (warming and control), the initial extracted soil properties showed declines in total C (TC) with depth and the highest total N (TN) within the middle depth (Table S1).
Incubation design
A total of 288 samples were analyzed for this study (6 plots × 2 field treatments × 3 depths × 2 incubation temperatures × 4 time points). Soil cores collected from each plot were sectioned by depth then split into ~10 g subsamples. Subsamples were put in open vials and eight of these vials were placed in a single 1 L incubation jar. Incubation jars were incubated at either 15 or 25 °C. At each time point one subsample was removed from each jar and destructively processed to obtain microbial community DNA. The accumulation of CO2 in each jar headspace was quantified using an infrared gas analyzer (IRGA, Li-820 Licor, Lincoln, Nebraska) at 0.9 L min−1 with constant flow maintained by a mass flow controller (Mass Flow meter GFM, Aalborg Instruments & Control) and data was recorded every 3 sec over 8 min by a datalogger (CR1000, Campbell Scientific, Logan UT). The headspace was purged when CO2 concentrations reached 10,000 ppm and C fluxes (Fc) were calculated as the rate of CO2 increase in the headspace of the jars over time after at least 4 cycles of 8.5 h each, expressed in μg CO2-C gCinitial−1 d−1. Fluxes were measured every 48 h during the first 2 weeks, twice a week up to 45 days of incubation, biweekly up to 180 days, then at least once per month until 3 years. Thorough details on the measurement of soil C fluxes were reported by Bracho et al. [23] and photographs of the incubation set-up are in Fig. S1.
Sampling and DNA extractions
After 2 weeks, 3 months, 9 months, and 3 years of incubation, subsample soils were removed from each incubation jar and stored at −80 °C until DNA extractions were performed for microbial analysis. To obtain total soil DNA, the PowerSoil® DNA isolation kit was used in accordance with the provided protocol (MoBio Laboratories, Inc, Carlsbad, California). In some samples, DNA of high purity (Nanodrop 260/280 and 260/230 absorbance ratios above 1.70) could not be obtained via the kit alone so a freeze-grind method [25] was used to obtain DNA that was subsequently purified with the PowerSoil® kit.
16S and ITS amplicon library preparation and illumina sequencing
Community DNA extracts were analyzed using targeted sequencing of the V3–V4 hypervariable region of the bacterial and archaeal 16S ribosomal RNA (rRNA) genes [26] and internal transcribed spacers (ITS), between 5.8S and 28S rRNA genes [27], for fungi. A total of 288 samples (3 depths, 6 field plots, warming and control field treatments, 15 and 25 °C incubations, 4-time points) were analyzed. Sequencing was performed using a 2-step PCR protocol and Illumina MiSeq high-throughput sequencing platform (Illumina, San Diego, CA, USA) [28]. Details on the PCR and sequencing primers, conditions, reagents, and sequence processing are available in Supporting Information. De-multiplexed Sequencing reads are available for download from NCBI Sequence Read Archive under BioProject PRJNA522791, accession numbers SAMN11233799- 11234070 (16S reads) and SAMN11267340-11267612 (ITS reads).
GeoChip analyses
We assessed the microbial functional gene structure using GeoChip 5.0, which contains over 60,000 probes targeting microbial functional genes relevant to environmental processes [29,30,31,32]. For this work, we focused on probes targeting genes involved in C-degradation only (24,886 probes). To generate these data, high-quality DNA (A260/280 ≥ 1.7, A260/230 ≥ 1.3) was fluorescently labeled and hybridized to GeoChip 5.0 60K microarrays. Scanned images of individual microarrays were denoised and normalized to remove poor-quality spots and transform signal intensities into relative abundances. Detailed methodologies for DNA labeling and hybridization, feature extraction, and normalization are provided in SI methods. A data table of normalized GeoChip signal intensities for all probes is available at http://www.ou.edu/ieg/publications/datasets.
Three-pool carbon modeling
To model and partition the SOC into fast, slow, and passive SOC pools we used a three-pool SOC decomposition model, described previously [33] and detailed in the supporting information. It should be noted, that in these soils all depths have high organic matter content and the 0–15 cm depth is a fibric O horizon, so slow and passive pools indicate recalcitrant chemical composition of organic matter and physical barriers to decomposition as well as minerally protected SOC. Briefly, estimates of the proportions and decomposition rate constants of different SOC fractions were modeled using the following equation:
where R is CO2 respiration rate (g CO2-C g−1 SOC day−1) at time t, C0 is initial SOC content (g SOC g−1 soil), f1, f2, f3, k1, k2, and k3 are the relative pool sizes and decomposition rate constants of the fast, slow, and passive SOC components. From these values additional parameters describing the SOC respiration kinetics were calculated and included; CO2 respiration rate from the decomposition of fast, slow, or passive SOC pools (R1, R2, R3), with units of g CO2-C g−1SOC day−1, the proportion of CO2 respiration rate from the decomposition of fast, slow, or passive SOC pools (fR1, fR2, fR3); cumulative CO2 respiration from the decomposition of fast, slow, or passive SOC pool or total cumulative CO2 respired (CR1, CR2, CR3, CRTOT) at time t, with units of g CO2-C g−1 SOC; and the percentage of cumulative CO2 respired from the decomposition of fast, slow, and passive SOC pool relative to the cumulative CO2 respiration from the decomposition of total SOC (fCR1, fCR2, fCR3) at time t. All calculations were based on the parameters estimated from the entire 3-year incubation determined at each time t, corresponding to the DNA extraction time points (2 weeks, 3 months, 9 months, and 3 years). All calculated model parameters are available at http://www.ou.edu/ieg/publications/datasets and rounded values are presented in Tables S2–S4.
Statistical analyses
Prior analyses showed that field warming had a negligible effect on community composition (16S rRNA gene amplicons) and functional potential (GeoChip) [23]. Therefore, we pooled samples from the two field treatments. GeoChip data and 16S and ITS OTU tables were refined prior to analyses using correlation analyses to discard non-significant probes or OTU’s (SI methods). To assess how the SOC parameters varied based on depth and incubation temperatures, ANOVA’s were applied to data subset by depth and incubation temperatures using R. To determine significance of variations between community profiles across soil depths and incubation temperatures non-metric multidimensional scaling (NMDS) plots and non-parametric multivariate dissimilarity tests based on distance matrices were employed (details in SI methods). Significant correlations between microbial community profiles and SOC decomposition parameters were identified using Mantel tests with Pearson correlations and Multiple Regression on distance Matrices (MRM) analyses based on distance matrices (SI methods). Random Forest analyses were employed to identify whether estimated SOC decomposition parameters could be predicted by the 16S, ITS, or GeoChip profiles and to assign values of predictor importance for each OTU, microbial class, or GeoChip probe for each significant prediction based on %IncMSE (the increase in mean squared error of prediction resulting from that OTU, probe, or class being permuted; SI methods).
Results
Soil SOC pools and decomposition kinetics across depth
Estimated SOC parameters (Tables S2–S4) varied significantly in soils from different depths and incubated at different temperatures (ANOVA, P ≤ 0.01, Table S5 and S6) Additionally, cumulative respiration from the decomposition of slow and passive SOC pools dominated the total soil respiration in all depths, under both incubation temperatures (15 °C data presented in Fig. 1 is representative of trends found at both incubation temperatures). The total cumulative respiration and the cumulative respiration attributed to each SOC pool were highest in the surface soil and decreased with soil depth (Fig. 1). Estimated SOC decomposition parameters exhibited significant differences over the depth profile for cumulative respiration, CO2 respiration rates, relative pools sizes, and decomposition rates for all three pools (ANOVA, P ≤ 0.01, Table S5). The percentage of cumulative CO2 respiration (slow and passive pools) significantly varied across depth at the 25 °C incubation, but not the 15 °C incubation. There were higher portions of SOC in the passive pool in the lower depths, whereas the surface depths contained the highest proportions of fast SOC pools (ANOVA, P ≤ 0.01, Table S5). Model parameters also varied in soils incubated at 15 °C compared to those incubated at 25 °C and were significantly different in the mid and lowest depths, but not at the surface depth (ANOVA, P ≤ 0.01, Table S6). For example, incubation temperature resulted in less variation in the surface soils with respect to fast SOC pool size and decomposition kinetics, whereas incubation temperature exhibited the highest significant variation in passive SOC pool sizes and kinetics in the deepest layer (ANOVA, P ≤ 0.01, Table S6). Specifically, the CO2 respiration (slow and passive pools) were affected by incubation temperature in all depths, but the CO2 respiration rate from the decomposition of the fast SOC pool was not impacted at any depth (ANOVA, P ≤ 0.01, Table S6). Altogether, incubation temperature had a greater effect on slow and passive as opposed to fast SOC decomposition parameters. Model parameters exhibited high variances across depth with a lesser effect of incubation temperature (Table S5 and S6).
Stacked bar plots show estimated cumulative respiration (CR) from each SOC pool and total measured cumulative respiration over the 3-year incubation. Estimated CR from the decomposition of the fast, slow and passive SOC pools were calculated using a 3-pool model. Measured CR corresponds to the CR quantified during the incubation. Data are from samples incubated at 15 °C. The samples incubated at 25 °C follow the same trend (not shown here, but data provided in Table S2)
Community dissimilarity was significant across depth and incubation temperature
The 16S rRNA gene profiles were significantly different between the deepest and surface depths at both temperatures (Table 1). In all depths, a significant effect of the incubation temperature on 16S rRNA gene profiles was observed based on almost all dissimilarity tests (Table 1). ITS profiles significantly varied with depth and incubation temperature as well (Table 1). Ordination (NMDS) illustrated a clear contrast between incubation temperatures and reduced dissimilarity (tighter clustering) in the 3-year 16S rRNA gene profiles compared to the other time points (Fig. 2a). The 16S rRNA gene profiles did not cluster based on depths (NMDS). However, ITS profiles were distinctly clustered based on depth, but showed no trends relating to incubation time or incubation temperature (Fig. 2b). The stress tests for these plots indicated the data should be evaluated with caution. GeoChip-based functional gene profiles significantly varied with respect to incubation temperature for all depths, as well as between the surface and deepest depth, for most dissimilarity tests (Table 1). Ordination of GeoChip data indicated functional gene profiles were more similar for communities incubated at the same temperature, except for the 3-year samples, which were distinctly clustered with respect to incubation temperature, though in the opposing ordination direction (Fig. 2c).
Microbial community profiles correlate with some estimated SOC Decomposition parameters
Random Forest and MRM analyses were run to identify correlations between the variances of the SOC decomposition parameters and community profiles (16S, ITS, and GeoChip). Results indicated a higher number of associations between SOC decomposition parameters and community profiles in the lowest depth (compared to the upper depths), for slow and passive SOC decomposition parameters (as opposed to fast SOC decomposition parameters), and when GeoChip profiles (vs. 16S or ITS profiles) were used (Table 2).
Overall, there were significant correlations between 16S profiles and SOC decomposition parameters (both pool sizes and respiration from those pools), particularly those of slow and passive SOC (Tables 2 and 3). From the lowest depth 16S rRNA gene profiles predicted cumulative respiration from the slow SOC pool (55% variance was explained based on Random Forest, Table 3). In all depths, 16S rRNA gene profiles predicted the cumulative respiration from the passive SOC pool and total cumulative respiration (Random Forest, >30% variance explained, Table 3). At the surface, the proportion of respired SOC attributed to the decomposition of the passive and slow pools exhibited variation that could be predicted with the 16S rRNA gene profiles (Random Forest, >30% variance explained, Table 3). In the lowest depth, the pool sizes of fast, slow and passive SOC were explained by 16S rRNA gene profiles (Random Forest, >30% variance explained). At the mid-depth only the variance of the slow pool was explained (Random Forest, 43% variance explained), and no pool sizes were reasonably predicted in the surface depth (Random Forest, <30% variance explained, Table 3). Compared to Random Forest analyses, MRM analyses revealed fewer significant correlations between 16S rRNA gene profiles and SOC decomposition parameters (Table 3). MRM identified significant correlations that were primarily found at the lowest depths (MRM, P < 0.05, Table 3). Cumulative respiration from the passive SOC pool in the lowest depth was the only model parameter that could be significantly associated with 16S profiles by both MRM and Random Forest (MRM, P = 0.04, Random Forest variance explained = 48%). Without sub-setting the community profiles by depth, there were significant, but weak correlations with the distance matrices of 16S rRNA gene profiles and SOC decomposition parameters (Mantel, P = 0.113, MRM, P < 0.05).
ITS profiles exhibited more significant correlations with SOC model parameters than did the 16S rRNA gene profiles. When samples from all depths were combined, distance matrices of ITS profiles and model parameters exhibited significant associations (P < 0.001 for both Mantel and MRM tests). When subset by depth, a higher number of slow and passive SOC decomposition parameters were significantly associated with ITS profiles than fast SOC decomposition parameters (MRM and Random Forest analyses, Tables 2 and 3). Overall, ITS profiles were more closely associated with cumulative respiration (passive, slow, and total) than with SOC pool sizes or decomposition rates (Tables 2 and 3). The pool size of fast SOC could be better explained by ITS profiles with increasing depth (Random Forest 18, 31, 42% variance explained, respectively, Table 3). In all depths, the cumulative respiration from the passive and slow pools as well as the total cumulative respiration were all explained by the ITS profiles (Random forest, ≥ 30% variance explained, Fig. S2, Table 3). In the surface soils, ITS profiles explained the variance in the proportion of cumulative respiration (passive pool), respiration rates (all pools), and the percentage of respiration from the decomposition of the slow pool (Random forest, ≥30% variance explained, Table 3). In both lower depths, the ITS profiles explained variances of the fast SOC pool sizes and in the lowest depth the respiration rate of the fast SOC was explained (Random forest, ≥30% variance explained, Table 3). Plot variance explained by ITS profiles was as high as the most influential SOC decomposition parameters (Random forest, ≥35% variance explained, Table 3). The cumulative respiration from the passive SOC pool and respiration rate of the slow SOC pool (surface soil) as well as the proportion of cumulative respiration from the fast SOC pool (lowest depth) were the only parameters significantly associated with the ITS profiles using both MRM and Random Forest analyses (Table 3).
Functional gene profiles developed using GeoChip microarrays showed a higher number of significant associations with the model parameters, than did 16S and ITS profiles (Table 2). GeoChip-based functional community dissimilarity correlated with the dissimilarity of the model parameters (Mantel test, P < 0.001; MRM analysis, P < 0.01). Altogether, the passive SOC decomposition parameters were more significantly correlated (83%) with GeoChip-based functional profiles across depths than were the slow (44%) or fast (28%) SOC decomposition parameters (Table 2). In all depths, there were significant associations between functional gene profiles and percentage of respiration, cumulative respiration, and the percentage of cumulative respiration attributed to the decomposition of the fast and passive SOC pools (MRM analyses, P ≤ 0.05, Table 3). Except for the fast SOC pool size in the surface soil, all SOC pool sizes could be explained by the functional profiles from all depths (Random Forest analyses, ≥30% variance explained, Table 3). In the surface and deepest layers the decomposition rate of the slow pool corresponded with functional profiles (Random Forest analyses, ≥30% variance explained, Table 3). The cumulative respiration (slow and passive pools), total cumulative respiration, and the proportion of cumulative respiration from the fast and passive pools also corresponded to functional profiles in all layers (Random Forest analyses, ≥30% variance explained, Table 3). Respiration rates and the proportion of respiration from the slow and passive SOC pools also could be explained by the functional profile for different depths (Random Forest analyses, ≥30% variance explained, Table 3). Only the lowest two depths had SOC decomposition parameters associated with C decomposition gene profiles using both MRM and Random Forest and these were predominantly slow and passive SOC decomposition parameters, not fast SOC decomposition parameters (Random Forest ≥ 30% variance explained; MRM P ≤ 0.05, Table 3).
Community markers that explained variance of SOC decomposition parameters
Several bacterial classes were predictors of model parameters. Only SOC decomposition parameters associated with 16S profiles based on Random Forest analyses (≥30% variance explained) were investigated to further to determine OTUs and classes that contributed the most to this association. As depth increased, the relative abundances of OTUs in the class Planctomycetes explained more variance of cumulative respiration (total, slow and passive pools) (heat map values increasing from 18.6–34.4, 2.9–38.1, and 34.4–39.2, respectively, Fig. 3). Planctomycetes exhibited some of the highest explanatory power consistently across 16S profile-associated model parameters (top 5% of heatmap values, Fig. 3). Class level analyses revealed Proteobacteria, Actinobacteria, and Acidobacteria to be associated with model parameters (Random Forest, heat map values in the upper 10%). Notably, Chlamydiea, Planctomycetcia, and Opitutae classes, which all belong to the PVC (Planctomycetes, Verrucomicrobia, and Chlamydiae) superphylum, were predictors of slow and passive, but not fast SOC decomposition parameters in the upper two layers (Plancotmycetica), at the surface (Opitutae), and across all three depths (Chlamydiea) (Fig. 4a). Actinobacteria exhibited associations with model parameters for all pools in the lowest depth, but not upper depths, a trend unique to this class (Fig. 4a). Deltaproteobacteria and Acidobacteria were identified as predictors of slow and passive SOC decomposition parameters, but not fast SOC decomposition parameters (Fig. 4a).
Heatmap of importance (%IncMSE) of bacterial classes and genera to predicting model parameters. Model parameters presented had ≥30% variance explained by 16S profiles. Model parameters are cumulative CO2 respiration from the slow (CR2) and passive (CR3) SOC pools, and total (CRtot); relative pool sizes of the fast (f1), slow (f2), and passive (f3) SOC pools, percentage of the cumulative CO2 respiration from the decomposition of the passive (fCR3) SOC pool, and percentages of the respiration rate from the decomposition of slow (fR2) and passive (fR3) SOC pools. SOC parameters are split by depth (A = 0–15 cm, B = 15–25 cm, C = 35–58 cm)
Ascomycota, Basidiomycota, and Zygomycota were the primary phyla comprising the fungal community in the incubated tundra soils. Ascomycota had the highest relative abundance and the genus Helotiales exhibited consistent dominance over time and depth (read abundance ≥ 18%, Fig. S4). Important OTUs, revealed by Random Forest were from of each of the 3 dominant fungal phyla, Sodariomycetes, Leotiomycetes, and Eurotiomycetes, and were important predictors of fast, slow, and passive SOC decomposition parameters (Random Forest, Fig. 4b and S3). Dothideomycetes and Mucoromycotina were predictors of slow and passive SOC decomposition parameters in the upper layers (Random Forest, Fig. 4b and S3). In the 15–25 cm soils Tremellomycetes were associated with fast SOC decomposition parameters only (Random Forest, Fig. 4b and S3). In the mid depth Microbotryomycetes were good predictors of all SOC decomposition parameters and Agaricomycetes predicted slow and passive parameters(Random Forest, Fig. 4b and S3). Pezizomycotina were predictors of fast SOC decomposition parameters, only in the lowest depth (Random Forest, Fig. 4b and S3).
GeoChip probes targeting enzymes involved in the decomposition of simple sugars were not good predictors of any of the model parameters (Random Forest). However, probes related to the decomposition of starch, other aromatics, chitin, hemicellulose, and pectin were identified as good predictors for fast, slow, and passive SOC decomposition parameters in all depths (Random Forest, Fig. 4c and S3). Probes targeting enzymes involved in the decomposition of agar, pesticides, and lignin were associated with fast SOC decomposition parameters whereas those involved in the decomposition of heparin, pectin, aromatics, and cellulose were associated with slow and passive SOC decomposition parameters, exclusively (Random Forest, Fig. 4c and S3).
Discussion
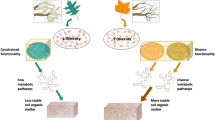
The potential for a significant positive feedback to climate warming exists if C in Arctic soils is decomposed by soil microbial communities, but whether and how microbes access and respire tundra SOC was elusive. This study generated in-depth profiling of tundra microbial communities during SOC turnover that was dominated by the decomposition of slow and passive SOC pools. The estimated SOC decomposition parameters generated for this study provide unique information on identifying microbial community characteristics related to the decomposition kinetics of stable tundra SOC. Here we showed how microbial community profiles (bacterial/archaeal, fungal, and functional genes) associate with these parameters and we identify taxa and traits that were best predictors of fast, slow, and passive SOC decomposition parameters (summary diagrams are in Figs. 5 and 6).
Model parameters for each depth that were predicted by one of the three community profiles, bacterial/archaeal (pink); fungal (yellow); or C decomposition functional genes (blue). These results are based on associations deemed significant using both a Random Forest approach (≥30% of the model parameter variance was explained by the community profile) and multiple regression on distance matrices (MRM) analyses (P < 0.05)
Bacterial/archaeal classes (pink), fungal classes (yellow), and functional gene substrate targets (blue) associated with relatively available SOC (fast model parameters) vs relatively stable SOC (slow and passive model parameters). Predictor importance was assigned to each class or gene probe category for all model parameters with ≥30% variance explained by the corresponding 16S, ITS, or GeoChip profile based on Random Forest analysis. Predictor importance was used to generate heatmaps with fast, slow, and passive groupings for the SOC parameters. A class or probe category was deemed important to prediction of an SOC category if heat map values were ≥4 (16S and ITS) or ≥5 (GeoChip). Only classes and substrates that met this threshold uniquely for either the fast SOC category or the slow/ passive categories are presented here
Overall, 16S-, ITS-, and GeoChip-based profiles were better predictors of cumulative respiration than of respiration rates, pool sizes, and decomposition rates. Likely, the DNA-based profiles were better suited for predicting net outcomes, rather than rates and fluxes. Fungal communities were better predictors of cumulative respiration and respiration rate, which, according to the exponential decay relation of SOC decomposition, are negatively correlated [34]. Hence, it is not surprising that fungal profiles could predict both sets of parameters. The C decomposition functional gene profiles from GeoChip were better predictors of model parameters than the taxonomic profiles, and in particular could predict SOC pool sizes. From these results, we could hypothesize that functional gene data could be more informative in predicting SOC quality or availability than community composition data. However, supplementation of functional gene data with fungal and bacterial/archaeal community composition profiles would be best for estimating SOC loss, as they explain divergent decomposition parameters.
The initial properties of the soil profile from the AK tundra site showed decreases in total SOC with increasing depth, with the highest N concentration and lowest C:N values residing in the 15–25 cm depth. Karhu et al. [11] found that higher soil C:N was related to enhanced microbial CO2 respiration under warming conditions. The AK soils tested herein had high C:N ratios and slow and passive SOC dominated the respiration. Hence, we infer that in these soils, microbial communities were effective in decomposing what would be considered stable SOC. Unsurprisingly, respiration from soils taken from different depths and incubation temperatures varied during the 3-year incubation. The total cumulative respiration was much higher in the surface soils than at the lower depths, irrespective of incubation temperature. With depth there was reduced total cumulative respiration and increasing proportions of slow and passive SOC pools, potentially arising from increased water-filled pore spaces and slow, anaerobic processes dominating SOC turnover with depth [35]. Additionally, increased mineral-organic associations occur with depth and have been shown to be critical stabilizers of SOC in soil but were not directly tested here [22, 36,37,38]. However, at all depths, the passive SOC pool offered a high contribution of cumulative respired CO2. Hence, the microbial mechanisms to access these pools were present along the depth profile in soils with a range of SOC pool sizes.
Overall, the fungal community composition data exhibited more associations with estimated SOC decomposition parameters in the upper depth and to fast SOC in the lowest depths. In contrast, the bacterial/archaeal community composition and functional gene profiles were associated with the estimated SOC decomposition parameters in the deeper layers. This could be related to biomasses of the communities, which were not tested here, but based on qPCR results from Blaud et al. [16], tundra soils exhibited greater bacterial and archaeal abundances when they had higher mineral content, whereas fungal abundances remained consistent.
Interestingly, the fast SOC decomposition parameters exhibited less relation to microbial community taxa and functional genes than the slow and passive SOC decomposition parameters (Figs. 5 and 6). This could reflect the ubiquity of genes and microbial taxa involved in fast SOC decomposition as opposed to specialized genes and taxa with the capacity to access slow and passive SOC. These “specialized” taxa and genes may be less consistently abundant but respond to SOC limitation with the ability to increase their prevalence in the community. Slow and passive SOC respiration was dominant across most of the incubation time. As such, the microbial communities appear to be able to access and respire what could be considered stable SOC. Estimated SOC parameters in the lowest depth frequently had closer associations to community metrics than the upper depths. Again, this may be indicating that as SOC is less easily accessible via mineral associations or other SOC stabilizers, microbial communities are more specialized, thus correlations are more easily detected and can be better used to predict SOC decomposition kinetics.
Many of the bacterial classes with strong contributions to predicting model parameters were not the most abundant organisms, but were classes belonging to the PVC superphylum (Fig. 6). Members of this superphylum are diverse in terms of habitat range, and lifestyles, and are grouped primarily owing to a shared evolutionary history [39]. Verrucomicrobia and Planctomycetes have been found in high-latitude peat bogs, which similar tundra, are characterized by high organic matter and water contents [40, 41]. Using 16S rRNA gene sequencing in forest soils Bai et al. [42] found that SOC temperature sensitivity (Q10) was positively related to copiotrophic guild relative abundances and inversely related to oligotrophic guilds. Here, the Q10 values calculated for the slow SOC pool was larger [23] and slow and passive SOC decomposition parameters were better predicted by Verrucomicrobia, likely K strategists [43] able to grow on low substrate concentrations. This may indicate ecotype variations in SOC decomposition driven by variations in soil edaphic properties and habitat. Interestingly, all Chlamydiae and some Verrucomicrobia are intracellular organisms, found in association with nematodes, ciliates, and amoebae [44,45,46]. The significance of these classes in predicting slow and passive SOC decomposition parameters could relate to essential nutrient cycling driven by soil invertebrates under low-nutrient conditions. Acidobacteria were abundant at all depths, are ubiquitous in soils, have been found broadly in Arctic soils with ranging properties, and have been assayed for their functional roles in SOC decomposition [20, 47]. In this study, Acidobacteria were found to be associated uniquely with slow and passive SOC decomposition parameters in the mid-layer and with fast SOC pool size in the lowest layer. Additionally, Actinobacteria exclusively associated with model parameters for fast, slow, and passive SOC pools in the lowest depth soil, which had higher mineral content and a larger proportion of old C. In previous works metagenome assemblies from arctic soils highlighted SOC catabolic potentials of Actinobacterial taxa related to diverse SOC sources [48, 49]. This could indicate roles of Acidobacteria and Actinobacteria in decomposing stable SOC as well as mineral-associated labile SOC. Altogether, this provides insights into the specialization of many tundra bacterial classes to access and respire stable SOC.
A detailed presentation of the dominant and rare fungal taxa at the CiPHER site was previously published based on soil samples assayed directly after field collection [50]. Interestingly, the dominant genus, Helotiales, found at this site maintained its dominance throughout the incubation. Ascomycota, Basidiomycota, and Zygomycota were the primary phyla comprising the fungal community in the tundra soils, with Ascomycota showing strong dominance. This falls in line with previous findings wherein Ascomycota was the dominant phylum in tundra communities throughout three seasons [51].
The rare and abundant taxa of fungi exhibited essential roles in SOC and litter decomposition [52]. Here, fungal classes from each of the three dominant phyla contained OTUs that explained the variances of model parameters across all depths. Dothideomycetes and Mucoromycotina, which contain many saprobes, were predictors of slow and passive SOC decomposition parameters in the upper layers. This suggests the influence of detritus, which was the primary component within the upper layers, on the decomposition kinetics of the fungal community. Mucoromycotina also contain many ectomycorrhizal taxa and tend to increase in diversity towards the poles [53, 54]. Members of the Mucoromycotina class have high extracellular enzyme production and have been well-studied with regards to lipases, a class of enzymes with efficient catalytic properties in hydrolyzing long chain C molecules [55]. Species richness within this class has been shown to be explained by soil C:N ratio [54], offering additional support that this class of fungi may be associated with SOC chemical composition and recalcitrance. In the 15–25 cm soils, TC remained high and TN was higher than in the other two depths. At this depth, Tremellomycetes were associated with fast SOC decomposition parameters. Members of this class have been previously detected in shallow tundra soils [56,57,58]. Microbotryomycetes were identified as good predictors of slow and passive SOC decomposition parameters and have been associated with mineral layers of tundra soil and increased in relative abundance in response to warming [59]. Hence, at the mid-depth, the fast SOC pool could be predicated by fungi within a class typically found in surface, organic matter rich soils. In contrast, the slow and passive SOC decomposition parameters were associated with fungi involved in accessing mineral-associated OM, highlighting the different strategies used by fungi within these classes to access different SOC pools. The class Agaricomycetes, was an important predictor of slow and passive SOC decomposition parameters at both lower depths and includes many ectomycorrhizal taxa. Diverse lineages within this class produce ligninolytic class II fungal peroxidases and other plant cell wall-decaying enzymes, indicating potential roles in the decay of wood and detritus [60]. Pezizomycotina were predictors of fast SOC decomposition parameters, but only in the lowest depth. While the species richness of this class was previously explained by a positive response to soil pH (increasing with neutral pH) [54], there are less reports on extracellular enzymatic activity by this class than those reported for the aforementioned fungal classes. The pH increased from 4.6 to 5.15 with depth in these samples, which may indicate that the potential of Pezizomycotina to decompose fast SOC and predict fast SOC decomposition parameters is pH dependent. Altogether, the fungi identified with the Random Forest analyses broadly reflect a diverse range of classes that can putatively decompose different SOC pools.
Our analyses revealed associations with model parameters and GeoChip probes targeting enzymes involved in the decomposition of more complex C substrates as opposed to simple sugars, a finding that was consistent across depths. Similarly, using shotgun metagenomic sequencing Mackelprang et al. [61] found cellulose, hemicellulose, and chitin decomposition genes to be significantly correlated with permafrost thaw and genes involved in simple sugar utilization to shift in response to thaw, but with a lesser fold change. Overall, probes targeting enzymes with extracellular activity and/or specialized capabilities had the greater predictive capacity for fast, slow, and passive SOC decomposition parameters. Because extracellular enzymes are energetically expensive for soil microorganisms they are often associated with slow-growing, oligotrophic life strategists, so we expected to find these associated with slow and passive SOC decomposition parameters [62]. However, agarase enzymes, identified in the upper soils in association with fast SOC decomposition parameters, have been demonstrated to be extracellularly secreted [63]. Interestingly, enzymes involved in heparin and pectin degradation were important predictors of slow and passive SOC decomposition parameters in the surface soils, but not of the fast parameters. Heparin is a highly, negatively charged biomolecule, hence chemically recalcitrant and microorganisms with heparinases have been studied, owing to their potential importance in SOC decomposition [64]. Microbial enzymes acting on pectins and heparins often employ elimination mechanisms (lyases) rather than hydrolytic pathways (hydrolases), more commonly utilized to break C–O, C–H, and C–C bonds [65]. Hence, the associations between the slow and passive SOC decomposition parameters and these enzymes reflect potential unique strategies for SOC decomposition processes in surface tundra soils. However, at lower depths, these enzymes no longer serve as the best predictors of slow and passive SOC decomposition parameters. Here, the deeper soils exhibit a stark contrast in functional gene predictive capacities for the fast and slow/passive SOC decomposition parameters. Lignin-targeting enzymes predicted fast SOC decomposition parameters. In contrast, the oxygenases, hydrolases, and aldolases, which are involved in aromatic compound-degradation, and cellulase and galactosidase enzymes, associated with cellulose degradation, were better predictors of the slow and passive SOC decomposition parameters and hence may become more important with increasing C-limitation.
These novel findings are the first to relate estimated SOC decomposition parameters to microbial community composition, phyla, and functional genes, providing unique insights into the associations of microbial taxa and traits with stable SOC turnover across a depth profile. First, fungal communities, which are well known for their capacity to decompose chemically recalcitrant SOC types, were associated with model parameters in surface layers. At lower depths, associations were more closely related to bacterial/archaeal and functional gene profiles, indicating that bacteria and archaea may play more strategic roles in accessing potentially mineral-associated SOC and have greater mobility over depth, likely through water-filled pore spaces, than do the fungal communities. Second, the PVC superphylum has not been reported as important to tundra SOC decomposition in the past, though classes within this superphylum were consistently indicated here by Random Forest analyses. This suggests that there is a previously overlooked role of this versatile superphylum in tundra SOC decomposition. Third, our analyses revealed that a suite of microbial classes and genes, associated with putative extracellular enzyme production, correlated to model parameters for fast, slow, and passive carbon. In addition, microbial community composition and functional gene structure could be better correlated to slow and passive model parameters than to fast model parameters, indicating an increased specialization of the microbial community to decompose SOC with increasing C-limitation. Altogether, in tundra soils microbial taxa and genes could predict SOC decomposition parameters and exhibited a potential of the community to decompose stable SOC. Provided that slow and passive SOC pools dominated the total soil respiration, we conclude that this functional potential was realized and is indicative of the vulnerability of old and stable tundra SOC to decomposition.
References
Koven CD, Riley WJ, Stern A. Analysis of permafrost thermal dynamics and response to climate change in the CMIP5 Earth System Models. J Clim. 2013;26:1877–900.
Schuur E, Abbott B, Bowden W, Brovkin V, Camill P, Canadell J, et al. Expert assessment of vulnerability of permafrost carbon to climate change. Clim Change. 2013;119:359–74.
Schuur E, McGuire AD, Schädel C, Grosse G, Harden J, Hayes D, et al. Climate change and the permafrost carbon feedback. Nature. 2015;520:171.
Schuur EA, Bockheim J, Canadell JG, Euskirchen E, Field CB, Goryachkin SV, et al. Vulnerability of permafrost carbon to climate change: Implications for the global carbon cycle. AIBS Bull. 2008;58:701–14.
Belshe E, Schuur E, Bolker B. Tundra ecosystems observed to be CO2 sources due to differential amplification of the carbon cycle. Ecol Lett. 2013;16:1307–15.
Commane R, Lindaas J, Benmergui J, Luus KA, Chang RY-W, Daube BC, et al. Carbon dioxide sources from Alaska driven by increasing early winter respiration from Arctic tundra. Proc Natl Acad Sci USA. 2017;114:5361–6.
Xue K, Yuan MM, Shi ZJ, Qin Y, Deng Y, Cheng L, et al. Tundra soil carbon is vulnerable to rapid microbial decomposition under climate warming. Nat Clim Change. 2016;6:595–600.
Zona D, Gioli B, Commane R, Lindaas J, Wofsy SC, Miller CE, et al. Cold season emissions dominate the Arctic tundra methane budget. Proc Natl Acad Sci USA. 2016;113:40–5.
Elberling B, Michelsen A, Schädel C, Schuur EA, Christiansen HH, Berg L, et al. Long-term CO2 production following permafrost thaw. Nat Clim Change. 2013;3:890.
Schädel C, Bader MK-F, Schuur EA, Biasi C, Bracho R, Čapek P, et al. Potential carbon emissions dominated by carbon dioxide from thawed permafrost soils. Nat Clim Change. 2016;6:950–3.
Karhu K, Auffret MD, Dungait JA, Hopkins DW, Prosser JI, Singh BK, et al. Temperature sensitivity of soil respiration rates enhanced by microbial community response. Nature. 2014;513:81.
Schädel C, Schuur EA, Bracho R, Elberling B, Knoblauch C, Lee H, et al. Circumpolar assessment of permafrost C quality and its vulnerability over time using long‐term incubation data. Glob Change Biol. 2014;20:641–52.
Nikrad MP, Kerkhof LJ, Häggblom MM. The subzero microbiome: microbial activity in frozen and thawing soils. FEMS Microbiol Ecol. 2016;92:fiw081.
Priemé A, Blok D, Skouw Haugwitz M, Vořίšková J, Elberling B. Arctic soil microbial sensitivity to seasonal dynamics and climate change. Microbial Biomass: A Paradigm Shift in Terrestrial Biogeochemistry. World Scientific. 2017;275–307.
Müller O, Bang‐Andreasen T, White RA III, Elberling B, Taş N, Kneafsey T, et al. Disentangling the complexity of permafrost soil by using high resolution profiling of microbial community composition, key functions and respiration rates. Environ Microbiol. 2018;20:4328–42.
Blaud A, Phoenix GK, Osborn AM. Variation in bacterial, archaeal and fungal community structure and abundance in High Arctic tundra soil. Polar Biol. 2015;38:1009–24.
Allison SD, Wallenstein MD, Bradford MA. Soil-carbon response to warming dependent on microbial physiology. Nat Geosci. 2010;3:336.
Coolen MJ, van de Giessen J, Zhu EY, Wuchter C. Bioavailability of soil organic matter and microbial community dynamics upon permafrost thaw. Environ Microbiol. 2011;13:2299–314.
Frank-Fahle BA, Yergeau É, Greer CW, Lantuit H, Wagner D. Microbial functional potential and community composition in permafrost-affected soils of the NW Canadian Arctic. PLoS ONE. 2014;9:e84761.
Taş N, Prestat E, Wang S, Wu Y, Ulrich C, Kneafsey T, et al. Landscape topography structures the soil microbiome in arctic polygonal tundra. Nat Commun. 2018;9:777.
Bottos EM, Kennedy DW, Romero EB, Fansler SJ, Brown JM, Bramer LM, et al. Dispersal limitation and thermodynamic constraints govern spatial structure of permafrost microbial communities. FEMS Microbiol Ecol. 2018;94:8.
Gentsch N, Wild B, Mikutta R, Čapek P, Diáková K, Schrumpf M, et al. Temperature response of permafrost soil carbon is attenuated by mineral protection. Glob change Biol. 2018;24:3401–15.
Bracho R, Natali S, Pegoraro E, Crummer KG, Schädel C, Celis G, et al. Temperature sensitivity of organic matter decomposition of permafrost-region soils during laboratory incubations. Soil Biol Biochem. 2016;97:1–14.
Pries CEH, Schuur EA, Crummer KG. Holocene carbon stocks and carbon accumulation rates altered in soils undergoing permafrost thaw. Ecosystems. 2012;15:162–73.
Zhou J, Bruns MA, Tiedje JM. DNA recovery from soils of diverse composition. Appl Environ Microbiol. 1996;62:316–22.
Peiffer JA, Spor A, Koren O, Jin Z, Tringe SG, Dangl JL et al. Diversity and heritability of the maize rhizosphere microbiome under field conditions. Proc Natl Acad Sci USA 2013;110:6548–53.
Zhou J, Deng Y, Shen L, Wen C, Yan Q, Ning D, et al. Temperature mediates continental-scale diversity of microbes in forest soils. Nat Commun. 2016;7:12083.
Wu L, Wen C, Qin Y, Yin H, Tu Q, Van Nostrand JD, et al. Phasing amplicon sequencing on Illumina Miseq for robust environmental microbial community analysis. BMC Microbiol 2015;15:125.
Tu Q, Yu H, He Z, Deng Y, Wu L, Van Nostrand JD, et al. GeoChip 4: a functional gene-array-based high-throughput environmental technology for microbial community analysis. Mol Ecol Resour. 2014;14:914–28.
Wu L, Liu X, Schadt CW, Zhou J. Microarray-based analysis of subnanogram quantities of microbial community DNAs by using whole-community genome amplification. Appl Environ Microbiol. 2006;72:4931–41.
Zhou J, Xue K, Xie J, Deng Y, Wu L, Cheng X, et al. Microbial mediation of carbon-cycle feedbacks to climate warming. Nat Clim Change. 2012;2:106–10.
Zhou J, Deng Y, Zhang P, Xue K, Liang Y, Van Nostrand JD et al. Stochasticity, succession, and environmental perturbations in a fluidic ecosystem. Proc Natl Acad Sci USA. 2014;111:E836–45.
Feng W, Liang J, Hale LE, Jung CG, Chen J, Zhou J, et al. Enhanced decomposition of stable soil organic carbon and microbial catabolic potentials by long‐term field warming. Glob change Biol. 2017;23:4765–76.
de Graaff. MA, Classen AT, Castro HF, Schadt CW. Labile soil carbon inputs mediate the soil microbial community composition and plant residue decomposition rates. New Phytol. 2010;188:1055–64.
Walz J, Knoblauch C, Böhme L, Pfeiffer EM. Regulation of soil organic matter decomposition in permafrost-affected Siberian tundra soils-Impact of oxygen availability, freezing and thawing, temperature, and labile organic matter. Soil Biol Biochem. 2017;110:34–43.
Gentsch N, Mikutta R, Shibistova O, Wild B, Schnecker J, Richter A, et al. Properties and bioavailability of particulate and mineral‐associated organic matter in A rctic permafrost soils, Lower Kolyma Region, R ussia. Eur J Soil Sci. 2015;66:722–34.
Gillabel J, Cebrian-Lopez B, Six J, Merckx R. Experimental evidence for the attenuating effect of SOM protection on temperature sensitivity of SOM decomposition. Glob Change Biol. 2010;16:2789–98.
Schrumpf M, Kaiser K, Guggenberger G, Persson T, Kögel-Knabner I, Schulze E-D. Storage and stability of organic carbon in soils as related to depth, occlusion within aggregates, and attachment to minerals. Biogeosciences. 2013;10:1675–91.
Wagner M, Horn M. The Planctomycetes, Verrucomicrobia, Chlamydiae and sister phyla comprise a superphylum with biotechnological and medical relevance. Curr Opin Biotechnol. 2006;17:241–9.
Dedysh SN, Pankratov TA, Belova SE, Kulichevskaya IS, Liesack W. Phylogenetic analysis and in situ identification of bacteria community composition in an acidic Sphagnum peat bog. Appl Environ Microbiol. 2006;72:2110–17.
Juottonen H, Galand PE, Tuittila ES, Laine J, Fritze H, Yrjälä K. Methanogen communities and Bacteria along an ecohydrological gradient in a northern raised bog complex. Environ Microbiol. 2005;7:1547–57.
Bai Z, Xie H, Kao-Kniffin J, Chen B, Shao P, Liang C. Shifts in microbial trophic strategy explain different temperature sensitivity of CO2 flux under constant and diurnally varying temperature regimes. FEMS Microbiol Ecol. 2017;93:1–12.
Noll M, Matthies D, Frenzel P, Derakshani M, Liesack W. Succession of bacterial community structure and diversity in a paddy soil oxygen gradient. Environ Microbiol. 2005;7:382–95.
Horn M, Collingro A, Schmitz-Esser S, Beier CL, Purkhold U, Fartmann B, et al. Illuminating the evolutionary history of chlamydiae. Science. 2004;304:728–30.
Petroni G, Spring S, Schleifer K-H, Verni F, Rosati G. Defensive extrusive ectosymbionts of Euplotidium (Ciliophora) that contain microtubule-like structures are bacteria related to Verrucomicrobia. Proc Natl Acad Sci USA. 2000;97:1813–17.
Vandekerckhove TT, Coomans A, Cornelis K, Baert P, Gillis M. Use of the Verrucomicrobia-specific probe EUB338-III and fluorescent in situ hybridization for detection of “Candidatus Xiphinematobacter” cells in nematode hosts. Appl Environ Microbiol. 2002;68:3121–5.
Rawat SR, Männistö MK, Bromberg Y, Häggblom MM. Comparative genomic and physiological analysis provides insights into the role of Acidobacteria in organic carbon utilization in Arctic tundra soils. FEMS Microbiol Ecol. 2012;82:341–55.
Johnston ER, Rodriguez-R LM, Luo C, Yuan MM, Wu L, He Z, Konstantinidis KT. Metagenomics reveals pervasive bacterial populations and reduced community diversity across the Alaska tundra ecosystem. Front Microbiol. 2016;7:579.
Tveit A, Schwacke R, Svenning MM, Urich T. Organic carbon transformations in high-Arctic peat soils: key functions and microorganisms. ISME J. 2013;7:299–311.
Penton CR, StLouis D, Cole JR, Luo Y, Wu L, Schuur EG, et al. Fungal diversity in permafrost and tallgrass prairie soils under experimental warming conditions. Appl Environ Microbiol. 2013;79:7063–72.
Schadt CW, Martin AP, Lipson DA, Schmidt SK. Seasonal dynamics of previously unknown fungal lineages in tundra soils. Science. 2003;301:1359–61.
Baldrian P, Kolařík M, Štursová M, Kopecký J, Valášková V, Větrovský T, Voříšková J. Active and total microbial communities in forest soil are largely different and highly stratified during decomposition. ISME J. 2012;6:248–58.
Benny GL, Humber RA, Voigt K. 8 Zygomycetous Fungi: Phylum Entomophthoromycota and Subphyla Kickxellomycotina, Mortierellomycotina, Mucoromycotina, and Zoopagomycotina. In: McLaughlin DJ, Spatafora JW, (eds). Systematics and Evolution: Part A. Berlin, Heidelberg: Springer Berlin Heidelberg; 2014. p. 209–50.
Tedersoo L, Bahram M, Põlme S, Kõljalg U, Yorou NS, Wijesundera R, et al. Global diversity and geography of soil fungi. science. 2014;346:1256688.
Kotogán A, Németh B, Vágvölgyi C, Papp T, Takó M. Screening for extracellular lipase enzymes with transesterification capacity in Mucoromycotina strains. Food Technol Biotechnol. 2014;52:73.
Higgins KL, Arnold AE, Miadlikowska J, Sarvate SD, Lutzoni F. Phylogenetic relationships, host affinity, and geographic structure of boreal and arctic endophytes from three major plant lineages. Mol phylogenetics Evol. 2007;42:543–55.
Hu W, Zhang Q, Li D, Cheng G, Mu J, Wu Q, et al. Diversity and community structure of fungi through a permafrost core profile from the Qinghai‐Tibet Plateau of China. J basic Microbiol. 2014;54:1331–41.
Zhang T, Yao Y-F. Endophytic fungal communities associated with vascular plants in the high arctic zone are highly diverse and host-plant specific. PLoS ONE. 2015;10:e0130051.
Deslippe JR, Hartmann M, Simard SW, Mohn WW. Long-term warming alters the composition of Arctic soil microbial communities. FEMS Microbiol Ecol. 2012;82:303–15.
Hibbett D, Bauer R, Binder M, Giachini A, Hosaka K, Justo A, et al. 14 Agaricomycetes. Systematics and evolution. Springer. 2014, 373-429.
Mackelprang R, Waldrop MP, DeAngelis KM, David MM, Chavarria KL, Blazewicz SJ, et al. Metagenomic analysis of a permafrost microbial community reveals a rapid response to thaw. Nature. 2011;480:368–71.
Ramirez KS, Craine JM, Fierer N. Consistent effects of nitrogen amendments on soil microbial communities and processes across biomes. Glob Change Biol. 2012;18:1918–27.
Hosoda A, Sakai M, Kanazawa S. Isolation and characterization of agar-degrading Paenibacillus spp. associated with the rhizosphere of spinach. Biosci, Biotechnol, Biochem. 2003;67:1048–55.
Moore EK, Hopmans EC, Rijpstra WIC, Sanchez Andrea I, Villanueva L, Wienk H, et al. Lysine and novel hydroxylysine lipids in soil bacteria: amino acid membrane lipid response to temperature and pH in Pseudopedobacter saltans. Front Microbiol. 2015;6:637.
Hovingh P, Linker A. [123] Heparinase and heparitinase from flavobacteria In: Ginsburg V (ed.). Degradation of Complex Carbohydrates. Methods in enzymology Elsevier, 1972, p. 902–11.
Acknowledgements
This work was funded by Biological Systems Research on the Role of Microbial Communities in Carbon Cycling Program grants DE-SC0004601 and DESC0010715, and the US Department of Energy, Terrestrial Ecosystem Sciences grant DE-SC0006982 and updated with DE-SC0014085.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Hale, L., Feng, W., Yin, H. et al. Tundra microbial community taxa and traits predict decomposition parameters of stable, old soil organic carbon. ISME J 13, 2901–2915 (2019). https://doi.org/10.1038/s41396-019-0485-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-019-0485-x
This article is cited by
-
Global patterns and edaphic-climatic controls of soil carbon decomposition kinetics predicted from incubation experiments
Nature Communications (2023)
-
DNA sequencing, microbial indicators, and the discovery of buried kimberlites
Communications Earth & Environment (2023)
-
Acidobacteria members harbour an abundant and diverse carbohydrate-active enzymes (cazyme) and secreted proteasome repertoire, key factors for potential efficient biomass degradation
Molecular Genetics and Genomics (2023)
-
Biological Interactions and Environmental Influences Shift Microeukaryotes in Permafrost Active Layer Soil Across the Qinghai-Tibet Plateau
Microbial Ecology (2023)
-
Unboxing the black box—one step forward to understand the soil microbiome: A systematic review
Microbial Ecology (2023)