Abstract
Microcystis is a genus of freshwater cyanobacteria, which causes harmful blooms in ecosystems worldwide. Some Microcystis strains produce harmful toxins such as microcystin, impacting drinking water quality. Microcystis colony morphology, rather than genetic similarity, is often used to classify Microcystis into morphospecies. Yet colony morphology is a plastic trait, which can change depending on environmental and laboratory culture conditions, and is thus an inadequate criterion for species delineation. Furthermore, Microcystis populations are thought to disperse globally and constitute a homogeneous gene pool. However, this assertion is based on relatively incomplete characterization of Microcystis genomic diversity. To better understand these issues, we performed a population genomic analysis of 33 newly sequenced genomes mainly from Canada and Brazil. We identified 17 Microcystis clusters of genomic similarity, five of which correspond to monophyletic clades containing at least three newly sequenced genomes. Four out of these five clades match to named morphospecies. Notably, M. aeruginosa is paraphyletic, distributed across 12 genomic clusters, suggesting it is not a coherent species. A few clades of closely related isolates are specific to a unique geographic location, suggesting biogeographic structure over relatively short evolutionary time scales. Higher homologous recombination rates within than between clades further suggest that monophyletic groups might adhere to a Biological Species-like concept, in which barriers to gene flow maintain species distinctness. However, certain genes—including some involved in microcystin and micropeptin biosynthesis—are recombined between monophyletic groups in the same geographic location, suggesting local adaptation.
Similar content being viewed by others
Introduction
Several categories of species concepts can be used as the basis to delimit and organize biological diversity, and two concepts in particular have been recently applied to bacteria and archaea. The Ecological Species Concept (ESC) proposes that speciation is driven by divergent natural selection between distinct ecological niches, while the Biological Species Concept (BSC) emphasizes gene flow (e.g., homologous recombination) as a cohesive force within species [1]. The Stable Ecotype Model (SEM) is a version of the ESC tailored to bacteria, under the general assumption that adaptive mutations spread more rapidly by clonal expansion than by recombination [2]. However, certain bacteria and archaea have relatively high rates of recombination, such that a BSC-like concept could apply—but strictly the BSC will never apply to bacteria that reproduce clonally and occasionally exchange genes across species boundaries. Several archaea and bacteria appear to fit a BSC-like model, showing higher recombination within species than between species [1, 3,4,5]. Barriers to recombination could be maintained by natural selection or genetic incompatibilities, or due to physical separation (i.e., allopatry). Allopatric speciation is thought to be rare for globally dispersed bacteria, but does appear to occur among geographically separated hotspring archaea [6]. Therefore, a plurality or spectrum of species concepts is probably necessary to fit the diverse lifestyles and recombination frequencies observed across microbes [1].
When two different species inhabit a common niche or geographic location, they may exchange genes beneficial to local adaptation. For example, Vibrio cholerae ‘core’’ housekeeping genes are freely recombined among V. cholerae from both Bangladesh and the American coast, to the exclusion of the sister species V. metecus [7]. However, in the integron (part of the genome subject to particularly frequent recombination), V. cholerae undergoes more genetic exchange with V. metecus from the same geographic location (USA) than with V. cholerae from a different location (Bangladesh). This suggests that species cohesion is maintained across most of the core genome, while certain ‘accessory’’ genes are exchanged across species boundaries to promote local adaptation. Identifying such locally adapted genes that cross-species boundaries can provide insight into the genetic basis of adaptation to different environments [8, 9].
Here, we consider the common bloom-forming cyanobacterium Microcystis as a model of speciation and local adaptation. Microcystis is a genus containing a great deal of genetic diversity [10] and capable of frequent recombination [11,12,13]. Previous genetic and genomic studies have suggested that Microcystis is globally distributed, with little geographic structure [10, 13,14,15]. Thus, it is plausible that Microcystis represents a single, globally distributed and homogeneous gene pool, adhering to a BSC-like model [1, 5, 13, 16]. However, multiple attempts have been made to classify Microcystis into several species, based on various morphological and genetic criteria [17,18,19].
Microcystis is able to form colonies or cell aggregates covered by exopolysaccharide or mucilage [20]. Several Microcystis strains are known to synthesize intracellular toxins, which are thought to be released to the environment primarily when cells die and lyse [21]. Altogether, toxins and cell decomposition followed by oxygen depletion threaten the health of humans and animals. Microcystis colony morphology, cell size, and the structure of mucilage have been used for decades as taxonomic criteria to classify Microcystis in morphospecies or morphotypes (International Code of Botanical Nomenclature) [18, 22]. However, morphospecies classifications are often inconsistent with genetic, genomic, and phylogenetic analyses [18, 23]. These inconsistencies may occur because Microcystis colonies can change morphology or become unicellular without any genotypic changes [18, 20, 24,25,26].
At least 51 morphospecies have been described within the Microcystis genus (http://www.algaebase.org/browse/taxonomy/?id=7066) (e.g., M. aeruginosa, M. panniformis, M. viridis, M. wesenbergii, M. flos-aquae, M. novacekii, and M. ichthyoblabe) [18, 27]. One of the most studied and frequently reported is M. aeruginosa. Under laboratory conditions, it has been shown that colonies of M. wesenbergii morphospecies could become morphologically similar to colonies of M. aeruginosa, after just a few hours of culture [24]. If a Microcystis strain undergoes colony morphology changes, it can then become over-classified into several morphospecies. Thus, the number of morphospecies may not reflect the number of Microcystis species based on other bacterial systematic approaches [18].
Because of these inconsistencies, several authors have attempted to reclassify Microcystis morphospecies using additional systematic approaches, like 16S rRNA gene sequence identity, DNA–DNA hybridization, phylogenetic analysis of conserved genes, and average nucleotide identity (ANI) [10, 12, 15, 18, 19, 28, 29]. For example, five Microcystis morphospecies (M. aeruginosa, M. ichthyoblabe, M. novacekii, M. viridis, and M. wesenbergii) were proposed to be reclassified as a single-bacterial species (M. aeruginosa) [18]. These five Microcystis strains showed 16S rRNA gene sequence identities higher than the usual cutoff value used to define bacterial species (>97%), DNA–DNA genome sequences hybridization values were also higher than the cutoff (>70%), and colony morphologies are generally similar [18, 24]. Other studies showed that M. aeruginosa morphospecies together with other morphospecies are a single-species complex with ANI values >95%, which is consistent with hybridization values >70%—a standard rule in bacterial species delineation [10, 15, 30]. Despite the high similarity in their core genomes, Microcystis are diverse in their gene content, resulting in large accessory genomes that can harbor genes related to the biosynthesis of harmful toxins or secondary metabolites [10, 15, 31]. Microcystis classification based on accessory genes (e.g., toxins and polysaccharides) has also been proposed [15, 29].
In this study, we present a population genomics analysis using 33 newly sequenced genomes (of which 19 were resequenced after several years in laboratory culture) belonging to six Microcystis morphospecies isolated mainly from Brazil and Canada over a 15-year period. We aimed to investigate the coherence of Microcystis morphospecies using phylogenomic and homologous recombination analyses. We identified five Microcystis monophyletic clades with at least three new sequenced genomes, two of which appear to be restricted to particular geographic regions. Consistent with a BSC-like model, three out of five clades show a preference for within-clade recombination. Meanwhile, M. aeruginosa morphospecies are paraphyletic and geographically unstructured, meaning that M. aeruginosa may in fact include multiples sub-species. In contrast with the general preference for recombination within clades, we also observed occasional HGT between clades. Many of these cross-species HGT events may be involved in local adaptation. Finally, we studied the profiles of genes related to the biosynthesis of secondary metabolites (such as microcystin) to determine if different Microcystis clades have a characteristic profile of biosynthetic genes.
Materials and methods
Genome sequencing, assembly, and binning
Over the past 15 years, we collected 30 Microcystis isolates from Brazil, Canada, and the United States. The Microcystis strains were initially characterized as morphospecies based on their colony morphology, according to Komárek [27, 32, 33]. Six morphospecies were identified: M. aeruginosa, M. flos-aquae, M. panniformis, M. wesenbergi, M. viridis, and M. novacekii. Isolates with unknown or ambiguous colony morphologies were indicated with M. or M. sp. designations. We performed the DNA extraction for these strains between 2006 and 2017 (see Table S1). The 30 Microcystis genomes were sequenced using the Illumina HiSeq 2500 platform with 125 bp paired-end reads. The genomic Illumina libraries (with average fragment size 360 bp) were prepared using the NEB (New England Biolabs®) low input protocol. We also sequenced the DNA of four single Microcystis colonies isolated manually under the microscope. Three of these colonies were new isolates recovered from Lake Champlain (Quebec, Canada) in 2017 without culture, while the fourth came from a culture that was also sequenced (in bulk) in this study, for a total of 33 new sequenced Microcystis genomes. Before DNA extraction and sequencing, each colony was washed 10–15 times with Z8 medium using a micropipette. DNA extraction was performed directly on each colony using the ChargeSwitch® gDNA Mini Bacteria Kit.
Of the 30 Microcystis isolates, 19 had been maintained in culture for several years until 2017. Thus, we extracted DNA and resequenced these 19 cultures in 2011, 2016, and 2017 to check for contamination and mutations in the Microcystis genome over time and differences between culture and colony sequences (Table S1). Altogether, the Microcystis genomes, of which 14 were sequenced once, 18 twice, and 1 three times comprised a total of 53 genome sequences. See Fig. S1 for a detailed workflow describing the genome datasets, and the subsequent bioinformatic analyses.
Sequences from cultures and colonies were assembled with the software IDBA-UD v1.1.3 [34], producing contigs belonging to both Microcystis and associated heterotrophic bacteria, which are naturally associated with Microcystis [35, 36]. The software Anvi’o v3.0 was used to filter, cluster, and bin the contigs belonging to Microcystis [37]. For each sample, reads were mapped back to contigs using the very sensitive mode of bowtie2 v2.3.0 [38]. For each contig, we assigned taxonomic affiliation using Centrifuge v1.0.3 [39]. Combining the coverage estimated from the mapping along with tetranucleotide frequencies, GC content, and taxonomic affiliation, contigs were further classified into bins using the anvi-profile procedure as implemented in Anvi’o [37, 39]. The associated bacterial bins were excluded and will be described in a forthcoming manuscript. The gene prediction and annotation were done for Microcystis genomes using Prodigal v2.6.3 and Prokka v1.12 packages, respectively [40,41,42].
Phylogenomic analysis
A core genome of 152 single-copy genes shared by 79 Microcystis genomes (53 and 26 genomes reported here and previously, respectively) and two outgroups (Anabaena variabilis ATCC29413 and Synechocystis sp. PCC6803) was identified using the software Roary and blastn-all. First, a core genome for the 79 Microcystis genomes (minimum value of 90% amino acid identity) was identified using Roary. The outgroups were initially excluded due to their high divergence from Microcystis. To identify the homologous genes in the outgroups, blast-all was used (Blastp similarities >60%). The common core genes between Roary and blast-all were selected and used to create a core gene alignment. Each homologous gene was aligned separately using muscle [43]. The concatenated and degapped alignment of length 129 835 bp was used for building a phylogenetic tree in RAxML v8.2.4, using the GTRGAMMA model, with 100 bootstraps [44]. Using the same method, another core phylogenetic tree was inferred without the previously published reference genomes. This concatenated core-alignment comprised 222 genes (211 589 bp degapped) (see Fig. S2).
Clustering analysis of Microcystis genomes
A multiple genome alignment of the 79 genomes (53 genomes reported here and the 26 reference genomes from NCBI) was performed using Mugsy 1.2.3 [45] (see Tables S1 and S2), yielding a core-alignment of 625 795 bp. The core-alignment (excluding duplicate/resequenced genomes) was used to perform genetic population structure and cluster analysis with hierBAPS (The hierarchical Bayesian Analysis of Population Structure) [46]. We used as input parameters two clustering levels and an expected number of cluster (k) equals to 10, 20, and 40. HierBAPS delineates the population using nested clustering. In this method, rare genotypes (distantly related to better sampled clades) often cluster together due to long-branch attraction [46]. As a result, HierBAPS clusters could be incongruent with the phylogeny inferred by maximum likelihood, which is less sensitive to long-branch attraction.
ANI between Microcystis genomes
The 79 genomes were compared using a python module (average_nucleotide_identity.py—https://github.com/widdowquinn/pyani) to estimate the ANI by Mummer and by BLAST (ANIm and ANIb values between genome pairs; see Fig. S3) [47]. Bacterial genomes with DNA–DNA hybridization (DDH) of at least 70%, are considered as the same species and usually show values of ANI >95%. Hence, a cutoff >95% for the ANI values between genome pairs is used to identify genomes within the same genomic cluster or species [30, 48, 49]. The pairwise identities were plotted using the R package ggplot (http://ggplot2.org/) and the function heatmap2 [50]. ANIb and ANIm values both greater than or equal to 96% were used as criteria to classify Microcystis genomes into 17 clusters. Five out of those clusters contained at least three new sequenced genomes and were used for pangenome and recombination analyses (Fig. S1). Additionally, four of these five clusters were also identified by HierBAPS analysis.
Pangenome analysis in Microcystis genomic clusters
Pangenomes were estimated for each Microcystis genomic cluster with at least three newly sequenced genomes, excluding duplicates and references (Fig. S1 and Table S3). A global pangenome estimation was generated using all the Microcystis genomes excluding two very short genomes (Ma_AC_P_00000000_S299 and Ma_QC_C_20070823_S18). These two genomes were excluded because of their reduced size compared to the average (10% and 30% for Ma_AC_P_00000000_S299 and Ma_QC_C_20070823_S18, respectively). Ma_QC_C_20070823_S18 had the lowest coverage (28X) and Ma_AC_P_00000000_S299 appeared to be contaminated with another cyanobacterium (Anabaena). Genomes were first annotated with Prokka v1.12 [42], then Roary v3.12.0 was used to generate the pangenomes. Specifically, the genomes in GGF3 format generated with Prokka were used as input to Roary. Roary was executed using a minimum percentage of amino acid identity of 90% for blastp, which was set-up according to the Roary recommendations [51].
We also did a pangenome analysis to find homologous and accessory genes within 53 Microcystis genomes from Brazil, Canada, USA (including the shorter ones), and the 26 reference genomes from the NCBI database. Roary allowed us to identify 370 clusters of homologs (shared in 79 genomes) and 1059 (shared in 76 up to 78 genomes). Roary also identified 23,728 accessory genes or genes shared by <76 genomes.
Single-nucleotide polymorphisms (SNPs) and deletion identification between duplicates genomes
The calling of SNPs within cultures was done using snippy v4.3.6 (https://github.com/tseemann/snippy) with default parameters. SNPs between resequenced genomes at different time points were identified using the genome sequenced at the first time point (T0) as reference (Table S1). We excluded small indels and multiple nucleotide polymorphism and focused only on SNPs. We also excluded SNPs with a fixed derived allele when mapping reads from T0 against the T0 assembly, which were likely errors due the assembly.
Homologous recombination rates across Microcystis genomic clusters
Using the 33 newly sequenced Microcystis genomes (excluding resequenced duplicates), we investigated rates of homologous recombination within and between genomic clusters or across the phylogenetic tree. To do this, we estimated the relative effect of recombination vs. mutation (r/m) rates using ClonalFrameML v1.11-3 [52]. Briefly, the degapped core genome alignment generated by Mugsy v2.2.1 (1 274 628 bp) [45] was split in several subalignments using pyfasta (https://pypi.org/project/pyfasta/). The subalignments corresponded to the genomic clusters defined the core phylogeny and ANI analyses described above. In order to estimate r/m between clusters, we also created subalignments for pair of clusters. ClonalFrameML was executed using a bootstrap of 100 replicates (emsim = 100). The input phylogenies given in ClonalFrameML were generated using RAxML with bootstrap of 100 replicates [44]. The transition/transversion ratios also used as an inputs in ClonalFrameML were estimated using PHYML v3.0 under the model of nucleotides substitution HKY85 [53]. ClonalFrameML analyses excluded the two smallest Microcystis genomes Ma_AC_P_00000000_S299 and Ma_QC_C_20070823_S18.
Identification of horizontally transferred and locally adapted genes
To identify genes transferred across species boundaries, we screened gene trees for instances of two distinct species (monophyletic clusters in the species tree) clustering together in the same monophyletic group, whereas they are normally distantly related in the species tree. As a signature of local adaptation, we additionally screened for such cross-species HGT events that occurred among two different species from the same country. For local adaptation analysis, we worked with the clusters generated with Roary and using 53 Microcystis genomes from Brazil, Canada, USA, and the 26 reference genomes from the NCBI database. Resequenced genomes were included in this analysis to ensure the full complement of genes was considered (i.e., to guard against assembly errors), although in practice they did not affect the identification of local HGT. Once the gene clusters were identified with Roary, the alignments of the nucleotide sequences in each gene cluster (core genes present in >75 strains (>95%) and accessory genes) were generated using the MAFFT software v7.271 [54]. Maximum likelihood phylogenetic trees for each alignment were inferred using FastTreeMP v2.1.8 and the generalized time-reversible model (GTR) for nucleotide substitution [55]. The trees were visualized with graphlan v0.9.7 [56]. The phylogenetic trees that showed local (geographic) adaptation signatures were identified using a Perl script (https://figshare.com/articles/Monophy_screening_tree_files_for_the_detection_of_local_adaptations/7661009/5) to screen phylogenetic trees and identify monophyletic groups with a particular level of bootstrap support (in our case, 90%). The script also allows monophyletic groups including particular combination of isolates (e.g., from different morphospecies or geographic locations) to be identified, with a given minimal branch length (in our case, this parameter was set to 0) and number of isolates (in our case, 4) within the group. A phylogeny was considered as positive for non-local HGT if Canadian and Brazilian isolates were together in the same clade supported by a bootstrap value >90%, while a phylogeny positive for the local HGT showed Brazilian or Canadian isolates, but not both in the same well-supported clade. The phylogenies with a signature of HGT were then manually curated to remove those consisting solely of HGT within a single-monophyletic cluster. Genes in the accessory genome and core genome were functionally annotated using the eggNOG database [570, 58]. The full HGT gene set is reported in Data Set S1.
Inferring secondary metabolic pathways in Microcystis genomes
We evaluated the metabolite profiles for individual Microcystis genomes using the package antiSMASH v4.0.2. The annotated genomes using Prokka (53 genomes reported here and the 26 reference genomes from NCBI) were used as input to AntiSMASH [59]. Two additional biosynthetic clusters absent in the AntiSMASH database (Aeruginosamide (NCBI accessions numbers CCH92964- CCH92969) and Microginin (NCBI accessions numbers CAQ48259-CAQ48262)) were added manually. Based on the antiSMASH results, we generated a matrix of presence–absence of genes related to the biosynthesis of secondary metabolites. The matrix was visualized using the R package ggplot and the function heatmap2 [50]. All-against-all BLASTP analysis was applied to find the best reciprocal hits between proteins in the database and the proteins in the Microcystis genomes [60]. The proteins with the best reciprocal hit were extracted; to be considered as present in the database, the amino acid identity had to be >60%, and >30% of the length of the sequences had to be aligned, with an e-value of 10−5.
Results
Phylogenetic coherence of named morphospecies
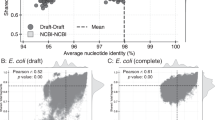
In order to assess the coherence of named Microcystis morphospecies, we sequenced 33 isolates of Microcystis, mainly from Canada and Brazil (see Table S1). These genomes were initially classified into six different morphospecies (M. aeruginosa, M. flos-aquae, M. panniformis, M. wesenbergi, M. viridis, and M. novacekii) and deposited in GenBank under Bioproject number PRJNA507251 (Table S1). Five out of 33 genomes were classified as unknown or ambiguous colony morphologies and indicated with M. or M. sp. designations. The size of the Microcystis assembled genomes ranged between 3.2 and 4.9 Mb with average GC content of 42.7%. The shared core genome consisted of 1260 genes and the pangenome contained 16,928 genes. We supplemented these newly sequenced genomes with 26 additional Microcystis genomes downloaded from GenBank and compared them based on their ANI. Most genomes have pairwise ANI values >95% with some exceptions: 198 out of 6241 pairwise comparisons have values between 93 and 94%, mostly involving comparison with Ma_AC_P_00000000_S299 (see Fig. S3 and Data Set S2). ANI values >95% are generally considered to include members of a single species. However, the Microcystis genomes do not constitute a single homogenous ANI cluster; rather, significant substructure is evident at >95% (Fig. S3).
To explore this substructure, we built a core genome phylogeny using 152 conserved genes also present in an outgroup (Fig. 1) and also clustered the aligned core genomes using ANIb and ANIm values both greater or equal to 96% [46]. Of the 33 newly sequenced genomes, 19 were resequenced after several years in culture. The resequenced genomes (at T1) differed from their ancestor (at T0) by an average of five point mutations (SNPs), excluding two outliers, Ma_QC_C_20070823_S18 and Ma_QC_Ch_20071001_S25 with 144 and 294 mutations, respectively (Table S4). The resequenced genomes always clustered with their ancestor in the phylogenetic tree, suggesting that evolution in the laboratory had little impact on the structure of the phylogeny (Table S4). The SNPs segregating at T1 all had derived alleles at frequencie >0.85, with the vast majority >0.95, suggesting that mutations were approaching fixation, either by drift or selection for laboratory conditions (Fig. S4). To distinguish between drift and selection, we identified mutations that occurred independently in multiple independent cultures of the same monophyletic morphospecies, suggesting laboratory adaptation. We found two parallel SNPs that occurred in all four cultures of M. panniformis, one SNP in all three M. flos-aquae/M. aeruginosa, and two SNPs in all three M. wesenbergii (Table S5). Overall, these results suggest limited but measurable laboratory adaptation is occurring in Microcystis cultures, but the resulting diversity is not sufficient to affect the global phylogeny (Fig. 1) or subsequent analyses.
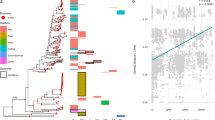
Phylogenetic tree of 53 Brazilian, Canadian, and USA Microcystis genomes and 26 globally sampled reference genomes. A core genome of 152 homologous genes shared by 79 Microcystis genomes and the outgroups (Anabaena variabilis ATCC29413 and Synechocystis sp. PCC6803) was used to infer the phylogeny (Methods). The five ANI clusters corresponding to monophyletic clades with at least three new sequenced genomes are highlighted in colored boxes. The genomes from the same isolate at a different time have a “D or T” at the end of their names and are indicated with a black asterisk. The genomes from a bulk culture and a single colony from the same cultured are indicated with a hash. The three genomes from uncultured colonies from Lake Champlain (Quebec, Canada) are indicated with a red asterisk and “Col” at the end of their name. The font colors indicate the geographical origin (Brazil: orange, Canada: blue, USA: purple, other: black). The abbreviated ANI cluster names correspond to M. panniformis (Mpa), M. flos-aquae/M. aeruginosa (Mfl_ae1 and Mf_ae2), M. wesenbergii (Mwe), M. viridis (Mvi), M. novacekii (Mn), M. sp. (Msp), and M. aeruginosa (Mae1, Mae2, Mae3, Mae4, Mae5, Mae6, Mae7, Mae8, Mae9, and Mae10). The color symbols correspond to ANI clusters and the shapes to the morphospecies: M. aeruginosa (square), M. flos-aquae (circle), M. panniformis (star), M. wesenbergi (three-point star), M. viridis (triangle), M. novacekii (cross), and ambiguous/unknown morphotypes M. or M. sp. (rhombus). The tree bar scale indicates the number of nucleotide substitutions per site
Additionally, we isolated and sequenced a single colony from one of the batch cultures. This colony genome (S217Col) clustered on the phylogeny with its parent culture (S217Cul) with a phylogenetic distance of zero (Fig. 1), suggesting that a single colony is representative of the entire culture.
The ANI analysis yielded 17 clusters, which we named according to their predominant morphospecies (e.g., Mae1, Mae2, etc.), indicated alongside the phylogenetic tree (Fig. 1). Unknown or ambiguous colony morphologies are indicated with M. or M. sp. designations (Fig. 1 and Table S1). Five out of 17 clusters have at least three newly sequenced genomes (excluding resequenced duplicates) and correspond to monophyletic groups: Mpa, Mfl_ae1, Mwe, Mae1, and Mae2 (highlighted in Fig. 1). Four out of five monophyletic clades (Mpa, Mfl_ae1, Mae1, and Mae2) were identical to hierBAPS clusters, while one clade (Mwe) is merged with another hierBAPS cluster, likely due to long-branch attraction [46, 61]. Four out of the five ANI clusters included precisely one morphospecies, whereas the Mfl_ae1 cluster includes mainly M. flos-aquae morphospecies, but also includes some M. aeruginosa (and this remains true after NCBI reference genomes from France and Japan are excluded). This supports the idea that morphology is generally stable, but can still vary among genetically near-identical isolates. Similarly, M. aeruginosa was paraphyletic and distributed across 12 ANI clusters. Based on these phylogenetic, ANI and population structure analyses, Microcystis appears to comprise at least five well-defined monophyletic groups (Mpa, Mfl_ae1, Mwe, Mae1, and Mae2), four of which are congruent with morphospecies (Mae1, Mae2, Mpa, and Mwe). M. viridis and M. novacekii, also corresponded to monophyletic groups, but were represented by <3 genomes. Thus, further sampling is needed to determine the coherence of these groups.
Microcystis is globally distributed, and it is thought that rapid dispersal abolishes any biogeographic associations [10, 14, 62]. Of the five well-sampled and well-supported clades, only two monophyletic clusters (Mae2 and Mpa) were specific to a particular geographic location (Canada or Brazil, respectively), suggesting possible local adaptation, recent clonal expansion, or reduced migration of these lineages. We note that Mpa was isolated from two different reservoirs in Brazil (MB and GB) and at two different time points, 2005 and 2008 (Fig. 1 and Table S1). Thus, Mpa may represent a Brazil-specific, but not a lake-specific clonal expansion. Two other “cosmopolitan” clades have geographically associated subclades: a Canada-associated group within Mfl_ae1 and two subclades within Mwe, respectively, associated with Canada or Brazil (Fig. 1). Only Mae1, a relatively diverse clade, appears to be truly cosmopolitan. Even if Microcystis are generally closely related at >95% ANI, there is clear and significant substructure within the genus, some of which is geographically associated over recent time scales.
We next investigated the pangenome content within each of the five ANI clusters with at least three newly sequenced genomes. The three clusters corresponding to M. panniformis, M. wesenbergii, and M. flos-aquae/M. aeruginosa had highly conserved core genomes (between 77% and 98% of genes shared by all members of the cluster) while M. aeruginosa morphospecies had much smaller core genomes (between 54% and 61% of genes shared by all members; see Table S3). This is consistent with the paraphyletic M. aeruginosa being an amalgam of high-genetic diversity and variable gene content.
Higher homologous recombination rates within than between clusters supports a BSC-like concept
We next asked if homologous recombination could explain the cohesion of the monophyletic groups. To address this question, we estimated the ratio of homologous recombination to mutation rates (r/m) within and between the five well-sampled and well-supported ANI groups. We found that 3/5 of these groups (M. panniformis, M. wesenbergii, and M. flos-aquae/M, aeruginosa) have r/m ratios 2–3x higher within than between clades (Fig. 2 and Table S6). Recombination rates were generally low for M. aeruginosa both within and between clades (Fig. 2). Overall, these results suggest that the cohesion of at least three monophyletic groups could be driven or reinforced by preferential recombination within vs. between groups, consistent with a BSC-like model of speciation. Conversely, the other two groups, consisting of M. aeruginosa, appeared to engage in relatively little recombination compared to mutation, and thus defied delineation based on the BSC.
Frequent local horizontal gene transfer (HGT)
From the core genome phylogenetic tree, certain monophyletic clades showed strong geographic preferences. For example, M. aeruginosa (Mae2) were found uniquely in Canada, while M. panniformis was found only in Brazil (Fig. 1). In contrast, M. wesenbergii is found in both Canada and Brazil (Fig. 1), suggesting a species coherence that transcends geography boundaries. As previously observed in Vibrio, different species in the same geographic region may exchange genes, possibly leading to local adaptation [7]. To identify potential locally adapted Microcystis genes, we screened gene trees for instances where two different named morphospecies (which formed distinct monophyletic groups in the species tree; Fig. 1) clustered together in the same monophyletic group (with bootstrap support >90%), consistent with cross-species HGT. As we were particularly interested in local HGT, we identified monophyletic groups of two distinct species, all isolated from the same region (i.e., Canada or Brazil, but not both). We screened a total of 25,157 core and accessory genes from 79 Microcystis genomes (53 reported here and 26 previously published). We considered 12,084 informative gene trees (that included four or more leafs). Of these trees, 593 (4.9% of the total) showed a pattern of non-local HGT (with Canadian and Brazilian isolates grouping together in the same well-supported clade), whereas slightly more (959 genes; 7.9% of the total) were consistent with local HGT. This suggests that geography, and possibly local adaptation, is an important factor in shaping rates of HGT. Local HGT events, on average, appear to be more recent than non-local events: in 77 out of 959 local HGTs, the phylogenetic distances within the recombined clade were equal to zero (Table 1), suggesting more recent HGT in local compared to non-local events (Fisher’s exact test, Odds ratio = 1.76, P = 0.012). Local HGTs also include more functionally annotated genes, compared to non-local HGTs, which involve mostly hypothetical genes (Table 1; Fisher’s exact test, Odds ratio = 2.93, P < 2.2e-16). While these differences could have many possible explanations, we speculate that non-local HGT events are enriched in phages and other poorly annotated mobile or selfish genetic elements, while local HGTs involve metabolically or ecologically relevant genes, which are more likely to have been studied and annotated. Consistent with this explanation, the non-local pangenome is dominated by genes involved in DNA replication, recombination and repair (COG category L, Fig. S5; X2 test, P < 0.05 after Bonferroni correction for multiple hypothesis testing), which is suggestive of self-replicating and recombining mobile elements (Fig. S5 and Data Set S1). Overall, these results suggest that local HGT events are relatively recent (and thus more frequently observed) and possibly more ecologically relevant (and less “selfish”) than non-local HGTs.
Figure 3 illustrates a few noteworthy examples of genes involved in local HGT among monophyletic clusters. For example, the phylogenetic trees of two neighboring genes encoding the hicA-hicB toxin-antitoxin system showed phylogenetic distances almost equal to zero (between 0 and 0.0008 substitutions per site), clustering Brazilian genomes of three different monophyletic ANI clusters into a single group (Fig. 3a, b), whereas these clusters are well-separated on the species tree (Fig. 1). This suggests that the toxin-antitoxin system has been subject to recent cross-species HGT in Brazil. The hicAB module is a mobile element that has been previously described in bacteria, archaea, plasmids, and phages [63, 64] and at least 31 hicB antitoxins and 21 hicA toxins have been reported in M. aeruginosa [65]. The hicAB module seems to act as a phage defense system, arresting cell growth in response to phage infection [65].
Phylogenetic trees of selected accessory genes showing evidence of local HGT a hicA, b hicB, c two Cas1 genes, and d mcyB. The symbols correspond to ANI clusters from Fig. 1. The font colors of indicate geographic origin. The clades showing geographic signatures (local HGT) are highlighted in orange (Brazil) and blue (Canada). The phylogenetic trees contain all the homologs (branches) of the gene family, except the mcyB tree, which is a subclade within a larger tree (Fig. S6). The subclade was chosen to better visualize the local HGT event. The bars below the trees indicate units of nucleotide substitutions per site
Another two genes encoding CRISPR cas1 endonucleases also showed a signature of recent local HGT, with phylogenetic distances almost equal to zero (between 0 and 0.0001), clustering Canadian genomes in a single clade (Fig. 3c). These two genes are neighbors located on the same contig, flanked by a hypothetical gene and a CRISPR-associated endoribonuclease (cas2). We also identified local HGT events involving other toxin-antitoxin genes, cyanotoxins (such as mcyB; Fig. 3d), endonucleases, and others (Data Set S1).
Clade-specific profiles of biosynthetic gene clusters
Having shown examples of cyanotoxin genes being involved in local HGT (Fig. 3d and Data Set S1), we sought to more broadly characterize the distribution of cyanotoxins and other biosynthetic gene clusters across Microcystis clades. Specifically, we asked whether Microcystis clades or morphospecies tended to have a characteristic profile of biosynthetic genes, despite potentially rapid gain and loss of these genes. The biosynthesis genes of secondary metabolites are usually found in gene clusters [66, 67]. Microcystis species can synthesize a variable number of secondary metabolites, many of which are toxic to humans and other animals [68, 69].
We identified 34 known secondary metabolite gene clusters within all the Microcystis genomes using the software AntiSMASH (Fig. 4 and Table S7). AntiSMASH identifies these genes based on a protein database and NRPS (nonribosomal peptide synthetases) and PKS (polyketide synthase) domain analysis [59]. Eight out of 34 secondary metabolic gene clusters were present and complete in at least one Microcystis genome (Fig. 4).
Distribution of biosynthetic gene clusters across Microcystis. a Phylogenetic tree of 53 Brazilian, Canadian, and USA Microcystis genomes and 26 reference genomes. b Presence and absence of the genes encoding secondary metabolites in each Microcystis genome are shown as a heatmap. Rows and columns represent the genomes and genes, respectively. The presence and absence of genes are indicated in blue and white, respectively. The shade of blue increases with the amino acid similarity to the reference database. The ten biosynthetic clusters are enclosed by colored rectangles and their names appear at the bottom of the figure
We observed that Microcystis genomes lacking the microcystin cluster (mcy) usually contained another gene cluster instead. For example, M. flos-aquae/M. aeruginosa (Mfl_ae1) lacked mcy but instead had genes related to the biosynthesis of anabaenopeptins (apn) (Fig. 4). However, other studies have found microcystin-producing strains of M. flos-aquae [70], suggesting that the genomes reported here likely undersample the diversity of biosynthetic gene clusters present in nature. Consistent with relatively high diversity within clades, genomes within the same clade tended to have similar, but non-identical profiles of gene clusters, with M. aeruginosa clades being among the most diverse. M. aeruginosa clades also tended to have a high coding potential for toxins, including the complete mcy and mdn (microviridin B) gene clusters. However, some genomes in M. aeruginosa subclade Mae1 lacked the mcy genes, consistent with loss or HGT. In contrast, M. wesenbergii encoded relatively few biosynthetic gene clusters, consistent with previously reported low toxin production and microcystin gene absence [70,71,72,73].
Viewed in aggregate, these biosynthetic gene clusters are part of the Microcystis accessory genome. However, certain gene clusters are core to specific monophyletic groups. In M. flos-aquae for example, anp genes were always present (core) and mcy genes were absent. These group-specific core gene clusters could provide potential niche adaptations and ecological distinctness. On the other hand, certain biosynthetic genes such as mcyB (Fig. 3d) are exchanged across species boundaries. Thus, biosynthetic genes may contribute to both species-specific and species-transcending adaptations [74].
Discussion
In this study, we investigated the correspondence among morphospecies and genome-informed species definitions using dozens of Microcystis isolates from both Northern and Southern hemispheres, primarily from Canada and Brazil. We assessed the genomic cohesion of Microcystis clades within Microcystis species by measuring the genome similarities (phylogeny, hierBAPS clustering and ANI values) and homologous recombination within and between clades.
We found that Microcystis genomes used in this study together with the reference genomes fell into a single-genomic complex (ANI values >95%). Previous studies have suggested a universal cutoff of 95% ANI as adequate for species delineation. These studies described a genetic discontinuity or bimodal distribution with peaks >95% (intra-species) and <83% (inter-species), but the mechanism for this discontinuity is unclear, and it is difficult to exclude sampling bias as a reason for the discontinuity [48, 75]. We observed that within a 95% ANI cluster of Microcystis, there is substantial genetic substructure, potentially containing distinct species or sub-species. Four of the 17 sub-clusters we identified corresponded to named morphospecies and had at least three new sequenced genomes, while the others were mostly composed of genomes classified as M. aeruginosa morphospecies. We concluded that M. aeruginosa is paraphyletic with a mixed geographical pattern, while the morphospecies M. panniformis and M. wesenbergii consisted of well-defined clades within Microcystis species complex (Fig. 1).
What are the mechanisms that can explain the genetic structure within Microcystis? Ecological selection (the ESC), barriers to gene flow (BSC-like), and biogeography (allopatric divergence) could all play a role. Previous studies based on a smaller sample of Microcystis genomes [10], or marker genes [14, 15] suggested that there are few if any biogeographic barriers in Microcystis to migrate, leading to a globally mixed population [76, 77]—and that Microcystis should be defined as a single species [17, 18]. Consistent with this, M. aeruginosa is globally distributed and paraphyletic in our core phylogeny. However, two monophyletic morphospecies tend to be geographically restricted, possibly due in this case to limited migration (at least on recent time scales) and/or local adaptation. The two morphospecies could also represent short-lived clonal expansions, or biases due to incomplete sampling. However, to minimize bias, our sampling was performed repeatedly over 15 years, with similar methods in both Brazil (four sampling locations) and Canada (10 sampling locations). Thus, although we cannot exclude the possibility of unsampled refugia in either Canada or Brazil, we sampled as thoroughly and consistently as possible. We also inferred more frequent homologous recombination within than between monophyletic clades (Fig. 2), consistent with a BSC-like model maintaining genetic distinctness [78,79,80]. While the coherence of these clades may thus be maintained by barriers to gene flow, we suspect that their divergence was initiated by selection for ecological distinctness [1]. Although the precise ecological differences between clades are unknown [73, 81], we found that each of the monophyletic clades had a distinct core genome and distinct profile of biosynthetic gene clusters (Fig. 4)—both of which could provide potential ecological adaptations. Further experimental study will be required to fully test the hypothesis of ecological distinctness among clades within Microcystis.
The BSC-like model requires more frequent recombination within than between species, but also allows occasional recombination of “globally adaptive” genes across species boundaries. Similar to previous observations in Vibrio [7], we inferred a significant proportion of cross-species HGT events occurred within the same geographic location, suggesting local environmental adaptation. Local HGT events tend to be phylogenetically more recent than non-local events, suggesting that they occur at relatively higher frequency. Local HGTs are also enriched in genes of annotated (non-hypothetical) function, including cyanotoxin genes such as microcystin (mcyB) and cyanopeptolin (mcnC, mcnF and mcnG). Microcystin genes are likely of ancient origin in cyanobacteria [82] and it has been suggested that they subsequently experienced significant homologous recombination and positive selection [83, 84]. Microcystin genes also show biogeographic patterns. For example, the mcyD gene has distinctive alleles found in Japanese Microcystis isolates but not elsewhere [85]. Our inference of local, cross-species HGT of toxin genes further supports the idea that they may be locally adapted.
Genes involved in phage defense systems were also involved in local HGT events, suggesting that local adaptation could be driven by local viruses. First, the hicAB operon appears to have been shared among at least three distinct ANI clusters (Mvi, Mpa, and Mn) in Brazil (Fig. 3), and has previously been suggested to be involved in phage defense and prone to HGT [63, 65]. Second, cas1, which encodes the most conserved protein in the CRISPR–Cas defense system [86], appears to have been exchanged among three distinct clusters of M. aeruginosa in Canada (Fig. 3). This is consistent with previous evidence, suggesting that CRISPR–Cas genes are subject to HGT and natural selection [86, 87]. Thus, local HGT could promote adaptation to local phages.
Taken together, our results resolve some of the longstanding confusion surrounding Microcystis species and suggest new avenues for future research. While all Microcystis genomes sampled to date are monophyletic and closely related, there is significant genetic substructure, suggesting the existence of several distinct species. The distinctiveness of these species appears to be maintained by barriers to gene flow, consistent with a BSC-like model [1, 5]. Whether gene flow barriers are mainly geographic, genetic, or ecological is a subject for future investigation. While different Microcystis species appear to inhabit different niches, as evidence by geographic preferences and distinct profile of biosynthetic gene clusters, the nature of their ecological distinctiveness should also be a subject of future field and laboratory studies.
Data availability
The raw reads and the Microcystis genomes contigs are available in GenBank under Bioproject number PRJNA507251 (Table S1).
References
Shapiro BJ. What microbial population genomics has taught us about speciation. In: Polz M, Rajora O (editors) Population genomics: Microorganisms Population genomics. Cham: Springer; 2018;31–47.
Wiedenbeck J, Cohan FM. Origins of bacterial diversity through horizontal genetic transfer and adaptation to new ecological niches. FEMS Microbiol Rev. 2011;35:957–76.
Whitaker RJ, Grogan DW, Taylor JW. Recombination shapes the natural population structure of the hyperthermophilic archaeon Sulfolobus islandicus. Mol Biol Evol. 2005;22:2354–61.
Cadillo-Quiroz H, Didelot X, Held NL, Herrera A, Darling A, Reno ML, et al. Patterns of gene flow define species of thermophilic Archaea. PLoS Biol. 2012;10:e1001265.
Bobay LM, Ochman H. Biological species are universal across Life’s domains. Genome Biol Evol. 2017;9:491–501.
Reno ML, Held NL, Fields CJ, Burke PV, Whitaker RJ. Biogeography of the Sulfolobus islandicus pan-genome. Proc Natl Acad Sci USA. 2009;106:8605–10.
Boucher Y, Cordero OX, Takemura A, Hunt DE, Schliep K, Bapteste E, et al. Local mobile gene pools rapidly cross species boundaries to create endemicity within global Vibrio cholerae populations. MBio. 2011;2:e00335–10.
van Elsas JD, Bailey MJ. The ecology of transfer of mobile genetic elements. FEMS Microbiol Ecol. 2002;42:187–97.
Whittaker KA, Rynearson TA. Evidence for environmental and ecological selection in a microbe with no geographic limits to gene flow. Proc Natl Acad Sci USA. 2017;114:2651–6.
Humbert JF, Barbe V, Latifi A, Gugger M, Calteau A, Coursin T, et al. A tribute to disorder in the genome of the bloom-forming freshwater cyanobacterium Microcystis aeruginosa. PLoS ONE. 2013;8:e70747.
Tanabe Y, Kaya K, Watanabe MM. Evidence for recombination in the microcystin synthetase (mcy) genes of toxic cyanobacteria Microcystis spp. J Mol Evol. 2004;58:633–41.
Tanabe Y, Kasai F, Watanabe MM. Multilocus sequence typing (MLST) reveals high genetic diversity and clonal population structure of the toxic cyanobacterium Microcystis aeruginosa. Microbiology. 2007;153:3695–703.
Tanabe Y, Sano T, Kasai F, Watanabe MM. Recombination, cryptic clades and neutral molecular divergence of the microcystin synthetase (mcy) genes of toxic cyanobacterium Microcystis aeruginosa. BMC Evol Biol. 2009;9:115.
van Gremberghe I, Leliaert F, Mergeay J, Vanormelingen P, Van der Gucht K, Debeer AE, et al. Lack of phylogeographic structure in the freshwater cyanobacterium Microcystis aeruginosa suggests global dispersal. PLoS ONE. 2011;6:e19561.
Harke MJ, Steffen MM, Gobler CJ, Otten TG, Wilhelm SW, Wood SA, et al. A review of the global ecology, genomics, and biogeography of the toxic cyanobacterium, Microcystis spp. Harmful Algae. 2016;54:4–20.
Shapiro BJ, Leducq JB, Mallet J. What is speciation? PLoS Genet. 2016;12:e1005860.
Otsuka S, Suda S, Li RH, Watanabe M, Oyaizu H, Matsumoto S, et al. Phylogenetic relationships between toxic and non-toxic strains of the genus Microcystis based on 16S to 23S internal transcribed spacer sequence. FEMS Microbiol Lett. 1999;172:15–21.
Otsuka S, Suda S, Shibata S, Oyaizu H, Matsumoto S, Watanabe MM. A proposal for the unification of five species of the cyanobacterial genus Microcystis Kutzing ex Lemmermann 1907 under the rules of the Bacteriological Code. Int J Syst Evol Microbiol. 2001;51:873–9.
Komárek J. Recent changes (2008) in cyanobacteria taxonomy based on a combination of molecular background with phenotype and ecological consequences (genus and species concept). Hydrobiologia. 2010;639:245–59.
Xiao M, Li M, Reynolds CS. Colony formation in the cyanobacterium Microcystis. Biol Rev Camb Philos Soc. 2018;93:1399–420.
Orr PT, Jones GJ. Relationship between microcystin production and cell division rates in nitrogen‐limited Microcystis aeruginosa cultures. Limnol Ocean. 1998;43:1604–14.
Compère P. The nomenclature of Cyanophyta under the Botanical Code. Arch Hydrobiol Suppl Algol Stud. 2005;117:31–7.
Zhang JY, Guan R, Zhang HJ, Li H, Xiao P, Yu GL, et al. Complete genome sequence and genomic characterization of Microcystis panniformis FACHB 1757 by third-generation sequencing. Stand Genom Sci. 2016;11:11.
Otsuka S, Suda S, Li RH, Matsumoto S, Watanabe MM. Morphological variability of colonies of Microcystis morphospecies in culture. J Gen Appl Microbiol. 2000;46:39–50.
Ma J, Brookes JD, Qin B, Paerl HW, Gao G, Wu P, et al. Environmental factors controlling colony formation in blooms of the cyanobacteria Microcystis spp. in Lake Taihu, China. Harmful Algae. 2014;31:136–42.
Wang WJ, Zhang YL, Shen H, Xie P, Yu J. Changes in the bacterial community and extracellular compounds associated with the disaggregation of Microcystis colonies. Biochem Syst Ecol. 2015;61:62–6.
Komárek J. A review of water-bloom forming Microcystis species, with regard to populations from Japan. Arch Hydrobiol Suppl Algol Stud. 1991;64:115–27.
Castenholz RW, Norris TB. Revisionary concepts of species in the cyanobacteria and their applications. Arch Hydrobiol Suppl Algol Stud. 2005;117:53–69.
Xu S, Sun Q, Zhou X, Tan X, Xiao M, Zhu W, et al. Polysaccharide biosynthesis-related genes explain phenotype-genotype correlation of Microcystis colonies in Meiliang Bay of Lake Taihu, China. Sci Rep. 2016;6:35551.
Konstantinidis KT, Tiedje JM. Towards a genome-based taxonomy for prokaryotes. J Bacteriol. 2005;187:6258–64.
Christiansen G, Molitor C, Philmus B, Kurmayer R. Nontoxic strains of cyanobacteria are the result of major gene deletion events induced by a transposable element. Mol Biol Evol. 2008;25:1695–704.
Komárek J, Komárková-Legnerová J, Santanna CL, Azevedo MTP, Senna PC. Two common Microcystis species (Chroococcales, Cyanobacteria) from tropical America, including M. panniformis sp. nov. Cryptogam Algol. 2002;23:159–77.
Komárek J, Komárková J. Review of the European Microcystis-morphospecies (Cyanoprokaryotes) from nature. Czech Phycol. 2002;2:1–24.
Peng Y, Leung HC, Yiu SM, Chin FY. IDBA-UD: a de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics. 2012;28:1420–8.
Meyer KA, Davis TW, Watson SB, Denef VJ, Berry MA, Dick GJ. Genome sequences of lower Great Lakes Microcystis sp reveal strain-specific genes that are present and expressed in western Lake Erie blooms. PLoS ONE. 2017;12:e0183859.
Li Q, Lin F, Yang C, Wang J, Lin Y, Shen M, et al. A large-scale comparative metagenomic study reveals the functional interactions in six bloom-forming Microcystis-epibiont communities. Front Microbiol. 2018;9:746.
Eren AM, Esen OC, Quince C, Vineis JH, Morrison HG, Sogin ML, et al. Anvi’o: an advanced analysis and visualization platform for ‘omics data. PeerJ. 2015;3:e1319.
Langmead B, Salzberg SL. Fast gapped-read alignment with Bowtie 2. Nat Methods. 2012;9:357–U54.
Kim D, Song L, Breitwieser FP, Salzberg SL. Centrifuge: rapid and sensitive classification of metagenomic sequences. Genome Res. 2016;26:1721–9.
Hyatt D, Chen GL, Locascio PF, Land ML, Larimer FW, Hauser LJ. Prodigal: prokaryotic gene recognition and translation initiation site identification. BMC Bioinforma. 2010;11:119.
Hyatt D, LoCascio PF, Hauser LJ, Uberbacher EC. Gene and translation initiation site prediction in metagenomic sequences. Bioinformatics. 2012;28:2223–30.
Seemann T. Prokka: rapid prokaryotic genome annotation. Bioinformatics. 2014;30:2068–9.
Edgar RC. MUSCLE: a multiple sequence alignment method with reduced time and space complexity. BMC Bioinforma. 2004;5:113.
Stamatakis A, Aberer AJ, Goll C, Smith SA, Berger SA, Izquierdo-Carrasco F. RAxML-Light: a tool for computing terabyte phylogenies. Bioinformatics. 2012;28:2064–6.
Angiuoli SV, Salzberg SL. Mugsy: fast multiple alignment of closely related whole genomes. Bioinformatics. 2011;27:334–42.
Cheng L, Connor TR, Siren J, Aanensen DM, Corander J. Hierarchical and spatially explicit clustering of DNA sequences with BAPS software. Mol Biol Evol. 2013;30:1224–8.
Richter M, Rossello-Mora R. Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA. 2009;106:19126–31.
Konstantinidis KT, Tiedje JM. Genomic insights that advance the species definition for prokaryotes. Proc Natl Acad Sci USA. 2005;102:2567–72.
Konstantinidis KT, Ramette A, Tiedje JM. Toward a more robust assessment of intraspecies diversity, using fewer genetic markers. Appl Environ Microbiol. 2006;72:7286–93.
Wickham H. Elegant graphics for data analysis. New York: Springer-Verlag; 2016.
Page AJ, Cummins CA, Hunt M, Wong VK, Reuter S, Holden MT, et al. Roary: rapid large-scale prokaryote pan genome analysis. Bioinformatics. 2015;31:3691–3.
Didelot X, Wilson DJ. ClonalFrameML: efficient inference of recombination in whole bacterial genomes. PLoS Comput Biol. 2015;11:e1004041.
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O. New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol. 2010;59:307–21.
Katoh K, Standley DM. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol. 2013;30:772–80.
Price MN, Dehal PS, Arkin AP. FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS ONE. 2010;5:e9490.
Asnicar F, Weingart G, Tickle TL, Huttenhower C, Segata N. Compact graphical representation of phylogenetic data and metadata with GraPhlAn. PeerJ. 2015;3:e1029.
Huerta-Cepas J, Szklarczyk D, Forslund K, Cook H, Heller D, Walter MC, et al. eggNOG 4.5: a hierarchical orthology framework with improved functional annotations for eukaryotic, prokaryotic and viral sequences. Nucl Acids Res. 2016;44:D286–93.
Huerta-Cepas J, Forslund K, Coelho LP, Szklarczyk D, Jensen LJ, von Mering C, et al. Fast genome-wide functional annotation through orthology assignment by eggNOG-Mapper. Mol Biol Evol. 2017;34:2115–22.
Blin K, Wolf T, Chevrette MG, Lu X, Schwalen CJ, Kautsar SA, et al. antiSMASH 4.0-improvements in chemistry prediction and gene cluster boundary identification. Nucl Acids Res. 2017;45:W36–41.
Altschul SF, Madden TL, Schaffer AA, Zhang J, Zhang Z, Miller W, et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucl Acids Res. 1997;25:3389–402.
Castillo-Ramirez S, Corander J, Marttinen P, Aldeljawi M, Hanage WP, Westh H, et al. Phylogeographic variation in recombination rates within a global clone of methicillin-resistant Staphylococcus aureus. Genome Biol. 2012;13:R126.
Belykh OI, Dmitrieva OA, Gladkikh AS, Sorokovikova EG. Identification of toxigenic Cyanobacteria of the genus Microcystis in the Curonian Lagoon (Baltic Sea). Oceanology+. 2013;53:71–9.
Makarova KS, Grishin NV, Koonin EV. The HicAB cassette, a putative novel, RNA-targeting toxin-antitoxin system in archaea and bacteria. Bioinformatics. 2006;22:2581–4.
Makarova KS, Wolf YI, Koonin EV. Comprehensive comparative-genomic analysis of type 2 toxin-antitoxin systems and related mobile stress response systems in prokaryotes. Biol Direct. 2009;4:19.
Makarova KS, Wolf YI, Snir S, Koonin EV. Defense islands in bacterial and archaeal genomes and prediction of novel defense systems. J Bacteriol. 2011;193:6039–56.
Welker M, von Dohren H. Cyanobacterial peptides—nature’s own combinatorial biosynthesis. FEMS Microbiol Rev. 2006;30:530–63.
Sussmuth RD, Mainz A. Nonribosomal peptide synthesis-principles and prospects. Angew Chem Int Ed Engl. 2017;56:3770–821.
Tillett D, Parker DL, Neilan BA. Detection of toxigenicity by a probe for the microcystin synthetase A gene (mcyA) of the cyanobacterial genus Microcystis: comparison of toxicities with 16S rRNA and phycocyanin operon (Phycocyanin Intergenic Spacer) phylogenies. Appl Environ Microbiol. 2001;67:2810–8.
Pearson LA, Dittmann E, Mazmouz R, Ongley SE, D’Agostino PM, Neilan BA. The genetics, biosynthesis and regulation of toxic specialized metabolites of cyanobacteria. Harmful Algae. 2016;54:98–111.
Via-Ordorika L, Fastner J, Kurmayer R, Hisbergues M, Dittmann E, Komarek J, et al. Distribution of microcystin-producing and non-microcystin-producing Microcystis sp. in European freshwater bodies: detection of microcystins and microcystin genes in individual colonies. Syst Appl Microbiol. 2004;27:592–602.
Yoshida M, Yoshida T, Satomi M, Takashima Y, Hosoda N, Hiroishi S. Intra-specific phenotypic and genotypic variation in toxic cyanobacterial Microcystis strains. J Appl Microbiol. 2008;105:407–15.
Xu Y, Wu Z, Yu B, Peng X, Yu G, Wei Z, et al. Non-microcystin producing Microcystis wesenbergii (Komarek) Komarek (Cyanobacteria) representing a main waterbloom-forming species in Chinese waters. Environ Pollut. 2008;156:162–7.
Otten TG, Paerl HW. Phylogenetic inference of colony isolates comprising seasonal Microcystis blooms in Lake Taihu, China. Micro Ecol. 2011;62:907–18.
Majewski J, Cohan FM. Adapt globally, act locally: the effect of selective sweeps on bacterial sequence diversity. Genetics. 1999;152:1459–74.
Jain C, Rodriguez RL, Phillippy AM, Konstantinidis KT, Aluru S. High throughput ANI analysis of 90K prokaryotic genomes reveals clear species boundaries. Nat Commun. 2018;9:5114.
Moreira C, Spillane C, Fathalli A, Vasconcelos V, Antunes A. African origin and europe-mediated global dispersal of the cyanobacterium Microcystis aeruginosa. Curr Microbiol. 2014;69:628–33.
Marmen S, Aharonovich D, Grossowicz M, Blank L, Yacobi YZ, Sher DJ. Distribution and habitat specificity of potentially-toxic Microcystis across climate, land, and water use gradients. Front Microbiol. 2016;7:271.
Fraser C, Hanage WP, Spratt BG. Recombination and the nature of bacterial speciation. Science. 2007;315:476–80.
Vos M, Didelot X. A comparison of homologous recombination rates in bacteria and archaea. ISME J. 2009;3:199–208.
Shapiro BJ, Polz MF. Ordering microbial diversity into ecologically and genetically cohesive units. Trends Microbiol. 2014;22:235–47.
Imai H, Chang KH, Kusaba M, Nakano S. Temperature-dependent dominance of Microcystis (Cyanophyceae) species: M. aeruginosa and M. wesenbergii. J Plankton Res. 2009;31:171–8.
Rantala A, Fewer DP, Hisbergues M, Rouhiainen L, Vaitomaa J, Borner T, et al. Phylogenetic evidence for the early evolution of microcystin synthesis. Proc Natl Acad Sci USA. 2004;101:568–73.
Mikalsen B, Boison G, Skulberg OM, Fastner J, Davies W, Gabrielsen TM, et al. Natural variation in the microcystin synthetase operon mcyABC and impact on microcystin production in Microcystis strains. J Bacteriol. 2003;185:2774–85.
Tooming-Klunderud A, Fewer DP, Rohrlack T, Jokela J, Rouhiainen L, Sivonen K, et al. Evidence for positive selection acting on microcystin synthetase adenylation domains in three cyanobacterial genera. BMC Evol Biol. 2008;8:256.
Moreira C, Vasconcelos V, Antunes A. Phylogeny of microcystins: evidence of a biogeographical trend? Curr Microbiol. 2013;66:214–21.
Makarova KS, Wolf YI, Alkhnbashi OS, Costa F, Shah SA, Saunders SJ, et al. An updated evolutionary classification of CRISPR-Cas systems. Nat Rev Microbiol. 2015;13:722–36.
Godde JS, Bickerton A. The repetitive DNA elements called CRISPRs and their associated genes: evidence of horizontal transfer among prokaryotes. J Mol Evol. 2006;62:718–29.
Acknowledgements
We are grateful to David Bird, Naíla Barbosa da Costa, Mylène Boyer, Julie Marleau, and Coralie Deladrière for assistance isolating strains and maintaining cultures. This work was supported by the Genome Québec and Genome Canada-funded ATRAPP Project (Algal blooms, Treatment, Risk Assessment, Prediction, and Prevention). NT was funded by a project from the European Union’s Horizon 2020 research and innovation program under the Marie Sklodowska-Curie grant agreement No 656647. Cultures collections (Brazilian and some Canadian strains) were partially obtained and maintained thanks to CNPq and FAPEMIG grants to AG. We also want to acknowledge the financial support of the National Research Council.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Pérez-Carrascal, O.M., Terrat, Y., Giani, A. et al. Coherence of Microcystis species revealed through population genomics. ISME J 13, 2887–2900 (2019). https://doi.org/10.1038/s41396-019-0481-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-019-0481-1
This article is cited by
-
The global speciation continuum of the cyanobacterium Microcoleus
Nature Communications (2024)
-
Trade-off in genome turnover events leading to adaptive evolution of Microcystis aeruginosa species complex
BMC Genomics (2023)
-
Intra-population genomic diversity of the bloom-forming cyanobacterium, Aphanizomenon gracile, at low spatial scale
ISME Communications (2023)
-
Single-colony sequencing reveals microbe-by-microbiome phylosymbiosis between the cyanobacterium Microcystis and its associated bacteria
Microbiome (2021)
-
Relationship between morphospecies and microcystin-producing genotypes of Microcystis species in Chinese freshwaters
Journal of Oceanology and Limnology (2021)