Abstract
Under controlled laboratory conditions, high and low ammonium availability are known to favor soil ammonia-oxidizing bacteria (AOB) and archaea (AOA) communities, respectively. However, whether this niche segregation is maintained under field conditions in terrestrial ecosystems remains unresolved, particularly at the global scale. We hypothesized that perennial vegetation might favor AOB vs. AOA communities compared with adjacent open areas devoid of perennial vegetation (i.e., bare soil) via several mechanisms, including increasing the amount of ammonium in soil. To test this niche-differentiation hypothesis, we conducted a global field survey including 80 drylands from 6 continents. Data supported our hypothesis, as soils collected under plant canopies had higher levels of ammonium, as well as higher richness (number of terminal restriction fragments; T-RFs) and abundance (qPCR amoA genes) of AOB, and lower richness and abundance of AOA, than those collected in open areas located between plant canopies. Some of the reported associations between plant canopies and AOA and AOB communities can be a consequence of the higher organic matter and available N contents found under plant canopies. Other aspects of soils associated with vegetation including shading and microclimatic conditions might also help explain our results. Our findings provide strong evidence for niche differentiation between AOA and AOB communities in drylands worldwide, advancing our understanding of their ecology and biogeography at the global scale.
Similar content being viewed by others
Introduction
Nitrification, the aerobic oxidation of ammonia to nitrate, is a core process in the global nitrogen (N) cycle [1, 2] that controls the availability of N for plants and microbes, regulating important ecosystem services such as food and fiber production and soil fertility [3,4,5,6,7,8,9]. Nitrification is a microbial driven process performed by archaea and bacteria holding the gene encoding ammonia monooxygenase (AMO), the enzyme responsible for the conversion of ammonia to nitrite. For over a century, ammonia-oxidizing bacteria (AOB) were thought to be the major driver for ammonia oxidation. However, the discovery of autotrophic ammonia-oxidizing archaea (AOA) changed the perception of the whole nitrification process, especially given the global presence of AOA, their resistance to extreme conditions (e.g., salinity or temperature), and their important role as drivers of soil nitrification [2, 10,11,12].
A strong niche differentiation has been suggested for AOB and AOA communities [2, 13], but yet to be found under field conditions in natural ecosystems across varying environmental conditions [14]. High and low ammonium concentrations have been found to favor AOB and AOA communities, respectively, in laboratory and soil microcosm studies [15, 16]. Thus, although experimental evidence indicates that AOA are often more abundant under oligotrophic conditions (e.g., media with low ammonium content; [15]), while AOB dominate more copiotrophic environments (e.g., media with high ammonium content) [17, 18], it is unknown whether the abundance and diversity of AOA and AOB respond in a predictable manner to changes in ammonium availability in terrestrial ecosystems under field conditions. Therefore, the validity of conclusions obtained from laboratory and microcosm studies remains to be tested under natural conditions, particularly at large biogeographic scales.
Strong physiological evidence supporting contrasting strategies between AOB and AOA is still lacking [2, 13]. AOB are adapted to grow under environmental conditions where ammonium content is high, while AOA are adapted to persist where ammonium content is generally low. AOA are expected to have a higher affinity for ammonium as a substrate than AOB, and to dominate low ammonium environments. Similar contrasting growth strategies are often reported for methanotrophic communities, which are known to follow contrasting life-strategies associated with their affinity for methane [19]. Enzymes encoded by particulate methane oxygenase (pmoA) and amoA genes belong to the same family and have similar oxidation functions. Thus, it would be expected that communities harboring both pmoA and amoA follow similar contrasting life strategies linked to resource availability. Additional evidence for contrasting growth strategies driven by resource availability are found in plant communities. For example, ecological theory and empirical evidence [20] suggest that plant species distribution on Earth is partially supported by the existence of contrasting fast–slow growth strategies associated with the availability of resources (e.g., light). As such, plant species with fast strategies invest in expensive photosynthetic machinery that only pays off when their energy source (light) is abundant, outcompeting slow-growers under high resource conditions [20,21,22]. Conversely, such a machinery is not efficient under low light conditions, where other plant species following a slow-growth strategy outcompete them [20, 22, 23]. Herein, we hypothesized that, similarly to what has been reported for plants, the predicted niche differentiation for AOA and AOB in the laboratory associated with different concentrations of ammonium should translate into recognizable patterns across contrasting environmental conditions in the field.
Recent studies suggest that plants might play an important role in driving the distribution of ammonia oxidizers in soils from local to regional scales [24, 25]. Potentially, different within-plot environments (e.g., plant canopies vs. open areas devoid of perennial vegetation) can reflect different substrate (ammonium) and organic matter conditions, and therefore support potential niche differentiation for soil microbial taxa [26]. In drylands, soils under plant canopies are known to have higher organic matter and ammonium contents compared to adjacent open areas located between plant patches [27]. Because of this, we hypothesized that AOB, which prefer nutrient-rich environments, would dominate under plant canopies (copiotrophic environments), which are known to support higher ammonium contents [27], while AOA would thrive in open environments between plant canopies (i.e., bare soil) characterized by a lower nutrient availability (oligotrophic environments). In other words, AOA are expected to be oligotrophic, stress tolerant and slow-growing organisms that cannot compete with copiotrophic fast-grower AOB under comparatively high ammonium conditions. On the contrary, AOB are not able to outcompete AOA organisms under low nutrient availability, which might limit their presence in the open areas between plant canopies.
To broadly test the niche differentiation hypothesis in realistic ecological settings, we conducted a survey in 80 dryland ecosystems from six continents. Our survey captured a wide range of vegetation attributes, climatic conditions, human influence and soil properties. Drylands offer a unique and important opportunity to evaluate our hypothesis for multiple reasons. First, they occupy ~45% of the Earth’s land surface [28] and maintain 38% of human population [29], hence are critical ecosystems to understand the global nitrogen cycle and the role of microorganisms involved on its multiple processes. More directly relevant to the questions herein, drylands are highly heterogeneous ecosystems that are typically formed by a matrix of discrete plant patches embedded in a matrix of open areas devoid of perennial vegetation [30]. The coexistence of these two distinct environments in drylands (soils under plant canopies and bare soil in open areas devoid of perennial vegetation) provides a unique opportunity to test the above introduced niche differentiation hypothesis for AOA and AOB in field conditions, and to address a critical research gap in the literature. Of course, plant canopies and substrate availability are not expected to be the only ecological predictor of AOA and AOB communities in global drylands. Their abundance and diversity is also likely to be directly and indirectly driven by factors including climate, human impacts (which are known to increase ammonium content; [31], soil properties, and micro-climatic conditions [32,33,34,35,36,37]. Consequently, attempts to identify the importance of niche differentiation driven by plant canopies in regulating the abundance and richness of AOA and AOB should also acknowledge them.
Materials and methods
Field site description and soil sampling
Field data and soil samples were collected from 80 drylands (30 × 30 plots) located in 12 countries from six continents [38] (Fig. 1). Vegetation and soil surveys were carried out between 2006 and 2012 using a standardized sampling protocol [39]. Plant cover in all locations ranged from 2 to 74%. A 13% of our plots were dominated by N-fixer plants (mostly Casuarina sp., Astragalus sp. and Prosopis sp.). Two composite (10–15 samples per site) soil samples were randomly taken (top 7.5 cm) under the canopy of the dominant perennial plant species (vegetation hereafter) and from open areas (bare soil hereafter). We conducted our soil surveys during the dry season in most of the study sites, when biological activity was the lowest. It is known that space is more important than seasonality in driving microbial community structure (e.g., see [40]). Because of this, we do not expect seasonality to be an important factor influencing our conclusions. We collected samples under perennial plant species rather than under annual plants because the former are expected to have the largest influence in dryland ecosystems (see [39] for a detailed explanation of the plant surveys conducted and a rationale for the use of perennial plants). Furthermore, and given the selected season for conducting the soil surveys, the presence of annual plants in our study sites was minimal. Each location in our study was considered as a replicate. As such, we had 80 replicates collected in open areas and 80 collected under plant canopies. Bare soil samples were collected far from plant canopies (>1m; [41] to minimize any impact from plant roots. Samples were immediately frozen at −20 °C (for molecular analyses) or air dried for 30 days (for physico-chemical analyses) after field collection and were stored for either 2–6 years (molecular) or 0–4 (physico-chemical) years before analyses (depending on the site).
Molecular analyses
Soil microbiological analyses were conducted on composite samples collected under bare soil and vegetated areas at each site. Soil DNA was extracted from 0.5 g of defrosted soil samples using the Powersoil® DNA Isolation Kit (Mo Bio Laboratories, Carlsbad, CA, USA) following manufacturer’s instructions. DNA concentration and quality were determined spectrophotometrically (NanoDrop ND-2000c, Thermo Scientific, MA, USA).
The abundance of AOA and AOB amoA genes was quantified on a Bio-Rad CFX96 Real-Time PCR System (Bio-Rad, Herculers, CA, USA) using the primer pairs CrenamoA23f/CrenamoA616r [42] and amoA-1F/amoA-2R [43], respectively. Each 20 μl quantitative PCR (qPCR) reaction contained 10 µl of SensiFAST SYBR No-ROX reagent (Bioline, Taunton, MA, USA), 0.2 µl of specific forward and reverse primers (10 µM), 2 µl of BSA (5.0 mg ml−1), 2 µl of diluted template DNA (5–10 ng), and 5.6 μl of nuclease free water. Primers and thermal cycling conditions for both AOA and AOB amoA genes are described in Supplementary Table S1. Melting curves were generated for each qPCR run, and fluorescence data were collected at temperatures above the melting temperature of the primers but below that of the target (78 °C for both amoA genes) to verify product specificity. Each qPCR reaction was run in triplicate. Standards for qPCR were constructed by cloning isolated AOA and AOB amoA genes into the pCR®4–TOPO vector (Invitrogen, Carlsbad, CA). A 10-fold serial dilution of plasmid was prepared to generate standard curves.
The community composition and diversity of ammonia oxidizers was characterized by terminal-restriction length polymorphism (T-RFLP) analysis using the fluorescently labeled primers FAM-CrenamoA23f/ CrenamoA616r and VIC-amoA-1F/amoA-2R for AOA and AOB, respectively. Our previous analyses suggested that TRFLP provides similar diversity measures to data from high-throughput sequencing for microbial communities [44, 45] and high throughput microarray for ammonia oxidizing communities [44]. The 50 μl PCR reactions contained 2.5 U of BioTaq DNA polymerase (Bioline), 0.5 μl of each primer (20 μM), 1 μl dNTP mix (20 mM), 5 μl 10 × NH4 reaction buffer, 2 μl BSA (20 mM), 2 μl MgCl2 solution (50 mM) and 2 μl of fivefold diluted template DNA (1–10 ng). Thermal cycling conditions for AOA and AOB were as follows: 5 min at 95 °C, 35 cycles of 30 s at 95 °C, 30 s at 56 °C, 1 min at 72 °C, followed by 72 °C for 10 min. The PCR products were purified using the Wizard SV Gel and PCR Clean-Up System (Promega, San Luis Obispo, CA, USA). The concentrations of PCR products were fluorometrically quantified using the Qubit dsDNA HS Assay Kit (Invitrogen, Carlsbad, CA, USA) per manufacturer’s instructions.
Restriction digestions were performed in a 10 μl mixture containing ~200 ng purified PCR products. We used the restriction enzymes HpyCH4V for AOA and MspI (BioLabs, Sydney, NSW, Australia) for AOB. Digests were incubated at 37 °C for 3 h, followed by 95 °C for 10 min to deactivate the restriction enzyme. Terminal-restriction fragments (T-RFs) were resolved on an ABI PRISM 3500 genetic analyzer (Applied Biosystems). The size and quantity of T-RFs were analyzed using the GeneMapper software version 5 (Applied Biosystems) with the advanced peak detection algorithm. Raw T-RFLP data were analyzed by using T-REX software [46]. We calculated the richness (number of TRFs) of AOA and AOB from these analyses. TRF data generated from the whole-community finger-print method were used to quantify the relative abundance of AOA and AOB taxa in our study. The T-RFLP technique is subject to the caveats of PCR-based techniques (e.g., PCR amplification biases, formation of PCR artifacts such as chimeric sequences). In addition, a single TRF could represent multiple phylogenetically related organisms and may not represent a true operational taxonomic unit; the T-RFLP fingerprinting technique is unable to detect populations in low abundance and T-RFs reflect the most abundant species. Even so, recent studies provided evidence that TRFLP and next generation sequencing (including 454 pyrosequencing and MiSeq) provide similar results in terms of richness estimation [31, 47]. Moreover, this technique is especially efficient for determining the diversity of specialized microbial groups using functional genes wherein the diversity is low, and the groups represent only a minor fraction of the overall microbial community [44, 48, 49].
Climate and human influence index
Aridity (1-aridity index) was estimated using the Global Aridity Index dataset (http://www.cgiar-csi.org/data/global-aridity-and-pet-database; [50]). We also obtained information on mean annual temperature (MAT) and rainfall and temperature seasonality (within-annual temperature variability; standard deviation×100) for all our sites from the WorldClim database (http://www.worldclim.org; [51]). Information on the Human Influence Index ([52] for each site was obtained. This index is based on eight measures of human presence: population density (km−2), distance to railroads, distance to major roads, distance to navigable rivers, distance to coastlines, distance to nighttime stable lights, and land use (urban areas, irrigated agriculture, rain-fed agriculture, and other cover types including forests, tundra, and deserts). Similar indexes have been successfully used in the past to evaluate the role of human impacts on single ecosystem functions, including N cycling, at the global scale e.g., [45, 53]) More importantly, this index is a good predictor of N deposition and N fertilization due to human activities, which are well-known to alter AOA and AOB communities [31] (Fig. S1).
Soil properties and N cycling functions
Total organic carbon (soil C), total N, Olsen P (soil P), C:N ratio and pH were measured in all soil samples, as explained in ref. [39]. In brief, pH was measured with a pH meter in a 1:2.5 mass: volume soil and water suspension. Total N was obtained using a CN analyzer (LECO CHN628 Series, LECO Corporation, St Joseph, MI, USA). The concentration of total organic C was determined as described in ref. [54]. Olsen P was measured using colorimetric analyses as explained in [39]. The concentration of dissolved organic N, ammonium –the main N source for AOA and AOB– and nitrate was measured as explained in ref. [55]. Electrical conductivity (salinity) was analyzed as explained in ref. [56]. Soil C ranged from 0.05 to 3.98% (bare soil) and 0.15 to 4.43 (vegetated). Electrical conductivity (salinity) ranged from 29.48 to 1356.32 µS/cm (bare soil) and 38.07–537.95 µS/cm (vegetated). Soil pH ranged from 6.12 to 8.98 (bare soil) and 6.33 to 8.72 (vegetated). Soil C:N ranged from 1.11 to 22.57 (bare soil) and 4.01–25.09 (vegetated).
Statistical analyses
We used one-way ANOVA to explore the differences between samples collected under plant vs. bare soil in community diversity (richness evaluated as the number of T-RFs) and abundance (as obtained with qPCR), and in soil properties and nutrients. PERMANOVA analyses were conducted to test for significant differences between vegetation and open areas (bare soil) for the community composition of AOA and AOB. Latitude and sine and cosine of longitude were included in these analyses as covariates to account for the spatial autocorrelation of our data. Non-metric multidimensional scaling (NMDS) analyses were used to identify the changes in the community composition of AOA and AOB taxa, and their association with vegetation cf. bare soil. We used the Euclidean distance matrix for these analyses. These analyses were carried out using PRIMER v 6113 and PERMANOVA+ (PRIMER-E, Plymouth, UK) [57].
We employed non-parametric partial Spearman rank correlations to explore the potential associations between vegetation (0 = bare soil and 1 = vegetation), N availability (DON, ammonium and nitrate), soil properties, climate and Human Influence Index with AOA and AOB richness and abundance after controlling for spatial influence (latitude and sine and cosine of longitude). We conducted further partial correlations to identify potential associations between AOA and AOB attributes. By using partial Spearman correlations, we aimed to identify the most important trends in our results while controlling for spatial autocorrelation. Spearman rank correlations measure the strength and direction of association between two ranked variables. Moreover, these correlations do not require normality of data, and linearity is not strictly an assumption (they can be run on a non-monotonic relationship to determine if there is a monotonic component to the association, and thus, to identify the most important trends between two variables). Moreover, unlike Pearson correlations, Spearman ranks can be used to associate two variables that are either ordinal, interval or ratio, and is robust to any effects from out-layers.
Results
Plant canopies regulate the diversity and abundance of AOA and AOB
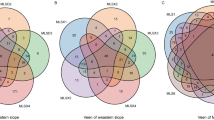
Our NMDS analysis revealed strong differences in the composition of AOA and AOB communities between samples collected under vegetation and bare soil areas (Fig. 2a, b). Taxonomic information for each T-RF is available in Table S2. The dominant AOB TRFs (60, 61, and 92) were reported previously to be related to cluster 3 of Nitrosospira belonging to β-proteobacteria (Table S2). The dominant AOA T-RFs 98/294 and 54 were related to Nitrososphaera and Nitrosotalea, respectively (Table S2).
AOA and AOB composition, richness and abundance in bare soil and vegetated areas. a and b show a 2-D nonmetric multidimensional scaling (NMDS) ordination of AOA a and AOB b. c and d show the abundance (log10 transformed) and richness of AOA c and AOB d in bare ground and vegetated areas (mean ± SE). P values as follows: ***P < 0.001; **P < 0.01; *P < 0.05 (n = 80). Abundance = gene copies g−1 soil (log10 transformed). Richness = number of phylotypes
We found that the abundance and richness of AOA were lower in vegetated patches than in bare soils (Fig. 2c). Conversely, the abundance and richness of AOB were significantly higher in vegetated vs. bare soil areas (Fig. 2d; See also Appendix S1 and Fig. S2 for a complementary approach). Vegetated areas showed as much as 60 times (3.7 folds on average; untransformed data) more abundance of AOB genes than bare soils within a given location. Moreover, bare soils had up to 91 times (6.5 folds on average; untransformed data) more abundance of AOA genes than vegetation within a given location. In general, similar results were found for the abundance and richness of AOA and AOB in locations dominated by grasses and shrubs (Figs. S3 and S4). More than 91% (73 out of 80) of the ecosystems studied had this type of vegetation.
Environmental factors regulating the richness and abundance of AOA and AOB communities
Plant canopies showed higher contents of soil C, ammonium and dissolved organic N (precursor of soil ammonium), and a higher soil C:N ratio, than bare soils (Fig. 3; P < 0.05). Of special interest, ammonium ranged from 0.1 to 21.12 mgNkg−1 soil across all samples. We did not find any significant differences between plant canopies and bare soil for nitrate and inorganic P content (Olsen P, soil P), electrical conductivity or soil pH (P > 0.05). Partial correlations further supported the positive associations between vegetation (plant canopies = 1 vs. bare soil = 0) and AOB richness and abundance (Table 1). Moreover, we also detected negative associations between vegetation and the abundance and richness of AOA communities (Table 1). Positive associations between vegetation and AOB can be, at least partially, explained by positive correlations between ammonium, organic matter (soil C) and dissolved organic N with the abundance and richness of AOB communities (Fig. 3; Table 1). The lower abundance of AOA under vegetation could be explained by the negative association between the amount of DON in soil (precursor of soil ammonium) and the abundance of AOA (Table 1). Moreover, and after controlling for spatial influence, negative associations were found between AOA and AOB richness (Table 1), and between AOA abundance and AOB richness (Table 1).
We also found significant associations between AOA and AOB communities with other environmental factors. For example, soil pH and mean annual temperature were negatively associated with the abundance and richness of AOA (Table 1), while temperature seasonality was positively associated with both AOA attributes (Table 1). On the contrary, mean annual temperature and temperature seasonality were positively and negatively associated with the abundance of AOB, respectively (Table 1). The Human Influence Index was positively associated with the abundance of AOB (Table 1).
Discussion
Our results provide novel evidence for niche differentiation between AOA and AOB communities in global drylands, driven by the presence of perennial plants, which is consistent with ecological strategy theories associated with the copiotroph–oligotroph dichotomy. As hypothesized, vegetation promoted nitrifying bacteria at the expense of nitrifying archaea. In particular, we found that the diversity and abundance of AOB were promoted under plant canopies, while amoA gene abundance and richness of oligotrophic AOA was enhanced in open areas devoid of perennial vegetation. These results were confirmed regardless of the methodological approach (qPCR and T-RFLP; see supplementary analyses in Appendix S1; Fig. S2) or vegetation type (grasses and shrubs; Figs. S3 and S4) considered, and were maintained after accounting for spatial influence. Moreover, the NMDS ordination of the AOA/AOB communities shows a clear separation between samples coming from bare soil areas and plant canopies. Our findings indicate that small-scale heterogeneity induced by plants, rather than large-scale changes in environmental conditions, largely regulate the diversity, abundance and co-occurrence network (Fig. S2) of nitrifying bacterial and archaeal communities in global drylands.
The clear differentiation in AOB and AOA communities observed between plant canopies and open areas might be, at least partially, explained by the influence of plant canopies on nutrient availability. Thus, higher and lower total abundance and richness of AOB and AOA under plant canopies (vs. bare soils) can be, at least partially, explained by their positive effect on organic matter (soil C), dissolved organic N (precursor of ammonium in soil) and ammonium contents (Fig. 3; Table 1; [27, 58, 59]. Also, plants can indirectly increase the amount of ammonium in the soil via increases in organic matter (ρ = 0.68; P < 0.001; n = 160), via litter and root inputs, and dissolved organic N (ρ = 0.24; P = 0.002; n = 160), which can release ammonium once mineralized. Thus, by increasing the amount of soil ammonium and dissolved organic N, plants could, potentially, promote the abundance and richness of AOB, and reduce the abundance of AOA (which is, in turn, positively correlated with the richness of AOA communities; AOA abundance ↔ AOA richness: ρ = 0.20; P = 0.010; Table 1). We also found that potential nitrification rates were higher in vegetated vs. bare soil areas (Fig. S5), and identified a significant correlation between nitrification rates and the abundance of AOB across all studied soils (ρ = 0.22; P = 0.005; n = 160). This result accords with a previous study that found a higher total abundance of AOB in rhizosphere vs. bulk soils of slow-growing plants with traits like low specific leaf area, low N content and high dry matter content, which typically dominate mesic environments [25]. Moreover, our results agree with laboratory and microcosm studies suggesting that AOA are outcompeted by AOB under high levels of ammonium availability [14, 15]. The findings presented here provide a much needed ecological context and field confirmation for the ammonium niche segregation of AOA and AOB suggested by these studies.
Despite the discussed role of ammonium as a potential driver of AOA and AOB communities, we would like to highlight that other aspects of soils (biogeochemical, but also moisture) associated with vegetation such as shading and microclimatic conditions, might also help explain the predominant role of vegetation in regulating the abundance of AOA and AOB abundance and richness. Thus, environmental factors other than nutrient enrichment under plant canopies could also influence the results observed. These include potential mutual exclusions between AOA and AOB communities (Table 1 and S3), improvement of micro-climatic conditions under plant canopies and to small-scale differences in other key but hard to measure (in the field and at the global scale) environmental factors such as rhizodeposition of labile C, litter decomposition and nitrification rates in situ. Plants can also potentially influence the abundance and richness of ammonia-oxidizing microbes by altering the micro-climatic conditions under their canopies, as vegetated areas are characterized by lower soil temperatures and higher soil moisture content compared to bare soils [58, 59]. In fact, mean annual temperatures and temperature seasonality showed contrasting correlations with AOA and AOB communities (Table 1). Thus, the greater abundance and richness of AOA in bare ground as compared to vegetated areas could be, at least in part, be attributed to their intrinsic physiological adaptation to the more extreme conditions (e.g., temperature seasonality) found in these areas (vs. plant canopies) in global drylands [60,61,62]). Of course, other environmental factors also influenced AOA and AOB communities. For example, soil properties, and pH in particular, negatively influenced the abundance and richness of AOA in global drylands (Fig. 3). Moreover, human influence showed positive associations with the abundance of AOB. Human activities are known to promote N deposition. For example, ammonium content was positively correlated with the Human Influence Index in the sites studied (ρ = 0.52; P < 0.001; see also Fig. S1), which might help explain the observed positive associations between this index and the abundance of AOB.
Further work, based on global monitoring efforts, should aim to evaluate the role of seasonality and peak of water availability on amoA communities and nitrification processes, as it is possible that AMO communities respond to seasonality and peaks of water availability in the global drylands where water is the main limiting factor. Moreover, we would like to stress that, because of the logistics involved in conducting a global survey such as that used here, samples in this study were collected over multiple years. Thus, whenever possible, future global research efforts aiming to study AOA and AOB communities should try to collect samples using shorter timeframes. Nonetheless, our work provides strong evidence of niche differentiation between AOA and AOB, and we would like to note that we do not expect major changes in our results for two major reasons. First, spatial and soil environmental influence have recently been highlighted to be far more important than seasonality in controlling soil microbial communities [40]. Second, the fact that we used a DNA-based rather than a RNA-based technique should provide a more complete picture about the abundance of both dormant and active organisms from soil fresh and relic DNA [63]. Thus, we believe that our results capture what happen in those soils along the year, rather than in a given moment of time.
In summary, our results support the hypothesis of a plant-driven niche differentiation for AOA and AOB communities in global drylands. We demonstrated that soils under plant canopies harbor a higher abundance, richness and dominant taxa of AOB in global drylands, at the expense of AOA communities. Plants indirectly influence the total abundance of AOA and AOB by increasing the amount of ammonium, dissolved organic N and soil C underneath their canopies. Taken together, our findings provide novel insights on the role of niche differentiation in driving the distribution of AOA and AOB in the field at the global scale, providing an ecological context to the experimental-based evidences for a central role of N availability in regulating AOA and AOB communities in terrestrial ecosystems.
Data accessibility
All the materials, raw data, and protocols used in the article are available upon request and without restriction, and all data will be made publicly available in a public repository (Figshare) upon publication.
References
Prosser JI. Autotrophic Nitrification in Bacteria. In: Advances in Microbial Physiology. 1990;30:125–81.
Hatzenpichler R. Diversity, physiology, and niche differentiation of ammonia-oxidizing archaea. Appl Environ Microbiol. 2012;78:7501–10.
Kowalchuk GA, Stephen JR. Ammonia-oxidizing bacteria: a model for molecular microbial ecology. Annu Rev Microbiol. 2001;55:485–529.
Hawkes CV, Wren IF, Herman DJ, Firestone MK. Plant invasion alters nitrogen cycling by modifying the soil nitrifying community. Ecol Lett. 2005;8:976–85.
Gruber N, Galloway JN. An Earth-system perspective of the global nitrogen cycle. Nature. 2008;451:293–6.
LeBauer DS, Treseder KK. Nitrogen limitation of net primary productivity in terrestrial ecosystems is globally distributed. Ecology. 2008;89:371–9.
Parker SS, Schimel JP. Soil nitrogen availability and transformations differ between the summer and the growing season in a California grassland. Appl Soil Ecol. 2011;48:185–92.
Schlesinger W, Bernhardt E. Biogeochemistry: an analysis of global change. 2013;3:688.
Yang WH, Ryals RA, Cusack DF, Silver WL. Cross-biome assessment of gross soil nitrogen cycling in California ecosystems. Soil Biol Biochem. 2017;107:144–55.
Venter JC, Remington K, Heidelberg JF, Halpern AL, Rusch D, Eisen JA, et al. Environmental genome shotgun sequencing of the Sargasso Sea. Science. 2004;304:66–74.
Könneke M, Bernhard AE, de la Torre JR, Walker CB, Waterbury JB, Stahl DA. Isolation of an autotrophic ammonia-oxidizing marine archaeon. Nature. 2005;437:543.
Treusch AH, Leininger S, Kletzin A, Schuster SC, Klenk HP, Schleper C. Novel genes for nitrite reductase and Amo-related proteins indicate a role of uncultivated mesophilic crenarchaeota in nitrogen cycling. Environ Microbiol. 2005;7:1985–95.
Maire V, Gross N, Börger L, Proulx R, Wirth C, Pontes LDS, Soussana JF, Louault F. Habitat filtering and niche differentiation jointly explain species relative abundance within grassland communities along fertility and disturbance gradients. New Phytol. 2012;196:497–509.
Prosser JI, Nicol GW. Archaeal and bacterial ammonia-oxidisers in soil: The quest for niche specialisation and differentiation. Trends Microbiol. 2012;20:523–31.
Verhamme DT, Prosser JI, Nicol GW. Ammonia concentration determines differential growth of ammonia-oxidising archaea and bacteria in soil microcosms. ISME J. 2011;5:1067–71.
Hink L, Gubry-Rangin C, Nicol GW, Prosser JI. The consequences of niche and physiological differentiation of archaeal and bacterial ammonia oxidisers for nitrous oxide emissions. ISME J. 2018;12:1084–93.
Di HJ, Cameron KC, Shen CS, Winefield CS, O'Callaghan M, Bowatte S, He JZ. Nitrification driven by bacteria and not archaea in nitrogen-rich grassland soils. Nat Geosci. 2009;2:621–4.
He JZ, Hu HW, Zhang LM. Current insights into the autotrophic thaumarchaeal ammonia oxidation in acidic soils. Soil Biol Biochem. 2012;55:146–54.
Nazaries L, Karunaratne SB, Delgado-Baquerizo M, Campbell CD, Singh BK. Environmental drivers of the geographical distribution of methanotrophs: Insights from a national survey. Soil Biol Biochem. 2018;127:264–79.
Reich PB. The world-wide ‘fast-slow’ plant economics spectrum: A traits manifesto. J Ecol. 2014;102:275–301.
Poorter L, Bongers L, Bongers F. Architecture of 54 moist‐forest tree species: traits, trade‐offs, and functional groups. Ecology. 2006;87:1289–301.
Sendall KM, Lusk CH, Reich PB. Trade-offs in juvenile growth potential vs. shade tolerance among subtropical rain forest trees on soils of contrasting fertility. Funct Ecol. 2016;30:845–55.
Sterck FJ, Poorter L, Schieving F. Leaf traits are good predictors of plant performance across 53 rain forest species. Ecology. 2006;87:1733–43.
Moreau D, Pivato B, Bru D, Busset H, Deau F, Faivre C, et al. Plant traits related to nitrogen uptake influence plant‐microbe competition. Ecology. 2015;96:2300–10.
Thion CE, Poirel JD, Cornulier T, De Vries FT, Bardgett RD, Prosser JI. Plant nitrogen-use strategy as a driver of rhizosphere archaeal and bacterial ammonia oxidiser abundance. FEMS Microbiol Ecol. 2016;92:fiw091.
Delgado-Baquerizo M, Morillas L, Maestre FT, Gallardo A. Biocrusts control the nitrogen dynamics and microbial functional diversity of semi-arid soils in response to nutrient additions. Plant Soil. 2013;372:643–54.
Ochoa-Hueso R, Eldridge DJ, Delgado-Baquerizo M, Soliveres S, Bowker MA, Gross N, et al. Soil fungal abundance and plant functional traits drive fertile island formation in global drylands. J Ecol. 2018;106:242–53.
Pravalie R. Drylands extent and environmental issues. A global approach. Earth-Science Rev. 2016;161:259–78.
Reynolds JF, Smith DMS, Lambin EF, Turner BL II, Mortimore M, et al. Global desertification: building a science for dryland development. Science. 2007;316:847–51.
Valentin C, D’Herbès JM, Poesen J. Soil and water components of banded vegetation patterns. Catena. 1999;37:1–24.
Delgado‐Baquerizo M, Maestre F. Human impacts and aridity differentially alter soil N availability in drylands worldwide. Glob Ecol. 2016;25:36-45.
Gubry-Rangin C, Hai B, Quince C, Engel M, Thomson BC, James P, et al. Niche specialization of terrestrial archaeal ammonia oxidizers. Proc Natl Acad Sci USA. 2011;108:21206–11.
Hu HW, Zhang LM, Dai Y, Di HJ, He JZ. pH-dependent distribution of soil ammonia oxidizers across a large geographical scale as revealed by high-throughput pyrosequencing. J Soils Sediments. 2013;13:1439–49.
Oton EV, Quince C, Nicol GW, Prosser JI, Gubry-Rangin C. Phylogenetic congruence and ecological coherence in terrestrial Thaumarchaeota. ISME J. 2016;10:85–96.
Hu HW, Chen D, He JZ. Microbial regulation of terrestrial nitrous oxide formation: understanding the biological pathways for prediction of emission rates. FEMS Microbiol Rev. 2015;39:729–49.
Bernhard AE, Landry ZC, Blevins A, José R, Giblin AE, Stahl DA. Abundance of ammonia-oxidizing archaea and bacteria along an estuarine salinity gradient in relation to potential nitrification rates. Appl Environ Microbiol. 2010;76:1285–9.
Bello MO, Thion C, Gubry-Rangin C, Prosser JI. Differential sensitivity of ammonia oxidising archaea and bacteria to matric and osmotic potential. Soil Biol Biochem. 2019;129:184–90.
Valencia E, Maestre FT, Le Bagousse-Pinguet Y, Quero JL, Tamme R, Börger L, et al. Functional diversity enhances the resistance of ecosystem multifunctionality to aridity in Mediterranean drylands. New Phytol. 2015;206:660–71.
Maestre FT, Delgado-Baquerizo M, Jeffries TC, Eldridge DJ, Ochoa V, Gozalo B, et al. Increasing aridity reduces soil microbial diversity and abundance in global drylands. Proc Natl Acad Sci USA. 2015;112:15684–9.
Carini P, Delgado-Baquerizo M, Hinckley EL, Brewer TE, Rue G, Vanderburgh C, et al. Unraveling the effects of spatial variability and relic DNA on the temporal dynamics of soil microbial communities. BioRxiv. 2018;402438.
Delgado-Baquerizo M, Covelo F, Maestre FT, Gallardo A. Biological soil crusts affect small-scale spatial patterns of inorganic N in a semiarid Mediterranean grassland. J Arid Environ. 2013;91:147–50.
Tourna M, Freitag TE, Nicol GW, Prosser JI. Growth, activity and temperature responses of ammonia-oxidizing archaea and bacteria in soil microcosms. Environ Microbiol. 2008;10:1357–64.
Rotthauwe JH, Witzel KP, Liesack W. The ammonia monooxygenase structural gene amoA as a functional marker: molecular fine-scale analysis of natural ammonia-oxidizing populations. Appl Environ Microbiol. 1997;63:4702–12.
Hu HW, Macdonald CA, Trivedi P, Holmes B, Bodrossy L, He JZ, Singh BK. Water addition regulates the metabolic activity of ammonia oxidizers responding to environmental perturbations in dry subhumid ecosystems. Environ Microbiol. 2015;17:444–61.
Delgado-Baquerizo M, Maestre FT, Eldridge DJ, Singh BK. Microsite differentiation drives the abundance of soil ammonia oxidizing bacteria along aridity gradients. Front Microbiol. 2016;7:505.
Culman SW, Bukowski R, Gauch HG, Cadillo-Quiroz H, Buckley DH. T-REX: software for the processing and analysis of T-RFLP data. BMC Bioinformatics. 2009;10:171.
van Dorst J, Bissett A, Palmer AS, Brown M, Snape I, Stark JS, et al. Community fingerprinting in a sequencing world. FEMS Microbiol Ecol. 2014;89:316–30.
Stralis-Pavese N, Sessitsch A, Weilharter A, Reichenauer T, Riesing J, Csontos J, et al. Optimization of diagnostic microarray for application in analysing landfill methanotroph communities under different plant covers. Environ Microbiol. 2004;6:347–63.
Singh BK, Nazaries L, Munro S, Anderson IC, Campbell CD. Use of multiplex terminal restriction fragment length polymorphism for rapid and simultaneous analysis of different components of the soil microbial community. Appl Environ Microbiol. 2006;72:7278–85.
Zomer RJ, Trabucco A, Bossio DA, Verchot LV. Climate change mitigation: A spatial analysis of global land suitability for clean development mechanism afforestation and reforestation. Agric Ecosyst Environ. 2008;126:67–8.
Hijmans RJ, Cameron SE, Parra JL, Jones PG, Jarvis A. Very high resolution interpolated climate surfaces for global land areas. Int J Climatol. 2005;7568:201.
Sanderson EW, Jaiteh M, Levy MA, Redford KH, Wannebo AV, Woolmer G. The Human Footprint and the Last of the Wild: The human footprint is a global map of human influence on the land surface, which suggests that human beings are stewards of nature, whether we like it or not. Bioscience. 2002;52:891–904.
Crowther TW, Glick HB, Covey KR, Bettigole C, Bettigole C, Maynard DS, et al. Mapping tree density at a global scale. Nature. 2015;525:201–5.
Anderson JM, Ingram JSI. Tropical Soil Biology and Fertility: A Handbook of Methods. Soil Sci. 1994;157:265.
Delgado-Baquerizo M, Gallardo A. Depolymerization and mineralization rates at 12 Mediterranean sites with varying soil N availability. A test for the Schimel and Bennett model. Soil Biol Biochem. 2011;43:693–6.
Maestre FT, Quero JL, Gotelli NJ, Escudero A, Ochoa V, Delgado-Baquerizo M, et al. Plant species richness and ecosystem multifunctionality in global drylands. Science. 2012;335:279–91.
Clarke KR, Gorley RN. PRIMER V6: User Manual-tutorial. 2006.
Maestre FT, Bautista S, Cortina J, Bellot J. Potential for using facilitation by grasses to establish shrubs on a semiarid degraded steppe. Ecol Appl. 2001;11:1641–55.
Maestre FT, Bautista S, Cortina J. Positive, negative, and net effects in Grass-Shrub interaction in Mediterranean Semiarid Grassland. Ecology. 2003;84:3186–97.
Valentine DL. Opinion: Adaptations to energy stress dictate the ecology and evolution of the archaea. Nat Rev. 2007;5:316.
Bates ST, Berg-Lyons D, Caporaso JG, Walters WA, Knight R, Fierer N. Examining the global distribution of dominant archaeal populations in soil. ISME J. 2011;5:908–17.
Zhalnina K, Dörr de Quadros P, AO Camargo F, Triplett EW. Drivers of archaeal ammonia-oxidizing communities in soil. Front Microbiol. 2012;3:210.
Carini P, Marsden PJ, Leff JW, Morgan EE, Strickland MW, Fierer N. Relic DNA is abundant in soil and obscures estimates of soil microbial diversity. Nat Microbiol. 2016;2:16242.
Acknowledgements
We acknowledge the help of Victoria Ochoa and Beatriz Gozalo with laboratory analyses, and of all the colleagues that collected soil samples in global drylands. This research is supported by the Australian Research Council projects (DP170104634 and DP190103714), by the European Research Council (BIOCOM project, ERC Grant agreement n°242658) and by the Spanish Ministerio de Economía y Competitividad (BIOMOD project, ref. CGL2013-44661-R). M.D-B. acknowledges support from the Marie Sklodowska-Curie Actions of the Horizon 2020 Framework Programme H2020-MSCA-IF-2016 under REA grant agreement n°702057. FTM acknowledges support from the European Research Council (BIODESERT project, ERC Grant agreement n°647038).
Author information
Authors and Affiliations
Contributions
Statement of authorship: M.D.-B. conceived the idea of this study. F.T.M. designed and coordinated field surveys. Laboratory analyses were done by C.T. and F.T.M. in consultation with B.K.S. and M.D.B. Data analyses were done by C.T., B.K.S. and H.-W.H; interpretation was done by all authors. Statistical modeling was done by M.D.B. The manuscript was written by M.D.-B. and C.T., edited by P.B.R., B.K.S. and F.T.M., and all authors contributed to the final draft.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Trivedi, C., Reich, P.B., Maestre, F.T. et al. Plant-driven niche differentiation of ammonia-oxidizing bacteria and archaea in global drylands. ISME J 13, 2727–2736 (2019). https://doi.org/10.1038/s41396-019-0465-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-019-0465-1
This article is cited by
-
Nitrification inhibitor 1-octyne inhibits growth of comammox Nitrospira but does not alter their community structure in an acidic soil
Journal of Soils and Sediments (2023)
-
Plant Species–Driven Distribution of Individual Clades of Comammox Nitrospira in a Subtropical Estuarine Wetland
Microbial Ecology (2023)
-
Physicochemical and Biotic Changes and the Phylogenetic Evenness of Microbial Community in Soil Subjected to Phytoreclamation
Microbial Ecology (2022)
-
Soil bacterial depth distribution controlled by soil orders and soil forms
Soil Ecology Letters (2022)
-
Distribution characteristics of ammonia-oxidizing microorganisms and their responses to external nitrogen and carbon in sediments of a freshwater reservoir, China
Aquatic Ecology (2022)