Abstract
Correlation analyses are often included in bioinformatic pipelines as methods for inferring taxon–taxon interactions. In this perspective, we highlight the pitfalls of inferring interactions from covariance and suggest methods, study design considerations, and additional data types for improving high-throughput interaction inferences. We conclude that correlation, even when augmented by other data types, almost never provides reliable information on direct biotic interactions in real-world ecosystems. These bioinformatically inferred associations are useful for reducing the number of potential hypotheses that we might test, but will never preclude the necessity for experimental validation.
Similar content being viewed by others
Introduction
Correlation (i.e., normalized covariance), the measure of statistical dependence between two variables, can be a useful summary of the associations between features across a dataset. Often, correlation refers to the linear relationship between two random variables, which can be captured by Pearson’s correlation coefficient, or nonparametric measures of dependence, like Spearman’s ρ, Kendall’s τ, or mutual information. The degree of dependence between variables can indicate a predictive relationship that can be exploited, whether or not these variables are causally related to one another. Overall, correlation is a useful statistical tool for identifying apparent interdependencies among many variables. Many researchers, implicitly or explicitly, use correlation structure in microbial community datasets to infer underlying ecological interactions. In general, these inferences are fraught with challenges.
While useful, correlation-based approaches are inherently limited when it comes to ecological interaction inference. Complex nonlinear dynamics, compositionality of sequencing data, environmental heterogeneity, latent confounders, indirect associations, and batch effects all hinder the usefulness of these correlation metrics when inferring direct species–species associations. A variety of newer metrics and methodologies have been developed in recent years to address some of these challenges [1,2,3,4,5,6,7,8,9]. However, newer methods are far from infallible, and the underlying assumptions of these approaches need to be carefully considered when applied to data. Any method that claims to accurately capture underlying biotic interactions of a system using longitudinal or cross-sectional correlation of taxon abundances or co-occurrences should be viewed with a generous dose of skepticism.
The proliferation of correlation-based methods for inferring ecological networks is understandable. In microbial ecology, we are often limited in our ability to directly observe interactions between microbial species. The most definitive work on microbial interactions has been done experimentally. For example, microscopy and staining techniques, along with stable isotope labeling, have been employed to observe co-localization and cross-feeding between methanotrophic archaea and sulfate reducing bacteria [10]. In addition to mutualistic interactions, direct bacterial antagonism through type VI secretion systems has been demonstrated using a combination of genomics, microscopy, and co-culturing assays [11]. Entire interaction networks have been determined in simplified microbial consortia consisting of a few species, where community membership can be manipulated to assess pairwise and higher-order interactions [12, 13]. While these experimental approaches represent gold standards for inferring interactions between microorganisms, they are difficult and time consuming. Furthermore, laboratory-based studies can fail to capture the environmental context in which natural interactions occur. Recent work has demonstrated just how important this context can be in mediating interactions [14]. Thus, it is not practical to apply these experimental methods to all potential interactions between thousands of taxa, many of which cannot be cultured. As such, there is a strong incentive for identifying bioinformatic methods for interaction inference.
While interactions are difficult to observe directly, relative fluctuations in population sizes can be readily quantified for thousands of bacterial phylotypes at once. Bioinformaticians have developed a wide array of tools to infer putative associations from these high-throughput measures of relative abundance [15, 16]. In general, these methods tend to generate correlation or covariance matrices, which are often used to infer hypothetical interactions. At their best, these inferences represent tentative hypotheses that can be combined with other data types to help experimentalists guide or constrain their work. At their worst, these inferences are fundamentally flawed due to incorrect assumptions about what they tell us about biotic interactions. In this perspective, we review the application of correlation-based methods in microbial ecology, the strengths and limitations of these analyses, the pitfalls surrounding how correlation can be misused or misinterpreted, and how we might augment these analyses to improve our inferences.
Theoretical considerations
Symmetric correlations and asymmetric interactions
To begin, we must recognize the inherent symmetry of correlation metrics and the frequent asymmetry of ecological interactions. It is impossible to identify the directedness of interactions from cross-sectional associations [3, 8, 17, 18]. By incorporating the ordering of events in time and space into an analysis, it becomes somewhat possible to infer directedness [8]. However, even when the order of events is incorporated into association analyses, biological, experimental, technical, and sampling noise can greatly reduce the sensitivity and accuracy of our inferences. Prior work has demonstrated that we are much more likely to detect strong, symmetric interactions, like obligate mutualisms or direct competition, and less likely to detect weaker, directed interactions, like parasitism or amensalism [16, 19, 20].
Dynamic models and mechanistic constraints can improve inferences
In principle, when the underlying biochemical processes that mediate microbial interactions are known, mechanistic models can be developed and tested against data. When applicable, this approach provides a powerful means of predicting population dynamics and inferring interaction structure. However, a priori knowledge of interaction mechanisms is generally not available. Even when some of these mechanistic details are known, building these models is surprisingly challenging, even for simple two-species systems [21]. Thus, while desirable, this approach is not generally applicable when taxon abundances are the only information available.
Lotka-Volterra (LV) models can be fit to longitudinal data, where fluctuations in taxon abundances reflect growth and death processes, without knowing the underlying mechanisms that mediate interactions. LV models are composed of nonlinear differential equations that describe temporal changes in species abundance that result from growth, death, and interspecies interactions. These models take into account the temporal ordering of events, can capture both positive and negative interactions, and can be used to model arbitrary numbers of directed interactions between species with the assumption that interactions are additive and pairwise. When log-transformed, LV models can be fit using linear regression, making the interaction terms somewhat analogous to correlation coefficients [8]. Depending on the number of species and the parameterization, these models can have fixed steady-states, limit cycles, or more complex behaviors. LV models can provide a useful means of inferring species interactions and predicting community dynamics in some contexts but have limitations. For instance, if growth dynamics are not captured by sampling or the assumptions of the model are violated (e.g., interactions are not additive and pairwise) the application of LV is inappropriate. Furthermore, theoretical and empirical studies have shown that LV models are fundamentally incapable of accurately capturing all types of pairwise interactions and can be a poor predictor of dynamics under realistic conditions [13, 22]. Thus, while these models are useful in certain systems, like in vitro communities, their application is not always appropriate and depends on the features of the system being studied [12, 13, 20].
In the basic two-species predator–prey form of the LV model (alternatively, the parasite–host model), the prey species x is described by the equation \(\frac{{dx}}{{dt}} = \alpha x - \beta xy\) and the predator y is described by \(\frac{{dy}}{{dt}} = \delta xy - \gamma y\), where α and δ are the growth rates and β and γ are the death rates for the prey and predator species, respectively. Over a wide range of parameter values in this system we observe oscillations in both predator and prey abundance as a function of time (Fig. 1a). As the prey population grows, the predator population has more food and also increases in abundance. However, predation eventually out-paces the growth of the prey population and drives the prey toward near-extinction, until there are too few prey to sustain the predator population. Once the predator population crashes, the few remaining prey are able to recover, and the cycle begins anew. Over the course of time, predator and prey populations transition between windows of positive covariance and negative covariance (Fig. 1a). Contemporaneous correlation is not capable of identifying this asymmetric interaction between x and y inherent to the underlying model [8]. However, if we time-lag x relative to y, we find that a lag exists where the two variables are consistently positively or negatively correlated over all time windows (Fig. 1b). By observing the temporal ordering of this time-lagged relationship, we see that the crash in the prey population is preceded by a spike in the predator population, which implies a directedness consistent with y predating upon x. These types of time-lagged interactions can be formally assessed using Granger causality, which captures the degree of linear prediction of one variable (say, species y) on the future values of another variable (say, species x) and can provide directed relationships [6]. Similarly, transfer entropy is a nonparametric extension of Granger causality that can be applied to infer nonlinear, time-asymmetric associations between variables [18]. While these approaches suggest direct causal relationships, they do not guarantee them. Latent factors, like pH, temperature, or another unmeasured species, could indirectly drive similar time-lagged population dynamics. However, if known a priori, these associations can be accounted for [6]. Another popular approach for inferring directed associations is extended local similarity analysis [7]. Like transfer entropy, this method provides a useful means of capturing both temporal relationships and nonlinear associations. All of these approaches work well in addressing the weakness of contemporaneous correlations for the simple two-species predator–prey relationship. However, in the more complicated scenario of multispecies virus–microbe interactions, time-lagged association inference methods have been shown to be incapable of accurately capturing the features of these complex networks [20].
Correlation alone cannot be used to infer drivers of species dynamics. a Lotka-Volterra (LV) predator-prey oscillatory dynamics. b Time-lagged LV predator–prey dynamics, with arrows indicating the time lag used for shifting the prey dynamics backwards in time. In both a and b blue rectangular boxes are used to indicate regions in time where the dynamics show significant positive correlation (r > 0, p < 0.05) and red boxes indicate significant negative correlation (r < 0, p < 0.05). The symbols above each time window reflect the color categorization, where “+” indicates a significant positive correlation, “−” indicates a significant negative correlation and “ø” indicates an insignificant correlation. Also shown is the overall correlation across all time windows. c Hypothetical two-species community with multiple drivers of oscillatory dynamics operating at different frequencies. For each of the hypothetical species, dynamics were simulated using a linear combination two sine functions with different amplitude and frequency. Noise was added to each abundance trajectory by sampling from a normal distribution. d Spectral decomposition (i.e., Fourier transform) of abundance data in (c) and species abundance relationships for both high and low-frequency signal components
Latent drivers of dynamics confound inference of species associations
Interspecies interactions are not the only drivers of dynamics. Complex population dynamics can arise due to latent variables. In particular, environmental drivers, like changes in nutrient availability or temperature, have a profound influence on microbial population dynamics. These drivers can operate over different spatial and temporal scales. When these drivers are not taken into account they can lead to inaccurate inferences of interspecies relationships. For example, marine bacterial populations can exhibit both low-frequency oscillations (e.g., seasonal changes) and high-frequency oscillations (e.g., species–species competition or day–night cycles). Martin-Plantera et al. [23] recently applied spectral decomposition methods to marine microbial communities to isolate the different frequencies embedded within species population dynamics. They found that low-frequency oscillations grouped species together that share a similar seasonal niche, which reflected environmental filtering and likely had nothing to do with species–species interactions. Higher-frequency oscillations revealed negative correlations between related species, which may be more reflective of biotic interaction, although these dynamics could also be driven by the environment [23]. Because the low-frequency seasonal signal was much stronger than the high-frequency signal, traditional correlation analyses were dominated by seasonal effects and missed the higher-frequency signals (e.g., see simulation data presented in Fig. 1c, d). While this kind of environmental filtering can mask putative species interactions, this information is still valuable for inferring shared environmental niches within a community and, when properly accounted for, can help researchers to decouple the biotic and abiotic components of community variance [6, 23, 24].
Neutral processes can drive covariance in the absence of species interactions and environmental drivers
In some scenarios fluctuations in species abundance cannot be attributed to interspecies interactions, changes in environmental factors, or niche constraints. In these cases, observed fluctuations may simply be due to stochastic variation in community structure. Neutral models simulate changes in community structure with stochastic birth, death, migration, and speciation. Methods have been developed that allow the application of neutral models to both cross-sectional and time-series data [25, 26]. These methods, along with other types of neutral models, can provide an effective null hypothesis when trying to fit interaction models like LV or when trying to infer species associations with correlative analyses [27].
Analytical considerations
Complex structure of microbiome data
Many of the assumptions of established statistical methods are violated by microbiome sequencing datasets. Microbial community species-abundance distributions are extremely fat-tailed, with a large number of low-abundance taxa detected in very few samples. Thus, microbiome data matrices are highly sparse. Unfortunately, we do not yet understand the functional form of this rare tail of microbial diversity, which makes imputation and normalization difficult. It is hard to assess whether zeros represent true absences of species or nondetection due to sampling limitations. The presence of these zeros introduces artifacts into rank-based correlation analyses [27]. Existing approaches have not yet addressed the ambiguity of zeros in amplicon and metagenomic sequencing datasets. In the absence of a clear consensus, more conservative approaches, like injecting random low-value pseudocounts to break zero rank ties or removing low-abundance taxa, seem to be the most appropriate when calculating correlations [27, 28].
Data transformations can introduce spurious correlations
When analyzing microbiome data from high-throughput sequencing platforms, differences in library sizes across samples must be dealt with prior to analysis. These differences in library sizes are technical artifacts and do not contain biological information. The most common normalizations are total sum scaling (i.e., converting counts to proportions by dividing each species count in a sample by the total sum of counts from within that sample) and subsampling [29], which both effectively convert counts into relative measures of abundance. Relative abundances are non-Euclidean and cannot vary independently from one another. Changes in the relative abundance of one species will necessarily influence the relative abundances of the other species due to the zero-sum constraint (Fig. 2). As such, relative measures of abundance violate the assumption of independence inherent to classical statistics.
Transformation from absolute to relative abundances introduces spurious correlations, which can be mitigated by employing log-ratio transformations (e.g., SparCC). a Simulated fluctuations in absolute and relative abundance across a set of samples for a hypothetical six-species community with one positive linear association. b Hypothetical six-species community with one negative and two positive linear associations. c Hypothetical fifteen-member community with three positive and two negative linear associations. For each of these model communities positive and negative associations are illustrated with yellow and dark blue connecting lines, respectively. Mean abundances of each species were chosen arbitrarily and random fluctuations were simulated by sampling from a Poisson distribution centered around a species’ mean abundance. Species associations were simulated using linear relationships where the abundance of species Y was made a function of its own random fluctuations about a mean and an additive component that increased or decreased its abundance with respective to another species x depending on the sign of the coefficient used. Hypothetical community correlation matrices were generated using Pearson correlation with absolute and relative abundance data. Also shown is the correlation matrix inferred from relative abundance data using SparCC with its default settings. Colored borders around cells in the correlation matrices indicate associations where the p values were <0.05 and the Benjamini–Hochberg false discovery rate (FDR) q values were <0.1. Red borders indicate significant associations not present in the model community (i.e., false positives), blue borders indicate significant associations present in the model community (i.e., true positives), and yellow boarders indicate nonsignificant associations present in the model (i.e., false negatives)
The most relevant repercussion to interaction inference in compositional data is the introduction of spurious correlation structure (Fig. 2). Compositionally aware methods for analyzing relative abundances were developed by Aitchison in the 1980s, based around log-ratio transformations of compositional features. Isometric log-ratio (ILR) transforms provide the most stringent way of breaking compositionality, but they can be difficult to interpret, because they involve comparing ratios of multiple data features, rather than pairwise associations between individual features. Recent work has extended these methods to microbiome data, improving the interpretability of ILR results by taking advantage of the placement of species on a phylogenetic tree (i.e., ratios of species from one branch of the tree over species on another branch) [30]. Others methods use log-ratio transform procedures that approximate pairwise linear correlations between individual taxon relative abundances [2]. This later approach, implemented in SparCC, is a popular choice for mitigating spurious, compositionally driven correlation structure (Fig. 2c) [2]. While SparCC provides a useful approach for dealing with compositionality, as with any method, it is important to keep the assumptions it relies upon in mind to avoid potential pitfalls. When SparCC’s sparsity assumption is violated (i.e., the assumption that there are very few underlying correlations) it can yield erroneous results (Fig. 2b). Performance is also hindered when there are few pairwise comparisons with which it can estimate the underlying feature variances and pairwise associations (Fig. 2a). When the sparsity assumption is not violated and there are more than a few pairwise comparisons with which it can produce estimates, SparCC is able to accurately recapitulate much of the known correlation structure from relative abundance data (Fig. 2c). While we highlight the use of SparCC, it is worth noting that there are several other valid choices for network inference that can mitigate the issue of compositionality. For a comprehensive review of network inference tools and their performance characteristics see the following reviews [16, 31]. Simulations and empirical analyses have shown that the correlation structure in compositional data begins to converge toward what we would expect from Euclidean data as the Shannon diversity of the system increases (i.e., as the effective number of species increases) [2]. Thus, compositional effects should be relatively weak in a typical, diverse gut microbiome, but these effects can completely overwhelm the correlation structure of the vaginal microbiome, which is often dominated by a single Lactobacillus species [2]. However, even in very diverse communities, the system is often positioned near the edge of the simplex (i.e., a single species is often dominant), which ensures that many low-magnitude compositional correlations will always be present. Overall, compositional effects inherent to microbiome data must be reckoned with prior to statistical inference.
Noninformative indirect associations are introduced when taxa engage in many pairwise interactions
When associations are obtained using correlative analyses, any species that interacts with more than one additional taxon can produce indirect associations between the taxa it interacts with (e.g., see significant indirect associations in Fig. 2b, c). This is a serious issue that can turn correlation networks into hairballs of interconnected features that are challenging to interpret. Both classical correlation methods and more contemporary approaches like SparCC are susceptible to indirect associations (Fig. 2b, c). To address this issue, newer methods like SPIEC-EASI and FlashWeave have been developed [1, 32]. These methods utilize the concept of conditional independence, which assesses how informative an association between two features is given information about all other features, to reduce the number of spurious indirect relationships inferred from the data.
Inferring associations between specific microbes and environmental properties, like host phenotype, can be confounded by dense correlation networks. In recent years correlative analyses have been used to associate specific microbes in the human gut microbiome with a wide array of diseases. These microbiome wide association studies (MWAS) have produced many putative connections between human gut microbes and host phenotypes. The issue with these studies is they often produce conflicting results and the number of associations generated by any given study can be so numerous that they thwart interpretation and complicate follow-up efforts [9]. Menon et al. (2018) recently demonstrated how correlations between microbes can produce spurious indirect associations in MWAS using simulated case control data and a hypothetical interaction network. To address this issue the authors developed a method based on the maximum entropy models in statistical physics, which they call direct association analysis [9]. Like SPIEC-EASI and FlashWeave, the author’s approach utilizes conditional independence to remove uninformative, indirect associations. When applied to data from a large inflammatory bowel disease study, the author’s method was able to reduce a set of almost one hundred putative associations between various microbiota and the disease previously obtained by a conventional differential abundance analysis to a more informative set of five species and four genera, several of which were supported by mechanistic insights from other studies [9]. Whether inferring interspecies associations or species associations with environmental properties, indirect effects should be considered and accounted for to avoid reporting spurious, noninformative relationships.
Biases due to batch effects
Microbiome data are prone to batch effects (i.e., biases), arising from both biological (e.g., geographic or genetic differences between otherwise similarly defined host cohorts) and technical variation (e.g., different DNA extraction methods or 16S primers) between batches of sequencing data [33, 34]. These effects are highly complex and nonlinear, potentially making parametric batch-correction methods designed for other ‘omics data types inappropriate for microbiome data [28]. If correlation analyses are run across batches, many of the strongest associations can be attributed to biases and batch effects rather than true biological signals [28]. Recent progress has been made in developing bias and batch-correction methods [28, 34]. However, the safest course of action is to restrict statistical analyses to within a given batch and compare the results of these independent analyses across batches.
Empirical considerations
Changes in relative abundance may not reflect population growth rates
Often times, the assumption of interaction inference methods is that relative changes in species population size are indicative of population growth or decay and can be used to infer growth or death rates. On its face, this seems to be a reasonable assumption. However, in the absence of absolute abundance information, we cannot distinguish whether one population of organisms is truly increasing, or whether this rise in relative abundance is occurring due to a concomitant decrease in the population size of another species. To address this issue, researchers can take measures of absolute biomass (e.g., quantitative polymerase chain reaction or cell counts) for the samples that they sequence [35], or they can use controlled spike-ins during sequencing to break the compositionality of the data [36]. Methods for directly inferring growth rates from shotgun metagenomic data have also been developed [37].
In addition to the challenges associated with relative abundance data, temporal and spatial scales should be considered prior to any analysis. For example, temporal sampling resolution in the human gut is limited by defecation frequency (~1 bowel movement per day), which is generally too coarse to capture microbial population dynamics (i.e., bacterial doubling times of 1–10 times per day), despite the common assumption that population-dynamics models can be fit to these data [3, 8, 38]. Consequently, most of the bacterial population dynamics in the gut happen internally. Thus, fecal samples represent the endpoint of dynamics. With the exception of major perturbations that reduce standing populations in the gut by several orders of magnitude and require days to recover from (e.g., due to antibiotics or diarrhea) [3], we probably cannot infer population growth rates from human fecal 16S amplicon sequencing data. Therefore, it is important to carefully consider whether or not the spatiotemporal scale of sampling can capture relevant dynamics for any system under investigation. If interaction model assumptions are violated by the input data, then any inferences dependent upon these assumptions are suspect (Fig. 3).
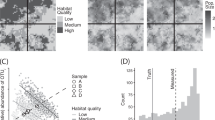
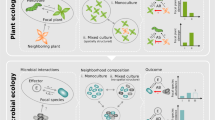
Sampling strategies should be optimized to span the appropriate spatial or temporal scales. Soils are notoriously heterogeneous environments. a Context-dependent interspecies interactions in a hypothetical soil community: blue and green species only interact during a perturbation event. b Infrequent sampling appropriately captures correlations from slower recovery process. c Infrequent sampling does a poor job of capturing correlation structure from a rapid recovery process
Environmental heterogeneity is usually the strongest driver of correlation structure in natural environments
In soils, drastic shifts in pH, carbon availability, and water content can occur over microns-to-centimeter scales. If environmental conditions vary over the spatial or temporal scales that are sampled, the organisms—often phylogenetically related—that are adapted to these conditions vary along with them [3, 23]. Cofluctuation of taxa due to variation in niche space is known as habitat filtering, and can be useful information about the niche requirements of species in an ecosystem. However, habitat filtering provides us with little-to-no information about direct species–species interactions. Habitat filtering is usually the dominant driver of correlation structure in natural microbial ecosystems and should be carefully considered when attempting to identify direct species–species interactions from ‘omics data.
Berry and Widder [39] showed that correlation networks generated from multispecies LV models only reflected true interactions under a narrow range of conditions, and that any amount of interaction complexity or environmental heterogeneity made correlation a poor predictor of interaction. Concordantly, recent empirical work from an intertidal ecosystem demonstrated that co-occurrence analyses were unable to recapitulate most known interactions in their system, with the exception of certain strong mutualistic or antagonistic interactions [19]. The deconvolution of direct species–species interactions from habitat filtering due to environmental heterogeneity is one of the most intractable challenges facing bioinformatic interaction inference in real-world ecosystems. Thus, researchers should be extremely skeptical and avoid explicit or implicit assumptions of species–species associations when applying the myriad methods that have been developed to infer putative “interactions”, “connectivity”, or “cohesion” from covariance structure in real-world systems [1,2,3,4, 8, 40].
Conclusion
We provide a few illustrative examples of the challenges associated with interpreting correlation networks in microbial ecology and highlight several methods that have been developed to address these challenges. For a more in depth discussion of the latest network inference methods, please see recent comprehensive reviews on the topic [15, 16, 31]. In this perspective, we focus on our various concerns regarding the use of correlation to infer biotic interactions. While correlation analyses are extremely useful for processing and digesting ‘omics data, they can also lead us astray in several important ways. We discuss how correlation metrics are inherently symmetric and cannot be used to identify asymmetric interactions without including additional information. We demonstrate how various types of community dynamics and interaction structures are fundamentally opaque to correlation analyses and how use of models that incorporate temporal and mechanistic details can aid inference of meaningful associations. We reveal how data transformations and analysis techniques can warp data and introduce spurious correlation structure that does not reflect the underlying biology and we introduce several methodological strategies to mitigate these issues. We note that indirect associations can be produced by environmental factors or taxa engaging in multiple interactions and present methods for addressing these latent confounders. Finally, we discuss how real-world ecosystems and the data we use to investigate them are messy and complex, and how this heterogeneity can confound our ability to infer species-species interactions. Even the simplest cases of interaction inference from correlations can fall apart. More often than not, the presence or absence of a correlation between variables tells the researcher almost nothing about biotic interactions.
Integrating other types of data into correlation analyses, like measures of potentially confounding environmental variables, accurate noise and bias estimates, absolute biomass, the ordering of events in space or time, multi-omic measurements, and mechanistic constraints can greatly improve our inferences. Perturbation experiments, which dislodge an environmental system from its steady state, can be used to generate more informative correlation structure [41, 42]. The use of mesocosms or microcosms helps to overcome the confounding influences of environmental heterogeneity and higher-order species interactions. However, even in these simplified systems, researchers should be supremely skeptical of inferred interactions. In the end, bioinformatic approaches only generate hypotheses. In order for these inferred interactions to be accepted as truth, the hard work of experimental validation is required.
References
Kurtz ZD, Müller CL, Miraldi ER, Littman DR, Blaser MJ, Bonneau RA. Sparse and compositionally robust inference of microbial ecological networks. PLoS Comput Biol. 2015;11:e1004226.
Friedman J, Alm EJ. Inferring correlation networks from genomic survey data. PLoS Comput Biol. 2012;8:e1002687.
Gibbons SM, Kearney SM, Smillie CS, Alm EJ. Two dynamic regimes in the human gut microbiome. PLoS Comput Biol. 2017;13:e1005364.
Herren CM, McMahon KD. Cohesion: a method for quantifying the connectivity of microbial communities. ISME J. 2017;11:2426–38.
Schmidt TSB, Rodrigues JFM, von Mering C. A family of interaction-adjusted indices of community similarity. ISME J. 2017;11:791–807.
Ai D, Li X, Liu G, Liang X, Xia LC. Constructing the Microbial Association Network from large-scale time series data using Granger causality. Genes. 2019;10:E216.
Xia LC, Steele JA, Cram JA, Cardon ZG, Simmons SL, Vallino JJ, et al. Extended local similarity analysis (eLSA) of microbial community and other time series data with replicates. BMC Syst Biol. 2011;5(Suppl 2):S15.
Fisher CK, Mehta P. Identifying keystone species in the human gut microbiome from metagenomic timeseries using sparse linear regression. PLoS ONE. 2014;9:e102451.
Menon R, Ramanan V, Korolev KS. Interactions between species introduce spurious associations in microbiome studies. PLoS Comput Biol. 2018;14:e1005939.
Orphan VJ, Turk KA, Green AM, House CH. Patterns of 15N assimilation and growth of methanotrophic ANME-2 archaea and sulfate-reducing bacteria within structured syntrophic consortia revealed by FISH-SIMS. Environ Microbiol. 2009;11:1777–91.
Russell AB, Wexler AG, Harding BN, Whitney JC, Bohn AJ, Goo YA, et al. A type VI secretion-related pathway in Bacteroidetes mediates interbacterial antagonism. Cell Host Microbe. 2014;16:227–36.
Venturelli OS, Carr AC, Fisher G, Hsu RH, Lau R, Bowen BP, et al. Deciphering microbial interactions in synthetic human gut microbiome communities. Mol Syst Biol. 2018;14:e8157.
Friedman J, Higgins LM, Gore J. Community structure follows simple assembly rules in microbial microcosms. Nat Ecol Evol. 2017;1:109.
Liu A, Archer AM, Biggs MB, Papin JA. Growth-altering microbial interactions are responsive to chemical context. PLoS ONE. 2017;12:e0164919.
Faust K, Raes J. Microbial interactions: from networks to models. Nat Rev Microbiol. 2012;10:538–50.
Weiss S, Van Treuren W, Lozupone C, Faust K, Friedman J, Deng Y, et al. Correlation detection strategies in microbial data sets vary widely in sensitivity and precision. ISME J. 2016;10:1669–81.
Trosvik P, de Muinck EJ, Stenseth NC. Biotic interactions and temporal dynamics of the human gastrointestinal microbiota. ISME J. 2015;9:533–41.
Barnett L, Barrett AB, Seth AK. Granger causality and transfer entropy are equivalent for Gaussian variables. Phys Rev Lett. 2009;103:238701.
Freilich MA, Wieters E, Broitman BR, Marquet PA, Navarrete SA. Species co-occurrence networks: can they reveal trophic and non-trophic interactions in ecological communities? Ecology. 2018;99:690–9.
Coenen AR, Weitz JS. Limitations of correlation-based inference in complex virus-microbe communities. mSystems. 2018;3:e00084–18.
Hart SFM, Mi H, Green R, Xie L, Pineda JMB, Momeni B, et al. Uncovering and resolving challenges of quantitative modeling in a simplified community of interacting cells. PLOS Biol. 2019;17:e3000135.
Momeni B, Xie L, Shou W. Lotka-Volterra pairwise modeling fails to capture diverse pairwise microbial interactions. Elife. 2017;6:e25051.
Martin-Platero AM, Cleary B, Kauffman K, Preheim SP, McGillicuddy DJ, Alm EJ, et al. High resolution time series reveals cohesive but short-lived communities in coastal plankton. Nat Commun. 2018;9:266.
Lima-Mendez G, Faust K, Henry N, Decelle J, Colin S, Carcillo F, et al. Ocean plankton. Determinants of community structure in the global plankton interactome. Science. 2015;348:1262073.
Harris K, Parsons TL, Ijaz UZ, Lahti L, Holmes I, Quince C. Linking statistical and ecological theory: Hubbell’s unified neutral theory of biodiversity as a hierarchical dirichlet process. Proc IEEE. 2017;105:516–29.
Washburne AD, Burby JW, Lacker D. Novel covariance-based neutrality test of time-series data reveals asymmetries in ecological and economic systems. PLoS Comput Biol. 2016;12:e1005124.
Connor N, Barberán A, Clauset A. Using null models to infer microbial co-occurrence networks. PLoS ONE. 2017;12:e0176751.
Gibbons SM, Duvallet C, Alm EJ. Correcting for batch effects in case-control microbiome studies. PLoS Comput Biol. 2018;14:e1006102.
Weiss S, Xu ZZ, Peddada S, Amir A, Bittinger K, Gonzalez A, et al. Normalization and microbial differential abundance strategies depend upon data characteristics. Microbiome. 2017;5:27.
Silverman JD, Washburne AD, Mukherjee S, David LA. A phylogenetic transform enhances analysis of compositional microbiota data. Elife. 2017;6:e21887.
Röttjers L, Faust K. From hairballs to hypotheses—biological insights from microbial networks. FEMS Microbiol Rev. 2018;42:761–80.
Tackmann J, Rodrigues JFM, von Mering C. Rapid inference of direct interactions in large-scale ecological networks from heterogeneous microbial sequencing data. https://doi.org/10.1101/390195.
Leek JT, Scharpf RB, Bravo HC, Simcha D, Langmead B, Johnson WE, et al. Tackling the widespread and critical impact of batch effects in high-throughput data. Nat Rev Genet. 2010;11:733–9.
McLaren MR, Willis AD, Callahan BJ. Consistent and correctable bias in metagenomic sequencing measurements. bioRxiv. https://doi.org/10.1101/559831.
Tsilimigras MCB, Fodor AA. Compositional data analysis of the microbiome: fundamentals, tools, and challenges. Ann Epidemiol. 2016;26:330–5.
Tourlousse DM, Yoshiike S, Ohashi A, Matsukura S, Noda N, Sekiguchi Y. Synthetic spike-in standards for high-throughput 16S rRNA gene amplicon sequencing. Nucleic Acids Res. 2017;45:e23.
Gao Y, Li H. Quantifying and comparing bacterial growth dynamics in multiple metagenomic samples. Nat Methods. 2018;15:1041–4.
Korem T, Zeevi D, Suez J, Weinberger A, Avnit-Sagi T, Pompan-Lotan M, et al. Growth dynamics of gut microbiota in health and disease inferred from single metagenomic samples. Science. 2015;349:1101–6.
Berry D, Widder S. Deciphering microbial interactions and detecting keystone species with co-occurrence networks. Front Microbiol. 2014;5:219.
Danczak RE, Johnston MD, Kenah C, Slattery M, Wilkins MJ. Microbial community cohesion mediates community turnover in unperturbed aquifers. mSystems 2018;3:e00066–18.
Carmody RN, Gerber GK, Luevano JM Jr, Gatti DM, Somes L, Svenson KL, et al. Diet dominates host genotype in shaping the murine gut microbiota. Cell Host Microbe. 2015;17:72–84.
Bender EA, Case TJ, Gilpin ME. Perturbation experiments in community ecology: theory and practice. Ecology. 1984;65:1–13.
Acknowledgements
AC, CD, and SMG were supported by a Washington Research Foundation Distinguished Investigator Award and by startup funds from the Institute for Systems Biology. AC and NSB were supported by Ecosystems and Networks Integrated with Genes and Molecular Assemblies (http://enigma.lbl.gov), a Scientific Focus Area Program at Lawrence Berkeley National Laboratory, is based upon work supported by the U.S. Department of Energy, Office of Science, Office of Biological & Environmental Research under Contract number DE-AC02-05CH11231. NSB was supported by the National Science Foundation under Grant Nos. OCE—1558924, CBET—1606206, MCB—1518261, DBI—1565166, and MCB—1616955 to NSB. The authors thank the editor and two anonymous peer reviewers for helping us to improve the quality of this Perspective.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Carr, A., Diener, C., Baliga, N.S. et al. Use and abuse of correlation analyses in microbial ecology. ISME J 13, 2647–2655 (2019). https://doi.org/10.1038/s41396-019-0459-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41396-019-0459-z
This article is cited by
-
The Co-occurrence Matrix and the Correlation Network of Phytophagous Insects Are Driven by Abiotic and Biotic Variables: the Case of Canola
Neotropical Entomology (2024)
-
Modeling Microbial Community Networks: Methods and Tools for Studying Microbial Interactions
Microbial Ecology (2024)
-
Rhizosphere phage communities drive soil suppressiveness to bacterial wilt disease
Microbiome (2023)
-
Relative abundance data can misrepresent heritability of the microbiome
Microbiome (2023)
-
From guest to host: parasite Cistanche deserticola shapes and dominates bacterial and fungal community structure and network complexity
Environmental Microbiome (2023)